Abstract
While persistence of fear memories is essential for survival, a failure to inhibit fear in response to harmless stimuli is a feature of anxiety disorders. Extinction training only temporarily suppresses fear memory recovery in adults, but it is highly effective in juvenile rodents. Maturation of GABAergic circuits, in particular of parvalbumin-positive (PV+) cells, restricts plasticity in the adult brain, thus reducing PV+ cell maturation could promote the suppression of fear memories following extinction training in adults. Epigenetic modifications such as histone acetylation control gene accessibility for transcription and help couple synaptic activity to changes in gene expression. Histone deacetylase 2 (Hdac2), in particular, restrains both structural and functional synaptic plasticity. However, whether and how Hdac2 controls the maturation of postnatal PV+ cells is not well understood. Here, we show that PV+- cell specific Hdac2 deletion limits spontaneous fear memory recovery in adult mice, while enhancing PV+ cell bouton remodeling and reducing perineuronal net aggregation around PV+ cells in prefrontal cortex and basolateral amygdala. Prefrontal cortex PV+ cells lacking Hdac2, show reduced expression of Acan, a critical perineuronal net component, which is rescued by Hdac2 re-expression. Pharmacological inhibition of Hdac2 before extinction training is sufficient to reduce both spontaneous fear memory recovery and Acan expression in wild-type adult mice, while these effects are occluded in PV+-cell specific Hdac2 conditional knockout mice. Finally, a brief knock-down of Acan expression mediated by intravenous siRNA delivery before extinction training but after fear memory acquisition is sufficient to reduce spontaneous fear recovery in wild-type mice. Altogether, these data suggest that controlled manipulation of PV+ cells by targeting Hdac2 activity, or the expression of its downstream effector Acan, promotes the long-term efficacy of extinction training in adults.
Similar content being viewed by others
Introduction
Anxiety and trauma-related disorders are associated with a huge socioeconomic burden, given the limited treatment options available [1]. Certain anxiety disorders are characterized by abnormally persistent emotional memories of fear-related stimuli, and impaired inhibition of learned fear is a feature in post-traumatic stress disorder (PTSD), anxiety disorders and phobias [2]. In rodents, pairing a neutral tone (conditioned stimulus, CS) with an aversive foot shock (unconditioned stimulus, US) leads to the formation of a robust fear memory that can last an entire lifetime [3]. The reduction of fear responses through fear extinction learning is at the heart of clinical exposure psychotherapies applied in the case of PTSD [4, 5]. In adults, extinction learning, or the gradual decrease of behavioral response to a CS that occurs when the stimulus is presented without reinforcement (e.g., without the aversive foot shock), is neither robust nor permanent since conditioned fear responses can recover spontaneously with the passage of time; in other words, extinction learning is unstable in adults [3, 6, 7]. Strategies aimed at enhancing the ability to inhibit responses to associations that are no longer relevant may have strong therapeutic value. Several studies suggested that the maturation of GABAergic neurotransmission contributes to the increased spontaneous recovery of fear memory following extinction learning in adults [8, 9]. Therefore, modulating GABAergic function is an attractive target to reduce spontaneous recovery of fear memories with time after extinction training in adults. However, global or prolonged reduction of GABAergic drive can generate undesirable effects such as epileptic activity. Therefore, ideal approaches targeting GABAergic transmission to foster adult brain plasticity need to be both cell type-specific and temporally controlled.
Amongst the different GABAergic interneuron types, reducing parvalbumin-expressing (PV+) interneuron activity or connectivity is sufficient to reinstate heightened plasticity in adult sensory cortices [10,11,12,13]. Recent studies showed that PV+ cells also play a role in fear memory extinction. First, fear responses during extinction learning are increased by chemogenetic activation of PV+ cells in the medial prefrontal cortex [14]. Second, extinction training induces remodeling of perisomatic PV+ synapses around excitatory neurons that were previously activated during fear conditioning in the basolateral amygdala (BLA) [15]. Third, PV+ cell-specific deletion of Nogo Receptor 1, a neuronal receptor for myelin-associated growth inhibitors [16], enhances BLA PV+ synapse remodeling and reduces the spontaneous recovery of fear memory after extinction training [17]. Fourth, fear extinction weakens the functional outputs of PV+ cells to pyramidal neurons in medial prefrontal cortex [14]. All together, these data suggest that GABAergic PV+ synapse remodeling, an activity-regulated process [18,19,20], may be implicated in extinction learning and long-term spontaneous recovery of fear memories.
Transcriptional mechanisms coupling synaptic activity to changes in gene expression drive cellular processes that mediate behavioral adaptations [21]. Accessibility of the chromatin to activate these transcriptional programs is modulated by changes in the state of histone acetylation of chromatin. In particular, blocking endogenous histone deacetylases (Hdacs) promotes both structural and functional synaptic plasticity processes [22, 23] that are required to modify behavior and improve performance during learning. Accordingly, administration of pan-Hdac inhibitors in adult mice reactivates visual cortical plasticity [24] and enhances extinction learning [25]. Several studies have shown that, of the many Hdacs expressed in the mammalian brain, Hdac2 plays a critical role in fear learning and memory [22, 23, 26]. A recent study showed that PV+ cell-specific deletion of Hdac2 was sufficient to decrease evoked IPSC and increase long-term depression in layer 2-3 of adult visual cortex [27], suggesting that deletion of Hdac2 in PV+ cells induced juvenile-like phenotypes both in inhibition and cortical plasticity. Therefore, manipulating Hdac2 expression in PV+ cells is an effective strategy to reduce their inhibitory output. Whether this would be sufficient to attenuate the recovery of fear memories after extinction training in adults is unknown.
PV+ interneurons are the only cortical cell type surrounded by perineuronal nets (PNNs), which are lattice-like aggregates of chondroitin sulfate proteoglycans-containing extracellular matrix [20]. The percentage of PV+ interneurons enwrapped by PNN increases during development in different brain regions, including those involved in fear memory formation and extinction [28]. Recent evidences show that PNNs stabilize afferent synaptic contacts onto mature cortical PV+ cells, thus limiting their plasticity [20, 29, 30]. Indeed, degradation of PNNs in adult BLA promotes fear memory erasure [28], a phenomena observed only in younger animals [8, 28, 31]. PNNs are dynamic structures, comprising of numerous structural components and remodeled by metalloproteases [32]. Whether the expression of distinct PNN components is controlled by epigenetic regulation specifically in PV+ cells, in turn modulating their role in cortical plasticity, is unknown.
Here, we report that Hdac2 deletion or pharmacological inhibition in PV+ cells attenuate spontaneous recovery of fear memory after fear extinction learning in adults. In particular, these manipulations promote a temporally downregulation of Acan, coding for aggrecan, a critical perineuronal net component, which we show to be expressed almost exclusively by PV+ cells in medial prefrontal cortex. Finally, we show that intravenous delivery of siRNA against Acan right before extinction training induces knock-down of Acan expression in prefrontal cortical PV+ cells and reduces spontaneous fear recovery with time in adult wild-type mice. Therefore, transiently changing the expression of a critical PNN component specifically in PV+ cells before extinction training but after fear memory acquisition is sufficient to reduce spontaneous fear memory recovery in adult mice.
Results
Hdac2 inhibition, in PV+ cells, leads to reduced spontaneous recovery of fear memory with time following extinction training in adult mice
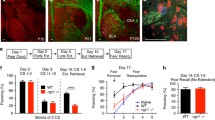
To evaluate whether and how Hdac2 deletion specifically in PV+ cells affects fear memory formation, extinction and spontaneous recovery, we crossed mice carrying a conditional allele of Hdac2 (Hdac2lox/lox, [22]), which allows cell-specific developmental stage-restricted manipulation of Hdac2, with mice expressing Cre recombinase under the control of the PV promoter (PV-Cre, [33]). In cortical GABAergic cells, PV starts to be expressed after the first postnatal week and peaks after the third week. This breeding scheme generated PV+-cell restricted homozygous (PV-Cre+/−;Hdac2lox/lox) mice and control PV-Cre−/− littermates (PV-Cre−/−;Hdac2lox/lox and PV-Cre−/−;Hdac2lox/+ mice, referred to hereafter as Hdac2lox). To confirm the specificity of Cre expression in PV-Cre+/−;Hdac2lox/lox mice, we bred them with RCEGFP reporter mice [34] and quantified GFP+/PV+ colocalized cells in somatosensory cortex (SSCX), medial prefrontal cortex (PFC) and basolateral amygdala (BLA). At P60, we observed that all GFP+ cells in all brain regions studied expressed PV, whereas the recombination rate was region specific (GFP+PV+/PV+ cells: SSCX: 82 ± 5%, n = 3 mice; PFC: 58 ± 8%, n = 5 mice; BLA: 38 ± 3%, n = 3 mice). Co-immunolabelling of PV and Hdac2 in coronal brain sections confirmed that Hdac2 expression levels were significantly reduced in PV+ cells in the conditional KO mice compared to their control littermates (Supplementary Fig. 1). We then assessed fear memory formation, extinction rate, spontaneous recovery and fear renewal in PV-Cre; Hdac2lox/lox mice (Fig. 1a). After fear conditioning, both PV-Cre; Hdac2lox/lox and Hdac2lox mice presented strong freezing behavior, suggesting that fear learning was not affected by Hdac2 deletion in PV+ cells (freezing time at early extinction (CS1-2), Mann–Whitney test, P = 0.1878, Fig. 1b). Compared to wild-type littermates, conditional KO mice showed similar extinction rate during training (Fig.1b). One week after extinction training, mice were re-exposed to the conditional stimulus (CS) either in the extinction context (retrieval test, to evaluate spontaneous fear memory recovery) or in the fear acquisition context (renewal test, to evaluate for generalization of extinction learning). Compared to wild-type littermates, conditional KO mice showed significantly less freezing both during the retrieval and renewal tests 7 days later (Fig. 1b), suggesting reduced spontaneous recovery of fear memory. The reduced fear expression was not due to loss of memory, since in absence of extinction training, freezing levels were undistinguishable between conditional KO and control mice 10 days after fear conditioning (Fig. 1c). Further, this was not due to altered motor or anxiety behavior, since we found no differences between the genotypes in the open field and elevated plus maze assays (Fig. 1d, e).
a Schematic representation of the experimental protocol. Ext. extinction. b Wild-type and PV-Cre; Hdac2lox/lox littermates show efficient fear extinction. Repeated two-way ANOVA; Fgenotype (1,22) = 4.430, P = 0.0470, Fextinction(11,242) = 8.864, P < 0.0001, Fgenotype*extinction (11,242) = 0.6427, P = 0.7911. Wild-type, but not PV-Cre; Hdac2lox/lox littermates, show spontaneous recovery and context-dependent renewal of fear memory. Unpaired two-tailed t-tests, P = 0.0205 for fear retrieval; P = 0.0008 for fear renewal. Number of mice: Hdac2lox/+ or Hdac2lox/lox n = 14; PV-Cre; Hdac2lox/lox n = 10. c In the absence of extinction training, PV-Cre; Hdac2lox/lox mice show stable fear memory 10 days after conditioning, as control littermates (Mann–Whitney test, P = 0.2509). Number of mice: Hdac2lox/+ or Hdac2lox/lox n = 9; PV-Cre; Hdac2lox/lox n = 7. d In the open field test, there is no significant difference in time spent in the center of the open field (unpaired two-tailed t-test, P = 0.4635) nor the total distance traveled (unpaired two-tailed t-test, P = 0.6083) between the two genotypes. Number of mice: Hdac2lox/+ or Hdac2lox/lox n = 12; PV-Cre; Hdac2lox/lox n = 10. e In the elevated plus maze test, there is no genotype-dependent difference in time spent (unpaired two-tailed t-test, P = 0.3279) nor the number of entries in the open arms (unpaired two-tailed t-test, P = 0.1449). Number of mice: Hdac2lox/+ or Hdac2lox/lox n = 14; PV-Cre; Hdac2lox/lox n = 12. f Schematic representation of the experimental protocol. Hdac2 inhibitor BRD6688 or the vehicle was injected intraperitoneally (i.p.) 6 h before the early extinction procedure. g One week after extinction training, freezing levels are significantly different in the spontaneous recovery test between vehicle- and BRD6688-injected Hdac2lox/lox (h) but not between vehicle- and BRD6688-injected PV-Cre; Hdac2lox/lox mice. g Extinction retrieval unpaired two-tailed t-test, P = 0.0030; fear renewal, unpaired two-tailed t-test, P = 0.1246. Number of Hdac2lox/lox mice injected with vehicle, n = 10 or BRD6688, n = 9. h Extinction retrieval unpaired two-tailed t-test P = 0.0850, fear renewal unpaired two-tailed t-test P = 0.6733. Number of PV-Cre; Hdac2lox/lox mice injected with vehicle, n = 10 or BRD6688, n = 8. Graph bars represent mean ± s.e.m. Circles represent individual mouse values. CS Conditioned stimulus, US Unconditioned stimulus. *P < 0.05, ***P < 0.001.
