Abstract
Epigenetic regulation is a pivotal factor during neuroblastoma (NB) pathogenesis and investigations into cancer epigenetics are actively underway to identify novel therapeutic strategies for NB patients. SUV39H1, a member of the H3K9 methyltransferase family, contributing to tumorigenesis across multiple malignancies. However, its specific role in NB remains unexplored. In this study, we conducted a high-throughput screen utilizing a compound library containing 288 epigenetic drugs, leading to the identification of chaetocin as the most potent NB inhibitor by targeting SUV39H1. Genetic manipulation and therapeutic inhibition of SUV39H1 significantly impacted proliferation, migration, cell cycle phases, and apoptosis in NB cells. Concurrently, chaetocin demonstrated robust anti-tumor efficacy in vivo with tolerable toxicity. RNA-seq unveiled that SUV39H1 knockdown and inhibition down-regulated cell cycle pathways, impacting vital genes such as AURKA. Besides, MCPIP1 emerged as a novel tumor suppressor following SUV39H1 inhibition, which decreased AURKA expression in NB. In detail, SUV39H1 mediated the enrichment of H3K9me3 at the promoter region of MCPIP1, repressing the MCPIP1-mediated degradation of AURKA and facilitating the subsequent accumulation of AURKA, which revealed the oncogenic role of SUV39H1 via the SUV39H1-MCPIP1-AURKA signaling axis in NB. Therapeutic inhibition of SUV39H1 using chaetocin emerges as an effective and safe strategy for NB patients.

Illustration of the oncogenic pathway regulated by SUV39H1 in NB.
Similar content being viewed by others
Introduction
Neuroblastoma (NB) is the most common extracranial malignant solid tumor in children, constituting approximately 8% of pediatric malignancies [1]. While recent advances in treatment, including high-intensity combinatorial treatments involving surgery, chemoradiotherapy and immunotherapy, have tremendously improved survival rates for low- and medium-risk NB patients, those with high-risk NB still exhibit therapeutic resistance, resulting in an overall survival (OS) rate of less than 40% [2]. Therefore, addressing the clinical challenges in high-risk NB remains critical in pediatric oncology research. And further understanding the molecular regulatory network governing the malignant progression of NB holds promise for new and novel therapeutic options.
Epigenetic regulation plays a vital role in the tumorigenesis of tumors [3], thus motivating many investigations on relevant therapies based on the specific epigenetic landscapes in pediatric tumors including neuroblastoma [4,5,6]. Major epigenetic modifications include abnormal DNA methylation, post-translational histone modification, chromatin remodeling, and abnormal expression of non-coding RNA. Current epigenetic therapy in NB mostly acts at DNA methylation by targeting of DNA methyltransferases (DNMT), histone modification by inhibiting histone acetylases/methylases, and epigenetic readers [4, 5]. Among the above targets, our high-throughput drug screen reveals that histone-modifying drugs exhibit inhibitory effects on NB cells. Chaetocin, as an epigenetic drug selectively for histone H3K9 methyltransferase SUV39H1, can exert the anti-tumor effect across diverse cancers, including melanoma [7], non-small cell lung cancer [8], ovarian cancer [9], gastric cancer [10], glioma [11] and liver cancer [12]. Nevertheless, its role and molecular mechanisms in NB remain unexplored.
SUV39H1, as a member of the H3K9 methyltransferase family, has been reported to be highly expressed in various malignancies, including hepatocellular carcinoma [13], breast cancer [14], colorectal cancer [15], cervical cancer [16, 17] and glioma [18], influencing tumor proliferation, metastasis, apoptosis, and patient prognosis. Although these results suggest that SUV39H1 may affect tumorigenesis through different regulatory mechanisms, the specific role of SUV39H1 in NB also remains elusive.
In light of these knowledge gaps, our study aims to investigate the clinical associations of SUV39H1 in NB patients and systematically unravel its role and regulatory mechanisms. Furthermore, we focus on delineating the anti-tumor effect of the SUV39H1-specific inhibitor chaetocin and evaluating its potential in pre-clinical models. Overall, our study provides the clinical and experimental basis for future translational research by enhancing our understanding of NB biology and precision treatment in NB patients.
Results
The high-throughput drug screen identifies the potent anti-NB effect of the SUV39H1 inhibitor chaetocin
The intricate characteristics of high-risk NB, including rare high-frequency mutations and genetic/epigenetic heterogeneity, complicate the development of more effective targeted therapy and limit the patient response to conventional treatments [4]. To address this clinical challenge, we aim to identify novel anti-NB drug candidates and explore the regulatory mechanism underlying the malignant progression of NB.
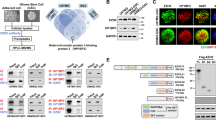
First, we carried out a high-throughput drug screen, evaluating the cytotoxicity of 288 small molecule candidates on NB cells. Two NB cell lines, SK-N-SH (MYCN non-amplified) and SK-N-BE (2) (MYCN amplified) that represented two classic NB types were selected for the screen. We identified 36 drugs exhibiting substantial tumor control (>50% tumor cell killing) across both cell lines. Remarkably, chaetocin, a highly specific SUV39H1 inhibitor, emerged as the most potent drug for NB cell killing (Fig. 1A, B). Building upon this discovery, we further investigated the specific effect and molecular mechanism of chaetocin and its downstream target SUV39H1 in both NB cells and NB tumor-bearing mice.
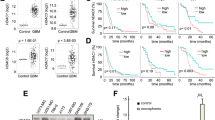
A Drug library screening identified 36 common effectively-anti-NB drugs in SK-N-SH after 5 μM various drugs treatment for 72 h, with DMSO as the control. B Drug library screening identified 36 common effectively-anti-NB drugs in SK-N-BE (2) after 5 μM various drugs treatment for 72 h, with DMSO as the control. C Bioinformatic analysis demonstrated the expression level of SUV39H1 between NB tissues (162 samples) and normal tissues (1280 normal tissues including 128 normal adrenal gland and 1152 normal brain samples) based on the combined cohort of TCGA, TARGET, and GTEx samples obtained from UCSC Xena browser (TCGA TARGET GTEx cohort, https://xenabrowser.net/datapages/?cohort=TCGA%20TARGET%20GTEx&removeHub=https%3A%2F%2Fxena.treehouse.gi.ucsc.edu%3A443). The Wilcox method was used for the difference analysis, with R package used for this bioinformatics analysis procedure. D R2 online website platform (http://r2.amc.nl) indicated the association of SUV39H1 with the prognosis of 786 NB patients (the tumor Neuroblastoma integrated platforms Cangelosi database), and the cut-off was determined using the scanning method. The log-rank test was performed for survival analysis. E qRT-PCR analysis revealed the expression level of SUV39H1 in 5 NB cell lines as compared to the normal HEK293T cell at the mRNA level. One-way ANOVA was performed for the statistical analysis. F WB analysis revealed the expression level of SUV39H1 in 5 NB cell lines as compared to the normal HEK293T cell at the protein level. Shown is one representative result of three independent experiments. G Representative pictures of IHC staining from 61 NB tumor tissues from SYSU cohort. H Quantitative analysis of SUV39H1 protein expression in NB tissues from SYSU cohort (NB patients expressing high or low levels of SUV39H1 was defined based on the IHC scores evaluated from the IHC staining: scores of 0–4 were classified as a low expression, whereas scores of 5–9 were classified as a high expression). The t-test was performed for statistical analysis. I Overall survival (left panel) and event-free survival (right panel) analysis of 61 NB patients from SYSU cohort expressing high or low levels of SUV39H1. The log-rank test was performed for survival analysis. Data were shown as the mean ± SEM, *p < 0.05, ***p < 0.001, ****p < 0.0001.
