Abstract
Environmental factors contribute to risk of bipolar disorder (BD), but how environmental factors impact the development of psychopathology within the context of elevated genetic risk is unknown. We herein sought to identify epigenetic signatures operating in the context of polygenic risk for BD in young people at high familial risk (HR) of BD. Peripheral blood-derived DNA was assayed using Illumina PsychArray, and Methylation-450K or -EPIC BeadChips. Polygenic risk scores (PRS) were calculated using summary statistics from recent genome-wide association studies for BD, major depressive disorder (MDD) and cross-disorder (meta-analysis of eight psychiatric disorders). Unrelated HR participants of European ancestry (n = 103) were stratified based on their BD-PRS score within the HR-population distribution, and the top two quintiles (High-BD-PRS; n = 41) compared against the bottom two quintiles (Low-BD-PRS; n = 41). The High-BD-PRS stratum also had higher mean cross-disorder-PRS and MDD-PRS (ANCOVA p = 0.035 and p = 0.024, respectively). We evaluated DNA methylation differences between High-BD-PRS and Low-BD-PRS strata using linear models. One differentially methylated probe (DMP) (cg00933603; p = 3.54 × 10−7) in VARS2, a mitochondrial aminoacyl-tRNA synthetase, remained significantly hypomethylated after multiple-testing correction. Overall, BD-PRS appeared to broadly impact epigenetic processes, with 1,183 genes mapped to nominal DMPs (p < 0.05); these displayed convergence with genes previously associated with BD, schizophrenia, chronotype, and risk taking. We tested poly-methylomic epigenetic profiles derived from nominal DMPs in two independent samples (n = 54 and n = 82, respectively), and conducted an exploratory evaluation of the effects of family environment, indexing cohesion and flexibility. This study highlights an important interplay between heritable risk and epigenetic factors, which warrant further exploration.
Similar content being viewed by others
Introduction
Bipolar disorder (BD) is a highly heritable mental illness, for which genetic factors explain ~60–85% of risk variance [1]; the remaining variance is explained by non-genetic factors, including environmental contributors. First-degree relatives of probands with BD have 5–10-fold increased risk of developing BD themselves [1,2,3], and are at increased risk of broader psychopathology [4, 5], including major depression, anxiety, behavioural and substance use disorders; therefore, young first-degree relatives of those with BD are considered high risk (HR) for later mental illness. The elucidation of clinical antecedents of the BD prodrome is an active research area [6,7,8,9,10], but specific precursors are heterogeneous, and biomarkers of risk trajectories are a research priority.
Genome-wide association studies (GWAS) have identified many single nucleotide polymorphisms (SNPs) associated with increased disease risk, each with small individual effect [11,12,13]. Many disease-associated SNPs are shared amongst psychiatric disorders, with substantial genetic correlation between BD and schizophrenia, and BD and major depressive disorder (MDD) [13,14,15,16,17]. GWAS have demonstrated the polygenic nature of BD, where common SNPs identified to date collectively account for ~25% of the estimated heritability [12,13,14,15]. Indeed, individuals with polygenic risk scores (PRS) at the extremes of a population distribution have substantially altered risk of developing a psychiatric disorder—those in the top 10% of the BD-PRS distribution have an odds ratio of 9.3 of developing BD compared to the lowest decile [13]—yet the predictive capacity of PRS is currently limited [18], with inadequate sensitivity and specificity as PRS will only capture part of the genetic contribution [19]. SNP-based heritability falls short of heritability estimates from family, twin and epidemiologic studies; [20] the latter approaches typically employ models that include additive genetic and common/unique environmental factors (and their interaction), whereas SNP-based heritability estimates typically only model additive genetic variance [20]. Moreover, some of the “missing heritability” may also reside in rare variants not captured by GWAS [21,22,23], and/or relate to other familial factors that are not encoded in nucleotide-level DNA sequence variation.
Family history is the strongest current predictor of future BD [7] and is sometimes posited as a proxy for genetic transmission, but likely reflects dynamic gene-environment interplay from preconception through the life span. Twin studies also indicate environmental contribution to BD [24]. Several environmental risk factors have been posited—including prenatal infections, childhood maltreatment, and psychological stress [25]—but the attributable impact of such factors is small and not disease-specific. Families that include a parent with BD have lower parent-reported cohesion compared to families with no parental psychiatric disorders [26], and while offspring-centred reports are less common, child reports of lower cohesion and adaptability, and higher conflict environments are also noted [27]; these factors potentially confound family history with environmental elements. Furthermore, children who experienced early maternal loss have a four-fold increased risk of BD [28], suggesting that early trauma and altered home-environment dynamics may influence disease trajectory. Indeed, children are influenced by both genes and environment provided by parents, which cannot be easily disambiguated to apportion causation [29]. Moreover, environmental effects are likely conditional on genetic factors, which are only partially appraised by existing gene-environment studies [30]. Also, genetic nurture may include indirect genetic effects from parental genes that are not transmitted [31]. Thus, there is increasing interest in the role of specific gene-environment interactions, as well as the mechanisms by which risk factors interact in the development of BD [32].
Environmental factors can potentially impact gene expression through epigenetic modulations [33]. One of the most studied epigenetic processes is DNA methylation; the addition of a methyl group to the 5′ cytosine of a cytosine-guanine sequence (CpG) [33]. To date, there are no large-scale epigenome-wide studies in BD [34], although several candidate-gene epigenetic studies [35, 36] and pharmaco-epigenomic studies of antipsychotic medicines [37] have been performed. Furthermore, GWAS signals for schizophrenia—a condition that shares genetic overlap with BD [17, 38]—are enriched in human-specific methylated regions [39], implying mechanistic overlap between genetic and epigenetic risk. Interestingly, a recent epigenetic element-based transcriptome-wide association study identified genes that contribute to BD heritability beyond those explained by GWAS-associated SNPs [40], suggesting that epigenetic regulation may further contribute to heritability. Epigenetic factors may even contribute to transgenerational genomic regulation [41], although this mechanism of genomic transmission in humans remains controversial [42].
To identify biomarkers for illness-onset, a prospective longitudinal approach is required. One small prospective study that compared HR participants who developed BD or MDD (n = 22) to those who remained well (n = 23) identified 22,543 nominally differentially methylated CpGs (p < 0.05) [43], although no probe passed epigenome-wide correction for multiple testing. To date, the characterisation of methylation differences relating to polygenic burden in individuals with familial risk for BD has not been performed. Thus, we undertook the identification of differentially methylated positions (DMPs) in young HR participants, stratified by genetic burden of common BD-associated risk alleles, to characterise epigenetic modifications operating beyond polygenic additive risk. Methylation signatures derived from the cumulative effect of multiple CpGs were then validated in independent samples of BD cases, HR, and controls. Finally, the impact of family environment measures on methylation signatures were explored, to further understand relationships between familial and environmental risk factors associated with the development of psychopathology.
Materials and methods
An overview of the approach is summarised in Fig. 1.
PRS polygenic risk score, PMPS poly-methylomic profile score, HR high-risk individuals, CON control, BD bipolar disorder, HRunwell high-risk individuals who developed sub-threshold or threshold BD at baseline or follow-up, HRwell high-risk individuals who remained well after follow-up, EA European ancestry based on components from multidimensional scaling analysis of SNP genotype data.
Study participants
Australian participants were aged 12–30 years and recruited as previously described [44, 45]. Briefly, HR and BD participants were recruited from families who had previously participated in BD family studies, specialised BD research clinics, mental health consumer organisations, or response to public notices. BD cases met DSM-IV criteria for BD type-I (BD-I) or type-II (BD-II). HR participants were the children or siblings of individuals with DSM-IV diagnoses of BD-I, BD-II, or schizoaffective disorder–bipolar-type (SABP), and did not personally have threshold diagnoses of these conditions at baseline (participants with sub-threshold BD “not otherwise specified” (BD-NOS) were not excluded). Control participants (CON) were recruited via print/electronic media, and noticeboards in universities and local communities. CON had no personal or familial (first-degree) history of BD-I, BD-II, recurrent unipolar disorder, SABP, schizophrenia, recurrent substance abuse or psychiatric hospitalisation, and no second-degree relative with a past mood-disorder hospitalisation or history of psychosis. Written informed consent was obtained from all participants, with additional parental consent for participants aged <16 years. This study was approved by University of New South Wales Human Research Ethics Committee (HREC Protocol 09/097).
