Abstract
The current pandemic of coronavirus disease 2019 (COVID-19) caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) infection has dramatically influenced various aspects of the world. It is urgent to thoroughly study pathology and underlying mechanisms for developing effective strategies to prevent and treat this threatening disease. It is universally acknowledged that cell death and cell autophagy are essential and crucial to maintaining host homeostasis and participating in disease pathogenesis. At present, more than twenty different types of cell death have been discovered, some parts of which have been fully understood, whereas some of which need more investigation. Increasing studies have indicated that cell death and cell autophagy caused by coronavirus might play an important role in virus infection and pathogenicity. However, the knowledge of the interactions and related mechanisms of SARS-CoV-2 between cell death and cell autophagy lacks systematic elucidation. Therefore, in this review, we comprehensively delineate how SARS-CoV-2 manipulates diverse cell death (including apoptosis, necroptosis, pyroptosis, ferroptosis, and NETosis) and cell autophagy for itself benefits, which is simultaneously involved in the occurrence and progression of COVID-19, aiming to provide a reasonable basis for the existing interventions and further development of novel therapies.
Similar content being viewed by others
Introduction
Coronavirus disease 2019 (COVID-19) pandemic caused by severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2)1 is disastrous to public health, the global economy, and society as a whole human. As of 8 June 2022, the cumulative number of the World Health Organization (WHO) reported cases is 530,896,347 with 6,301,020 deaths (for constantly updated information, see: https://covid19.who.int). SARS-CoV-2 infects the host cells by binding with angiotensin-converting enzyme-2 (ACE2)2 or CD1473 receptors on the cell membrane and being assisted with transmembrane serine protease 2 (TMPRSS2) and protease Furin.4 Besides, SARS-CoV-2 also utilizes phosphatidylinositol 3-phosphate 5-kinase (PIKfyve), two-pore channel subtype 2 (TPC2), cathepsin L (CTSL) -mediated endocytosis for entry.5 Once entering into the host cell, SARS-CoV-2 replicates RNAs and expresses proteins, assembles into complete viral particles, and eventually releases virions, concurrently leading to cell death and cellular content release.6 The ACE2 receptor is widely expressed in the lungs, kidneys, gastrointestinal system, cardiovascular system, central nervous system, etc.7 From mild to severe, the clinical manifestations of COVID-19 include asymptomatic infection, pulmonary and extra-pulmonary symptoms (such as fever, fatigue, cough, diarrhea, nausea, loss of taste and smell, etc.), acute respiratory distress syndrome (ARDS), acute kidney injury (AKI), multiple organ failure, and even death.8
It is generally acknowledged that cell death and cell autophagy are fundamental cellular activities, playing crucial roles in maintaining homeostasis and disease pathogenesis. Over the past decades, more than twenty types of cell death have been identified by the Nomenclature Committee on Cell Death (NCCD),9 to which apoptosis, necroptosis, and pyroptosis are paid more attention. Previous studies have suggested that viruses can delicately regulate cell death through multiple mechanisms in different types of cells. A growing body of evidence indicates that these processes are considered double-edged swords during virus infection.10,11 On the one hand, the replication and spread of the virus are impeded due to the clearance of infected cells via cell death. However, on the other hand, dysregulated cell death causes uncontrol cell damage and disordered immune response. Meanwhile, viruses can hijack cell autophagy to be advantageous for replication niches, immune evade, and extracellular release.12 Currently, the relationship between SARS-CoV-2, cell death, and cell autophagy is not well established, but to some extent, could be referred to previous findings of SARS-CoV and Middle East respiratory syndrome coronavirus (MERS-CoV).13,14 Herein, we discuss cell death and cell autophagy involved in the pathogenesis and intervention therapies of COVID-19, providing a potential direction for the repurposing of old drugs and the development of novel treatments.
SARS-CoV-2: structure and characteristics
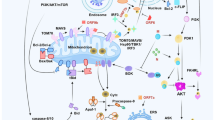
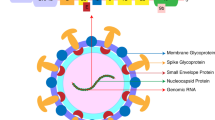
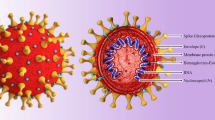
Coronaviruses are classified into four genera, namely alpha-, beta-, gamma-, and delta-coronavirus, with the genomes ranging from 26 to 32 kb. SARS-CoV-2 is an enveloped, positive-sense single-stranded RNA (+ssRNA) with a length of 29.9 kb. This genomic RNA consists of four structural regions (which encode Spike (S), Envelope (E), Membrane (M), Nucleocapsid (N) protein), and thirteen open reading frames (ORFs) (which encode nonstructural proteins (NSP1-16) and accessory proteins).15 All of them are involved in virus entry, translation of replication machinery, virus replication, translation of structure proteins, virion assembly, and virus release.16 Furthermore, the next-generation sequencing presented that SARS-CoV-2 shares 79.4% homology to SARS-CoV and 50% to MERS‐CoV.16
Apoptosis and SARS-CoV-2
Apoptosis core machinery
Apoptosis is a common programmed cell death (PCD) with immunological silence studied by J. Kerr first via electron microscopy in 1972.17 The morphological alterations of apoptosis are characterized by cell shrinkage, condensation and fragmentation of chromatin, and forming of apoptotic bodies. Apoptosis is vital in maintaining cellular homeostasis and regulating physiological and pathological events.18,19 Apoptosis can be triggered mainly by the extrinsic and intrinsic pathways. Interacting with the extracellular death ligands, including Fas ligand (FasL or CD95), tumor necrosis factor-alpha (TNF-α), TNF-related apoptosis-inducing ligand (TRAIL), and interferon-gamma (IFN-γ), the death receptors (DRs) immediately recruit Fas-associated protein with death domain (FADD) or TNF receptor 1-associated death domain protein (TRADD) and pro-cysteine-aspartic protease (procaspase)−8/10 to assemble the death-inducing signaling complex (DISC), causing cleavage of procaspase-8, and then, active caspase-8 directly cleaves procaspase-3/6/7 to induce cell apoptosis. Meanwhile, a truncated form of the BH3 interacting domain death agonist (BID) processed by active caspase-8 can translocate into mitochondria, leading to B-cell lymphoma-2 (Bcl-2)-associated X (BAX)/Bcl-2 homologous killer (BAK) mediated-mitochondrial outer membrane permeabilization (MOMP) for releasing apoptotic factors cytochrome c (cyt c) into the cytoplasm. Afterward, the apoptosome composed of cyt c, the apoptotic protease activating factor-1 (Apaf-1), and caspase-9 activates executioner caspases-3/6/7. The intrinsic pathways stimulated by DNA damage and stress can directly cause mitochondrial damage and MOMP to release cyt c. The other subsequent events have been described as above.20
Apoptosis inhibition by SARS-CoV-2
As of current evidence, it is shown that apoptosis is an important event during SARS-CoV-2 infection. SARS-CoV-2 infection can, directly and indirectly, upregulate the expression of cellular FADD-like interleukin-1 (IL-1)-converting enzyme-inhibitory protein (c-FLIP), which modulate death effector domains to suppress caspase-8/10 activation.21 The upregulation of miRNA-155 at the beginning of virus infection was reported by the small RNA profiling analysis of SARS-CoV-2 infected human cell lines22 for reducing the expression of forkhead transcription factor O 3a (FoXO3a) with the ability to reverse the inhibition of c-FLIP.23 It is acknowledged that the nuclear factor kappa B (NF-κB) pathway can upregulate transcriptional expression of some important apoptosis inhibitors, such as c-FLIP, X-linked inhibitor of apoptosis proteins (XIAPs), and Bcl-2.24 Activation of the (NF-κB) pathway has been identified upon SARS-CoV-2 infection.25,26 Besides, SARS-CoV-2 proteins including NSP13 and ORF9c were indicated to interact with the NF-κB pathway according to proteomic analyses.15 SARS-CoV-2 S protein was also reported to activate Toll-like receptor 2 (TLR2)-dependent NF-κB signaling pathway in human and mouse lung epithelial cells.27 Hence, it is reasonable speculation that SARS-CoV-2 would utilize apoptosis inhibition to avoid elimination, so as to obtain enough time and place for replication at the early stage.
Apoptosis induction by SARS-CoV-2
Accumulative studies have demonstrated that SARS-CoV-2 proteins greatly participate in apoptosis induction in various aspects. Similar to the previous findings of SARS-CoV ORF3a,28,29 SARS-CoV-2 ORF3a is also identified as a viroporin, with the ability to form an ion channel on the cell membrane to disturb intracellular homeostasis, which is responsible for apoptosis and promote virus release.30,31 Another study showed that SARS-CoV-2 ORF3a could efficiently induce apoptosis by directly cleaving and activating caspase-8 for the extrinsic pathway and cross-talk to the intrinsic pathway via tBID, leading to the cyt c release and caspase-9 activation.32 Besides, SAR-CoV-2 ORF7b was reported to promote the expression of TNF-α and induce TNF-α-dependent apoptosis in HEK293T cells and Vero E6 cells.33
Apart from the accessory proteins, the structural proteins also can trigger apoptosis by multiple mechanisms. SARS-CoV-2 E protein is sub-located in the endoplasmic reticulum (ER)-Golgi intermediate compartment (ERGIC) and serves as an ion channel,34 which is in accordance with the previous study of SARS-CoV E protein.35 Inhibition of E protein with a channel inhibitor BE-33 reduced both viral load and cell damage of SARS-CoV-2 infection in the hACE2 mouse model. Furthermore, the SARS-CoV-2 M protein regulates cell apoptosis in two ways. Firstly, SARS-CoV-2 M protein directly inhibits the ubiquitination of Bcl-2 ovarian killer (BOK) (a pro-apoptotic protein36) and promotes its translocating to the mitochondria, leading to H292 cell apoptosis via the intrinsic way.37 In addition, the pulmonary damage caused by M protein overexpression can be alleviated by the knockdown of BOK. Secondly, M protein suppresses 3-phosphoinositide-dependent protein kinase 1 (PDK1)-protein kinase B (PBK)/AKT axis, thereby decreasing the activity of its substrates, including forkhead transcription factor (FKHRL1) and apoptosis signal-regulating kinase (ASK).38 The reduction in phosphorylated FKHRL1 boosts its translocation to the nucleus for upregulating FasL expression on the membrane for the extrinsic apoptosis. Meanwhile, the inhibition of caspase-9 is reversed by dephosphorylated ASK, triggering the intrinsic apoptosis. Besides, SARS-CoV-2 S protein can activate reactive oxygen species (ROS)-inhibited phosphoinositide 3-kinase (PI3K)/AKT/ mammalian target of rapamycin (mTOR) pathways,39 subsequently inducing autophagy-triggered apoptosis and inflammation.40,41,42 It is noteworthy that SARS-CoV-2 has weaker pro-apoptotic activity than SARS-CoV,32,43 which may partially explain the more cases of asymptomatic patients with widespread viral transmission. Mutations of SARS-CoV-2 ORF3a attenuate cytosolic form-related apoptosis, whereas SARS-CoV ORF3a could well leverage this way. In addition, previous studies have shown that SARS-CoV can promote cell apoptosis in various other ways, including ORF3a- and ORF7a-induced p38 mitogen-activated protein kinase (MAPK) activation,44,45 ORF3a- and ORF6-induced ER stress,46,47 and N protein-upregulated JNK and p38 MAPK activities,48 all of which should be further investigated on SARS-CoV-2. The underlying mechanisms that SARS-CoV-2 regulates cell apoptosis are summarized in Fig. 1.
The mechanisms of SARS-CoV-2 regulating cell apoptosis. Activated death receptors recruit and cleave caspase-8/10, subsequently activating caspase-3/6/7 to induce extrinsic cell apoptosis. Stress- and DNA damage-stimulated cytochrome c directly cleaves caspase-9 to activate caspase-3/6/7 for intrinsic cell apoptosis. And tBID caused by caspase-8 also induces intrinsic apoptotic way. SARS-CoV-2 infection-upregulated c-FLIP and NF-κB inhibit apoptosis in infected cells. In contrast, SARS-CoV-2 ORF7b- and M protein-upregulated death ligands and SARS-CoV-2 ORF3a-activated caspase-8 promote extrinsic apoptosis. And SARS-CoV-2 S protein regulates Bcl-2 and BAX to induce the intrinsic apoptotic pathway. SARS-CoV-2 M protein-stabilized BOK and -directly activated caspase-9 both induce intrinsic apoptosis. Besides, cellular ion imbalance caused by SARS-CoV-2 ORF3a and E protein is involved in infected cell apoptosis. SARS-CoV-2 severe acute respiratory syndrome coronavirus 2, S spike, E envelope, M membrane, ORF open reading frame, NF-κB nuclear factor kappa B, c-FLIP cellular Fas-associated protein with death domain-like interleukin-1-converting enzyme-inhibitory protein, Bcl-2 B-cell lymphoma-2, XIAP X-linked inhibitor of apoptosis protein, ROS reactive oxygen species, PI3K phosphoinositide 3-kinase, AKT protein kinase B, mTOR mammalian target of rapamycin, TNF-α tumor necrosis factor-alpha, TRAIL TNF-related apoptosis-inducing ligand, FADD Fas-associated protein with death domain, TRADD TNF receptor 1-associated death domain protein, Caspase cysteine-aspartic protease, tBID truncated BH3 interacting domain death agonist, BOK Bcl-2 ovarian killer, BAX Bcl-2-associated X, BAK Bcl-2 homologous killer, MOMP mitochondrial outer membrane permeabilization, Apaf-1 apoptotic protease activating factor-1, ERGIC endoplasmic reticulum-Golgi intermediate compartment. Created with BioRender
Apoptosis during SARS-CoV-2 infection
Increasing evidence has demonstrated that apoptosis is involved in the pathogenesis of COVID-19 might be due to massive cell damage and tissue injury, such as in the lung, kidney, liver, pancreas, nervous system, immune system, etc. Apoptosis was detected by TdT-mediated dUTP nick end labeling (TUNEL) staining in lung samples from human patients and the non-human primate (NHP) model of SARS-CoV-2 infection which is responsible for accelerating the progression of ARDS.42 Further study showed that the intrinsic and extrinsic pathways triggered apoptosis in various cell types, including alveolar type 1 and 2 cells (AT1s and AT2s), vascular endothelial cells (ECs), macrophages, and T cells in the infected NHP lungs. A number of studies on SARS-CoV-2-induced apoptosis in respiratory epithelial cells and ECs align with the above findings.49,50,51,52 A large number of the renal tubular epithelial cells also occur apoptosis, leading to AKI in the mouse model with SARS-CoV-2 infection.53 In addition, SARS-CoV-2 induced β cell apoptosis via the JNK-MAPK apoptotic pathway, resulting in attenuated insulin production and secretion and exacerbation of diabetes.54 In addition, the phosphoproteomics data of human islets revealed that apoptosis-related signaling pathways were upregulated during SARS-CoV-2 infection.
Dysregulation of immune response is considered as a very important characteristic in SARS-CoV-2 infection.25,55 Lymphopenia, manifested with reduced absolute numbers of CD4+ and CD8+ T cells,56 has been related to the severity of COVID-19 patients.57 Two studies found that FasL expression on CD4+ and CD8+ T cells in COVID-19 patients is significantly higher than that in healthy controls, contributing to cell apoptosis,58 which correlates with lymphopenia and severe condition.59 Meanwhile, apoptosis-related caspase activation was also observed in T cells of infected patients.60 Moreover, the single-cell RNA sequencing (scRNA-seq) analysis of CD3+ T cells from COVID-19 patients indicted the upregulation of cell death-related genes.61 In this study, dysfunctional mitochondria with irregular shapes and incomplete cristae accompanied by cyt c release were exhibited by confocal and transmission electron microscopy, linking to apoptosis in T cells, which could be rescued by caspases inhibitor. Besides, surface FasL expression on B cells in the lymph nodes (LNs) of COVID-19 indicated SARS-CoV-2-promoted B-cell apoptosis.62
The decreased proportion of dendritic cells (DCs) in COVID-19 patients is another immune feature, which also correlates with the disease severity of SARS-CoV-2 infection.56,63,64 Damaged mitochondria and caspase-3 activation-dependent apoptosis were found in monocyte-derived macrophages (MDMs) and DCs, which could be prevented via anti-IFNγ treatment.65 Another study further indicated that IFNγ-induced macrophage cell death is mediated by caspase-8.66 On the one hand, active caspase-8 directly processes caspase-3 for the extrinsic cell apoptosis; on the other hand, it downregulates the expression of Bcl-2 to promote mitochondrial-driven cell apoptosis. Interestingly, upregulated inducible nitric oxide synthase (iNOS) promotes the cleavage of caspase-8 and instability of anti-apoptotic protein myeloid cell leukemia 1 (Mcl-1) to reinforce cell apoptosis. Besides, the conditional-independence network analysis suggested that apoptotic gene signatures (including BRCA2, CASP3, CASP8, BID, BAK1, and XBP1) significantly increased in plasmacytoid DCs (pDCs) from COVID-19, which is related to low-frequency pDCs and disease severity.67 Another scRNA-seq data was in accordance with this study.68 Generally speaking, pDCs are the major subset of secreted type I IFN (including IFN-α and IFN-β) in response to viral infection.69
Apoptosis-targeted therapies
As described above, two completely opposite approaches should be considered for the treatment according to the different stages of SARS-CoV-2 infection. The impeded apoptosis of host cells is advantageous for viral replication at the early stage, which emerges as a potential target for COVID-19 treatment. Therefore, inducing apoptosis cascade via blocking the suppressor (such as c-FLIP and NF-κB) might be beneficial for the host. In this regard, the histone deacetylase inhibitors (such as LBH589) with inhibitory effect on c-FLIP can be applied to COVID-19 patients.70 Many anti-inflammatory and antioxidative agents, such as phillyrin,71 dexamethasone,72 hydroxychloroquine,73 macrolide antibiotics,74 and N-acetylcysteine,75 have adverse effects on SARS-CoV-2 replication partially due to inhibition of the NF-κB pathway.76 It should be noticed that inhibiting NF-κB would contribute to significant side effects and adverse reactions for the reason that suppression of immune response.