Then, based on comprehensive bioinformatics analysis on public NB transcriptome cohorts and the R2 online platform, we verified that SUV39H1 was highly expressed in NB cells and correlated with the inferior prognosis of NB patients (Fig. 1C, D). Compared to the normal control HEK293T cell [19, 20], SUV39H1 was overexpressed in 5 NB cell lines both at the mRNA and protein levels (Fig. 1E, F). IHC staining of SUV39H1 in 61 NB tumor tissues from our independent patient cohort validated the negative correlation between SUV39H1 expression and prognosis of NB patients (Fig. 1G–I), indicating that SUV39H1 might play a pivotal role in NB progression and serve as a promising therapeutic target for NB patients (Tables 1 and 2).
Chaetocin exhibits anti-tumor effects on NB cells both in vitro and in vivo
To comprehensively understand the impact of chaetocin on NB biology, we conducted functional experiments which demonstrated that chaetocin inhibited proliferation (Fig. 2A, B) and migration of NB cells (Fig. 2C, D). In addition, chaetocin treatment also induced NB cell apoptosis (Fig. 2E) and caused cell cycle arrested (Fig. 2F). Given the SUV39H1’s role as a histone methyltransferase (HMTase) for histone H3 lysine 9 trimethylation (H3K9me3) [21], we hypothesized that inhibition of SUV39H1 activity via chaetocin would reduce H3K9me3 expression in NB cells. IF staining and WB results collectively demonstrated that H3K9me3 in the NB cell nucleus was downregulated after chaetocin treatment (Fig. 2G, H). Chaetocin treatment also achieved good tumor control in the NOD SCID mouse model (Fig. 2I), exhibiting slower tumor growth kinetics compared to the vehicle treatment group (Fig. 2J, K). Importantly, there was no significant difference in the mouse weight between the two groups (Fig. 2L), indicating the in vivo safety of chaetocin at the effective dose. The H&E and IHC analysis of H3K9me3 indicated a decrease of H3K9me3 in mice tumors after chaetocin treatment (Fig. 2M).
A CCK-8 analysis identified the IC50 values of SK-N-SH and SK-N-BE (2) after chaetocin treatment for 48 h. The cell viability rate was calculated as (ODtest − ODblank)/(ODcontrol − ODblank)*100. The Graphpad software was used to fit the growth curve and calculate the IC50 value. B Colony formation analysis of NB cells after chaetocin treatment with different concentrations for 48 h, and the colonies of SK-N-SH (after 14 days) and SK-N-BE (2) (after 12 days) were collected and stained with crystal violet. C, D Wound healing and transwell assay illustrated the migration abilities of NB cells after treatment with chaetocin for 24 h. E Apoptosis analysis by flow cytometry in NB cells after treated with chaetocin for 48 h. F Cell cycle distribution changes caused by chaetocin treatment for 12 h. G IF staining of H3K9me3 in NB cells located in nucleus with or without chaetocin treatment for 48 h. H WB analysis illustrated levels of the H3K9me3 protein after chaetocin treatment for 48 h. I Pictures of harvested tumors in NOD SCID mice in the control (n = 5) or chaetocin (n = 5) treatment group. J Tumor volumes of NB xenograft models with control (n = 5) or chaetocin treatment (n = 5). K Tumor weight of tumors from the control (n = 5) or the chaetocin group (n = 5). L Body weights of NB-bearing mice after chaetocin treatment (n = 5) or not (n = 5). M H&E and IHC analysis of H3K9me3 in tumors from the control or the chaetocin group. Shown is one representative picture of indicated groups (n = 5 for each group). B–E, G, J The one-way ANOVA with multiple comparisons to the control was performed for statistical analysis. F, H The two-way ANOVA was performed for statistical analysis. K The t-test was performed for statistical analysis. The experiments were repeated three times with similar results. Data were shown as the mean ± SEM, ns: non significance, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
SUV39H1 exerts an oncogenic role in NB cells both in vitro and in vivo
Next, the specific impact of chaetocin’s target SUV39H1 on the biological functions of NB cells was elucidated through genetic knockdown of SUV39H1. We revealed that NB cells exhibited substantial regression in proliferative and migrating capabilities upon SUV39H1 knockdown (Fig. 3A–D). Consistent with the chaetocin treatment results, SUV39H1 knockdown also induced higher apoptosis in NB cells (Fig. 3E) and more cycling cells arrested at the G2 phase (Fig. 3F). SUV39H1 knockdown also decreased the nuclear H3K9me3 at the protein level (Fig. 3G, H). The oncogenic role of SUV39H1 was further confirmed in vitro using NB cells with stable SUV39H1 overexpression (Supplementary Fig. 2).
A CCK-8 analysis identified the proliferation abilities of SK-N-SH and SK-N-BE (2) after genetic knockdown of SUV39H1. B Colony formation analysis of NB cells after SUV39H1 inhibition with shRNAs for 14 days (SK-N-SH) or 12 days (SK-N-BE (2)). C, D Wound healing (24 h) and transwell assay (24 h) demonstrated the migration abilities of NB cells after SUV39H1 stably suppression with shRNAs. E Changes of apoptosis in NB cells after SUV39H1 knock-down with siRNAs. F Changes of the cell cycle distribution based on SUV39H1 inhibition with shRNAs. G IF staining of H3K9me3 in NB cells located in nucleus after stably inhibiting SUV39H1 with shRNAs. H WB analysis revealed changes of H3K9me3 protein with SUV39H1 stable suppression by shRNAs. I Pictures of harvested tumors in NOD SCID mice from independent groups (n = 5 for each group) with or without SUV39H1 genetic knockdown. J Tumor volumes of NB xenograft models with or without SUV39H1 genetic knockdown (n = 5 for each group). K Tumor weight of tumors with or without SUV39H1 genetic knockdown (n = 5 for each group). L H&E and IHC analysis of H3K9me3 tumors with or without SUV39H1 genetic knockdown (n = 5 for each group). Shown is one representative picture of indicated groups. B, D, E, G, K The one-way ANOVA with multiple comparisons to the control was performed for statistical analysis. A, F, H, J The two-way ANOVA was performed for statistical analysis. The experiments were repeated three times with similar results. Data were shown as the mean ± SEM, ns: non significance, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
In vivo validation using the NOD SCID mouse model demonstrated that SUV39H1 knockdown significantly inhibited NB growth (Fig. 3I). The tumor growth kinetics in the SUV39H1 knockdown group was tremendously slower than that in the control group (Fig. 3J), and tumor weight was also reduced in the SUV39H1 knockdown group (Fig. 3K). The H&E and IHC analysis of H3K9me3 indicated a decrease of H3K9me3 in mice tumors after SUV39H1 genetic knockdown (Fig. 3L).
SUV39H1 induces cell cycle up-regulation in NB cells
To further explore the molecular mechanisms governed by SUV39H1 in NB cells, we performed the RNA-sequencing (RNA-seq) in NB cells after chaetocin treatment and SUV39H1 knockdown. Significant differentially expressed genes (DEGs) from both SUV39H1 knockdown and chaetocin-treated NB cells were depicted in the Venn diagram (Fig. 4A). Metascape software [22] was used to perform the KEGG pathway analysis of the most deregulated genes in the knockdown and treatment groups. Notably, the top 3 pathways exhibiting significant changes were enriched in cell division, cell cycle, and the PID PLK1 pathway (Fig. 4B), suggesting a potential role of SUV39H1 in NB oncogenesis through the regulation of cell cycle processes.