High-risk participants from the United States were aged 12–21 years, and recruited from Indiana University, University of Michigan, Johns Hopkins University, and Washington University in St. Louis (with site-specific IRB approval), as previously described [46]. The US sites used recruitment criteria and clinical assessments identical to those used at the Australian site.
Clinical assessments
Structured interviews were administered by staff with extensive clinical background and after specific training in each instrument, which comprised a battery of structured clinical interviews, self-report questionnaires and clinician rated assessments. The Family Interview for Genetic Studies (FIGS) [47] was administered to each participant or a member of their family to determine family history of mood or psychotic disorder. To determine psychiatric diagnoses for participants aged 12–21, both participants and their parent/s completed the Kiddie-Schedule for Affective Disorders and Schizophrenia for School-Aged Children–Present and Lifetime Version (K-SADS-BP; v2, July 2009) [46, 48]. To determine psychiatric diagnoses for participants aged 22-30 (including BD-probands), the Diagnostic Interview for Genetic Studies (DIGS; v4.0/BP, July 2005) [49] was administered. Consensus DSM-IV diagnoses were determined by two clinicians (i.e., psychiatrists with child specialty training, clinical psychologists, or clinical social workers) using best-estimate methodology [50], using the K-SADS or DIGS, FIGS, and medical records (where available). Functional capacity was determined via Global Assessment of Functioning (GAF) [51] or Clinical Global Impression (CGI) [52] rating scales.
A subset of participants completed the Family Adaptability and Cohesion Evaluation Scales (FACES-II) [53], a 30-item questionnaire from which a total score across the adaptability and cohesion subscales was used to represent family environment (described in Supplementary Material).
One to four follow-up clinical interviews were performed on HR and control participants to identify emergent psychopathology. Diagnoses were assigned a confidence rating on a 4-point scale; only those that met full DSM-IV criteria received a confidence rating ≥3. As previously described by Frankland et al. [8], participants with a best-estimate diagnosis of BD were categorised as either threshold converters (i.e. with BD-I or BD-II; confidence level 3–4) or sub-threshold converters (i.e. with BD-NOS; confidence level of 1–2); the latter diagnosis being made when participants did not meet the minimum 4 day duration criterion, but otherwise met full symptom criteria for hypomania. Participants (HR or control) reaching threshold or sub-threshold criteria at follow-up assessment for diagnosis of BD-I, BD-II, BD-NOS, with a confidence of ≥2 were considered “BD-syndromic”, and those with no clinical diagnoses after follow-up were considered “disorder-free”.
Genotyping and polygenic risk scores
Peripheral blood samples were collected, DNA extracted, and genotyping performed on PsychArray-24 BeadChip as previously described [54] (details in Supplementary methods). An independent sample of BD cases (n = 264; described in [13] as “neuc1”) and controls (n = 1115) [55] were employed to infer optimal p value threshold for generation of PRS (pT; Supplementary Methods, Fig. S1).
Polygenic risk scores (PRS) were generated using summary statistics from the PGC GWAS for BD (n = 31,358 controls and n = 20,352 cases; excluding 6,201 participants from GAIN, BMAU, FAT2, MICH datasets which contained parents of some of the study participants) [12]. Data for MDD (n = 561,190 controls and n = 246,363 cases) [56] and cross-disorder (n = 494,162 controls and n = 232,964 cases with anorexia nervosa, attention-deficit/hyperactivity disorder, autism spectrum disorder, BD, MDD, obsessive-compulsive disorder, schizophrenia, and Tourette syndrome) [17] were also used. PRS were calculated in a single analysis for all participants simultaneously using PRSice v2.2.13 [57], employing unrelated individuals (identity-by-descent pi_hat < 0.1; Supplementary Methods) identified as European-ancestry by multidimensional scaling analysis (Supplementary Methods), and comparing BD cases to controls to define optimal pT to explain the largest variance (Nagelkerke pseudo-R2); phenotypes for HR were coded unknown/missing (-9). Polygenic scoring employed linkage disequilibrium clumping, removal of strand-ambiguous SNPs, and SNPs with MAF < 0.05 or low imputation quality (INFO < 0.8), prior to PRS computation and use in subsequent analyses.
Demographic comparisons
Group differences were examined using univariate general linear ANCOVA models, using SPSS Statistics for Windows v26 (IBM corp., Armonk, NY). For PRS group level comparisons in the extended cohort, generalised estimation equations (GEE) were employed to account for relationships within families which contained multiple relatives, and included genotype-derived MDS components (C1 and 2) as covariates.
Methylation analysis
Epigenome-wide methylation profiling
Methylation quantification was performed on DNA derived from peripheral blood in four batches: batches 1 and 2 (both n = 96) employed the Illumina HumanMethylation450K array, and batches 3 and 4 (n = 127 and 96, respectively) employed the Illumina MethylEPIC BeadChip. The R package meffil [58] was used to extract signal intensities and initial quality control (QC) within each batch (Supplementary Material). The R package Shinymethyl [59] was used to conduct principal component analysis and visualise outliers within each batch. After removal of 11 technical replicates and 20 samples failing QC procedures, 384 participants remained (round 1 n = 88, round 2 n = 96, round 3 n = 118, round 4 n = 82) – 425,453 CpG probes that passed QC and were represented on both 450k and EPIC chips then underwent further filtering for EWAS as outlined below. To avoid potential bias in PRS due to ethnicity [60], the cohort was restricted to unrelated European-ancestry participants (Fig. S2), which were divided into Discovery and Validation sets (described below and in Fig. 1). Normalisation of methylation β values employed meffil, including technical covariates (sentrix position and sentrix group; the latter unique across batches) [58]. Normalisation of batches 1–3 was performed simultaneously across all three batches, and formed the Discovery Set and Validation Set 1, whereas batch 4 was normalised separately as Validation Set 2.
Tissue source, smoking status and cell count estimates
DNA was derived from three peripheral blood sources: whole blood, ficoll/buffy-coat, and isolated lymphoblasts (details in Supplementary Material). To account for inter-individual differences in cellular proportions, estimation of six cell types (B cells, CD4T, CD8T, granulocytes, monocytes and natural killer) was performed using meffil’s gse35069 profile [58]. Tissue source and cell count estimates (Fig. S3) were included as covariates, as described below.
Lifetime tobacco use collected via KSADS was available for ~60% of participants in this study, and current smoking status was available for ~16% (n = 36 out of 218 individuals; n = 5 current users, all in Validation Set 2). To predict smoking status in all participants, normalised β values at cg05575921 were used (Fig. S4), where β < 0.75 were classified as probable smokers [61].
Discovery set: HR youth stratified by BD-PRS
To stratify HR participants on the basis of their personal burden of BD-associated SNP variants, the distribution of BD-PRS within the HR sample was divided into quintiles; this stratification was undertaken on the basis that individuals with PRS in the extremes of a population distribution have substantially altered risk of developing a psychiatric disorder [13]. Quintiles were chosen to balance maximisation of the available sample, with exclusion of individuals with intermediate scores reflecting ‘average’ SNP burden. Thus two comparator groups were defined: the top two quintiles (40% of participants) formed the High-BD-PRS group (n = 41) and the bottom two quintiles (40% of participants) formed the Low-BD-PRS group (n = 41) (Table S1, Fig. 2).
BD-PRS was standardised using the mean and standard deviation of all participants with genotype data (including an independent sample of BD cases and controls) regardless of availability of methylation data (n = 1699 controls, 354 HR, 355 BD; M = 0.0077695, SD = 7.543 × 10−5). The boxes and whiskers inside the violins indicate the 25–75th percentiles, and 1.5 times the interquartile range, respectively. The dashed rectangles indicate the stratification of HR participants into High-BD-PRS vs. Low-BD-PRS strata, as defined by the top and bottom two quintiles (i.e. 40% of the HR distribution), for the epigenome-wide association study. HR high risk, BD bipolar disorder, CON control, PRS polygenic risk score.
Discovery EWAS: high vs. low-BD-PRS within HR youth
CpG selection
Probes were restricted to those which were: (1) variable in our data (n = 249,436)—defined by >5% variability in the difference between the 10th to 90th percentile range of β values across batches 1–3 [62], and (2) reported to be variable in blood (n = 244,724), as defined by Edgar et al. [63], and 3) correlated between blood and brain (n = 42,364; with mean r ≥ ±0.3 across prefrontal cortex, superior temporal gyrus and entorhinal cortex) as defined by Hannon et al. [62]. Restricting the discovery EWAS to variable blood-brain correlated probes had benefits in: (1) reducing the EWAS search space, and (2) focusing on CpGs that are likely to be functionally relevant in the primary disease-affected tissue. Importantly, ‘probe SNPs’ as annotated in the Illumina manifest file were removed prior to EWAS, to exclude direct effects of sequence variation out to 50 bp (n = 6,455). The final probe set included 35,907 CpGs, and β values for each probe were transformed into M-values using the ‘beta2m’ function in R package lumi [64].