In the later stage, SARS-CoV-2 strongly induces apoptosis in various host cells, resulting in a large amount of tissue damage and loss of function, which accelerates the development of disease and even death. Thereby, impeding apoptotic pathways by blocking DRs signaling and caspase cascade, has been proposed as a promising strategy to reduce viral widespread and alleviate disease progression. Emricasan, an irreversible pan-caspase inhibitor, was reported to protect human cortical neural progenitors from caspase-3-induced cell death during Zika virus infection,77 which is considered as a possible drug for COVID-19 patients. However, the clinical trial on emricasan for mild-COVID-19 treatment has been terminated for the reason of difficulty in recruiting patients (NCT04803227). Unfortunately, the majority of caspase inhibitors are only utilized in cell and animal experiments, thus requiring further experiments explored in clinical trials.
It has been well established that TNF-α plays a vital role in the pathogenesis of infectious diseases, inflammatory processes, and malignant tumors.78 Increasing studies have shown that elevated circulating levels of TNF-α are positively associated with disease severity and death of COVID-19 patients.79,80,81 A recent study displayed that co-treatment of TNF-α and IFN-γ could substantially induce cell death, including apoptosis in PMA-differentiated macrophage-like THP-1 cells.82 Thus, targeting TNF-α and its receptor is a promising therapy for SARS-CoV-2 infection. Infliximab is a mouse-human chimeric monoclonal antibody to effectively block TNF-α and prevent its binding to receptors, which has obtained the US Food and Drug Administration (FDA)-approval for treatment with rheumatoid arthritis, psoriatic arthritis, ankylosing spondylitis, and plaque psoriasis. A single-arm phase II trial indicated that Infliximab treatment decreased pathological inflammatory cytokines in severe and critical hospitalized COVID-19 patients (NCT04425538).83 Other clinical trials of Infliximab are under investigation (NCT04922827, NCT05220280, NCT04593940,84 NCT05273242). Adalimumab, a humanized anti-TNF-α monoclonal antibody, has been utilized with remdesivir and dexamethasone for severe COVID-19 patients in a randomized controlled trial. Nevertheless, no positive results were observed in mortality, mechanical ventilation requirement, hospital and ICU stay duration, and even radiologic changes.85 Additionally, Etanercept serves as a TNF-α/β inhibitor and is undergoing clinical trial (NCT05080218). Intriguingly, it is found that anti-TNF-α treatment would have negative effects on antibody response in response to SARS-CoV-2 infection.86
Necroptosis and SARS-CoV-2
Necroptosis core machinery
Necroptosis is a form of PCD and caspase-independent lytic cell death, which was first observed by J. Yuan in 2005.87 Morphological changes of necroptosis include early loss of plasma membrane integrity, organelle swelling, and nuclear condensation. Binding with death ligands (including TNF-α, FasL, and IFN-γ), the DRs recruit TRADD, FADD, caspase-8, receptor-interacting protein kinase 1 (RIPK1) and RIPK3 to assemble the necrosome. Inactivation of caspase-8 caused autophosphorylation of RIPK1-RIPK3, which further phosphorylate mixed lineage kinase domain-like protein (MLKL). pMLKL oligomerize to form pores in the cytomembrane leading to cell death and release of damage-associated molecular patterns (DAMPs) and cytokines that can stimulate immune responses.88,89 The TLRs signaling pathway and viral RNA-Z-DNA/RNA binding protein-1 (ZBP1, also known as DAI) axis also can directly interact and activate RIPK3 to induce necroptosis.90,91
Necroptosis interplay with SARS-CoV-2 and -targeted therapies
The interaction between the host cell necroptosis and SARS-CoV-2 has not yet been totally elucidated. In a recent study, RIPK1 activation was found in human COVID-19 lung samples.92 Treatment with Necrostatin-1(Nec1, a known RIPK1 inhibitor) reduced viral load, inflammation, and damage in cells and mouse models infected with SARS-CoV-2. It is demonstrated that necroptosis plays an important role in the pathogenesis of COVID-19. Furthermore, SARS-CoV-2 NSP12 can directly interact with and promote RIPK1 activation. Another study showed the fluorescence staining of pMLKL pores on the cytomembrane in SARS-CoV-2-infected cells and human postmortem lungs in another study.93 SARS-CoV-2 can induce RIPK3 to promote cell necroptosis, which is significantly attenuated by RIPK3 inhibitors. Interestingly, a previous study reported that RIPK3-induced SARS-CoV ORF3a oligomerization augments membrane insertion and ion channel function of 3a, finally resulting in multimodal cell death in a human lung cell line (including necroptosis).94 The interaction of SARS-CoV-2 with cell necroptosis is concluded in Fig. 2.
The interaction of SARS-CoV-2 and cell necroptosis. Activated death receptors phosphorylate RIPK1 and RIPK3 in the absence of active caspase-8, and consecutively phosphorylate MLKL to form pores in the cell membrane, eventually leading to cell necroptosis. SARS-CoV-2 ORF7b and M protein-upregulated death ligands, SARS-CoV-2 NSP12-activated RIPK1, and SARS-CoV-2-related RIPK3 activation all promote necroptosis in infected cells. Meanwhile, the SARS-CoV-2-stimulated TLRs pathway and viral ZBP1 axis also directly activate RIPK3 to induce necroptosis. Besides, RIPK3-induced SARS-CoV-2 ORF3a oligomerization enhances its functions. SARS-CoV-2 severe acute respiratory syndrome coronavirus 2, M membrane, ORF open reading frame, NSP nonstructural protein, IFN-γ interferon-gamma, TRADD TNF receptor 1-associated death domain protein, FADD Fas-associated protein with death domain, Caspase cysteine-aspartic protease, RIPK receptor-interacting protein kinase, Nec1 necrostatin-1, MLKL mixed lineage kinase domain-like protein, TLR Toll-like receptor, ZBP1/DAI Z-DNA/RNA binding protein-1. Created with BioRender
Although blocking the virus replication via cell death, necroptosis conversely accelerates viral widespread and the release of cellular contents by cell rupture. It was reported that the serum RIPK3 levels are higher in severe patients than in mild ones, suggesting that RIPK3 may be involved in the progression of COVID-19 pneumonia to ARDS.95,96 Thus, targeting the components of the necroptotic pathway might be a therapeutic approach for COVID-19.97 Both the RNA-sequencing data and proteome-wide data analysis revealed that RIPK1 is a potential drug target for COVID-19 patients.98,99 The FDA-approved drug primidone has the property to reduce RIPK1-mediated necroptosis in vitro and TNFα-induced inflammation in vivo,100 which may be beneficial to patients with COVID-19. The drugs targeting RIPK3 (GSK872) and MLKL (necrosulfonamide) have not yet been further investigated in COVID-19.
Pyroptosis and SARS-CoV-2
Pyroptosis core machinery
Pyroptosis is a lytic form of PCD driven by inflammasome activation, which was first termed by B. Cookson in 2001.101 In morphology, cell swelling and rupture, DNA condensation and fragmentation, and membrane pores formation are observed in pyroptosis. Pyroptosis plays a crucial role in inflammation and immunity,102 thus involved in the development of diverse diseases.103 Pyroptosis can be induced by mainly two inflammasome pathways: the caspase-1-dependent canonical pathway and the caspase-4/5/11-dependent noncanonical pathway. In the priming phase, pathogen-associated molecular patterns (PAMPs) and damage-associated molecular patterns (DAMPs) are recognized by various receptors (such as TLRs, TNFR, and IL-1R1), resulting in NF-κB activation to promote pro-inflammatory cytokines production including nucleotide-binding oligomerization domain-like receptor containing pyrin domain 3 (NLRP3), procaspase-1, pro-IL-1β, and pro-IL-18. In the activation phase, PAMPs- and DAMPs-activated NLRP3 recruits the apoptosis-associated speck-like protein containing a CARD (ASC) and procaspase-1 to form inflammasome specks generating active caspase-1 that further cleaves pro-IL-1β, pro-IL-18, and gasdermin-D (GSDMD). GSDMD-N inserts into cytomembrane to form GSDMD pores, leading to cell pyroptosis and contents liberation (such as IL-1β, IL18, IL-6, TNFα, IL-8, DAMPs, etc).104 In addition, lipopolysaccharide (LPS)-induced activation of caspase-4/5/11 can directly cleave GSDMD to form pores and trigger cell pyroptosis.
Pyroptosis regulation by SARS-CoV-2 and host system
It is well known that the activation of inflammasomes is the rate-limiting step of cell pyroptosis, which can be affected by a wide range of factors, such as toxins, viruses, bacteria,105 ATP change,106 ion imbalance,107,108 ROS production,109 mitochondrial dysfunction,110 lysosomal destruction,111 etc. Growing studies have indicated that various proteins encoded by SARS-CoV-2 regulate inflammasome activity through multiple mechanisms. As described above, SARS-CoV-2 ORF3a and E protein are identified as viroporins for ion disorder. SARS-CoV-2 ORF3a induces K+ efflux and Ca2+ influx112 whereas E protein only promotes Ca2+ influx,34 resulting in a decrease in cellular K+ level and an increase in Ca2+ level. Previous studies showed that ion imbalance not only directly activates NLRP3 inflammasome but also augments the generation of mitochondrial ROS. Furthermore, SARS-CoV-2 ORF3a works as a bridge for promoting the interaction of never in mitosis gene a (NIMA)-related kinase 7 (NEK7) and NLRP3, leading to IL-1β release in lung epithelial cell line A549.112 The NEK7 is an essential mediator of NLRP3 activation.113 In addition, lysosomal dysfunction caused by SARS-CoV-2 ORF3a also has been reported, manifesting K+ efflux, Ca2+ influx, and cathepsins leakage.114 Apart from ORF3a, SARS-CoV-2 N protein directly interacts with NLRP3 protein, further promoting the interaction of NLRP3 with ASC and assembly of the inflammasome.115 Besides, SARS-CoV-2 NSP6 impairs lysosomal acidification via directly interacting with ATP6AP1 (a vacuolar ATPase proton pump component), leading to autophagic flux stagnation, which facilitates NLRP3 inflammasome activation and pyroptosis in lung epithelial cell lines.116,117 Intriguingly, in turn, SARS-CoV-2 also possesses the ability to diminish host cell pyroptosis, which might be utilized to strive for replication time at the early stage. SARS-CoV-2 NSP1 and NSP13 both can directly inhibit NLRP3 inflammasome-induced active caspase-1 in macrophage-like THP-1 cells.118 NSP5 antagonizes pyroptosis by cleaving GSDMD at Q193-G194 junction into two inactive fragments with ineffective pores formation.119 Moreover, SARS-CoV-2 N protein protects GSDMD from caspase-1 cleavage via occupying the linker region of GSDMD in infected human monocytes.120
The human ACE2 receptor is widely expressed in many cell types, whose primary function is converting angiotensin II (Ang II) to angiotensin 1–7.121 Due to ACE2 being occupied mainly by SARS-CoV-2, Ang II degradation is attenuated, leading to Ang II accumulation. Hyperactivation of Ang II-angiotensin 1 receptor (AT1R) signaling induces mitochondrial dysfunction with ROS production, resulting in NLRP3 inflammasome-related pyroptosis,122,123 which is further confirmed in line with recent results of hematopoietic stem cells (HSCs) and endothelial progenitor cells (EPCs) damaged by SARS-CoV-2 infection.124
A number of studies indicated that the complement system is activated during SARS-CoV-2 infection, releasing C3a, C5a, and membrane attack complex (MAC), which is mediated by mannan-binding lectin (MBL) - MBL-associated serine protease (MASP).125,126,127 For example, SARS-CoV-2 N protein activates the MASP2-mediated complement pathway whereas S protein activates the alternative complement pathway.128 Previous studies have demonstrated that complement cascade can augment the activation of NLRP3 inflammasome and cell pyroptosis. The C3a-mediated ATP efflux initiates P2 x 7, an ATP-gated ion channel for K+ efflux.129 The C5a upregulates the expression of double-stranded RNA-dependent protein kinase (PKR),130 an important regulator of NLRP3 inflammasome activation.131 Additionally, the MAC can insert into the cytomembrane to form transmembrane pores for Ca2+ influx.132
Furthermore, absent in melanoma 2 (AIM2) inflammasome also activates caspase-1 and promotes cell pyroptosis, which was detected in SARS-CoV-2-infected human monocytes.133 It has been previously shown that cytoplasmic double-stranded DNA, such as mitochondrial DNA (mtDNA), oxidized mtDNA,134,135 and cell-free DNA (cfDNA)136 can directly trigger AIM2 inflammasome-mediated pyroptosis. During SARS-COV-2 infection, massive dysfunctional mitochondria would release mtDNA and oxidized mtDNA (induced by a large amount of ROS),137 whereas dead cell releases cfDNA.138 The complex associations among SARS-CoV-2, host system and cell pyroptosis are summarized in Fig. 3.
