Abstract
Machine learning approaches using structural magnetic resonance imaging (sMRI) can be informative for disease classification, although their ability to predict psychosis is largely unknown. We created a model with individuals at CHR who developed psychosis later (CHR-PS+) from healthy controls (HCs) that can differentiate each other. We also evaluated whether we could distinguish CHR-PS+ individuals from those who did not develop psychosis later (CHR-PS-) and those with uncertain follow-up status (CHR-UNK). T1-weighted structural brain MRI scans from 1165 individuals at CHR (CHR-PS+, n = 144; CHR-PS-, n = 793; and CHR-UNK, n = 228), and 1029 HCs, were obtained from 21 sites. We used ComBat to harmonize measures of subcortical volume, cortical thickness and surface area data and corrected for non-linear effects of age and sex using a general additive model. CHR-PS+ (n = 120) and HC (n = 799) data from 20 sites served as a training dataset, which we used to build a classifier. The remaining samples were used external validation datasets to evaluate classifier performance (test, independent confirmatory, and independent group [CHR-PS- and CHR-UNK] datasets). The accuracy of the classifier on the training and independent confirmatory datasets was 85% and 73% respectively. Regional cortical surface area measures-including those from the right superior frontal, right superior temporal, and bilateral insular cortices strongly contributed to classifying CHR-PS+ from HC. CHR-PS- and CHR-UNK individuals were more likely to be classified as HC compared to CHR-PS+ (classification rate to HC: CHR-PS+, 30%; CHR-PS-, 73%; CHR-UNK, 80%). We used multisite sMRI to train a classifier to predict psychosis onset in CHR individuals, and it showed promise predicting CHR-PS+ in an independent sample. The results suggest that when considering adolescent brain development, baseline MRI scans for CHR individuals may be helpful to identify their prognosis. Future prospective studies are required about whether the classifier could be actually helpful in the clinical settings.
Similar content being viewed by others
Introduction
The clinical high risk (CHR) paradigm is widely used with the goal of improving early detection of and prevention of psychotic disorders [1]. Individuals are considered at CHR for psychosis if they meet criteria for attenuated positive symptom syndrome (APSS), brief intermittent (limited) psychotic syndrome (BLIPS), and/or genetic risk and deterioration syndrome (GRDS) based on semistructured interviews [2,3,4,5]. The CHR state is present in 1.7% of the general population and 19.2% of clinical samples [6]. CHR individuals have a higher risk of developing psychosis (0.15 at 1 year) comparing to healthy controls, the transition risk increased from 0.09 at half years to 0.27 at 4 years [7]. However, most CHR subjects who do not transition to psychosis will continue to meet CHR criteria or experience attenuated psychosis symptoms at follow-up and only 33% will eventually remit [7, 8].
The CHR state, is also associated with alterations in proxy measures of brain structure [9,10,11,12,13,14,15]. Previous structural magnetic resonance imaging (MRI) studies reported a progressive decrease in gray matter volume in the medial and superior temporal and medial frontal cortex during the transition period among CHR individuals [14,15,16,17]. Gray matter volume continued to decrease several years after disease onset [15, 16, 18]. Cortical surface area (SA) and cortical thickness (CT), which can be extracted using FreeSurfer software [19,20,21], are also crucial predictors of important life outcome [22] and associated with neurological, psychological, and behavioral traits [23]. SA is strongly correlated with grey matter volume compared to CT, suggesting SA and CT are unique structural features in the grey matter cortex [24, 25]. Recent study indicated that the multivariate architectures with respect to the makeup of the genetic factors were distinct across cortical surface area and thickness [22]. This is in line with the radial unit hypothesis [26] that the expansion of cortical surface area is driven by the proliferation of neural progenitor cells, whereas numbers of neurogenetic division of these cells for thickness [23]. Widespread lower CT has also been identified in cross-sectional MRI data in individuals at CHR in a large-scale pooled analysis of the Enhancing Neuro Imaging Genetics through Meta-Analysis (ENIGMA) CHR Working Group [27]. Among these widespread alterations, frontal cortical and temporal regions (e.g., fusiform, superior temporal, and paracentral) have been relatively consistently associated with CHR status [9,10,11, 28,29,30], with these regions also exhibiting lower CT in individuals with established schizophrenia [29]. In addition to regional changes, individuals with CHR, have showed greater neuroanatomical variability in global SA, CT, and subcortical volume compared to HC [31]. Furthermore, longitudinal studies have shown reductions of cortical thickness in the paracentral, superior temporal, and fusiform gyrus have been reported to be associated with psychosis conversion in those at CHR [13, 14, 32]. Recent work has indicated that whole-brain sMRI patterns of schizophrenia forecasted 2-year psychosocial impairments in individuals with CHR [33], suggesting that alterations in brain structure may predict real-life outcomes.
Adolescent development is a crucial time window that is associated with brain-wide changes, including reductions in cortical thickness and volume [34, 35]. Cortical characteristics such as gray matter volume, cortical surface area, and cortical thickness decline by about 10% during adolescence [36]. On the other hand, white matter volume was reported peaking in young adulthood [36]. Since the period from adolescence to early adulthood is a high risk time window for psychosis onset [32], age-related anatomical deviations from typically-occuring declines may hold valuable information to predict later psychosis conversion, especially in frontal and temporal regions that have been implicated in CHR [27, 32, 37,38,39] and schizophrenia [40,41,42,43,44,45]. Further, greater brain age deviations were found to be associated with a higher risk for psychosis over time [11, 38]. Importantly, these results suggest that the adolescent brain development pattern of CHR individuals may differ from that of HCs. Indeed, the ENIGMA CHR Working Group has reported that CHR compared to HC participants exhibit altered non-linear age associations with cortical thickness [27], suggesting that cross-sectional between-group differences in sMRI metrics may involve altered adolescent development, trait characteristics associated with psychosis liability, and/or progressive brain pathology around the onset of psychosis [32, 39, 46].
An increasing number of studies have attempted to use (cross-sectional) sMRI data to predict outcome or case-control status. These prior studies show that machine learning approaches are informative for differentiating individuals with schizophrenia from HCs [47,48,49,50,51,52]. Similar findings were observed in different clinical stages of psychosis, including first episode schizophrenia and CHR individuals [48, 49]. A major limitation, however, is the need for large and diverse sample sizes to establish a well-tuned classifier that also provides generalized predictive performance [12, 53]. Since single sites cannot typically provide the necessary sample sizes [49, 54, 55], multisite consortia data may be advantageous if site effects are adequately accounted for (e.g., via cross-site harmonization procedures) [49, 54, 56]. For example, without harmonization, a prior study failed to build a useful model with multi site data [38]. In the current study, we aimed to investigate whether cross-sectional sMRI data can be used to build a classifier to differentiate the neuroanatomical developmental patterns of HCs relative to participants who later developed a psychotic disorder (CHR-PS+) as biomarkers for future psychosis conversion. As altered developmental processes are implicated in psychosis risk, we considered the potential non-linear effects of age and sex to gain optimal predictive accuracy of trained classifiers.