Surrogate variable analyses: To identify and account for residual unmeasured variation that was not corrected by normalisation of technical covariates (sentrix position/chip/batch), the ‘be’ method in R package SVA [65] was used to identify surrogate variables considering BD-PRS stratum as the variable of interest (Low-BD-PRS = 0, High-BD-PRS = 1). Four significant surrogate variables (p < 0.05) were identified and included in downstream analysis.
EWAS: DMPs were identified using the R package limma [66], employing a linear regression model, with normalised M-values as outcome and BD-PRS stratum as predictor (Low-BD-PRS = 0, High-BD-PRS = 1). Covariates included sex, age, two ethnicity components (MDS C1 and C2; Supplementary Material), tissue source, blood-cell count estimates and four surrogate variables.
Bias correction
The R package bacon [67] was applied to correct test-statistic inflation, to minimise shift in distribution of effect sizes and risk of false-positive findings. Empirical null estimates were generated using a Gibbs Sampling algorithm in a Bayesian framework, based on 5000 iterations with burn-in period of 2000. DMPs whose inflation- and bias-corrected p values exceeded p < 1.39 × 10−6 (Bonferroni adjustment for 35,907 tests at α = 0.05) were considered epigenome-wide significant.
Pathway analyses
DMPs at two p value thresholds (the Q–Q plot inflection point of p < 0.002, and p < 0.05; uncorrected) were mapped to genes using two complementary methods: (1) physical mapping of probes based on UCSC hg19 base-pair coordinates (as per Illumina manifest file), and (2) functional mapping of cis-regulatory regions using the Genomic Regions Enrichment of Annotations Tool (GREAT; v4.0.4) [68]. GREAT enables functional mapping of DMPs in non-coding regulatory regions (i.e. outside coding-sequence boundaries)—we employed ‘basal plus extension’ mode that includes genes 5 kb upstream, 1 kb downstream, plus 5 kb distal binding. The combined output of both methods created a gene list for pathway interrogation.
Finally, Functional Mapping and Annotation of Genome-Wide Association Studies (FUMA) [69] was employed with default parameters (i.e. not excluding MHC region) to examine enrichment of genes harbouring DMPs in biological function and pathway categories, and enrichment of gene expression across tissues [70]. Enrichments were considered significant if false discovery rate (FDR) q ≤ 0.05.
Supplementary EWAS: post-hoc sensitivity analysis for interpretation of DMP functional enrichments
As the discovery EWAS employed a restricted set of 35,907 CpG probes that were blood-brain correlated [62], we reasoned that probe pre-selection might bias functional enrichments in genes mapped to DMPs. Therefore, a supplementary EWAS was conducted using 214,352 probes that passed QC and were variable in blood [63] and variable in our data (but without applying the blood-brain correlation filter), employing identical procedures as the primary EWAS. The multiple-testing correction threshold was adjusted to p < 2.33 × 10−7 for 214,352 tests at α = 0.05 [71].
Validation using poly-methylomic profile scores
A ‘poly-methylomic profile score’ (PMPS) is a quantitative metric reflecting the degree of methylation at multiple sites across the epigenome (similar to calculating a PRS from GWAS summary statistics for genotypic data), as previously described [72]. A PMPS was calculated for each participant using effect sizes and p-values for DMPs from the discovery PRS-stratified EWAS, and applied to two independent validation sets (Fig. 1), as described below. Normality was assessed via Shapiro-Wilk tests in SPSS, and effect sizes reported as partial eta squared (η2p).
Validation Set 1
To determine whether PMPS provided a replicable index of BD-PRS, we employed 67 control participants [European-ancestry, unrelated (pi_hat < 0.1)] derived from batch 1–3, thereby minimising technical variability between the Discovery Set (who were all HR) and Validation Set 1. As controls have an overlapping BD-PRS distribution to both BD and HR (Fig. 2), we reasoned that methylation changes that reflect BD-PRS variability should also be observable in controls.
Controls were divided into two strata based on their BD-PRS distribution: the top two quintiles (High-BD-PRS; n = 27) and the bottom two quintiles (Low-BD-PRS; n = 27). To obtain the optimal methylation signature, PMPS were calculated utilising three p value thresholds from discovery EWAS summary statistics (pT < 0.002, pT < 0.01 and pT < 0.05; representing n = 72, n = 389 and n = 1957 CpGs, respectively). Association with PMPS (dependent variable) was examined using univariate GLM, including BD-PRS strata as factor, with age, sex, tissue source, and blood-cell counts as covariates.
Validation Set 2
To determine whether the PMPS indexed effects related to the development of psychopathology, 82 independent unrelated individuals of European ancestry were selected from batch 4. BD cases were recruited in Australia, and 68% of HR and controls were from US sites. A ‘BD-syndromic’ group (n = 44) comprised BD cases (n = 20) plus HR individuals who developed BD-related psychopathology (n = 24), and a ‘disorder-free’ group (n = 38) comprised controls (n = 26) plus HR who remained well (n = 12).
PMPS were generated using pT < 0.05 (n = 1,891 CpGs, after QC exclusions), and compared between clinical groups: BD vs. CON, and extended ‘BD-syndromic’ vs. ‘disorder-free’ groups, using univariate GLM, as described above. BD-PRS and a BD-PRS×Group interaction term were also added to the model.
Sixty-six individuals in Validation Set 2 (80%) completed the FACES-II scale [53]. Exploratory regression analysis evaluating the impact of family environment, BD-PRS and psychopathology-group (and their interaction terms) on PMPS, employed a general linear model including age and sex as covariates.
Results
Polygenic risk scores
BD-PRS at pT = 0.105 optimally distinguished the independent BD group (n = 264) from controls (n = 1115), with p = 5.36 × 10−12 and R2 = 0.057 (n = 214,928 SNPs; Fig. S1). The optimal MDD-PRS (for BD vs. controls) was at pT = 0.017 (n = 187,029 SNPs, p = 1.89 × 10−8, R2 = 0.037), and the cross-disorder-PRS was optimal at pT = 0.229 (n = 269,339 SNPs, p = 1.76 × 10−10, R2 = 0.049). At a group level, BD-PRS distinguished HR (n = 355) from both BD and control groups, where HR had higher mean BD-PRS than controls (n = 1699; GEE p = 0.0003) and lower BD-PRS than BD (n = 319; GEE p = 0.0006). The BD-PRS exhibited an overlapping population distribution across clinical groups (Fig. 2).
Group comparisons: High vs. Low-BD-PRS strata within HR participants
Within the HR group, the High-BD-PRS stratum had significantly higher mean MDD-PRS (ANCOVA F = 5.32, p = 0.024, η2p = 0.064) and cross-disorder-PRS (ANCOVA F = 4.57, p = 0.035, η2p = 0.055) than the low-BD-PRS stratum. Epigenetically inferred smoking status analysis identified only two probable smokers, and these were equally represented in PRS group strata (Fig. S4). There was no difference in global functioning (GAF) between High-BD-PRS vs. Low-BD-PRS groups (M ± SD = 86.0 ± 9.3 vs. 84.8 ± 8.4, p = 0.54) (Fig. S6).
Differentially methylated positions: high vs. low-BD-PRS strata within HR
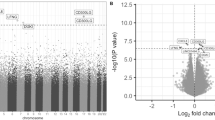
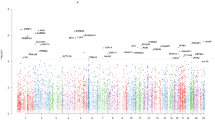
EWAS was performed using a linear regression framework, using the final probe set of 35,907 variable CpGs that were blood-brain correlated. Following minor correction for bias and inflation (−0.0022 and 0.97, respectively; corrected to −0.00049 and 1, respectively), one probe (cg00933603, p = 3.54 × 10−7) exceeded the epigenome-wide association threshold (pT < 1.39 × 10−6; FDR = 0.026) (Fig. 3). The quantile-quantile inflection point was defined at p < 0.002. Post-hoc adjustment for epigenetically-inferred smoking exposure (using βcg05575921 as a covariate) did not substantially alter the findings (cg00933603, p = 3.10 × 10−7; Table S2, Fig. S5).
