Abstract
Background
Distant metastasis and recurrence remain the main obstacle to nasopharyngeal carcinoma (NPC) treatment. However, the molecular mechanisms underlying NPC growth and metastasis are poorly understood.
Methods
LHX2 expression was examined in NPC cell lines and NPC tissues using quantitative reverse transcription-polymerase chain reaction, western blotting and Immunohistochemistry assay. NPC cells overexpressing or silencing LHX2 were used to perform CCK-8 assay, colony-formation assay, EdU assay, wound-healing and invasion assays in vitro. Xenograft tumour models and lung metastasis models were involved for the in vivo assays. The Gene Set Enrichment Analysis (GSEA), ELISA assay, western blot, chromatin immunoprecipitation (ChIP) assay and Luciferase reporter assay were applied for the downstream target mechanism investigation.
Results
LIM-homeodomain transcription factor 2 (LHX2) was upregulated in NPC tissues and cell lines. Elevated LHX2 was closely associated with poor survival in NPC patients. Ectopic LHX2 overexpression dramatically promoted the growth, migration and invasion of NPC cells both in vitro and in vivo. Mechanistically, LHX2 transcriptionally increased the fibroblast growth factor 1 (FGF1) expression, which in turn activated the phosphorylation of STAT3 (signal transducer and activator of transcription 3), ERK1/2 (extracellular regulated protein kinases 1/2) and AKT signalling pathways in an autocrine and paracrine manner, thereby promoting the growth and metastasis of NPC. Inhibition of FGF1 with siRNA or FGFR inhibitor blocked LHX2-induced nasopharyngeal carcinoma cell growth, migration and invasion.
Conclusions
Our study identifies the LHX2-FGF1-FGFR axis plays a key role in NPC progression and provides a potential target for NPC therapy.
Similar content being viewed by others
Introduction
Nasopharyngeal carcinoma (NPC), a malignant tumour arising from the nasopharynx epithelium, has the highest incidence rate in Southeast Asia, especially in Southern China [1]. Most patients diagnosed with NPC are in advanced stages, and ~30% of them eventually develop distant metastasis [2]. Although, advances in intensity-modulated radiotherapy and a broader application of chemotherapy have improved the local and regional control of NPC but recurrence and distance metastasis eventually lead to treatment failure [3, 4]. Therefore, a better understanding of the molecular mechanisms that underly the recurrence and metastasis of NPC is essential in developing efficient therapeutic strategies.
Aberrant oncogenes expression universally exists in NPC and is a key feature in cancer initiation and progression [5]. However, the lack of reliable biomarkers for early detection and survival prediction of NPC limits NPC therapy. The development of high-throughput technologies and bioinformatics have identified candidate biomarkers that play important roles in NPC development, such as Serine Peptidase Inhibitor Kazal Type 6 (SPINK6), HOP Homeobox (HOPX) and Shisa Family Member 3 (SHISA3) [6,7,8], and transcription factors (TFs) that may offer therapeutic targets. The LIM-homeodomain transcription factor 2 (LHX2) was first identified as a critical transcription factor for B and T-lymphoid cell line differentiation. LHX2 has important roles in the development of the eye, bone, forebrain and olfactory sensory neurons [9,10,11,12]. Recent studies revealed LHX2 promotes tumour development and is highly expressed in a variety of human cancer types, including pilocytic astrocytoma, chronic myelogenous, leukaemia and pancreatic cancer [13,14,15]. However, the role of LHX2 in NPC is not fully understood.
Secreted proteins, such as transforming growth factor-β (TGFβ), Wnt family member 5A (Wnt5A) and epidermal growth factor (EGF), are dysregulated in the microenvironment of NPC and promote the growth, epithelial–mesenchymal transition (EMT) and metastasis of NPC in an autocrine or paracrine manner [16,17,18]. Fibroblast growth factor (FGF) signalling is fundamental for a variety of biological processes, including cell growth, tissue repair and tumour metastasis [19]. FGF1 is a member of the FGF family and stimulates downstream signalling cascades via binding and phosphorylating the FGF receptor (FGFR). Accumulating evidence indicates that FGF1 promotes tumour development by facilitating cell proliferation, migration, invasion and angiogenesis [20,21,22]. Aberrant expression of FGF1 promotes the development of various human cancers, such as breast cancer, bladder cancer and hepatocellular carcinoma [23,24,25,26,27]. However, the underlying role of FGF1 in NPC progression remains elusive.