Here, we combined data from 21 sites harmonized through the ENIGMA CHR Working Group using ComBat [57] to minimize differences related to site-, scanner- and scanning protocols using an Empirical Bayes method. Second, to model non-linear age effects, we fitted generalized additive models (GAMs) [58, 59] to the HC data, and then applied the fitted GAMs to obtain non-linear age- and sex-corrected features for the entire sample [60]. More specifically, we estimated the model in HCs and applied it to individuals at CHR to capture deviations from the expected patterns of physiological aging. As for patients with early-onset psychosis [61] and schizophrenia [41] have been reported to have abnormally low estimated intracranial volume (ICV), all procedures were performed after adjusting the MRI features for effects of ICV. Third, we developed an XGBoost [62] classifier using only HCs and CHR-PS+ to determine deviation in neuroanatomical developmental patterns as potential predictors of future psychosis conversion. Finally, we tested the predictive performance of the classifier with the left-out site data, to avoid the potential for information leakage between the training and test data.
We hypothesized that CHR-PS+ individuals would be distinguishable from HCs based on features derived from structural MRI features, based on the assumption that those CHR individuals who are most likely to convert to psychosis would show the greatest baseline anatomical alterations. Second, we expected our classifier to label individuals at CHR who had not developed a psychotic disorder (CHR-PS-) at follow-up, and individuals at CHR who did not complete follow-up visits, resulting in missing information about their transition status (CHR-UNK), as HCs. Third, we expected the classifiers to perform similarly in independent confirmatory datasets, and expected to find associations between classifications and symptom severity.
Methods
Participants
We included data from a total of 1165 CHR individuals (144 CHR-PS+, 793 CHR-PS−, and 228 CHR-UNK individuals) and 1029 healthy controls (HCs) from 21 ENIGMA Clinical High Risk for Psychosis Working Group sites (Table 1). As previous study showed that using CHR psychometric instruments to assess the CHR state in clinical samples is associated with an excellent overall prognostic performance [63], we combined two assessments directly as previous studies [27, 31, 64]. CHR status was assessed using the full version of the Comprehensive Assessment of At-Risk Mental States (CAARMS [65]; n = 650) or the Structured Interview for Prodromal Syndromes (SIPS [66, 67]; n = 799). Site-specific inclusion and exclusion criteria, the available scale scores in premorbid IQ, symptom severity, global functioning, and antipsychotic use at scan are the same as in a prior publication (Supplementary Table S1) [27]. All sites obtained local institutional review board approval prior to data collection. Written informed consent was obtained from every participant, or from the participant’s guardian for participants younger than 18 years. All studies were conducted in accordance with the Declaration of Helsinki [68].
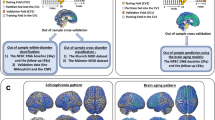
We applied a two-step approach [49] to evaluate the performance of the models by dividing the data into four datasets: training, test, independent confirmatory, and independent group datasets (Fig. 1). Test and independent confirmatory datasets were used as external validation datasets. First, the training and test datasets comprised the data from CHR-PS+ and HC from 20 sites except for Toyama, which was used as the independent confirmatory dataset. We chose this dataset because the Toyama site contributed the largest HC sample and excluding this dataset reduced sample imbalance between groups in building a machine learning classifier. Ninety percent of the data were randomly sorted as the training dataset, and the remaining 10% as the test dataset. A Kolmogorov–Smirnov test did not show any significant differences between training and test datasets in any structural features. The independent confirmatory dataset comprised the data from HCs and CHRs at the Toyama site; this data was completely excluded from the training partition, and was used to perform an independent first-step evaluation without site information leakage. To evaluate the classifier on unseen new data, we defined the CHR-PS− and CHR-UNK individuals in all sites as the independent group dataset to perform the second step.
MRI data acquisition and preprocessing
Image acquisition and Processing
Participating sites contributed to T1-weighted MRI brain scans from 31 MRI scanners, including 29 3-T scanners and 2 1.5-T scanners (Supplementary Table S2). Detailed scan protocols and the number of scans for each protocol are described in the Supplementary Materials. After processing the data using FreeSurfer analysis software at each site [19,20,21], we extracted structural features from 153 regions of interest (ROI) including 68 regional measures of cortical thickness, 68 surface area (SA), 16 subcortical volume, and one intracranial volume according to the Desikan-Killiany atlas [69]. We implemented the ENIGMA consortium quality assessment pipeline [40, 41, 70,71,72,73] and 8 samples were excluded for lacking 20% of the ROIs data. Remaining missing values (1.20%) were imputed using a k-Nearest Neighbor (k = 3) approach.
ComBat harmonization
ComBat [57] is a harmonization method used to remove scanner and protocol effects based on an adjusted general linear model harmonization method. Based on recent work demonstrating that neuroComBat harmonization increases statistical power within a mega-analytic framework, primary analyses were conducted within a mega-analysis framework using data that were corrected for site and scanner associations using neuroComBat harmonization [74]. Further analyses were conducted using Python version 3.7.12. We applied the extracted cortical thickness, surface area, subcortical volume, and intracranial volume measures with participants’ age and sex as covariates, along with protocol and site effects. To confirm that group status had no significant influence on the ComBat harmonization steps, we also conducted ComBat harmonization using the training dataset only (see Supplementary Materials).
Features engineering
First, we fitted a general linear model to regress out effects of intracranial volume. Next, we fitted GAMs to only the HC data to estimate non-linear effects of age and sex for every structural feature; then we applied the fitted GAMs to obtain non-linear age- and sex-corrected features. To verify the absence of information leakage and the stability of the GAMs, we also repeated this procedure 1000 times on randomly sub-sampled HC data to estimate the GAMs (see Supplementary Materials and Fig. S1).
XGBoost
XGBoost is a scalable tree boosting algorithm [62]. We applied standardization for the structural features to building a classifier. The use of input data standardization, optimization of the hyperparameters of the classifier (eta, min_child_weight, max_depth, subsample, colsample_bytree) were tuned using GridSearchCV implemented in the ‘scikit-learn’ module (version 1.0.2) in Python (https://scikit-learn.org/stable/auto_examples/release_highlights/plot_release_highlights_1_0_0.html) [75]. We plotted the weights of the classifier to determine the importance of the features for generalization. The classifier was optimized using a tenfold cross-validated grid search over a defined parameter grid. Data from the HC group were randomly downsampled to the same sample size as the CHR-PS+ group in each fold. To reduce downsampling bias, downsampling and grid search were repeated 1000 times and stratified tenfold for the training data. Then, we applied tenfold cross-validation and 1000 permutations to evaluate the significance of the cross-validation scores of the model with the best hyperparameters for the training dataset. The best cross-validation accuracy score was averaged across 1000 repeats. Permutation tests were conducted by shuffling the labels in the training data, and the permutation-based p-value was calculated [76]. The final model with the best hyperparameters was trained using the entire training dataset. Finally, the trained classifier was applied to the test set and the independent confirmatory dataset with the best parameters tuned by grid search. The predict probability was calculated by the trained classifier for each sample. Predict probability ranges from 0 to 1, with smaller values indicating more likely classification as CHR-PS+. The cut-off point for the predictive performance was set to 0.5.