A Quantile-Quantile plot, indicating observed vs. expected p values from 35,907 probes. Bias and inflation corrected to −0.00049 and 1, respectively. The dashed line indicates the inflection point at p < 0.002. B Manhattan plot indicating the genomic location of differentially methylated probes. The vertical line indicates the multiple-testing correction threshold at α = 0.05 for epigenome-wide association, based on 35,907 probes (p < 1.39 × 10−6). The location of the top DMP, cg00933603, is indicated. Covariates included age, sex, MDS C1 and C2, tissue source, six blood-cell components, and four surrogate variables. log logarithm; P p value.
Putative biological impact of differentially methylated genes
The DMPs above the inflection point (p < 0.002; n = 82; Table S2) and nominally significant DMPs (p < 0.05; n = 1957) were physically and functionally mapped to 66 and 1260 genes, respectively. Genes with DMPs above the inflection included MLC1, ESR1 KCKN5, L1CAM, CPEB1 and GABBR2, previously associated with BD [73] (Table S2; Fig. S7). EWAS-significant probe cg00933603 lies in exon 2 of VARS2, and has a cis-mQTL SNP rs2532928 (formerly known as rs116537083) [74], that correlates with VARS2 cortical expression (GTEx v8; [75] p = 9.1 × 10−29; Fig. S8).
In FUMA analyses, the 1183 DMPs were enriched in genes previously associated with several relevant neurobehavioral phenotypes in the GWAS catalog, including schizophrenia, chronotype and general risk tolerance, which were represented in the top 15 enrichments (adjusted p < 0.01; Table 1 and Table S3). The genes that mapped to DMPs were mostly upregulated in brain regions, including the hypothalamus, anterior cingulate cortex and amygdala (adjusted p = 4.40 × 10−16, 7.08 × 10−16 and 1.69 × 10−15, respectively; Fig. S9). Top gene ontology enrichment categories from FUMA included synaptic and neuronal cellular components; canonical pathways involving synaptic function/interactions and signal transmission; and biological processes relating to cell-cell adhesion and signalling functions (Table 2, Table S4).
Supplementary EWAS: post-hoc sensitivity analysis for interpretation of DMP functional enrichments
No DMP survived multiple-testing correction in the supplementary EWAS using 214,352 probes regardless of blood-brain correlation (Fig. S10). The top DMP from the primary PRS-EWAS (cg00933603; p = 3.54 × 10−7) was the third top DMP (p = 4.38 × 10−6) in this secondary EWAS (Table S5).
Pathway analysis in FUMA showed that brain tissues continue to exhibit the strongest enrichment with an equivalently-sized gene list derived from DMPs regardless of blood–brain correlation. The top brain regions were cortex, anterior cingulate cortex and amygdala (p = 1.58 × 10−14, p = 1.35 × 10−11 and p = 2.39 × 10−11, respectively; Fig. S11). Gene ontology enrichments implicate neuronal-related functions and GWAS catalog overlaps, with chronotype represented in the top 15 enrichments (Tables 1 and 2, Tables S6 and S7).
Test of generalisability of epigenetic signature using poly-methylomic profile score
Validation Set 1
PMPS calculated using three p value thresholds (pT) were normally distributed (Shapiro–Wilk p > 0.05). At the most stringent threshold (pT = 0.002), no association between PMPS and BD-PRS strata in controls was observed (F = 1.00, p = 0.323, η2p = 0.023). However, the PMPS at pT = 0.05 (n = 1,891 CpGs) showed stronger associations with BD-PRS strata (F = 11.57, p < 0.002, η2p = 0.212) than the PMPs at pT = 0.01 (F = 7.11, p = 0.011, η2p = 0.142); PMPS was higher in controls with High-BD-PRS (pT = 0.05; M ± SE = 11.84 ± 1.27) compared to low BD-PRS (M ± SE = 8.17 ± 1.34). Thus we considered pT = 0.05 to be optimal for use in Validation Set 2 (Table S9).
Validation Set 2
Associations between PMPS and PRS or clinical status in BD and CON groups
PMPS at pT = 0.05 was normally distributed within each clinical group (Shapiro-Wilk p > 0.05). PMPS did not differ between BD and control groups in a basic model that did not covary for BD-PRS (F = 2.39, p = 0.131, η2p = 0.066; Table S9). Following the inclusion of BD-PRS and BD-PRSxGroup term in the model, a significant association between PMPS and BD group was observed (F = 7.20, p = 0.011, η2p = 0.184; Table S9), in conjunction with a significant BD-PRSxGroup interaction (F = 7.07, p = 0.012, η2p = 0.181) and a suggestive effect of BD-PRS (F = 3.99, p = 0.054, η2p = 0.111).
Associations between PMPS and PRS or clinical status in ‘BD-syndromic’ and ‘disorder-free’ groups
Extended-clinical group definitions were utilised based on BD-psychopathology in HR (Table S8). HR meeting threshold or sub-threshold diagnoses (HRunwell) demonstrated lower functional capacity (CGI scores) than HR who remained well (HRwell) (Fig. S12). Overall, the ‘BD-syndromic’ group had poorer global functioning (GAF) than the ‘disorder-free’ group (M ± SD = 74.0 ± 14.7 vs. 92.7 ± 6.0, p = 2.75 × 10−6) (Fig. S6).
PMPS was not associated with psychopathology-group without BD-PRS in the model (F = 0.39, p = 0.533, η2p = 0.006). Following the inclusion of BD-PRS and BD-PRS × Group term in the model, a significant association between PMPS and BD group was observed (F = 5.55, p = 0.021, η2p = 0.076; Table S9), in conjunction with a significant BD-PRS × Group interaction (F = 5.52, p = 0.022, η2p = 0.075) and a non-significant effect of BD-PRS (F = 2.33, p = 0.132, η2p = 0.033).
A post-hoc sensitivity analysis of the impact of psychiatric medication use on PMPS was conducted within the BD-syndromic group, which showed no evidence of medication effects on PMPS (Supplementary Results). Exploratory regression analysis in the subset of participants with FACES data (n = 66; omnibus model p = 0.013), showed that PMPS was not associated with FACES (Wald-χ2 = 0.671, p = 0.413), but BD-PRS, psychopathology-group, and BD-PRS × Group were significant predictors of PMPS (Wald-χ2 = 9.55, p = 0.002; Wald-χ2 = 9.29, p = 0.002; Wald-χ2 = 10.17, p = 0.001; respectively), while the FACES × Group interaction was non-significant (Wald-χ2 = 3.24, p = 0.072).
Sensitivity analysis for population stratification
To ensure our results were robust to residual population stratification within our selected study samples, the most divergent four ‘nearest neighbour’ individuals were identified by pairwise IBS distances, and excluded if Z > 4 (n = 5 individuals total; n = 1 discovery EWAS, n = 3 validation set 1, n = 1 validation set 2). Removal of one participant from discovery EWAS with outlier values on C4 did not substantially alter the findings (effect size r = 0.9999; absolute β difference ~1.43 ± 1.49%). For validation set 1, three ‘nearest neighbour’ exclusions resulted in a slight attenuation of association between PMPS at pT = 0.05 and BD-PRS strata (F = 8.946, p = 0.005, η2p = 0.183). For validation set 2, ‘nearest neighbour’ exclusions (n = 1) revealed almost identical results and effect sizes (η2p ± 0.001).
Discussion
After stratifying HR individuals based on their personal burden of BD-associated common genetic variants indexed by BD-PRS and restricting the epigenome search space to variable probes with blood-brain correlation, we found a single epigenome-wide significant differentially methylated probe (cg00933603), located in an active regulatory element in exon 2 of the VARS2 gene, which lies in the major histocompatibility complex region (hg19/chr6:25–34 Mb). VARS2 encodes a mitochondrial aminoacyl-tRNA synthetase involved in mitochondrial protein synthesis. Mitochondrial abnormalities are evident in BD [76] and other psychiatric illnesses [77], and loss-of-function mutations in VARS2 have been previously associated with mitochondrial encephalopathies [78], epilepsy [79], and schizophrenia [80]. Furthermore, an EWAS of depressive symptoms in 724 monozygotic Danish twins identified a differentially methylated region in a putative active enhancer of VARS2, which spanned 9-probes including cg00933603 [81]. In our study, VARS2 was hypomethylated in individuals with a high polygenic burden for BD. Hypomethylation in the 5′ region of a gene can promote gene expression [82], which in the case of VARS2, might play a role in phasic dysregulation of mitochondrial bioenergetics associated with BD [83]. Further characterisation of this ubiquitously expressed functionally relevant gene in the pathophysiology of BD is required.