To ensure that the observed phenotype was caused by the genetic deletion of Hdac2 rather than any side-effect of the Cre insertion in the PV locus [33], we generated a second mouse line in which control littermates also carried the PV-Cre allele, while the Hdac2 locus remained wild-type (PV-Cre+/−;Hdac2+/+ mice). Consistent with our previous observations, these conditional KO mice showed significantly reduced freezing during the retrieval, but not the renewal, test compared to their control littermates (Supplementary Fig. 2a). In absence of extinction training, both genotypes showed strong fear memory 10 days after fear learning (Supplementary Fig. 2b). Overall, these data suggest that the deletion of Hdac2 specifically in postnatal PV+ cells is sufficient to reduce spontaneous recovery of fear memories over time in adults.
Reduced spontaneous recovery of fear memories after extinction training in adult PV-Cre; Hdac2lox/lox mice could rely on different mechanisms recruited during fear memory acquisition in the conditional KO mice compared to control littermates. For example, Gogolla et al. [28] showed that enzymatic degradation of PNN in the BLA of adult mice before but not after fear memory acquisition increased fear memory erasure susceptibility. Alternatively, Hdac2 deletion in PV+ cells may specifically affect the long-term retention of extinction memories. To distinguish between these two hypotheses, we used BRD6688, a small molecule with kinetic selectivity for Hdac2 as compared to Hdac1, a close family member showing 92% homology within the catalytic domain with Hdac2 [35, 36]. We first asked whether pharmacological inhibition of Hdac2 after fear learning but before the onset of extinction training was sufficient to reduce spontaneous fear memory recovery in wild-type mice (Fig. 1f). Compared to vehicle-injected mice, BRD6688-injected mice presented less freezing behavior at the retrieval, but not at the renewal, test (Fig. 1g). Therefore, combining a brief Hdac2 inhibition with extinction training rendered the extinction memory stronger, increasing its persistence over time. Circuits other than PV+ cells may contribute to the promoting effect of Hdac2 inhibition on the reduced spontaneous recovery of fear memory after extinction training. If the observed effect was mostly mediated by Hdac2 expression in PV+ cells, we would expect that spontaneous recovery of fear memory in wild-type mice would be reduced by Hdac2 inhibition, whereas the same treatment should have no effects in conditional KO mice. We found this to be the case; indeed, PV-Cre; Hdac2lox/lox mice injected with either vehicle or BRD6688 did not show any significant difference in the freezing behavior during the retrieval test (Fig. 1h), thus supporting the hypothesis that the state of chromatin acetylation regulated by Hdac2 in PV+ cells plays a critical role in limiting the spontaneous recovery of fear memory with time after extinction training.
Postnatal deletion of Hdac2 in PV+ cells enhances the remodeling of their synaptic terminals
During their maturation process, PV+ cells form numerous perisomatic synapses on pyramidal cell somata over the first 5 postnatal weeks in rodent neocortex [37]. To investigate whether Hdac2 conditional deletion in PV+ cells affected their efferent connectivity, we quantified putative perisomatic synapses formed by PV+ cells onto pyramidal cells by co-immunolabeling with gephyrin (scaffolding protein associated with GABA-A receptor, postsynaptic) and PV, in PFC and BLA. We found that the density of PV+ boutons colocalizing with gephyrin was significantly reduced in both PFC and BLA of naive conditional KO mice compared to their control littermates (Fig. 2a, b). This difference was not due to decreased PV expression, since the density of perisomatic PV+ boutons was not significantly different between the two genotypes (Fig. 2b).
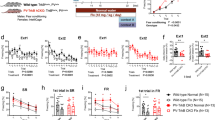
a PFC coronal sections of PV-Cre; Hdac2lox/lox and control littermates immunolabeled for PV (cyan) and gephyrin (magenta). White arrows indicate perisomatic PV-positive boutons colocalizing with Gephyrin (PV+Gephyrin+ boutons), while yellow arrows indicate perisomatic PV-positive boutons that do not colocalise with Gephyrin (PV+Gephyrin- boutons). Scale bar, 5 µm. b The density of perisomatic PV+ boutons is not significantly different neither in PFC nor in BLA between the two genotypes. PFC: Mann–Whitney test, P = 0.6825, Hdac2lox/lox n = 4; PV-Cre; Hdac2lox/lox n = 5;. BLA: Mann–Whitney test, P = 0.1014, Hdac2lox/lox n = 7; PV-Cre; Hdac2lox/lox n = 6. c The density of perisomatic PV+ boutons co-localizing with gephyrin puncta is significantly reduced in both PFC and BLA of conditional KO mice compared to their littermates. PFC: Mann–Whitney test, P = 0.0159, Hdac2lox/lox n = 4; PV-Cre; Hdac2lox/lox n = 5. BLA: Mann–Whitney test, P = 0.0140, Hdac2lox/lox n = 7; PV-Cre; Hdac2lox/lox n = 6). d Coronal sections of PFC of PV-Cre; Hdac2lox/lox and control littermates labeled with WFA (PNN, cyan) and PV (magenta). White arrows indicate PV-positive cells surrounded by PNN (PV+PNN+ cell bodies), while yellow arrows point to PV-positives cell that are not surrounded by PNN (PV+PNN- cell bodies). Scale bar, 40 µm. e The percentage of PV+ cell bodies surrounded by WFA-stained PNN is significantly reduced in the PFC and BLA, but not in the SSCX, of conditional KO mice compared to their littermates. PFC: Mann–Whitney test, P = 0.0173, Hdac2lox/lox, n = 6; PV-Cre; Hdac2lox/lox, n = 5. BLA: Mann–Whitney test, P = 0.0087, Hdac2lox/lox, n = 6; PV-Cre; Hdac2lox/lox, n = 6. SSCX: Mann–Whitney test, P = 0.6753, Hdac2lox/lox, n = 6; PV-Cre; Hdac2lox/lox n = 6. Data represent mean ± s.e.m. Black circle represent individual data points. *P < 0.05, **P < 0.01.
Previous reports showed that remodeling of perisomatic inhibitory synapses was critical for efficient extinction of fear memories [15, 17]. To investigate whether PV+ cell synapse remodeling was affected by Hdac2 deletion, we analyzed PV+ cell perisomatic synapses in both PFC and BLA 24 h after the end of fear extinction training in PV-Cre; Hdac2lox/lox mice and their control littermates. While naive conditional mutant mice showed decreased percentage of perisomatic PV+ boutons colocalizing with gephyrin puncta compared to control mice in both PFC and BLA (Fig. 2c, Supplementary Fig. 3a, b, e), no significant difference could be detected between the two genotypes following extinction training (Supplementary Fig. 3c, d, e). These results suggest that fear extinction training promoted more extensive PV+ cell synapse remodeling in PV-Cre; Hdac2lox/lox mice than in control littermates, reinforcing the hypothesis that changes in PV+ cell-mediated inhibition is critical for reducing the spontaneous recovery of fear memory over time [15].
Postnatal deletion of Hdac2 in PV+ cells reduces PNN aggregation around their somata
PNN appearance around PV+ cell somata and dendrites is developmentally regulated, and correlates with increased inhibition and critical period closure in sensory cortices [38]. PV+ cell enwrapping by PNN has been shown to be a limiting factor for adult brain plasticity [20, 29, 30]. In particular, chondroitin sulfate proteoglycans assembly into PNNs promotes the formation of extinction-resistant memories [28]. Since we observed reduced spontaneous recovery of fear memory over time following extinction training in adult PV-Cre; Hdac2lox/lox mice, we reasoned that this process may be accompanied by PNN reduction in conditional KO mice. Using WFA staining to label PNNs, we indeed found a significant decrease in the percentage of PV+ cells surrounded by PNNs in PFC and BLA, two regions directly implicated in fear behavior regulation, in naive mutant mice compared to control littermates (Fig. 2c, d). Of note, we observed that PNNs were not localized exclusively around PV+ cells in the BLA (data not shown), consistent with previous reports [39, 40]. Unexpectedly, we did not observe a significant difference in the percentage of PNN+PV+ cells in the SSCX of PV-Cre; Hdac2lox/lox compared to Hdac2lox mice (Fig. 2d), suggesting that epigenetic regulation of PNN organization is region-specific in the adult mouse cortex.
Hdac2 regulates Aggrecan expression in PV+ cells
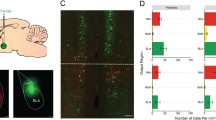
WFA specifically binds to CAG-chains [41] in all lecticans. Since we observed decreased WFA labeling surrounding PV+ cell somata in brain regions involved in fear regulation in PV-Cre; Hdac2lox/lox mice (Fig. 2c, d), we next sought to identify which lecticans are cell-autonomously expressed by PV+ cells, and could thus be directly affected by PV+ cell-specific Hdac2 deletion. Using Drop-Seq, we transcriptionally profiled PFC cells from 6 weeks-old wild-type mice. Samples from 5 biological replicates yielded 19393 cells that were sequenced to a median depth of 15,555 aligned reads/cell (representing a median of 1427 genes/cell) and controlled for quality (Supplementary Fig. 4). Cells were then clustered and represented in t-Distributed Stochastic Neighbor Embedding (t-SNE) plots (Fig. 3a). Cell clusters were defined as glutamatergic neurons, GABAergic neurons or non-neuronal cells based on Slc17a7, Gad1 and Gad2 detection (Supplementary Fig. 5). Using the Dropviz database as a template for murine cortical transcriptomes, clusters were then associated with well-known neuronal and non-neuronal cell groups. Among the latter, we distinguished astrocytes, oligodendrocytes/polydendrocytes, microglial cells/macrophages as well as endothelial cells. Within neuronal cell types, VGLUT+ glutamatergic neurons and Gad1+/Gad2+ interneurons were identified, including PV-expressing interneurons (cluster 9, Supplementary Fig. 6).
a Single cell RNA-sequencing (DropSeq) of PFC cells of P40-43 wild-type mice (n = 8 mice). t-SNE visualization of identified cell types; cluster 9 refers to PV+ cells. b–d Expression localization of critical components of PNN structure and remodeling, including lecticans (b Acan, Bcan, Ncan, and Vcan), metalloproteases known to be expressed by PV+ cells (c Adamts8, Adamts15, and Mme) and other critical PNN components (d Halpn1, Halpn4, and Tnr).
Next, we analyzed which cell types express mRNAs coding for known PNN structural components and remodeling enzymes (metalloproteases). We confirmed that the metalloproteases Adamts8, Adamts15, and Mme were highly enriched in PV+ cells, (Fig. 3c and Supplementary Fig. 6), as previously reported [42, 43]. Of the 4 analyzed lecticans, Neurocan was the most abundant and was expressed by different neuronal cell types and astrocytes. Brevican was also expressed by different cell types, including PV+ cells, but it was highly enriched in astrocytes and in CPSG4-expressing NG2+ polydendrocytes. The latter were also the major source of Versican (Fig. 3b and Supplementary Fig. 6). Acan, on the other hand, was mostly expressed by PV+ cells, and to a much lesser extent, by polydendrocytes (Fig. 3b and Supplementary Fig. 6). None of the other PNN components analyzed (Hapln1, Hapln4, Tnr) showed cell type-specific expression (Fig. 3d and Supplementary Fig. 6).