The predictive performance of the classifier was evaluated using an independent group dataset (CHR-PS− and CHR-UNK). We compared the classifiers built from four different feature sets: (i) only cortical thickness values, (ii) only surface area values, (iii) only subcortical volumes only, and (iv) all features. The classifier with the best predictive performance for the independent confirmatory dataset was used for subsequent analysis.
Statistical analysis
Evaluation metrics
First, the classifier was evaluated using the test, independent confirmatory, and independent group datasets by the given scores of the tuned classifier. We calculated the confusion matrix, macro, and weighted average accuracies to evaluate the classifier because the data used were imbalanced (see Supplementary Materials) [49].
Predictive performance of the classifier
The predictive performance of the classifier was defined as its performance on unseen data (in the independent confirmatory/group datasets) and was assessed using standard evaluation metrics. Chi-squared tests were applied to the classified labels of the test, independent confirmatory, and independent group datasets. Since we conducted a total of 6 comparisons, a Bonferroni’s correction was applied to adjust for the multiple statistical comparisons (p < 0.05/6 = 0.008). Predict probabilities generated by the XGBoost classifier were also tested using a nonparametric analysis of variance for all samples. To confirm little difference in the predictive performance between the assessments of CHR state, we tested the difference of predict probabilities including SIPS or CAARMS as a covariate. We also tested the difference in the rates of individuals predicted as CHR-PS+ that were assessed by either SIPS or CAARMS using a Chi-squared test. A GAM was used to assess non-linear relationships between age and the predictive performance of the classifier. Moreover, we conducted 4 comparisons (HCs vs. CHR-PS+, CHR-PS+ vs. CHR-PS-, CHR-PS+ vs. CHR-UNK and CHR-PS+ vs. CHR-PS- and CHR-UNK) of the decision curve analysis [77,78,79] using ‘dcurves’ package (version 0.4.0) in R software to estimate the classifier as well. Net benefit was calculated across a range of threshold probabilities [80] in comparison to getting MRI measurements to get a prediction for all patients or no patients. As threshold probabilities were set up to 50% (i.e., chance level), net benefit = sensitivity × prevalence – (1 – specificity) × (1 – prevalence) × 50%.
Relationship between predict probability and demographic and clinical characteristics
We tested the difference in the predictive performance with respect to sex and the existence of APSS, BLIPS, and GRDS using t tests (p < 0.05/3 = 0.016). Pearson’s correlation analyses were also conducted between standardized IQ and the predict probability. Bonferroni’s correction was applied to the subscores (p < 0.05/4 = 0.0125). To determine the relationship between the predict probability and symptom severity, Pearson’s correlation analyses were performed using the SIPS and CAARMS subscores for CHR-PS+, CHR-PS-, and CHR-UNK groups. We tested z-score normalized positive, negative and general subscores of the SIPS and CAARMS using Pearson’s correlation coefficients. Bonferroni’s correction was applied to the SIPS or CAARMS subscores (for SIPS: positive, negative, disorganization and general symptoms, p < 0.05/4 = 0.0125; for CAARMS: positive symptoms, cognitive change, emotional disturbance, negative symptoms, behavioral change, motor/physical changes, p < 0.05/6 = 0.0083). To determine the potential effect of antipsychotic medication on the classification, we also tested the difference in predict probabilities between those with and without medication use for each CHR subgroup using a t-test.
Results
Model evaluation
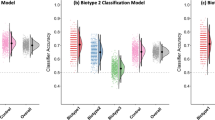
A non-linear effect of age, sex, and age x sex interaction on SA was found in HCs, as shown in Fig. 2. The classifier using only non-linear fitted SA features (i.e., fit to HCs, applied to all) obtained the best performance in differentiating HC and CHR-PS+ groups (Supplementary Table S3). For the SA model, the best cross-validation accuracy within the training dataset was 85% (± 0.00008). The permutation test showed that the classifier performed significantly better than chance level (50%, p < 0.001). The accuracies with the best estimator for the test and independent confirmatory datasets were 68% and 73% (Fig. 3B), respectively. Regions with the top ten largest features weights were the superior temporal, insula, superior frontal, superior parietal, fusiform, isthmus of cingulate, parahippocampal gyri, and postcentral gyri to differentiate HC from CHR-PS+ groups (Fig. 3A, Supplementary Table S3). For SA in the right superior temporal gyrus, which was the strongest contributing feature of the classifier, the ComBat harmonized feature showed no significant difference among the groups (p > 0.05), while ComBat harmonized and non-linear age- and sex-adjusted feature revealed a difference between CHR-PS+ and CHR-PS- (t = 2.137, p = 0.0327), and CHR-PS+ and CHR-UNK (t = 2.140, p = 0.0325; Fig. 4).
A Weighted surface area features of XGBoost classification in Desikan-Killiany atlas. B Predictive performance of HC and CHR-PS+ groups was evaluated using the independent confirmatory dataset, and CHR-PS- and CHR-UNK groups using the independent group dataset. C Box and scatter plot of predict probabilities of XGBoost. P-values of post hoc comparisons were corrected using a Bonferroni method (***p < 0.001, **p < 0.01, *p < 0.05). D Best fit for the association of age with the predict probability in a GAM. Shading around the line indicates the standard error. E Decision curve analysis showed the benefits of XGBoost predicting the risk of psychosis conversion according to MRI scan.
For a confirmatory analysis, machine learning classifiers using 152 sMRI raw brain characteristics showed poorer performance compared to the corresponding age- and sex-adjusted machine learning classifiers (Supplementary Materials). We also tried to build classifiers to differentiate CHR from HCs or CHR-PS+ from CHR-PS-, however, those ones only showed approximate chance level (50%) accuracies.
Predictive performance of the classifier for the test, independent confirmatory, and independent group datasets
A chi-squared test showed a significant difference in the classified labels for the independent confirmatory, and independent group datasets, respectively (X2(1, n = 151) = 6.34, p = 0.012 and X2(1, n = 1021) = 4.39, p = 0.036). Further residual analysis showed that the HC group was significantly more likely to be classified as HCs than the CHR-PS+ group (73% vs. 30%, corrected p = 0.004, Fig. 3B). For the independent group dataset, no difference between CHR-PS- and CHR-UNK groups was found (73% vs. 80%, corrected p = 0.029).
