Abstract
Large-scale fecal shotgun metagenomic sequencing revealed the high abundance of Parvimonas micra in colorectal cancer (CRC) patients. We investigated the role and clinical significance of P. micra in colorectal tumorigenesis. The abundance of P. micra was examined in 309 fecal samples and 165 colon biopsy tissues of CRC patients and healthy subjects. P. micra was significantly enriched in fecal samples from 128 CRC patients compared to 181 healthy subjects (P < 0.0001); and in colon tissue biopsies from 52 CRC patients compared to 61 healthy subjects (P < 0.0001). Multivariate analysis showed that P. micra is an independent risk factor of poor survival in CRC patients (Hazard Ratio: 1.93). P. micra strain was isolated from feces of a CRC patient. Apcmin/+ mice gavaged with P. micra showed significantly higher tumor burden and tumor load (both P < 0.01). Consistently, gavage of P. micra significantly promoted colonocyte proliferation in conventional mice, which was further confirmed by germ-free mice. P. micra colonization up-regulated genes involved in cell proliferation, stemness, angiogenesis and invasiveness/metastasis; and enhanced Th17 cells infiltration and expression of Th17 cells-secreted cytokines (Il-17, Il-22, and Il-23) in the colon of Apcmin/+, conventional and germ-free mice. P. micra-conditioned medium significantly promoted the differentiation of CD4+ T cells to Th17 cells (IL-17+CD4+ phenotype) and enhanced the oncogenic Wnt signaling pathway. In conclusion, P. micra promoted colorectal tumorigenesis in mice by inducing colonocyte proliferation and altering Th17 immune response. P. micra may act as a prognostic biomarker for poor survival of CRC patients.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is one of the leading causes of cancer-related deaths worldwide [1]. The progression of CRC involves a variety of factors including genetic mutations, epigenetic changes, and environmental alteration [2]. Accumulated evidence from us and others has suggested that the gut microbiota contributes to CRC development [3,4,5]. Gut commensal microbiota plays multiple roles in maintaining host health and inducing diseases [6]. A balanced microbiota could produce essential nutrients, prompt efficient host nutrient absorption, aid development of a mature and competent host immune system, and prevent pathogen colonization [7]. An unbalanced gut microbiota, commonly termed as dysbiosis, could result in inflammation, intestinal barrier failure, mucosal tissue damage, oncogene upregulation, and altered microenvironment, which all favor the development of CRC [8, 9]. Of note, the enrichment of opportunistic pathogenic bacteria in the gut microbiota of CRC patients has been constantly reported in previous studies which identified several individual species that contribute to CRC development. For instance, Fusobacterium nucleatum has been widely studied as a CRC-promoting microbe in both human and animal models by modulating tumor-immune microenvironment and autophagy [3, 10]. Peptostreptococcus anaerobius promotes colorectal carcinogenesis and modulates the tumor immune microenvironment by activating PI3K/Akt signaling via its PCWBR2 adhesin [5, 11]. Identifying microbial pathogens that are important contributors of colorectal tumorigenesis is imperative to the manipulation of gut microbiota for CRC prevention and treatment.
By conducting a large-scale meta-analysis of fecal shotgun metagenomic sequencing from four cohorts of different ethnicities (Chinese, French & German, Austrian and American), we identified the enrichment of Parvimonas micra in CRC patients compared to healthy controls [12, 13]. P. micra, formerly known as Peptostreptococcus micros or Micromonas micros, is a gram-positive, anaerobic and opportunistic pathogen commonly found in the human oral cavity [14]. P. micra is frequently isolated from a wide range of human infections, including orofacial odontogenic infections, periodontitis lesions, endodontic abscesses, and purulent pleurisy [15, 16]. Meanwhile, increasing evidence has reported the correlation between oral microbes and CRC [17], of which P. micra was found to be associated with the consensus molecular subtype 1 of CRC [18]. We therefore speculated that P. micra may be involved in the development of CRC. In this study, we investigated the functional importance of P. micra in multiple mouse models of CRC as well as its clinical implication. Our results altogether confirmed the CRC-promoting role of P. micra and its potential to be a prognostic marker for CRC patients.
Results
P. micra is significantly enriched in fecal samples and tissue biopsies of CRC patients
From our in-house fecal metagenomic cohort (118 CRC patients and 128 healthy subjects), we demonstrated that P. micra was significantly enriched in fecal samples of CRC patients compared to healthy subjects (P < 0.001) (Fig. 1A). The enrichment of P. micra in patients with CRC was validated in two independent metagenomic cohorts (French cohort: 89 CRC patients and 66 healthy subjects, P < 0.01; Austrian cohort: 46 CRC patients and 63 healthy subjects, P < 0.01) (Fig. 1B). To further verify the findings from metagenomic sequencing data, the abundance of P. micra was detected by real-time quantitative polymerase chain reaction (PCR) in additional fecal samples (CRC, n = 128; healthy subjects, n = 181). Our result revealed that P. micra was significantly enriched in CRC patients compared to healthy subjects (P < 0.0001) (Fig. 1C).
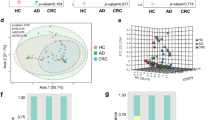
A The level of P. micra in faecal samples of CRC patients (n = 118) and healthy subjects (n = 128) in Chinese by metagenomic sequencing; B P. micra level in stool samples from two public cohorts of French (CRC: n = 89 and healthy subjects: n = 66) and Austrian (CRC: n = 46; healthy subjects: n = 63) by metagenomic sequencing; C Real time qPCR analysis of faecal P. micra level in CRC patients (n = 128) and healthy subjects (n = 181) of Chinese cohort; D Real time qPCR analysis of paired CRC tumor and tumor-adjacent mucosal samples from CRC patients (n = 52) and mucosal samples from healthy subjects (n = 61); E Kaplan–Meier curves showed that high stool P. micra abundance was significantly associated with shortened survival in patients with CRC.
