Abstract
UTX (also known as KDM6A), a histone 3 lysine 27 demethylase, is among the most frequently mutated epigenetic regulators in myelodysplastic syndrome (MDS) and acute myeloid leukemia (AML). Recent studies have suggested that UTX mutations promote MDS and AML by blocking the differentiation of hematopoietic stem and progenitor cells (HSPCs). Here, we performed an epigenetic drug library screening for small molecules able to release the differentiation block on HSPCs induced by UTX deficiency. We found that SP2509, a selective inhibitor of LSD1, specifically promoted the differentiation of Utx-null HSPCs while sparing wild-type HSPCs. Transcriptome profiling showed that Utx loss reduced the expression of differentiation-related and tumor suppressor genes, correlating with their potential roles in HSPC self-renewal and leukemogenesis. In contrast, SP2509 treatment reversed these changes in gene expression in Utx-null HSPCs. Accordingly, Utx loss decreased H3K4 methylation level probably through the COMPASS-like complex, while LSD1 inhibition by SP2509 partially reversed the reduction of H3K4 methylation in Utx-deficient HSPCs. Further, SP2509 promoted the differentiation of Utx-null AML cells in vitro and in vivo and, therefore, extended the survival of these leukemic mice. Thus, our study identified a novel strategy to specifically target both premalignant and malignant cells with Utx deficiency for differentiation therapy and provided insights into the molecular mechanisms underlying the role of Utx in regulating HSPCs and related diseases.
Similar content being viewed by others
Introduction
Genes encoding epigenetic modifiers are among the most frequently mutated in hematopoietic malignancies.1,2 It has been suggested that dysfunctions in these genes, such as TET2, DNMT3A, and IDH1/2, promote leukemogenesis by blocking the differentiation of hematopoietic stem and progenitor cells (HSPCs).3,4,5,6,7 Our previous studies identified Mixed Lineage Leukemia 3 (MLL3, officially known as KMT2C) on chromosome 7q as a tumor suppressor gene.8 Haploinsufficiency of Mll3 impairs HSPC differentiation and leads to (myelodysplastic syndrome) MDS-like phenotypes. Mll3 (or Mll4), H3K4 mono- and dimethyltransferase, forms the COMPASS-like complex with an H3K27 demethylase Utx (Ubiquitously transcribed tetratricopeptide repeat on chromosome X, officially named as KDM6A) and other components.9,10,11,12 Therefore, the COMPASS-like complex can regulate target gene expressions by simultaneously modifying both H3K4 and H3K27.13,14
Interestingly, UTX is highly frequently mutated in human blood malignancies and solid cancers.15,16,17,18,19,20 For example, UTX is mutated in approximately 10% chronic myelomonocytic leukemia (CMML) and CMML-derived acute myeloid leukemia (AML) patients and 20% acute lymphoblastic leukemia (ALL) patients.5,14,21 UTX is essential for multiple biological processes including stem cell self-renewal, embryogenesis, and posterior development.22,23,24,25 In cancer biology, UTX is considered as a tumor suppressor gene of various cancers. UTX knockout has been reported to promote pancreatic or lung cancer development.26,27 Also, its deficiency could lead to both myeloid and lymphoid malignancies.28,29,30,31 It is known that UTX is required for hematopoiesis. Without it, the frequency and absolute number of both LT-HSCs and ST-HSCs increases due to the impaired cell differentiation.5,31
Differentiation therapy has been approved as being effective for hematopoietic malignancies, such as all-trans-retinoic acid and arsenic trioxide for acute promyelocytic leukemia.32,33,34 Given the critical role of UTX in the regulation of HSPCs, we hypothesized that releasing the differentiation block due to UTX deficiencies could benefit patients with UTX mutations. To explore potential therapeutic strategies for malignancies with UTX deficiencies, we performed an epigenetic drug library screening. We found that SP2509, a putative LSD1 inhibitor,35 specifically promoted the differentiation of UTX knockout HSPCs while sparing wild-type HSPCs. Mechanistically, UTX, likely through the COMPASS-like complex, facilitates the expression of differentiation-related genes and tumor suppressors associated with increased H3K4 methylation. LSD1 is an H3K4 demethylase.36 LSD1 inhibition restores the balance in the H3K4 methylation of target genes in UTX-mutant cells and releases them from differentiation block. Intriguingly, SP2509 had similar effects on UTX-null AML cells and extended the lifespan of animals with UTX-deficient-driven leukemia.
Results
Epigenetic drug library screening identified candidate small molecules for specifically promoting the differentiation of Utx-null HSPCs
Given the functions of the tumor suppressor Utx in the differentiation of HSPCs through the modifications of histone 3, we decided to perform a high-throughput functional screening of an epigenetic drug library in wild-type (WT) and Utx-null HSPCs (Fig. 1a). We hypothesized that candidate small molecules could reverse the epigenetic abnormalities resulting from Utx loss and specifically release the differentiation block on Utx-deficient HSPCs. cKit+ HSPCs were isolated using MACS beads from wild-type (WT) or Utxf/f;Mx1-cre (KO) C57BL/6 mice37,38 7 days after pIpC treatment. The epigenetic drug library has 276 compounds including a majority of currently available agonists and antagonists of epigenetic regulatory enzymes (Fig. 1b). The working concentrations of these drugs were 10 μM.