In this study, we find LHX2 expression is upregulated in NPC and associated with poor survival. LHX2 promotes cellular growth, migration and invasion via promoting the FGF1 transcription, which further activates the STAT3, ERK and AKT signal pathways in an autocrine and paracrine mechanism.
Materials and methods
Cell culture and clinical specimens
Six human NPC cell lines (SUNE1, CNE1, CNE2, 5–8F, 6-10B and HONE1) were cultured in RPMI-1640 (Invitrogen, USA) supplemented with 10% foetal bovine serum (FBS; Gibco). The normal nasopharyngeal epithelial cell line, NP69, was cultured in keratinocyte/serum-free medium (Invitrogen) supplemented with bovine pituitary extract (BD Bioscience, San Diego, CA, USA). In all, 16 fresh frozen NPC biopsy samples and 6 normal nasopharyngeal epithelial tissues and 98 paraffin-embedded (FFPE) NPC tissues were collected from the Affiliated Cancer Hospital and Institute of Guangzhou Medical University. None of the patients had received anti-tumour treatment before biopsy collection. The study was approved by the Institutional Ethical Review Board of the Affiliated Cancer Hospital and Institute of Guangzhou Medical University, and informed consent was obtained from all patients.
RNA extraction, reverse transcription and real-time RT-PCR
Total RNA was extracted using TRIzol reagent (Invitrogen) and reverse-transcribed into cDNA using a Reverse Transcription Kit (Promega, Madison, WI). Quantitative PCR reactions were performed using SYBR Green Real-Time PCR Master Mix Kit (Invitrogen). The experiment was performed in triplicate and the relative expression was calculated with the 2−ΔΔCT method. GAPDH was applied as endogenous controls. Primer sequences are shown in Supplementary Table 2.
Western blot analysis
Protein was extracted with lysis buffer (Beyotime, Shanghai, China) and then quantified by BCA Protein Quantitation Kit (Beyotime, Shanghai, China). Equal amounts of protein (30 μg) were separated by SDS-polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to polyvinylidene fluoride (PVDF) membranes (Millipore, Billerica, MD) Later on. The membrane was blocked with 5% bovine serum albumin (BSA) and incubated with primary antibodies overnight at 4 °C. After incubation with secondary antibodies for 1 h at room temperature, the proteins were detected by enhanced chemiluminescence reagents. Primary antibodies against β-catenin (51067-2-AP), GAPDH (60004-1-Ig), and E-cadherin (20874-1-AP) were purchased from Proteintech Group. Antibodies against LHX2 (ab243030), Vimentin (ab8978), FGF1 (ab179455), ZEB1 (ab203829), FGFR1 (ab76464), FGFR2 (ab109372), FRS (ab183492) and TWIST1 (ab50581) were purchased from Abcam. Antibodies against STAT3 (#9139), Phospho-STAT3 (#9145), ERK (#4695), Phospho-ERK (#4370), AKT (#2920), Phospho-AKT (#4060), Phospho-GSK3 (#5558), Phospho-FGFR (#3471) and Phospho-FRS2 (#3861) were purchased from Cell Signalling Technology.