For the overall sample, a chi-square test showed a significant difference in the classified labels between the four groups (X2(3, 1172) = 15.12, p = 0.002). Further residual analysis showed a significant difference in the classified labels between CHR-PS+ and the other three groups (Bonferroni corrected p’s < 0.05; Fig. 3B). For the predict probability, an Kruskal-Wallis test showed a significant difference between the four groups (H = 278.86, p < 0.001). Post-hoc comparisons showed that CHR-PS+ group was different from all other groups and that the CHR-PS- group was in between CHR-PS+ and HC groups (HC > CHR-PS- > CHR-PS+), while the predict probability did not differ between CHR-UNK and HCs (CHR-UNK > CHR-PS- > CHR-PS+; Bonferroni corrected p’s < 0.05; Fig. 3C). The difference changed little after controlling the methods of the CHR assessments as a covariate (CHR group: F(2, 1161) = 192.25, p < 0.001; Assessment method: F(1, 1161) = 0.00, p > 0.05), and CHR individuals predicted as CHR-PS+ did not differ between participants assessed with SIPS versus CAARMS, X2(1, 1449) = 2.59, p > 0.05; (Supplementary Table S4). Although the classifier was built according to the features after controlling for non-linear age effect, a GAM analysis demonstrated that the predict probability was associated with age (F = 11.33, p = 0.003), and differed between CHR-PS+ and HC (t = 20.72, p < 0.001), CHR-PS+ and CHR-PS- (t = 17.83, p < 0.001), and CHR-PS+ and CHR-UNK (t = 17.64, p < 0.001; Fig. 3D). No significant age × group interaction was found in the predict probability.
The estimated decision curve for all comparisons (HCs vs. CHR-PS+, CHR-PS+ vs. CHR-PS-, CHR-PS+ vs. CHR-UNK and CHR-PS+ vs. CHR-PS- and CHR-UNK) showed that in clinical setting, compared to MRI measurement for all patients or no MRIs at all, getting a prediction from current classifier/model leads to higher net benefit to discoverer transition of CHR (Fig. 3E).
Relationship between predict probability and demographic and clinical characteristics
We observed no effects of sex or APSS, BLIPS, or GRDS status, on the predict probability (p > 0.05). No significant correlations were found between standardized IQ and the predict probability for each group. No significant correlation was found between symptom severity scores and predict probability. No significant difference was found for the antipsychotics use was found among each CHR group (p > 0.05).
Discussion
To the best of our knowledge, the current study is the one of a few to apply a machine learning approach to discriminate HC and CHR-PS+ groups in a large multisite sample [12]. To evaluate the classifier, we employed a two-step approach using an independent confirmatory dataset, obtained at a different site and using a different protocol from the ones used to build the classifier; we also used an independent group dataset including CHR-PS- and CHR-UNK groups. Although previous study reported 94% accuracy [12], we have achieved 85% accuracy in the 2-class classification in the training dataset using non-linear adjustment of SA features for age and sex. The patterns of neuroanatomical alterations were also useful in identifying CHR-PS- individuals. Specially, of the CHR groups, the CHR-UNK group was the most likely to be classified as HC by the classifier, than those in other CHR groups, showing no difference in the predict probability from HC.
In this study, we were able to differentiate HC from CHR-PS+ group with 85% and 68% accuracy in the training and test sets, respectively. The performance accuracy achieved by the classifier on the independent confirmatory dataset was 73%. In contrast to prior studies [12, 38, 53], we successfully built a model with promising predictive performance for new data. Our findings suggest that ComBat is not only useful to increase statistical power [55, 57, 74] but also crucial for improving the accuracy in building a machine learning model out of multisite data. As expected, the majority of CHR-PS- and CHR-UNK individuals were classified as HCs. Moreover, no significant associations were found between the predict probability and sex or IQ, or antipsychotics use for each CHR group. We suggest that a machine learning classifier trained to identify differences between CHR-PS+ and healthy controls may be helpful to identify UHR individuals at risk for conversion.
In line with prior studies of cortical alterations in CHR [11, 37,38,39, 43], we found that the pattern of SA features, including the superior temporal, insula, superior frontal, superior parietal, fusiform, isthmus of cingulate, and parahippocampal gyri, contributed to identifying CHR-PS+ from HCs (Fig. 2A, Fig. 4). These findings align with previous work reporting (right) superior temporal gyrus functional alterations may underlie deficits in (non-)emotional multisensory integration in schizophrenia patients [81] and working memory-related dysfunction in CHR [82]. CHR individuals who converted or presented with greater clinical symptom within a 2-year follow-up exhibited smaller SA in the rostral anterior cingulate, lateral and medial prefrontal regions, and parahippocampal gyrus [11]. SA is more closely related to volume than cortical thickness [24], and the volume of the isthmus of cingulate gyrus has been reported to be different in resilient and non-resilient CHR individuals [39]. The neuroanatomical alteration/ deviance pattern of SA found in the current study between HCs and CHR-PS+ groups are consistent with findings from other studies, which implicate the volume of superior temporal, frontal and fusiform regions in CHR transitions [27, 38] and schizophrenia [29, 41, 42]. Our initial ENIGMA CHR study showed the differences mainly in CT, and for two regions (i.e., the paracentral lobule and fusiform gyrus), the non-linear pattern of the age trajectory differed between HC and CHR [27, 83]. However, the former study was focused on the statistical significance, and the current study is focused on predictive performance. As traditional significance approaches do not capture predictive variable sets [84], resulting in SA serving better building a predictive model. It is possible that the current study engineered the features that made the differences in SA more prominent, by using GAM to estimate the brain age gap in a non-linear manner. Moreover, as the result of GAM eliminating the non-linear adolescent development of SA in differentiating HCs and CHR-PS+, our classifier achieved promising generalization of predictive performance.
Although we did not find any difference in predict probability between APSS, BLIPS, or GRDS status, it is important to note that previous studies demonstrated CHR subgroup-specific changes in sMRI metrics [85], such as subcortical volume reductions in left anterior frontal, right caudate, right hippocampus, and amygdala in CHR with a genetic risk, while CHR with attenuated psychotic symptoms exhibited right middle temporal cortical reduction [86]. Moreover, studies have shown that transition rates may differ between CHR subgroups [87]. These findings underscore the importance of using adequate sampling of CHR participants across subgroups and different clinical stages. Such efforts may result in more accurate predictive models in the future.
