Abstract
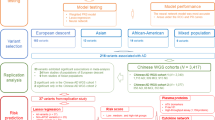
Late-onset Alzheimer’s disease (LOAD) is the most common multifactorial neurodegenerative disease among elderly people. LOAD is heterogeneous, and the symptoms vary among patients. Genome-wide association studies (GWAS) have identified genetic risk factors for LOAD but not for LOAD subtypes. Here, we examined the genetic architecture of LOAD based on Japanese GWAS data from 1947 patients and 2192 cognitively normal controls in a discovery cohort and 847 patients and 2298 controls in an independent validation cohort. Two distinct groups of LOAD patients were identified. One was characterized by major risk genes for developing LOAD (APOC1 and APOC1P1) and immune-related genes (RELB and CBLC). The other was characterized by genes associated with kidney disorders (AXDND1, FBP1, and MIR2278). Subsequent analysis of albumin and hemoglobin values from routine blood test results suggested that impaired kidney function could lead to LOAD pathogenesis. We developed a prediction model for LOAD subtypes using a deep neural network, which achieved an accuracy of 0.694 (2870/4137) in the discovery cohort and 0.687 (2162/3145) in the validation cohort. These findings provide new insights into the pathogenic mechanisms of LOAD.
Similar content being viewed by others
Introduction
Alzheimer’s disease (AD) is the most common cause of dementia among elderly people [1, 2]. The majority of AD cases are sporadic late-onset AD (LOAD), diagnosed in people ≥65 years [3]. The diagnosis of LOAD is characterized by the accumulation of amyloid-beta (Aβ) plaques and tau neurofibrillary tangles in the neurodegenerative brain [4], but it is a heterogeneous disease and the symptoms vary among patients. Previous studies have reported some subtypes of LOAD [5, 6]. Bredesen reported three LOAD subtypes (inflammatory, non-inflammatory, and cortical) after a metabolic profiling analysis of patients with cognitive decline [5]. Byun et al. identified four LOAD subtypes with heterogeneous patterns of regional atrophy from MRI-measured volumes of the hippocampus and cortical regions (both impaired, hippocampal atrophy only, cortical atrophy only, and both spared) [7]. However, these findings were derived from small samples, limiting their interpretation.
Genome-wide association studies (GWAS) have been highly successful in identifying genetic risk factors associated with LOAD [8]. The strongest genetic risk factor for LOAD is the apolipoprotein E ε4 (APOE4) polymorphism [9]. Others include bridging integrator 1 (BIN1) [10], clusterin (CLU) [10], and triggering receptors expressed on myeloid cells 2 (TREM2) [11], also identified from LOAD GWAS analyses. Risk prediction models using these genetic risk factors have also been developed for LOAD using various computational approaches [12, 13], including machine learning techniques [14]. With a large sample size, determining genetic risk will likely not only contribute to our understanding of the underlying pathologic mechanisms of LOAD but will also provide new insights into the classification of distinct LOAD subtypes. However, no previous studies have been conducted on genetic risks for the determination of LOAD subtypes.
Here, we comprehensively investigated the genetic architecture of LOAD based on Japanese GWAS data from a large number of patients and controls and examined the presence of LOAD subtypes using the energy landscape [15]. Each subject was represented as a binary vector based on the association signals obtained from GWAS. The energy landscape was visualized with disconnectivity graphs. We revealed the presence of two distinct groups of LOAD patients. Subsequent analysis of representative association signals from each group provided evidence that one group was characterized by both major risk genes for developing LOAD and immune-related genes, and the other group was characterized by genes associated with kidney disorders. We further found that LOAD patients had significantly decreased levels of serum albumin and hemoglobin, which are used to assess kidney function. These results suggest that impaired kidney function could be involved in LOAD pathogenesis. Our findings will contribute to a better understanding of the mechanisms driving heterogeneity in LOAD and will provide novel insight into potential subtype-specific therapies as a step toward precision medicine.
Materials and methods
Ethics statements
This study was approved by the ethics committee of the National Center for Geriatrics and Gerontology (NCGG). The design and performance of the current study involving human subjects were clearly described in a research protocol. All participation was voluntary, and all participants completed informed consent in writing before registering with the NCGG Biobank.