In particular, Aggrecan, coded by Acan, plays an obligatory role in PNN formation. Indeed, Acan downregulation leads to the disappearance of PNN aggregation around PV+ cells, both in brain-wide homozygous Acan mutant mice and following virally-induced Acan deletion [27]. To confirm the Drop-Seq finding and further investigate whether Acan expression is regulated by Hdac2 in PV+ cells, we analyzed Acan transcripts by RNAscope hybridization in situ in PFC, BLA and SSCX of adult PV-Cre; Hdac2lox/lox mice and control littermates (Fig. 4). We confirmed that a majority of PFC PV+ cells expressed Acan in wildtype mice (percentage of PV+Acan+/PV+ cells = 71.4 ± 3.6%, n = 7 mice). We further found that PFC PV+ cells expressing detectable level of Acan mRNA expressed also higher levels of Pvalb mRNA compared to PV+ cells negative for Acan (Supplementary Fig. 7).
a Representative images of fluorescent RNAscope in situ hybridization against Acan (Aggrecan, magenta) and Pvalb (PV, cyan) in prefrontal cortex of PV-Cre; Hdac2lox/lox and control littermates (DAPI labeling is in gray). Somata expressing Pvalb mRNA are outlined. Scale bar, 5 µm. b The mean number of Acan dots detected in Pvalb+ cells is significantly lower in the PFC, but not in BLA or SSCX, of conditional knockout mice compared to their control littermates. PFC: Mann–Whitney test, P = 0.0173, PV-Cre; Hdac2+/+ n = 5, PV-Cre;Hdac2lox/lox n = 6. BLA: Mann–Whitney test, P = 0.3874, PV-Cre; Hdac2+/+ n = 6, PV-Cre;Hdac2lox/lox n = 6. SSCX: Mann–Whitney test, P = 0.1797, PV-Cre; Hdac2+/+ n = 6, PV-Cre; Hdac2lox/lox n = 6. c Prefrontal cortex coronal sections labeled for Aggrecan (cyan) and PV (magenta) in PV-Cre; Hdac2lox/lox and control littermates. White arrows indicate PV-positive cells surrounded by Aggrecan (PV+Aggrecan+ cell bodies), while yellow arrows indicate PV-positive cells that are not surrounded by Aggrecan (PV+Aggrecan− cell bodies). Scale bar, 10 µm. d The percentage of PV+ cell bodies surrounded by aggrecan is significantly reduced in the PFC, but not BLA or SSCX, of conditional knockout mice compared to their control littermates. PFC: Mann–Whitney test, P = 0.0012, BLA: Mann–Whitney test P = 0.0513, SSCX: Mann–Whitney test P = 0.6037. For all three brain regions, Hdac2lox/lox or PV-Cre; Hdac2+/+ n = 7; PV-Cre; Hdac2lox/lox n = 6. Data represent mean ± s.e.m. Black circle represent individual data points. *P < 0.05, **P < 0.01.
Interestingly, in PV-Cre; Hdac2lox/lox mice, Acan expression was significantly reduced specifically in the PFC, but not in the SSCX or the BLA, compared to control littermates (Fig. 4a, b). This decrease in Acan expression was accompanied by a significant decrease in the percentage of PV+ cells surrounded by Aggrecan immunostaining specifically in the PFC (Fig. 4c, d). Unaltered Aggrecan expression levels in SSCX of mutant mice is consistent with the observation that the percentage of PNN+PV+ cells in the SSCX was not affected (Fig. 2c, d). Conversely, the percentage of PNN+PV+ cells was reduced in the BLA in absence of parallel changes in the percentage of Aggrecan+PV+ cells (Fig. 2c, d; Fig. 4). Taken together, these data strongly point towards a strict requirement for Aggrecan in PNN formation specifically around prefrontal PV+ cells.
The effects of PV+ cell-specific Hdac2 deletion on PNN aggregation can be due to a direct regulation of the expression of critical PNN components by Hdac2, or due to secondary homeostatic effects caused by long-term Hdac2 deletion. To test whether Hdac2 regulates Aggrecan expression in PV+ cells, we virally re-expressed Hdac2 with EGFP or EGFP alone specifically in PFC PV+ cells of adult PV-Cre; Hdac2lox/lox and PV_Cre; Hdac2+/+ mice for three weeks using an inverted double-floxed Hdac2 coding region driven by a ubiquitous EF-1α promoter (Fig. 5a). Consistent with our previous data (Fig. 4d), we observed a significant reduction in the percentage of PV+ cells enwrapped by Aggrecan in conditional KO compared to wildtype littermates when both were injected with the control virus (AAV_DIO_EGFP; Fig. 5b, c). On the other hand, injection of AAV_DIO_Hdac2-T2A-EGFP was sufficient to rescue the percentage of Aggrecan+ PV+ cells (Fig. 5c) and to increase (albeit not significantly) the number of PV+ cells surrounded by WFA-stained PNN in PV-Cre; Hdac2lox/lox mice (Fig. 5d, e).
a Schematic representation of Cre-dependent viral vectors and experimental approach. b Immunolabelling of Aggrecan (magenta), GFP (gray) and PV (cyan) in PFC coronal sections from PV-Cre; Hdac2+/+ and PV-Cre; Hdac2lox/lox mice injected with the control AAV (AAV_DIO_EGFP) and from PV-Cre; Hdac2lox/lox mice injected with the Cre-dependent Hdac2 expressing virus (AAV_DIO_Hdac2_T2A_EGFP). White arrows point to the somata of AAV-transfected (GFP-positive) PV cells surrounded by Aggrecan (PV+GFP+Aggrecan+ cell bodies), while yellow arrows indicate AAV-transfected (GFP-positive) PV cells that are not surrounded by Aggrecan (PV+GFP+Aggrecan− cell bodies). Scale bar, 10 µm. c Percentage of PV+GFP+ cell bodies surrounded by Aggrecan. Kruskall–Wallis with Dunn’s posthoc test, P = 0.0008. d Labeling for WFA (PNN, magenta), GFP (gray) and PV (cyan) in the three experimental groups. White arrows indicate PV+GFP+ cell bodies enwrapped by PNN (PV+GFP+PNN+ somata), while yellow arrows indicate PV+GFP+ cell bodies that are not enwrapped by PNN (PV+GFP+PNN− somata). Scale bar, 10 µm. e Percentage of PV+GFP+ cell bodies surrounded by WFA-stained PNN in the prefrontal cortex. Kruskall–Wallis with Dunn’s posthoc test, P = 0.2603. Number of mice: PV-Cre; Hdac2+/+ + AAV_DIO_EGFP, n = 4, PV-Cre; Hdac2lox/lox + AAV_DIO_EGFP, n = 6, PV-Cre; Hdac2lox/lox + AAV_DIO_Hdac2_T2A_EGFP, n = 5. Data represent mean ± s.e.m. Black circle represent individual data points. *P < 0.05.
To further support the hypothesis that Hdac2 activity in PV+ cells regulates Aggrecan levels, we explored the effects of acute Hdac2 pharmacological inhibition. We found a significant decrease in the number of Acan mRNA molecules in PFC PV+ cell somata (Fig. 6a, b) and in the proportion of PV+ cells enwrapped by Aggrecan+ nets (Fig. 6d, e) 17 h after BRD6688 i.p. injection in wild-type mice. Conversely, the same treatment was unable to further reduce Acan/Aggrecan expression in PV-Cre; Hdac2lox/lox mice (Fig. 6c, f), suggesting that the effect of BRD6688 treatment was mostly mediated by Hdac2 expression in PV+ cells,
a Representative images of fluorescent RNAscope in situ against Acan (Aggrecan, magenta) and Pvalb (PV, cyan) around a DAPI (gray) nucleus in PFC of Hdac2lox/lox mice 17 h after i.p. injection of either the Hdac2 inhibitor BRD6688 or the vehicle solution. Somata expressing Pvalb mRNA are outlined. Scale bar, 5 µm. b The mean number of Acan dots present around a Pvalb+ cell is reduced in BRD6688-treated wild-type (Mann–Whitney test, P = 0.0303) (c) but not conditional knockout mice (Mann–Whitney test, P = 0.6786). Number of Hdac2lox/lox injected with vehicle, n = 5 or BRD6688, n = 6. Number of PV-Cre; Hdac2lox/lox mice injected with vehicle n = 3 or BRD6688, n = 5. d PFC coronal sections immunolabeled for Aggrecan (cyan) and PV (magenta) in Hdac2lox/lox mice 17 h after i.p. injection of either the Hdac2 inhibitor BRD6688 or the vehicle. White arrows indicate PV-positive cells surrounded by Aggrecan (PV+Aggrecan+ cell bodies), while yellow arrows indicate PV-positive cells that are not surrounded by Aggrecan (PV+Aggrecan− cell bodies). Scale bar, 10 µm. e The percentage of PV+ cell bodies surrounded by Aggrecan is reduced by BDR6688 injection in control (Mann–Whitney test, P = 0.0317) (f) but not conditional knockout mice (Mann–Whitney test, P = 0.8286). Number of Hdac2lox/lox injected with vehicle, n = 5 or BRD6688, n = 4. Number of PV-Cre;Hdac2lox/lox mice injected with vehicle, n = 4 or BRD6688, n = 4. Graph bars represent mean ± s.e.m. Circles represent individual mouse values. *P < 0.05.
Overall, these results show that PV+ cells are the major source of Aggrecan in the PFC and that Hdac2 modulates PNN aggregation by regulating Acan transcription in PV+ cells.
Reduced Hdac2 activity in PFC PV+ cells promotes dynamic changes in Acan expression following extinction training
Class 1 Hdac (including Hdac2) inhibition has been shown to prime the expression of neuroplasticity-related genes [23]. We thus asked whether Acan expression might be more dynamic in Hdac2−/− PV+ cells following extinction training. To answer this question, we collected brains from PV-Cre; Hdac2lox/lox mice and control littermates 3 h after the end of extinction training (Fig. 7a, b) and quantified Acan mRNA expression in PV+ cells from the PFC (Fig. 7c, d). We found that, although Acan expression was reduced in PFC PV+ cells from naive mutant mice compared to wild-type littermates (Fig. 4a, b), its expression reached wild-type levels after extinction training (Fig. 7c, d). Next, we asked whether acute pharmacological Hdac2 inhibition could promote Acan dynamic changes triggered by extinction learning. While naive mice injected with BRD6688 showed reduced Acan levels (Fig. 6a, b), mice injected with BRD6688 following fear acquisition but before the onset of extinction training showed no significant difference between Acan levels compared to vehicle-treated controls 3 h after the end of extinction training (Fig. 7g, h).
a Schematic representation of the experimental protocol. Brains of PV-Cre; Hdac2lox/lox mice and their control littermates were collected 3 h after late extinction training for in situ hybridization. b Both PV-Cre; Hdac2lox/lox mice and controls show efficient fear extinction. Repeated two-way ANOVA; Fgenotype (1,12) = 7.554, P = 0.0177, Fextinction(11,132) = 9.148, P < 0.0001, Fgenotype*extinction(11,132) = 1.107, P = 0.3605. c Representative images of fluorescent RNAscope in situ against Acan (Aggrecan, magenta) and Pvalb (PV, cyan) around a DAPI (gray) nucleus in PFC of PV-Cre; Hdac2+/+ and PV-Cre; Hdac2lox/lox mice 3 h after late extinction training. Somata expressing Pvalb mRNA are outlined. Scale bar, 5 µm. d The mean number of Acan dots present around a prefrontal Pvalb+ cell 3 h after late extinction training is comparable between PV-Cre; Hdac2+/+ and PV-Cre; Hdac2lox/lox mice (Mann–Whitney test, P = 0.8112). Number of mice: PV-Cre; Hdac2+/+ n = 6; PV-Cre; Hdac2lox/lox n = 8. e Schematic representation of the experimental protocol. Brains of BRD6688-injected or vehicle-injected mice 6 h before extinction training have been collected 3 h after late extinction training for in situ hybridization. f Both BRD6688-injected and vehicle-injected controls show efficient fear extinction. Repeated two-way ANOVA; Ftreatment (1,6) = 1.093, P = 0.3362, Fextinction(1166) = 3.902, P = 0.0002, Ftreatment*extinction (1166) = 0.7446, P = 0.6922. g Representative images of fluorescent RNAscope in situ against Acan (Aggrecan, magenta) and Pvalb (PV, cyan) around a DAPI (gray) nucleus in PFC of BRD6688-injected and vehicle-injected mice 3 h after late extinction training. Somata expressing Pvalb mRNA are outlined. Scale bar, 5 µm. h The mean number of Acan dots present around a prefrontal Pvalb+ cell 3 h after late extinction training is comparable between BRD6688-injected and vehicle-injected mice (Mann–Whitney test, P = 0.2). Number of mice: vehicle-injected n = 4; BRD6688-injected n = 4.
Altogether, these results indicate that either PV+ cell-specific Hdac2 deletion or pharmacological inhibition allows for dynamic changes in Acan expression following extinction training.