Gut mucosa microbiota is associated with colorectal carcinogenesis [19]. We therefore examined the abundance of P. micra in 61 mucosal samples of normal colon and 52 paired samples of colorectal tumor biopsies and adjacent normal mucosa. P. micra was significantly enriched in both tumor (P < 0.0001) and adjacent normal mucosa (P < 0.05) of CRC patients compared to healthy subjects, of which its abundance was the greatest in tumor tissues (Fig. 1D). Collectively, we confirmed the enrichment of P. micra in both fecal and mucosal samples of patients with CRC.
P. micra is an independent predictor of poor outcome in CRC patients
We then analyzed the association of fecal P. micra abundance with clinicopathological features. There was no significant correlation between P. micra abundance and age, gender, or tumor-node-metastasis (TNM) stage (Table S1). Whereas P. micra was associated with increased risk of cancer-related death by univariate Cox regression analysis (Hazard Ratio (HR): 1.99, 95% confidence interval (CI): 1.10 to 3.60, P = 0.022; Table 1). After adjustment of confounding factors including age, gender and TNM stage, P. micra was identified as an independent risk factor of shortened 5-year survival in CRC by multivariate Cox regression analysis (HR: 1.93, 95% CI: 1.04 to 3.60, P = 0.037; Table 1). As shown in the Kaplan–Meier survival curves, patients with high abundance of P. micra had significantly shorter 5-year survival than those with low abundance of P. micra (P = 0.02, Fig. 1E). These results suggested that P. micra could be a predictor of poor prognosis in patients with CRC.
P. micra promotes colorectal tumorigenesis in transgenic Apc min/+ mice
A strain of P. micra was successfully isolated from the fecal sample of a CRC patient (Fig. 2A and Fig. S1A). Chromatogram analysis showed that our isolated strain had nearly identical sequence to the 16 S ribosomal RNA (rRNA) gene sequence of P. micra in the NCBI RefSeq database (Fig. S1B). Growth dynamics our P. micra strain were shown in Fig. S2A. Given that P. micra was enriched in the colon mucosa of CRC patients (Fig. 1D), we co-cultured P. micra with HT-29, a human colon adenocarcinoma cell line with epithelial morphology. P. micra could adherent to HT-29 cells under anaerobic incubation compared to broth control (P < 0.0001), indicating the capability of P. micra to directly interacting with the gut epithelial cell (Fig. S2B).
A Isolation of P. micra in faecal sample of a CRC patient; B Schematic diagram showing the experimental design and timeline of Apcmin/+ mice model (broth control group n = 8, E. coli MG1655 group n = 10 and P. micra group n = 10); C Faecal quantitation of total bacteria and P. micra in Apcmin/+ mouse model after antibiotic treatment; D Relative abundance of P. micra in mice under different treatments; E Representative colonic morphologies in mice gavaged with P. micra, E. coli or broth control; F The colonic tumor numbers and loads of Apcmin/+ mice under different treatments; G P. micra promoted the formation of colonic dysplasia. Representative histologic images of colon tissues of mice by H&E staining and statistical analysis of colon samples according to the histologic score. The low and high resolution histological images of the adenocarcinoma in P. micra-treated colon tissues were shown.
To evaluate the role of P. micra in colorectal tumorigenesis, we used Apcmin/+ mice which is a commonly used transgenic mouse model of spontaneous CRC (Fig. 2B). Before oral gavage of P. micra, the resident microbiota was depleted using a cocktail of broad-spectrum antibiotics (ampicillin 0.2 g/L, vancomycin 0.1 g/L, neomycin 0.2 g/L, and metronidazole 0.2 g/L) for 2 weeks (Fig. 2B). Microbiota depletion was confirmed by quantitative PCR which showed a 10-fold reduction of total bacterial DNA in mice feces (Fig. 2C). Apcmin/+ mice were then orally gavaged with P. micra, a non-pathogenic E. coli strain MG1655, or broth control 3 times per week for 8 weeks (Fig. 2B). The fecal abundance of P. micra was increased after P. micra inoculation (Fig. 2D). Apcmin/+ mice were euthanized after 10 weeks and the colon of each mouse was examined macroscopically and histologically. We observed significantly increased tumor number and tumor load in mice inoculated with P. micra, compared to mice with E. coli and broth control (both P < 0.01) (Fig. 2E, F). P. micra-gavaged Apcmin/+ mice also showed significantly higher incidence of high-grade dysplasia (16.7%) and low-grade dysplasia (50.0%), compared to mice gavaged with E. coli or broth control (Fig. 2G). Collectively, these results indicated that P. micra plays a cancer-promoting role by accelerating colorectal tumorigenesis in Apcmin/+ mice.
P. micra promotes colonocyte proliferation in conventional C57BL/6 mice
To ascertain the role of P. micra in natural non-disease condition, we gavaged P. micra or broth control to conventional C57BL/6 mice without microbiota depletion, and mice were harvested after 36 weeks of gavage (Fig. 3A). Although no visible histological differences could be observed in mice colon (Fig. S3), increased proliferation of colon epithelial cells was identified in P. micra-gavaged mice compared to controls as indicated by a higher proportion of Ki-67 positive cells (P. micra = 17.0%, control = 12.5%; P < 0.05) (Fig. 3B) and proliferating cell nuclear antigen (PCNA) positive cells (P. micra = 42.7%, control = 30.1%; P < 0.05) (Fig. 3C).