The predict probability given by the classifier based on the neuroanatomical deviance showed significant differences among the HC or CHR-UNK, CHR-PS-, and CHR-PS+ groups at baseline (HC, CHR-UNK > CHR-PS- > CHR-PS+; Fig. 3C). The results suggest that predict probability is a useful index allowing us to better understand how neuroanatomical deviance is associated with psychosis conversion. This further implies that the neuroanatomical deviance was already observed at baseline in CHR-PS+ group. Moreover, in contrast to previous working reporting a positive association between age conversion rates [83], our observed association between predict probability and age (Fig. 3D) could suggest that the likelihood of a HC prediction increases with age. One possibility for this finding is the distribution of age across groups. Specifically, participants older than 30 years old were sparsely distributed in all groups in the current study, which may result in spurious associations between age and predict probability. To understand the exact nature of the association between age and predict probability, more data of CHR participants of older ages is necessary. These results suggest that psychosis-related brain characteristics may decrease according to brain development which may effect on the onset of psychosis.
Our study has several limitations. First, to harmonize site effects, ComBat was applied to both HC and CHR subjects which by assuming a common covariate model (that is typically preserved by ComBat) might potentially lead to an information leak [88]. However, without traveling subject harmonization, ComBat was considered the most appropriate method for testing a classifier on individual samples from multi-site datasets [41,42,43]. Second, we could not test the effect of psychosis-by-age interaction on the predict probability as longitudinal MRI data were not available. Longitudinally tracking neuroanatomical changes around the onset of psychosis would offer more detailed information to understand the progressive brain pathology. Third, substance use of cannabis or alcohol was not available for the current study which is reported associated with increased risk of developing depression in young adulthood [89]. Fourth, while we note that a classifier that can distinguish between CHR-PS+ and CHR-PS- status is clinically useful, we did not explicitly train our classifier to distinguish between CHR-PS+ and CHR-PS-. Previous work suggests that the magnitude of differences in MRI metrics between CHR-PS+ and CHR-PS- are small. Although there exist no well-validated methods to decide on the minimal sample size to create a reliable classifier, considering these subtle differences, the sample size of CHR-PS+ may likely be insufficient. Increased availability of CHR data may enable the development of such a classifier in the future.
In conclusion, we successfully trained a 2-class XGBoost classifier (HC versus CHR-PS+) and showed promising predictive performance on a multi-site dataset after considering age and sex differences. This classifier successfully identified 73% of CHR-PS- individuals as HC, and further 80% of CHR individuals who were not follow-up for the onset. These results suggest that when considering adolescent brain development, baseline MRI scans for CHR individuals may be helpful to identify their prognosis. Especially, the superior temporal, insula, superior frontal areas contributed most in differentiating CHR-PS+ from HCs. In light previous work reporting that alterations in these regions have implicated in psychosis onset, these areas could be informative in improving understanding of pathophysiology linked to psychosis onset. Future prospective studies are required about what and how the psychosis-related brain characteristics change according to the adolescent development, and whether the classifier could be helpful in the clinical settings.
Code availability
The Python code used to build the classifier is openly available on GitHub: https://github.com/yh-zhu/MolPsy_2024_ENIGMA.git.
References
Fusar‐Poli P, Correll CU, Arango C, Berk M, Patel V, Ioannidis JPA. Preventive psychiatry: a blueprint for improving the mental health of young people. World Psychiatry. 2021;20:200–21.
Miller TJ, McGlashan TH, Rosen JL, Somjee L, Markovich PJ, Stein K, et al. Prospective diagnosis of the initial prodrome for schizophrenia based on the structured interview for prodromal syndromes: preliminary evidence of interrater reliability and predictive validity. Am J Psychiatry. 2002;159:863–5.
Yung AR, Nelson B, Stanford C, Simmons MB, Cosgrave EM, Killackey E, et al. Validation of “prodromal” criteria to detect individuals at ultra high risk of psychosis: 2 year follow-up. Schizophr Res. 2008;105:10–17.
Seiler N, Nguyen T, Yung A, O’Donoghue B. Terminology and assessment tools of psychosis: a systematic narrative review. Psychiatry Clin Neurosci. 2020;74:226–46.
Catalan A, Salazar De Pablo G, Vaquerizo Serrano J, Mosillo P, Baldwin H, Fernández‐Rivas A, et al. Annual research review: prevention of psychosis in adolescents – systematic review and meta‐analysis of advances in detection, prognosis and intervention. J Child Psychol Psychiatry. 2021;62:657–73.
Salazar De Pablo G, Woods SW, Drymonitou G, De Diego H, Fusar-Poli P. Prevalence of individuals at clinical high-risk of psychosis in the general population and clinical samples: systematic review and meta-analysis. Brain Sci. 2021;11:1544.
Salazar De Pablo G, Radua J, Pereira J, Bonoldi I, Arienti V, Besana F, et al. Probability of transition to psychosis in individuals at clinical high risk: an updated meta-analysis. JAMA Psychiatry. 2021;78:970.
Salazar De Pablo G, Soardo L, Cabras A, Pereira J, Kaur S, Besana F, et al. Clinical outcomes in individuals at clinical high risk of psychosis who do not transition to psychosis: a meta-analysis. Epidemiol Psychiatr Sci. 2022;31:e9.
Tomyshev AS, Lebedeva IS, Akhadov TA, Omelchenko MA, Rumyantsev AO, Kaleda VG. Alterations in white matter microstructure and cortical thickness in individuals at ultra-high risk of psychosis: a multimodal tractography and surface-based morphometry study. Psychiatry Res Neuroimaging. 2019;289:26–36.
Kwak YB, Kim M, Cho KIK, Lee J, Lee TY, Kwon JS. Reduced cortical thickness in subjects at clinical high risk for psychosis and clinical attributes. Aust N. Z J Psychiatry. 2019;53:219–27.
Chung Y, Allswede D, Addington J, Bearden CE, Cadenhead K, Cornblatt B, et al. Cortical abnormalities in youth at clinical high-risk for psychosis: findings from the NAPLS2 cohort. Neuroimage Clin. 2019;23:101862.
Koutsouleris N, Meisenzahl EM, Davatzikos C, Bottlender R, Frodl T, Scheuerecker J, et al. Use of neuroanatomical pattern classification to identify subjects in at-risk mental states of psychosis and predict disease transition. Arch Gen Psychiatry. 2009;66:700.
Del Re EC, Stone WS, Bouix S, Seitz J, Zeng V, Guliano A, et al. Baseline cortical thickness reductions in clinical high risk for psychosis: brain regions associated with conversion to psychosis versus non-conversion as assessed at one-year follow-up in the shanghai-at-risk-for-psychosis (SHARP) study. Schizophr Bull. 2021;47:562–74.
Pantelis C, Velakoulis D, McGorry PD, Wood SJ, Suckling J, Phillips LJ, et al. Neuroanatomical abnormalities before and after onset of psychosis: a cross-sectional and longitudinal MRI comparison. Lancet. 2003;361:281–8.
Takahashi T, Wood SJ, Yung AR, Soulsby B, McGorry PD, Suzuki M, et al. Progressive gray matter reduction of the superior temporal gyrus during transition to psychosis. Arch Gen Psychiatry. 2009;66:366.