Immunohistochemistry assay (IHC)
Immunohistochemistry assay (IHC) was performed on paraffin-embedded sections of clinical NPC tissues and xenograft mice tissues. In brief, the tissues were deparaffinized with xylene and rehydrated in a graded ethanol series; the endogenous peroxidase activity was blocked with 3% (v/v) hydrogen peroxide; and the slides were next operated for antigen retrieval under the citrate-mediated high-temperature. The samples were washed once with PBS, and incubated with the primary antibodies at 4 °C overnight. The sections were washed with PBS, incubated with a peroxidase-conjugated secondary antibody, and stained with diaminobenzidine (DAB). The images were captured and scored by two experienced pathologists. The staining intensity was scored as follows: 0, no staining; 1, weak, light yellow; 2, moderate, yellow brown; and 3, strong, brown. Percentage of positive cells was evaluated as: 0, negative; 1, 1–25%; 2, 26–50%; 3, 51–75% and 4, 76–100%. The final staining score was calculated by multiplying the intensity and percentage scores. The antibodies for immunostaining assay were as follows: LHX2 (1:250, Abcam, ab243030, UK), FGF1(1:250, Abcam, ab179455, UK), Ki67 (1:300, Abcam, ab16667, UK), E-cadherin (1:300, Proteintech, 20874-1-AP, USA) and Vimentin (1:200, Abcam, ab8978, UK), Phospho-STAT3 (1:200, CST, #9145), Phospho-ERK (1:200, CST, #4370), Phospho-AKT (1:200, CST, #4060) according to the manufacturer’s instructions.
Lentiviral construction and transduction
The full length of LHX2 and FGF1 were synthesised and cloned into the lentiviral plasmid, pSin-EF2-puromycin (Addgene, Cambridge, MA, USA), and pSin-EF2-LHX2/FGF1-puromycin or the negative control pSin-EF2-puromycin vector was then co-transfected into 293T cells with the VSVG and PSPAX packaging plasmid (Addgene, Cambridge, MA, USA) using Lipofectamine 3000 Transfection Reagent (Invitrogen). The supernatant was obtained and used to infect HONE1 and SUNE1 cells. Stable clones were selected using 0.5 μg/ml puromycin.
Plasmid and small interfering RNA(siRNA) construction and transfection
The short hairpin RNA targeting LHX2 (sh-LHX2) (Supplementary Table 3) was synthesised and cloned into pLKO.1. siRNA targeting FGF1 (siFGF1) (Supplementary Table 3) was synthesised by RiboBio (Guangzhou, China). For LHX2 and FGF1 knockdown, HONE1 and SUNE1 cells were transfected with sh-LHX2 plasmids (4 μg) or siFGF1 (50 nM) and their corresponding control vectors with Lipofectamine 3000 Transfection Reagent (Invitrogen), according to the introduction, and harvested for assays 48 h after transfection.
CCK-8 assay, clone-formation assay and EdU assay
For the CCK-8 assay, 2 × 103 cells were placed into 96-well plates, incubated with CCK-8 reagent (Dojindo, Japan) and absorbance (450 nm) was measured every 24 h. For the colony-formation assay, 6 × 102 cells were seeded into six-well plates, with new medium every 3 days. After 10 days, colonies were then fixed in methanol, stained with 0.1% crystal violet, and the colonies in each group were counted and the relative colonies were normalised to the control group. Following the standard EdU (5-ethynyl-2’-deoxyuridine) Assay protocol, tumour cells were added to EdU (Invitrogen, Cat. A10044, USA) as recommended dose and imaged by fluorescence microscopy.
Wound-healing assay
Cells in six-well culture plates were cultured with serum-free medium for 12 h. A P200 pipette tip was used to scrape the cells in a straight line. The detached cells were gently washed with PBS twice and the remaining cells were cultured in serum-free medium. After incubation for 36 h, the width of the scratch was detected, and the migration rate in each group (migration rate = (original width of the wound width after cell migration)/original width of the wound).was quantified and normalised to the control group.
Migration and invasion assays
Cells were digested and suspended in serum-free medium, and 3 × 104 cells were placed into the upper 8-μm pore transwell chambers (Corning, NY, USA), which were coated with or without Matrigel (BD Biosciences, San Diego, CA, USA). Culture medium with 10% FBS was placed into the lower chamber to act as a chemoattractant. After incubation for 12 h or 24 h, cells were fixed in paraformaldehyde and stained with 0.1% crystal violet. Cells on the undersides of the filters were observed and counted under ×200 magnification. The relative migrated or invaded cells were normalised to the control group.
Conditioned medium collection
HONE1 and SUNE1 cells were transfected with an FGF1 overexpression plasmid and the corresponding empty vector. After transfection (48 h), the medium was replaced, and the cells were cultured for another 48 h. The supernatant was collected and centrifuged to remove cell debris. The conditioned medium was used for further functional studies, including CCK-8, colony formation, wound healing and transwell migration and invasion assays. All experiments were conducted in triplicate.