Epigenetic drug library screening for compounds specifically promoting the differentiation of Utx-null HSPCs. a Schematic showing the experimental design of epigenetic drug screening: HSPCs were isolated from young adult Utxf/f (WT) or Utxf/f; Mx-1-cre (KO) C57BL/6 mice and then treated with compounds from an epigenetic drug library (Selleckchem, 276 compounds in total) individually. After 72-h treatment, cells were analyzed by flow cytometry with cKit and Mac-1 antibodies. b Summary of drug classification in the epigenetic drug library. The library consists of compounds directly targeting epigenetic enzymes and other related factors. c, d Relative mean fluorescence intensities (MFIs) of Mac-1 (c) and of cKit staining (d), measured by flow cytometry, in Utx WT and KO HSPCs treated with individual compounds from the epigenetic drug library or vehicle. e Relative proportions of cKit+ populations in Utx WT and KO HSPCs treated as in c and d
The outputs of the screen were stemness and differentiation status of HSPCs, measured by flow cytometry using cKit and Mac-1 staining, respectively. To identify compounds specifically enhancing the differentiation of Utx KO HSPCs while sparing WT HSPCs, we ranked the compounds by differences in the mean fluorescence intensity (MFI) of Mac-1 or cKit staining between Utx WT and KO HSPCs (Fig. 1c, d). Consistent with previous reports that Utx deficiency blocks HSPC differentiation, vehicle-treated Utx KO HSPCs expressed lower levels of Mac-1, a myeloid cell marker, than WT HSPCs, indicating impaired differentiation in KO cells. In contrast, Utx KO HSPCs expressed higher levels of cKit, a standard marker for HSPCs, than WT HSPCs.
The top three compounds that induced the most pronounced changes in Mac-1 expression between Utx KO and WT HSPCs were BIX01294, Bisindolylmaleimide IX, and SP2509 (Fig. 1c). BIX01294 is an inhibitor of histone methyltransferase G9a, and it therefore reduces H3K9m2 levels at an IC50 of 2.7 μM.39 Bisindolylmaleimide IX is a pan-PKC inhibitor, which can inhibit PKC-α, PKC-βI, PKC-βII, PKC-γ, and PKC-ε with low nanomolar IC50.40 SP2509 is a selective inhibitor of the histone demethylase LSD1 with an IC50 of 3 nM.35,41 On the other hand, the top three compounds that mostly reduced cKit expression in Utx KO HSPCs compared to that in WT HSPCs were azacytidine, decitabine, and SP2509 (Fig. 1d). Azacytidine and decitabine are already used in patients with various hematopoietic malignancies, and they induce differentiation of or directly kill leukemic blasts.42,43 The identification of azacytidine and decitabine validated our screening. However, we were more interested in the less studied SP2509, especially because it was the only compound identified to affect both Mac-1 and cKit expression. Actually, SP2509 was also the only candidate identified when the compounds were ranked by different proportions of cKit+ populations between Utx KO and WT HSPCs (Fig. 1e). Therefore, we focused on SP2509 in the subsequent studies.
Validating SP2509 as a specific inducer of differentiation in Utx-deficient cells
To validate the function of SP2509 on Utx KO HSPCs, we performed another independent experiment similar to the library screening (Fig. 1a). First, in vehicle-treated group, Utx KO HSPCs remained more cKit+ cells than that in Utx WT HSPCs (32.4% vs 23.4%; Fig. 2a), while less Mac-1+ cells in Utx KO HSPCs than that in WT HSPCs (56.2% vs 76.3%; Fig. 2b), indicating that Utx loss impaired the differentiation of HSPCs. With SP2509 treatment, Utx WT HSPCs did not have much changes compared to vehicle-treated group. Twenty-six percent of SP2509-treated Utx WT HSPCs remained cKit+, whereas 69.7% became Mac-1+ (Fig. 2c, d). In striking contrast, only 9.38% of Utx KO HSPCs remained cKit+ after SP2509 treatment, which was approximately 2.5-fold lower than that in vehicle-treated KO HSPCs and significantly less than that in WT HSPCs (Fig. 2e, f). Conversely, 90% of Utx KO HSPCs became Mac-1+ with SP2509 treatment, which was approximately one-fold higher than that in vehicle-treated Utx KO HSPCs and significantly more than that in WT HSPCs (Fig. 2g, h). Consistently, the MFI of cKit staining in Utx KO HSPCs decreased from 2489 for vehicle treatment to 681 for SP2509 treatment, a 2.65-fold reduction, while there was no significant difference between vehicle- and SP2509-treated WT HSPCs (Fig. 2i, j). The MFI of Mac-1 staining in Utx KO HSPCs increased from 1450 for vehicle treatment to 16,326 for SP2509 treatment, a 10.2-fold increase, which was dramatically more than the difference between vehicle- and SP2509-treated Utx WT HSPCs (Fig. 2k, l). Therefore, SP2509 was validated as a specific differentiation inducer for Utx KO HSPCs while sparing WT HSPCs.
Validation of SP2509 in specifically inducing the differentiation of Utx KO HSPCs. a, b Representative flow plots showing cKit (a) and Mac-1 (b) staining in vehicle-treated Utx WT or KO HSPCs from three independent experiments. c, d Flow plots showing cKit (c) and Mac-1 (d) staining in SP2509-treated Utx WT and KO HSPCs. e–h Percentage of cKit+ (e) or Mac-1+ (g) populations in Utx WT and KO HSPCs treated with vehicle or SP2509. Corresponding fold changes of SP2509 vs vehicle-treated Utx WT or KO HSPCs are shown in f (cKit) and h (Mac-1). n = 3. i–l MFI of cKit (i) or Mac-1 (k) staining in Utx WT and KO HSPCs treated with vehicle or SP2509. Corresponding fold changes are shown in j (cKit) and l (Mac-1). n = 3. Error bars represent s.d. *p < 0.05; **p < 0.01; ***p < 0.001
SP2509 treatment reversed the global effect of Utx deficiency on the transcriptome of HSPCs
To understand the molecular mechanisms underlying the SP2509-induced differentiation of Utx KO HSPCs, we used RNA sequencing (RNA-seq) to analyze the transcriptomes of Utx WT and KO HSPCs treated with vehicle or SP2509. Cells were harvested after 12-h treatment to capture the direct effects of SP2509 treatment on gene expression. In each group, there were three independent treated samples. Unsupervised grouping analysis showed that all of the three samples were grouped together in each group (Fig. 3a), indicating a high quality of the RNA-seq data. Principal component analysis (PCA) also revealed tight grouping of the three replicates in each group. The location of Utx WT and KO HSPCs treated with vehicle or SP2509 on the PCA plot clearly demonstrated the differences between Utx WT and KO HSPCs and those between vehicle- and SP2509-treated HSPCs (Fig. 3b).