Clinical samples
Genome-wide genotyping data from the blood samples of a total of 7284 subjects used in this study and their associated clinical data were downloaded from the NCGG Biobank database. Of the total, 4139 subjects, composed of 1947 AD subjects and 2192 cognitively normal (CN) subjects, were genotyped using the Affymetrix Japonica Array (called the discovery cohort). The remaining 3145 subjects, composed of 847 AD subjects and 2298 CN subjects, were genotyped using the Infinium Asian Screening Array (called the validation cohort). The AD subjects were diagnosed with probable or possible AD using the criteria of the National Institute on Aging and Alzheimer’s Association workgroups [16, 17]. The CN subjects had subjective cognitive complaints, but normal cognition (score >23) on a neuropsychological assessment with a comprehensive neuropsychological test, the Mini-Mental State Examination. The diagnosis of all subjects was conducted based on medical history, physical examination and diagnostic tests, neurological examination, neuropsychological tests, and brain imaging with magnetic resonance imaging or computerized tomography by experts including neurologists, psychiatrists, geriatricians, or a neurosurgeon, all experts in dementia who are familiar with its diagnostic criteria. All subjects were ≥60 years of age.
Quality control in the GWAS
Genotype imputation was conducted by using IMPUTE2 [18] with the 3.5k Japanese reference panel developed by the Tohoku Medical Megabank Organization (https://www.megabank.tohoku.ac.jp/english/) for the discovery cohort and the 5.6k in-house reference panel for the validation set. We used imputed variants with an INFO score ≥0.4. Quality control was performed in each dataset separately after imputation using PLINK software [19]. We first applied quality control (QC) filters: (1) sex inconsistencies (--check-sex); (2) inbreeding coefficient (--het 0.1); (3) genotype missingness (--missing 0.05); (4) PI_HAT >0.25, where PI_HAT is a statistic for the proportion of identity by descent (--genome); and (5) exclusion of outliers from the clusters of East Asian populations in a principal component analysis (PCA) that was conducted together with 1000 Genomes Phase 3 data. We next applied QC filters to the genetic markers (single-nucleotide polymorphisms [SNPs] and short insertions and deletions [Indels]): (1) genotyping efficiency or call rate (--geno 0.95), (2) minor allele frequency (--freq 0.001), and (3) Hardy–Weinberg equilibrium (--hwe 0.001). To generate a set of independent SNPs, we further performed linkage disequilibrium–based SNP pruning using the statistical analysis program PLINK, version 1.90b [20] with a window size of 50 SNPs, a step of 5 SNPs, and a pairwise r2 threshold of 0.1 (--indep-pairwise 50 5 0.1).
Disconnectivity graph
The autosomal variants that passed QC criteria as described above were assessed with a logistic regression model, adjusting for sex and age with PLINK software (--logistic) [19] in the following way:
The coefficient and p-value of each variant were obtained from the discovery set. Using the variants with p < 0.01 weighted by their coefficients, a PCA analysis was performed. The principal component (PC) scores were binarized (i.e., −1 or +1) by the mean of the eigenvector values. Each subject was represented by a binary vector \(V_k\) = (\(\sigma _{1,}\sigma _{2, \cdots ,}\sigma _N\)) of \(2^N\), the appearance probability \(P(V_k)\) of which was applied to the pairwise maximum entropy model (i.e., Boltzmann distribution). Energy values of \(2^N\) binary vectors were compared to identify local energy minimums. By using a disconnectivity graph labeling the energy local minimum states, energy landscapes [15] can be visualized using the R packages akima and rgl (Supplementary Fig. 1).
RNA-sequencing data analysis
All RNA-sequencing (RNA-seq) data were downloaded from the NCGG Biobank database [21]. The quality of the read sequences (fastq files) was assessed by using FastQC (version 0.11.7). The low-quality reads (<Q20) and trimmed reads with adapter sequences (shorter than 50 bp) were excluded by using Cutadapt (version 1.16). The remaining clean sequenced reads were mapped to the human reference genome (GRCh37) with STAR (version 2.5.2b) [22]. Quantification, in transcripts per million, was performed with RSEM (version 1.3.0).