Immunofluorescence
Transfected cells cultured in 24-well plates were fixed and incubated with β-catenin primary antibodies (1:200, Proteintech, 17565-1-AP, Chicago, USA) overnight at 4 °C. After washing with PBS, fluorescent secondary antibodies (1:200; Proteintech, SA00013-2) were added to the cells and incubated at room temperature for 1 h. Images were captured after staining with DAPI solution.
Xenograft tumour models and metastasis models
All animal research procedures were performed according to the detailed rules of the Animal Care and Use Ethics Committee of Guangzhou Medical University. Five-week-old female nude mice were purchased from Guangdong Medical Laboratory Animal Center (Guangzhou, China). Mice were divided into four groups at random (five mice in each group), and 2 × 106 LHX2-overexpressed NPC cell lines and their control cell lines were injected into the dorsal flank of the mice. One week after injection, mice were treated with AZD4547 (12.5 mg/kg/d) or equal volume of vehicle every other day by oral gavage for 3 weeks. The volume of the tumour was detected every 3 days. The tumour weight and volume were monitored. Volume (mm3) = (L×W2)/2 (L: length, W: width). Animals were sacrificed 30 days later, and the tumours were excised for further examination.
For the tumour metastasis model, 1 × 106 LHX2-overexpressed or Vector NPC cell lines were injected into the tail vein of nude mice (five mice in each group) and treated with AZD4547 (12.5 mg/kg/d) or equal volume of vehicle every other day by oral gavage. Animals were sacrificed 30 days later, lungs were removed, and hematoxylin eosin (H&E) staining was conducted.
Luciferase reporter assay
The pGL3 luciferase reporter plasmids, containing wild-type or mutant promoter region of FGF1, were constructed. Transfected cells were co-transfected with the indicated luciferase reporters and Renilla luciferase reporter for 24 h. The cells were harvested, and the luciferase activity was detected using a Dual-Luciferase Reporter Assay Kit (Promega).
Chromatin immunoprecipitation assay
The chromatin immunoprecipitation (ChIP) assays were performed as previously described [7]. In brief, LHX2 overexpression or control cells were prepared for cross-linking and sheared to 200–500 bp by sonication. The chromatin fraction was immunoprecipitated with anti-LHX2 antibody (1:50; Abcam, UK) or IgG (1:50; Abcam, UK; negative control). Real-time PCR assays were conducted to detect the enrichment of the FGF1 promoter occupancy. The sequences of the ChIP primers are shown in Supplementary Table 1.
The gene set enrichment analysis
The Gene Set Enrichment Analysis (GSEA) software (version 2.0.13, ww.broadinstitute.org/gsea/) identified LHX2 expression-related pathways in GSE12452 and GSE53819 datasets (https://www.ncbi.nlm.nih.gov/geo/). In brief, the enrichment score for each gene set was calculated using the Kolmogorov–Smirnov statistic to identify the metastasis-related pathway and FGF associated with LHX2 expression.
Statistical analysis
Statistical analysis was performed with SPSS 22.0 software (SPSS, Chicago, USA). Statistical analysis of the data was performed using the Student’s t test or analysis of variance (ANOVA). Spearman correlation analysis was applied to analyse the relationship between LHX2 and FGF1 mRNA expression in the Gene Expression Omnibus (GEO) datasets and the TCGA datasets. All tests were two-tailed; P <0.05 were considered statistically significant.