Acan knock-down during extinction training is sufficient to limit the spontaneous recovery of fear memories with time
Finally, we asked whether the observed dynamic changes in Acan levels caused by Hdac2 inhibition were sufficient to attenuate the spontaneous recovery of fear memories following extinction training over time. To answer this question, we sought to transiently decrease Acan expression after fear acquisition but before extinction training. To this purpose, we used a chimeric fusion protein comprising a double-stranded RNA binding domain fused to a brain targeting peptide, namely TARBP-BTP, to deliver siRNA to the brain [44, 45]. Previous data showed that using TARBP-BTP fusion protein as a carrier facilitated siRNA delivery across the brain-blood barrier to the brain [44]. We observed a significantly reduced number of Acan mRNA molecules in PFC PV+ cells compared to the control group 48 h after TARBP-BTP/Acan-siRNA intravenous injection in wild-type adult mice (Fig. 8a–c), demonstrating that siRNAs were efficiently delivered to the brain parenchyma and could enter PV+ cells. We then injected either TARBP-BTP/Acan-siRNA or TARBP-BTP/CtrlNeg-siRNA in wild-type mice, 24 h after fear learning, and then subjected them to fear extinction training 24 h after injection (Fig. 8d). Compared to control TARBP-BTP/CtrlNeg-siRNA mice, TARBP-BTP/Acan-siRNA-injected mice presented significantly less freezing behavior during the retrieval test (Fig. 8e). Therefore, Acan knock-down in PV+ cells prior to extinction training was sufficient to render extinction memories stronger and limits the spontaneous recovery of fear memories with time.
a Schematic representation of the experimental protocol. Brains of adult Hdac2lox mice were collected 48 h after intravenous injection (i.v.) of either TARBP-BTP:siCtrlNeg or TARBP-BTP:siAcan. b Representative images of fluorescent RNAscope in situ against Acan (Aggrecan, magenta) and Pvalb (PV, cyan) around a DAPI (gray) nucleus in PFC of Hdac2lox mice 48 h after the i.v. injection. Somata expressing Pvalb mRNA are outlined. Scale bar, 5 µm. c The mean number of Acan dots in PFC Pvalb+ cells 48 h after injection of TARBP-BTP:siAcan is significantly decreased compared to those in PFC Pvalb+ cells from TARBP-BTP:siCtrlNeg injected mice (Unpaired two tailed t-test, P = 0.0139). Number of mice: Hdac2lox + TARBP-BTP:siCtrlNeg n = 3; Hdac2lox + TARBP-BTP:siAcan n = 4. d Schematic representation of the experimental protocol. Ext. extinction. e TARBP-BTP:siCtrlNeg-injected and TARBP-BTP:siAcan-injected mice show similar fear extinction rate. Repeated two-way ANOVA; Ftreatment (1,19) = 1.109, P = 0.3054, Fextinction(11,209) = 18.85, P < 0.0001, Ftreatment*extinction (11,209) = 1.052, P = 0.4021. One week after extinction training, freezing levels in the retrieval test are significantly lower in TARBP-BTP:siCtrlNeg-injected compared to TARBP-BTP:siAcan-injected mice. Extinction retrieval unpaired two-tailed t-test, P = 0.0349; fear renewal, unpaired two-tailed t-test, P = 0.5608. Number of Hdac2lox mice injected with TARBP-BTP:siCtrlNeg, n = 11 or TARBP-BTP-siAcan, n = 10. Graph bars represent mean ± s.e.m. Circles represent individual mouse values. CS conditioned stimulus. *P < 0.05.
Discussion
The maturation of GABAergic circuits, in particular of PV+ cells, is one of the factors that restricts critical period plasticity in the adult brain [11, 12, 38]. Hence, controlled manipulation of cortical PV+ cell function can reinstate heightened plasticity, creating an opportunity for therapeutic intervention [46, 47]. To this purpose, it is essential to gain a better understanding of the molecular mechanisms determining PV+ cell maturation state in the adult brain. Here, we showed that the epigenetic landscape of PV+ cells regulates their maturation state, thereby affecting adult neuroplasticity. In particular, our results demonstrate that PV+ cell-specific Hdac2 deletion reduces PNN aggregation around PV+ cell somata and PV+ cell synapse density in naive mice, while promoting PV+ cell structural synapse remodeling following extinction training. We further found that Hdac2 levels controlled the expression of the lectican Aggrecan, a critical regulator of PNN aggregation, by regulating its transcription specifically in PFC PV+ cells. These anatomical changes are accompanied by decreased spontaneous recovery of fear memories with time in mutant mice. Finally, siRNA-mediated brain-targeted Acan reduction after fear acquisition coupled with extinction training was sufficient to reduce the spontaneous recovery of fear memory with time in adult wild-type mice, indicating a potential therapeutic target for enhancing extinction-dependent plasticity.
A previous study by Gogolla and collaborators [28] showed that degradation of PNNs by chondroitinase ABC injection in adult BLA rendered subsequently acquired fear memories susceptible to erasure following extinction learning. However, the same treatment had no effect when applied after fear learning, but before extinction training. In contrast, our data showed that global Acan reduction after fear learning was sufficient to reduce the spontaneous recovery of fear memories one week after extinction training. It is possible that PNN remodeling in PFC and not, or not exclusively, in the BLA, is essential for long term retention of extinction learning. Alternatively, the dynamic regulation of PNN expression, more than its absolute levels, may play a key role in this process. Indeed, a transient decrease in Acan expression after fear learning but before extinction training attenuated the spontaneous recovery of fear memory over time, suggesting that dynamic changes of PNNs could be a critical step in this process. To test this hypothesis, we choose to deliver siRNA against Acan using a peptidic carrier that crosses the brain blood barrier Weather transient Acan changes and PNN remodeling selectively in PFC are sufficient to affect the spontaneous recovery of fear memory, or whether PNN remodeling needs to occur in the BLA too, remain to be established.
Numerous studies have addressed the role of epigenetic modifiers, through pharmacological inhibition or genetic deletion of different Hdacs, on fear memory formation and extinction [48]. In particular, an elegant study demonstrated that pharmacological inhibition of Hdac2 during contextual fear extinction prevents the spontaneous recovery of freezing behavior during the recall of remote fear memories by upregulating neuroplasticity-related gene expression [23]. However, whether Hdac2 plays a different role in different neuron types thus leading to distinct behavioral outputs, is not clear. A previous study showed that Hdac2 deletion in post-mitotic forebrain glutamatergic neurons (using CaMKII-Cre mice) led to a robust acceleration of extinction learning, which was however followed by a normal spontaneous recovery of fear memories 5 days after the end of the extinction [26]. Conversely, our data showed that Hdac2 deletion specifically in postnatal PV+ cells did not affect the rate of extinction learning but led to decreased spontaneous recovery of fear memories with time. The specific role of Hdac2 expressed by PV+ cells in allowing spontaneous fear recovery was further supported by the fact that the beneficial effect of Hdac2 inhibitor and PV+ cell-specific deletion of Hdac2 on preventing the recovery of fear memories were not additive. These different results could be explained by partly different roles played by glutamatergic and PV+ neurons in fear extinction learning and spontaneous fear memory recovery. It is also conceivable that Hdac2 regulates a different set of genes in different cell types, thus leading to different cellular responses.
The main problem with fear extinction is the reappearance, or spontaneous recovery, of the original fear after extinction training. To explain this phenomena, two opposite models of extinction have been proposed and strongly debated: one posits that the extinction training leads to the erasure of the fear memory itself [17, 28], while the second proposes that extinction creates new inhibitory learning, which controls the fear memory [49, 50]. It has been suggested that extinction learning could be seen as an equilibrium between erasure and inhibition, where any behavioral manipulation which influences fear expression would either promote a functional erasure of the original fear trace or have permissive effects on the new inhibitory memory [51]. In the field of cognitive neuro-epigenetics, it has been proposed that “there exists an equilibrium in epigenetic states that can be pushed in one direction or another” [51]. The data presented in this work support the hypothesis that increasing chromatin acetylation, likely at specific genes, enhances the retention of fear extinction memories over time. In particular, we identified PV+ cell-epigenetic state as a major determinant in the spontaneous recovery of fear memories over time. Whether Hdac2 inhibition in PV+ cells leads to stronger inhibitory memory opposing the fear memory, or to the erasure of the original fear memory is not clear. However, the lack of generalization of fear extinction memory, as observed by similar freezing times during the renewal test in PV-Cre;Hdac2lox/lox versus PV-Cre;Hdac2+/+ mice (Supplementary Fig. 2) or following Hdac2 inhibition after fear memory acquisition (Fig. 1g), argues against the erasure of the original fear memory, instead supporting the hypothesis that PV+ cell heightened plastic remodeling could contribute to forming stronger (or more stable) inhibitory memory [15].
The development of cortical PV+ cells is a prolonged process which plateaus only by the end of adolescence [37]. During the postnatal maturation phase, cortical PV+ cells form exuberant innervation fields characterized by baskets of perisomatic synapses. In addition, during their maturation phase, PV+ cell somata and dendrites are progressively enwrapped in PNNs, thought to stabilize their synaptic inputs [19, 52] and protect them from oxidative stress [53]. Our data suggest that PV+ cell-epigenetic modifications likely regulate their maturation state, since PV-Cre;Hdac2lox/lox mice showed significantly reduced PNN condensation around PV+ cells and decreased density of PV+ cell perisomatic synapses in the PFC and BLA, two brain regions implicated in fear behavior regulation. Whether these structural changes in PV+ cells synaptic density correlate with reduced PV+ cell-mediated functional inhibition in the BLA or/and PFC remains to be determined. Nevertheless, our data is consistent with the observation that PV+ cell-restricted Hdac2 deletion reduced evoked inhibitory responses and enhanced long-term depression in the visual cortex of adult mice, which are phenotypes typically associated with younger mice [27].
We further found that PFC PV+ cells expressing detectable level of Acan mRNA also expressed higher levels of Pvalb mRNA compared to PV+ cells negative for Acan (Supplementary Fig. 7). Higher number of PV+ cells expressing high levels of PV protein have been associated with reduced memory and structural synaptic plasticity in the hippocampus [18] and visual cortex [13]. This observation supports the hypothesis that reduced Acan expression in the PFC of conditional KO mice (Fig.4) or in wild-type mice injected with BRD6688 (Fig.6) may lead to a PV+ cell network configuration favoring plasticity. In accordance with this hypothesis, Chen et al. [14] showed that extinction learning increased the fraction of PV+ cells expressing low PV levels, which are thought to be more plastic [18]. They further showed that chemogenetic-mediated manipulations of PV+ cell activity affected mouse freezing during extinction learning, with more active PV+ cells leading to higher freezing. However, this study did not explore whether and how these manipulations affected long-term spontaneous recovery of fear memories.
Trouche et al. [15] showed that extinction training induced structural remodeling of perisomatic PV+ cell synapses specifically around excitatory neurons that had been previously activated during the encoding of the original fear memory (fear neurons) in the BLA. In addition, Bhagat et al. [17] reported that PV+ cell-specific deletion of Nogo Receptor 1 enhanced BLA PV+ cell synaptic structural remodeling and reduced spontaneous recovery of fear memory over time. In PV-Cre;Hdac2lox/lox mice, we observed increased remodeling of PV+ cell synapses in both BLA and PFC following extinction training. Elegant work demonstrated that PFC PV+ cell firing inversely correlates with freezing behavior and that their optogenetic activation reduces freezing behavior [54]. Altogether, this data support the hypothesis that reinforcing PFC PV+ cell-mediated inhibitory drive by the end of extinction training could contribute to the maintenance of the extinction memory, thereby leading to reduced fear response freezing at the retrieval test.