The sophisticated associations of SARS-CoV-2, host system, and cell pyroptosis. Activated TLRs, TNFR, and IL-1R1 pathways induce NF-κB activation to promote pro-inflammatory cytokines production including pro-IL-1β, pro-IL-18, procaspase-1, and NLRP3. The PAMPs and DAMPs caused by SARS-CoV-2 immediately activate the NLRP3 inflammasome to cleave procaspase-1 that subsequently cleaves pro-IL-1β and pro-IL-18. Meanwhile, caspase-1-cleaved GSDMD inserts into the membrane to form pores, leading to cell pyroptosis and contents release. SARS-CoV-2 N protein, NSP5, NSP1, and NSP13 inhibit infected cell pyroptosis via impeding the caspase-1 or GSDMD. On the contrary, more commonly, SARS-CoV-2 promotes cell pyroptosis in various ways. SARS-CoV-2 ORF3a and N protein directly activate the NLRP3 inflammasome. The ion imbalance caused by SARS-CoV-2 ORF3a and E protein and lysosomal dysfunction caused by SARS-CoV-2 ORF3a, ORF7a, and NSP16 both promote the NLRP3 inflammasome activation. In addition, the MAVS pathway activated by SARS-CoV-2 infection causes mitochondria damage and the activation of NF-κB and IRF. Besides, SARS-CoV-2-related ROS production and mtDNA release also activate the NLRP3 inflammasome. Meanwhile, dsDNA and mtDNA-induced AIM2 inflammasome and LPS-induced caspase-4/5/11 also induce cell pyroptosis. From the perspective of the host, the Ang II-AT1R axis and the MBL-MASP-mediated complement system are involved in infected cell pyroptosis. SARS-CoV-2 severe acute respiratory syndrome coronavirus 2, N nucleocapsid, E envelope, ORF open reading frame, NSP nonstructural protein, TLR Toll-like receptor, TNFR tumor necrosis factor receptor, IL-1R1 interleukin-1 receptor 1, NF-κB nuclear factor kappa B, IRF transcription factors interferon regulatory factor, TNF-α tumor necrosis factor-alpha, HMGB1 high mobility group box 1, GSDMD gasdermin-D, Caspase cysteine-aspartic protease, AIM2 absent in melanoma, dsDNA double-stranded DNA, LPS lipopolysaccharide, PAMPs pathogen-associated molecular patterns, DAMPs damage-associated molecular patterns, NLRP3 nucleotide-binding oligomerization domain-like receptor containing pyrin domain 3, ROS reactive oxygen species, mtDNA mitochondrial DNA, Ang II angiotensin II, AT1R Ang II-angiotensin 1 receptor, MBL-MASP mannan-binding lectin (MBL)-MBL-associated serine protease, MAC membrane attack complex, PKR double-stranded RNA-dependent protein kinase, ERGIC endoplasmic reticulum-Golgi intermediate compartment. Created with BioRender
Pyroptosis during SARS-CoV-2 infection
Mounting evidence has implied that inflammasome activation and cell pyroptosis play pivotal roles in the pathogenesis of COVID-19. For one thing, this process causes massive cell damage and tissue injury. For another, it releases excessive inflammatory cytokines and DAPMs, termed as cytokine storm, strongly accelerating the development of COVID-19. Activated NLRP3 and AIM2 inflammasome-dependent pyroptosis were observed via fluorescence staining in infected circulating monocytes and lung tissue-resident macrophages from COVID-19 patients,139 in which NLRP3 activation was further confirmed through virus stimulation in other studies.133,140 In another study, compared to healthy controls, cell pyroptosis and cytokine storm occurred in CD163+ pro-inflammatory macrophages in lung samples from severe COVID-19 patients.141 SARS-CoV-2 pseudovirus-treated macrophage-like THP-1 cells also activated caspase-1, cleaved GSDMD, and secreted cytokines. Apart from the classical pathway, SARS-CoV-2-derived ssRNA can facilitate TLR8-dependent NLRP3 inflammasome activation and cytokines release in human macrophages without pyroptosis.142 Surprisingly, HSCs and EPCs are also damaged by NLRP3 inflammasome-mediated pyroptosis during SARS-CoV-2 infection,143,144 destroying hematopoietic and coagulation functions. Furthermore, it has been reported that SARS-CoV-2 triggers NLRP3 inflammasome-mediated pyroptosis in the endothelium,145 thus leading to endothelial dysfunction in the respiratory and myocardial system.146,147
Importantly, SARS-CoV-2 infection initiates cytokine storm, in which inflammasome-dependent cell pyroptosis has been proven as the very important inducer, manifested by elevated levels of pro-inflammatory cytokines and chemokines,148 such as IL-6, IL-1β, TNFα, IL-8, IL-10, CXCL10, MCP1, which strongly correlated with the COVID-19 severity and mortality.79,149,150 In this context, these inflammatory mediators strongly promote cell migration and activation, further emphasizing the inflammatory environment and tissue damage. Meanwhile, multiple types of cell death enhance epithelial and endothelial cell permeability, providing convenience for cell migration and fluid leakage, aggravating the progression of acute lung injury (ALI) and ARDS.151,152 It has been stated that SARS-CoV-2 spreads into the cardiovascular system via the circulatory or lymphatic system,153 as does in the lung, inducing uncontrolled cell deaths and triggering the pro-inflammatory cascade,154,155 resulting in myocarditis, loss of cardiac functions, and vascular diseases.156,157
Pyroptosis-targeted therapies
As of current evidence, inflammasome-mediated cell pyroptosis dramatically affects the development and progression of COVID-19, supporting it as a promising therapeutic target, particularly in severe cases. Two strategies could be under consideration: inhibition of the basic components (such as NLRP3 inflammasome, GSDMD, and cytokines) and inhibition of the enhancer (such as ion imbalance, ROS production, and Ang II-AT1R axis) of cell pyroptosis. A recent study showed that targeting NLRP3 inflammasome by gene knockout or selective inhibitor MCC950 ameliorated overactive inflammatory response and pathology of lung in mice models with SARS-CoV-2 infection,158 as does in the previous study of influenza A.159 A multicenter, randomized, controlled phase II trial of DFV890 (a selective NLRP3 inflammasome inhibitor) treatment for COVID-19 with pneumonia and impaired respiratory function exhibited negative results (NCT04382053). There is no significant difference between the two groups: standard of care (SoC) plus DFV890 treatment and only SoC. A phase II clinical trial of selective NLRP3 inflammasome inhibitor Dapansutrile for moderate COVID-19 with early cytokine release syndrome is recruiting (NCT04540120). Colchicine, a microtubule polymerization inhibitor, has been identified with the ability to inhibit NLRP3 inflammasome160,161 and investigated in clinical trials for COVID-19.162 A randomized, double-blind, placebo-controlled clinical trial in moderate to severe COVID-19 patients suggested that colchicine reduces oxygen supplement and hospitalization.163 Besides, colchicine was reported to decrease the rate of hospitalization and death of non-hospitalized patients with COVID-19 in a randomized, double-blind, placebo-controlled phase III trial (NCT04322682).164 However, the large-scale clinical trials of colchicine are disappointing. A randomized, controlled phase II/III trial in hospitalized COVID-19 patients indicated that colchicine could not reduce mechanical ventilation requirement, length of hospital stay, and 28-day mortality (NCT04381936),165 in line with results of another clinical trial (NCT04328480).166 Thus, the efficacies of colchicine by the single or combined treatment should be further studied.
GSDMD-dependent pores work as the executor of cell pyroptosis and cytokine release, implying the treatment possibility for inhibiting GSDMD. The FDA-approved disulfiram (DSF) 167 and dimethyl fumarate (DMF) 168 both have the property of inhibiting GSDMD pore formation in cells and animal models. It is noteworthy that DSF also can suppress virus replication by inhibiting the polyprotein proteases (including main protease (Mpro) and papain-like protease (PLpro)), NSP13, and NSP14 of SARS-CoV-2.169 Meanwhile, DMF exhibits antiviral activity by upregulating nuclear factor erythroid-2 related factor 2 (NRF2) antioxidant gene expression170 in SARS-CoV-2-infected cells.171 A retrospective study showed lower SARS-CoV-2 infection risk in alcohol use disorder patients with DSF treatment than without DSF treatment (hazard ratio (HR) 0.66).172 A randomized, double-blind, placebo-controlled phase II trial to evaluate the efficacy and safety of DSF in moderate COVID-19 patients has been completed but no result has been published yet (NCT04594343). Other clinical trials of DSF (NCT04485130) and DMF (NCT04381936, NCT04792567) in COVID-19 are under evaluation.
Cell pyroptosis causes numerous inflammatory cytokines release, among which IL-1β can in turn boost inflammasome activation in a positive feedback way.173,174 Thereby, that gives powerful support for direct inhibition of cytokines for COVID-19 treatment.175 Anakinra is an IL-1 receptor antagonist (IL-1RA) to block the binding of IL-1 with their receptor, which has obtained FDA approval for the treatment of rheumatoid arthritis and neonatal onset multisystem inflammatory disease. Anakinra inhibited NLRP3 inflammasome-dependent pyroptosis, alleviated the lung pathology, and eventually reduced the mortality in the SASR-CoV-2-infected mice model.176 A variety of clinical studies evaluating the effects of anakinra for COVID-19 treatment have been conducted. In a retrospective study, compared to historical controls, anakinra reduced mechanical ventilation requirement and mortality (HR 0.22) in severe COVID-19.177 Furthermore, a non-randomized, single-arm phase II trial suggested that anakinra regulated the inflammatory system, attenuated the rate of ARDS (HR 0.3), and reduced 30-day mortality (HR 0.49) in COVID-19 (NCT4357366).178 Nevertheless, there is no significant difference between anakinra treatment and SoC for mild to moderate COVID-19 patients in a randomized controlled trial (NCT04341584).179 FDA-approved Canakinumab directly neutralizes IL-1β and has been investigated in clinical trials. Canakinumab showed positive effects on inflammatory markers, oxygen improvement, and final outcomes of COVID-19 patients according to the results of several retrospective studies.180,181,182,183 However, a randomized, double-blind, placebo-controlled phase III study indicated that canakinumab exhibited no benefit on mechanical ventilation requirement and mortality in hospitalized patients with severe COVID-19 (NCT04362813).184 In addition, Canakinumab has been evaluated in the COVID-19 patients concurrently with cardiac injury or type 2 diabetes, whose results have not been published (NCT04365153, NCT04510493).
Extensive studies have suggested that IL-6 greatly takes part in the progression of cytokine storm185 and the pathogenesis of COVID-19.186 The increased concentration of IL-6 has been identified as a negative prognosis factor for the severity and death rate of COVID-19 patients,187,188 giving support for the strategy of IL-6 blockage for COVID-19 treatment.186 Tocilizumab and sarilumab, FDA-approved humanized monoclonal antibodies, antagonize both soluble and membrane-bound IL-6 receptors to prevent IL-6 binding, in which the former has been utilized in cytokine release syndrome, giant cell arteritis, polyarticular juvenile idiopathic arthritis, rheumatoid arthritis, systemic sclerosis-associated interstitial lung disease. A large number of clinical trials of Tocilizumab and sarilumab have been conducted, whereas the efficacies are still controversial. According to the data of a randomized, SoC-controlled phase II trial, Tocilizumab utilized in COVID-19 pneumonia had no benefit on disease progression (NCT04346355).189 Concurrently, another randomized, double-blind, placebo-controlled trial also indicted that Tocilizumab was ineffective in reducing invasive mechanical ventilation and death of moderate COVID-19 patients (NCT04356937).190 On the contrary, a randomized, double-blind phase III trial displayed that lower percentage of mechanical ventilation requirement and 28-day mortality (HR 0.56) in Tocilizumab-treated COVID-19 pneumonia compared to placebo-controls (NCT04372186).191 Tocilizumab significantly had positive effects on the median number of organ support-free days (HR 1.64) and 90-day survival (HR 1.64) of severe COVID-19 with ICU support in a randomized, multifactorial adaptive platform trial (NCT02735707).192 Surprisingly, a large-scale trial gives the potent support for the reason that reduction in relative mortality (rate ratio (RR) 0.86), increase in discharge alive (RR 1.23), and decrease in invasive mechanical ventilation or death (RR 0.85) of Tocilizumab treated hospitalized COVID-19 compared to that in SoC-controls (NCT04381936).193 The chimeric monoclonal antibodies (such as Clazakizumab, Olokizumab, Siltuximab, and Sirukumab) inhibit IL-6 activity by reducing the binding of IL-6 with IL-6R. However, two randomized, double-blind, placebo-controlled phase II trials showed that Clazakizumab did not improve oxygen requirement and mortality of patients with severe SARS-CoV-2 infection (NCT04348500, NCT04343989). Equally disappointing results have emerged in the clinical trial of Olokizumab (NCT04380519). Meanwhile, a multicenter, randomized phase III trial indicated that there was no difference in time to clinical improvement (TTI) and mortality between Siltuximab-treated severe COVID-19 and controls (NCT04330638).194 Other phase II clinical trials of Clazakizumab (NCT04494724, NCT04363502) or Olokizumab (NCT05187793), Siltuximab (NCT04329650) for COVID-19 are under evaluation.
The loss of cellular ion homeostasis has been proven to promote NLRP3 inflammasome activation and cell pyroptosis during SARS-CoV-2 infection, thus ion-channel inhibitors (such as Amantadine, Memantine,195 Rimantadine,196 Tretinoin197) could be utilized in treatment with COVID-19 patients. Amantadine displayed antiviral activity via directly blocking the ion channel encoded by SARS-CoV-2 E protein in vitro.198 It is being investigated in a randomized, double-blind, controlled phase III trial for COVID-19 (NCT04894617).
Targeting overactivation of the Ang II-AT1R axis represents another strategy to attenuate NLRP3 inflammasome activation and cell pyroptosis in COVID-19. On one hand, ACE inhibitors (ACEIs, such as Lisinopril, Captopril) reduce the conversion of Ang I into Ang II, leading to a decrease in the level of Ang II. On the other hand, angiotensin receptor blockers (ARBs, such as Losartan, Irbesartan, Telmisartan, Olmesartan) can inhibit Ang binding with their receptors, leading to a reduction in the Ang-ATR signaling. Previous experiments199,200,201,202 and observational studies203,204 both provided the support that ACEIs and ARBs might be beneficial for COVI-19 patients. A randomized, SoC-controlled phase IV trial showed that telmisartan reduced mechanical ventilation requirement and death on days 15 and 30 in hospitalized patients with COVID-19 (NCT04355936).205 However, it is a pity that two randomized, double-blind, placebo-controlled phase II clinical trials of losartan had no significant effect on outpatients and mild hospitalized patients with COVID-19 (NCT04311177, NCT04340557).206,207 Therefore, more clinical trials to exactly evaluate the effects of ACEIs and ARB on COVID-19 treatment are ongoing (NCT04345406, NCT04591210, NCT02735707, NCT04920838, NCT04466241).
NETosis and SARS-CoV-2
NETosis in SARS-CoV-2 infection
NETosis is a ROS-dependent PCD driven by neutrophil extracellular traps (NETs) in response to various infections, which was first described by A. Zychlinsky in 2004.208 Peptidylarginine deiminase 4 (PAD4), neutrophil elastase (NE), GSDMD, and free DNA greatly participate in the whole process of NETosis.209 NETs are complex networks comprised of DNA containing histones, myeloperoxidase (MPO), and NE.210
Cytokine storm has been characterized in COVID-19, particularly in severe patients. Previous studies have suggested that the pro-inflammatory cytokines (such as IL-1β, IL-6, C-X-C motif chemokine 8 (CXCL-8)/IL-8) can potently recruit and activate neutrophils.211,212 Accumulative studies have shown a significant increase in the numbers and activation of neutrophils, which are associated with severity during SARS-CoV-2 infection.213,214 The increased neutrophils and NETosis were found in blood samples from COVID-19 patients via transcriptomics, proteomics, and cfDNA analyses.215 In addition, the autocrine loop of IL-8 is identified in severe COVID-19 patients for augmentation of neutrophil migration and activation.216 Meanwhile, the concentrations of NETs in plasma, tracheal aspirate, and lung specimens from COVID-19 patients were higher compared to the healthy controls.217 SARS-CoV-2 can directly stimulate NETosis of healthy neutrophils, which is dependent on ACE2, TMPRSS2, and PAD4. As a result, the accumulation of NETs leads to apoptosis in lung epithelial cells. Furthermore, a recent study has indicated that SARS-CoV-2 cunningly manipulate histones (especially H3 and H4) released by NETosis to bridge subunit 2 of the S protein and sialic acid on the cell surface and promote membrane fusion, finally boosting their infectivity.218 Taken together, NETosis have two-edged sword activities during virus infection.219,220 The NETs trap viral particles to inhibit their transmission efficiently. However, the NETs also lead to cell damage, cytokine storm,221 and even immunothrombosis.222 The immunothrombosis is comprised of activated leukocytes with platelets and plasma coagulation factors and is involved in systemic diseases of COVID-19.126,223,224 The NETosis occurred during SARS-CoV-2 infection is displayed in Fig. 4.
NETs, released by SARS-CoV-2-induced NETosis, are comprised of DNA, histones, MPO, and NE. The IL-8-CXCR2, IL-6-IL-6R, and IL-1β-IL-1R1 axis play important roles in recruiting and activating neutrophils. In addition, SARS-CoV-2 hijacks histones to interact with S2 and sialic acid for enhanced infection. SARS-CoV-2 severe acute respiratory syndrome coronavirus 2, S2 subunit 2 of SARS-CoV-2 spike protein, ACE2 angiotensin-converting enzyme-2, TMPRSS2 transmembrane serine protease 2, IL-8 interleukin-8, CXCR2 C-X-C motif chemokine receptor 2, sIl-6R soluble IL-6 receptor, IL-1R1 interleukin-1 receptor 1, GSDMD gasdermin-D, PDEs phosphodiesterases, cAMP cyclic adenosine monophosphate, PAD4 peptidylarginine deiminase 4, NE neutrophil elastase, MPO myeloperoxidase, NETs neutrophil extracellular traps. Created with BioRender
NETosis-targeted therapies
Thereby, it has been proposed that inhibiting neutrophils and NETosis might be feasible approaches for COVID-19 treatment.225 Certainly, the primary strategy focuses on the vital molecules involved in NETosis, such as PAD4, NE, and GSDMD. The PAD4 inhibitors (such as Cl-amidine and GSK484) prevent NETosis by reducing histone citrullination in the treatment of severe inflammatory diseases.226 Sivelestat, a potent NE inhibitor approved in Korea and Japan, can ameliorate acute lung injury.227 A retrospective cohort study showed that sivelestat exhibited positive outcomes in patients with ARDS,228 indicating that NE inhibitor might be beneficial for patients with COVID-19. In addition, A randomized, placebo-controlled phase Ib/II clinical trial of Alvelestat treatment for COVID-19 with ARDS has been completed with no results published (NCT04539795). The degradation and elimination of NETs using DNases represent an appropriate pathway to reduce NETs’ burden. Dornase alfa is a FAD-approved recombinant human DNase I and showed significant effects of antiviral activity and disease relief of SARS-CoV-2 infection in vitro and in vivo.229 Currently, there are several clinical trials related dornase alfa for COVID-19 (NCT04432987, NCT04409925, NCT04409925, NCT04445285, NCT04402970, etc). In a non-randomized phase III clinical trial (NCT04402970), inhaled dornase alfa yielded positive outcomes of improved oxygenation and decreased NETs in COVID-19 patients.230
In addition to therapies directly antagonizing the forming and degradation, it is feasible to hinder the components involved in amplification loops of NETs. It is acknowledged that IL-8-C-X-C motif chemokine receptor 2 (CXCR2) axis powerfully recruits neutrophils and promotes NETs formation.231 However, the clinical trial of anti-IL-8 monoclonal antibody BMS-986253 for COVID-19 has been terminated according to an interim analysis that suggested its ineffectiveness (NCT04347226). Furthermore, there are no registered trials of CXCR2 inhibitors for the treatment of COVID-19 patients.