Beyond the top DMP, disease-associated genetic variants may broadly impact epigenetic processes, supporting the supposition that DNA methylation may mediate genetic risk [84], potentially via long-range epigenetic networks [85]. Of 1,183 genes that mapped to nominally significant DMPs, some were previously associated with BD or schizophrenia, including MLC1, ESR1, KCKN5, L1CAM, CPEB1 and GABBR2 [73]. In addition, 693 of these genes contained CpGs independently nominally associated with later development of BD and MDD in individuals at high familial risk of mood disorder [43]. Although overlap between the specific DMPs implicated in that prior study and our present analysis was low, gene-level convergence provides supporting evidence for specific differentially methylated genes in developing affective psychopathology.
While the subgrouping of HR participants was primarily focused on genetic predisposition to BD, the BD-PRS strata did not exclusively index BD-associated risk—a higher burden of cross-disorder and MDD-associated variants was observed—which supports a relative lack of specificity of PRS. It is therefore possible that the methylation signatures identified herein may show pleiotropic effects across these related psychiatric disorders.
Functional analyses indicated that genes differentially expressed in the brain were over-represented amongst the differentially methylated genes, particularly in the hypothalamus and anterior cingulate cortex (ACC). The hypothalamus is part of the hypothalamic-pituitary-adrenal (HPA) axis, the key stress-response system, which has been shown to be dysregulated in mood disorders [86]. In addition, the ACC plays an important role in cognitive functions and emotional regulation. Structural imaging meta-analyses have shown grey matter reduction in ACC [87], and smaller hippocampal volumes associated with BD [88, 89], thus methylation changes impacting these tissues are potentially of relevance. Overlap between genes mapped to DMPs and genes implicated in relevant phenotypes via GWAS—including BD and schizophrenia, chronotype, and risk tolerance—suggest convergence of epigenetic and genetic signals. Sleep disruption and chronotype have long been posited as hallmark features of BD [90], as well as targets of psychiatric medication [91], and may influence medication response [92]. Risky behaviour is a common symptom in manic phases and is genetically correlated with BD [93], highlighting relevant pathways within our defined epigenetic signals.
Restricting the discovery EWAS to putatively functionally relevant CpGs that reflect methylation status in the primary disease-affected tissue [62, 63] enabled reduction of the EWAS search space and application of a more permissive epigenome-wide significance threshold than previously modelled [71], but may consequentially bias enrichment analyses. Our secondary EWAS employed all variable CpGs, regardless of blood-brain correlation, and revealed similar over-representation in brain tissue, pathway enrichments, and overlap of top DMPs with an equivalent-sized gene list; supporting the primary methodological approach.
Utilising nominally differentially methylated CpGs from PRS-stratified EWAS in generating a poly-methylomic profile score (PMPS) [72]—akin to calculating PRS from GWAS summary statistics for genotypic data [14]—we validated the impact of BD-PRS on methylation signature. We found significant associations between BD-PRS and PMPS in controls, demonstrating the generalisability of the PMPS as an index of PRS, regardless of family history of BD. Consistent with the dependency of the PMPS on PRS background, when BD-PRS was included in the statistical models testing differences between groups in PMPS, a higher mean poly-methylomic profile score was observed in those with BD-symptomatology compared to disorder-free individuals. This highlights the important interplay between environmental, epigenetic and genetic risk factors, which may lead to the development of psychopathology, and is consistent with a multifactorial liability model [94].
While there was no significant effect of FACES on PMPS in our exploratory analysis, this study cannot rule out the potential influence of family environment on methylation signature, given the small sample size with available FACES that was used. The PMPS appears to at least partially reflect environmental differences related to the burden of BD-associated variants, however, increasing the sample size for discovery EWAS may further elucidate the mechanisms underlying the development of psychopathology. Longitudinal investigation of epigenetic markers in HR individuals based on their clinical status over time may also reveal useful epigenetic signatures of the BD prodrome.
Limitations
This study was restricted to participants of European-ancestry, due to substantial ethnicity-specific effects on PRS [60]. We note that residual population substructure not indexed by the covariates we employed (i.e. the first two MDS components or four surrogate variables of methylation data) may influence the results reported herein. However, expanding the existing 15 essential covariates to include additional MDS components would further reduce the ‘subjects per variable’ ratio, and compromise model stability and accuracy of estimation of regression coefficients [95]. This trade-off in design of our statistical model for analysis of this cohort may lead to under-correction of fine-scale population structure that is evident in European population [96, 97], although sensitivity analysis showed little effect of excluding potential outliers. To permit the study design to include both discovery and validation samples, sub-setting of the cohort was required, which limited sample size. While we employed a Bayesian method to reduce potential for EWAS bias [67], the methylation effects reported herein require validation. We mapped DMPs both physically and functionally [68], to comprehensively define genes associated with DMPs in both coding and non-coding genomic regions. However, genes were not prioritised for inclusion in enrichment analysis based on the number nor location of DMPs within each mapped gene, nor was the list restricted to one gene per DMP, thus enrichment signals may have been impacted by inclusion of genes whose expression was not affected by DMPs. Furthermore, in silico mapping of potentially cross-reactive CpG probes to homologous genome locations [98] may impact gene annotations and enrichments. Moreover, FUMA does not account for the number of CpGs per gene, therefore enrichment results may be biased towards categories with larger genes that contain more probes (i.e., with greater stochastic chance to be differentially methylated) [99].
One universal limitation of epigenome-wide studies in psychiatric disorders is the accessibility to brain tissue, as there is considerable variation in DNA methylation across tissues and cell types [100, 101]. We addressed this design limitation by covarying for inter-individual variability in blood-cell components, and focusing on CpGs which are blood-brain correlated [62]. However, use of blood-derived DNA may exclude relevant brain-specific CpGs that are not variable in this surrogate tissue. We acknowledge that: (1) participants were at different follow-up points, with potential limitations on capacity for defining ‘syndromic’ status that is dependent on length of clinical follow-up, and (2) PMPS reflects baseline methylation signatures, and timing of emergence of symptoms in relation to proximity to blood draw is not accounted for. Furthermore, while only two participants in the discovery EWAS were epigenetically inferred to be probable smokers (and were equally represented in PRS strata), smoking exposure may be a relevant confounder for future replication. Finally, psychiatric medication use was not controlled for due to the small numbers of individuals exposed, and although sensitivity analysis indicated non-significant effect of medication on PMPS, this remains an important caveat.
Although discovery EWAS in HR participants stratified by BD-PRS can reveal epigenetic markers associated with genetic risk for developing BD, many genetic variants that may impact BD risk are not indexed by PRS [18]. Indeed, familial risk encompasses all classes of genomic variation—including structural and rare variants—which may track more closely than PRS with disease-status in families with strong family history [21]. While we excluded annotated ‘probe SNPs’ from our analysis [102], the existence of polymorphic genotypes (e.g., rare or recently discovered DNA variants) that are present in our cohort and underlie CpGs could potentially lead to bias [103]. Conversely, we note that standard clumping procedures employed herein (based on r2) for generation of PRS included multiple MHC variants, thus PRS group differences may be inflated by multiple genetic effects from this region, which may amplify apparent effect sizes of methylation differences in the MHC region. The MHC region was also not excluded in FUMA enrichment analysis. Finally, limited availability of family environment data impacted power to detect effects, which require larger samples to definitively characterise. Thus extension to larger cohorts, and potential examination of the impact of other environmental factors, including stressful life-events, should be considered in future.
Conclusions
To our knowledge, this is the first report to investigate DNA methylation differences amongst individuals at high risk of BD, in the context of their personal polygenic background for BD. While only a single site in the VARS2 gene exceeded epigenome-wide significance, many CpG sites were nominally differentially methylated, which related to neurological pathways and functions associated with risk of psychopathology. Methylation profiling in independent validation sets confirmed the relationships between methylation signatures and genetic background, finding that methylation profiles may also partially reflect differences in family environment. Further larger-scale studies are needed to examine the impact of environmental factors in the relationship between familial risk and development of psychopathology.
Data availability
The data that support the findings of this study are available in the UNSW data-archive “ResData” at www.dataarchive.unsw.edu.au, under Research Data Management Plan reference number D0237303, or are available on request from the authors.
References
Smoller JW, Finn CT. Family, twin, and adoption studies of bipolar disorder. Am J Med Genet C Semin Med Genet. 2003;123C:48–58.
Craddock N, Jones I. Genetics of bipolar disorder. J Med Genet. 1999;36:585–94.
Craddock N, Sklar P. Genetics of bipolar disorder. Lancet. 2013;381:1654–62.