A P. micra or broth control were gavaged to C57BL/6 conventional mice and harvested at 9 months; B Immunohistochemistry showing Ki-67-positive cells in the colon of conventional mice gavaged with P. micra or broth control; C Immunohistochemistry showing PCNA-positive cells in the colon of conventional mice gavaged with P. micra or broth control; D P. micra or broth control were gavaged to MNU-treated C57BL/6 conventional mice and harvested at 54 weeks; E Immunohistochemistry showing Ki-67-positive cells in the colon of MNU-treated conventional mice gavaged with P. micra or broth control.
We validated our in vivo results by establishing a carcinogen N-methyl-N-nitrosourea (MNU)-induced CRC mouse model (Fig. 3D). Consistently, the number of Ki-67 positive cells was significantly increased in P. micra-gavaged MNU mice (P. micra = 34.2%, control = 27.1%; P < 0.05) (Fig. 3E). Taken together, these results illustrated that P. micra promotes cell proliferation of colon epithelial cells.
P. micra promotes colonocyte proliferation in germ-free mice
Germ-free mice were then used to examine the pure role of P. micra in the initiation and development of CRC. P. micra or broth control was given to germ-free mice, and mice were harvested at different time points of 20 weeks, 32 weeks, or 36 weeks (Fig. 4A). To examine the spatial distribution and intestinal localization of P. micra, we performed fluorescence in situ hybridization (FISH) using P. micra oligonucleotide probes on the colon tissues of germ-free mice. The germ-free status was confirmed by microbial FISH staining of the colon tissues using EUB338 universal bacterial probe (Fig. S4A), while the mono-colonization status of P. micra was accessed by co-staining of EUB338 universal bacterial probe and P. micra-specific probe. As shown in Fig. 4B, the intestinal colonization of P. micra was observed in germ-free mice gavaged with P. micra but not in the control mice. Whereas our results of FISH testified that germ-free mice were mono-colonized by a single bacterium (which is P. micra) without any other environmental microbes following P. micra gavage (Fig. S4B). Consistent to the findings from conventional mice, P. micra significantly induced colonocyte proliferation in the colon of P. micra-gavaged germ-free mice at 20 weeks (P < 0.05), 32 weeks (P < 0.05), and 36 weeks (P < 0.001), as compared to control mice (Fig. 4C). The ability of P. micra in promoting cell proliferation in the colon tissues of P. micra-gavaged germ-free mice was further confirmed by increased PCNA protein expression as compared with control mice (Fig. 4D). The findings from germ-free mice collectively suggested that P. micra could initiate CRC progression by inducing colonocyte proliferation.
A Design of P. micra gavage experiment in germ-free mice; B Representative FISH images of colon tissue sections from BHI or P. micra-infected mice 36 weeks post-infection (Blue: nuclear; Green: P. micra probe); C Immunohistochemistry showing Ki-67-positive cells in the colon of germ-free mice at 20 weeks, 32 weeks, and 36 weeks after P. micra gavage. The proportion of Ki-67-positive cells was calculated in five fields of high-power microscopic field under microscopes. D Immunohistochemistry staining of PCNA protein in the colon of germ-free mice at 36 weeks after bacteria gavage; E MTT assay and F colony formation of HT29 cell line treated with broth control, E. coli conditioned medium (CM) and P. micra CM; G Western blot analysis of activated β-catenin and Cyclin D1 and real time PCR analysis of c-myc mRNA in HT29 cells treated with broth control or P. micra CM.
P. micra induces cell proliferation in vitro
To investigate its pro-tumorigenic functions, the conditioned medium of P. micra was treated with HT-29 CRC cell line. P. micra-conditioned medium significantly promoted proliferation of HT-29 cells compared with E. coli-conditioned medium and broth control, as evidenced by 3-(4,5-Dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT) viability assay (Fig. 4E) and colony formation assay (Fig. 4F).
The oncogenic Wnt signaling pathway plays an important role in CRC. We therefore evaluated whether Wnt signaling pathway was involved in the development of P. micra-induced CRC. The treatment of P. micra-conditioned medium could significantly induce the protein levels of activated β-catenin and Cyclin D1 as well as the mRNA level of c-myc in HT-29 cells (Fig. 4G). These results hence revealed that P. micra could activate the Wnt/β-catenin signaling pathway in epithelial cells to contribute CRC progression.
Altered expression of cell proliferation-related genes in P. micra-induced tumorigenesis
To uncover the mechanism underlying the pro-tumorigenic role of P. micra, we performed mouse Cancer Gene expression PCR Array on the colon tissues of germ-free mice (Fig. 5A). Genes involved in cell proliferation (Tbx2, Mki67, Mcm2, Cdc20), invasion and metastasis (Cdh2, Foxc2, Snai1), stemness (Sirt1, Bmi1), and angiogenesis (Pgf, Tek, Angpt1, Fit1) were observed to have more than 2-fold increase in expression in mice gavaged with P. micra, compared to the control mice (Fig. 5A, B). On the other hand, genes related to apoptotic pathways (FasL, Casp7, Map2k3) and DNA damage and repair (Gadd45g) were downregulated in P. micra-gavaged mice (Fig. 5A, B).
A Significant up-regulation of 74 transcripts and down-regulation of 10 transcripts by the Mouse Cancer Pathway Finder PCR Array in colon tissues of germ-free mice inoculated with P. micra for 32 weeks; B A systematic diagram showing major oncogenic pathways implicated by differentially expressed genes identified by the PCR array.