qRT-PCR validation of genes
cDNA was synthesized by using a PrimeScript II 1st Strand cDNA Synthesis Kit (Takara Bio, Shiga, Japan). quantitative RT-PCR (qRT-PCR) analysis was performed by using TaqMan Fast Advanced Master Mix (Thermo Fisher Scientific, Waltham, MA) and TaqMan Probes (Thermo Fisher Scientific), in accordance with the manufacturer’s instructions and the Quantstudio7 Flex Real-Time PCR System (Thermo Fisher Scientific). Target genes and their corresponding TaqMan probes were RELB (Hs00232399_m1) and FBP1 (Hs00983323_m1). The qRT-PCR conditions were one cycle of 50 °C for 2 min, 95 °C for 20 s followed by 40 cycles of 95 °C for 1 s, and 60 °C for 20 s. Human beta-actin (ACTB, Hs99999903_m1) was preselected as a reference gene for the normalization of target gene expression levels. Gene expression levels from qRT-PCR were calculated relative to that of the reference gene ACTB by using the semiquantitative method [23]. Gene expression levels of RELB were obtained for 16 randomly selected subjects with heterozygous genotypes and 20 randomly selected subjects with homozygous genotypes. Those of FBP were obtained for eight randomly selected subjects with heterozygous genotypes and 20 randomly selected subjects with homozygous genotypes.
Construction of a LOAD subtype prediction model
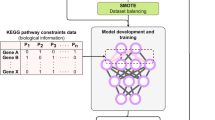
All data were strictly separated into a discovery cohort and a validation cohort. A PCA analysis was performed using variants with GWAS p < 0.01 in the discovery cohort. PC scores obtained from the PCA analysis were used to construct a disconnectivity graph. To estimate the PC scores (PC1 to PC10) from the variant data, prediction models were developed by using a deep neural network model. The neural networks comprised six hidden layers of 512 neurons along with randomized leaky rectified linear unit (RReLU) activation and 50% dropout. The networks were trained for 100 to 1500 epochs at 100 intervals with a batch size of 32 using the discovery cohort.
Estimated PC scores were binarized (i.e., −1 or +1) by the mean of eigenvector values used to construct the disconnectivity graph. Each subject was represented by a binary vector Vk = (\(\sigma _{1,}\sigma _{2, \cdots ,}\sigma _N\)) of \(2^N\), from which the position on the disconnectivity graph was detected. Prediction models were assessed by the rate at which LOAD cases and CN subjects were predicted in the red and green spheres of the same groups on the disconnectivity graph. The LOAD subtype prediction models were evaluated on the independent validation cohort. These models were constructed using the open-source machine learning library PyTorch (version 1.7.0).
Results
GWAS of the Japanese subjects
The discovery cohort consisted of 1947 LOAD cases and 2192 CN subjects. The average ages of the subjects from whom the LOAD and CN blood samples were obtained was 79.1 years (SD = 6.3 years) and 70.8 years (SD = 5.9 years), respectively, and the female-to-male ratios were 2.13:1 and 1.19:1, respectively. We examined association signals from the discovery cohort. A total of 9,252,465 genetic markers (SNPs and Indels) passed stringent QC filters for both genotypes and samples after genotype imputation. The QC-passed genetic markers were further pruned for linkage disequilibrium to obtain a set of independent markers. A subset of 2,741,829 genetic markers was used for subsequent analysis.
The validation cohort consisted of 847 LOAD cases and 2298 CN subjects. The average ages of the subjects from whom the LOAD and CN samples were obtained was 79.4 years (SD = 6.4 years) and 75.4 years (SD = 4.3 years), respectively, and the female-to-male ratios were 1.91:1 and 1.27:1, respectively.
Disconnectivity graph construction
Of the 2,741,829 genetic markers from the discovery cohort, 22,128 showed association signals with GWAS p < 0.01. Using the association signals, we performed a PCA. According to the PC scores, subjects were binarized to 27 binary vectors. Local energy minimum states were obtained from a comparison of energy values among the binary vectors, and the energy landscape was visualized with disconnectivity graphs labeling their local energy minimum states. In Fig. 1a, LOAD cases and CN subjects are represented as red and green spheres whose sizes correspond to the difference in LOAD and CN frequencies (i.e., the red spheres represent that LOAD cases are more frequent than controls and the green spheres represent that CN subjects are more frequent than LOAD cases). Both LOAD cases and CN subjects created clusters, and we revealed the presence of two distinct groups in LOAD and in CN (Fig. 1b).
The energy landscape was visualized by using 3D (a) and 2D (b) disconnectivity graphs, where all samples were classified into two groups. LOAD cases and CN subjects are represented in red and green spheres, and their sizes correspond to the difference of LOAD and CN frequencies at each node. b Gray circles represent to the same frequency in LOAD and CN subjects.