Phosphodiesterases (PDEs) regulate the levels of cyclic adenosine monophosphate (cAMP) and cyclic guanosine monophosphate (cGMP) and play a vital role in inflammation.232 The cAMP-specific PDE4 is highly expressed and promotes inflammation in neutrophils. Thus, several PDE inhibitors are considered as possible drugs for COVID-19 treatment.233 The phase III clinical trials of PDE4 selective-inhibitor apremilast and ensifentrine (NCT04590586, NCT02735707) for COVID-19 treatment are ongoing. Dipyridamole and Pentoxifylline are FDA-approved nonspecific PDE inhibitors. It is reported that dipyridamole suppresses SARS-CoV-2 replication in infected cells and contributes to disease relief with significantly reduced D-dimer in COVID-19 patients.234 The randomized phase II (NCT04391179) and III (NCT04410328) clinical trials to evaluate the outcomes of dipyridamole for COVID-19 patients have been completed without results published. The decreased serum lactate dehydrogenase (LDH) level and increased lymphocyte count were observed in COVID-19 patients receiving pentoxifylline treatment.235 The efficacy of pentoxifylline for COVID-19 is being evaluated in the clinical trial (NCT04433988).
Ferroptosis and SARS-CoV-2
Ferroptosis in SARS-CoV-2 infection
Ferroptosis is a new-type PCD mainly caused by ROS-dependent lipid peroxidation, which was first systematically described by R. Stockwell in 2012.236 The morphology of ferroptosis is indicated by smaller mitochondria with increased membrane densities, reduced crista, and ruptured outer membrane. Ferroptosis has been implicated in numerous organ injuries and degenerative pathologies.237 Ferroptosis can be triggered through two main pathways: the extrinsic pathway (also known as the transporter-dependent pathway) and the intrinsic pathway (also known as the enzyme-regulated pathway). Iron uptake is initiated by transferrin receptor 1 (TfR1)-dependent endocytosis and Fe2+ is released into cytoplasm via six-transmembrane epithelial antigen of prostate 3 (Steap3) and divalent metal transporter 1 (DMT1). Intracellular iron is stored in ferritin (which can further release Fe2+ via ferritinophagy) and exported to the extracellular by ferroportin. Meanwhile, iron overload leads to mitochondrial ROS generation. Polyunsaturated fatty acids (PUFAs) are synthesized to phospholipid hydroperoxides (PLOOHs) mediated by acyl-CoA synthetase long-chain family member 4 (ACSL4), lysophosphatidylcholine acyltransferase 3 (LPCAT3), and lipoxygenases (LOXs), termed as lipid peroxidation, resulting in ferroptosis. The cellular antioxidant system exhibits ferroptosis-suppressing effects, particularly the cystine-glutathione (GSH)-glutathione peroxidase 4 (GPX4) axis with potent inhibition of lipid peroxidation.
Mounting evidence has suggested that ferroptosis takes part in the pathogenesis of COVID-19. Ferroptosis was detected in samples of human hearts238 and hamsters’ lungs239,240 with SARS-CoV-2 infection. The changes of iron metabolism markers in blood samples (including decreased serum iron, and increased ferritin) demonstrated iron overload and are associated with severe COVID-19.241,242,243,244 The secretion of IL-6 is highly induced by inflammation upon SARS-COV-2 infection. Previous studies have shown the effects of IL-6 on iron metabolism. On the one hand, IL-6 can directly boost transferrin uptake and ferritin expression245; on the other hand, IL-6 can induce the synthesis of hepcidin,246 an inhibitor of ferroportin, consequently leading to cellular iron accumulation. Several studies showed that increased serum hepcidin correlated with the severity of COVID-19.244,247 In addition, the scRNA-seq data of PBMC, T cells, and B cells from COVID-19 patients revealed that ferroptosis-related genes (including GPX4, FTH1, FTL, and SAT1) were increased in the acute phase and decreased in the recovery phase.55 Besides, SARS-CoV-2 was reported to downregulate the GPX4 mRNA level in infected Vero E6 cells.248 The induction of ferroptosis by SARS-CoV-2 could be reversed through two ACSL4 inhibitors in vitro.249 The ferroptosis triggered by SARS-CoV-2 is shown in Fig. 5.
The ferroptosis upon SARS-CoV-2 infection. Ferroptosis is triggered by lipid peroxidation resulting from the imbalance among iron metabolism, ROS generation, and antioxidant system (especially the cystine-GSH-GPX4 axis). The high level of IL-6 in COVID-19 directly promotes the expression of iron accumulation proteins (including transferrin and ferritin) and indirectly inhibits the exported protein ferroportin via hepcidin, leading to cell ferroptosis. Meanwhile, SARS-CoV-2 infection exhibits a negative effect on GPX4. SARS-CoV-2 severe acute respiratory syndrome coronavirus 2, IL-6 interleukin-6, TfR1 transferrin receptor 1, Steap3 six-transmembrane epithelial antigen of prostate 3, DMT1 divalent metal transporter 1, ROS reactive oxygen species, PUFAs polyunsaturated fatty acids, PLOOHs phospholipid hydroperoxides, ACSL4 acyl-CoA synthetase long-chain family member 4, LPCAT3 lysophosphatidylcholine acyltransferase 3, LOXs lipoxygenases, GSH glutathione, GPX4 glutathione peroxidase 4. Created with BioRender
Ferroptosis-targeted therapies
Given the harmfulness of ferroptosis in COVID-19, it is reasonable to protect cells from ferroptosis via three aspects: alleviate iron overload250; augment the cystine-GSH-GPX4 axis; inhibit lipid peroxidation. Deferoxamine, an FDA-approved injective iron chelator, binds with free iron and iron from lysosomal ferritin, forming a stable complex removed by kidneys.251,252 Deferiprone is another iron chelator approved by FAD for use of oral, being more powerful for the chelation of iron in the heart compared to deferoxamine. The clinical trials of iron chelation therapy for COVID-19 are ongoing (NCT04333550, NCT04361032, NCT04389801). Intriguingly, the available data suggested that the selenium level is positively associated with clinical complications and outcomes,253,254 which supports selenium supplementation in COVID-19 patients.255 Selenium was reported to induce GPX4 expression to protect cells from ferroptosis in the model of stroke256 and further applied for COVID-19 (NCT04869579, NCT04798677, NCT04751669, NCT04323228). Ebselen, a synthetic selenium compound, can not only impede ferroptosis by acting as a GPX mimic but also inhibit virus replication by interacting with SARS-CoV-2 Mpro.257,258 N-acetylcysteine (NAC), a precursor of GSH, possesses anti-ferroptosis activity through directly reinforcing the cystine-GSH-GPX4 axis259 and reducing the IL-6-promoted ROS production,260 accounting for a part of its effects for COVID-19 treatment. Additionally, it has been proven that NAC can regulate T cell response261 and inflammation.261 However, in a randomized, double-blind, placebo-controlled trial, high-dose NAC failed to prevent the progression of ARDS in COVID-19 patients.262 More clinical trials of NAC as monotherapy and combination therapy for COVID-19 are still ongoing (NCT04374461, NCT04928495, NCT04792021, NCT04703036, NCT05074121, NCT04455243).
It is well known that lipid peroxidation is the most important step in ferroptosis, thus blocking key enzymes (such as ACSL4 and LOX263) of this process as a potential approach for COVID-19 treatment. Pioglitazone and Troglitazone, two FDA-approved thiazolidinediones for diabetes, can reduce cell ferroptosis and viral replication by selectively inhibiting ACSL4 activity during SARS-CoV-2 infection.264,265 Two randomized phase IV clinical trials of pioglitazone utilized in patients with COVID-19 concurrently with type 2 diabetes are recruiting (NCT04604223 and NCT04535700).
Autophagy and SARS-CoV-2
Autophagy core machinery
Autophagy (also named type II PCD) is a highly conserved catabolic process to degrade and recycle misfolded proteins and damaged organelles to maintain cell homeostasis and eliminate intracellular pathogens, which was first elucidated in yeast by Ohsumi 1992.266,267 The morphological characteristics of autophagy manifest cytoplasmic vacuoles and autophagic bodies. Autophagy is classified into three types: macroautophagy (referred to as autophagy), microautophagy, and selective autophagy. In canonical autophagy, subcellular membranes (such as ER, Golgi complex, mitochondria, and endosomes) engulf the undesired components to form a double-membrane structure named phagophore. The phagophore membrane expands and elongates until it closes in on itself to form the autophagosome, which further fuse with lysosome to form the autophagolysosome for degradation under acidic conditions.268 The whole process is regulated by a series of proteins encoded by autophagy‐related genes (ATGs),269 mainly including: (i) the unc-51 like autophagy activating kinase 1 (ULK1) complex; (ii) the phosphatidylinositol 3-phosphate class III lipid kinase (PIK3C3) complex; (iii) the ubiquitin-like ATG12-conjunction system; (iv) the ubiquitin-like microtubule-associated protein 1 light chain 3 (MAP1LC3, termed as LC3)-conjunction system. Besides, the undesired components are labeled by sequestosome 1 (SQSTM1, also named as p62) complex and brought into the autophagosome for degradation. In addition, the fusion step is mediated by the homotypic fusion and protein sorting (HOPS) complex and N-ethylmaleimide sensitive factor attachment protein receptor (SNARE), and vesicle associated membrane protein 8 (VAMP8) complex.
Autophagy induction by SARS-CoV-2
Growing studies have suggested that the interplay of SARS-CoV-2 and autophagy is extremely intricate, which has not been fully established. The double-membrane vesicles (DMVs) were detected by transmission electron microscopy270 and further confirmed via integrative imaging (including FIB-SEM analysis, electron tomography, and 3D rendering)271 in SARS-CoV-2-infected nasal, bronchial, and pulmonary human epithelial cells. SARS-CoV-2 NSP6, which co-localizes in ER, has been reported to promote rearrangements of ER membranes to DMVs production.272 Thus, like other coronaviruses,273 SARS-CoV-2 can hijack autophagy flux to form DMVs derivated from ER membranes. As of current studies, these DMVs provide the niches for viral RNA replication and immune escape. In addition, recent evidence has concluded that viral proteins encoded by SARS-CoV-2 can promote autophagy to regulate the host’s immune response. It has been reported that SARS-CoV-2 M protein translocates to mitochondria via mitochondrial Tu translation elongation factor (TUFM) and interacts with LC3-II through its LC3-interacting region (LIR), thus inducing mitochondrial autophagy (termed as mitophagy).274 In a similar way, SARS-CoV-2 ORF10 directly interacts with a mitochondrial receptor Bcl-2 interacting protein 3 like (BNIP3L)/NIX (also containing LIR) to promote mitophagy.275 On the one hand, activated mitophagy causes the degradation of mitochondrial antiviral signaling protein (MAVS). Previous studies have indicated that MAVS can activate transcription factors interferon regulatory factor 3 (IRF3) and NF-κB to upregulate the expression of immune response and inflammation-related genes.276 Thus, SARS-CoV-2 M and ORF10 abate type I IFN response and pro-inflammatory cytokines by inducing mitophagy-mediated MAVS degradation. On the other hand, mitophagy decreases the release of mtDNA and ROS, thus inhibiting NLRP3 inflammasome-mediated cell pyroptosis and cytokine secretion. Additionally, type I IFN production is also diminished due to SARS-CoV-2 NSP13 recruiting TBK1 (via RecA domain) to p62 (via ZBD domain) for autophagic degradation of TBK1. Interestingly, the direct interaction of SARS-CoV-2 ORF8 with major histocompatibility complex I (MHC I) leads to MHC I lysosomal degradation, reduction of antigen presentation, and evading immune surveillance in SARS-CoV-2-infected cells and mice model.277
Autophagy inhibition by SARS-CoV-2
A great number of reports have demonstrated that autophagy is an anti-viral process, for reason that it not only impedes the viral life cycle by eliminating viral particles and viral components (also known as virophage) but also mediates antigen presentation to induce adaptive immunity. Therefore, SARS-CoV-2 could inhibit autophagy through multifarious mechanisms to facilitate its replication and transmission.278 SARS-CoV-2 ORF3a prohibits cell autophagy in a variety of ways. SAS-CoV-2 ORF3a interacting with autophagy regulator UV irradiation resistance-associated gene (UVRAG)279 promotes PI3KC3-C1 but blocks PI3KC3-C2, leading to incomplete autophagy.280 Incomplete autophagy represents the inhibition of autophagosome-lysosome fusion, that is autophagosome induction but autolysosome reduction, also termed as the failure of autophagosome turnover. SAS-CoV-2 ORF3a-induced ER stress can also trigger incomplete autophagy by activating transcription factor 6 (ATF6)- and inositol-requiring enzyme 1 (IRE-1)-dependent unfolded protein response (UPR).281 Moreover, another two studies have demonstrated that SARS-CoV-2 ORF3a directly interacts with and sequestrates VPS39 in endosomes and lysosomes, resulting in the dysfunctional HOPS complex. For one thing, VPS39 sequestration disturbs the interaction of HOPS with Ras-related protein Rab-7 (RAB7).282 For another, invalid HOPS blocks the assembly of SNARE complex via directly interacting with STX17,283 thereby preventing the fusion of autophagosomes with lysosomes. In addition, SARS-CoV-2 ORF3a can promote viral egress through lysosomal exocytosis.114 This process is regulated by Ca2+ channel TRPML3, lysosomal trafficker BORC-ARL8b complex, and exocytosis-related STX4-SNAP23-VAMP7 SNARE complex. Like ORF3a, SARS-CoV-2 ORF7a also co-localizes with late endosomes and disturbs lysosomal acidification, resulting in blockage of autophagosome turnover.284 Besides, SARS-CoV-2 NSP15 was reported to affect the production of autophagosomes via interacting with the mTOR axis, which could be reversed by rapamycin (an mTOR inhibitor).284 It is known that the mTOR axis exerts inhibitory effects on autophagy via prohibiting the ULK1 complex.285 Meanwhile, SARS-CoV-2 PLpro protein directly cleaves ULK1 to the destruction of ULK1-ATG13 complex formation.286 The complicated mechanisms of cell autophagy mediated by SARS-CoV-2 are summarized in Fig. 6.
The complicated mechanisms of SARS-CoV-2 manipulating cell autophagy. The phagophore derived from subcellular membranes expands and elongates to form the autophagosome, further fusing with lysosome to form the autophagolysosome for degradation, whose processes are closely regulated by ATGs. SARS-CoV-2 hijacks ER-derivated DMVs for replication and immune escape, which are enhanced by SARS-CoV-2 NSP6. SARS-CoV-2 M protein (via TUFM), ORF10 (via NIX), and NSP13 can induce mitochondrial autophagy to reduce the activation of NF-κB and IRF. Besides, SARS-CoV-2 ORF8 promotes MHC I degradation. In contrast, SARS-CoV-2 also plays an important role in autophagy inhibition. SARS-CoV-2 PLpro-cleaved ULK1 and SARS-CoV-2 NSP15-activated mTOR cause ULK1 complex inhibition. SARS-CoV-2 ORF3a and SARS-CoV-2 infection both regulate the PIK3C3 complex, all of which lead to autophagy inhibition. SARS-CoV-2 ORF3a, ORF7a, and NSP6 impede the fusion of lysosome with autophagosome via damaging lysosomal acidification. Besides, SARS-CoV-2 ORF3a directly impairs the interactions among membrane fusion complex HOPS, SNARE and VAMP8. SARS-CoV-2 severe acute respiratory syndrome coronavirus 2, M membrane, ORF open reading frame, NSP nonstructural protein, PLpro papain-like protease, ULK1 unc-51 like autophagy activating kinase 1, ATG autophagy‐related gene, FIP200 focal adhesion kinase family interacting protein of 200 kD, PIK3C3 complex phosphatidylinositol 3-phosphate class III lipid kinase complex, UPRs unfolded protein responses, ATF6 activating transcription factor 6, IRE-1 inositol-requiring enzyme 1, MAPK mitogen-activated protein kinase, ERK extracellular-signal regulated kinase, PI3K phosphoinositide 3-kinase, AKT protein kinase B, mTOR mammalian target of rapamycin, UVRAG UV irradiation resistance-associated gene, LC3 microtubule-associated protein 1 light chain 3, PE phosphatidylethanolamine, SQSTM1 sequestosome 1, NBR1 neighbor of BRCA1 gene 1, Ambra1 Beclin-1-regulated autophagy, HOPS homotypic fusion and protein sorting, RAB7 Ras-related protein Rab7, STX17 syntaxin 17, SNAP29 synaptosome associated protein 29, VAMP8 vesicle associated membrane protein 8, MAVS mitochondrial anti-viral signaling protein, NF-κB nuclear factor kappa B, IRF transcription factors interferon regulatory factor, TUFM mitochondrial Tu translation elongation factor, NIX Bcl-2 interacting protein 3 like, DMVs double-membrane vesicles, MHC I major histocompatibility complex I, ER endoplasmic reticulum. Created with BioRender
Taken together, cell autophagy has a double face during SARS-CoV-2 infection. On the one hand, autophagy exerts its anti-viral activity by induction of virophage and regulation of immune response; On the other hand, autophagy is manipulated by SARS-CoV-2 to favor their replication and transmission, immunosurveillance escape, and pro-inflammatory response. Furthermore, accumulative autophagosomes would induce the Bcl-2/BAX-mediated cell apoptosis and the NLRP3 inflammasome-dependent cell pyroptosis (see previous sections), resulting in cell damage and tissue injury.