P. micra induces Th17 immune cells infiltration
The dysregulated immunity and chronic inflammation are known to be crucially involved in the development of gut microbiota-modulated CRC [6]. To explore immune response modulated by P. micra, we performed mouse Inflammatory Response and Autoimmunity PCR Array on colon tissues of P. micra-gavaged germ-free mice. Significant upregulation of 48 genes and downregulation of 6 genes were observed in germ-free mice after gavage with P. micra for 32 weeks, compared to the control mice (Fig. 6A). Differentially expressed genes included interleukin (Il)-17a, Il-22, and Il-23a, which encode 3 cytokines secreted by T helper (Th)-17 cells. The upregulated genes were involved in chemotaxis of various immune cells including neutrophil chemotaxis (Cxcl1, Cxcl2, Cxcl5, Cxcl9, Cxcr2, Cxcr4, and Ccl20), T-lymphocyte chemotaxis (Ccr4, Ccl17, Ccl19, Ccl22, Ccl24, Cxcl9, Cxcl10, and Cxcl11), and monocyte chemotaxis (Ccl1, Ccl2, Ccl3, Ccl4, Ccl5, Ccl7, and Ccl8) (Fig. 6B). Quantitative PCR confirmed the upregulated expressions of major pro-inflammatory cytokines including Il-17a, Tnf-α, Il-6 and Cxcr1 in the colon tissues of P. micra-gavaged mice (Fig. 6C), hence indicating that P. micra may contribute to dysregulated immunity by increasing secretion of inflammatory cytokines.
A Significant up-regulation of 48 transcripts and down-regulation of 10 transcripts by the Mouse Inflammatory Response and Autoimmunity PCR Array in the colon tissues of germ-free mice inoculated with P. micra for 32 weeks; B A systematic diagram showing major inflammatory pathways implicated by up-regulated genes identified by the PCR array; C qPCR validation was performed to confirm changes in expression of genes including Tnf, Il17a, Il16, and Cxcl1 in germ-free mice gavaged with P. micra.
Intestinal Th17 cells are critical for maintaining tissue homeostasis [20]. As Th17-related cytokines including Il-17a, Il-22, and Il-23a were found to be upregulated (Fig. 6), we characterized Th17 immune cells in the colonic lamina of P. micra-gavaged germ-free mice. The number of colonic Th17 (CD4+Il-17+) immune cells was significantly higher in P. micra-gavaged germ-free mice (P < 0.05) compared to the corresponding control mice (Fig. 7A). The number of colonic Th17 cells was also significantly increased in P. micra-gavaged conventional mice (P < 0.05) (Fig. 7B and Fig. S5). Consistently, immunohistochemistry confirmed the enhanced colonic infiltration of Th17 cells in P. micra-gavaged germ-free mice (P < 0.05) (Fig. 7C).
A Flow cytometric analysis of Th17 cells in the colon of germ-free mice gavaged with P. micra or broth control; B Flow cytometric analysis of Th17 cells in the colon of C57BL/6 conventional mice gavaged with P. micra or broth control; C Immunohistochemistry staining of IL-17 in the colon of germ-free mice gavaged with P. micra or broth control; D The effect of P. micra-conditioned medium (CM) on Th17 cell differentiation compared to broth control or E. coli CM.
To investigate if P. micra directly promotes the expansion of Th17 cells, we isolated CD4+ T cells from mouse spleen and treated them with P. micra-conditioned medium. Il-17 expression was evaluated by flow cytometry 5 days after the treatment. P. micra-conditioned medium significantly promoted cell differentiation to Il-17+CD4+ phenotype compared to broth control (P < 0.001) or E. coli-conditioned medium (P < 0.0001) (Fig. 7D). Taken together, these results suggested that the pro-tumorigenic effect of P. micra is associated with increased Th17 cell infiltration and differentiation in the colon, thereby promoting the establishment of a pro-inflammatory microenvironment which favors colorectal tumorigenesis.
Discussion
We and others previously reported the enrichment of P. micra in fecal samples of CRC patients by shotgun metagenomic sequencing [13, 21]. In this study, increased abundance of P. micra in CRC patients were validated by multiple cohorts of fecal metagenomes and our in-house cohort of fecal samples. We confirmed that P. micra was significantly enriched in feces and tumor tissues of CRC patients compared with healthy subjects [22, 23]. Moreover, we identified that the fecal abundance of P. micra is an independent risk factor of poor survival in patients with CRC by multivariate Cox regression analysis (HR 1.93). In particular, the disease-free survival with high fecal abundance of P. micra was significantly shorter than those with low P. micra abundance. An accurate evaluation of prognosis is essential for clinicians to decide appropriate treatment for achieving maximized efficacy. Our results suggested that P. micra could serve as a new prognostic biomarker for CRC patients.
The role of P. micra in promoting colorectal tumorigenesis was elucidated in multiple mouse models including microbiota-depleted Apcmin/+, conventional C57BL/6, and germ-free mice. We found that P. micra potentiates CRC progression as indicated by increased colon tumor number, tumor load, and incidence of histological dysplasia in P. micra-gavaged Apcmin/+ mice. We further tested whether P. micra could contribute to CRC initiation by using conventional C57BL/6 mice without any carcinogen treatment. After gavage for 9 months, mice with P. micra inoculation had significant increased proliferation in colon epithelial cells which is an essential event for the development of precancerous colorectal lesion [24]. Such increase in epithelial cell proliferation was also observed in germ-free mice receiving a single gavage of P. micra, indicating that the pro-tumorigenic role of P. micra is independent from other gut microbes. In keeping with these findings, the expressions of genes related to cell proliferation, angiogenesis and invasion/metastasis were significantly upregulated in P. micra-gavaged mice compared to the control mice. These results collectively demonstrated that P. micra could accelerate colorectal tumorigenesis by promoting colonocyte proliferation.