How and to what extent PV+ cells control the development, integrity and dynamic of the PNNs surrounding their somata and dendrites, in a cell-autonomous fashion, is still an open question. By using single-cell transcriptomic analysis, we found that PFC PV+ cells express a variety of PNN molecular components and metalloproteases. While most of these factors were expressed by other cell types as well, Acan transcription appeared to be mostly restricted to PV+ cells, consistent with previous studies [55, 56]. In vitro [57, 58] and in vivo [30] experiments revealed that Aggrecan, coded by Acan, plays a critical role in PNN formation. For instance, brain-wide targeting of Acan (Nes_Cre; Acanlox/lox mice) or virally-mediated local Acan deletion in adult visual cortex led to complete loss of WFA-stained PNN, while conditional deletion of one Acan allele was sufficient to reduce WFA staining intensity [30]. Therefore, the reduced Aggrecan levels we observed in PV_Cre; Hdac2lox/lox or BRD6688-injected mice likely contributes to impaired aggregation of other extracellular matrix molecules into PNNs. Our data showed that PV+ cell-specific Hdac2 deletion reduced Acan/Aggrecan expression levels in PV+ cells, while re-introduction of Hdac2 in Hdac2−/− mutant cells rescued Acan/Aggrecan expression levels, thus suggesting that Hdac2 controls Acan levels in PV+ cells. By determining Aggrecan expression levels, PV+ cells may cell-autonomously modulate PNN dynamics and integrity, thereby regulating the stability of their synaptic inputs [19, 52, 59]. Further, a recent study showed that 4 days of monocular deprivation was sufficient to produce a strong shift in ocular dominance in adult Acanlox/lox mice injected with AAV-Cre, suggesting that Acan expression in the adult brain acts as a brake on neuroplasticity [30]. Therefore, decreasing Acan levels in PV-Cre; Hdac2lox/lox or BRD6688-injected mice likely promotes neuroplasticity, which in turn contributes to stronger retention of extinction memory and reduced spontaneous recovery of fear memory over time. Consistent with this hypothesis, our data show that a brief Acan reduction coupled with extinction training was sufficient to decrease the spontaneous recovery of fear memory. Furthermore, we showed that PV+ cell-specific Hdac2 deletion reduced Acan transcription selectively in PFC, but not in somatosensory cortex, PV+ cells. This surprising regional specificity suggests that PV+ cell circuits in different adult cortical areas might differ more, at least in term of epigenetic regulation, than what was previously assumed. In particular, region-specific dynamic epigenetic regulation of PV+ cells could increase plasticity of PFC networks compared to primary sensory areas in the adult brain.
In contrast to what we observed in the PFC, the reduced numbers of PV+ cells enwrapped by PNN in the BLA of PV-Cre; Hdac2lox/lox mice was not accompanied by reduced Acan/Aggrecan expression, indicating that different molecular organizers might be at play in the two regions. In BLA, PNN enwrap not only PV+ cells, but also a population of excitatory neurons [60], therefore PNNs components produced by excitatory neurons may contribute to PNN aggregation around PV+ cells. Finally, consistent with previous findings [42], we observed that the expression of the metalloproteases Adamts8, Adamts15 and Mme was highly enriched in, but not restricted to, PV+ cells. Increased metalloproteases expression might lead to Aggrecan cleavage and PNN disassembly. Whether their expression levels are regulated, directly or indirectly, by Hdac2 remains to be explored.
Finally, we observed that while Acan gene expression in PFC PV+ cells was lower in naive PV_Cre; Hdac2lox/lox and in BRD6688-injected wild-type mice compared to their respective control groups, the difference was lost after extinction training, suggesting that Hdac2 might limit dynamic changes in Acan expression. Increased Acan expression likely leads to increased PNN aggregation and stabilization around PV+ cells [23, 45, 46]. It has been suggested that new PNN formation plays a critical role in memory consolidation, by modulating PV+ cell activity; [61] therefore, it is possible that the ability to dynamically regulate Acan, and thus PNN formation/stability, and not absolute levels of Acan, determines the degree of spontaneous fear memory recovery over time. A recent study showed that Hdac2 S-nitrosylation was upregulated in the hippocampus during the extinction of contextual fear memories. Hdac2 S-nitrosylation caused Hdac2 dislocation from the chromatin and increased chromatin acetylation, thereby priming the expression of neuroplasticity-regulated genes (“transcriptionally permissive state”) [23]. Whether the same epigenetic mechanism is at play in PFC PV+ cells during fear extinction learning and whether Aggrecan expression is downstream of such regulation, remains to be investigated. Nevertheless, our findings suggest that Hdac2 levels and/or activity regulate Acan expression specifically in PFC PV+ cells, but not in somatosensory or BLA PV+ cells, in adult mice. Therefore, any future study aimed at dissecting the molecular mechanisms linking Hdac2 and Acan expression must specifically target adult PFC PV+ cells.
Taken together, our data support a model in which the acetylation of PV+ cell chromatin primes the Acan gene in PFC to be dynamically expressed during extinction learning. Increased Aggrecan expression in turn promotes PNN condensation [23, 45, 46], and modulates PV+ cell-mediated inhibition in the PFC contributing to reduced fear expression over time, following extinction training. In addition, acetylation of PV+ cell chromatin regulates the ability of PV+ cell synapses to remodel their output following extinction training, which could promote the selective inhibition of fear memory traces. Since spontaneous recovery of fear response following extinction training remains an important challenge in exposure therapy, any manipulation that can potentiate the plasticity of PV+ cell networks, whether it is through modulating chromatin accessibility or the expression of PNN components, could in turn foster adult brain plasticity and be of future therapeutic interest.
Material and methods
Animals
Hdac2lox/lox (The Jackson laboratory (JAX), B6.Cg-Hdac2tm1.1Rdp/J, 022625) mice, in which exons 5 and 6 of Hdac2 gene, encoding the HDAC-catalytic core of the protein, are flanked by loxP sites, were crossed to the PV-Cre (JAX, B6.129P2-Pvalbtm1(cre)Arbr/J, 017320) line to generate PV-Cre+/−;Hdac2lox/lox (Hdac2 cKO) mice. These lines were maintained through crosses of PV-Cre+/−;Hdac2lox/+ females and Hdac2lox/lox males to generate PV-Cre+/−;Hdac2lox/lox and PV-Cre−/−;Hdac2lox/lox or PV-Cre−/−;Hdac2lox/+ control littermates, or PV-Cre+/+;Hdac2lox/+ females and Hdac2lox/+ males to generate PV-Cre+/−;Hdac2lox/lox and PV-Cre+/−;Hdac2+/+ control littermates, on a mixed 129sv/C57BL/6J background. Cell specificity of Cre-mediated recombination was analyzed by breeding PV-Cre with RCEEGFP mice (JAX, Gt(ROSA)26Sortm1.1(CAG-EGFP)Fsh/Mjax, 32037). All animals were maintained under a light-dark cycle (12 h light–12 h dark) in a temperature and humidity-controlled room. Food and water were available ad libitum. All procedures described here had been approved by the Comité Institutionnel de Bonnes Pratiques Animales en Recherche (CIBPAR) of the Research Center of Sainte-Justine Hospital in accordance with the principles published in the Canadian Council on Animal’s Care’s (Guide to the Care and Use of Experimental Animals).
Mice genotyping
DNA was extracted from mouse tails and genotyped to detect the presence of Cre alleles and Hdac2 conditional and wild-type alleles. Polymerase chain reaction (PCR) was performed using either a set of 2 separated primers (F1 5’-TGGTATGTGCATTTGGGAGA-3’ and R1 5’-ATTTCACAGCCCCAGCTAAGA-3’) to identify the Hdac2 floxed versus the wildtype allele or a set of 3 separate primers (5’-ATTTGGGAGAAGGCCGTTT-3’, 5’-AATTTCACAGCCCCAGCTAAG-3’ and 5′-CGAAATACCTGGGTAGATAAAGC-3′) to assure the absence of the Hdac2 null allele. Band sizes were of 720 bp for the wild-type, 560 bp for the floxed and 380 bp for the null allele. The 3 separate primers used to detect Cre in PV_Cre mice were: F1 (5’-CAGCCTCTGTTCCACATACACTCC-3’), F2 (5’- GCTCAGAGCCTCCATTCCCT-3’) and R1 (5’-TCACTCGAGAGTACCAAGCAGGCAGGA GATATC-3’) which generated 400 bp and 526 bp (mutant and wild-type) bands. To detect the presence of the RCE allele, 3 separate primers namely, RCE-Rosa1 (5’-CCCAAAGTCGCTCTGAGTTGTTATC-3’), RCE-Rosa2 (5’GAAGGAGCGGGAGAAATGGATATG-3,) and RCE-Cag3 (5’-CCAGGCGGGC CATTTACCGTAAG-3’) were used, which generated 350 bp and 550 bp bands.
Immunohistochemistry
Mice of both sexes were anesthetized, then perfused intracardially with PBS followed by 4% (w/v) paraformaldehyde (PFA) in PBS. Intact brains were extracted and post-fixed in 4% PFA/PBS overnight at 4 °C. The tissue was then cryoprotected in 30% (w/v) sucrose (Sigma) in PBS, sectioned coronally at 40 μm on a cryostat (Leica VT100) and stored as floating sections in PBS. For immunohistological analysis, brain sections were blocked in 10% normal goat serum (NGS, Invitrogen, 10000C) in PBS containing 1% (v/v) Triton X-100 for 2 h at room temperature. Primary antibodies were diluted in 5% goat serum in PBS containing 0.1% (v/v) Triton X-100 and incubated 24–48 h at 4 °C. Slices were then washed in PBS (3 ×10’), incubated in the appropriate Alexa-conjugated antibodies in 5% NGS, 0,1% Triton in PBS for 2 h at room temperature, washed again in PBS (3 ×10’), and mounted in Vectashield (Vector Lab, H-1000) before imaging.
The primary antibodies used in this study and their working concentrations are as follows: rabbit monoclonal anti-Hdac2 (1:500; Abcam, 32117, RRID:AB_732777); mouse monoclonal anti-PV (1:2000; Swant, 235, RRID:AB_10000343); rabbit polyclonal anti-PV (1:4000; Swant, PV27, RRID:AB_2631173); guinea pig polyclonal anti-PV (1:1000; Synaptic Systems, 195004, RRID:AB_2156476); mouse monoclonal anti-gephyrin (1:500; Synaptic Systems, 147021, RRID:AB_22325461); rabbit polyclonal anti-aggrecan (1:500; Millipore, AB1031, RRID:AB_90460); chicken polyclonal anti-GFP (1:1000; Abcam, 13970, RRID:AB_300798). To label PNNs, a solution of biotin-conjugated lectin Wisteria floribunda (WFA) (10 µg/ml; Sigma-Aldrich, L1516) was added in the primary antibody solution.
The secondary antibodies used in this study and their working concentrations are as follows: goat anti-chicken Alexa488 conjugated (1:1000; Abcam, ab150169), goat anti-rabbit Alexa633 conjugated (1:400; Life technologies, A21072), goat anti-mouse Alexa555 conjugated (1:1000; Cell Signaling, 4409S), goat anti-mouse Alexa647 conjugated (1:1000; Cell signaling, 4410S), goat anti-rabbit Alexa555 conjugated (1:400; Life technologies, A21430), goat anti-guinea pig Alexa647 conjugated (1:400; Life technologies, AB21450) and Alexa 568-conjugated streptavidin (1:500; Invitrogen, S-11226).
All immunohistological experiments were performed on at least three different sections per brain region per animal. No mice were excluded from the following analysis.
Confocal imaging
All images were acquired using Leica confocal microscopes (SP8 or SP8-STED). We imaged somatosensory (SSCX) and prefrontal cortex (PFC) layer 5 and basolateral amygdala (BLA) using 20X multi-immersion (NA 0.75) and 63X oil (NA1.4) objectives. We focussed on layer 5 of SSCX and PFC because PV+ cells are more abundant in this layer. The 20X objective was used to acquire images to analyze the percentage of: GFP+PV+/PV+ cells (recombination rate), GFP+PV+/GFP+ cells (Cre-recombination specificity), PNN+PV+/PV+ cells and Agg+PV+/PV+ cells. The 63X objective was used to acquire images to analyze the perisomatic innervation (number of perisomatic PV+ boutons and gephyrin+ punctas). At least three confocal images from three different brain sections were acquired per brain region with z-step size of 2 µm (20X) and 0.5 µm (63X). All the confocal parameters were maintained constant throughout the acquisition of an experiment.
Image analysis
The number of positive cells (GFP+, PV+, PNN+ and/or Agg+) were manually identified and counted using ImageJ-Fiji software. To quantify the number of PV+, gephyrin+ and PV+/gephyrin+ punctas, images were exported as TIFF files and analyzed using Neurolucida software. PV+ boutons and gephyrin+ punctas were independently identified around the perimeter of a targeted cell after selecting the focal plane with the highest soma circumference. At least 4 perisomatic innervated somata were selected in each confocal image. Investigators were blind to the genotypes or conditions during the analysis. Each experiment included both controls and mutant or treated mice.
Behavioral testing
For all experiments, a camera was mounted above the arena; images were captured and transmitted to a computer running the Smart software (Panlab, Harvard Apparatus) or FreezeFrame software IMAQ 3 (Version 3.0.1.0). The sequence of animals tested was randomized by the genotype. Care was taken to test litters containing both the genotypes specific to the breeding. Room lights were kept low during all procedures.