Autophagy-targeted therapies
Although the intricate relation is still obscure, cell autophagy is important for SARS-CoV-2 infection and host response, providing rationality for autophagy mediators as a therapeutic strategy. There are a large number of drugs under clinical investigation. It should be noted that most of them exert anti-viral activity by regulating autophagy as well as other mechanisms. Chloroquine (CQ) and its less toxic derivative, hydroxychloroquine (HCQ), two FDA-approved anti-malarial drugs, exhibit autophagy inhibition, immunosuppressive functions, anti-inflammatory properties as well as anti-viral activity, which has been proposed against SARS-CoV,287 HIV,288 and Zika.289 The previous study has shown that they can inhibit autophagic flux by disturbing autophagosome-lysosome fusion290 and endosomal/lysosomal acidification,291 resulting in accumulative autophagosomes-induced cell death and cessation of virus replication. In addition, they could impede SARS-CoV-2 entry by restricting the glycosylation of ACE2 and S protein292 and inhibiting acidification-dependent endocytosis.293 Several in vitro studies showed that CQ and HCQ made inhibitory effects on SARS-CoV-2-infected Vero E6 cells.293,294,295 A prospective observational study indicated that CQ provided positive effects on reduced viral load and time to clinical recovery (TTCR) in COVID-19 patients, compared with historical controls.296 A randomized clinical trial showed that HCQ treatment improved the TTCR and absorption of pneumonia in COVID-19 patients, compared with SoC-treated patients.297 In addition, another non-randomized, small-size trial suggested that HCQ decreased viral load in COVID-19, whose effect could be reinforced by azithromycin.298 These positive results pay the way to further investigation of CQ and HCQ in large-scale and randomized clinical trials. Nevertheless, the majority of them presented disappointing outcomes. In a randomized, SoC-controlled phase II/III clinical trial, HCQ exhibited no significant effect on the 28-day mortality and hospital stay duration in hospitalized patients with COVID-19 (NCT04381936).299 Another randomized, double-blind phase III trial also obtained negative results that no significant difference at day 14 in hospitalized COVID-19 treated with HCQ and placebo (NCT04332991).300 In light of current results, it is indicated that the anti-viral effects of CQ/HCQ are controversial. Given its diverse effects, CQ/HCQ are still considered as potential drugs, thus employment of combination therapies and pre/post-exposure prophylaxis might be the further direction. Furthermore, autophagy inhibition can be achieved by inhibiting autophagy-related complexes, among which targeting the ULK1 complex and PIK3C3 complex are most studied. Consequently, several studies showed that PIK3C3 complex inhibitors potently reduced SARS-CoV-2 replication,301,302 among which commonly applied to impede autophagy are VPS34-IN1, VVPS34-IN1, 3-methyladenine,303 wortmannin, LY294002,304 PT210,305 and GSK-2126458,306. Inhibitors of the ULK1 complex (including ULK-101,307 compound 6,308 MRT68921,309 and SBI-0206965310) are also identified as autophagy inhibitors and need further investigation for antagonizing SARS-CoV-2 infection.
In the light of uncontrolled inflammatory response in severe COVID-19, the administration of autophagy activators for anti-inflammation and anti-virus also might be helpful. It is known that mTOR signaling plays a vital role in mediating cell autophagy. AKT and MAPK-activated mTOR can inhibit autophagy, whereas AMPK and p53-suppressed mTOR induce autophagy. The proteotranscriptomic data displayed activation of the PI3K/AKT/mTOR pathway in response to SARS-CoV-2 infection.311 In line with this result, the mTOR activation was confirmed by in-depth analyses, and AKT inhibitor MK-2206 inhibited SARS-CoV-2 replication.312 Rapamycin (also known as Sirolimus), an FDA-approved mTOR inhibitor, showed positive anti-viral activity on porcine epidemic diarrhea virus,313 transmissible gastroenteritis virus,314 H1N1,315 and MERS-CoV316 in previous studies. The clinical trials investing the effects of sirolimus treatment for COVID-19 are ongoing (NCT04461340, NCT04341675, NCT04482712). Metformin, an FDA-approved antidiabetic drug, promotes autophagy through upregulation of AMPK and downregulation of mTOR.317,318 Furthermore, increasing evidence has demonstrated that it can directly block viral entry and infection, and mediate inflammatory and immune responses,319,320 which provides a basis for rational application in COVID-19 patients. In a retrospective analysis, 30-day mortality was significantly decreased (HR 0.48) in nursing home patients with metformin treatment, compared to that in patients with no metformin-containing regimens.321 Other retrospective studies showed similar results in hospitalized patients with SARS-CoV-2 infection.322,323 A randomized, double-blind, placebo-controlled phase II trial of metformin for hospitalized COVID-19 patients with ARDS has been completed with no result published yet (NCT04625985). Other clinical trials of metformin-treated with COVID-19 are ongoing (NCT04604678, NCT04510194, NCT04727424). Additionally, a number of drugs (such as resveratrol, trehalose, ivermectin, nitazoxanide, etc) with the ability to induce autophagy to anti-virus have been under investigation. However, the underlying mechanisms of these drugs for COVID-19 treatment still have been obscure.
Conclusions and perspectives
Taken together, SARS-CoV-2 can regulate cell death and cell autophagy through various mechanisms and participate in the occurrence and development of COVID-19. From the above description, it is not hard to conclude that cell death and autophagy are a double-edged sword. On the one hand, they can directly prevent the propagation of viruses through the reduction in replicative vehicles. On the other hand, they would lead to tissue destruction, widespread virus, cytokine storm, and incompetent immune response. At different times of infection, SARS-CoV-2 can craftily manipulate cell death and autophagy to serve itself. Thus, it should be carefully decided whether and when cell death and autophagy should be inhibited or stimulated. With the help of massive inhibitors and agonists targeting these pathways, further researches are able to dive into the diverse influences of cell death and autophagy on COVID-19 development. In addition, high-throughput methods including single-cell sequencing, proteomics and metabonomics, and spatial omics may facilitate the discovery of novel pathways in COVID-19 and the cross-talk among different cell death and autophagy pathways. A good question is whether disparate cell death and autophagy pathways are induced in different strains of SARS-CoV-2. More delicate observations are needed to figure out the relationship between SARS-CoV-2 mutations and altered cell death and autophagy pathways.
In view of the important roles of cell death and autophagy, a variety of related therapeutic strategies have been explored and put into practice in clinical trials, whereas the results are controversial (Table 1). Thereby, the interactions among SARS-CoV-2 infection, host response, and therapies need more in-depth and comprehensive research. Meanwhile, it is indeed crucial to estimate the therapeutic window for COVID-19 patients according to the course and severity of the disease. The real-time and dynamic information on clinical manifestations, laboratory parameters, and imaging findings, combined with relevant high-throughput methods may be helpful in finding the critical point for therapies. It may be hard but beneficial to create a guide to balance ameliorating symptoms and preventing viruses from spreading in patients.
References
Zhu, N. et al. A novel coronavirus from patients with pneumonia in China, 2019. N. Engl. J. Med. 382, 727–733 (2020).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271–280 (2020). e278.
Wang, K. et al. CD147-spike protein is a novel route for SARS-CoV-2 infection to host cells. Signal Transduct. Target Ther. 5, 283 (2020).
Hoffmann, M., Kleine-Weber, H. & Pohlmann, S. A multibasic cleavage site in the spike protein of SARS-CoV-2 is essential for infection of human lung cells. Mol. Cell 78, 779–784 (2020). e775.
Ou, X. et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry and its immune cross-reactivity with SARS-CoV. Nat. Commun. 11, 1620 (2020).
V’Kovski, P. et al. Coronavirus biology and replication: implications for SARS-CoV-2. Nat. Rev. Microbiol. 19, 155–170 (2021).
Gheblawi, M. et al. Angiotensin-converting enzyme 2: SARS-CoV-2 receptor and regulator of the Renin-angiotensin system: celebrating the 20th Anniversary of the discovery of ACE2. Circ. Res. 126, 1456–1474 (2020).
Rodriguez-Lago, I. et al. Characteristics and prognosis of patients with inflammatory bowel disease during the SARS-CoV-2 pandemic in the basque country (Spain). Gastroenterology 159, 781–783 (2020).
Galluzzi, L. et al. Molecular mechanisms of cell death: recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death Differ. 25, 486–541 (2018).
Imre, G. The involvement of regulated cell death forms in modulating the bacterial and viral pathogenesis. Int Rev. Cell Mol. Biol. 353, 211–253 (2020).
Demarco, B., Chen, K. W. & Broz, P. Cross talk between intracellular pathogens and cell death. Immunol. Rev. 297, 174–193 (2020).
Zhao, Z. et al. The interplay between emerging human coronavirus infections and autophagy. Emerg. Microbes Infect. 10, 196–205 (2021).
Tan, Y. J., Lim, S. G. & Hong, W. Regulation of cell death during infection by the severe acute respiratory syndrome coronavirus and other coronaviruses. Cell Microbiol. 9, 2552–2561 (2007).
Fung, T. S. & Liu, D. X. Human coronavirus: host-pathogen interaction. Annu Rev. Microbiol. 73, 529–557 (2019).
Gordon, D. E. et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 583, 459–468 (2020).
Abdalla, A. E. et al. Insight into the emerging role of SARS-CoV-2 nonstructural and accessory proteins in modulation of multiple mechanisms of host innate defense. Bosn. J. Basic Med. Sci. 21, 515–527 (2021).
Kerr, J. F., Wyllie, A. H. & Currie, A. R. Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. Br. J. Cancer 26, 239–257 (1972).
Singh, R., Letai, A. & Sarosiek, K. Regulation of apoptosis in health and disease: the balancing act of BCL-2 family proteins. Nat. Rev. Mol. Cell Biol. 20, 175–193 (2019).
Carneiro, B. A. & El-Deiry, W. S. Targeting apoptosis in cancer therapy. Nat. Rev. Clin. Oncol. 17, 395–417 (2020).
Bedoui, S., Herold, M. J. & Strasser, A. Emerging connectivity of programmed cell death pathways and its physiological implications. Nat. Rev. Mol. Cell Biol. 21, 678–695 (2020).
Lavrik, I. N. & Krammer, P. H. Regulation of CD95/Fas signaling at the DISC. Cell Death Differ. 19, 36–41 (2012).
Wyler, E. et al. Transcriptomic profiling of SARS-CoV-2 infected human cell lines identifies HSP90 as target for COVID-19 therapy. iScience 24, 102151 (2021).
Li, X., Liu, K., Zhou, W. & Jiang, Z. MiR-155 targeting FoxO3a regulates oral cancer cell proliferation, apoptosis, and DDP resistance through targeting FoxO3a. Cancer Biomark. 27, 105–111 (2020).
Kucharczak, J., Simmons, M. J., Fan, Y. & Gelinas, C. To be, or not to be: NF-kappaB is the answer-role of Rel/NF-kappaB in the regulation of apoptosis. Oncogene 22, 8961–8982 (2003).
Blanco-Melo, D. et al. Imbalanced host response to SARS-CoV-2 drives development of COVID-19. Cell 181, 1036–1045 e1039 (2020).
Sun, J. et al. Comparative transcriptome analysis reveals the intensive early stage responses of host cells to SARS-CoV-2 infection. Front. Microbiol. 11, 593857 (2020).
Khan, S. et al. SARS-CoV-2 spike protein induces inflammation via TLR2-dependent activation of the NF-kappaB pathway. bioRxiv 10, e68563 (2021).
Lu, W. et al. Severe acute respiratory syndrome-associated coronavirus 3a protein forms an ion channel and modulates virus release. Proc. Natl Acad. Sci. USA 103, 12540–12545 (2006).
Chan, C. M. et al. The ion channel activity of the SARS-coronavirus 3a protein is linked to its pro-apoptotic function. Int J. Biochem. Cell Biol. 41, 2232–2239 (2009).
Issa, E. et al. SARS-CoV-2 and ORF3a: nonsynonymous mutations, functional domains, and viral pathogenesis. mSystems. https://doi.org/10.1101/2021.03.16.435700 (2020).
Bianchi, M., Borsetti, A., Ciccozzi, M. & Pascarella, S. SARS-Cov-2 ORF3a: mutability and function. Int J. Biol. Macromol. 170, 820–826 (2021).
Ren, Y. et al. The ORF3a protein of SARS-CoV-2 induces apoptosis in cells. Cell Mol. Immunol. 17, 881–883 (2020).
Yang, R. et al. SARS-CoV-2 accessory protein ORF7b mediates tumor necrosis factor-alpha-induced apoptosis in cells. Front Microbiol. 12, 654709 (2021).
Xia, B. et al. SARS-CoV-2 envelope protein causes acute respiratory distress syndrome (ARDS)-like pathological damages and constitutes an antiviral target. Cell Res. 31, 847–860 (2021).
Nieto-Torres, J. L. et al. Severe acute respiratory syndrome coronavirus E protein transports calcium ions and activates the NLRP3 inflammasome. Virology 485, 330–339 (2015).
Llambi, F. et al. BOK is a non-canonical BCL-2 family effector of apoptosis regulated by ER-associated degradation. Cell 165, 421–433 (2016).
Yang, Y. et al. SARS-CoV-2 membrane protein causes the mitochondrial apoptosis and pulmonary edema via targeting BOK. Cell Death Differ. https://doi.org/10.1038/s41418-022-00928-x (2022).
Ren, Y. et al. SARS-CoV-2 membrane glycoprotein M triggers apoptosis with the assistance of nucleocapsid protein N in cells. Front. Cell Infect. Microbiol. 11, 706252 (2021).
Jiang, Y. et al. ROS-dependent activation of autophagy through the PI3K/Akt/mTOR pathway is induced by hydroxysafflor yellow A-Sonodynamic therapy in THP-1 macrophages. Oxid. Med. Cell. Longev. 2017, 1–16 (2017).
Vucicevic, L. et al. Compound C induces protective autophagy in cancer cells through AMPK inhibition-independent blockade of Akt/mTOR pathway. Autophagy 7, 40–50 (2011).
Eisenberg-Lerner, A., Bialik, S., Simon, H. U. & Kimchi, A. Life and death partners: apoptosis, autophagy and the cross-talk between them. Cell Death Differ. 16, 966–975 (2009).
Li, F. et al. SARS-CoV-2 spike promotes inflammation and apoptosis through autophagy by ROS-suppressed PI3K/AKT/mTOR signaling. Biochim. Biophys. Acta Mol. Basis Dis. 1867, 166260 (2021).
Lopes, L. R. Functional and tissue enrichment analyses suggest that SARS-CoV-2 infection affects host metabolism and catabolism mediated by interference on host proteins. Braz. J. Microbiol. 52, 1151–1159 (2021).
Padhan, K., Minakshi, R., Towheed, M. A. B. & Jameel, S. Severe acute respiratory syndrome coronavirus 3a protein activates the mitochondrial death pathway through p38 MAP kinase activation. J. Gen. Virol. 89, 1960–1969 (2008).
Kopecky-Bromberg, S. A., Martinez-Sobrido, L. & Palese, P. 7a protein of severe acute respiratory syndrome coronavirus inhibits cellular protein synthesis and activates p38 mitogen-activated protein kinase. J. Virol. 80, 785–793 (2006).
Chan, E. et al. Molecular and genetic characterisation of the SARS coronavirus auxiliary protein X1 in drosophila. Hong. Kong Med. J. 14, 14–16 (2008).
Ye, Z., Wong, C. K., Li, P. & Xie, Y. A SARS-CoV protein, ORF-6, induces caspase-3 mediated, ER stress and JNK-dependent apoptosis. Biochim Biophys. Acta 1780, 1383–1387 (2008).
Surjit, M. et al. The SARS coronavirus nucleocapsid protein induces actin reorganization and apoptosis in COS-1 cells in the absence of growth factors. Biochem J. 383, 13–18 (2004).
Nelli, R. K. et al. Enhanced apoptosis as a possible mechanism to self-limit SARS-CoV-2 replication in porcine primary respiratory epithelial cells in contrast to human cells. Cell Death Disco. 7, 383 (2021).
Zhu, N. et al. Morphogenesis and cytopathic effect of SARS-CoV-2 infection in human airway epithelial cells. Nat. Commun. 11, 3910 (2020).
Bridges, J. P., Vladar, E. K., Huang, H. & Mason, R. J. Respiratory epithelial cell responses to SARS-CoV-2 in COVID-19. Thorax 77, 203–209 (2022).
Krishnamachary, B. et al. Extracellular vesicle-mediated endothelial apoptosis and EV-associated proteins correlate with COVID-19 disease severity. J. Extracell. Vesicles 10, e12117 (2021).
Wang, W. et al. SARS-CoV-2 N protein induces acute kidney injury via Smad3-dependent G1 cell cycle arrest mechanism. Adv. Sci. (Weinh.) 9, e2103248 (2022).
Wu, C. T. et al. SARS-CoV-2 infects human pancreatic beta cells and elicits beta cell impairment. Cell Metab. 33, 1565–1576 (2021). e1565.