SP2509 treatment reversed the global effect of Utx deficiency on the transcriptome of HSPCs. a, b Unsupervised clustering (a) and principal component analysis (PCA) (b) of RNA-seq data from Utx WT and KO HSPCs treated with vehicle or SP2509 for 12 h. c Heatmap of differentially expressed genes in Utx WT or KO HSPCs measured by RNA-seq. d Heatmap of differentially expressed genes in Utx KO HSPCs treated with vehicle or SP2509 for 12 h
Interestingly, compared to the differentially expressed genes in WT HSPCs, 177 genes were significantly downregulated in Utx KO HSPCs (p < 0.05, log2[KO/WT] < −1), while only 34 genes were significantly upregulated (Fig. 3c). The overwhelming repression of gene expression is consistent with the histone modification functions of Utx and its associated COMPASS-like complex. Similar results were also observed in Mll3 (the core component of COMPASS-like complex)-deficient HSPCs.8
In striking contrast, 99 genes in SP2509-treated Utx KO HSPCs were significantly upregulated (p < 0.05, log2[KO/WT] < −1) compared to those in vehicle-treated Utx KO HSPCs, while only nine genes were significantly downregulated (Fig. 3d). Notably, the overwhelming upregulation of gene expression by SP2509 was directly contradictory to the effect of Utx loss in HSPCs. Accordingly, the effects of SP2509 treatment on the global transcriptome was specific to Utx KO HSPCs, and the up- and downregulated genes in SP2509-treated vs vehicle-treated Utx WT HSPCs were largely balanced.
SP2509 enhanced the expression of differentiation-related genes and tumor suppressors repressed in Utx KO HSPCs
To understand the functions of SP2509-regulated genes, we performed gene set enrichment analysis (GSEA). Consistent with the differentiation block due to Utx deficiency44 (Fig. 2a–d), hematopoietic cell lineage-related pathways and multiple immune-related pathways were the most enriched pathways among the downregulated genes in Utx KO HSPCs. Transcriptional signatures produced by Utx KO (compared to Utx WT) were negatively correlated with the RPS14 pathway45-associated gene signature (normalized enrichment score [NES] = −1.717, false discovery rate [FDR] = 0.004). Nevertheless, the transcriptional signatures in response to SP2509 treatment were positively correlation with the RPS14 pathway ([NES] = 1.694, [FDR] = 0.006) (Fig. 4a). Similarly, the hallmark IL2/STAT5 pathway-related gene set was negatively enriched in Utx KO HSPCs ([NES] = −1.561, [FDR] = 0.017). In turn, this gene set was positively enriched in Utx KO HSPCs treated with SP2905 ([NES] = 1.410, [FDR] = 0.170) (Fig. 4b). Further, the transcriptional signature of Utx KO HSPCs was negatively enriched in genes involved in the inflammatory response pathway (NES = −1.615, FDR q = 0.013), while the Bmi1_DN.V1_UP46 gene set, related to stemness, was enriched in SP2590-treated Utx KO HSPCs (NES = 1.651, FDR q = 0.010). Hence, Utx deficiency inhibited the expression of differentiation-related genes, which was reversed by SP2509 treatment.
Utx deficiency repressed the expression of differentiation-related genes and tumor suppressors, while SP2509 treatment reversed this inhibition. a–f Gene set enrichment analysis (GSEA) of genes in the RPS14_DN. V1_UP gene set (a), the Hallmark_IL2_STAT5_signaling gene set (b), the GO_Inflammatory_Response and the BMI1_DN. V1_UP gene set (c), the p53_DN. V1_DN, and the Hallmark_p53_pathway gene set (d), the Hallmark_Myc_Targets_V1 gene set (e), the GO_Ribosomal_Subunit and the GO_Ribosome_Biogenesis gene set (f) in Utx KO HSPCs compared to that in Utx WT HSPCs, and Utx KO HSPCs treated with SP2509 compared to that in cells treated with vehicle (Veh). g Relative mRNA expression level of Car1, Car2, Gata1, Gata2, Fabp4, and Arg1 in Utx WT or KO HSPCs, measured by qPCR. n = 3. h Relative mRNA expression of Fabp4, Igfbp7, Jag1, Arg1, Epha2, and Bax in Utx KO HSPCs treated with vehicle (Veh) or SP2509. n = 3. Error bars represent s.d. *p < 0.05; **p < 0.01; ***p < 0.001
Interestingly, the transcriptional signature produced by Utx KO was negatively correlated with the tumor suppressor p53 pathway-related gene set ([NES] = −1.600, [FDR] = 0.017) and positively correlated with the Myc pathway-related gene set ([NES] = 1.876, [FDR] = 0.006), which might explain the functions of Utx in numerous human cancers.27,30,31,44,47 In contrast, the transcriptome in SP2509-treated KO HSPCs was significantly positively enriched in p53 pathway-related genes ([NES] = 1.859, [FDR] = 0.000) and negatively correlated with Myc pathway-related genes ([NES] = −2.490, [FDR] = 0.000) (Fig. 4e, f). Additionally, the ribosome biogenesis-related gene set was positively enriched in Utx KO HSPCs ([NES] = 1.90, [FDR] = 0.12). Conversely, this gene set was negatively enriched in SP2590-treated Utx KO HSPCs ([NES] = 1.90, [FDR] = −2.218) (Fig. 4f). Thus, SP2509 treatment rescued the expression of tumor suppressors and oncogenes perturbed by Utx deficiency in HSPCs.