Open field
Each subject (P60) was gently placed at the center of the open-field arena and recorded by a video camera for 10 min. The recorded video file was analyzed with the SMART video tracking system (v3.0, Harvard Apparatus). The open field arena was cleaned with 70% ethanol and wiped with paper towels between each trial. The time spent in the center (45% of the surface) versus the periphery was calculated. Locomotor activity was indexed as the total distance traveled (m).
Elevated plus maze
The apparatus consisted of two open arms without walls across from each other and perpendicular to two closed arms with walls joining at a central platform. Each subject (P60) was placed at the junction of the two open and closed arms and allowed to explore the maze during 5 min while video recorded. The recorded video file was analyzed with the SMART video tracking system (v3.0, Harvard Apparatus) to evaluate the percentage of time spent in the open arms (open/(open + closed) × 100) and the number of entries in the open arms as a measure of anxiety-related behavior.
Fear conditioning and fear extinction
Fear conditioning and extinction were conducted in an isolated behavior room using standard operant boxes in two different contexts (contexts A and B, respectively). The context A consisted of white walls, a grid floor, and was washed with 70% ethanol before and after each session. The context B consisted of black and white stripped walls, a white plexiglass floor, and was washed with 1% acetic acid before and after each session. Two different fear conditioning protocols [28] were used in this study, as described in details below.
PV-Cre+/−; Hdac2lox/lox males and their PV-Cre−/−; Hdac2lox/lox or PV-Cre−/−; Hdac2lox/+ control male littermates, or Hdac2lox wildtype males, were conditioned using the following protocol. One day prior to fear conditioning (Day 0), mice were allowed to freely explore the context A for habituation. On day 1, mice were conditioned using 5 pairings of the conditioned stimulus (CS duration 5 s, white noise, 80 dB) with a co-terminating unconditioned stimulus (US, 1 s foot-shock, 0.6 mA, inter-trial interval: 30–60 s). On days 2 and 3, conditioned mice were submitted to extinction training in context B. During this training, 12 CS were presented at 30 s intervals on each day. Retrieval of extinction memory and context-dependent fear renewal were tested 7 days later in context B and A, respectively, using 4 presentations of the CS. The retrieval test allowed to quantify the spontaneous recovery of fear memory associated with the cue (CS, conditional stimulus), since it is performed in the same context as extinction learning. The renewal test was performed using the same sound and context of fear acquisition (day 1). The goal of this test was to verify weather extinction of cue-based fear memory was generalized to the context as well, a behavior that is common in very young animals [8, 28, 31].
PV-Cre+/−; Hdac2lox/lox males and their PV-Cre+/−; Hdac2+/+ control male littermates were conditioned using the following protocol. On day 0, mice were allowed to freely explore the context A for habituation. On day 1, mice were conditioned using 5 pairings of the conditioned stimulus (CS duration 30 s, 50 ms pips repeated at 0.9 Hz, 2 ms rise and fall, pip frequency: 7.5 kHz, 80 dB) with an unconditioned stimulus (US, 1 s foot-shock, 0.6 mA, inter-trial interval: 30–150 s). The onset of the US coincided with the offset of the CS. On days 2 and 3, conditioned mice were exposed to extinction training in context B. During this training, mice were exposed to 12 CS at 30 s intervals on each day. Retrieval of extinction and context-dependent fear renewal were tested 7 days later in context B and A, respectively, using 4 presentations of the CS.
Mice behavior was video-recorded with FreezeFrame software. An experimenter blind to the genotype or experimental conditions scored freezing behavior (defined by complete immobility with the exception of respiratory movements) by measuring the time spent freezing during 30 s following CS presentation. Results presented are the mean of time spent freezing after the presentation of the first 2 CS on the first day of extinction (day 2, CS1-2 = early extinction), the last 2 CS on the second day of extinction (day 3, CS 11-12 = late extinction), the first 2 CS during the retrieval test in the extinction context B (day 10, CS 1-2 = retrieval) and the first 2 CS during the renewal test in the fear acquisition context A (day 10, CS 1-2 = renewal).
Mice were excluded from analysis if they did not show effective fear extinction (freezing more than 80% of time at the end of extinction training, Late extinction = day 3, CS 11-12): Fig. 1b: n = 3 control mice excluded, n = 0 PV-Cre+/−; Hdac2lox/lox mouse excluded, Fig. 1g: n = 0 vehicle-injected Hdac2lox mouse excluded, n = 1 BRD6688-injected Hdac2lox mouse excluded, Supplementary Fig. 2: n = 0 PV-Cre+/−; Hdac2+/+ mouse excluded, n = 1 PV-Cre+/−; Hdac2lox/lox mouse excluded. Mice were also excluded from analysis if they did not show effective fear acquisition (freezing less than 15% of time at the onset of extinction training, early extinction = day 2, CS 1-2): Supplementary Fig. 2: n = 0 PV-Cre+/−; Hdac2+/+ mouse excluded, n = 1 PV-Cre+/; Hdac2lox/lox mouse excluded.
Viral vector and stereotaxic injections
pAAV.EF1α-DIO-Flag-Hdac2_T2A_GFP (titre – 1E13 GC/mL) was cloned from pCIG-HDAC2-IRES-eGFP [62] and produced as AAV2/9 serotype by the Canadian Neurophotonics Platform. pAAV_EF1α_DIO_eYFP control virus was produced by the same platform (titre – 5E13 GC/mL [injected at 1:6 dilution]). PV-Cre; Hdac2+/+ or PV-Cre; Hdac2lox/lox mice were used at postnatal day (P) 42-50 for all surgeries. Unilateral viral injections were performed at the following stereotaxic coordinates: +1.9 mm Anterior-Posterior (from Bregma), +0.4 mm Medio-Lateral and 1.6 mm Depth from the cortical surface. Surgical procedures were standardized to minimize the variability of AAV injections. Glass pipette with the virus was lowered at the right depth and kept for 2 min before starting the injections. Each injection volume was kept to 46 nl, and a total of 8 injections per mouse were done for each virus. To ensure minimal leakage into surrounding brain areas and to optimize viral spread in the desired area, injection pipettes were kept in the same position following the injection for 5 min. They were withdrawn at the rate of 0.25 mm/min following the injection. Mice were kept for 3 weeks following viral injections to allow for optimal gene expression. The efficiency of AAV injection was evaluated in each mouse by GFP+ cell density. No mouse was excluded in this analysis.
PFC dissection and dissociation
Mice of both sexes aged P40-P43, were anesthetized with ketamine/xylazine (4:1), transcardically perfused with ice-cold aCSF solution of the following composition (in mM): 62.5 NaCl, 70 sucrose, 2.5 KCl, 25 NaHCO3, 1.25 NaH2PO4, 7 MgCl2, 1 CaCl2, 20 HEPES and 20 glucose. Brains were then quickly removed, transferred to ice-cold aCSF solution and fixed to a stage allowing to cut 300 μm slices with a VT1200 S, Leica vibratome. Slices were individually transferred to a cutting surface in ice-cold aCSF, where frontal cortex was micro dissected from the slices. Dissected tissue was then dissociated into cell suspension [63] using the Papain dissociation system (Worthington) as per manufacturer’s instructions, with the addition of trehalose as described in [64]. Post trituration, cells were then counted using iNCYTO C-Chip hemocytometers, and resuspended at 150–200 cells/μl in DropSeq Buffer (in mM): x1 HBSS, 7 MgCl2, 1 CaCl2, 20 glucose, 5% Trehalose and 10% BSA.
Single cell capture and sequencing
Resuspended cells were run through a custom Drop-Seq setup allowing the capture of single cells and barcoded beads (ChemGenes) within nanoliter droplets as described in the original Drop-seq protocol (http://mccarrolllab.com/dropseq/) [65]. The setup closely resembled the original one [65], with minor modifications. In particular, Medfusion 3500 syringe pumps (Smiths Medical) instead of KD Scientific Legato 100 pumps were used, as well as a different RNase inhibitor (SUPERase• In; catalog no. AM2694; Thermo Fisher Scientific).
Single-cell suspensions (150–200 cells/µl) were run alongside barcoded beads (140 beads µl/1) in 1.7 ml of cell lysis buffer (Tris 200 mM, EDTA 20 mM, 20% Sarkosyl, 20% Ficoll, DTT 50 mM; also containing 50 µl of RNase inhibitor), to produce beads attached to single-cell transcriptomes. Beads attached to single-cell transcriptomes were subsequently washed, reverse-transcribed, treated with exonuclease and counted using a Fuchs-Rosenthal hemocytometer (INCYTO C-Chip Cat-Nr:82030–474), as described in the original Drop-seq protocol [65]. Bead counting was performed using water and bead counting solution (10% polyethylene glycol, 2.5 M NaCl) in a 1:1 ratio as described in Gierahn et al. 2017 [66]. For PCR amplification 4000 beads (approximately 200 single-cell transcriptomes attached to microparticles) were used as input for each PCR. Individual PCR reactions were then pooled to achieve the desired number of single-cell transcriptomes attached to microparticles (STAMPS). In this experiment we used 2000 STAMPS which were cleaned up with AMPure XP (Beckmann Coulter Life Sciences) beads at a ratio of 0.6X prior to sequencing.
Correct size distribution and concentration of complementary DNA was determined using a Bioanalyzer High Sensitivity DNA assay (catalog no. 5067–4626; Agilent Technologies). Library preparation was performed using the Nextera XT DNA Library Preparation Kit (Illumina) with 600 pg input according to the Drop-seq protocol using Illumina i7 primers (N701—N706) together with a custom P5 primer (GCCTGTCCGCGGAAGCAGTGGTATCAACGCAGAGTAC) in the library PCR amplification (see the Drop-seq protocol [65]). Libraries were quality-controlled for size and concentration using the Bioanalyzer High Sensitivity DNA assay. Libraries were quantified using the Universal Library Quantification Kit (KAPA) before sequencing on a NextSeq 550 system (Illumina) at the Institute for Research in Immunology and Cancer (IRIC) in Montreal. Sequencing was performed on a NextSeq 500 system with the read settings R1 = 20 bp, R2 = 63 bp and index = 8 bp.
Computational analysis of single-cell transcriptomic data
Demultiplexing of raw Illumina sequencing data was performed with bcl2fastq v.2.17 by the Institute for Research in Immunology and Cancer (IRIC) in Montreal. Fastq files were quality controlled (Supplementary Fig. 4) and processed using the Drop-seq computational pipeline, similar to earlier report [67] (Drop-seq alignment cookbook v.1.2, January 2016) using Drop-seq tools v.1.12, to produce digital gene expression (DGE) matrices for each sample [65]. DGE matrices were processed in R (3.5.2) using Seurat (v.2.2.0) [68, 69]. Cells with less than 250 genes expressed and genes detected in less than 3 cells were filtered out from further analysis. Cell cycle scores for filtered cells were calculated using the function CellCycleScoring from Seurat with cell cycle gene lists from Nestorowa et al. [70] (https://satijalab.org/seurat/articles/cell_cycle_vignette.html). Expression values were normalized and scaled to 10,000 transcripts with Seurat’s Normalize function. Normalized expression values were scaled using the ScaleData function from Seurat, regressing out the number of UMIs, Mouse ID, percent mitochondrial genes as well as S and G2M scores from cell cycle prediction.
Cells from eight different mice were then aligned with canonical correlation analysis (CCA) [69]. To do so, we used the RunMultiCCA function on the top 1000 highly variable genes overlapping between the samples. We subsequently ran the AlignSubspace function in Seurat, using the experiment as a grouping variable with an appropriate number of CCA dimensions. These dimensions were derived from the inflection point of the elbow plot generated by Seurat ElbowPlot function, resulting in 15 dimensions. The output from this analysis was then analyzed with RunTSNE. followed by FindClusters function, with a resolution parameter of 1.2. The process resulted in identification of 18 total clusters that were subsequently plotted in 2 tSNE dimensions. Biological cell types in the 18 resulting clusters were annotated manually based on known markers for brain cell types. Violin Plots were plotted using the VlnPlot function from Seurat with normalized gene expression values.
Cell type identification
Final cell types were attributed based on the identified top markers in each cluster. Clusters containing VGLUT1 were considered to be pyramidal Neurons, while clusters containing GAD1 and GAD2 were considered to be GABAergic interneurons (Fig. 3a, Supplementary Fig. 5). For more in depth analysis of neuron types as well as identification of non-neuronal cell subtypes, we took the top 30 and 50 markers from each cluster and compared them to the DropVis database [71]. The number of markers shared for each cluster was then calculated, and a percentage identity with DropViz clusters calculated.