A Overlapped DEGs from the RNA-seq after chaetocin treatment (50 nM, 48 h) and SUV39H1 genetic knockdown with siRNA (48 h after transfection) in SK-N-SH cell, and there were three biological replicates for each sample. B KEGG analysis of DEGs with the same change trend under the above two conditions. C Heatmap showed the crucial genes enriched at the cell cycle pathway after SUV39H1 inhibition by chaetocin treatment or genetic knockdown. D, E qRT-PCR analysis verified genes with significant changes in the cell cycle pathway at the mRNA level. F, G WB analysis verified genes with significant changes in the cell cycle pathway at the protein level. H Spearman correlation analysis (https://www.cbioportal.org/) of 143 NB samples based on the TARGET database indicated the positive correlation between SUV39H1 and AURKA. I Rescue experiments of cell cycle analysis (12 h) indicated that the cell cycle arrest caused by SUV39H1 knockdown could be reversed by AURKA overexpression. D–G, I The two-way ANOVA was performed for statistical analysis. The experiments were repeated three times with similar results. Data were shown as the mean ± SEM, ns: non significance, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
While SUV39H1’s role in regulating cell cycle processes has been reported in various cancers [15, 23], its specific function in NB remains understudied. Given our earlier findings that inhibiting SUV39H1 resulted in cell cycle arrest at the G2 phase in NB cells, we sought to delve into the molecular basis behind this phenomenon. The heatmap in Fig. 4C showed the key genes based on the RNA-seq, which were enriched at the cell cycle pathway, including AURKA, CDK1, CCNB1, CCNA2, TPX2, WEE1, PLK1, CDC25B, and CDC25C. Then these cell cycle regulation associated genes were validated at the mRNA level using qRT-PCR (Fig. 4D, E). Subsequently, critical phase conversion proteins, such as AURKA, CDK1, CCNB1, and CCNA2, were further verified by WB (Fig. 4F, G).
These results collectively indicated that inhibition of SUV39H1 affected cell cycle pathways in NB cells. More importantly, Spearman correlation analysis using the TARGET database indicated a strong positive correlation between SUV39H1 and AURKA (Fig. 4H). Rescue experiments revealed that re-introduction of AURKA partially reversed the G2 phrase arrest caused by SUV39H1 knockdown (Fig. 4I). Above all, the downregulation of AURKA, a master regulator critical for cell division, highlighted the significance of SUV39H1 in orchestrating NB cell cycle dynamics.
MCPIP1 emerges as a potential downstream regulator of SUV39H1
Since the preceding results verified the H3K9me3-dependent regulatory model of SUV39H1 on its targeted genes shown in Fig. 2 and 3, and H3K9me3 is a marker of transcriptional inhibition [17, 21, 24, 25], we speculated that inhibiting SUV39H1 might attenuate the transcriptional inhibitory effect from H3K9me3 on its target genes, thereby facilitating the transcriptional re-activation of specific genes. To explore this further, we analyzed our RNA-seq data and public datasets and identified the top 20 significantly up-regulated genes in both SUV39H1 knockdown and chaetocin treatment conditions (Fig. 5A). From this, MCPIP1 emerged as an important candidate for subsequent investigations. While MCPIP1 has been identified as a tumor suppressor in previous studies [26,27,28], its explicit regulatory mechanism in NB cells remains unclear. Therefore, we corroborated that SUV39H1 inhibition significantly up-regulated MCPIP1 at both the mRNA and protein levels, consistent with RNA-seq results (Fig. 5B–D). Spearman correlation analysis using the TARGET database also indicated a negative correlation between MCPIP1 and SUV39H1 in clinical samples (Fig. 5E). Down-regulation of MCPIP1 after SUV39H1 inhibition was confirmed across 5 NB cell lines via qRT-PCR (Fig. 5F). The clinical significance of MCPIP1 was explored through Kaplan–Meier survival analysis, revealing that low MCPIP1 expression is associated with an inferior prognosis for NB patients (Fig. 5G). Functional in vitro experiments validated the tumor-suppressing role of MCPIP1 in NB progression. Ectopic overexpression of MCPIP1 resulted in diminished proliferative and migrating capacities (Supplementary Fig. 3), the increasingly induced apoptosis, and aberrant cell cycle arrest in NB cells (Fig. 5H–M).
A Heatmap showed the common top 20 upregulated genes after SUV39H1 inhibition by chaetocin treatment or genetic knockdown. B qRT-PCR analysis confirmed the upregulation of MCPIP1 in NB cells after chaetocin treatment at 25 nM or 50 nM for 48 h. C qRT-PCR analysis confirmed the upregulation of MCPIP1 in NB cells after SUV39H1 genetic knockdown with shRNAs. D WB analysis confirmed the upregulation of MCPIP1 in NB cells after chaetocin treatment for 48 h or genetic knockdown of SUV39H1 with shRNAs. E Spearman correlation analysis of 143 NB samples based on the TARGET database indicated the negative correlation between MCPIP1 and SUV39H1. F qRT-PCR analysis revealed the expression level of MCPIP1 in 5 NB cell lines as compared to the normal HEK293T cell. G R2 online website platform indicated the association of MCPIP1 with the prognosis of NB patients, and the cut-off was determined using the scanning method. H qRT-PCR confirmed the transfection efficiency of overexpressing MCPIP1 in NB cells after transfection for 48 h. I WB analysis validated the transfection efficiency of overexpressing MCPIP1 transfection for 72 h in NB cells. J CCK-8 analysis identified the proliferation abilities of NB cells after MCPIP1 overexpression at 0, 24, 48, 72 and 96 h. K Colony formation analysis of NB cells after MCPIP1 overexpression (14 days for SK-N-SH and 12 days for SK-N-BE (2)). L Changes of apoptosis in NB cells after overexpressing MCPIP1. M Changes of the cell cycle distribution based on MCPIP1 overexpression. B, C, M The two-way ANOVA was performed for statistical analysis. F, J The one-way ANOVA with multiple comparisons to the control was performed for statistical analysis. H, K, L The t-test was performed for statistical analysis. The experiments were repeated three times with similar results. Data were shown as the mean ± SEM, ns: non significance, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
MCPIP1 down-regulates AURKA in NB cells
A study by El Village Bieta Boratyn’s team showed that overexpression of MCPIP1 could cut AURKA transcripts at the mRNA level through its ribonuclease function, promoting the degradation of AURKA [29]. Based on the above findings, we hypothesized that MCPIP1 could modulate cell cycle process of NB cells through negative regulation of AURKA. Thus, we overexpressed MCPIP1 (oeMCPIP1) in SK-N-SH and SK-N-BE (2) cells, and found that AURKA was downregulated as expected (Fig. 6A). Results of WB further confirmed this phenomenon at the protein level (Fig. 6B). When we overexpressed AURKA (Fig. 6C, D) in oeMCPIP1-NB cells, we found that the proliferation inhibition caused by MCPIP1-overexpression could be partially reversed (Fig. 6E, F), indicating that the tumor suppressive function of MCPIP1 in NB was partially dependent on its degradation of AURKA.
A qRT-PCR analysis confirmed the downregulation of AURKA in NB cells after MCPIP1 overexpression. B WB analysis confirmed the downregulation of AURKA in NB cells after MCPIP1 overexpression. C qRT-PCR confirmed the transfection efficiency of overexpressing AURKA in NB cells after transfection for 48 h. D WB confirmed the transfection efficiency of overexpressing AURKA in NB cells after transfection for 72 h. E NB cells were overexpressed with MCPIP1, AURKA was subsequently overexpressed then, and CCK-8 analysis was performed to identify the proliferation abilities of NB cells. F NB cells were overexpressed with MCPIP1, AURKA was subsequently overexpressed then, and colony formation assay was performed to identify the colony formation abilities of NB cells. A, E The two-way ANOVA was performed for statistical analysis. C The t-test was performed for statistical analysis. F The one-way ANOVA with multiple comparisons to the control was performed for statistical analysis. The experiments were repeated three times with similar results. Data were shown as the mean ± SEM, *p < 0.05, ***p < 0.001, ****p < 0.0001.