Huang, L. et al. Dynamic blood single-cell immune responses in patients with COVID-19. Signal Transduct. Target Ther. 6, 110 (2021).
Liao, M. et al. Single-cell landscape of bronchoalveolar immune cells in patients with COVID-19. Nat. Med. 26, 842–844 (2020).
Zhao, Q. et al. Lymphopenia is associated with severe coronavirus disease 2019 (COVID-19) infections: a systemic review and meta-analysis. Int J. Infect. Dis. 96, 131–135 (2020).
Bellesi, S. et al. Increased CD95 (Fas) and PD-1 expression in peripheral blood T lymphocytes in COVID-19 patients. Br. J. Haematol. 191, 207–211 (2020).
Cizmecioglu, A. et al. Apoptosis-induced T-cell lymphopenia is related to COVID-19 severity. J. Med Virol. 93, 2867–2874 (2021).
Andre, S. et al. T cell apoptosis characterizes severe covid-19 disease. Cell Death Differ. https://doi.org/10.1038/s41418-022-00936-x (2022).
Thompson, E. A. et al. Metabolic programs define dysfunctional immune responses in severe COVID-19 patients. Cell Rep. 34, 108863 (2021).
Xiang, Q. et al. SARS-CoV-2 induces lymphocytopenia by promoting inflammation and decimates secondary lymphoid organs. Front. Immunol. 12, 661052 (2021).
Kuri-Cervantes, L. et al. Comprehensive mapping of immune perturbations associated with severe COVID-19. Sci. Immunol. 5, eabd7114 (2020).
Arunachalam, P. S. et al. Systems biological assessment of immunity to mild versus severe COVID-19 infection in humans. Science 369, 1210–1220 (2020).
Zheng, J. et al. Severe acute respiratory syndrome coronavirus 2-induced immune activation and death of monocyte-derived human macrophages and dendritic cells. J. Infect. Dis. 223, 785–795 (2021).
Simpson, D. S. et al. Interferon-gamma primes macrophages for pathogen ligand-induced killing via a caspase-8 and mitochondrial cell death pathway. Immunity 55, 423–441 e429 (2022).
Liu, C. et al. Time-resolved systems immunology reveals a late juncture linked to fatal COVID-19. Cell 184, 1836–1857 e1822 (2021).
Schulte-Schrepping, J. et al. Severe COVID-19 is marked by a dysregulated myeloid cell compartment. Cell 182, 1419–1440 e1423 (2020).
Fitzgerald-Bocarsly, P., Dai, J. & Singh, S. Plasmacytoid dendritic cells and type I IFN: 50 years of convergent history. Cytokine Growth Factor Rev. 19, 3–19 (2008).
Scuto, A. et al. The novel histone deacetylase inhibitor, LBH589, induces expression of DNA damage response genes and apoptosis in Ph- acute lymphoblastic leukemia cells. Blood 111, 5093–5100 (2008).
Shin, D. et al. Papain-like protease regulates SARS-CoV-2 viral spread and innate immunity. Nature 587, 657–662 (2020).
Kandasamy, M. NF-kappaB signalling as a pharmacological target in COVID-19: potential roles for IKKbeta inhibitors. Naunyn Schmiedebergs Arch. Pharm. 394, 561–567 (2021).
Liang, N. et al. Immunosuppressive effects of hydroxychloroquine and artemisinin combination therapy via the nuclear factor-kappaB signaling pathway in lupus nephritis mice. Exp. Ther. Med. 15, 2436–2442 (2018).
Bleyzac, N., Goutelle, S., Bourguignon, L. & Tod, M. Azithromycin for COVID-19: more than just an antimicrobial? Clin. Drug Investig. 40, 683–686 (2020).
Assimakopoulos, S. F. & Marangos, M. N-acetyl-cysteine may prevent COVID-19-associated cytokine storm and acute respiratory distress syndrome. Med. Hypotheses 140, 109778 (2020).
Yang, L. et al. The signal pathways and treatment of cytokine storm in COVID-19. Signal Transduct. Target Ther. 6, 255 (2021).
Gracia-Sancho, J. et al. Emricasan ameliorates portal hypertension and liver fibrosis in cirrhotic rats through a hepatocyte-mediated paracrine mechanism. Hepatol. Commun. 3, 987–1000 (2019).
Palladino, M. A., Bahjat, F. R., Theodorakis, E. A. & Moldawer, L. L. Anti-TNF-alpha therapies: the next generation. Nat. Rev. Drug Disco. 2, 736–746 (2003).
Del Valle, D. M. et al. An inflammatory cytokine signature predicts COVID-19 severity and survival. Nat. Med. 26, 1636–1643 (2020).
Qin, C. et al. Dysregulation of immune response in patients with coronavirus 2019 (COVID-19) in Wuhan, China. Clin. Infect. Dis. 71, 762–768 (2020).
Costela-Ruiz, V. J. et al. SARS-CoV-2 infection: the role of cytokines in COVID-19 disease. Cytokine Growth Factor Rev. 54, 62–75 (2020).
Karki, R. et al. Synergism of TNF-alpha and IFN-gamma triggers inflammatory cell death, tissue damage, and mortality in SARS-CoV-2 infection and cytokine shock syndromes. Cell 184, 149–168 e117 (2021).
Hachem, H. et al. Rapid and sustained decline in CXCL-10 (IP-10) annotates clinical outcomes following TNFalpha-antagonist therapy in hospitalized patients with severe and critical COVID-19 respiratory failure. J. Clin. Transl. Sci. 5, e146 (2021).
Kreuzberger, N. et al. SARS-CoV-2-neutralising monoclonal antibodies for treatment of COVID-19. Cochrane Database Syst. Rev. 9, CD013825 (2021).
Fakharian, A. et al. Evaluation of adalimumab effects in managing severe cases of COVID-19: a randomized controlled trial. Int Immunopharmacol. 99, 107961 (2021).
Chanchlani, N. et al. Adalimumab and infliximab impair SARS-CoV-2 antibody responses: results from a therapeutic drug monitoring study in 11 422 biologic-treated patients. J. Crohns Colitis 16, 389–397 (2022).
Degterev, A. et al. Chemical inhibitor of nonapoptotic cell death with therapeutic potential for ischemic brain injury. Nat. Chem. Biol. 1, 112–119 (2005).
Pasparakis, M. & Vandenabeele, P. Necroptosis and its role in inflammation. Nature 517, 311–320 (2015).
Galluzzi, L., Kepp, O., Chan, F. K. & Kroemer, G. Necroptosis: mechanisms and relevance to disease. Annu Rev. Pathol. 12, 103–130 (2017).
Petrie, E. J., Czabotar, P. E. & Murphy, J. M. The structural basis of necroptotic cell death signaling. Trends Biochem. Sci. 44, 53–63 (2019).
Yang, D. et al. ZBP1 mediates interferon-induced necroptosis. Cell Mol. Immunol. 17, 356–368 (2020).
Xu, G. et al. SARS-CoV-2 promotes RIPK1 activation to facilitate viral propagation. Cell Res. 31, 1230–1243 (2021).
Li, S. et al. SARS-CoV-2 triggers inflammatory responses and cell death through caspase-8 activation. Signal Transduct. Target Ther. 5, 235 (2020).
Yue, Y. et al. SARS-coronavirus open reading frame-3a drives multimodal necrotic cell death. Cell Death Dis. 9, 904 (2018).
Nakamura, H. et al. Serum levels of receptor-interacting protein kinase-3 in patients with COVID-19. Crit. Care 24, 484 (2020).
Shashaty, M. G. S. et al. Plasma receptor interacting protein kinase-3 levels are associated with acute respiratory distress syndrome in sepsis and trauma: a cohort study. Crit. Care 23, 235 (2019).
Cao, L. & Mu, W. Necrostatin-1 and necroptosis inhibition: pathophysiology and therapeutic implications. Pharm. Res. 163, 105297 (2021).
Feng, L. et al. Proteome-wide data analysis reveals tissue-specific network associated with SARS-CoV-2 infection. J. Mol. Cell Biol. 12, 946–957 (2020).
Belyaeva, A. et al. Causal network models of SARS-CoV-2 expression and aging to identify candidates for drug repurposing. Nat. Commun. 12, 1024 (2021).
Riebeling, T. et al. Primidone blocks RIPK1-driven cell death and inflammation. Cell Death Differ. 28, 1610–1626 (2021).
Cookson, B. T. & Brennan, M. A. Pro-inflammatory programmed cell death. Trends Microbiol. 9, 113–114 (2001).
Bergsbaken, T., Fink, S. L. & Cookson, B. T. Pyroptosis: host cell death and inflammation. Nat. Rev. Microbiol 7, 99–109 (2009).
Yu, P. et al. Pyroptosis: mechanisms and diseases. Signal Transduct. Target Ther. 6, 128 (2021).
Kelley, N., Jeltema, D., Duan, Y. & He, Y. The NLRP3 Inflammasome: an overview of mechanisms of activation and regulation. Int. J. Mol. Sci. https://doi.org/10.3390/ijms20133328 (2019).
He, Y., Hara, H. & Nunez, G. Mechanism and regulation of NLRP3 inflammasome activation. Trends Biochem Sci. 41, 1012–1021 (2016).
Di Virgilio, F. et al. The P2X7 receptor in infection and inflammation. Immunity 47, 15–31 (2017).
Petrilli, V. et al. Activation of the NALP3 inflammasome is triggered by low intracellular potassium concentration. Cell Death Differ. 14, 1583–1589 (2007).
Lee, G. S. et al. The calcium-sensing receptor regulates the NLRP3 inflammasome through Ca2+ and cAMP. Nature 492, 123–127 (2012).
Heid, M. E. et al. Mitochondrial reactive oxygen species induces NLRP3-dependent lysosomal damage and inflammasome activation. J. Immunol. 191, 5230–5238 (2013).
Zhou, R., Yazdi, A. S., Menu, P. & Tschopp, J. A role for mitochondria in NLRP3 inflammasome activation. Nature 469, 221–225 (2011).
Tseng, W. A. et al. NLRP3 inflammasome activation in retinal pigment epithelial cells by lysosomal destabilization: implications for age-related macular degeneration. Invest Ophthalmol. Vis. Sci. 54, 110–120 (2013).
Xu, H. et al. SARS-CoV-2 viroporin encoded by ORF3a triggers the NLRP3 inflammatory pathway. Virology 568, 13–22 (2022).
He, Y. et al. NEK7 is an essential mediator of NLRP3 activation downstream of potassium efflux. Nature 530, 354–357 (2016).
Chen, D. et al. ORF3a of SARS-CoV-2 promotes lysosomal exocytosis-mediated viral egress. Dev. Cell 56, 3250–3263 e3255 (2021).
Pan, P. et al. SARS-CoV-2 N protein promotes NLRP3 inflammasome activation to induce hyperinflammation. Nat. Commun. 12, 4664 (2021).
Sun, X. et al. SARS-CoV-2 non-structural protein 6 triggers NLRP3-dependent pyroptosis by targeting ATP6AP1. Cell Death Differ. https://doi.org/10.1038/s41418-021-00916-7 (2022).
Wu, W. K. K. & Yue, J. Autophagy in host-microbe interactions. Semin Cell Dev. Biol. 101, 1–2 (2020).
Kim, N. E., Kim, D. K. & Song, Y. J. SARS-CoV-2 Nonstructural proteins 1 and 13 suppress Caspase-1 and the NLRP3 inflammasome activation. Microorganisms 9, 494 (2021).
Shi, F. et al. Coronaviruses Nsp5 antagonizes porcine gasdermin D-mediated pyroptosis by cleaving pore-forming p30 fragment. mBio 13, e0273921, (2022).
Ma, J. et al. SARS-CoV-2 nucleocapsid suppresses host pyroptosis by blocking Gasdermin D cleavage. EMBO J. 40, e108249 (2021).
Singh, K. D. & Karnik, S. S. Angiotensin receptors: structure, function, signaling and clinical applications. J Cell Signal. 1, 111 (2016).
Zhao, M. et al. Angiotensin II stimulates the NLRP3 inflammasome to induce podocyte injury and mitochondrial dysfunction. Kidney Dis. (Basel) 4, 83–94 (2018).
Mori, J., Oudit, G. Y. & Lopaschuk, G. D. SARS-CoV-2 perturbs the renin-angiotensin system and energy metabolism. Am. J. Physiol. Endocrinol. Metab. 319, E43–E47 (2020).
Ratajczak, M. Z. et al. SARS-CoV-2 entry receptor ACE2 is expressed on very small CD45(-) precursors of hematopoietic and endothelial cells and in response to virus spike protein activates the Nlrp3 inflammasome. Stem Cell Rev. Rep. 17, 266–277 (2021).
Chouaki Benmansour, N., Carvelli, J. & Vivier, E. Complement cascade in severe forms of COVID-19: recent advances in therapy. Eur. J. Immunol. 51, 1652–1659 (2021).
Magro, C. et al. Complement associated microvascular injury and thrombosis in the pathogenesis of severe COVID-19 infection: a report of five cases. Transl. Res. 220, 1–13 (2020).
Zhang, Y. et al. Inflammatory response cells during acute respiratory distress syndrome in patients with coronavirus disease 2019 (COVID-19). Ann. Intern. Med. 173, 402–404 (2020).
Gao, T. et al. Highly pathogenic coronavirus N protein aggravates lung injury by MASP-2-mediated complement over-activation. medRxiv. https://doi.org/10.1101/2020.03.29.20041962 (2020).
Asgari, E. et al. C3a modulates IL-1beta secretion in human monocytes by regulating ATP efflux and subsequent NLRP3 inflammasome activation. Blood 122, 3473–3481 (2013).
Yu, S. et al. The complement receptor C5aR2 promotes protein kinase R expression and contributes to NLRP3 inflammasome activation and HMGB1 release from macrophages. J. Biol. Chem. 294, 8384–8394 (2019).
Boriushkin, E. et al. p58(IPK) suppresses NLRP3 inflammasome activation and IL-1beta production via inhibition of PKR in macrophages. Sci. Rep. 6, 25013 (2016).
Triantafilou, K., Hughes, T. R., Triantafilou, M. & Morgan, B. P. The complement membrane attack complex triggers intracellular Ca2+ fluxes leading to NLRP3 inflammasome activation. J. Cell Sci. 126, 2903–2913 (2013).
Ferreira, A. C. et al. SARS-CoV-2 engages inflammasome and pyroptosis in human primary monocytes. Cell Death Disco. 7, 43 (2021).
Moriyama, M. et al. Influenza virus-induced oxidized DNA activates inflammasomes. iScience. https://doi.org/10.1016/j.isci.2020.101270 (2020).
Shimada, K. et al. Oxidized mitochondrial DNA activates the NLRP3 inflammasome during apoptosis. Immunity 36, 401–414 (2012).
Bae, J. H. et al. Circulating cell-free mtDNA contributes to AIM2 inflammasome-mediatedchronic inflammation in patients with type 2 diabetes. Cells. https://doi.org/10.3390/cells8040328 (2019).
Singh, K. K., Chaubey, G., Chen, J. Y. & Suravajhala, P. Decoding SARS-CoV-2 hijacking of host mitochondria in COVID-19 pathogenesis. Am. J. Physiol. Cell Physiol. 319, C258–C267 (2020).
Andargie, T. E. et al. Cell-free DNA maps COVID-19 tissue injury and risk of death and can cause tissue injury. JCI Insight. 6, e147610 (2021).
Junqueira, C. et al. SARS-CoV-2 infects blood monocytes to activate NLRP3 and AIM2 inflammasomes, pyroptosis and cytokine release. Res Sq. https://doi.org/10.1101/2021.03.06.21252796 (2021).
Eisfeld, H. S. et al. Viral glycoproteins induce NLRP3 inflammasome activation and pyroptosis in macrophages. Viruses. 13, 2076 (2021).
Zhang, J. et al. Pyroptotic macrophages stimulate the SARS-CoV-2-associated cytokine storm. Cell Mol. Immunol. 18, 1305–1307 (2021).
Campbell, G. R., To, R. K., Hanna, J. & Spector, S. A. SARS-CoV-2, SARS-CoV-1, and HIV-1 derived ssRNA sequences activate the NLRP3 inflammasome in human macrophages through a non-classical pathway. iScience 24, 102295 (2021).
Ratajczak, M. Z. & Kucia, M. SARS-CoV-2 infection and overactivation of Nlrp3 inflammasome as a trigger of cytokine “storm” and risk factor for damage of hematopoietic stem cells. Leukemia 34, 1726–1729 (2020).
Kucia, M. et al. An evidence that SARS-Cov-2/COVID-19 spike protein (SP) damages hematopoietic stem/progenitor cells in the mechanism of pyroptosis in Nlrp3 inflammasome-dependent manner. Leukemia 35, 3026–3029 (2021).
Paul, O. et al. Vascular inflammation in lungs of patients with fatal coronavirus disease 2019 (COVID-19) infection: possible role for the NLRP3 inflammasome. medRxiv. https://doi.org/10.1101/2021.03.19.21253815 (2021).