Demographic data of the two LOAD groups
The discovery cohort was classified into two distinct groups: group 1 was composed of 918 AD cases and 895 CN subjects, and group 2 contained 1029 AD cases and 1297 CN subjects. The demographic data of the subjects in each group are shown in Table 1. No differences were observed between the two groups in the average age and female-to-male ratio, but statistically significant differences were found in the number of APOE4 alleles between the LOAD subjects in each group (Fisher’s exact test p = 3.70 × 10−12) and between the CN subjects in each group (Fisher’s exact test p = 1.06 × 10−8), with the proportion of subjects with APOE4 higher in group 1 than in group 2. When visualizing the energy landscape by APOE4 and APOE2 information, significant positive correlations between LOAD frequency and APOE4 frequency were observed, with a higher correlation coefficient in group 1 (Kendall’s τ = 0.48, p = 6.72 × 10−7) than in group 2 (Kendall’s τ = 0.41, p = 1.59 × 10−6) (Supplementary Fig. 2).
Representative association signals of each group and their functional annotations
To identify distinctive association signals in each group, we focused on representative clusters where LOAD and CN frequencies differed by more than 5% (i.e., the red or green spheres in Fig. 1 were large). We observed six representative clusters (three red and three green spheres) in group 1, which contained 20,792 genetic markers from 848 subjects (391 LOAD cases and 457 CN samples), and five representative clusters (two red and three green spheres) in group 2, which contained 19,929 genetic markers from 1047 subjects (305 LOAD cases and 742 CN samples). Association signals were detected with logistic regression, adjusting for sex and age. From group 1, four genetic markers, located on or near four genes (APOC1, APOC1P1, RELB, and CBLC) showed statistically significant associations at Bonferroni-corrected p < 0.05; from group 2, 12 genetic markers, located on or near four genes (AXDND1, FBP1, AOPEP, and MIR2278) had significant associations (Table 2). Group 1 was characterized by major risk genes for developing LOAD (APOC1 and APOC1P1) [24] and immune-related genes (RELB and CBLC) [25,26,27], whereas group 2 was characterized by genes associated with kidney disorders, such as nephrotic syndrome (AXDND1, FBP1, and MIR2278) [28, 29]. Our results were consistent with previous findings of associations between kidney disease and AD [30] and between the immune system and AD [31].
Closest gene expression of association signals using blood samples
We used blood samples to examine whether the association signals detected affect expression of their closest genes. Of the eight genes, the Human Protein Atlas (HPA) database (https://www.proteinatlas.org) indicated that two (RELB = 2.0 and FBP1 = 33.1) are expressed in blood cells, where expressed genes were defined as genes with normalized transcripts per million ≥2 in total human peripheral blood mononuclear cells. Although MIR2278 and APOC1P1 have not been registered in the HPA database, the HPA database showed that APOC1, AOPEP, RELB, and FBP1 are expressed in brain tissues (Fig. 2a).
a The expression of six genes in the blood (red) and brain tissues (yellow) was checked in the HPA database. The x-axis represents the resulting transcript expression values, denoted as normalized transcripts per million (nTPM). b RNA-seq data of 126 subjects and 151 subjects were available from the NCGG Biobank database for RELB and FBP1, respectively. The effects of association signals on the expression of their nearby genes were examined by using linear regression adjusted for age and sex. The association signal rs146190016 significantly increased RELB expression (p = 0.019), but the signal rs550833079 did not significantly change FBP1 expression (p = 0.37). c We used quantitative RT-PCR (qRT-PCR) data to validate eQTL results for these genes (RELB and FBP1). The results were consistent with the RNA-seq results for both genes. Data in (b) and (c) are represented as box and whisker plots, depicting minimum, lower quartile (Q1), mean (Q2), upper quartile (Q3), and maximum values.
Association signals of RELB and FBP1 (rs146190016 and rs550833079) were located within their genes. The whole-blood RNA-seq data are available from the NCGG Biobank database. We examined the effect of the association signals on the expression of these genes by using linear regression adjusting for age and sex. We downloaded data on RELB for 126 of the 848 subjects in group 1 and data on RBP1 for 151 of the 1047 subjects in group 2 from the NCGG Biobank database. Although rs550833079 was not associated with a significant expression change in FBP1 (p = 0.37), rs146190016 contributed to a statistically significant increase of RELB expression (p = 0.019, Table 2 and Fig. 2b). To validate the expression Quantitative Trait Locus (eQTL) results for these genes, we used qRT-PCR data, which were consistent with the RNA-seq results for both genes (Fig. 2c).