Takahashi T, Wood SJ, Yung AR, Phillips LJ, Soulsby B, McGorry PD, et al. Insular cortex gray matter changes in individuals at ultra-high-risk of developing psychosis. Schizophr Res. 2009;111:94–102.
Pantelis C, Yücel M, Bora E, Fornito A, Testa R, Brewer WJ, et al. Neurobiological markers of illness onset in psychosis and schizophrenia: the search for a moving target. Neuropsychol Rev. 2009;19:385–98.
Kasai K, Shenton ME, Salisbury DF, Hirayasu Y, Lee C-U, Ciszewski AA, et al. Progressive decrease of left superior temporal gyrus gray matter volume in patients with first-episode schizophrenia. Am J Psychiatry. 2003;160:156–64.
Fischl B, Sereno MI, Dale AM. Cortical surface-based analysis. II: inflation, flattening, and a surface-based coordinate system. Neuroimage. 1999;9:195–207.
Fischl B, Dale AM. Measuring the thickness of the human cerebral cortex from magnetic resonance images. Proc Natl Acad Sci USA. 2000;97:11050–5.
Fischl B, Salat DH, Busa E, Albert M, Dieterich M, Haselgrove C, et al. Whole brain segmentation. Neuron. 2002;33:341–55.
Grotzinger AD, Mallard TT, Liu Z, Seidlitz J, Ge T, Smoller JW. Multivariate genomic architecture of cortical thickness and surface area at multiple levels of analysis. Nat Commun. 2023;14:946.
Grasby KL, Jahanshad N, Painter JN, Colodro-Conde L, Bralten J, Hibar DP, et al. The genetic architecture of the human cerebral cortex. [published correction appears in Science. 2021 Oct 22;374:eabm7211]. Science. 2020;367:eaay6690.
Winkler AM, Kochunov P, Blangero J, Almasy L, Zilles K, Fox PT, et al. Cortical thickness or grey matter volume? The importance of selecting the phenotype for imaging genetics studies. Neuroimage. 2010;53:1135–46.
Panizzon MS, Fennema-Notestine C, Eyler LT, Jernigan TL, Prom-Wormley E, Neale M, et al. Distinct genetic influences on cortical surface area and cortical thickness. Cereb Cortex. 2009;19:2728–35.
Rakic P. Specification of cerebral cortical areas. Science. 1988;241:170–6.
ENIGMA Clinical High Risk for Psychosis Working Group, Jalbrzikowski M, Hayes RA, Wood SJ, Nordholm D, Zhou JH, et al. Association of structural magnetic resonance imaging measures with psychosis onset in individuals at clinical high risk for developing psychosis: an enigma working group mega-analysis. JAMA Psychiatry. 2021;78:753.
Ding Y, Ou Y, Pan P, Shan X, Chen J, Liu F, et al. Brain structural abnormalities as potential markers for detecting individuals with ultra-high risk for psychosis: a systematic review and meta-analysis. Schizophr Res. 2019;209:22–31.
Zhao Y, Zhang Q, Shah C, Li Q, Sweeney JA, Li F, et al. Cortical thickness abnormalities at different stages of the illness course in schizophrenia: a systematic review and meta-analysis. JAMA Psychiatry. 2022;79:560–70.
Collins MA, Ji JL, Chung Y, Lympus CA, Afriyie-Agyemang Y, Addington JM, et al. Accelerated cortical thinning precedes and predicts conversion to psychosis: the NAPLS3 longitudinal study of youth at clinical high-risk. Mol Psychiatry. 2023;28:1182–9.
Baldwin H, Radua J, Antoniades M, Haas SS, Frangou S, Agartz I, et al. Neuroanatomical heterogeneity and homogeneity in individuals at clinical high risk for psychosis. Transl Psychiatry. 2022;12:297.
Cannon TD, Chung Y, He G, Sun D, Jacobson A, van Erp TGM, et al. Progressive reduction in cortical thickness as psychosis develops: a multisite longitudinal neuroimaging study of youth at elevated clinical risk. Biol Psychiatry. 2015;77:147–57.
Koutsouleris N, Pantelis C, Velakoulis D, McGuire P, Dwyer DB, Urquijo-Castro M-F, et al. Exploring links between psychosis and frontotemporal dementia using multimodal machine learning: dementia praecox revisited. JAMA Psychiatry. 2022;79:907.
Dima D, Modabbernia A, Papachristou E, Doucet GE, Agartz I, Aghajani M, et al. Subcortical volumes across the lifespan: Data from 18,605 healthy individuals aged 3–90 years. Hum Brain Mapp. 2022;43:452–69.
Frangou S, Modabbernia A, Williams SCR, Papachristou E, Doucet GE, Agartz I, et al. Cortical thickness across the lifespan: Data from 17,075 healthy individuals aged 3–90 years. Hum Brain Mapp. 2022;43:431–51.
Bethlehem RAI, Seidlitz J, White SR, Vogel JW, Anderson KM, Adamson C, et al. Brain charts for the human lifespan. Nature. 2022;604:525–33.
Chung Y, Addington J, Bearden CE, Cadenhead K, Cornblatt B, Mathalon DH, et al. Adding a neuroanatomical biomarker to an individualized risk calculator for psychosis: a proof-of-concept study. Schizophr Res. 2019;208:41–3.
Chung Y, Addington J, Bearden CE, Cadenhead K, Cornblatt B, Mathalon DH, et al. Use of machine learning to determine deviance in neuroanatomical maturity associated with future psychosis in youths at clinically high risk. JAMA Psychiatry. 2018;75:960–8.
de Wit S, Wierenga LM, Oranje B, Ziermans TB, Schothorst PF, van Engeland H, et al. Brain development in adolescents at ultra-high risk for psychosis: longitudinal changes related to resilience. Neuroimage Clin. 2016;12:542–9.
van Erp TGM, Walton E, Hibar DP, Schmaal L, Jiang W, Glahn DC, et al. Cortical brain abnormalities in 4474 individuals with schizophrenia and 5098 control subjects via the enhancing neuro imaging genetics through meta analysis (ENIGMA) consortium. Biol Psychiatry. 2018;84:644–54.
ENIGMA Schizophrenia Working Group, van Erp TGM, Hibar DP, Rasmussen JM, Glahn DC, Pearlson GD, et al. Subcortical brain volume abnormalities in 2028 individuals with schizophrenia and 2540 healthy controls via the ENIGMA consortium. Mol Psychiatry. 2016;21:547–53.
for the ENIGMA Consortium, Thompson PM, Jahanshad N, Ching CRK, Salminen LE, Thomopoulos SI, et al. ENIGMA and global neuroscience: a decade of large-scale studies of the brain in health and disease across more than 40 countries. Transl Psychiatry. 2020;10:100.
Rimol LM, Nesvåg R, Hagler DJ, Bergmann Ø, Fennema-Notestine C, Hartberg CB, et al. Cortical volume, surface area, and thickness in schizophrenia and bipolar disorder. Biol Psychiatry. 2012;71:552–60.