Mesman E, Nolen WA, Reichart CG, Wals M, Hillegers MH. The Dutch bipolar offspring study: 12-year follow-up. Am J Psychiatry. 2013;170:542–9.
Perich T, Lau P, Hadzi-Pavlovic D, Roberts G, Frankland A, Wright A, et al. What clinical features precede the onset of bipolar disorder? J Psychiatr Res. 2015;62:71–7.
Van Meter AR, Burke C, Youngstrom EA, Faedda GL, Correll CU. The bipolar prodrome: meta-analysis of symptom prevalence prior to initial or recurrent mood episodes. J Am Acad Child Adolesc Psychiatry. 2016;55:543–55.
Duffy A, Vandeleur C, Heffer N, Preisig M. The clinical trajectory of emerging bipolar disorder among the high-risk offspring of bipolar parents: current understanding and future considerations. Int J Bipolar Disord. 2017;5:37.
Frankland A, Roberts G, Holmes-Preston E, Perich T, Levy F, Lenroot R, et al. Clinical predictors of conversion to bipolar disorder in a prospective longitudinal familial high-risk sample: focus on depressive features. Psychol Med. 2018;48:1713–21.
Duffy A, Goodday S, Keown-Stoneman C, Grof P. The emergent course of bipolar disorder: observations over two decades from the canadian high-risk offspring cohort. Am J Psychiatry. 2019;176:720–9.
Sandstrom A, MacKenzie L, Pizzo A, Fine A, Rempel S, Howard C, et al. Observed psychopathology in offspring of parents with major depressive disorder, bipolar disorder and schizophrenia. Psychol Med. 2020;50:1050–6.
Sklar P, Ripke S, Scott LJ, Andreassen OA, Cichon S, Craddock N, et al. Large-scale genome-wide association analysis of bipolar disorder identifies a new susceptibility locus near ODZ4. Nat Genet. 2011;43:977–83.
Stahl EA, Breen G, Forstner AJ, McQuillin A, Ripke S, Trubetskoy V, et al. Genome-wide association study identifies 30 loci associated with bipolar disorder. Nat Genet. 2019;51:793–803.
Mullins N, Forstner AJ, O'Connell KS, Coombes B, Coleman JR, Qiao Z, et al. Genome-wide association study of more than 40,000 bipolar disorder cases provides new insights into the underlying biology. Nat Genet. 2021;53:817–29.
Purcell SM, Wray NR, Stone JL, Visscher PM, O’Donovan MC, Sullivan PF, et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature. 2009;460:748–52.
Lee SH, Ripke S, Neale BM, Faraone SV, Purcell SM, Perlis RH, et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat Genet. 2013;45:984–94.
Anttila V, Bulik-Sullivan B, Finucane HK, Walters RK, Bras J, Duncan L, et al. Analysis of shared heritability in common disorders of the brain. Science. 2018;360:eaap8757.
Cross-Disorder Group of the Psychiatric Genomics Consortium. Genomic relationships, novel loci, and pleiotropic mechanisms across eight psychiatric disorders. Cell. 2019;179:1469–1482.e11.
Fullerton JM, Nurnberger JI. Polygenic risk scores in psychiatry: will they be useful for clinicians? F1000Research. 2019;8:F1000 Faculty Rev−1293.
Wray NR, Lin T, Austin J, McGrath JJ, Hickie IB, Murray GK, et al. From basic science to clinical application of polygenic risk scores: a primer. JAMA Psychiatry. 2021;78:101–9.
Pettersson E, Lichtenstein P, Larsson H, Song J, Agrawal A, Børglum A, et al. Genetic influences on eight psychiatric disorders based on family data of 4 408 646 full and half-siblings, and genetic data of 333 748 cases and controls. Psychol Med. 2019;49:1166–73.
Toma C, Shaw AD, Allcock RJN, Heath A, Pierce KD, Mitchell PB, et al. An examination of multiple classes of rare variants in extended families with bipolar disorder. Transl Psychiatry. 2018;8:65.
Toma C, Shaw AD, Overs BJ, Mitchell PB, Schofield PR, Cooper AA, et al. De novo gene variants and familial bipolar disorder. JAMA Netw Open. 2020;3:e203382.
Jia X, Goes FS, Locke AE, Palmer D, Wang W, Cohen-Woods S, et al. Investigating rare pathogenic/likely pathogenic exonic variation in bipolar disorder. Mol Psychiatry. 2021;26:5239–50.
Yeung MW, Waye MYM. Genetics and epigenetics of bipolar disorder. J Biochem Mol Biol Post Genom Era. 2012;2:71.
Rowland TA, Marwaha S. Epidemiology and risk factors for bipolar disorder. Ther Adv Psychopharmacol. 2018;8:251–69.
Stapp EK, Mendelson T, Merikangas KR, Wilcox HC. Parental bipolar disorder, family environment, and offspring psychiatric disorders: a systematic review. J Affect Disord. 2020;268:69–81.
Shalev A, Merranko J, Goldstein T, Miklowitz DJ, Axelson D, Goldstein BI, et al. A longitudinal study of family functioning in offspring of parents diagnosed with bipolar disorder. J Am Acad Child Adolesc Psychiatry. 2019;58:961–70.
Mortensen PB, Pedersen C, Melbye M, Mors O, Ewald H. Individual and familial risk factors for bipolar affective disorders in Denmark. Arch Gen Psychiatry. 2003;60:1209–15.
Hart SA, Little C, van Bergen E. Nurture might be nature: cautionary tales and proposed solutions. NPJ Sci Learn. 2021;6:2.
Uher R, Zwicker A. Etiology in psychiatry: embracing the reality of poly‐gene‐environmental causation of mental illness. World Psychiatry. 2017;16:121–9.
Wu Y, Zhong X, Lin Y, Zhao Z, Chen J, Zheng B, et al. Estimating genetic nurture with summary statistics of multigenerational genome-wide association studies. Proc Natl Acad Sci USA. 2021;118:e2023184118.
Hu B, Cha J, Fullerton JM, Hesam-Shariati S, Nakamura K, Nurnberger JI, et al. Genetic and environment effects on structural neuroimaging endophenotype for bipolar disorder: a novel molecular approach. Transl Psychiatry. 2022;12:137.
Jirtle RL, Skinner MK. Environmental epigenomics and disease susceptibility. Nat Rev Genet. 2007;8:253–62.
Legrand A, Iftimovici A, Khayachi A, Chaumette B. Epigenetics in bipolar disorder: a critical review of the literature. Psychiatr Genet. 2021;31:1–12.
Sugawara H, Murata Y, Ikegame T, Sawamura R, Shimanaga S, Takeoka Y, et al. DNA methylation analyses of the candidate genes identified by a methylome‐wide association study revealed common epigenetic alterations in schizophrenia and bipolar disorder. Psychiatry Clin Neurosci. 2018;72:245–54.
Starnawska A, Demontis D, McQuillin A, O’Brien NL, Staunstrup NH, Mors O, et al. Hypomethylation of FAM63B in bipolar disorder patients. Clin Epigenetics. 2016;8:52.
Burghardt KJ, Khoury AS, Msallaty Z, Yi Z, Seyoum B. Antipsychotic medications and DNA methylation in schizophrenia and bipolar disorder: a systematic review. Pharmacotherapy. 2020;40:331–42.
Lichtenstein P, Yip BH, Björk C, Pawitan Y, Cannon TD, Sullivan PF, et al. Common genetic determinants of schizophrenia and bipolar disorder in Swedish families: a population-based study. Lancet. 2009;373:234–9.
Banerjee N, Polushina T, Bettella F, Giddaluru S, Steen VM, Andreassen OA, et al. Recently evolved human-specific methylated regions are enriched in schizophrenia signals. BMC Evol Biol. 2018;18:1–11.
Yao S, Guo Y, Ding J, Wu H, Hao R, Rong Y, et al. Epigenetic element-based transcriptome-wide association study identifies novel genes for bipolar disorder. Schizophr Bull. 2021;47:1642–52.
Lahiri DK, Maloney B, Bayon BL, Chopra N, White FA, Greig NH, et al. Transgenerational latent early-life associated regulation unites environment and genetics across generations. Epigenomics 2016;8:373–87.
Horsthemke B. A critical view on transgenerational epigenetic inheritance in humans. Nat Commun. 2018;9:2973.
Walker RM, Sussmann JE, Whalley HC, Ryan NM, Porteous DJ, McIntosh AM, et al. Preliminary assessment of pre‐morbid DNA methylation in individuals at high genetic risk of mood disorders. Bipolar Disord. 2016;18:410–22.