Assessment of kidney function
Five measurements for assessment of kidney function (creatinine, cystatin C, eGFR, albumin [Alb], and hemoglobin [Hb]) were measured in routine blood tests. We obtained these data for 387 (342 LOAD cases and 45 CN subjects) from group 1 and 355 (269 LOAD cases and 86 CN subjects) from group 2. The difference in the measurements between LOAD and CN was determined with the Wilcoxon rank-sum test. We found that albumin and hemoglobin reached a false discovery rate (FDR) of significance (FDRAlb = 0.012 in group 1; FDRAlb = 1.30 × 10−6, FDRHb = 4.53 × 10−6 in group 2, Fig. 3), although the remaining three measures showed no statistically significant difference between LOAD and CN in either group. These results support the hypothesis that impaired kidney function can lead to LOAD pathogenesis.
We examined five markers of kidney function measured in routine blood tests (creatinine, cystatin C, eGFR, albumin [Alb], and hemoglobin [Hb]). The differences in the results between LOAD and CN were tested with the Wilcoxon rank-sum test. Data were represented as box and whisker plots, depicting minimum, lower quartile (Q1), mean (Q2), upper quartile (Q3), and maximum values. *FDR < 0.05, **FDR < 0.001.
As type 2 diabetes mellitus (T2D) is known to increase the risk for dementia, including AD [32], we examined the level of HbA1c measured in routine blood tests. We obtained the data for 388 (343 LOAD cases and 45 CN subjects) from group 1 and 355 (269 LOAD cases and 86 CN subjects) from group 2. The difference in levels between LOAD and CN was determined with the Wilcoxon rank-sum test, but there were no statistically significant differences between LOAD and CN subjects in both groups (P = 0.687 in group 1; P = 0.696 in group 2, Supplementary Fig. 3).
LOAD subtype prediction model
A disconnectivity graph was constructed based on PC scores, obtained from the PCA analysis using 22,128 variants with GWAS p < 0.01 in the discovery cohort. Of them, 17,743 variants were common with those in the independent validation cohort (see the Methods). The genomic variants could predict LOAD subtypes of patients. We applied a deep neural network with six hidden layers of 512 neurons along with RReLU activation, 50% dropout, and a batch size of 32 to predict PC scores from variant data of the discovery cohort (Fig. 4a). The predicted PC scores were binarized by the mean of eigenvector values used for the construction of the disconnectivity graph. Each subject was represented by a binary vector \(V_7\) = (\(\sigma _{1,}\sigma _{2, \cdots ,}\sigma _7\)) of 27, from which the location on the disconnectivity graph was determined. Prediction models were assessed by the rate at which LOAD cases and CN subjects were predicted in the red and green spheres of the same groups on the disconnectivity graph. We constructed neural network models with several hidden layers (Layer = 1, 2, …,7) and assessed them using the independent validation cohort (Supplemental Fig. 4). Our LOAD subtype prediction model (i.e., a deep neural network model comprised of six hidden layers) achieved an accuracy of 0.694 (2870/4137) in 900 epochs in the discovery cohort (group 1, 1228/1812 = 0.678; group 2, 1642/2325 = 0.706) and of 0.687 (2162/3145) in the validation cohort (group 1, 1025/1508 = 0.680; group 2, 1137/1637 = 0.695, Fig. 4b).
a We applied a deep neural network with six hidden layers of 512 neurons along with RReLU activation, 50% dropout, and a batch size of 32 to predict PC scores from variant data of the discovery cohort. b The networks were trained for 100 to 1500 epochs at 100 intervals using the discovery cohort. The best model achieved an accuracy of 0.694 in 900 epochs in the discovery cohort and 0.687 in the independent validation cohort.
Discussion
Using the genotyping data from a case-control cohort of >4100 subjects, we examined genetic risks for the determination of LOAD subtypes and revealed the presence of two distinct groups of LOAD patients. Clinical characteristics of the groups showed a significant difference in the frequency of APOE4 alleles, although no differences were observed in the average age and the female-to-male ratio between groups. To understand the genetic difference between groups, we further investigated representative association signals in each group. Four representative genes were detected from each group: group 1, APOC1, APOC1P1, RELB, and CBLC; group 2, AXDND1, FBP1, AOPEP, and MIR2278.