Quantitative PCR analysis of selected differentiation-related genes, such as Car1, Car2, Gata1, and Gata2, from GSEA in Utx KO HSPCs confirmed significant decreased expression compared to that in Utx WT HSPCs (p < 0.01) (Fig. 4g). Accordingly, SP2509-treated Utx KO HSPCs displayed significantly increased expression of selected differentiation-related genes and tumor suppressors including Bax compared to vehicle-treated Utx KO HSPCs (p < 0.05) (Fig. 4h).
SP2509 treatment reversed the H3K4 methylation-mediated repression of differentiation-related genes and tumor suppressors in Utx KO HSPCs by inhibiting LSD1
Given the H3K4 methylation activity of Utx and its associated COMPASS-like complex and the H3K4 demethylation activity of LSD1, the putative target of SP2509,48 we further studied the mechanisms of SP2509-induced differentiation in Utx KO HSPCs. As shown in western blot (Fig. 5a, b), the total H3K4m3 level was reduced in Utx KO HSPCs compared to that in Utx WT HSPCs. Although Utx itself does not have a direct H3K4-modifying activity, it associates with Mll3/4 in COMPASS-like complex, which are H3K4 mono- and dimethyltransferase. Actually, we did observe a significant reduction in H3K4m1 levels, but the levels of H3K4m2 were similar between Utx KO and WT HSPCs. Consistently, differentiation-related genes, such as Car1, Car2 and Gata1, and the tumor suppressor Bax, displayed reduced H3K4m1 levels in Utx KO HSPCs compared to those in WT HSPCs44 (Fig. 5c–f), which led to the decreased expression levels of these genes (Fig. 4h). Our results suggested that Utx loss might impair the H3K4 methyltransferase activities of Mll3/4.
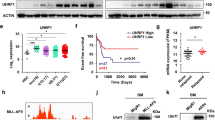
SP2509 treatment reversed Utx deficiency inducing the histone H3K4 methylation change on globe and specific differentiation-related or tumor suppressor genes in HSPCs. a Western blotting of H3K4m1, H3K4m2, and H3K4m3 in cell lysates from Utx WT or KO HSPCs treated with vehicle (V) or SP2509 (SP) for 12 h. b Quantification of H3K4m1, H3K4m2, and H3K4m3 levels in a. c–f H3K4m1 peaks on Car1(c), Car2 (d), Gata1(e), and Bax (f) in Utx WT or KO HSPCs measured by ChIP-seq. Data analyzed from Gozdecka et al.44 g H3K4m3 peaks on Bax in Utx WT and KO ES cells measured by ChIP-seq. Data analyzed from Dhar et al.49 h MLL4 peaks on BAX in vehicle or OG86-treated leukemic cells measured by ChIP-seq. Data analyzed from Maiques-Diaz et al.50
In contrast, LSD1, the target of SP2509, is an H3K4 and H3K9 demethylase. Indeed, LSD1 inhibition by SP2509 increased H3K4m3 levels in Utx KO HSPCs (Fig. 5a, b). The total levels of H3K4m2 were not significantly changed in Utx KO HSPCs treated with SP2509, but the level of H3K4m1 was further reduced, suggesting that a majority of these loci might have been further methylated to H3K4m3. The level of H3K4m3 in Bax gene was reduced in Utx KO ES cells compared to that in Utx WT cells49 (Fig. 5g). Interestingly, treatment with OG86, another LSD1 inhibitor, increased the binding of MLL4 to the promoter of BAX in leukemic cells50 (Fig. 5h), suggesting that LSD1 and the COMPASS-like complex might directly compete for binding on the promoters of differentiation-related genes and tumor suppressors. Thus, LSD1 inhibition reversed the reduction in H3K4 methylation in Utx-deficient cells and increased the expression of differentiation-related genes and tumor suppressors.
SP2509 treatment promoted the differentiation of Utx-deficient AML cells in vitro and in vivo and extended the survival of leukemic mice
We further tested whether SP2509 would have similar inhibitory effect on Utx-deficient malignant cells as it did on Utx KO HSPCs. Utx−/−; shp53-mCherry;shNf1-GFP AML cells (from unpublished independent work) were treated with or without 10 μM SP2509 for 3 days. By flow cytometry, we found that SP2509 treatment dramatically increased the expression of the differentiation marker Mac-1 in leukemia cells (Fig. 6a). The percentage of Mac-1+ population and the MFI of Mac-1 staining in SP2509-treated Utx-null AML cells were approximately two-fold higher than those in vehicle-treated cells. Accordingly, the expression of the stemness marker cKit was reduced by SP2509 (Fig. 6b). Consistent with the release of differentiation block, SP2509 significantly repressed the growth of Utx-null AML cells (Fig. 6c). Thus, similar to its function in premalignant Utx-deficient HSPCs, SP2509 inhibited Utx-null AML cells in vitro by promoting their differentiation. These data validated our screening strategy to target Utx deficiency in malignant cells.