Cheon E, Bearden CE, Sun D, Ching CRK, Andreassen OA, Schmaal L, et al. Cross disorder comparisons of brain structure in schizophrenia, bipolar disorder, major depressive disorder, and 22q11.2 deletion syndrome: a review of ENIGMA findings. Psychiatry Clin Neurosci. 2022;76:140–61.
Koike S, Uematsu A, Sasabayashi D, Maikusa N, Takahashi T, Ohi K, et al. Recent advances and future directions in brain mr imaging studies in schizophrenia: toward elucidating brain pathology and developing clinical tools. Magn Reson Med Sci. 2021;21:539–52.
Shahab S, Mulsant BH, Levesque ML, Calarco N, Nazeri A, Wheeler AL, et al. Brain structure, cognition, and brain age in schizophrenia, bipolar disorder, and healthy controls. Neuropsychopharmacology. 2019;44:898–906.
Koutsouleris N, Riecher-Rössler A, Meisenzahl EM, Smieskova R, Studerus E, Kambeitz-Ilankovic L, et al. Detecting the psychosis prodrome across high-risk populations using neuroanatomical biomarkers. Schizophr Bull. 2015;41:471–82.
Yassin W, Nakatani H, Zhu Y, Kojima M, Owada K, Kuwabara H, et al. Machine-learning classification using neuroimaging data in schizophrenia, autism, ultra-high risk and first-episode psychosis. Transl Psychiatry. 2020;10:278.
Zhu Y, Nakatani H, Yassin W, Maikusa N, Okada N, Kunimatsu A, et al. Application of a machine learning algorithm for structural brain images in chronic schizophrenia to earlier clinical stages of psychosis and autism spectrum disorder: a multiprotocol imaging dataset study. Schizophr Bull. 2022;48:563–74.
Rozycki M, Satterthwaite TD, Koutsouleris N, Erus G, Doshi J, Wolf DH, et al. Multisite machine learning analysis provides a robust structural imaging signature of schizophrenia detectable across diverse patient populations and within individuals. Schizophr Bull. 2018;44:1035–44.
Li A, Zalesky A, Yue W, Howes O, Yan H, Liu Y, et al. A neuroimaging biomarker for striatal dysfunction in schizophrenia. Nat Med. 2020;26:558–65.
Koutsouleris N, Worthington M, Dwyer DB, Kambeitz-Ilankovic L, Sanfelici R, Fusar-Poli P, et al. Toward generalizable and transdiagnostic tools for psychosis prediction: an independent validation and improvement of the NAPLS-2 risk calculator in the multisite PRONIA Cohort. Biol Psychiatry. 2021;90:632–42.
Zarogianni E, Storkey AJ, Borgwardt S, Smieskova R, Studerus E, Riecher-Rössler A, et al. Individualized prediction of psychosis in subjects with an at-risk mental state. Schizophr Res. 2019;214:18–23.
Yamashita A, Yahata N, Itahashi T, Lisi G, Yamada T, Ichikawa N, et al. Harmonization of resting-state functional MRI data across multiple imaging sites via the separation of site differences into sampling bias and measurement bias. PLOS Biol. 2019;17:e3000042.
Maikusa N, Zhu Y, Uematsu A, Yamashita A, Saotome K, Okada N, et al. Comparison of traveling‐subject and ComBat harmonization methods for assessing structural brain characteristics. Hum Brain Mapp. 2021;42:5278–87.
Bayer JMM, Thompson PM, Ching CRK, Liu M, Chen A, Panzenhagen AC, et al. Site affects how-to & when: an overview of retrospective techniques to accommodate site effects in multi-site neuroimaging analyses. Front Neurol. 2022;13:923988.
Fortin J-P, Cullen N, Sheline YI, Taylor WD, Aselcioglu I, Cook PA, et al. Harmonization of cortical thickness measurements across scanners and sites. NeuroImage. 2018;167:104–20.
Wood SN Generalized Additive Models: An Introduction with R, 2nd ed. Chapman and Hall/CRC; 2017.
Hastie T, Tibshirani R. Generalized additive models. Stat Sci. 1986;1:297–310.
Pomponio R, Erus G, Habes M, Doshi J, Srinivasan D, Mamourian E, et al. Harmonization of large MRI datasets for the analysis of brain imaging patterns throughout the lifespan. NeuroImage. 2020;208:116450.
Gurholt TP, Lonning V, Nerland S, Jørgensen KN, Haukvik UK, Alloza C, et al. Intracranial and subcortical volumes in adolescents with early‐onset psychosis: a multisite mega‐analysis from the ENIGMA consortium. Hum Brain Mapp. 2022;43:373–84.
Chen T, Guestrin C. XGBoost: a scalable tree boosting system. Proc. 22nd ACM SIGKDD Int. Conf. Knowl. Discov. Data Min. 785–94, (2016).
Oliver D, Arribas M, Radua J, Salazar De Pablo G, De Micheli A, Spada G, et al. Prognostic accuracy and clinical utility of psychometric instruments for individuals at clinical high-risk of psychosis: a systematic review and meta-analysis. Mol Psychiatry. 2022;27:3670–8.
ENIGMA Clinical High Risk for Psychosis Working Group, Allen P, Baldwin H, Bartholomeusz CF, Chee MW, Chen X, et al. Normative modeling of brain morphometry in clinical high risk for psychosis. JAMA Psychiatry. 2023. https://doi.org/10.1001/jamapsychiatry.2023.3850.
Yung AR, Yung AR, Yuen HP, Mcgorry PD, Phillips LJ, Kelly D, et al. Mapping the onset of psychosis: the comprehensive assessment of at-risk mental states. Aust N Z J Psychiatry. 2005;39:964–71.
Miller TJ, Woods SW, Corcoran CM, Davidson L. Symptom assessment in schizophrenic prodromal states. Psychiatr Q:15.
Miller TJ, McGlashan TH, Rosen JL, Cadenhead K, Ventura J, McFarlane W, et al. Prodromal assessment with the structured interview for prodromal syndromes and the scale of prodromal symptoms: predictive validity, interrater reliability, and training to reliability. Schizophr Bull. 2003;29:703–15.
World Medical Association. World Medical Association Declaration of Helsinki: Ethical Principles for Medical Research Involving Human Subjects. JAMA. 2013;310:2191.
Desikan RS, Segonne F, Fischl B, Quinn BT, Dickerson BC, Blacker D, et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage. 2006;31:968–80.
Ching CRK, Gutman BA, Sun D, Villalon Reina J, Ragothaman A, Isaev D, et al. Mapping subcortical brain alterations in 22q11.2 deletion syndrome: effects of deletion size and convergence with idiopathic neuropsychiatric illness. Am J Psychiatry. 2020;177:589–600.