In general, microbial communities in the oral cavity and gut are well-segregated due to the physical distance between these two regions, while the gastric acidity also limits the growth and further migration of translocated oral commensal microbes [25]. However, the oral microbiota could translocate to the intestinal mucosa when the oral-gut barrier is in dysfunction [26]. A previous study reported that the oral microbiota could overcome the physical barrier to invade and colonize in the gut, eventually reshaping the gut microbial community in germ-free mice [27]. In humans, many studies have revealed the presence and enrichment of oral commensals in patients with CRC, particularly Fusobacterium nucleatum which can promote colorectal tumorigenesis by generating a pro-inflammatory tumor immune microenvironment [10]. Given that physical barriers in the body are mostly likely weakened and impaired under pathological conditions, opportunistic pathogen including P. micra derived from oral cavity may move into the gut and potentially contribute to disease progression.
CRC-associated bacteria evoke colorectal tumorigenesis via various means. F. nucleatum modulates host immunity and tumor microenvironment by inducing mucin secretion and inflammatory cytokine TNF-α expression, while P. anaerobius promotes intracellular cholesterol biosynthesis to induce colon cell proliferation [5, 10]. Given the diverse mechanisms through which bacteria could promote tumorigenesis, we performed functional investigation to gain insights into the mechanisms employed by P. micra in CRC development. Our array analysis revealed that the pro-tumorigenic effect of P. micra is associated with altered immune responses and enhanced secretion of inflammatory cytokines in the gut. In particular, Th17-mediated cytokines, including Il17a, Il22, and Il23a, were significantly upregulated by P. micra. Il-23 and Il-17 signalling were found to be activated by microbial products and correlated with tumor growth [28]. The increased infiltration of Th17 cells in the colon tissues of P. micra-gavaged mice from multiple models was confirmed by flow cytometry and immunohistochemistry. In supporting our findings, previous studies have reported the roles of Th17-mediated inflammation in promoting tumor growth and progression in colorectal tumorigenesis [28, 29]. Taken together, our results suggested that P. micra promotes CRC development via enhancing Th17-mediated immune response.
In conclusion, we identified that P. micra is a novel pathogenic microbe to promote CRC initiation and development. P. micra drives colorectal tumorigenesis through enhancing Th17 immune responses and oncogenic factors in the colon. For clinical implication, the fecal abundance of P. micra may act as a prognostic marker of poor survival for CRC patients.
Materials and methods
Patient recruitment and samples collection
Fecal samples (128 CRC patients and 181 healthy subjects) were retrieved from the research stool bank, collected from individuals undergoing colonoscopy at Shaw Endoscopy Centre, Prince of Wales Hospital, The Chinese University of Hong Kong. The inclusion and exclusion criteria were described previously [30]. Mucosal biopsies (52 paired CRC tumor and tumor adjacent mucosal samples and 61 normal tissues) were obtained from individuals who had undergone standardized colonoscopy examinations previously [13, 19]. All samples were stored at −80 °C immediately after collection until further analysis. Metagenomic sequencing data of fecal samples from our in-house cohort (Chinese: 118 CRC patients and 128 healthy subjects) and two independent European cohorts (French: 89 CRC patients and 66 healthy subjects; Austrian: 46 CRC patients and 63 healthy subjects) were retrieved and analyzed. Written informed consent was obtained from all subjects and ethics was approved by The Joint Chinese University of Hong Kong - New Territories East Cluster Clinical Research Ethics Committee.
Extraction of bacterial DNA from feces and tissues and P. micra quantification by quantitative polymerase chain reaction
200 mg of fecal samples were thawed on ice, and fecal DNA was extracted using ZR Faecal DNA MiniPrep Kit (Zymo Research, CA). The DNA quantity was determined using NanoDrop 2000c Spectrophotometer (Thermo Fisher Scientific, Waltham, MA). To extract bacterial DNA from human mucosal biopsies, glass beads (<100 µm, Sigma, St. Louis, MO) and QIAamp DNA Mini Kit (QIAGEN, Valencia, CAA) were used. 0.5 ng DNA was used for real time PCR analysis using Universal SYBR Greem Master (Roche, Risch-Rotkreuz, Switerland). The primers were listed in Table S2.
Isolation of P. micra from faecal samples of CRC patients
Fecal samples from CRC patients with high relative abundance of P. micra were used for bacterial isolation. Samples were spread on horse blood agar plates and incubated in anaerobic chamber. The identity of the candidate colony (P. micra strain 512) was determined by gram staining and amplification of 16 S rRNA gene using universal primers targeting hypervariable regions V1-V4 and V6 of the 16 S rRNA gene, with further confirmation by P. micra-specific primers and Sanger sequencing. The bacteria were cultured in anaerobe basal broth (CM0957; Thermo Fisher Scientific) in anaerobic jar (Hardy Diagnostics, Santa Maria, CA). The anaerobic condition was created by the application of Anaerogen (AN0035; Thermo Fisher Scientific). Another strain (P. micra Smith) was bought from American Type Culture Collection (33270,ATCC, Manassas, VA).