SUV39H1 regulates the cell cycle progression of NB cells through the MCPIP1-AURKA signaling axis
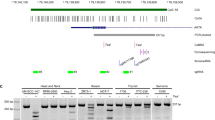
Our preceding results identified the tumor-promoting role of SUV39H1 in NB cells, and revealed that SUV39H1 inhibition dramatically activated MCPIP1 expression. Next, we postulated that SUV39H1 might regulate MCPIP1 expression through H3K9me3-dependent epigenetic silencing. The ChIP-PCR assay was conducted with primers that were specifically designed against about every 300 bp of the MCPIP1 promoter (−3000 bp to 0 bp) (Fig. 7A). The result indicated that H3K9me3 bound to the MCPIP1 promoter at regions of 4, 5, and 6 (−1987 bp to −1015 bp), with significant enrichment discrepancies between the control and treatment groups (Fig. 7B, C). This indicated that SUV39H1 transcriptionally silenced MCPIP1 through the H3K9me3 epigenetic modification.
A The binding site of H3K9me3 on the promoter of MCPIP1. B, C ChIP-qPCR analysis indicated the H3K9me3 enrichment at MCPIP1 promoter after chaetocin treatment at 50 nM for 48 h or SUV39H1 inhibition with shRNAs. D NB cells were overexpressed with SUV39H1, MCPIP1 was subsequently overexpressed then, and CCK-8 analysis was performed to identify the proliferation abilities of NB cells. E NB cells were overexpressed with SUV39H1, MCPIP1 was subsequently overexpressed then, and colony formation assay was performed to identify the colony formation abilities of NB cells. F NB cells were overexpressed with SUV39H1, MCPIP1 was subsequently overexpressed then, and WB analysis showed the AURKA expression level. B, C, D The two-way ANOVA was performed for statistical analysis. E The one-way ANOVA with multiple comparisons to control was performed for statistical analysis. The experiments were repeated three times with similar results. Data were shown as the mean ± SEM, *p < 0.05, **p < 0.01, ***p < 0.001.
Given AURKA’s significance in the cell cycle and its decreased expression after SUV39H1 inhibition, we speculated that SUV39H1 might regulate AURKA through targeted inhibition of MCPIP1, and then played a crucial role in the cell cycle regulation of NB cells. To verify our hypothesis, we re-introduced MCPIP1 in SUV39H1-overexpression cells. Both CCK-8 and colony formation assays demonstrated that overexpression of MCPIP1 partially reversed the oncogenic role of SUV39H1 in NB cells, consistent across SK-N-SH and SK-N-BE (2) cell lines (Fig. 7D, E). Besides, the WB result revealed that SUV39H1 overexpression led to the up-regulation of AURKA, while the re-introduction of MCPIP1 partially reversed this effect (Fig. 7F).
These results collectively support the inference that the regulation of SUV39H1 on downstream AURKA is contingent on the epigenetic silencing of MCPIP1. In essence, SUV39H1 orchestrates cell cycle progression in NB cells through the MCPIP1-AURKA signaling axis.
Discussion
NB is the most common extracranial malignant solid tumor in children that undergoes defective differentiation due to genomic and epigenetic damage [1, 2]. There still remains a challenge for patients with high-risk NB. Novel therapeutic strategies targeting specific genetic/epigenetic alterations might provide promising outcome for NB patients [4,5,6].
In the present study, high-throughput screen of a pharmacy library containing 288 small molecules helped to identify the significant killing effect of chaetocin on NB cells. Based on our preliminary screening results, several compounds showed significant anti-NB activities including LMK235, CUDC-907, Panobinostat, Dorsomorphin and Mitomycin C, etc. According to reported literatures, the exact roles and mechanisms of several compounds mentioned above had already been investigated in NB [19, 30,31,32,33,34], with an unelucidated role of chaetocin in NB yet. It was revealed that the anti-NB effect of chaetocin was significantly in both SK-N-SH and SK-N-BE (2). Besides, the target gene of chaetocin, SUV39H1, has become a popular gene that widely studied recently because of its gradually discovered roles in tumor development and anti-tumor immunity [18, 35,36,37,38,39]. However, the role of SUV39H1 in NB has not been reported yet. Given the above reasons, we selected chaetocin and SUV39H1 as the subjects of interest, focusing on the function and mechanism behind them in this study. As the highly selective inhibitor of SUV39H1, chaetocin exerted anti-NB effect both in the MYCN non-amplified cell line SK-N-SH and in the MYCN amplified cell line SK-N-BE (2). Both bioinformatic analysis and experiment validation indicated the high expression level and the prognostic value of SUV39H1 in NB. Subsequent experiments both in vitro and in vivo revealed the oncogenetic role of SUV39H1 in NB. Targeted inhibition of SUV39H1 in NB showed a significant anti-tumor effect, and SUV39H1 might be a potential therapeutic target for NB. However, the mechanism of SUV39H1 overexpression in NB remains unclear, and more studies are needed to clarify it in the future.
As a member of the H3K9 methyltransferase family, SUV39H1 is highly expressed in breast cancer [14], colorectal cancer [15], cervical cancer [16, 17], glioma [18], et al, and it is reported to participate in regulating the growth, proliferation, metastasis, and the apoptosis resistance of tumor cells. One recent study reported that SUV39H1 was involved in the ferroptosis process in renal clear cell carcinoma via epigenetically regulating DDP4 [35], and it also regulated T cell differentiation [40, 41] and affected the anti-tumor immunity in several tumors [42,43,44]. However, SUV39H1 was also reported to role as a tumor suppressor in acute myeloid leukemia [45]. It can be concluded that the reported effects of SUV39H1 diverged from different tumors excepting NB. In this study, we reported for the first time the high expression level of SUV39H1 and its oncogenetic effect in NB, and the regulatory mechanism was also investigated subsequently.
Based on the results from the RNA-seq, MCPIP1 was one of the top 20 up-regulated genes, which was confirmed to be a tumor suppressor in NB: it inhibited NB cell proliferation and migration, promoted the apoptosis and mediated the cell cycle arrest. It was also illustrated that SUV39H1 inhibition reactivated the transcriptional expression of MCPIP1, which was due to the reduced H3K9me3 enrichment at the promoter region of MCPIP1. This phenomenon may be one of the reasons for the generally low expression level of MCPIP1 in NB cell lines, which needs further investigation in the future.
AURKA (aurora kinase A) belongs to the mitotic serine/threonine kinase family and is localized at the centrosome with highly conserved in evolution. AURKA plays an indispensable role in many biological processes, including centrosome maturation and separation, spindle assembly, chromosome arrangement, etc. [46]. Studies have shown that AURKA was overexpressed in various tumors including neuroblastoma [46, 47], and indicated a poor prognosis in patients. Iwona Nowak’s team has demonstrated that MCPIP1 can promote the degradation of AURKA in NB cell lines at the transcriptional level through its ribonuclease function [29]. We further verified that MCPIP1 caused the decreased expression level of AURKA at mRNA and protein levels. Rescue experiments confirmed that the suppressive role of MCPIP1 was partially dependent on its degradation of AURKA. Besides, as an important regulation gene in the cell cycle signal pathway, AURKA was significantly downregulated after SUV39H1 suppression, and it could reverse the cell cycle arrest caused by SUV39H1 inhibition. Therefore, we confirmed that SUV39H1 inhibits the expression of MCPIP1 through epigenetic silencing, thus weakens the mRNA degradation of AURKA, enhancing the effect of AURKA on the process of NB cell cycle, and then promotes the proliferation of NB cells.