Results
LHX2 is highly expressed in NPC and correlated with poor clinical outcome
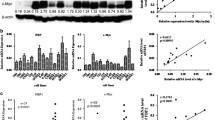
Genomic expression profiling of the TCGA database reveals LHX2 level is increased in many solid tumours (Supplementary Fig. S1A), and higher expression of LHX2 predicted poor survival in several solid tumours (Supplementary Fig. S1B), suggesting its potential oncogenic role in tumour development. To verify the expression of LHX2 in NPC, we analysed the expression level of LHX2 between NPC and normal tissues in three microarray-based high-throughput NPC datasets (GSE12452, GSE53819 and GSE64634). LHX2 mRNA expression was significantly upregulated in NPC clinical specimens (Fig. 1a). Next, we examined LHX2 expression in NPC cell lines and a normal nasopharyngeal epithelial cell line. Consistently, LHX2 protein level was significantly upregulated in NPC cell lines (Fig. 1b). Furthermore, western blot and IHC analysis showed LHX2 was dramatically upregulated in NPC tissues compared to normal tissues (Fig. 1c, d). Notably, LHX2 protein level was substantially higher in NPC tissues with regional lymph node and distant metastasis than in tissues without metastasis (Fig. 1e). Moreover, the LHX2 level was higher in NPC tissues of the T3/T4 stage than T1/T2 stage (Supplementary Fig. S2). By analysing the relationship between LHX2 expression and clinical data in a cohort of 98 paraffin-embedded NPC biopsy samples, we found a higher LHX2 level was significantly associated with T, N and TNM staging (P < 0.05, Table 1). Moreover, Kaplan–Meier survival analysis demonstrated that patients with higher LHX2 expression have significantly worse overall survival (P < 0.05, Fig. 1f). Taken together, these data suggest that LHX2 is upregulated in NPC and associated with poor survival.
a Box plots showing LHX2 mRNA expression level is higher in nasopharyngeal carcinoma tissues than in normal tissues in GSE12452 (left), GSE53819 (middle) and GSE64634 (right). b Western blot analysis shows LHX2 expression level is higher in NPC cell lines than in normal nasopharyngeal epithelial cell lines. Each experiment was independently repeated at least three times. c Western blot analysis shows LHX2 expression level is higher in nasopharyngeal carcinoma tissues than in normal tissues. Each experiment was independently repeated at least three times. d Immunohistochemical staining and (e) statistical analysis of LHX2 in normal tissues, primary NPC tissues with (LN + ) or without (LN-) lymph node metastasis and primary NPC tissues with distant metastasis. f Kaplan–Meier survival curves indicate high expression of LHX2 is associated with poor survival in NPC patients. Data shown as mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001.
LHX2 promotes NPC cells growth both in vitro and in vivo
To evaluate the effect of LHX2 on NPC cell growth, HONE1 and SUNE1 were transduced with lentivirus expressing LHX2 (Fig. 2a and Supplementary Fig. S3A). The CCK-8 and colony-formation assays demonstrated that NPC cells overexpressing LHX2 achieved significantly faster viability (Fig. 2b, c and Supplementary Fig. S3B). Similarly, 5-ethynyl-2’-deoxyuridine (EdU) assays also indicated LHX2 promotes the proliferation of HONE1 and SUNE1 cells (Supplementary Fig. S3C). In contrast, LHX2 inhibition decreased NPC cell proliferation (Fig. 2d–f and Supplementary Fig. S3D–F).
a Western blot analysis of LHX2 protein expression in HONE1 and SUNE1 cells stably overexpressing LHX2. Each experiment was independently repeated at least three times. b The CCK-8 assay and c the colony-formation assay of HONE1 and SUNE1 cells stably overexpressing LHX2. Each experiment was independently repeated at least three times. d Western blot analysis of LHX2 protein expression in HONE1 and SUNE1 cells transfected with LHX2 shRNAs or control cells. Each experiment was independently repeated at least three times. e The CCK-8 assay and f the colony-formation assay of HONE1 and SUNE1 cells transfected with LHX2 shRNAs or control. Each experiment was independently repeated at least three times. g, h Images of Xenograft tumours of nude mice 30 days after injecting with HONE1 and SUNE1 cells stably overexpressing LHX2 or vector (n = 5 in each group). i Tumour volume growth curves. j Average xenograft tumour weights. k Immunohistochemistry assay of LHX2 and Ki67 protein expression in xenograft tumours. Data shown as mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001.
To further explore the roles of LHX2 on NPC growth in vivo, xenograft tumour models were established. Tumour growth promotion was observed in LHX2-overexpressing cells compared to control cells (Fig. 2g–i). In parallel, the average weight of tumours was significantly higher in LHX2-overexpressed tumours than in control tumours (Fig. 2j). IHC analysis of xenograft tissues showed an increased expression of cell proliferation marker Ki67 in LHX2-overexpressing groups compared with control groups (Fig. 2k). Collectively, these findings indicate that LHX2 promotes NPC cell growth both in vitro and in vivo.