Fluorescent multiplex RNAscope
To evaluate Acan mRNA expression in naive condition, mice of both sexes were cervically dislocated, brains were dissected, fast-frozen in cold isopentane (~−70 °C) and conserved at −80 °C. To evaluate Acan mRNA expression after extinction training, adult males were cervically dislocated 3 h after the end of extinction training on day 4 (late extinction).
Coronal brain sections (20 µm) were cut using a cryostat (Leica Microsystems) and mounted on Superfrost Plus Gold Glass Slides (Fisher Scientific, 22-035-813). Slides were subsequently stored at −80 °C. Probes against Acan mRNA (439101), which codes for aggrecan, and Pvalb mRNA (421931-C2), which codes for PV, as well as other reagents for ISH and DAPI labeling, were purchased from Advanced Cell Diagnostics (ACD). Tissue pre-treatment, hybridization, amplification and detection were performed according to the RNAscope Sample Preparation and Pretreatment Guide for Fresh Frozen Tissue using RNAscope Fluorescent Multiplex Assay manual (ACD). During RNAscope hybridization, positive probes (320881), negative probes (320871), and acan/Pvalb probes were processed simultaneously. Briefly, the slides were removed from −80 °C and immediately post-fixed in 4% PFA/PB for 15 min before dehydration in 5 min consecutive baths (70% EtOH, 50% EtOH, 2 ×100% EtOH). The slides were air dried and hydrophobic barrier was created around each section with ImmEdge Pen (310018). Protease IV (322336) was added to each section and incubated at room temperature for 30 min followed by two washes in distilled water. For detection, probes were added to each section and incubated for 2 h at 40 °C. Unbound probes were subsequently washed away by rinsing slides in 1X wash buffer (310091) 2 × 2 min. AMP reagents (AMP1 (320852), AMP2 (320853), AMP3 (320854), AMP4A (C1 probes-Alexa-488 and C2 probes-Atto-550) (320855)) were added to each section and incubates for as per the manufacturer’s instructions (RNAscope Fluorescent Multiplex Reagent Kit PART 2 Manual), and washed in wash buffer for 2 × 2 min. Sections were stained with DAPI (320858) for 30 s, and then mounted with Prolong Diamond Antifade Mountant (Invitrogen P36961). Images were acquired with a SP8-STED confocal microscope (Leica Microsystems), using a 63X (NA 1.4) objective, voxel size: 0.06 µm × 0.06 µm × 0.299 µm, and deconvolved using Huygen’s express deconvolution option. These experiments were performed using tissue from three different mice for each genotype and/or treatment. To determine the number of Acan dots per PV+ cell, images were analyzed using a custom-made ImageJ-Fiji macro. Briefly, a dark region was selected on each focal plane to measure the mean gray value (MGV) as background (BCKG). Then, for each channel and each focal plane, the BCKG value was subtracted, and intensity value of each pixel was readjusted by multiplying it with 65535/(65535-BCKG). Then, each PV+ cell was manually selected based on the presence of Pvalb puncta, filtered with the Gaussian Blur 3D option and analyzed using the 3D objects counter (threshold = 4500, minimal voxel size = 6). Results are presented as mean of the number of objects per PV+ cell ±s.e.m. No mice were excluded from these analysis.
Drug treatment
BRD6688 (Glixx Lab, #GLXC-05908) was dissolved in DMSO (1% of the total resultant solution) and then diluted in 30% Cremophor/69% in physiological saline (H2O containing 0.9% NaCl (Hospira, 04888)), for a final dosage solution of 1 mg/kg. Vehicle solutions consisted of the aforementioned solution without the compound. Solutions were prepared immediately before injection and administered by an investigator blind to the genotype to either Hdac2lox wildtype or PV-Cre+/−; Hdac2lox/lox via intraperitoneal injection 6 h prior to extinction training performed by a second investigator blind for both the genotype and treatment. Mice were randomly allocated to vehicle or BRD6688 injection.
siRNA-delivering peptides
TARBP-BTP RNA-binding protein was produced by the National Research Council Canada (NRC), Montreal, Quebec, Canada, as described previously [44]. The original pET28a-His-pTARBP-BTP coding plasmid was provided by CSIR-CCMB. TARBP-BTP:siRNA complex were prepared as previously described [44, 45]. Silencer Select Acan mouse targeting siRNA (Life technologies, #s61116) and Silencer Select Negative Control siRNA (Life technologies, #4390843) were resuspended at 200 µm with nuclease-free water. For in vivo delivery, TARBP-BTP and siRNA were drop wise mixed together at 5:1 mole ratio. The complex was incubated 20 min at room temperature and administered intravenously (i.v.) via the lateral tail vein at a dose of 20 nmol:4 nmol TARBP-BTP:siRNA in a total volume of 150 µL per mouse. 48 h after injection, 3 P60 Hdac2lox mice injected with siCtrlNeg and 4 P60 Hdac2lox mice injected with siAcan were cervically dislocated, brains were dissected and fast-frozen in cold isopentane (~−70 °C) and conserved at −80 °C before proceeding with Fluorescent multiplex RNAscope. To investigate Acan knockdown effect on spontaneous recovery of fear memory after extinction training, 21 Hdac2lox were fear-trained by a second investigator blind for the treatment using the following protocol. On day 0, mice were allowed to freely explore the context A for habituation. On day 1, mice were conditioned using 5 pairings of the conditioned stimulus (CS, duration 5 s, white noise, 80 dB) with a co-terminating unconditioned stimulus (US, 1 s foot-shock, 0.6 mA, inter-trial interval: 30–60 s). On day 2, 11 mice were i.v. injected with TARBP-BTP:siCtrlNeg and 10 mice were i.v. injected with TARBP-BTP:siAcan at a dose of 20 nmol:4 nmol per mouse. On days 3 and 4, conditioned mice were subjected to extinction training in context B. During this training, mice were presented with 12 CS at 30 s intervals on each day. Spontaneous recovery of fear memory (retrieval of extinction memory test) and context-dependent fear renewal (renewal test) were tested 7 days later in context B and A, respectively, using 4 presentations of the CS. Data analysis was performed by an investigator blind to the treatment and as described above. Mice were randomly allocated to TARBP-BTP:siCtrlNeg or TARBP-BTP:siAcan injection.
Statistical analysis
No statistical methods were used to predetermine sample size. Each experiment included wild type and mutant littermate mice, or mice injected with vehicle/ TARBP-BTP:siCtrlNeg and BRD6688/ or TARBP-BTP:siAcan. Experiments were performed using multiple litters. Statistical analysis was performed with Prism 6.01 (GraphPad software). Data were systematically tested for normal distribution, with the D’Agostino & Pearson omnibus normality test. Differences between two groups of normally distributed data with homogenous variances were analyzed using parametric student’s t-test, while not normally distributed data were analyzed with the Mann–Whitney test. To evaluate genotype effects on PV+Gephyrin+ perisomatic puncta in naive mice and in mice following extinction training, two-ways ANOVA test was performed followed by Sidak posthoc test (variables: genotype, treatment). To evaluate the effect of virally-mediated re-expression of Hdac2 on Aggrecan and PNN, Kruskall–Wallis test was performed followed by Dunn’s multiple comparison test. For fear conditioning analysis, we evaluated the extinction rate by Repeated measures two-ways ANOVA, while freezing time at extinction retrieval and fear renewal tests were analyzed with unpaired two-tailed t-test, if normally distributed and Mann–Whitney if not. Results were considered significant for values of P < 0.05. Data are presented as mean ± standard error of mean.
Data availability
All data are available upon request.
References
Sareen J. Posttraumatic stress disorder in adults: impact, comorbidity, risk factors, and treatment. Can J Psychiatry. 2014;59:460–7.
Lebois LAM, Seligowski AV, Wolff JD, Hill SB, Ressler KJ. Augmentation of extinction and inhibitory learning in anxiety and trauma-related disorders. Annu Rev Clin Psychol. 2019;15:257–84.
Myers KM, Davis M. Mechanisms of fear extinction. Mol Psychiatry. 2007;12:120–50.
Rothbaum BO, Davis M. Applying learning principles to the treatment of post-trauma reactions. Ann NY Acad Sci. 2003;1008:112–21.
Rodriguez BI, Craske MG, Mineka S, Hladek D. Context-specificity of relapse: effects of therapist and environmental context on return of fear. Behav Res Ther. 1999;37:845–62.
Bouton ME, Westbrook RF, Corcoran KA, Maren S. Contextual and temporal modulation of extinction: behavioral and biological mechanisms. Biol Psychiatry. 2006;60:352–60.
Quirk GJ. Memory for extinction of conditioned fear is long-lasting and persists following spontaneous recovery. Learn Mem. 2002;9:402–7.
Kim JH, Richardson R. A developmental dissociation of context and GABA effects on extinguished fear in rats. Behav Neurosci. 2007;121:131–9.
Makkar SR, Zhang SQ, Cranney J. Behavioral and neural analysis of GABA in the acquisition, consolidation, reconsolidation, and extinction of fear memory. Neuropsychopharmacology. 2010;35:1625–52.
Cisneros-Franco JM, de Villers-Sidani E. Reactivation of critical period plasticity in adult auditory cortex through chemogenetic silencing of parvalbumin-positive interneurons. Proc Natl Acad Sci USA. 2019;116:26329–31.
Ribic A, Crair MC, Biederer T. Synapse-selective control of cortical maturation and plasticity by parvalbumin-autonomous action of SynCAM 1. Cell Rep. 2019;26:381–93. e386.
Baho E, Chattopadhyaya B, Lavertu-Jolin M, Mazziotti R, Awad PN, Chehrazi P, et al. p75 neurotrophin receptor activation regulates the timing of the maturation of cortical parvalbumin interneuron connectivity and promotes juvenile-like plasticity in adult visual cortex. J Neurosci. 2019;39:4489–510.
Winkel F, Ryazantseva M, Voigt MB, Didio G, Lilja A, Pou ML, et al. Pharmacological and optical activation of TrkB in Parvalbumin interneurons regulate intrinsic states to orchestrate cortical plasticity. Mol Psychiatry. 2021;26:7247–56.
Chen YH, Hu NY, Wu DY, Bi LL, Luo ZY, Huang L, et al. PV network plasticity mediated by neuregulin1-ErbB4 signalling controls fear extinction. Mol Psychiatry. 2022;27:896–906.
Trouche S, Sasaki JM, Tu T, Reijmers LG. Fear extinction causes target-specific remodeling of perisomatic inhibitory synapses. Neuron. 2013;80:1054–65.
Brittis PA, Flanagan JG. Nogo domains and a Nogo receptor: implications for axon regeneration. Neuron. 2001;30:11–14.
Bhagat SM, Butler SS, Taylor JR, McEwen BS, Strittmatter SM. Erasure of fear memories is prevented by Nogo Receptor 1 in adulthood. Mol Psychiatry. 2016;21:1281–9.
Donato F, Rompani SB, Caroni P. Parvalbumin-expressing basket-cell network plasticity induced by experience regulates adult learning. Nature. 2013;504:272–6.
Favuzzi E, Marques-Smith A, Deogracias R, Winterflood CM, Sánchez-Aguilera A, Mantoan L, et al. Activity-dependent gating of parvalbumin interneuron function by the perineuronal net protein brevican. Neuron. 2017;95:639–55.
Pizzorusso T, Medini P, Berardi N, Chierzi S, Fawcett JW, Maffei L. Reactivation of ocular dominance plasticity in the adult visual cortex. Science. 2002;298:1248–51.
Yap EL, Greenberg ME. Activity-regulated transcription: bridging the gap between neural activity and behavior. Neuron. 2018;100:330–48.
Guan JS, Haggarty SJ, Giacometti E, Dannenberg JH, Joseph N, Gao J, et al. HDAC2 negatively regulates memory formation and synaptic plasticity. Nature. 2009;459:55–60.
Graff J, Joseph NF, Horn ME, Samiei A, Meng J, Seo J, et al. Epigenetic priming of memory updating during reconsolidation to attenuate remote fear memories. Cell. 2014;156:261–76.
Putignano E, Lonetti G, Cancedda L, Ratto G, Costa M, Maffei L, et al. Developmental downregulation of histone posttranslational modifications regulates visual cortical plasticity. Neuron. 2007;53:747–59.