Nagashima, S. et al. Endothelial dysfunction and thrombosis in patients with COVID-19-brief report. Arterioscler Thromb. Vasc. Biol. 40, 2404–2407 (2020).
Stenmark, K. R. et al. Mechanisms of SARS-CoV-2-induced lung vascular disease: potential role of complement. Pulm. Circ. 11, 20458940211015799 (2021).
Rodrigues, T. S. et al. Inflammasomes are activated in response to SARS-CoV-2 infection and are associated with COVID-19 severity in patients. J Exp. Med. 218, e20201707 (2021).
Laing, A. G. et al. A dynamic COVID-19 immune signature includes associations with poor prognosis. Nat. Med. 26, 1623–1635 (2020).
Lucas, C. et al. Longitudinal analyses reveal immunological misfiring in severe COVID-19. Nature 584, 463–469 (2020).
Toldo, S. et al. Inflammasome formation in the lungs of patients with fatal COVID-19. Inflamm. Res. 70, 7–10 (2020).
Wang, L. L., Yang, J. W. & Xu, J. F. Severe acute respiratory syndrome coronavirus 2 causes lung inflammation and injury. Clin. Microbiol. Infect. 28, 513–520 (2021).
Guo, Y.-R. et al. The origin, transmission and clinical therapies on coronavirus disease 2019 (COVID-19) outbreak—an update on the status. Mil. Med. Res. 7, 11 (2020).
Li, H., Zhang, S., Li, F. & Qin, L. NLRX1 attenuates apoptosis and inflammatory responses in myocardial ischemia by inhibiting MAVS-dependent NLRP3 inflammasome activation. Mol. Immunol. 76, 90–97 (2016).
Seth, R. B., Sun, L., Ea, C. K. & Chen, Z. J. Identification and characterization of MAVS, a mitochondrial antiviral signaling protein that activates NF-kappaB and IRF 3. Cell 122, 669–682 (2005).
Jin, Y. et al. Endothelial activation and dysfunction in COVID-19: from basic mechanisms to potential therapeutic approaches. Signal Transduct. Target Ther. 5, 293 (2020).
Tschope, C. et al. Myocarditis and inflammatory cardiomyopathy: current evidence and future directions. Nat. Rev. Cardiol. 18, 169–193 (2021).
Zeng, J. et al. Specific inhibition of the NLRP3 inflammasome suppresses immune overactivation and alleviates COVID-19 like pathology in mice. EBioMedicine 75, 103803 (2022).
Tate, M. D. et al. Reassessing the role of the NLRP3 inflammasome during pathogenic influenza A virus infection via temporal inhibition. Sci. Rep. 6, 27912 (2016).
Otani, K. et al. Colchicine prevents NSAID-induced small intestinal injury by inhibiting activation of the NLRP3 inflammasome. Sci. Rep. 6, 32587 (2016).
Magupalli, V. G. et al. HDAC6 mediates an aggresome-like mechanism for NLRP3 and pyrin inflammasome activation. Science. 369, eaas8995 (2020).
Mikolajewska, A. et al. Colchicine for the treatment of COVID-19. Cochrane Database Syst. Rev. 10, CD015045 (2021).
Lopes, M. I. et al. Beneficial effects of colchicine for moderate to severe COVID-19: a randomised, double-blinded, placebo-controlled clinical trial. RMD Open. 7, e001455 (2021).
Tardif, J.-C. et al. Efficacy of colchicine in non-hospitalized patients with COVID-19. medRxiv. https://doi.org/10.1101/2021.01.26.21250494 (2021).
Colchicine in patients admitted to hospital with COVID-19 (RECOVERY): a randomised, controlled, open-label, platform trial. Lancet Respir. Med. 9, 1419-1426, (2021).
Diaz, R. et al. Effect of colchicine vs usual care alone on intubation and 28-day mortality in patients hospitalized with COVID-19: a randomized clinical trial. JAMA Netw. Open 4, e2141328 (2021).
Hu, J. J. et al. FDA-approved disulfiram inhibits pyroptosis by blocking gasdermin D pore formation. Nat. Immunol. 21, 736–745 (2020).
Humphries, F. et al. Succination inactivates gasdermin D and blocks pyroptosis. Science 369, 1633–1637 (2020).
Chen, T. et al. Synergistic inhibition of SARS-CoV-2 replication using disulfiram/ebselen and remdesivir. ACS Pharm. Transl. Sci. 4, 898–907 (2021).
Mills, E. L. et al. Itaconate is an anti-inflammatory metabolite that activates Nrf2 via alkylation of KEAP1. Nature 556, 113–117 (2018).
Olagnier, D. et al. SARS-CoV2-mediated suppression of NRF2-signaling reveals potent antiviral and anti-inflammatory activity of 4-octyl-itaconate and dimethyl fumarate. Nat. Commun. 11, 4938 (2020).
Fillmore, N. et al. Disulfiram use is associated with lower risk of COVID-19: a retrospective cohort study. PLoS One 16, e0259061 (2021).
Yaqinuddin, A. & Kashir, J. Novel therapeutic targets for SARS-CoV-2-induced acute lung injury: targeting a potential IL-1beta/neutrophil extracellular traps feedback loop. Med. Hypotheses 143, 109906 (2020).
Abbate, A. et al. Interleukin-1 and the inflammasome as therapeutic targets in cardiovascular disease. Circ. Res. 126, 1260–1280 (2020).
Cauchois, R. et al. Early IL-1 receptor blockade in severe inflammatory respiratory failure complicating COVID-19. Proc. Natl Acad. Sci. USA 117, 18951–18953 (2020).
Xiong, S. et al. Interleukin-1RA mitigates SARS-CoV-2–induced inflammatory lung vascular leakage and mortality in humanized K18-hACE-2 mice. Arteriosclerosis, Thrombosis, Vasc. Biol. 41, 2773–2785 (2021).
Huet, T. et al. Anakinra for severe forms of COVID-19: a cohort study. Lancet Rheumatol. 2, e393–e400 (2020).
Kyriazopoulou, E. et al. An open label trial of anakinra to prevent respiratory failure in COVID-19. Elife. 10, e66125 (2021).
Tharaux, P.-L. et al. Effect of anakinra versus usual care in adults in hospital with COVID-19 and mild-to-moderate pneumonia (CORIMUNO-ANA-1): a randomised controlled trial. Lancet Respir. Med. 9, 295–304 (2021).
Mastroianni, A. et al. Early use of canakinumab to prevent mechanical ventilation in select COVID-19 patients: a retrospective, observational analysis. Int J. Immunopathol. Pharm. 35, 20587384211059675 (2021).
Katia, F. et al. Efficacy of canakinumab in mild or severe COVID-19 pneumonia. Immun., Inflamm. Dis. 9, 399–405 (2021).
Generali, D. et al. Canakinumab as treatment for COVID-19-related pneumonia: a prospective case-control study. Int J. Infect. Dis. 104, 433–440 (2021).
Landi, L. et al. Blockage of interleukin-1beta with canakinumab in patients with covid-19. Sci. Rep. 10, 21775 (2020).
Caricchio, R. et al. Effect of canakinumab vs placebo on survival without invasive mechanical ventilation in patients hospitalized with severe COVID-19: a randomized clinical trial. JAMA 326, 230–239 (2021).
Copaescu, A. et al. The role of IL-6 and other mediators in the cytokine storm associated with SARS-CoV-2 infection. J. Allergy Clin. Immunol. 146, 518–534 e511 (2020).
Potere, N. et al. The role of IL-6 and IL-6 blockade in COVID-19. Expert Rev. Clin. Immunol. 17, 601–618 (2021).
Wu, C. et al. Risk factors associated with acute respiratory distress syndrome and death in patients with coronavirus disease 2019 pneumonia in Wuhan, China. JAMA Intern Med. 180, 934–943 (2020).
Chen, X. et al. Detectable serum severe acute respiratory syndrome coronavirus 2 viral load (RNAemia) is closely correlated with drastically elevated interleukin 6 level in critically Ill patients with coronavirus disease 2019. Clin. Infect. Dis. 71, 1937–1942 (2020).
Salvarani, C. et al. Effect of tocilizumab vs standard care on clinical worsening in patients hospitalized with COVID-19 pneumonia: a randomized clinical trial. JAMA Intern Med. 181, 24–31 (2021).
Stone, J. H. et al. Efficacy of Tocilizumab in patients hospitalized with Covid-19. N. Engl. J. Med. 383, 2333–2344 (2020).
Salama, C. et al. Tocilizumab in patients hospitalized with Covid-19 pneumonia. N. Engl. J. Med. 384, 20–30 (2021).
Investigators, R.-C. et al. Interleukin-6 receptor antagonists in critically Ill patients with Covid-19. N. Engl. J. Med. 384, 1491–1502 (2021).
Group, R. C. Tocilizumab in patients admitted to hospital with COVID-19 (RECOVERY): a randomised, controlled, open-label, platform trial. Lancet 397, 1637–1645 (2021).
Declercq, J. et al. Effect of anti-interleukin drugs in patients with COVID-19 and signs of cytokine release syndrome (COV-AID): a factorial, randomised, controlled trial. Lancet Respir. Med. 9, 1427–1438 (2021).
Zhou, Y. et al. Efficacy of ion-channel inhibitors amantadine, memantine and rimantadine for the treatment of SARS-CoV-2 In Vitro. Viruses. 13, 2082 (2021).
Breitinger, U., Ali, N. K. M., Sticht, H. & Breitinger, H. G. Inhibition of SARS CoV envelope protein by flavonoids and classical viroporin inhibitors. Front Microbiol 12, 692423 (2021).
Dey, D., Borkotoky, S. & Banerjee, M. In silico identification of Tretinoin as a SARS-CoV-2 envelope (E) protein ion channel inhibitor. Comput Biol. Med 127, 104063 (2020).
Toft-Bertelsen, T. L. et al. Amantadine inhibits known and novel ion channels encoded by SARS-CoV-2 in vitro. Commun Biol. 4, 1402 (2021).
Reus, P. et al. Characterization of ACE inhibitors and AT1R antagonists with regard to their effect on ACE2 expression and infection with SARS-CoV-2 using a Caco-2 cell model. Life (Basel). 11, 810 (2021).
Nejat, R. et al. Losartan inhibits SARS-CoV-2 replication in vitro. J. Pharm. Pharm. Sci. 24, 390–399 (2021).
Rahmani, W. et al. Attenuation of SARS-CoV-2 infection by losartan in human kidney organoids. iScience 25, 103818 (2022).
Al-Karmalawy, A. A. et al. Molecular docking and dynamics simulation revealed the potential inhibitory activity of ACEIs against SARS-CoV-2 targeting the hACE2 receptor. Front Chem. 9, 661230 (2021).
Derington, C. G. et al. Angiotensin II receptor blocker or angiotensin-converting enzyme inhibitor use and COVID-19-related outcomes among US Veterans. PLoS One 16, e0248080 (2021).
Oussalah, A. et al. Long-term ACE inhibitor/ARB use is associated with severe renal dysfunction and acute kidney injury in patients with severe COVID-19: results from a referral center cohort in the northeast of france. Clin. Infect. Dis. 71, 2447–2456 (2020).
Duarte, M. et al. Telmisartan for treatment of Covid-19 patients: an open multicenter randomized clinical trial. EClinicalMedicine 37, 100962 (2021).
Puskarich, M. A. et al. A multi-center phase II randomized clinical trial of losartan on symptomatic outpatients with COVID-19. EClinicalMedicine 37, 100957 (2021).
Geriak, M. et al. Randomized prospective open label study shows no impact on clinical outcome of adding losartan to hospitalized COVID-19 patients with mild hypoxemia. Infect. Dis. Ther. 10, 1323–1330 (2021).
Brinkmann, V. et al. Neutrophil extracellular traps kill bacteria. Science 303, 1532–1535 (2004).
Thiam, H. R., Wong, S. L., Wagner, D. D. & Waterman, C. M. Cellular mechanisms of NETosis. Annu Rev. Cell Dev. Biol. 36, 191–218 (2020).
Yipp, B. G. & Kubes, P. NETosis: how vital is it? Blood 122, 2784–2794 (2013).
Mitroulis, I., Kambas, K. & Ritis, K. Neutrophils, IL-1beta, and gout: is there a link? Semin Immunopathol. 35, 501–512 (2013).
Tamassia, N. et al. Cytokine production by human neutrophils: revisiting the “dark side of the moon”. Eur. J. Clin. Invest 48, e12952 (2018).
Masso-Silva, J. A. et al. Increased peripheral blood Neutrophil activation phenotypes and neutrophil extracellular trap formation in critically Ill coronavirus disease 2019 (COVID-19) patients: a case series and review of the literature. Clin. Infect. Dis. 74, 479–489 (2022).
deKay, J. T. et al. DEspR(high) neutrophils are associated with critical illness in COVID-19. Sci. Rep. 11, 22463 (2021).
LaSalle, T. J. et al. Longitudinal characterization of circulating neutrophils uncovers distinct phenotypes associated with disease severity in hospitalized COVID-19 patients. bioRxiv. https://doi.org/10.1101/2021.10.04.463121 (2021).
Kaiser, R. et al. Self-sustaining IL-8 loops drive a prothrombotic neutrophil phenotype in severe COVID-19. JCI Insight. 6, e150862 (2021).
Veras, F. P. et al. SARS-CoV-2-triggered neutrophil extracellular traps mediate COVID-19 pathology. J. Exp .Med. 217, e20201129 (2020).
Hong, W. et al. Histones released by NETosis enhance the infectivity of SARS-CoV-2 by bridging the spike protein subunit 2 and sialic acid on host cells. Cell. Mol. Immunol. 19, 577–587 (2022).
Zuo, Y. et al. Neutrophil extracellular traps in COVID-19. JCI Insight. 5, e138999 (2020).
Nathan, C. Neutrophils and COVID-19: Nots, NETs, and knots. J Exp Med. 217, e20201439 (2020).
Aymonnier, K. et al. Inflammasome activation in neutrophils of patients with severe COVID-19. Blood Adv. 6, 2001–2013 (2022).
Middleton, E. A. et al. Neutrophil extracellular traps contribute to immunothrombosis in COVID-19 acute respiratory distress syndrome. Blood 136, 1169–1179 (2020).
Dolhnikoff, M. et al. Pathological evidence of pulmonary thrombotic phenomena in severe COVID-19. J. Thromb. Haemost. 18, 1517–1519 (2020).
Morris, G. et al. Preventing the development of severe COVID-19 by modifying immunothrombosis. Life Sci. 264, 118617 (2021).
Chiang, C. C., Korinek, M., Cheng, W. J. & Hwang, T. L. Targeting neutrophils to treat acute respiratory distress syndrome in coronavirus disease. Front Pharm. 11, 572009 (2020).
Curran, A. M., Naik, P., Giles, J. T. & Darrah, E. PAD enzymes in rheumatoid arthritis: pathogenic effectors and autoimmune targets. Nat. Rev. Rheumatol. 16, 301–315 (2020).
Sahebnasagh, A. et al. Neutrophil elastase inhibitor (sivelestat) may be a promising therapeutic option for management of acute lung injury/acute respiratory distress syndrome or disseminated intravascular coagulation in COVID-19. J. Clin. Pharm. Ther. 45, 1515–1519 (2020).
Maki, C. et al. Evaluation of appropriate indications for the use of sivelestat sodium in acute respiratory distress syndrome: a retrospective cohort study. Acute Med Surg. 7, e471 (2020).
Okur, H. K. et al. Preliminary report of in vitro and in vivo effectiveness of dornase alfa on SARS-CoV-2 infection. N. Microbes N. Infect. 37, 100756 (2020).
Holliday, Z. M. et al. Non-randomized trial of dornase Alfa for acute respiratory distress syndrome secondary to Covid-19. Front Immunol. 12, 714833 (2021).
Waugh, D. J. & Wilson, C. The interleukin-8 pathway in cancer. Clin. Cancer Res 14, 6735–6741 (2008).
Maurice, D. H. et al. Advances in targeting cyclic nucleotide phosphodiesterases. Nat. Rev. Drug Disco. 13, 290–314 (2014).
Giorgi, M. et al. Phosphodiesterase inhibitors: could they be beneficial for the treatment of COVID-19? Int. J. Mol. Sci. 21, 5338 (2020).
Liu, X. et al. Potential therapeutic effects of dipyridamole in the severely ill patients with COVID-19. Acta Pharm. Sin. B 10, 1205–1215 (2020).
Maldonado, V. et al. Pentoxifylline decreases serum LDH levels and increases lymphocyte count in COVID-19 patients: Results from an external pilot study. Int Immunopharmacol. 90, 107209 (2021).
Dixon, S. J. et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell 149, 1060–1072 (2012).
Jiang, X., Stockwell, B. R. & Conrad, M. Ferroptosis: mechanisms, biology and role in disease. Nat. Rev. Mol. Cell Biol. 22, 266–282 (2021).
Jacobs, W. et al. Fatal lymphocytic cardiac damage in coronavirus disease 2019 (COVID-19): autopsy reveals a ferroptosis signature. ESC Heart Fail. https://doi.org/10.1002/ehf2.12958 (2020).