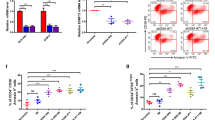
SP2509 inhibited Utx-mutated AML by promoting differentiation in vitro and in vivo. a Flow cytometry measuring protein expression in Utx−/−; shp53-mCherry;shNf1-GFP AML cells treated with vehicle or SP2509. Left, representative flow plots of AML cells showing the expression of Mac-1. Right, percentage of Mac-1+ populations (top), and the MFI of Mac-1 staining (bottom). b MFI of cKit staining in AML cells treated with vehicle or SP2509. c Relative cell number of AML cells treated with vehicle or SP2509. d Schematic showing Utx−/−; shp53-mCherry;shNf1-GFP mice with AML treated with vehicle or SP2509. C57BL/6 mice were sublethally irradiated and then transplanted with Utx−/−; shp53-mCherry; shNf1-GFP AML cells. Once AML was established, these mice were treated with vehicle or 25 mg/kg SP2509 twice per week via i.p. injection. The mice were monitored by flow cytometry, CBC, and blood smear. e, f Representative flow plots showing the expression of Mac-1 in peripheral blood AML cells in vehicle- or SP2509-treated mice after 1 (e) or 2 (f) weeks. g, h Percentage of Mac-1+ (g) and MFI of Mac-1 staining (h) of AML cells in the peripheral blood of vehicle- or SP2509-treated mice. i Tumor burden in the peripheral blood of vehicle- or SP2509-treated mice, measured by the percentage of GFP+mCherry+ populations. j Whole blood cell counts in vehicle- or SP2509-treated mice after 2.5 weeks. k Kaplan–Meier survival curves of mice with AML treated with vehicle or SP2509. n = 6, **p < 0.01. l Blood smear showing leukemic cells in the peripheral blood of vehicle- or SP2509-treated mice
Then, the potential therapeutic effect of SP2509 on Utx-deficient AML was tested in vivo. Utx−/−; shp53-mCherry;shNf1-GFP AML cells were transplanted into sublethally irradiated congenic recipient mice. Seven days after transplantation, all of the recipient mice had similar proportions of GFP+-mCherry+ AML cells in their peripheral blood measured by flow cytometry. These leukemic mice were randomly assigned into two groups: one was treated with 25 mg/kg SP2509 twice per week by intraperitoneal injection, while the other one was treated with vehicle. These mice were monitored for disease progression by complete blood cell (CBC) count, blood smear staining, and flow cytometry (Fig. 6d). One week after treatment, we observed approximately 50% Mac-1+ AML cells in the peripheral blood of SP2509-treated mice compared to 25% in the peripheral blood of vehicle-treated mice (Fig. 6e). This differentiation-promoting effect of SP2509 was more obvious after a 2-week treatment. The majority of AML cells were Mac-1+ in SP2509-treated mice, and the MFI of Mac-1 staining in SP2509-treated leukemic cells was approximately five-fold higher than that in vehicle-treated cells (Fig. 6f–h). We did not observe significant weight loss in mice treated with SP2509, suggesting that SP2509 was well tolerated by the mice (data not shown). These data demonstrated that SP2509 promoted the differentiation of Utx-mutated tumor cells in vivo.
More importantly, SP2509 prevented the progression of AML in vivo. The tumor burden of recipient mice was measured using the percentage of GFP+-mCherry+ cells in peripheral blood. Utx-mutated AML was very aggressive, and leukemic cells progressively proliferated in vehicle-treated mice. The tumor burden in this group increased from 2% to 70% in 2.5 weeks. In contrast, tumor burden increased much slower in SP2509-treated mice, as only 20% AML cells detected in the peripheral blood after 2.5 weeks (Fig. 6i). Interestingly, more leukemic cells in SP2509-treated mouse peripheral blood were observed than that in vehicle-treated mice 1 week after treatment. This caveat might be because of early proliferation associated with differentiation, which we observed within other AML models, such as IDH2-mutated AML.6 After 2.5-week treatment, whole blood cell count in vehicle-treated mice were 25–65k/μL, but those in SP2509-treated mice were largely normal (Fig. 6j). Furthmore, all of the vehicle-treated mice died within 17 days after treatment, while five out of six SP2509-treated mice survived over 3 weeks (Fig. 6k). The surviving mice had few blast cells in their peripheral blood (Fig. 6l). Thus, SP2509 exerted marked therapeutic effects on Utx-mutated tumors in vivo.
In summary, Utx loss reduces H3K4 methylation, presumably through the COMPASS-like complex, and therefore represses the expression of differentiation-related genes and tumor suppressors, which leads to a differentiation block in HSPCs eventually giving rise to tumorigenesis (Fig. 7a). LSD1 competes with MLL3/4 on the promoters of these genes. Thus, the inhibition of LSD1 by SP2509 reverses the reduction in H3K4 methylation in Utx-deficient cells, which gives rise to an increased expression of differentiation-related genes and tumor suppressors and specifically promotes the differentiation of Utx-mutated cells (Fig. 7b).
A working model of the molecular mechanism by which SP2509 induces the differentiation of Utx-deficient cells. a Schematic showing that the H3K4 methylation activity of MLL3/4 in the COMPASS-like complex is impaired by Utx deficiency, which leads to a reduced expression of differentiation-related genes and tumor suppressors. b Schematic showing that in Utx-deficient cells, the small molecule SP2509 inhibits the demethylation activity of LSD1 on H3K4 and therefore rescues the expression of differentiation-related genes and tumor suppressors
Discussion
Accumulating evidence suggests that epigenetic abnormalities are common and critical for human cancers, especially hematopoietic malignancies. Many cancer-associated epigenetic alterations promote tumorigenesis by impairing the differentiation of tissue stem and/or progenitor cells. Thus, differentiation therapy would be especially effective for cancers involving epigenetic dysregulation. Since any given epigenetic modification, including histone methylation, can regulate the expression of a number of targets, the direct reversal of epigenetic abnormalities due to a driver epigenetic modifier would be more efficient than targeting individual targets. The balance between epigenetic writers and erasers provides opportunities to identify therapeutic targets specific for cancers with given epigenetic abnormalities.