Sun D, Ching CRK, Lin A, Forsyth JK, Kushan L, Vajdi A, et al. Large-scale mapping of cortical alterations in 22q11.2 deletion syndrome: Convergence with idiopathic psychosis and effects of deletion size. Mol Psychiatry. 2020;25:1822–34.
Schmaal L, Veltman DJ, van Erp TGM, Sämann PG, Frodl T, Jahanshad N. et al. Subcortical brain alterations in major depressive disorder: findings from the ENIGMA Major Depressive Disorder working group. Mol Psychiatry. 2016;21:806–12.
Schmaal L, Hibar DP, Sämann PG, Hall GB, Baune BT, Jahanshad N. et al. Cortical abnormalities in adults and adolescents with major depression based on brain scans from 20 cohorts worldwide in the ENIGMA Major Depressive Disorder Working Group. Mol Psychiatry. 2017;22:900–9.
Radua J, Vieta E, Shinohara R, Kochunov P, Quidé Y, Green MJ, et al. Increased power by harmonizing structural MRI site differences with the ComBat batch adjustment method in ENIGMA. NeuroImage. 2020;218:116956.
Pedregosa F, Varoquaux G, Gramfort A, Michel V, Thirion B, Grisel O, et al. Scikit-learn: machine Learning in Python. Mach Learn PYTHON:6.
Ojala M, Garriga GC permutation tests for studying classifier performance. Ninth IEEE Int. Conf. Data Min., Miami Beach, FL, USA: IEEE. 908–13; 2009.
Fusar-Poli P, Rutigliano G, Stahl D, Davies C, Bonoldi I, Reilly T, et al. Development and validation of a clinically based risk calculator for the transdiagnostic prediction of psychosis. JAMA Psychiatry. 2017;74:493.
Vickers AJ, Elkin EB. Decision curve analysis: a novel method for evaluating prediction models. Med Decis Mak. 2006;26:565–74.
Pfeiffer RM, Gail MH. Estimating the decision curve and its precision from three study designs. Biom J. 2020;62:764–76.
Vickers AJ, Van Calster B, Steyerberg EW. A simple, step-by-step guide to interpreting decision curve analysis. Diagn Progn Res. 2019;3:18.
Tseng HH, Bossong MG, Modinos G, Chen KM, McGuire P, Allen P. A systematic review of multisensory cognitive–affective integration in schizophrenia. Neurosci Biobehav Rev. 2015;55:444–52.
Dutt A, Tseng HH, Fonville L, Drakesmith M, Su L, Evans J, et al. Exploring neural dysfunction in ‘clinical high risk’ for psychosis: a quantitative review of fMRI studies. J Psychiatr Res. 2015;61:122–34.
Fusar-Poli P, Bonoldi I, Yung AR, Borgwardt S, Kempton MJ, Valmaggia L, et al. Predicting psychosis: meta-analysis of transition outcomes in individuals at high clinical risk. Arch Gen Psychiatry. 2012;69:10.
Lo A, Chernoff H, Zheng T, Lo S-H. Why significant variables aren’t automatically good predictors. Proc Natl Acad Sci. 2015;112:13892–7.
Fusar-Poli P, Salazar De Pablo G, Correll CU, Meyer-Lindenberg A, Millan MJ, Borgwardt S, et al. Prevention of psychosis: advances in detection, prognosis, and intervention. JAMA Psychiatry. 2020;77:755.
Nenadic I, Dietzek M, Schönfeld N, Lorenz C, Gussew A, Reichenbach JR, et al. Brain structure in people at ultra-high risk of psychosis, patients with first-episode schizophrenia, and healthy controls: a VBM study. Schizophr Res. 2015;161:169–76.
Fusar-Poli P, Cappucciati M, Borgwardt S, Woods SW, Addington J, Nelson B, et al. Heterogeneity of psychosis risk within individuals at clinical high risk: a meta-analytical stratification. JAMA Psychiatry. 2016;73:113.
Solanes A, Gosling CJ, Fortea L, Ortuño M, Lopez-Soley E, Llufriu S, et al. Removing the effects of the site in brain imaging machine-learning – Measurement and extendable benchmark. NeuroImage. 2023;265:119800.
Gobbi G, Atkin T, Zytynski T, Wang S, Askari S, Boruff J, et al. Association of cannabis use in adolescence and risk of depression, anxiety, and suicidality in young adulthood: a systematic review and meta-analysis. JAMA Psychiatry. 2019;76:426.
Acknowledgements
This research was supported in part by AMED (Grant Number JP18dm0307001, JP18dm0307004, and JP19dm0207069), JST Moonshot R&D (JPMJMS2021), JSPS KAKENHI (JP23H03877 and JP21H02851), Takeda Science Foundation and SENSHIN Medical Research Foundation. This study was also supported by the International Research Center for Neurointelligence (WPI-IRCN), the University of Tokyo.
Funding
Open Access funding provided by The University of Tokyo.
Author information
Authors and Affiliations
Consortia
Contributions
Author contributions: YZ and SK had full access to all of the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis. Study concept and design: YZ, NM and SK. The whole ENIGMA consortium collected the data. Acquisition, analysis, interpretation of data, or funding (in alphabetical order): IA, OAA, PB, IB, XC, SC, CMC, BHE, AF, RRGG, BYG, LBG, SSH, HKH, RAH, YH, KH, KK, NK, MK, TDK, JSK, SML, IL, JL, RLL, DHM, PMcG, RM, MM, PM, TN, DN, MAO, JMR, JIR, WR, DFS, DS, LS, GS, TT, CKT, JT, AT, AST, PJU, TGV, TAMJvanA, JAW, LTW, JHZ and SK. Statistical analysis: YZ and SK. Drafting of the manuscript: YZ and SK. Administrative, technical or material support: JR, PGS, PFP and PMT. Critical revision of the manuscript for important intellectual content: DH and MJ. All authors approved the contents of the paper.
Corresponding author
Ethics declarations
Competing interests
Author OAO conducted a consultant to cortechs.ai, received speaker’s honorarium from Lundbeck, Janssen, Sunovion. Author BYG has been the leader of a Lundbeck Foundation Centre of Excellence for Clinical Intervention and Neuropsychiatric Schizophrenia Research (CINS) (January 2009 – December 2021), which was partially financed by an independent grant from the Lundbeck Foundation based on international review and partially financed by the Mental Health Services in the Capital Region of Denmark, the University of Copenhagen, and other foundations. All grants are the property of the Mental Health Services in the Capital Region of Denmark and administrated by them. Other authors have no conflict of interest to declare that are relevant to the content of this article.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhu, Y., Maikusa, N., Radua, J. et al. Using brain structural neuroimaging measures to predict psychosis onset for individuals at clinical high-risk. Mol Psychiatry 29, 1465–1477 (2024). https://doi.org/10.1038/s41380-024-02426-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41380-024-02426-7
- Springer Nature Limited