Bednash, J. S. et al. Syrian hamsters as a model of lung injury with SARS-CoV-2 infection: Pathologic, physiologic, and detailed molecular profiling. Transl. Res 240, 1–16 (2022).
Suresh, V. et al. Quantitative proteomics of hamster lung tissues infected with SARS-CoV-2 reveal host factors having implication in the disease pathogenesis and severity. FASEB J. 35, e21713 (2021).
Zhao, K. et al. Serum iron level as a potential predictor of coronavirus disease 2019 severity and mortality: a retrospective study. Open Forum Infect. Dis. 7, ofaa250 (2020).
Shah, A. et al. Systemic hypoferremia and severity of hypoxemic respiratory failure in COVID-19. Crit. Care 24, 320 (2020).
Mehta, P. et al. COVID-19: consider cytokine storm syndromes and immunosuppression. Lancet 395, 1033–1034 (2020).
Zhou, C. et al. Increased serum levels of hepcidin and ferritin are associated with severity of COVID-19. Med Sci. Monit. 26, e926178 (2020).
Kobune, M. et al. Interleukin-6 enhances hepatic transferrin uptake and ferritin expression in rats. Hepatology 19, 1468–1475 (1994).
Nemeth, E. et al. IL-6 mediates hypoferremia of inflammation by inducing the synthesis of the iron regulatory hormone hepcidin. J. Clin. Invest 113, 1271–1276 (2004).
Nai, A. et al. Hepcidin levels predict Covid-19 severity and mortality in a cohort of hospitalized Italian patients. Am. J. Hematol. 96, E32–E35 (2021).
Wang, Y. et al. SARS-CoV-2 suppresses mRNA expression of selenoproteins associated with ferroptosis, endoplasmic reticulum stress and DNA synthesis. Food Chem. Toxicol. 153, 112286 (2021).
Kung, Y. A. et al. Acyl-Coenzyme a synthetase long-chain family member 4 is involved in viral replication organelle formation and facilitates virus replication via ferroptosis. mBio 13, e0271721, (2022).
Habib, H. M., Ibrahim, S., Zaim, A. & Ibrahim, W. H. The role of iron in the pathogenesis of COVID-19 and possible treatment with lactoferrin and other iron chelators. Biomed. Pharmacother. 136, 111228 (2021).
Chen, X., Yu, C., Kang, R. & Tang, D. Iron metabolism in ferroptosis. Front Cell Dev. Biol. 8, 590226 (2020).
Temraz, S., Santini, V., Musallam, K. & Taher, A. Iron overload and chelation therapy in myelodysplastic syndromes. Crit. Rev. Oncol. Hematol. 91, 64–73 (2014).
Zhang, J. et al. Association between regional selenium status and reported outcome of COVID-19 cases in China. Am. J. Clin. Nutr. 111, 1297–1299 (2020).
Moghaddam, A. et al. Selenium deficiency is associated with mortality risk from COVID-19. Nutrients. 12, 2098 (2020).
Kieliszek, M. & Lipinski, B. Selenium supplementation in the prevention of coronavirus infections (COVID-19). Med Hypotheses 143, 109878 (2020).
Alim, I. et al. Selenium drives a transcriptional adaptive program to block ferroptosis and treat stroke. Cell 177, 1262–1279 e1225 (2019).
Jin, Z. et al. Structure of M(pro) from SARS-CoV-2 and discovery of its inhibitors. Nature 582, 289–293 (2020).
Bermano, G., Meplan, C., Mercer, D. K. & Hesketh, J. E. Selenium and viral infection: are there lessons for COVID-19? Br. J. Nutr. 125, 618–627 (2021).
Codenotti, S. et al. Cell growth potential drives ferroptosis susceptibility in rhabdomyosarcoma and myoblast cell lines. J. Cancer Res Clin. Oncol. 144, 1717–1730 (2018).
Han, F., Li, S., Yang, Y. & Bai, Z. Interleukin-6 promotes ferroptosis in bronchial epithelial cells by inducing reactive oxygen species-dependent lipid peroxidation and disrupting iron homeostasis. Bioengineered 12, 5279–5288 (2021).
Scheffel, M. J. et al. N-acetyl cysteine protects anti-melanoma cytotoxic T cells from exhaustion induced by rapid expansion via the downmodulation of Foxo1 in an Akt-dependent manner. Cancer Immunol. Immunother. 67, 691–702 (2018).
de Alencar, J. C. G. et al. Double-blind, randomized, placebo-controlled trial with N-acetylcysteine for treatment of severe acute respiratory syndrome caused by coronavirus disease 2019 (COVID-19). Clin. Infect. Dis. 72, e736–e741 (2021).
Ayola-Serrano, N. C. et al. The role of 5-lipoxygenase in the pathophysiology of COVID-19 and its therapeutic implications. Inflamm. Res 70, 877–889 (2021).
Chandra, A. et al. Exploring potential inhibitor of SARS-CoV2 replicase from FDA approved drugs using insilico drug discovery methods. J. Biomol. Struct. Dyn. https://doi.org/10.1080/07391102.2020.1871416 (2021).
Carboni, E., Carta, A. R. & Carboni, E. Can pioglitazone be potentially useful therapeutically in treating patients with COVID-19? Med Hypotheses 140, 109776 (2020).
Takeshige, K. et al. Autophagy in yeast demonstrated with proteinase-deficient mutants and conditions for its induction. J. Cell Biol. 119, 301–311 (1992).
Dikic, I. & Elazar, Z. Mechanism and medical implications of mammalian autophagy. Nat. Rev. Mol. Cell Biol. 19, 349–364 (2018).
Yu, L., Chen, Y. & Tooze, S. A. Autophagy pathway: cellular and molecular mechanisms. Autophagy 14, 207–215 (2018).
Levine, B. & Kroemer, G. Biological functions of autophagy genes: a disease perspective. Cell 176, 11–42 (2019).
Pizzorno, A. et al. Characterization and treatment of SARS-CoV-2 in nasal and bronchial human airway epithelia. Cell Rep. Med 1, 100059 (2020).
Cortese, M. et al. Integrative imaging reveals SARS-CoV-2-induced reshaping of subcellular morphologies. Cell Host Microbe 28, 853–866 e855 (2020).
Benvenuto, D. et al. Evolutionary analysis of SARS-CoV-2: how mutation of Non-Structural Protein 6 (NSP6) could affect viral autophagy. J. Infect. 81, e24–e27 (2020).
Du Toit, A. Coronavirus replication factories. Nat. Rev. Microbiol 18, 411 (2020).
Hui, X. et al. SARS-CoV-2 promote autophagy to suppress type I interferon response. Signal Transduct. Target Ther. 6, 180 (2021).
Li, X. et al. SARS-CoV-2 ORF10 suppresses the antiviral innate immune response by degrading MAVS through mitophagy. Cell Mol. Immunol. 19, 67–78 (2022).
Sharma, A., Kontodimas, K. & Bosmann, M. The MAVS immune recognition pathway in viral infection and sepsis. Antioxid. Redox Signal 35, 1376–1392 (2021).
Zhang, Y. et al. The ORF8 protein of SARS-CoV-2 mediates immune evasion through down-regulating MHC-Iota. Proc. Natl Acad. Sci. USA https://doi.org/10.1073/pnas.2024202118 (2021).
Koepke, L. et al. Manipulation of autophagy by SARS-CoV-2 proteins. Autophagy 17, 2659–2661 (2021).
Liang, C. et al. Autophagic and tumour suppressor activity of a novel Beclin1-binding protein UVRAG. Nat. Cell Biol. 8, 688–699 (2006).
Qu, Y. et al. ORF3a-mediated incomplete autophagy facilitates severe acute respiratory syndrome coronavirus-2 replication. Front Cell Dev. Biol. 9, 716208 (2021).
Bhardwaj, M., Leli, N. M., Koumenis, C. & Amaravadi, R. K. Regulation of autophagy by canonical and non-canonical ER stress responses. Semin Cancer Biol. 66, 116–128 (2020).
Zhang, Y. et al. The SARS-CoV-2 protein ORF3a inhibits fusion of autophagosomes with lysosomes. Cell Disco. 7, 31 (2021).
Miao, G. et al. ORF3a of the COVID-19 virus SARS-CoV-2 blocks HOPS complex-mediated assembly of the SNARE complex required for autolysosome formation. Dev. Cell 56, 427–442 e425 (2021).
Hayn, M. et al. Systematic functional analysis of SARS-CoV-2 proteins uncovers viral innate immune antagonists and remaining vulnerabilities. Cell Rep. 35, 109126 (2021).
Munson, M. J. & Ganley, I. G. MTOR, PIK3C3, and autophagy: Signaling the beginning from the end. Autophagy 11, 2375–2376 (2015).
Mohamud, Y. et al. The papain-like protease of coronaviruses cleaves ULK1 to disrupt host autophagy. Biochem Biophys. Res Commun. 540, 75–82 (2021).
Vincent, M. J. et al. Chloroquine is a potent inhibitor of SARS coronavirus infection and spread. Virol. J. 2, 69 (2005).
Savarino, A. et al. Effects of chloroquine on viral infections: an old drug against today’s diseases. Lancet Infect. Dis. 3, 722–727 (2003).
Gratton, R. et al. Autophagy in Zika virus infection: a possible therapeutic target to counteract viral replication. Int. J. Mol. Sci. 28, 1048 (2019).
Mauthe, M. et al. Chloroquine inhibits autophagic flux by decreasing autophagosome-lysosome fusion. Autophagy 14, 1435–1455 (2018).
Al-Bari, M. A. A. Targeting endosomal acidification by chloroquine analogs as a promising strategy for the treatment of emerging viral diseases. Pharm. Res Perspect. 5, e00293 (2017).
Savarino, A. et al. New insights into the antiviral effects of chloroquine. Lancet Infect. Dis. 6, 67–69 (2006).
Liu, J. et al. Hydroxychloroquine, a less toxic derivative of chloroquine, is effective in inhibiting SARS-CoV-2 infection in vitro. Cell Disco. 6, 16 (2020).
Wang, M. et al. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res 30, 269–271 (2020).
Gorshkov, K. et al. The SARS-CoV-2 cytopathic effect is blocked with autophagy modulators. bioRxiv. https://doi.org/10.1101/2020.05.16.091520 (2020).
Huang, M. et al. Preliminary evidence from a multicenter prospective observational study of the safety and efficacy of chloroquine for the treatment of COVID-19. Natl Sci. Rev. 7, 1428–1436 (2020).
Chen, Z. et al. Efficacy of hydroxychloroquine in patients with COVID-19: results of a randomized clinical trial. medRxiv. https://doi.org/10.1101/2020.03.22.20040758 (2020).
Gautret, P. et al. Hydroxychloroquine and azithromycin as a treatment of COVID-19: results of an open-label non-randomized clinical trial. Int J. Antimicrob. Agents 56, 105949 (2020).
Group, R. C. et al. Effect of hydroxychloroquine in hospitalized patients with Covid-19. N. Engl. J. Med. 383, 2030–2040 (2020).
Self, W. H. et al. Effect of hydroxychloroquine on clinical status at 14 days in hospitalized patients with COVID-19: a randomized clinical trial. JAMA 324, 2165–2176 (2020).
Yuen, C. K. et al. Suppression of SARS-CoV-2 infection in ex-vivo human lung tissues by targeting class III phosphoinositide 3-kinase. J. Med Virol. 93, 2076–2083 (2021).
Williams, C. G. et al. Inhibitors of VPS34 and fatty-acid metabolism suppress SARS-CoV-2 replication. Cell Rep. 36, 109479 (2021).
Seglen, P. O. & Gordon, P. B. 3-Methyladenine: specific inhibitor of autophagic/lysosomal protein degradation in isolated rat hepatocytes. Proc. Natl Acad. Sci. USA 79, 1889–1892 (1982).
Blommaart, E. F. et al. The phosphatidylinositol 3-kinase inhibitors wortmannin and LY294002 inhibit autophagy in isolated rat hepatocytes. Eur. J. Biochem 243, 240–246 (1997).
Miller, S. et al. Shaping development of autophagy inhibitors with the structure of the lipid kinase Vps34. Science 327, 1638–1642 (2010).
Knight, S. D. et al. Discovery of GSK2126458, a highly potent inhibitor of PI3K and the mammalian target of rapamycin. ACS Med Chem. Lett. 1, 39–43 (2010).
Martin, K. R. et al. A potent and selective ULK1 inhibitor suppresses autophagy and sensitizes cancer cells to nutrient. Stress. iScience. 8, 74–84 (2018).
Lazarus, M. B., Novotny, C. J. & Shokat, K. M. Structure of the human autophagy initiating kinase ULK1 in complex with potent inhibitors. ACS Chem. Biol. 10, 257–261 (2015).
Petherick, K. J. et al. Pharmacological inhibition of ULK1 kinase blocks mammalian target of rapamycin (mTOR)-dependent autophagy. J. Biol. Chem. 290, 11376–11383 (2015).
Egan, D. F. et al. Small molecule inhibition of the autophagy kinase ULK1 and identification of ULK1 substrates. Mol. Cell 59, 285–297 (2015).
Appelberg, S. et al. Dysregulation in Akt/mTOR/HIF-1 signaling identified by proteo-transcriptomics of SARS-CoV-2 infected cells. Emerg. Microbes Infect. 9, 1748–1760 (2020).
Gassen, N. C. et al. Analysis of SARS-CoV-2-controlled autophagy reveals spermidine, MK-2206, and niclosamide as putative antiviral therapeutics. bioRxiv. https://doi.org/10.1101/2020.04.15.997254 (2020).
Ko, S. et al. Rapamycin-induced autophagy restricts porcine epidemic diarrhea virus infectivity in porcine intestinal epithelial cells. Antivir. Res. 146, 86–95 (2017).
Guo, L. et al. Autophagy negatively regulates transmissible gastroenteritis virus replication. Sci. Rep. 6, 23864 (2016).
Wang, C. H. et al. Adjuvant treatment with a mammalian target of rapamycin inhibitor, sirolimus, and steroids improves outcomes in patients with severe H1N1 pneumonia and acute respiratory failure. Crit. Care Med. 42, 313–321 (2014).
Kindrachuk, J. et al. Antiviral potential of ERK/MAPK and PI3K/AKT/mTOR signaling modulation for Middle East respiratory syndrome coronavirus infection as identified by temporal kinome analysis. Antimicrob. Agents Chemother. 59, 1088–1099 (2015).
Gao, C. et al. Metformin induces autophagy via the AMPK-mTOR signaling pathway in human hepatocellular carcinoma cells. Cancer Manag Res. 12, 5803–5811 (2020).
Samuel, S. M., Varghese, E. & Busselberg, D. Therapeutic potential of metformin in COVID-19: reasoning for its protective role. Trends Microbiol. 29, 894–907 (2021).
Chen, X. et al. Immunomodulatory and antiviral activity of metformin and its potential implications in treating coronavirus disease 2019 and lung injury. Front Immunol. 11, 2056 (2020).
Xian, H. et al. Metformin inhibition of mitochondrial ATP and DNA synthesis abrogates NLRP3 inflammasome activation and pulmonary inflammation. Immunity 54, 1463–1477 e1411 (2021).
Lally, M. A. et al. Metformin is associated with decreased 30-day mortality among nursing home residents infected with SARS-CoV2. J. Am. Med Dir. Assoc. 22, 193–198 (2021).
Bramante, C. T. et al. Observational study of metformin and risk of mortality in patients hospitalized with Covid-19. medRxiv. https://doi.org/10.1101/2020.06.19.20135095 (2020).
Luo, P. et al. Metformin treatment was associated with decreased mortality in COVID-19 patients with diabetes in a retrospective analysis. Am. J. Trop. Med Hyg. 103, 69–72 (2020).
Acknowledgements
This work was financially supported by the National Natural Science Foundation of China (81201788).
Author information
Authors and Affiliations
Contributions
X.L. and Z.Z. wrote the manuscript and created the figures. Z.W. and P.G.-C. revised the initial manuscript and figures. H.S. provided the conceptual idea and revised the manuscript. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, X., Zhang, Z., Wang, Z. et al. Cell deaths: Involvement in the pathogenesis and intervention therapy of COVID-19. Sig Transduct Target Ther 7, 186 (2022). https://doi.org/10.1038/s41392-022-01043-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41392-022-01043-6
- Springer Nature Limited
This article is cited by
-
Progesterone modulates the immune microenvironment to suppress ovalbumin-induced airway inflammation by inhibiting NETosis
Scientific Reports (2024)
-
COVID-19 related neurological manifestations in Parkinson’s disease: has ferroptosis been a suspect?
Cell Death Discovery (2024)
-
Apoptosis and pyroptosis in the nasal mucosa of Syrian hamster during SARS-CoV-2 infection and reinfection
Apoptosis (2024)
-
A Machine Learning Approach to Identify Potential miRNA-Gene Regulatory Network Contributing to the Pathogenesis of SARS-CoV-2 Infection
Biochemical Genetics (2024)
-
Mining phase separation-related diagnostic biomarkers for endometriosis through WGCNA and multiple machine learning techniques: a retrospective and nomogram study
Journal of Assisted Reproduction and Genetics (2024)