Roberts G, Lenroot R, Frankland A, Yeung P, Gale N, Wright A, et al. Abnormalities in left inferior frontal gyral thickness and parahippocampal gyral volume in young people at high genetic risk for bipolar disorder. Psychol Med. 2016;46:2083–96.
Overs BJ, Lenroot RK, Roberts G, Green MJ, Toma C, Hadzi-Pavlovic D, et al. Cortical mediation of relationships between dopamine receptor D2 and cognition is absent in youth at risk of bipolar disorder. Psychiatry Res Neuroimaging. 2021:111258.
Nurnberger JI, McInnis M, Reich W, Kastelic E, Wilcox HC, Glowinski A, et al. A high-risk study of bipolar disorder: childhood clinical phenotypes as precursors of major mood disorders. Arch Gen Psychiatry. 2011;68:1012–20.
Maxwell ME. Family Interview for Genetic Studies (FIGS): a manual for FIGS. Clinical Neurogenetics Branch, National Institute of Mental Health; 1992.
Kaufman J, Birmaher B, Brent D, Rao U, Flynn C, Moreci P, et al. Schedule for Affective Disorders and Schizophrenia for School-age Children-present and Lifetime Version (K-SADS-PL): initial reliability and validity data. J Am Acad Child Adolesc Psychiatry. 1997;36:980–8.
Nurnberger JI Jr, Blehar MC, Kaufmann CA, York-Cooler C, Simpson SG, Harkavy-Friedman J, et al. Diagnostic interview for genetic studies. Rationale, unique features, and training. NIMH Genetics Initiative. Arch Gen Psychiatry 1994;51:849–59.
Leckman JF, Sholomskas D, Thompson WD, Belanger A, Weissman MM. Best estimate of lifetime psychiatric diagnosis: a methodological study. Arch Gen Psychiatry. 1982;39:879–83.
American Psychiatric Association, Diagnostic and statistical manual of mental disorders. 4th ed. (DSM-IV). Washington DC: American Psychiatric Press Inc; 1994.
Busner J, Targum SD. The clinical global impressions scale: applying a research tool in clinical practice. Psychiatry. 2007;4:28–37.
Olson DH, Portner J, Bell RQ. FACES II: family adaptability and cohesion evaluation scale. St Paul: University of Minnesota, Department of Family Social Science; 1982.
Wilcox HC, Fullerton JM, Glowinski AL, Benke K, Kamali M, Hulvershorn LA, et al. Traumatic stress interacts with bipolar disorder genetic risk to increase risk for suicide attempts. J Am Acad Child Adolesc Psychiatry. 2017;56:1073–80.
Jamshidi J, Williams LM, Schofield PR, Park HR, Montalto A, Chilver MR, et al. Diverse phenotypic measurements of wellbeing: Heritability, temporal stability and the variance explained by polygenic scores. Genes Brain Behav. 2020;19:e12694.
Howard DM, Adams MJ, Clarke T-K, Hafferty JD, Gibson J, Shirali M, et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat Neurosci. 2019;22:343–52.
Euesden J, Lewis CM, O’Reilly PF. PRSice: polygenic risk score software. Bioinformatics 2015;31:1466–8.
Min JL, Hemani G, Davey Smith G, Relton C, Suderman M. Meffil: efficient normalization and analysis of very large DNA methylation datasets. Bioinformatics 2018;34:3983–9.
Fortin J-P, Fertig E, Hansen K shinyMethyl: interactive quality control of Illumina 450k DNA methylation arrays in R. F1000Research. 2014;3:175.
Curtis D. Polygenic risk score for schizophrenia is more strongly associated with ancestry than with schizophrenia. Psychiatr Genet. 2018;28:85–89.
Dawes K, Andersen A, Reimer R, Mills JA, Hoffman E, Long JD, et al. The relationship of smoking to cg05575921 methylation in blood and saliva DNA samples from several studies. Sci Rep. 2021;11:21627.
Hannon E, Lunnon K, Schalkwyk L, Mill J. Interindividual methylomic variation across blood, cortex, and cerebellum: implications for epigenetic studies of neurological and neuropsychiatric phenotypes. Epigenetics 2015;10:1024–32.
Edgar RD, Jones MJ, Robinson WP, Kobor MS. An empirically driven data reduction method on the human 450K methylation array to remove tissue specific non-variable CpGs. Clin Epigenetics. 2017;9:11.
Du P, Kibbe WA, Lin SM. lumi: a pipeline for processing Illumina microarray. Bioinformatics 2008;24:1547–8.
Leek JT, Johnson WE, Parker HS, Jaffe AE, Storey JD. The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics. 2012;28:882–3.
Smyth GK. Limma: linear models for microarray data, in Bioinformatics and computational biology solutions using R and bioconductor. Springer; 2005. p. 397–420.
van Iterson M, van Zwet EW, Heijmans BT, BIOS Consortium. Controlling bias and inflation in epigenome-and transcriptome-wide association studies using the empirical null distribution. Genome Biol. 2017;18:19.
McLean CY, Bristor D, Hiller M, Clarke SL, Schaar BT, Lowe CB, et al. GREAT improves functional interpretation of cis-regulatory regions. Nat Biotechnol. 2010;28:495–501.
Watanabe K, Taskesen E, Van Bochoven A, Posthuma D. Functional mapping and annotation of genetic associations with FUMA. Nat Commun. 2017;8:1–11.
Watanabe K, Mirkov MU, de Leeuw CA, van den Heuvel MP, Posthuma D. Genetic mapping of cell type specificity for complex traits. Nat Commun. 2019;10:1–13.
Saffari A, Silver MJ, Zavattari P, Moi L, Columbano A, Meaburn EL, et al. Estimation of a significance threshold for epigenome-wide association studies. Genet Epidemiol. 2018;42:20–33.
Watkeys OJ, Cohen-Woods S, Quidé Y, Cairns MJ, Overs B, Fullerton JM, et al. Derivation of poly-methylomic profile scores for schizophrenia. Prog Neuropsychopharmacol Biol Psychiatry. 2020;101:109925.
Piñero J, Ramírez-Anguita JM, Saüch-Pitarch J, Ronzano F, Centeno E, Sanz F, et al. The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res. 2020;48:D845–D855. D1
McRae AF, Marioni RE, Shah S, Yang J, Powell JE, Harris SE, et al. Identification of 55,000 Replicated DNA Methylation QTL. Sci Rep. 2018;8:17605.
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat Genet. 2013;45:580–5.
Scaini G, Andrews T, Lima CN, Benevenuto D, Streck EL, Quevedo JJM. Mitochondrial dysfunction as a critical event in the pathophysiology of bipolar disorder. Mitochondrion. 2021;57:23–36.
Anglin RE, Mazurek MF, Tarnopolsky MA, Rosebush PI. The mitochondrial genome and psychiatric illness. Am J Med Genet B Neuropsychiatr Genet. 2012;159:749–59.
Diodato D, Melchionda L, Haack TB, Dallabona C, Baruffini E, Donnini C, et al. VARS2 and TARS2 mutations in patients with mitochondrial encephalomyopathies. Hum Mutat. 2014;35:983–9.
Ruzman L, Kolic I, Radic Nisevic J, Ruzic Barsic A, Skarpa Prpic I, Prpic I. A novel VARS2 gene variant in a patient with epileptic encephalopathy. Ups J Med Sci. 2019;124:273–7.
Minică CC, Genovese G, Hultman CM, Pool R, Vink JM, Neale MC, et al. The weighting is the hardest part: On the behavior of the Likelihood Ratio Test and the Score Test under a data-driven weighting scheme in sequenced samples. Twin Res Hum Genet. 2017;20:108–18.
Starnawska A, Tan Q, Soerensen M, McGue M, Mors O, Borglum AD, et al. Epigenome-wide association study of depression symptomatology in elderly monozygotic twins. Transl Psychiatry. 2019;9:214.
Jones PA. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet. 2012;13:484–92.
Morris G, Walder K, McGee SL, Dean OM, Tye SJ, Maes M, et al. A model of the mitochondrial basis of bipolar disorder. Neurosci Biobehav Rev. 2017;74:1–20. Pt A
Starnawska A, Demontis D. Role of DNA methylation in mediating genetic risk of psychiatric disorders. Front Psychiatry. 2021;12:596821.
Lemire M, Zaidi SH, Ban M, Ge B, Aïssi D, Germain M, et al. Long-range epigenetic regulation is conferred by genetic variation located at thousands of independent loci. Nat Commun. 2015;6:6326.