UTX, an H3K27 demethylase and a key component of the COMPASS-like complex, is a putative tumor suppressor in multiple human cancers. Our study, as well as previous reports, showed that UTX deficiency impairs the differentiation of HSPCs. In this study, we screened an epigenetic drug library using Utx KO and WT HSPCs. We found that SP2509, an inhibitor of the H3K4 and H3K9 demethylase LSD1, specifically promoted the differentiation of Utx-null HSPCs while sparing Utx WT HSPCs. Based on the effect of SP2509 on Utx-deficient premalignant cells, we further tested it in Utx-mutated tumor cells in vitro and in vivo. Notably, SP2509 had similar differentiation-promoting effect on AML cells, and, more importantly, it inhibited leukemia progression in vivo and significantly extended the lifespan of mice with AML. Mechanistically, Utx mutations repress the expression of differentiation-related genes and tumor suppressors by impairing H3K4 methylation on these genes, while SP2509 inhibits LSD1 and reverses the impairments in H3K4 methylation in Utx-mutated cells. Thus, we identified a highly specific inhibitor for abnormal cells with deficiencies in Utx and/or the COMPASS-like complex. Interestingly, SP2509 and some other LSD1 inhibitors have been suggested to be effective in some leukemias and other human cancers.41,51 SP2577, the clinical formulation of SP2509, is in a phase I clinical trial for patients with relapsed or refractory Ewing sarcoma. Further validation of SP2509 and other inhibitors with similar functions in human cancer cells with UTX mutations or COMPASS-like complex dysfunctions would pave the way for their potential applications for the treatment of patients with such epigenetic abnormalities.
Methods
Mice
Utxf/f mice (purchased from The Jackson Labratory, 024177) were crossed with Mx1-Cre mice (purchased from Jackson Lab, 005673). The expression of Cre was induced by the intraperitoneal injection of pIpC (Sigma P1530, 10 mg/kg) into 5- to 6-week-old mice every other day for a total of 10 days.
Isolation of HSPCs and cell culture
WBM cells were freshly isolated from Utxf/f and Utxf/f; Mx1-Cre mice, and then HSPCs were sorted with mouse CD117 magnetic microbeads (Miltenyi Biotec, 130-091-224). Primary mouse cells were cultured in BCM medium (50% IMDM + 50% DMEM + 10% FBS + 5% PS + 0.34% BME) with 10% Stem Cell Medium (SCM) including 10% FBS, IL3 (10 ng/mL; 1:1000), IL6 (10 ng/mL; 1:500), and SCF (50 ng/mL; 1:1000).
Drug screening
The epigenetics compound library (including 276 compounds) used for drug screening was purchased from Selleckchem (Catalog# L1900). The compounds were generally provided in DMSO at 10 μM concentrations. After HSPCs were isolated from bone marrow and cultured in vitro for 24 h, they were seeded at equal numbers in five low-attachment 96-well plates (the primary number of cells was 40,000 in 100 μL medium per well; in one 96-well plate, 60 wells were filled with cells, while other surrounding wells were filled with PBS). Then, 276 kinds of compounds at a final concentration of 10 μM were used separately (marked in pink), and DMSO was used to treat control cells (marked in gray). After 3 days of treatment, cell features were determined by flow cytometry.
Flow cytometry
Cells were stained with antibodies against Mac-1 (Clone: M1/70, BD101224) and cKit (Clone: 2B8, BD105812) after 3 days of treatment. All flow cytometry analyses were performed with an LSR Fortessa instrument (BD), and the data were analyzed using FlowJo software.
RNA-sequencing analysis
RNA-seq libraries were prepared using NEBNext® Ultra™ RNA Library Prep Kit for Illumina® and were sequenced with an Illumina HiSeq™ X sequencing machine with 150-bp paired-end reads. The RNA-seq reads were aligned to a reference genome (GRCm38) by STAR_2.6.0a.52 Transcript abundance was normalized and measured in fragments per kb of exon per million fragments mapped (FPKM). Differential gene expression was analyzed by DESeq2.53 Genes with absolute fold changes greater than 0.5 and FDR ≤ 0.05 were considered differentially expressed genes. Heatmaps of differentially expressed genes were constructed and normalized by Z scores. Samples distances were calculated by PCA and the Euclidean distance. GSEA54,55 was employed to determine statistically significant similarities and differences between two given clusters by identifying a priori-defined gene sets.
ChIP-seq data collection and visualization
Data were downloaded from the GEO database under accession codes GSE63222,50 GSE101307,44 and GSE76692.49 In addition, data were visualized by Integrative Genomics Viewer56 (IGV).
Western blotting
1M cells were harvested and lysed in SDS buffer (50 mM Tris-HCl (pH 6.8), 2% (w/v) SDS, 150 mM NaCl, 1% NP-40, 40 mM DTT) followed by sonication with an Ultrasonic Cell Disruptor. Then, lysate proteins were separated by 15% SDS–PAGE gels and then transferred to PVDF membranes. Western blots were performed using antibodies against H3K4m1 (Abcam, ab8895), H3K4m2 (Abcam, ab7766), H3K4m3 (Abcam, ab8580), or total histone 3 (H3, HuaBio, EM30605).
qPCR assay
Expression levels of genes (one set of the genes was selected from downregulated genes in the microarray data comparing the Utx KO group to the WT group, and the other set of genes was selected from upregulated genes in the microarray data comparing SP2509 treatment to vehicle treatment in the Utx KO group) were quantified by using SYBR Green Master Mix and the Quant Studio 3 Real-Time PCR System (Thermo Fisher Scientific). qPCR primer sequences are listed in Table 1.