Animal experiments
Resident microbiota of male Apcmin/+ mice (6 weeks old, n = 8–10/group) were depleted using a cocktail of broad-spectrum antibiotics (ampicillin 0.2 g/L, vancomycin 0.1 g/L, neomycin 0.2 g/L, and metronidazole 0.2 g/L) in drinking water for 2 weeks, before oral gavage with 1 × 108 colony forming unit (CFU) of P. micra, E. coli MG1655 or broth control every 3 days for 8 weeks. Randomization was used to allocate the mice to different groups. E. coli MG1655 is a non-pathogenic human commensal intestinal bacterium which could not induce dysplasia [5, 31]. It was cultured in Brain Heart Infusion broth under aerobic condition. Mice were euthanized after 10 weeks, and their colon tissues were collected and examined.
In a separated model, conventional C57/BL6 male mice (7, 8 weeks old, n = 6–10/group) were gavaged with 1 × 108 CFU of P. micra or broth control every 3 days for up to 9 months. Another group of mice (7, 8 weeks old) were administrated with 240 ppm MNU in the drinking water biweekly for 10 weeks to induce colon tumorigenesis. Mice were then randomly gavaged with 1 × 108 CFU of P. micra or broth control and euthanized at week 54.
Germ-free male mice (7, 8 weeks old) were randomly gavaged with 1 × 108 CFU of P. micra or broth control twice per week. Mice were euthanized at 20, 32, and 36 weeks respectively (n = 5–7/group). All animal experiments were performed in accordance with guidelines approved by the Animal Experimentation Ethics Committee of the Chinese University of Hong Kong.
Histopathology
Mouse colon specimens were formalin-fixed and paraffin-embedded for histologic examination. Sections of 4 μm were stained with hematoxylin and eosin (H&E) for histologic diagnosis by an experienced pathologist (Dr. Alvin Ho-Kwan Cheung) who was blind to the group information. Dysplasia was defined according to the latest World Health Organization’s Classification of Tumors of the Digestive System.
Immunohistochemistry staining
Paraffin-embedded tissues were used for analyzing the expressions of Ki-67 (#9449 S, Cell Signaling Technology, Danvers, MA), PCNA (#2586, Cell Signaling Technology) and IL-17 (ab79056, Cell Signaling Technology) by immunohistochemistry. Slides were incubated with primary antibody (1:100) at 4 °C overnight. Signals were developed with IHC HRP/DAB kit (Millipore, Burlington, MA). The proportion of Ki-67, PCNA, or IL-17-positive cells was determined by counting immunostaining-positive cells, as a percentage to the total number of nuclei in the field. At least 1000 cells were counted in 5 random microscopic fields.
Fluorescence in situ hybridization
FISH-labeled P. micra Alexa Fluor 488-conjugated specific probe (GCCGCCGATCTAACCGCA) (Guangzhou exon biotechnology Co Ltd, Guangzhou, China) was used to detect P. micra colonization in paraffin-embedded sections. A Cy3-conjugated EUB338 universal bacterial probe (GCTGCCTCCCGTAGGAGT), as the positive control, was labelled with Spectrum-Red (Guangzhou exon biotechnology Co Ltd). 5 μm paraffin-embedded sections were hybridized in the hybridization buffer (1:50 in 25% hybridization buffer for P. micra; 1:50 in 45% hybridization buffer for EUB338 universal bacterial probe). After incubation overnight in a dark humid chamber at 40 °C, each slide was rinsed with sterile wash buffer and mounted with ProLong Gold Antifade Mountant with DAPI (Thermo Fisher Scientific).
Cell culture
HT-29 colon cancer cell line was obtained from ATCC and grown in Dulbecco’s Modified Eagle’s Medium medium (Thermo Fisher Scientific) supplemented with 10% fetal bovine serum (Sigma-Aldrich). To obtain bacterial conditioned medium, P. micra CM or E. coli CM was centrifuged at 4500 g for 15 mins and filtered through 0.22 μm filter. The filtered medium was diluted to 12.5% CM with cell culture medium (DMEM + 10% FBS). For MTT assay, cells were seeded in 96-well plates (1000 cells/well). The cell culture medium was then replaced with 100 µL 12.5% P. micra CM or E. coli CM. Cell viability was examined by MTT assay. For colony formation assay, HT-29 cells were plated in 6-well plates and treated with P. micra CM or E. coli CM. After culturing for 10–14 days, cells were fixed with 70% ethanol and stained with 0.5% crystal violet solution. Colonies with more than 50 cells per colony were counted. All experiments were conducted in triplicates.
Bacterial attachment assay
The bacterial attachment assay was performed as described previously described [11]; 1 × 106 HT-29 colon cancer cells were grown in a 6-well plate and co-cultured with bacteria for 2 h (multiplicity of infection = 100) under anaerobic condition. After co-culture, cells were washed with phosphate-buffered saline (PBS) for three times and lysed with 1% Triton X-100 for 5 min. Cells with attached P. micra colonies were then recovered on horse blood agar plates under anaerobic condition, and the number of bacteria colonies that are adherent onto epithelial cells was counted after 4 days.
Western blot
The total proteins isolated from cells or tissue samples and protein concentration were measured by DC Protein Assay (Bio-Rad Laboratories). 10 µg of proteins from each sample were separated on 10% SDS–PAGE and then transferred onto PVDF membranes. After blocking with 5% bovine serum albumin, blots were incubated with primary antibodies overnight at 4 °C. The primary antibodies used are as follows: activated β-catenin (05-665, Sigma), Cyclin D1 (#2922 S, Cell Signaling Technology), β-Actin (#4970, Cell Signaling Technology). Membranes were then incubated with secondary antibodies for 1 h at room temperature. Membranes were exposed to Pierce ECL Western Blotting Substrate (GE Healthcare). Band intensities were determined using ImageJ (National Institutes of Health).