The four genes in group 1 (APOC1, APOC1P1, RELB, and CBLC) are located near the APOE region, and APOC1 and APOC1P1 are in linkage disequilibrium with the APOE gene. However, previous studies have shown that they are APOE-independent risk factors for AD [24]. RELB is a member of the nuclear factor kappa B (NF-κB) family [25]. Activated NF-κB has been found in neurons and glial cells in the brains of AD patients [33]. CBLC (Cbl proto-oncogene C) is a member of the Cbl family of E3 ubiquitin ligases. Recent studies have shown that CBLC is likely to indirectly modulate the stimulatory function of NF‐κB complexes [34]. Nho et al. reported that RELB and CBLC were significantly associated with LOAD-related cerebrospinal fluid Aβ1–42 with adjustment for APOE genotype [35]. These results suggest that the representative genes in group 1 are APOE-independent risk factors for LOAD and are likely to contribute to Aβ accumulation through direct or indirect NF‐κB activation.
On the other hand, three of the four representative genes in group 2 (AXDND1, FBP1, and MIR2278) are likely to be associated with kidney diseases. Several variants in AXDND1 have been registered as mutations involved in nephrotic syndrome in the GeneCards [28] and ClinVar databases [36]. FBP1 (fructose-bisphosphatase1) is a gluconeogenesis regulatory enzyme, which is suppressed in kidney tumors [37]. Although AOPEP has not been shown to be associated with kidney diseases, MIR2278 (microRNA 2278) can target STAT5A [38], which has been reported to contribute to the pathogenesis of kidney disease [39]. We further examined markers for kidney function measured in routine blood test results. Statistically significant differences between LOAD and CN were found in albumin and hemoglobin concentrations. These results suggest that impaired kidney function in group 2 LOAD patients could have led to AD pathogenesis. Wu et al. reported that AD severity was significantly associated with serum albumin and hemoglobin [40].
In this study, we revealed the presence of two distinct groups of LOAD patients through the genetic architecture of LOAD based on Japanese GWAS data, clearly revealing the presence of LOAD subtypes. We also used a deep neural network to create a LOAD subtype prediction model that achieved high accuracies in the discovery and validation cohorts, as our model can predict LOAD subtypes and phenotype simultaneously (i.e., LOAD group 1, LOAD group 2, CN group 1, and CN group 2). We believe that our findings will contribute to the understanding of LOAD and provide new insights into its pathogenic mechanisms. Many clinical trials using anti-Aβ antibody drugs, even Lecanemab, have not shown convincing results. This might be due to the existence of LOAD subtypes with different genetic backgrounds. Additional association studies between LOAD and CN using whole-genome sequencing data from large samples including non-Japanese will undoubtedly lead to further understanding of LOAD subtypes in the future.
Data availability
All GWAS summary datasets used or analyzed in the current study are available from the corresponding author on reasonable request.
Code availability
We used open-source program languages R (version 3.4.3) and Ruby (version 2.4.0) and the machine learning library PyTorch (version 1.7.0) to analyze data and create plots. Code is available upon request from the corresponding author.
References
Hardy J, Selkoe DJ. The amyloid hypothesis of Alzheimer’s disease: progress and problems on the road to therapeutics. Science. 2002;297:353–6.
Prince M, Bryce R, Albanese E, Wimo A, Ribeiro W, Ferri CP. The global prevalence of dementia: a systematic review and metaanalysis. Alzheimers Dement. 2013;9:63–75. e62
Rabinovici GD. Late-onset Alzheimer disease. Continuum. 2019;25:14–33.
Huber CM, Yee C, May T, Dhanala A, Mitchell CS. Cognitive decline in preclinical Alzheimer’s disease: amyloid-beta versus tauopathy. J Alzheimers Dis. 2018;61:265–81.
Bredesen DE. Metabolic profiling distinguishes three subtypes of Alzheimer’s disease. Aging. 2015;7:595–600.
Ferreira D, Nordberg A, Westman E. Biological subtypes of Alzheimer disease: a systematic review and meta-analysis. Neurology. 2020;94:436–48.
Byun MS, Kim SE, Park J, Yi D, Choe YM, Sohn BK, et al. Heterogeneity of regional brain atrophy patterns associated with distinct progression rates in Alzheimer’s disease. PLoS ONE. 2015;10:e0142756.
Bellenguez C, Kucukali F, Jansen IE, Kleineidam L, Moreno-Grau S, Amin N, et al. New insights into the genetic etiology of Alzheimer’s disease and related dementias. Nat Genet. 2022;54:412–36.