Bao A-M, Swaab DF. The human hypothalamus in mood disorders: the HPA axis in the center. IBRO Rep. 2019;6:45–53.
Ellison-Wright I, Bullmore E. Anatomy of bipolar disorder and schizophrenia: a meta-analysis. Schizophr Res. 2010;117:1–12.
Haukvik UK, Gurholt TP, Nerland S, Elvsashagen T, Akudjedu TN, Alda M, et al. In vivo hippocampal subfield volumes in bipolar disorder-A mega-analysis from The Enhancing Neuro Imaging Genetics through Meta-Analysis Bipolar Disorder Working Group. Hum Brain Mapp. 2022;43:385–98.
Ching CRK, Hibar DP, Gurholt TP, Nunes A, Thomopoulos SI, Abe C, et al. What we learn about bipolar disorder from large-scale neuroimaging: Findings and future directions from the ENIGMA Bipolar Disorder Working Group. Hum Brain Mapp. 2022;43:56–82.
Melo MCA, Abreu RLC, Linhares Neto VB, de Bruin PFC, de Bruin VMS. Chronotype and circadian rhythm in bipolar disorder: A systematic review. Sleep Med Rev. 2017;34:46–58.
Lane JM, Vlasac I, Anderson SG, Kyle SD, Dixon WG, Bechtold DA, et al. Genome-wide association analysis identifies novel loci for chronotype in 100,420 individuals from the UK Biobank. Nat Commun. 2016;7:10889.
McCarthy MJ, Wei H, Nievergelt CM, Stautland A, Maihofer AX, Welsh DK, et al. Chronotype and cellular circadian rhythms predict the clinical response to lithium maintenance treatment in patients with bipolar disorder. Neuropsychopharmacology 2019;44:620–8.
Karlsson Linnér R, Biroli P, Kong E, Meddens SFW, Wedow R, Fontana MA, et al. Genome-wide association analyses of risk tolerance and risky behaviors in over 1 million individuals identify hundreds of loci and shared genetic influences. Nat Genet. 2019;51:245–57.
Stapp EK, Fullerton JM, Musci RJ, Zandi PP, McInnis MG, Mitchell PB, et al. Multifactorial threshold model in bipolar disorder: the interaction of family environment with polygenic risk. 2022. Submitted.
Austin PC, Steyerberg EW. The number of subjects per variable required in linear regression analyses. J Clin Epidemiol. 2015;68:627–36.
Cook JP, Mahajan A, Morris AP. Fine-scale population structure in the UK Biobank: implications for genome-wide association studies. Hum Mol Genet. 2020;29:2803–11.
Bycroft C, Freeman C, Petkova D, Band G, Elliott LT, Sharp K, et al. The UK Biobank resource with deep phenotyping and genomic data. Nature 2018;562:203–9.
Chen Y-A, Lemire M, Choufani S, Butcher DT, Grafodatskaya D, Zanke BW, et al. Discovery of cross-reactive probes and polymorphic CpGs in the Illumina Infinium HumanMethylation450 microarray. Epigenetics 2013;8:203–9.
Maksimovic J, Oshlack A, Phipson B. Gene set enrichment analysis for genome-wide DNA methylation data. Genome Biol. 2021;22:173.
Reinius LE, Acevedo N, Joerink M, Pershagen G, Dahlén S-E, Greco D, et al. Differential DNA methylation in purified human blood cells: implications for cell lineage and studies on disease susceptibility. PLOS one. 2012;7:e41361.
Gutierrez-Arcelus M, Ongen H, Lappalainen T, Montgomery SB, Buil A, Yurovsky A, et al. Tissue-specific effects of genetic and epigenetic variation on gene regulation and splicing. PLOS Genet. 2015;11:e1004958.
Price ME, Cotton AM, Lam LL, Farre P, Emberly E, Brown CJ, et al. Additional annotation enhances potential for biologically-relevant analysis of the Illumina Infinium HumanMethylation450 BeadChip array. Epigenetics Chromatin. 2013;6:4.
LaBarre BA, Goncearenco A, Petrykowska HM, Jaratlerdsiri W, Bornman MSR, Hayes VM, et al. MethylToSNP: identifying SNPs in Illumina DNA methylation array data. Epigenetics Chromatin. 2019;12:79.
Acknowledgements
This research was supported by the Australian National Medical and Health Research Council (NHMRC) Project Grants 1066177 (JMF) and 1063960 (JMF), Program Grant 1037196 (PBM, PRS) and Investigator Grants 1176716 (PRS) and 1177991 (PBM), and philanthropic support from the Lansdowne Foundation and Mrs Betty Lynch OAM (dec). Collection of samples and phenotyping at the US sites was supported by NIH Collaborative R01s MH068009 (JIN), MH073151, and MH068006 (MGM). Indiana University provided funding for the methylation analysis of participant samples collected at the US sites. SHS was supported by a University International Postgraduate Award from the University of New South Wales, and this work forms part of the first author’s doctoral dissertation. JMF was supported by the Janette Mary O’Neil Research Fellowship. EKS received support from the Intramural Research Program at the National Institute of Mental Health. MGM was supported in part by the HC Prechter Bipolar Research Program at the University of Michigan. DNA of Australian participants was extracted by Genetic Repositories Australia, an Enabling Facility that was supported by NHMRC Enabling Grant 401184. DNA of US participants was extracted by Rutgers University Cell and DNA repository. The authors wish to acknowledge the use of the services and facilities of the Australian Genome Research Facility (AGRF), for generation of methylation data. We thank Jessica Johnson and Doug Ruderfer for assistance with sample preparation and PsychArray genotyping at Mt Sinai School of Medicine, New York, USA. We thank Leanne M. Williams (School of Medicine, Stanford University, CA, USA) and Justine M. Gatt (School of Psychology, University of New South Wales, Sydney, Australia; and Neuroscience Research Australia, Sydney, Australia) for access to genotype data for the independent cohort of controls (PI: Williams; TWIN-E study) for determination of optimal p-value threshold for PRS in the high-risk participants. Recruitment of the TWIN-E cohort was supported by an Australian Research Council Linkage Grant LP0883621 (LMW, JMG, PRS) and genotyping was supported by National Health and Medical Research Council (NHMRC) Program Grant 1037196 (PBM, PRS). This cohort collection utilised Twins Research Australia, a national resource supported by NHMRC Centre of Research Excellence Grant (1079102). Finally, we thank and acknowledge all research participants who gave their time to contribute to the “Adolescents at High Risk for Familial Bipolar Disorder” and “Bipolar Kids and Sibs” research programmes, whose generosity and ongoing support has made this research possible.
Author information
Authors and Affiliations
Contributions
The study was conceptualised by SHS, BJO, CT, PRS, and JMF. Analysis methodology was developed by SHS, BJO and JMF, and administrative efforts and resources were provided by JMF, PRS, JIN, and PBM. HJE provided DNA samples from US participants, and DNA samples were received and prepared by SHS and KDP for methylation typing, and by KDP and HJE for genotyping. Formal analysis was conducted by SHS as part of a PhD thesis dissertation, with supervision by JMF and PRS. The PMPS scoring procedure was developed by OJW, with supervision by MJG and JMF. Genotype cleaning, imputation and quality control were performed by CT and JMF, prior to generation of polygenic scores by SHS. Clinical and family environment data were provided by GR, EKS, HCW, MGM, LAH, JIN, and PBM. The original draft was written by SHS, with primary review & editing by BJO and JMF, before being circulated for feedback and editing by all coauthors, each of whom approved the final manuscript. HJE provided particularly helpful critical reading and advice. Funding relevant to the current work was acquired by SHS (UNSW University International Postgraduate Award scholarship; with the support of JMF and PRS), and was supported by funding acquired by JMF, PRS, PBM, MGM, and JIN.
Corresponding author
Ethics declarations
Competing interests
JIN is an investigator for Janssen, with interests unrelated to the current work. MGM has consulted with Janssen Pharmaceuticals and received research support from Janssen, both unrelated to this work. All other authors have no potential or perceived conflicts of interest to declare.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hesam-Shariati, S., Overs, B.J., Roberts, G. et al. Epigenetic signatures relating to disease-associated genotypic burden in familial risk of bipolar disorder. Transl Psychiatry 12, 310 (2022). https://doi.org/10.1038/s41398-022-02079-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-022-02079-6
- Springer Nature Limited
This article is cited by
-
Putative Risk Biomarkers of Bipolar Disorder in At-risk Youth
Neuroscience Bulletin (2024)