PCR array
Mouse Cancer Finder PCR array PM033ZC (Qiagen), including 84 genes representative of 9 different biological pathways involved in transformation and tumorigenesis, and Mouse Inflammatory Response and Autoimmunity PCR Array PAMM-077 (Qiagen), including 84 key genes of inflammatory cytokines and chemokines as well as their receptors, were performed according to the manufacturer’s protocol.
Flow cytometry
Colon tissues were dissected and incubated in Hank’s balanced salt solution with 0.1 mg/mL collagenase D (Roche) and 50 U/mL DNase I (Roche) for 30 min at 37 °C. Lamina propria leukocytes was isolated as previously described [4]. Cells were then resuspended in staining solution (PBS with 2% fetal calf serum) for flow cytometry after staining with surface markers CD45 (103139, Biolegend, San Diego, CA), CD3 (100205, Biolegend), CD4 (100528, Biolegend), intracellular markers Il-17 (506903, Biolegend) and interferon (IFN)-γ (505806, Biolegend).
CD4+ T cells isolation and treatment
CD4+ T cells were isolated from mouse spleen by EasySep™ Mouse CD4+ T Cell Isolation Kit (STEMCELL Technologies, Vancouver, Canada). CD4+ T cells were treated with P. micra CM or broth control, and flow cytometry analysis of Il-17 (506908, Biolegend) and CD4 (100528, Biolegend) was performed 5 days after treatment.
Statistical analysis
All statistical tests were performed using GraphPad or R Software. Multiple group comparisons were analyzed by one-way analysis of variance (ANOVA) followed by post hoc tests (Bonferroni’s multiple comparisons test). The variables of the 2 sample groups were computed by Mann-Whitney U test or Student t-test. Two-way analysis of variance was performed to compare the difference in cell growth curves. The comparation of colonic dysplasia in different groups was analyzed by Chi-squared analysis. HR of survival associated with P. micra was estimated by using a univariate Cox regression model, and a multivariate Cox regression model was constructed to estimate the adjusted HR for high P. micra abundance. The overall survival in relation to P. micra abundance was evaluated by log-rank test. Data were presented as mean ± standard deviation. P < 0.05 was considered statistically significant.
References
Siegel RL, Miller KD, Goding Sauer A, Fedewa SA, Butterly LF, Anderson JC, et al. Colorectal cancer statistics, 2020. CA Cancer J Clin. 2020;70:145–64.
Wong SH, Yu J. Gut microbiota in colorectal cancer: mechanisms of action and clinical applications. Nat Rev Gastroenterol Hepatol. 2019;16:690–704.
Yu T, Guo F, Yu Y, Sun T, Ma D, Han J, et al. Fusobacterium nucleatum promotes chemoresistance to colorectal cancer by modulating autophagy. Cell. 2017;170:548–63.
Wong SH, Zhao L, Zhang X, Nakatsu G, Han J, Xu W, et al. Gavage of fecal samples from patients with colorectal cancer promotes intestinal carcinogenesis in germ-free and conventional mice. Gastroenterology. 2017;153:1621–33.
Tsoi H, Chu ESH, Zhang X, Sheng J, Nakatsu G, Ng SC, et al. Peptostreptococcus anaerobius Induces Intracellular Cholesterol Biosynthesis in Colon Cells to Induce Proliferation and Causes Dysplasia in Mice. Gastroenterology. 2017;152:1419–33.
Belkaid Y, Hand TW. Role of the microbiota in immunity and inflammation. Cell. 2014;157:121–41.
Zheng D, Liwinski T, Elinav E. Interaction between microbiota and immunity in health and disease. Cell Res. 2020;30:492–506.
Irrazabal T, Belcheva A, Girardin SE, Martin A, Philpott DJ. The multifaceted role of the intestinal microbiota in colon cancer. Mol Cell. 2014;54:309–20.
Louis P, Hold GL, Flint HJ. The gut microbiota, bacterial metabolites and colorectal cancer. Nat Rev Microbiol. 2014;12:661–72.
Kostic AD, Chun E, Robertson L, Glickman JN, Gallini CA, Michaud M, et al. Fusobacterium nucleatum potentiates intestinal tumorigenesis and modulates the tumor-immune microenvironment. Cell Host Microbe. 2013;14:207–15.
Long X, Wong CC, Tong L, Chu ESH, Ho Szeto C, Go MYY, et al. Peptostreptococcus anaerobius promotes colorectal carcinogenesis and modulates tumour immunity. Nat Microbiol. 2019;4:2319–30.
Dai Z, Coker OO, Nakatsu G, Wu WKK, Zhao L, Chen Z, et al. Multi-cohort analysis of colorectal cancer metagenome identified altered bacteria across populations and universal bacterial markers. Microbiome. 2018;6:70.
Yu J, Feng Q, Wong SH, Zhang D, Liang QY, Qin Y, et al. Metagenomic analysis of faecal microbiome as a tool towards targeted non-invasive biomarkers for colorectal cancer. Gut. 2017;66:70–8.
van Dalen PJ, van Steenbergen TJ, Cowan MM, Busscher HJ, de Graaff J. Description of two morphotypes of Peptostreptococcus micros. Int J Syst Bacteriol. 1993;43:787–93.
Sillanpaa J, Nallapareddy SR, Singh KV, Ferraro MJ, Murray BE. Adherence characteristics of endocarditis-derived Streptococcus gallolyticus ssp. gallolyticus (Streptococcus bovis biotype I) isolates to host extracellular matrix proteins. FEMS Microbiol Lett. 2008;289:104–9.