Leukemia modeling and in vivo treatment
All animal studies were approved by the Institutional Animal Care and Use Committees of Sichuan University. Approximately 2 × 106 Utx−/−; shp53-mCherry;shNf1-GFP AML cells were transplanted by tail-vein injections into sublethally irradiated (4.5 Gy) 8-week-old female C57BL/6 mice. Drug treatments were initiated on day 7 after the cells were transplanted, and six mice were treated in each group: vehicle vs 25 mg/kg SP2509. SP2509 in solvent buffer (20% PEG-40, 20% dimethyl sulfoxide, 60% sterile water) or vehicle was intraperitoneally administered twice per week (Tuesday and Thursday) for 3 weeks as previously administrated.41 The leukemia progression in recipient mice was monitored by CBC, flow cytometry, and blood smear once per week.
References
Cancer Genome Atlas Research, N. et al. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013).
Shih, A. H., Abdel-Wahab, O., Patel, J. P. & Levine, R. L. The role of mutations in epigenetic regulators in myeloid malignancies. Nat. Rev. Cancer 12, 599–612 (2012).
Figueroa, M. E. et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell 18, 553–567 (2010).
Challen, G. A. et al. Dnmt3a is essential for hematopoietic stem cell differentiation. Nat. Genet. 44, 23–31 (2011).
Lu, C. et al. IDH mutation impairs histone demethylation and results in a block to cell differentiation. Nature 483, 474–478 (2012).
Chen, C. et al. Cancer-associated IDH2 mutants drive an acute myeloid leukemia that is susceptible to Brd4 inhibition. Genes Dev. 27, 1974–1985 (2013).
Ko, M. et al. Ten-Eleven-Translocation 2 (TET2) negatively regulates homeostasis and differentiation of hematopoietic stem cells in mice. Proc. Natl Acad. Sci. USA 108, 14566–14571 (2011).
Chen, C. et al. MLL3 is a haploinsufficient 7q tumor suppressor in acute myeloid leukemia. Cancer Cell 25, 652–665 (2014).
Agger, K. et al. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature 449, 731–734 (2007).
De Santa, F. et al. The histone H3 lysine-27 demethylase Jmjd3 links inflammation to inhibition of polycomb-mediated gene silencing. Cell 130, 1083–1094 (2007).
Hong, S. et al. Identification of JmjC domain-containing UTX and JMJD3 as histone H3 lysine 27 demethylases. Proc. Natl Acad. Sci. USA 104, 18439–18444 (2007).
Lan, F. et al. A histone H3 lysine 27 demethylase regulates animal posterior development. Nature 449, 689–694 (2007).
Shilatifard, A. The COMPASS family of histone H3K4 methylases: mechanisms of regulation in development and disease pathogenesis. Annu. Rev. Biochem. 81, 65–95 (2012).
Hu, D. et al. The MLL3/MLL4 branches of the COMPASS family function as major histone H3K4 monomethylases at enhancers. Mol. Cell. Biol. 33, 4745–4754 (2013).
van Haaften, G. et al. Somatic mutations of the histone H3K27 demethylase gene UTX in human cancer. Nat. Genet. 41, 521–523 (2009).
Gui, Y. et al. Frequent mutations of chromatin remodeling genes in transitional cell carcinoma of the bladder. Nat. Genet. 43, 875–878 (2011).
Jankowska, A. M. et al. Mutational spectrum analysis of chronic myelomonocytic leukemia includes genes associated with epigenetic regulation: UTX, EZH2, and DNMT3A. Blood 118, 3932–3941 (2011).
Wartman, L. D. et al. Sequencing a mouse acute promyelocytic leukemia genome reveals genetic events relevant for disease progression. J. Clin. Invest. 121, 1445–1455 (2011).
Jones, D. T. et al. Dissecting the genomic complexity underlying medulloblastoma. Nature 488, 100–105 (2012).
Liu, J. et al. A functional role for the histone demethylase UTX in normal and malignant hematopoietic cells. Exp. Hematol. 40, 487–498.e483 (2012).
Graubert, T. & Walter, M. J. Genetics of myelodysplastic syndromes: new insights. Hematology 2011, 543–549 (2011).
Welstead, G. G. et al. X-linked H3K27me3 demethylase Utx is required for embryonic development in a sex-specific manner. Proc. Natl Acad. Sci. USA 109, 13004–13009 (2012).
Thieme, S. et al. The histone demethylase UTX regulates stem cell migration and hematopoiesis. Blood 121, 2462–2473 (2013).
Mansour, A. A. et al. The H3K27 demethylase Utx regulates somatic and germ cell epigenetic reprogramming. Nature 488, 409–413 (2012).
Shpargel, K. B., Starmer, J., Yee, D., Pohlers, M. & Magnuson, T. KDM6 demethylase independent loss of histone H3 lysine 27 trimethylation during early embryonic development. PLoS Genet. 10, e1004507 (2014).
Bailey, P. et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 531, 47–52 (2016).
Wu, Q. et al. In vivo CRISPR screening unveils histone demethylase UTX as an important epigenetic regulator in lung tumorigenesis. Proc. Natl Acad. Sci. USA 115, E3978–e3986 (2018).
Ntziachristos, P. et al. Contrasting roles of histone 3 lysine 27 demethylases in acute lymphoblastic leukaemia. Nature 514, 513–517 (2014).
Van der Meulen, J. et al. The H3K27me3 demethylase UTX is a gender-specific tumor suppressor in T-cell acute lymphoblastic leukemia. Blood 125, 13–21 (2015).