Liu CC, Liu CC, Kanekiyo T, Xu H, Bu G. Apolipoprotein E and Alzheimer disease: risk, mechanisms and therapy. Nat Rev Neurol. 2013;9:106–18.
Santos LRD, Almeida JFF, Pimassoni LHS, Morelato RL, Paula F. The combined risk effect among BIN1, CLU, and APOE genes in Alzheimer’s disease. Genet Mol Biol. 2020;43:e20180320.
Olive C, Ibanez L, Farias FHG, Wang F, Budde JP, Norton JB, et al. Examination of the effect of rare variants in TREM2, ABI3, and PLCG2 in LOAD through multiple phenotypes. J Alzheimers Dis. 2020;77:1469–82.
Sims R, Hill M, Williams J. The multiplex model of the genetics of Alzheimer’s disease. Nat Neurosci. 2020;23:311–22.
Jia L, Li F, Wei C, Zhu M, Qu Q, Qin W, et al. Prediction of Alzheimer’s disease using multi-variants from a Chinese genome-wide association study. Brain. 2021;144:924–37.
De Velasco Oriol J, Vallejo EE, Estrada K, Tamez Pena JG, Disease Neuroimaging Initiative TA. Benchmarking machine learning models for late-onset alzheimer’s disease prediction from genomic data. BMC Bioinform. 2019;20:709.
Ezaki T, Watanabe T, Ohzeki M, Masuda N. Energy landscape analysis of neuroimaging data. Philos Trans A Math Phys Eng Sci. 2017;375:20160287.
McKhann GM, Knopman DS, Chertkow H, Hyman BT, Jack CR Jr, Kawas CH, et al. The diagnosis of dementia due to Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement. 2011;7:263–9.
Albert MS, DeKosky ST, Dickson D, Dubois B, Feldman HH, Fox NC, et al. The diagnosis of mild cognitive impairment due to Alzheimer’s disease: recommendations from the National Institute on Aging-Alzheimer’s Association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement. 2011;7:270–9.
Howie B, Fuchsberger C, Stephens M, Marchini J, Abecasis GR. Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat Genet. 2012;44:955–9.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Slifer SH. PLINK: key functions for data analysis. Curr Protoc Hum Genet. 2018;97:e59.
Shigemizu D, Mori T, Akiyama S, Higaki S, Watanabe H, Sakurai T, et al. Identification of potential blood biomarkers for early diagnosis of Alzheimer’s disease through RNA sequencing analysis. Alzheimers Res Ther. 2020;12:87.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Marone M, Mozzetti S, De Ritis D, Pierelli L, Scambia G. Semiquantitative RT-PCR analysis to assess the expression levels of multiple transcripts from the same sample. Biol Proced Online. 2001;3:19–25.
Prendecki M, Florczak-Wyspianska J, Kowalska M, Ilkowski J, Grzelak T, Bialas K, et al. Biothiols and oxidative stress markers and polymorphisms of TOMM40 and APOC1 genes in Alzheimer’s disease patients. Oncotarget. 2018;9:35207–25.
Li J, Chen S, Chen W, Ye Q, Dou Y, Xiao Y, et al. Role of the NF-kappaB family member RelB in regulation of Foxp3(+) regulatory T cells in vivo. J Immunol. 2018;200:1325–34.
Qiao G, Zhao Y, Li Z, Tang PQ, Langdon WY, Yang T, et al. T cell activation threshold regulated by E3 ubiquitin ligase Cbl-b determines fate of inducible regulatory T cells. J Immunol. 2013;191:632–9.
Liu YC. Ubiquitin ligases and the immune response. Annu Rev Immunol. 2004;22:81–127.
Stelzer G, Rosen N, Plaschkes I, Zimmerman S, Twik M, Fishilevich S, et al. The GeneCards suite: from gene data mining to disease genome sequence analyses. Curr Protoc Bioinformatics 2016;54:1.30.31–33.
Wang L, Man S, Bian Y. Bioinformatics analysis of biomarkers of aristolochic acid-induced early nephrotoxicity in embryonic stem cells. Exp Ther Med. 2021;21:508.
Zhang CY, He FF, Su H, Zhang C, Meng XF. Association between chronic kidney disease and Alzheimer’s disease: an update. Metab Brain Dis. 2020;35:883–94.
Jevtic S, Sengar AS, Salter MW, McLaurin J. The role of the immune system in Alzheimer disease: etiology and treatment. Ageing Res Rev. 2017;40:84–94.