Koeth LM, Good CE, Appelbaum PC, Goldstein EJ, Rodloff AC, Claros M, et al. Surveillance of susceptibility patterns in 1297 European and US anaerobic and capnophilic isolates to co-amoxiclav and five other antimicrobial agents. J Antimicrob Chemother. 2004;53:1039–44.
Flemer B, Warren RD, Barrett MP, Cisek K, Das A, Jeffery IB, et al. The oral microbiota in colorectal cancer is distinctive and predictive. Gut. 2018;67:1454–63.
Purcell RV, Visnovska M, Biggs PJ, Schmeier S, Frizelle FA. Distinct gut microbiome patterns associate with consensus molecular subtypes of colorectal cancer. Sci Rep. 2017;7:11590.
Nakatsu G, Li X, Zhou H, Sheng J, Wong SH, Wu WK, et al. Gut mucosal microbiome across stages of colorectal carcinogenesis. Nat Commun. 2015;6:8727.
Wagner A, Wang C, Fessler J, DeTomaso D, Avila-Pacheco J, Kaminski J, et al. Metabolic modeling of single Th17 cells reveals regulators of autoimmunity. Cell. 2021;184:4168–85.
Yachida S, Mizutani S, Shiroma H, Shiba S, Nakajima T, Sakamoto T, et al. Metagenomic and metabolomic analyses reveal distinct stage-specific phenotypes of the gut microbiota in colorectal cancer. Nat Med. 2019;25:968–76.
Xu J, Yang M, Wang D, Zhang S, Yan S, Zhu Y, et al. Alteration of the abundance of Parvimonas micra in the gut along the adenoma-carcinoma sequence. Oncol Lett. 2020;20:106.
Loftus M, Hassouneh SA, Yooseph S. Bacterial community structure alterations within the colorectal cancer gut microbiome. BMC Microbiol. 2021;21:98.
La Vecchia S, Sebastian C. Metabolic pathways regulating colorectal cancer initiation and progression. Semin Cell Dev Biol. 2020;98:63–70.
Segata N, Haake SK, Mannon P, Lemon KP, Waldron L, Gevers D, et al. Composition of the adult digestive tract bacterial microbiome based on seven mouth surfaces, tonsils, throat and stool samples. Genome Biol. 2012;13:R42.
Park SY, Hwang BO, Lim M, Ok SH, Lee SK, Chun KS, et al. Oral-Gut microbiome axis in gastrointestinal disease and cancer. Cancers. 2021;13:2124.
Li B, Ge Y, Cheng L, Zeng B, Yu J, Peng X, et al. Oral bacteria colonize and compete with gut microbiota in gnotobiotic mice. Int J Oral Sci. 2019;11:10.
Grivennikov SI, Wang K, Mucida D, Stewart CA, Schnabl B, Jauch D, et al. Adenoma-linked barrier defects and microbial products drive IL-23/IL-17-mediated tumour growth. Nature. 2012;491:254–8.
Huber S, Gagliani N, Zenewicz LA, Huber FJ. Bosurgi L, Hu B, et al. IL-22BP is regulated by the inflammasome and modulates tumorigenesis in the intestine. Nature. 2012;491:259–63.
Wong SH, Kwong TNY, Chow TC, Luk AKC, Dai RZW, Nakatsu G, et al. Quantitation of faecal Fusobacterium improves faecal immunochemical test in detecting advanced colorectal neoplasia. Gut. 2017;66:1441–8.
Li Q, Hu W, Liu WX, Zhao LY, Huang D, Liu XD, et al. Streptococcus thermophilus Inhibits Colorectal Tumorigenesis Through Secreting beta-Galactosidase. Gastroenterology. 2021;160:1179–93.
Funding
This project was supported by National Key R&D Program of China (2020YFA0509200/2020YFA0509203); RGC-Research Impact Fund Hong Kong (R4032-21F); RGC-GRF Hong Kong (14163817); Vice-Chancellor’s Discretionary Fund CUHK and CUHK direct grant.
Author information
Authors and Affiliations
Contributions
L.Z. and X.Z. performed experiments, analysed data and drafted the manuscript. Y.Z., K.F., H.C-H.L and T.W-Y.L. performed experiments. A.H-K.C. performed the histological evaluation. O.O.C., W.K-K.W., S.H.W. and J.J-Y.S commented on the study and revised the manuscript. HW provided germ-free mice. K.F.T. supervised the study. J.Y. designed, supervised the study and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, L., Zhang, X., Zhou, Y. et al. Parvimonas micra promotes colorectal tumorigenesis and is associated with prognosis of colorectal cancer patients. Oncogene 41, 4200–4210 (2022). https://doi.org/10.1038/s41388-022-02395-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41388-022-02395-7
- Springer Nature Limited
This article is cited by
-
The microbial landscape of colorectal cancer
Nature Reviews Microbiology (2024)
-
DeepPIG: deep neural network architecture with pairwise connected layers and stochastic gates using knockoff frameworks for feature selection
Scientific Reports (2024)
-
Peptostreptococcus anaerobius mediates anti-PD1 therapy resistance and exacerbates colorectal cancer via myeloid-derived suppressor cells in mice
Nature Microbiology (2024)
-
The oral–gut microbiome axis in health and disease
Nature Reviews Microbiology (2024)
-
The Microbiome Matters: Its Impact on Cancer Development and Therapeutic Responses
Journal of Microbiology (2024)