Li, X. et al. UTX is an escape from X-inactivation tumor-suppressor in B cell lymphoma. Nat. Commun. 9, 2720 (2018).
Zheng, L. et al. Utx loss causes myeloid transformation. Leukemia 32, 1458–1465 (2018).
Nowak, D., Stewart, D. & Koeffler, H. P. Differentiation therapy of leukemia: 3 decades of development. Blood 113, 3655–3665 (2009).
Zhu, J., Chen, Z., Lallemand-Breitenbach, V. & de The, H. How acute promyelocytic leukaemia revived arsenic. Nat. Rev. Cancer 2, 705–713 (2002).
Wang, Z. Y. & Chen, Z. Acute promyelocytic leukemia: from highly fatal to highly curable. Blood 111, 2505–2515 (2008).
Sankar, S. et al. Mechanism and relevance of EWS/FLI-mediated transcriptional repression in Ewing sarcoma. Oncogene 32, 5089–5100 (2013).
Shi, Y. et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119, 941–953 (2004).
Wang, C. et al. UTX regulates mesoderm differentiation of embryonic stem cells independent of H3K27 demethylase activity. Proc. Natl Acad. Sci. USA 109, 15324–15329 (2012).
Kuhn, R., Schwenk, F., Aguet, M. & Rajewsky, K. Inducible gene targeting in mice. Science 269, 1427–1429 (1995).
Kubicek, S. et al. Reversal of H3K9me2 by a small-molecule inhibitor for the G9a histone methyltransferase. Mol. Cell 25, 473–481 (2007).
Davies, S. P., Reddy, H., Caivano, M. & Cohen, P. Specificity and mechanism of action of some commonly used protein kinase inhibitors. Biochem. J. 351, 95–105 (2000).
Fiskus, W. et al. Highly effective combination of LSD1 (KDM1A) antagonist and pan-histone deacetylase inhibitor against human AML cells. Leukemia 28, 2155–2164 (2014).
Silverman, L. R. et al. Randomized controlled trial of azacitidine in patients with the myelodysplastic syndrome: a study of the cancer and leukemia group B. J. Clin. Oncol. 20, 2429–2440 (2002).
Kantarjian, H. M. et al. Multicenter, randomized, open-label, phase III trial of decitabine versus patient choice, with physician advice, of either supportive care or low-dose cytarabine for the treatment of older patients with newly diagnosed acute myeloid leukemia. J. Clin. Oncol. 30, 2670–2677 (2012).
Gozdecka, M. et al. UTX-mediated enhancer and chromatin remodeling suppresses myeloid leukemogenesis through noncatalytic inverse regulation of ETS and GATA programs. Nat. Genet. 50, 883–894 (2018).
Ebert, B. L. et al. Identification of RPS14 as a 5q- syndrome gene by RNA interference screen. Nature 451, 335–339 (2008).
Wiederschain, D. et al. Contribution of polycomb homologues Bmi-1 and Mel-18 to medulloblastoma pathogenesis. Mol. Cell. Biol. 27, 4968–4979 (2007).
Andricovich, J. et al. Loss of KDM6A activates super-enhancers to induce gender-specific squamous-like pancreatic cancer and confers sensitivity to BET inhibitors. Cancer Cell 33, 512–526.e518 (2018).
Sze, C. C. & Shilatifard, A. MLL3/MLL4/COMPASS family on epigenetic regulation of enhancer function and cancer. Cold Spring Harbor Perspect. Med. 6 https://doi.org/10.1101/cshperspect.a026427 (2016).
Dhar, S. S. et al. An essential role for UTX in resolution and activation of bivalent promoters. Nucleic Acids Res. 44, 3659–3674 (2016).
Maiques-Diaz, A. et al. Enhancer activation by pharmacologic displacement of LSD1 from GFI1 induces differentiation in acute myeloid leukemia. Cell Rep. 22, 3641–3659 (2018).
Pishas, K. I. et al. Therapeutic targeting of KDM1A/LSD1 in Ewing sarcoma with SP-2509 engages the endoplasmic reticulum stress response. Mol. Cancer Ther. 17, 1902–1916 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Mootha, V. K. et al. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 34, 267–273 (2003).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Acknowledgements
We thank all Chen and Liu laboratory members for their invaluable contributions to discussion and technical support. This work was supported by the National Key R&D Program of China (2017YFA0505600) and the National Natural Science Foundation of China (81570150 and 81670182).
Author information
Authors and Affiliations
Contributions
B.W., C.C., X.S. and Y.L. conceived the project, designed experiments, and wrote the manuscript. B.W., M.C., K.S., J.X. and J.Z. performed experiments and analyzed data. X.P. and X.C. performed RNA-seq and ChIP-seq analysis.
Corresponding authors
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, B., Pan, X., Chen, X. et al. Epigenetic drug library screening identified an LSD1 inhibitor to target UTX-deficient cells for differentiation therapy. Sig Transduct Target Ther 4, 11 (2019). https://doi.org/10.1038/s41392-019-0040-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41392-019-0040-2
- Springer Nature Limited
This article is cited by
-
Repression of LSD1/KDM1A activity improves the response of liver cancer cells to the lenvatinib
Discover Oncology (2024)
-
JAK/BCL2 inhibition acts synergistically with LSD1 inhibitors to selectively target ETP-ALL
Leukemia (2022)
-
KMT2C deficiency promotes small cell lung cancer metastasis through DNMT3A-mediated epigenetic reprogramming
Nature Cancer (2022)
-
Lysine-specific demethylase 1 inhibition enhances autophagy and attenuates early-stage post-spinal cord injury apoptosis
Cell Death Discovery (2021)
-
SRT1720 inhibits the growth of bladder cancer in organoids and murine models through the SIRT1-HIF axis
Oncogene (2021)