Stafford JM, Raybuck JD, Ryabinin AE, Lattal KM. Increasing histone acetylation in the hippocampus-infralimbic network enhances fear extinction. Biol Psychiatry. 2012;72:25–33.
Morris MJ, Mahgoub M, Na ES, Pranav H, Monteggia LM. Loss of histone deacetylase 2 improves working memory and accelerates extinction learning. J Neurosci. 2013;33:6401–11.
Nott A, Cho S, Seo J, Tsai LH. HDAC2 expression in parvalbumin interneurons regulates synaptic plasticity in the mouse visual cortex. Neuroepigenetics. 2015;1:34–40.
Gogolla N, Caroni P, Luthi A, Herry C. Perineuronal nets protect fear memories from erasure. Science. 2009;325:1258–61.
Carulli D, Pizzorusso T, Kwok JC, Putignano E, Poli A, Forostyak S, et al. Animals lacking link protein have attenuated perineuronal nets and persistent plasticity. Brain. 2010;133:2331–47.
Rowlands D, Lensjø KK, Dinh T, Yang S, Andrews MR, Hafting T, et al. Aggrecan directs extracellular matrix-mediated neuronal plasticity. J Neurosci. 2018;38:10102–13.
Kim JH, Richardson R. A developmental dissociation in reinstatement of an extinguished fear response in rats. Neurobiol Learn Mem. 2007;88:48–57.
Fawcett JW, Fyhn M, Jendelova P, Kwok JCF, Ruzicka J, Sorg BA. The extracellular matrix and perineuronal nets in memory. Mol Psychiatry. 2022;27:3192–203.
Hippenmeyer S, Vrieseling E, Sigrist M, Portmann T, Laengle C, Ladle DR, et al. A developmental switch in the response of DRG neurons to ETS transcription factor signaling. PLoS Biol. 2005;3:e159.
Sousa VH, Miyoshi G, Hjerling-Leffler J, Karayannis T, Fishell G. Characterization of Nkx6-2-derived neocortical interneuron lineages. Cereb Cortex. 2009;19:i1–10.
Segre CV, Chiocca S. Regulating the regulators: the post-translational code of class I HDAC1 and HDAC2. J Biomed Biotechnol. 2011;2011:690848.
Wagner FF, Zhang YL, Fass DM, Joseph N, Gale JP, Weïwer M, et al. Kinetically selective inhibitors of histone deacetylase 2 (HDAC2) as cognition enhancers. Chem Sci. 2015;6:804–15.
Chattopadhyaya B, Di Cristo G, Higashiyama H, Knott GW, Kuhlman SJ, Welker E, et al. Experience and activity-dependent maturation of perisomatic GABAergic innervation in primary visual cortex during a postnatal critical period. J Neurosci. 2004;24:9598–611.
Beurdeley M, Spatazza J, Lee HH, Sugiyama S, Bernard C, Di Nardo AA, et al. Otx2 binding to perineuronal nets persistently regulates plasticity in the mature visual cortex. J Neurosci. 2012;32:9429–37.
Hartig W, Bruckner G, Brauer K, Schmidt C, Bigl V. Allocation of perineuronal nets and parvalbumin-, calbindin-D28k- and glutamic acid decarboxylase-immunoreactivity in the amygdala of the rhesus monkey. Brain Res. 1995;698:265–9.
Morikawa S, Ikegaya Y, Narita M, Tamura H. Activation of perineuronal net-expressing excitatory neurons during associative memory encoding and retrieval. Sci Rep. 2017;7:46024.
Hartig W, Brauer K, Bigl V, Bruckner G. Chondroitin sulfate proteoglycan-immunoreactivity of lectin-labeled perineuronal nets around parvalbumin-containing neurons. Brain Res. 1994;635:307–11.
Rossier J, Bernard A, Cabungcal JH, Perrenoud Q, Savoye A, Gallopin T, et al. Cortical fast-spiking parvalbumin interneurons enwrapped in the perineuronal net express the metallopeptidases Adamts8, Adamts15 and Neprilysin. Mol Psychiatry. 2015;20:154–61.
Levi C, Brooks JM, Chen J, Su J, Fox MA. Cell-specific and developmental expression of lectican-cleaving proteases in mouse hippocampus and neocortex. J Comp Neurol. 2015;523:629–48.
Haroon MM, Dar GH, Jeyalakshmi D, Venkatraman U, Saba K, Rangaraj N, et al. A designed recombinant fusion protein for targeted delivery of siRNA to the mouse brain. J Control Rel. 2016;228:120–31.
Haroon MM, Saba K, Boddedda VH, Kumar JM, Patel AB, Gopal V. Delivery of BACE1 siRNA mediated by TARBP-BTP fusion protein reduces beta-amyloid deposits in a transgenic mouse model of Alzheimer’s disease. J Biosci. 2019;44:1.
Bavelier D, Levi DM, Li RW, Dan Y, Hensch TK. Removing brakes on adult brain plasticity: from molecular to behavioral interventions. J Neurosci. 2010;30:14964–71.
Dehorter N, Del Pino I. Shifting developmental trajectories during critical periods of brain formation. Front Cell Neurosci. 2020;14:283.
Ganai SA, Ramadoss M, Mahadevan V. Histone Deacetylase (HDAC) Inhibitors - emerging roles in neuronal memory, learning, synaptic plasticity and neural regeneration. Curr Neuropharmacol. 2016;14:55–71.
Bouton ME. Context, time, and memory retrieval in the interference paradigms of Pavlovian learning. Psychol Bull. 1993;114:80–99.
Craske MG, Treanor M, Conway CC, Zbozinek T, Vervliet B. Maximizing exposure therapy: an inhibitory learning approach. Behav Res Ther. 2014;58:10–23.
Marshall PR, Bredy TW. Neuroepigenetic mechanisms underlying fear extinction: emerging concepts. Psychopharmacology. 2019;236:133–42.
Carceller H, Guirado R, Ripolles-Campos E, Teruel-Marti V, Nacher J. Perineuronal nets regulate the inhibitory perisomatic input onto parvalbumin interneurons and gamma activity in the prefrontal cortex. J Neurosci. 2020;40:5008–18.
Cabungcal JH, Steullet P, Morishita H, Kraftsik R, Cuenod M, Hensch TK, et al. Perineuronal nets protect fast-spiking interneurons against oxidative stress. Proc Natl Acad Sci USA. 2013;110:9130–5.
Courtin J, Chaudun F, Rozeske RR, Karalis N, Gonzalez-Campo C, Wurtz H, et al. Prefrontal parvalbumin interneurons shape neuronal activity to drive fear expression. Nature. 2014;505:92–96.
Tasic B, Yao Z, Graybuck LT, Smith KA, Nguyen TN, Bertagnolli D, et al. Shared and distinct transcriptomic cell types across neocortical areas. Nature. 2018;563:72–78.
Devienne G, Picaud S, Cohen I, Piquet J, Tricoire L, Testa D. et al. Regulation of perineuronal nets in the adult cortex by the activity of the cortical network. J Neurosci. 2021;41:5779–90.
Giamanco KA, Morawski M, Matthews RT. Perineuronal net formation and structure in aggrecan knockout mice. Neuroscience. 2010;170:1314–27.
Kwok JC, Carulli D, Fawcett JW. In vitro modeling of perineuronal nets: hyaluronan synthase and link protein are necessary for their formation and integrity. J Neurochem. 2010;114:1447–59.
Faini, G, Aguirre, A, Landi, S, Lamers, D, Pizzorusso, T, Ratto, GM et al. Perineuronal nets control visual input via thalamic recruitment of cortical PV interneurons. Elife. 2018;7. https://doi.org/10.7554/eLife.41520.
Baker KD, Gray AR, Richardson R. The development of perineuronal nets around parvalbumin gabaergic neurons in the medial prefrontal cortex and basolateral amygdala of rats. Behav Neurosci. 2017;131:289–303.
Shi W, Wei X, Wang X, Du S, Liu W, Song J, et al. Perineuronal nets protect long-term memory by limiting activity-dependent inhibition from parvalbumin interneurons. Proc Natl Acad Sci USA. 2019;116;27063–73.
Nott A, Nitarska J, Veenvliet JV, Schacke S, Derijck AA, Sirko P, et al. S-nitrosylation of HDAC2 regulates the expression of the chromatin-remodeling factor Brm during radial neuron migration. Proc Natl Acad Sci USA. 2013;110:3113–8.
Munoz-Manchado AB, et al. Diversity of interneurons in the dorsal striatum revealed by single-cell RNA sequencing and PatchSeq. Cell Rep. 2018;24:2179–90 e2177.
Saxena A, Wagatsuma A, Noro Y, Kuji T, Asaka-Oba A, Watahiki A, et al. Trehalose-enhanced isolation of neuronal sub-types from adult mouse brain. Biotechniques. 2012;52:381–5.
Macosko EZ, Basu A, Satija R, Nemesh J, Shekhar K, Goldman M, et al. Highly parallel genome-wide expression profiling of individual cells using nanoliter droplets. Cell. 2015;161:1202–14.
Gierahn TM, Wadsworth MH 2nd, Hughes TK, Bryson BD, Butler A, Satija R, et al. Seq-Well: portable, low-cost RNA sequencing of single cells at high throughput. Nat Methods. 2017;14:395–8.
Wunnemann F, Ta-Shma A, Preuss C, Leclerc S, van Vliet PP, Oneglia A, et al. Loss of ADAMTS19 causes progressive non-syndromic heart valve disease. Nat Genet. 2020;52:40–47.
Satija R, Farrell JA, Gennert D, Schier AF, Regev A. Spatial reconstruction of single-cell gene expression data. Nat Biotechnol. 2015;33:495–502.
Butler A, Hoffman P, Smibert P, Papalexi E, Satija R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol. 2018;36:411–20.
Nestorowa S, Hamey FK, Pijuan Sala B, Diamanti E, Shepherd M, Laurenti E, et al. A single-cell resolution map of mouse hematopoietic stem and progenitor cell differentiation. Blood. 2016;128:e20–31.
Saunders A, Macosko EZ, Wysoker A, Goldman M, Krienen FM, de Rivera H, et al. Molecular diversity and specializations among the cells of the adult mouse brain. Cell. 2018;174:1015–30.e1016.
Acknowledgements
We thank Dr. Elsa Rossignol for her insightful suggestions and critical reading of the manuscript. We would like to thank Josianne Nunes-Carriço, Antônia Samia Fernandes do Nascimento and Antoine Farley for their technical assistance. We acknowledge the Comité Institutionnel de Bonne Pratiques Animales en Recherche (CIBPAR), and all the personnel of the animal facility of the Research Center of CHU Sainte-Justine (Université de Montreal), as well as Dr. Elke Küster-Schöck at the plateforme d’imagerie microscopique (PIM) for instrumental technical support. We thank J. Huber at the IRIC genomic platform for performing the Illumina sequencing for the Drop-seq libraries and P. Gendron at the IRIC Bioinformatics platform for data demultiplexing. We thank the Canadian Neurophotonic Platform for viral production and Drs. A.Nott and L.H. Tsai for providing plasmids. We also would like to thank Andrei Vlad Varlan, Kayleigh Gauvin and Théo Badra for their help with data analysis, CCMB’s central facility for plasmid DNA sequencing, and the National Research Council Canada for TARBP-BTP carrier protein production and purification.
Funding
This work was supported by the Canadian Institutes of Health Research (GP, GDC), Natural Sciences and Engineering Research Council of Canada (GDC), EraNet-Neuron (DECODE!, GDC). ML-J was supported by le Fonds de Recherche du Québec en Santé (FRQS).
Author information
Authors and Affiliations
Contributions
MLJ, GDC designed the experiments. MLJ, BC, PC, DC, SL, FD, DR performed the experiments. MLJ, BC, PC, FW, FD, DR, HA, GP and GDC analyzed data. GA and GP provided critical technical support. KS, VG, ABP contributed reagents. MLJ, BC, GP, GDC wrote the manuscript. All authors read and corrected the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lavertu-Jolin, M., Chattopadhyaya, B., Chehrazi, P. et al. Acan downregulation in parvalbumin GABAergic cells reduces spontaneous recovery of fear memories. Mol Psychiatry 28, 2946–2963 (2023). https://doi.org/10.1038/s41380-023-02085-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-023-02085-0
- Springer Nature Limited
This article is cited by
-
A disinhibitory microcircuit of the orbitofrontal cortex mediates cocaine preference in mice
Molecular Psychiatry (2024)