Conventional treatments for NB include chemotherapy, radiotherapy and surgery. However, traditional treatment strategies may lack specificity with low effectiveness, and may cause serious side effects [12]. Chaetocin is a natural metabolite isolated from trichomyces thiodiketopyrazines with a variety of biological activities and pharmacological functions. As early as 2005, chaetocin was found to inhibit the activity of histone lysine methyltransferase SUV39H1 [48]. However, as the study progressed, researchers gradually discovered that the compound can also regulate many other proteins apart from SUV39H1 [12, 49,50,51]. Nevertheless, due to its extensive anti-tumor activity both in vivo and in vitro, it still has stimulated a wide range of research interests of scientists [12]. However, the role and mechanism of chaetocin in NB have not been reported before. In our study, we identified for the first time the significant anti-tumor effect of chaetocin on NB and the toxicity in vivo is tolerable at the working dose. This suggests that, as a highly selective inhibitor of SUV39H1, chaetocin has potential translational application value in anti-NB therapy. It is worth discussing that in the studies reported so far, most researchers used chaetocin as an inhibitor of SUV39H1, but there is no commercial small molecule inhibitor singly targeting SUV39H1. Given the current background, we have the confidence to believe the necessary of developing small molecule inhibitors specifically targeting SUV39H1, which may provide individualized treatment options for tumor patients with high level of SUV39H1.
Conclusions
To conclude, our study identifies that SUV39H1 is significantly overexpressed and plays an oncogenic role in NB cells. Mechanistically, we elucidate that high expression of SUV39H1 in NB cells transcriptionally silences its target MCPIP1 through epigenetic modification of MCPIP1, marked by H3K9me3 enrichment at the MCPIP1 promoter region, which subsequently reduces MCPIP1-mediated degradation of AURKA. This molecular process enhances cell cycle processes in NB cells, thus promoting the malignant phenotypes. In short, SUV39H1 promotes NB progression through the MCPIP1-AURKA signaling axis. Chaetocin treatment diminishes the H3K9 methyltransferase activity of SUV39H1 and suppresses the cell cycle progression of NB cells, and thus might generate clinical benefit to NB patients as an effective and safe epigenetic therapy (Graphical abstract).
Materials and methods
Cell lines
Five NB cell lines used in the study, including SK-N-SH, SK-N-BE (2), SH-SY5Y, IMR32, and SK-N-AS, were purchased from COBIOER BIOSCIENCES (Nanjing, China). SK-N-BE (2) and SH-SY5Y were cultured in MEM/F12 (1:1) (Gibco, USA) supplemented with 10% fetal bovine serum (FBS, Gibco, USA); SK-N-SH and IMR32 were cultured in MEM (Gibco, USA) supplemented with 10% FBS (Gibco, USA); SK-N-AS was cultured in DMEM (Gibco, USA) supplemented with 10% FBS (Gibco, USA). All mediums were supplemented with 1% penicillin–streptomycin solution (Gibco, USA). All cell lines had been authenticated by STR and are free from mycoplasma.
Cell counting kit-8 (CCK-8) assay
NB cells were seeded in 96-well plates with three replicates for each sample. After the indicated treatment, 10% CCK-8 solution (APExBIO, USA) was added to the plate and continued to be incubated for 3 h at 37 °C in the cell incubator. Then, the absorbance was detected with the spectrophotometry at 450 nm.
Compounds and drug screening
The small molecule compound library containing 288 molecular drugs was purchased from Selleck (Shanghai, China). Two representative NB cell lines were selected for the high-throughput screening: SK-N-SH (MYCN non-amplified) and SK-N-BE (2) (MYCN amplified). Cells pre-inoculated into 96-well plates at a density of 5000 cells/well. The working concentration of the drug was 5 μM, with DMSO as the control. After 72 h, cell viability was detected by CCK-8 and the survival rate was calculated [52]. An effective drug is defined as the tumor survival rate lower than 50%. The drugs that showed co-efficacy in both two cell lines were screened out, and the natural antifungal product chaetocin, with a significant killing effect was identified.
Bioinformatics analysis
The combined cohort of TCGA, TARGET, and GTEx samples were obtained from UCSC Xena browser (TCGA TARGET GTEx cohort, https://xenabrowser.net/datapages/?cohort=TCGA%20TARGET%20GTEx&removeHub=https%3A%2F%2Fxena.treehouse.gi.ucsc.edu%3A443), and there were 162 NB samples and 1280 normal tissues (128 normal adrenal gland and 1152 normal brain tissues) [19]. The Wilcox method was used for the difference analysis, with R package used for the bioinformatics analysis of the SUV39H1 expression levels between NB samples and normal tissues. For the Kaplan–Meier survival analysis, overall survival (OS) and event-free survival (EFS) curves between NB patients expressing high/low levels of SUV39H1 or MCPIP1 were generated through the R2 online website platform (http://r2.amc.nl), followed by the log-rank test. The scanning method was used to determine the cut-off [53, 54]. For the Spearman correlation analysis based on the TARGET database, the mRNA expression levels of indicated genes were analyzed based on the RNA-seq results from 143 NB samples (https://www.cbioportal.org/).
Immunohistochemistry (IHC) assay
The 61 NB tissue specimens obtained from our center (all patients were <18 years old with the initial diagnosis as NB, and the follow-up data was complete) were fixed in formalin, embedded in paraffin blocks, and sectioned with a thickness of 4 μm. Then, IHC staining was performed following instructions as previously described [19]. Anti-SUV39H1 rabbit pAb (Abclonal, China, A3277) was used in this assay. We used a fluorescence microscope to capture images (Olympus BX61, Japan). The intensity was scored as follows: 0—negative; 1—weak; 2—moderate; 3—strong. The proportion of positive cells was scored as follows: 0—less than 25%; 1—25% to 50%; 2—50% to 75%; 3—75% to 100%. The product of the above two scores indicated the composite staining score. Two expression staining levels were defined: scores of 0–4 were classified as a low expression, whereas scores of 5–9 were classified as a high expression.
RNA isolation and quantitative real-time polymerase chain reaction (qRT-PCR) analysis
The total RNA from the cultured cells was extracted with the Trizol reagent (Life Technologies, USA). The RNA was reverse transcribed into cDNA using the Prime Script RT Master Mix (TaKaRa, Japan) following the manufacturer’s instructions. Then, qRT-PCR was conducted with a Power SYBR Green Master Mix (Dongsheng Biotech, China) using primers listed in Supplementary Table 1, with the β-actin as an endogenous control.
Western blotting (WB) analysis and antibodies
The Whole Cell Lysis Assay Kit (KeyGEN, China) was used to extract the whole-cell lysates. The BCA method (Thermo Fisher Scientific, USA) was applied to detect the protein concentration. β-actin (1: 1000, Proteintech, China, 66009) was used as the internal control. An equal amount of each protein sample was separated with SDS-PAGE and transferred onto a PVDF membrane. Then, the membrane was blocked in 5% skim milk and incubated with primary antibodies overnight, followed by incubating with indicated secondary antibody for 1 h at room temperature. Primary antibodies used this study were listed in Supplementary Table 2. The signal detection was performed with the ECL chemiluminescence detection system (Bio-Rad, USA).
Colony formation assay
Two thousand cells were seeded in a 6-well plate. After 10–14 days, the cell colonies were fixed in 4% polyformaldehyde and stained with 0.1% crystal violet for 15 min. After staining, the colonies were washed with tap water to remove excessive crystal violet. Images were taken by a camera, and the number of colonies was quantified by ImageJ (NIH).
Wound healing assay
NB cells were seeded in 6-well plates at a density of 5 × 104 cells/well with scratch plug-in components. The cells were cultured with the culture medium containing 0.5% FBS in the incubator overnight. The scratch plug-in components were taken out the next day, and an even wound formed then. Images of samples were captured at appropriate time with the microscope (Leica, USA) at ×20 magnification. In order to exclude the effect of cell proliferation in wound healing assay, the CCK-8 assay for cell proliferation was performed among indicated cells with the same condition used in wound healing assay, and the results indicated that there was no significant statistical difference among indicated cells, confirming that changes in cell migration ability were independent of cell proliferation (Supplementary Fig. 1).