Bangen KJ, Werhane ML, Weigand AJ, Edmonds EC, Delano-Wood L, Thomas KR, et al. Reduced regional cerebral blood flow relates to poorer cognition in older adults with type 2 diabetes. Front Aging Neurosci. 2018;10:270.
Kaltschmidt B, Uherek M, Volk B, Baeuerle PA, Kaltschmidt C. Transcription factor NF-kappaB is activated in primary neurons by amyloid beta peptides and in neurons surrounding early plaques from patients with Alzheimer disease. Proc Natl Acad Sci USA. 1997;94:2642–7.
Dong L, Li YZ, An HT, Wang YL, Chen SH, Qian YJ, et al. The E3 ubiquitin ligase c-Cbl inhibits microglia-mediated CNS inflammation by regulating PI3K/Akt/NF-kappaB pathway. CNS Neurosci Ther. 2016;22:661–9.
Nho K, Kim S, Horgusluoglu E, Risacher SL, Shen L, Kim D, et al. Association analysis of rare variants near the APOE region with CSF and neuroimaging biomarkers of Alzheimer’s disease. BMC Med Genomics. 2017;10:29.
Landrum MJ, Chitipiralla S, Brown GR, Chen C, Gu B, Hart J, et al. ClinVar: improvements to accessing data. Nucleic Acids Res. 2020;48:D835–D844.
Liu M, Pan Q, Xiao R, Yu Y, Lu W, Wang L. A cluster of metabolism-related genes predict prognosis and progression of clear cell renal cell carcinoma. Sci Rep. 2020;10:12949.
Kaymaz BT, Gunel NS, Ceyhan M, Cetintas VB, Ozel B, Yandim MK, et al. Revealing genome-wide mRNA and microRNA expression patterns in leukemic cells highlighted “hsa-miR-2278” as a tumor suppressor for regain of chemotherapeutic imatinib response due to targeting STAT5A. Tumour Biol. 2015;36:7915–27.
Fragiadaki M, Lannoy M, Themanns M, Maurer B, Leonhard WN, Peters DJ, et al. STAT5 drives abnormal proliferation in autosomal dominant polycystic kidney disease. Kidney Int. 2017;91:575–86.
Wu JJ, Weng SC, Liang CK, Lin CS, Lan TH, Lin SY, et al. Effects of kidney function, serum albumin and hemoglobin on dementia severity in the oldest old people with newly diagnosed Alzheimer’s disease in a residential aged care facility: a cross-sectional study. BMC Geriatr. 2020;20:391.
Acknowledgements
We thank the NCGG Biobank and the NCGG Integrated Database for Dementia Research (iDDR) for providing the study materials, clinical information, and technical support. This study was supported by grants from AMED (Grant Numbers JP18kk0205009 and JP18kk0205012 to SN, grant numbers JP21dk0207045 and JP22dk0207060 to SN and KO, grant numbers JP21de0107002 and JP21dk0207052 to SN and DS, and grant number JP21km0405501 to KO); Research Funding for Longevity Sciences from the NCGG (21-23 to KO and 21-24 to DS); JSPS KAKENHI (grant number JP21H02470 to DS); The Hori Science and Arts Foundation (to DS); The Chukyo Longevity Medical Research and Promotion Foundation (to DS); Japan Foundation For Aging and Health (to SN); and a grant from the Japanese Ministry of Health, Labor, and Welfare for Research on Dementia (to KO). We also thank ELSS editors for English corrections.
Author information
Authors and Affiliations
Contributions
DS developed the method and performed the analyses; SA, AY, and MF provided technical assistance; MS performed the qRT-PCR validation of gene expression; DS wrote the manuscript; and DS, KO, YN, and SN organized this work. All authors contributed to and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shigemizu, D., Akiyama, S., Suganuma, M. et al. Classification and deep-learning–based prediction of Alzheimer disease subtypes by using genomic data. Transl Psychiatry 13, 232 (2023). https://doi.org/10.1038/s41398-023-02531-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-023-02531-1
- Springer Nature Limited
This article is cited by
-
The HLA-DRB1*09:01-DQB1*03:03 haplotype is associated with the risk for late-onset Alzheimer’s disease in APOE \({{\varepsilon }}\)4–negative Japanese adults
npj Aging (2024)
-
Comprehensive Systematic Computation on Alzheimer's Disease Classification
Archives of Computational Methods in Engineering (2024)