Transwell assay
A density at 1 × 105 cells/well of NB cells was seeded in 24-well plates in 200 μL FBS-free culture medium, and was then plated into the upper chamber with 8 μm pores. Accordingly, 650 μL medium containing 10% FBS was added into the lower chamber. After continuously incubated for about 24 h, the migrated cells were fixed by 4% paraformaldehyde for 20 min and followed by stained with crystal violet for 10 min. Images of the migrated cells were captured with microscopy (Leica, USA). For each well, 5 randomized fields within the view were selected for imaging, and the numbers were counted.
Apoptosis analysis
Cell apoptosis of NB cells was analyzed by flow cytometry using the Annexin V-Alexa Fluor 647/7-AAD Kit (4A Biotech, China) according to the manufacturer’s instructions. In brief, 5 × 105 of indicated NB cells were prepared, and incubated with 5 μL Annexin V-Alexa Fluor 647 for 15 min in the dark, then 10 μL of 7-AAD was added into each tube, immediately analyzed with the flow cytometry (SP6800, Sony, Japan) within 1 h.
Cell cycle analysis
The cell cycle phases of NB cells were analyzed by flow cytometry using the PI Cell Cycle Detection Kit (4A Biotech, China). Briefly, cells with the indicated treatment were fixed in 70% ethanol at 4 °C overnight. Then cells were washed and stained with 50 ug/mL propidium iodide (PI) in solution of 2 mg/mL RNase [55, 56]. Flow cytometry (SP6800, Sony, Japan) was performed on the cells. Modfit software was used for the cell cycle distribution analysis.
Immunofluorescence (IF) staining and imaging
NB cells were prepared in the glass-bottom cell culture dishes (NEST, Wuxi NEST Biotechnology Co., Ltd., China) and fixed with 4% paraformaldehyde for 20 min at room temperature. Then, cells were permeabilized with 0.1% Triton X-100 for 5 min following by blocked in 5% BSA for 1 h. The primary anti-SUV39H1 (Santa Cruz Biotechnology, USA) antibody was used and incubated with cells for 1 h at room temperature. NB cells were washed with PBS and incubated with Alexa Fluor 594-conjugated secondary antibodies (FuDe, China) for 1 h in the dark. DAPI (Beyotime, Shanghai, China) was added and incubated for 5 min to stain the cell nuclei. The confocal microscope (LSM880, Carl Zeiss, Oberkochen, Germany) was used for image acquisition.
Xenograft tumorigenesis in vivo
The NOD/SCID mice aged 3–4 weeks were purchased from Vital River Laboratory Animal Technology (Beijing, China). A mixture of SK-N-BE (2) cells (8 × 106 cells/mice) and thawed Matrigel (Corning, USA) (at a ratio of 3:1) was subcutaneously injected into the right axilla area of the mice [55, 57]. When the tumor volume reached 100 mm3 (tumor volumes = short diameter*short diameter*long diameter/2), the mice were divided randomly into the vehicle group and the treatment group. The treatment group was given 100 uL of chaetocin (0.5 mg/kg) intraperitoneally every 2 days. Body weight and tumor volumes of mice were recorded every other day. Any mice with tumor reached the maximum permitted diameter of 2 cm was regarded as the humane endpoint. After euthanized, the subcutaneous tumors of the mice were isolated and weighed immediately. The in vivo experiments were performed with the approval of the Ethics Committee of Sun Yat-sen University. Investigators were not blinded in this experiment procedure.
RNA sequencing and data analysis
Biological triplicate RNA-seq analysis was performed on NB cells of SUV39H1 inhibition: siNC-NB cells (3 samples), siSUV39H1-NB cells (3 samples), DMSO-NB cells (3 samples), and chaetocin-NB cells (50 nM, 3 samples). Total RNA was extracted with TRIzol reagents (Ambion, Life Technologies, USA). Then, cDNA library building and RNA sequencing (RNA-seq) were carried out via a commercially available service [58] (Huada Gene, Wuhan, China). RNA-seq reads were aligned to the hg38 genome, using |log2FC| ≥ 0.5 and FDR ≤ 0.01 as the cutoff criteria to identify the DEGs. Data can be accessed via GSE254118.
siRNA transfection
Human siRNA oligos were purchased from Ribo (Shanghai, China). For transfection, NB cells were seeded onto 6-well plates and transfected with 20 nM SUV39H1(siSUV39H1) or scrambled control (siNC) using Lipofectamine 3000 (Invitrogen, USA) according to the manufacturer’s instructions. After 24 h, the RNA was harvested and analyzed by qRT-PCR to verify the transfect efficiency. The transfected cells were cultured for up to 48 h for further experiment studies.
Lentiviral transduction of NB cells
For the stably knockdown of SUV39H1, the SUV39H1 and control shRNA plasmids (Cat#:LPP-HSH062782-LVRU6GH-200) were purchased from GeneCopoeia (Guangzhou, China). For the stably overexpression of SUV39H1, MCPIP1 and AURKA, the pCDH-SUV39H1-3xflag-puro plasmid (Cat#:OENM_001282166), the pcDNA3.1-ZC3H12A-3xFlag (Cat#:OENM_025079) and the HY22766 pCDH-CMV-AURKA-EF1-Puro-3xFlag (Cat#:OENM_198433) plasmids were purchased from DaHong Biosciences (Guangzhou, China). The lentiviral transduction system construction was conducted as the previous description [19, 58], with the core plasmids: packaging plasmid psPAX2: envelope plasmid pMD2.G at a ratio of 10:7.5:3. HEK293T cells were used for the lentiviral particle production. The SK-N-SH and SK-N-BE (2) cells were infected with viral supernatant (diluted at 1:1 with fresh media), followed by puromycin selection at 1 μg/mL for 48 h.
Chromatin immunoprecipitation (ChIP) assays
ChIP assay was performed using a SimpleChiP (R) Kit (Magnetic Beads) (Cell Signaling Technology, USA, 91820s) according to the manufacturer’s protocol. NB cells were seeded in 15 cm cell culture dish and were collected following by fixed with 1% formaldehyde at room temperature for 10 min. Subsequently, glycine was added to stop this reaction, and cells were sonicated to shear the DNA into fragments at 200–1000 bp. Then, sheared DNA was incubated with antibodies against H3K9me3 (Cell Signaling Technology, USA) or IgG control (Santa Cruz Biotechnology, USA, sc-2027). DNA-protein-antibody complexes were incubated with Protein A/G magnetic beads, and were washed sequentially with gradient salt buffer, eluted with 1% SDS/NaHCO3, and finally, enriched DNA fragments were subjected to qPCR analysis. IgG was used as the negative control. The primers used for the ChIP-qPCR assays are listed in Supplementary Table 3.
Statistical analysis
Statistical analyses were performed with SPSS 22.0 software (SPSS, Chicago, IL, USA) by Student’s t test, one or two-way ANOVA, or χ2 test. The variance analyzed in this study was assumed as similar with normal distribution. The Kaplan–Meier survival curves were generated based on a log-rank test to compare the survival rates between different groups. All functional assays were independently repeated at least three times, and the results were expressed as mean ± SEM. p < 0.05 was considered statistically significant.
Data availability
Data of the finding from the present study are available at https://www.researchdata.org.cn/.
References
Zafar A, Wang W, Liu G, Wang X, Xian W, Mckeon F, et al. Molecular targeting therapies for neuroblastoma: progress and challenges. Med Res Rev. 2021;41:961–1021.
Matthay KK, Maris JM, Schleiermacher G, Nakagawara A, Mackall CL, Diller L, et al. Neuroblastoma. Nat Rev Dis Prim. 2016;2:16078.
Gong L, Luo J, Zhang Y, Yang Y, Li S, Fang X, et al. Nasopharyngeal carcinoma cells promote regulatory T cell development and suppressive activity via CD70-CD27 interaction. Nat Commun. 2023;14:1912.
Schulte JH, Lim S, Schramm A, Friedrichs N, Koster J, Versteeg R, et al. Lysine-specific demethylase 1 is strongly expressed in poorly differentiated neuroblastoma: implications for therapy. Cancer Res. 2009;69:2065–71.
Jubierre L, Jimenez C, Rovira E, Soriano A, Sabado C, Gros L, et al. Targeting of epigenetic regulators in neuroblastoma. Exp Mol Med. 2018;50:1–12.
Fetahu IS, Taschner-Mandl S. Neuroblastoma and the epigenome. Cancer Metastasis Rev. 2021;40:173–89.
Bae JS, Han M, Yao C, Chung JH. Chaetocin inhibits IBMX-induced melanogenesis in B16F10 mouse melanoma cells through activation of ERK. Chem Biol Interact. 2016;245:66–71.
Liu X, Guo S, Liu X, Su L. Chaetocin induces endoplasmic reticulum stress response and leads to death receptor 5-dependent apoptosis in human non-small cell lung cancer cells. Apoptosis. 2015;20:1499–507.
Isham CR, Tibodeau JD, Bossou AR, Merchan JR, Bible KC. The anticancer effects of chaetocin are independent of programmed cell death and hypoxia, and are associated with inhibition of endothelial cell proliferation. Br J Cancer. 2012;106:314–23.
Liao X, Fan Y, Hou J, Chen X, Xu X, Yang Y, et al. Identification of chaetocin as a potent non-ROS-mediated anticancer drug candidate for gastric cancer. J Cancer. 2019;10:3678–90.
Maleszewska M, Steranka A, Kaminska B. The effects of selected inhibitors of histone modifying enzyme on C6 glioma cells. Pharm Rep. 2014;66:107–13.
Jiang H, Li Y, Xiang X, Tang Z, Liu K, Su Q, et al. Chaetocin: a review of its anticancer potentials and mechanisms. Eur J Pharm. 2021;910:174459.
Gou D, Liu R, Shan X, Deng H, Chen C, Xiang J, et al. Gluconeogenic enzyme PCK1 supports S-adenosylmethionine biosynthesis and promotes H3K9me3 modification to suppress hepatocellular carcinoma progression. J Clin Invest. 2023;133:e161713.
Dong C, Wu Y, Wang Y, Wang C, Kang T, Rychahou PG, et al. Interaction with Suv39H1 is critical for Snail-mediated E-cadherin repression in breast cancer. Oncogene. 2013;32:1351–62.
Lu C, Klement JD, Yang D, Albers T, Lebedyeva IO, Waller JL, et al. SUV39H1 regulates human colon carcinoma apoptosis and cell cycle to promote tumor growth. Cancer Lett. 2020;476:87–96.
Yang Y, Liu R, Qiu R, Zheng Y, Huang W, Hu H, et al. CRL4B promotes tumorigenesis by coordinating with SUV39H1/HP1/DNMT3A in DNA methylation-based epigenetic silencing. Oncogene. 2015;34:104–18.
Zhang L, Tian S, Zhao M, Yang T, Quan S, Yang Q, et al. SUV39H1-DNMT3A-mediated epigenetic regulation of Tim-3 and galectin-9 in the cervical cancer. Cancer Cell Int. 2020;20:325.
Xin DE, Liao Y, Rao R, Ogurek S, Sengupta S, Xin M, et al. Chaetocin-mediated SUV39H1 inhibition targets stemness and oncogenic networks of diffuse midline gliomas and synergizes with ONC201. Neuro Oncol. 2024;26:735–748.
Li M, Sun C, Bu X, Que Y, Zhang L, Zhang Y, et al. ISL1 promoted tumorigenesis and EMT via Aurora kinase A-induced activation of PI3K/AKT signaling pathway in neuroblastoma. Cell Death Dis. 2021;12:620.
Veschi V, Liu Z, Voss TC, Ozbun L, Gryder B, Yan C, et al. Epigenetic siRNA and chemical screens identify SETD8 inhibition as a therapeutic strategy for p53 activation in high-risk neuroblastoma. Cancer Cell. 2017;31:50–63.
Padeken J, Methot SP, Gasser SM. Establishment of H3K9-methylated heterochromatin and its functions in tissue differentiation and maintenance. Nat Rev Mol Cell Biol. 2022;23:623–40.
Zhou Y, Zhou B, Pache L, Chang M, Khodabakhshi AH, Tanaseichuk O, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun. 2019;10:1523.
Park SH, Yu SE, Chai YG, Jang YK. CDK2-dependent phosphorylation of Suv39H1 is involved in control of heterochromatin replication during cell cycle progression. Nucleic Acids Res. 2014;42:6196–207.
Qi Y, Wang D, Huang W, Wang B, Huang D, Xiong F, et al. CyclinD1 inhibits dicer and crucial miRNA expression by chromatin modification to promote the progression of intrahepatic cholangiocarcinoma. J Exp Clin Cancer Res. 2019;38:413.
Bian E, Chen X, Xu Y, Ji X, Cheng M, Wang H, et al. A central role for MeCP2 in the epigenetic repression of miR-200c during epithelial-to-mesenchymal transition of glioma. J Exp Clin Cancer Res. 2019;38:366.
Szukala W, Lichawska-Cieslar A, Pietrzycka R, Kulecka M, Rumienczyk I, Mikula M, et al. Loss of epidermal MCPIP1 is associated with aggressive squamous cell carcinoma. J Exp Clin Cancer Res. 2021;40:391.
Boratyn E, Nowak I, Karnas E, Ryszawy D, Wnuk D, Polus A, et al. MCPIP1 overexpression in human neuroblastoma cell lines causes cell-cycle arrest by G1/S checkpoint block. J Cell Biochem. 2020;121:3406–25.
Miekus K, Kotlinowski J, Lichawska-Cieslar A, Rys J, Jura J. Activity of MCPIP1 RNase in tumor associated processes. J Exp Clin Cancer Res. 2019;38:421.
Nowak I, Boratyn E, Student S, Bernhart SF, Fallmann J, Durbas M, et al. MCPIP1 ribonuclease can bind and cleave AURKA mRNA in MYCN-amplified neuroblastoma cells. RNA Biol. 2021;18:144–56.
Waldeck K, Cullinane C, Ardley K, Shortt J, Martin B, Tothill RW, et al. Long term, continuous exposure to panobinostat induces terminal differentiation and long term survival in the TH-MYCN neuroblastoma mouse model. Int J Cancer. 2016;139:194–204.
Xiao L, Somers K, Murray J, Pandher R, Karsa M, Ronca E, et al. Dual targeting of chromatin stability by the curaxin CBL0137 and histone deacetylase inhibitor panobinostat shows significant preclinical efficacy in neuroblastoma. Clin Cancer Res. 2021;27:4338–52.
Tang Y, Zhang J, Hu Z, Xu W, Xu P, Ma Y, et al. PRKAA1 induces aberrant mitophagy in a PINK1/Parkin-dependent manner, contributing to fluoride-induced developmental neurotoxicity. Ecotoxicol Environ Saf. 2023;255:114772.
Goldstein MN, Plurad S. Drug-induced differentiation of human neuroblastoma: transformation into ganglion cells with mitomycin-C. Results Probl Cell Differ. 1980;11:259–64.
Lee JH, Kim KT. Regulation of cyclin-dependent kinase 5 and p53 by ERK1/2 pathway in the DNA damage-induced neuronal death. J Cell Physiol. 2007;210:784–97.
Wang J, Yin X, He W, Xue W, Zhang J, Huang Y. SUV39H1 deficiency suppresses clear cell renal cell carcinoma growth by inducing ferroptosis. Acta Pharm Sin B. 2021;11:406–19.
Jain N, Zhao Z, Koche RP, Antelope C, Gozlan Y, Montalbano A, et al. Disruption of SUV39H1-mediated H3K9 methylation sustains CAR T-cell function. Cancer Discov. 2024;14:142–57.
Lopez-Cobo S, Fuentealba JR, Gueguen P, Bonte PE, Tsalkitzi K, Chacon I, et al. SUV39H1 ablation enhances long-term CAR T function in solid tumors. Cancer Discov. 2024;14:120–41.
Kong M, Zhou J, Kang A, Kuai Y, Xu H, Li M, et al. Histone methyltransferase Suv39h1 regulates hepatic stellate cell activation and is targetable in liver fibrosis. Gut. 2024;73:810–24.
Xin DE, Liao Y, Rao R, Ogurek S, Sengupta S, Xin M, et al. Chaetocin-mediated SUV39H1 inhibition targets stemness and oncogenic networks of diffuse midline gliomas and synergizes with ONC201. Neuro Oncol. 2024;26:735–48.
Allan RS, Zueva E, Cammas F, Schreiber HA, Masson V, Belz GT, et al. An epigenetic silencing pathway controlling T helper 2 cell lineage commitment. Nature. 2012;487:249–53.
Pace L, Goudot C, Zueva E, Gueguen P, Burgdorf N, Waterfall JJ, et al. The epigenetic control of stemness in CD8(+) T cell fate commitment. Science. 2018;359:177–86.
Shen JZ, Qiu Z, Wu Q, Finlay D, Garcia G, Sun D, et al. FBXO44 promotes DNA replication-coupled repetitive element silencing in cancer cells. Cell. 2021;184:352–69.
Jain N, Zhao Z, Koche RP, Antelope C, Gozlan Y, Montalbano A, et al. Disruption of SUV39H1-mediated H3K9 methylation sustains CAR T cell function. Cancer Discov. 2024;14:142–157.
Fang Y, Zhang MC, He Y, Li C, Fang H, Xu PP, et al. Human endogenous retroviruses as epigenetic therapeutic targets in TP53-mutated diffuse large B-cell lymphoma. Signal Transduct Target Ther. 2023;8:381.
Chu Y, Chen Y, Guo H, Li M, Wang B, Shi D, et al. SUV39H1 regulates the progression of MLL-AF9-induced acute myeloid leukemia. Oncogene. 2020;39:7239.
Du R, Huang C, Liu K, Li X, Dong Z. Targeting AURKA in cancer: molecular mechanisms and opportunities for cancer therapy. Mol Cancer. 2021;20:15.
Dahlhaus M, Burkovski A, Hertwig F, Mussel C, Volland R, Fischer M, et al. Boolean modeling identifies Greatwall/MASTL as an important regulator in the AURKA network of neuroblastoma. Cancer Lett. 2016;371:79–89.
Greiner D, Bonaldi T, Eskeland R, Roemer E, Imhof A. Identification of a specific inhibitor of the histone methyltransferase SU(VAR)3-9. Nat Chem Biol. 2005;1:143.
Tibodeau JD, Benson LM, Isham CR, Owen WG, Bible KC. The anticancer agent chaetocin is a competitive substrate and inhibitor of thioredoxin reductase. Antioxid Redox Signal. 2009;11:1097–106.
Cherblanc FL, Chapman KL, Brown R, Fuchter MJ. Chaetocin is a nonspecific inhibitor of histone lysine methyltransferases. Nat Chem Biol. 2013;9:136–7.
Reece KM, Richardson ED, Cook KM, Campbell TJ, Pisle ST, Holly AJ, et al. Epidithiodiketopiperazines (ETPs) exhibit in vitro antiangiogenic and in vivo antitumor activity by disrupting the HIF-1alpha/p300 complex in a preclinical model of prostate cancer. Mol Cancer. 2014;13:91.
Zhang S, Zhao Y, Wang S, Li M, Xu Y, Ran J, et al. Discovery of novel diarylamides as orally active diuretics targeting urea transporters. Acta Pharm Sin B. 2021;11:181–202.
Lee JY, Jonus HC, Sadanand A, Branella GM, Maximov V, Suttapitugsakul S, et al. Identification and targeting of protein tyrosine kinase 7 (PTK7) as an immunotherapy candidate for neuroblastoma. Cell Rep Med. 2023;4:101091.
Bosse KR, Raman P, Zhu Z, Lane M, Martinez D, Heitzeneder S, et al. Identification of GPC2 as an oncoprotein and candidate immunotherapeutic target in high-risk neuroblastoma. Cancer Cell. 2017;32:295–309.
Yang Z, Wang T, Wu D, Min Z, Tan J, Yu B. RNA N6-methyladenosine reader IGF2BP3 regulates cell cycle and angiogenesis in colon cancer. J Exp Clin Cancer Res. 2020;39:203.
Boyanapalli SS, Li W, Fuentes F, Guo Y, Ramirez CN, Gonzalez XP, et al. Epigenetic reactivation of RASSF1A by phenethyl isothiocyanate (PEITC) and promotion of apoptosis in LNCaP cells. Pharm Res. 2016;114:175–84.
Tian X, Zhou D, Chen L, Tian Y, Zhong B, Cao Y, et al. Polo-like kinase 4 mediates epithelial-mesenchymal transition in neuroblastoma via PI3K/Akt signaling pathway. Cell Death Dis. 2018;9:54.
Chen B, Chen H, Lu S, Zhu X, Que Y, Zhang Y, et al. KDM5B promotes tumorigenesis of Ewing sarcoma via FBXW7/CCNE1 axis. Cell Death Dis. 2022;13:354.
Acknowledgements
This study was supported by grants from the National Key Research and Development Program of China (No. 2022YFC2705005), the National Natural Science Foundation of China (No. 82303160; 82203303; 82302940), the Basic and Applied Basic Research Foundation of Guangdong Province (No. 2021A1515110234), the Medical Science and Technology Research Fund Project of Guangdong Province (No. A2022079), and the Medical Scientific Research Foundation of Guangdong Province of China (No. A2023072).
Author information
Authors and Affiliations
Contributions
MZL: editing conception, design, experiment conduction, verification of the underlying data, data analysis and curation, methodology, investigation, writing and editing. FFS and JW: editing conception, design, verification of the underlying data, methodology, investigation, validation and editing. SYL, YQ, MJS, HMC, XYX, WJX, JZ and JTH: formal analysis, methodology, investigation, formal analysis and visualization. YZ: editing conception, design, validation, supervision, funding acquisition and resources, writing and editing. YZZ: editing conception, design, supervision, funding acquisition and resources, editing. All authors have read and approved the submission of this manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All methods used in the present study were in accordance with the relevant guidelines and regulations. This study was approved by the Institutional Review Board (IRB) of the Sun Yat‑sen University Cancer Center (Approval no. B2020‑101‑Y01). Informed consent was obtained from the parents of the participants. All the animal experiments in vivo were conducted consist with the Animal Care and Use Committee of the Sun Yat‑sen University Cancer Center, which were approved by the Animal Ethics Committee of Sun Yat‑sen University (Approval no. SYSU-IACUC-2023-000004).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, M., Sun, F., Wang, J. et al. SUV39H1 epigenetically modulates the MCPIP1-AURKA signaling axis to enhance neuroblastoma tumorigenesis. Oncogene (2024). https://doi.org/10.1038/s41388-024-03164-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41388-024-03164-4
- Springer Nature Limited