LHX2 promotes NPC cells migration, invasion and EMT
GSEA based on the GSE12452 and GSE53819 databases revealed that LHX2 was associated with NPC metastasis (Fig. 3a). Wound-healing and transwell assays indicated overexpression of LHX2 significantly promoted the migration and invasion of NPC cells (Fig. 3b, c and Supplementary Fig. S4A, B). However, inhibition of LHX2 suppressed the migratory and invasive abilities of NPC cells (Fig. 3d, e and Supplementary Fig. S4C, D). Moreover, we explored the role of LHX2 in tumour metastasis in vivo via a lung metastasis model. The results showed ectopic LHX2 overexpression significantly increased the number of metastatic nodules, as confirmed by stained lung sections (Fig. 3f–h).
a GSEA enrichment plots demonstrated that metastasis was associated with upregulation of LHX2 in GSE12452 (left and middle) and GSE53819 (right). b Wound healing and (c) transwell migration and invasion assays in HONE1 cells stably overexpressing LHX2 or empty vector. Each experiment was independently repeated at least three times. d Wound healing and (e) transwell migration and invasion assays in HONE1 cells transfected with LHX2 shRNAs or control. Each experiment was independently repeated at least three times. f–h Representative images of metastatic lungs (f), representative HE images (g) and numbers of metastatic foci per lung (h) injected with HONE1 and SUNE1 cells stably overexpressing LHX2 or empty vector (n = 5 in each group). i Western blot analysis of EMT-related protein expression in HONE1 and SUNE1 cells stably overexpressing LHX2 or empty vector. Each experiment was independently repeated at least three times. j IHC analysis of E-cadherin and vimentin protein expression in xenograft tumours. Data shown as mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001.
EMT is a crucial process for tumour metastasis. We examined whether LHX2 was associated with EMT in NPC. Western blot analysis showed that LHX2 overexpression decreased the expression of E-cadherin and increased the expression of vimentin, Zinc Finger E-Box Binding Homeobox 1 (ZEB1) and Twist Family BHLH Transcription Factor 1 (TWEST1) (Fig. 3i). In addition, a decreased level of E-cadherin and an increased level of Vimentin were determined in LHX2-overexpressing tumours (Fig. 3j). These results reveal that LHX2 promotes cell migration, invasion, metastasis and EMT in NPC cells.
LHX2 activates pro-survival and β-catenin signalling pathways through transcriptionally regulating FGF1 expression
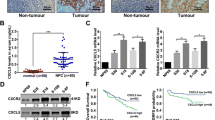
Considering LHX2 is a transcription factor, we referred to the accessible online TF ChIP-seq database (http://dc2.cistrome.org/) to explore its potential downstream targets. We found FGF1 was a putative target of LHX2 that LHX2 motif (Fig. 4a) could bind to its promoter. Consistently, overexpression of LHX2 increased FGF1 expression (protein and mRNA) while inhibition of LHX2 could reduce FGF1 expression (Fig. 4b, c and Supplementary Fig. S5A, B). Furthermore, ELIAS assay showed overexpression of LHX2 increased FGF1 expression in the supernatant, while the reverse was observed in LHX2-silenced NPC cells (Fig. 4d, e). FGF1 also has higher mRNA expression in NPC clinical specimens (Supplementary Fig. S5C). To confirm the LHX2-binding sites on FGF1, deletions and selective mutations were generated into the FGF1 promoter sequences. We identified that the FGF1 promoter binding site 2 was the putative LHX2-binding site (Fig. 4f), and ChIP real-time PCR analysis confirmed the affinity of LHX2 to the FGF1 promoter (Fig. 4g).
a LHX2 motif. b, c Western blot analysis of FGF1 expression in HONE1 and SUNE1 cells with LHX2 overexpression (b) or knockdown (c). Each experiment was independently repeated at least three times. d, e ELISA assay of FGF1 expression in the supernatant derived from HONE1 and SUNE1 cells with LHX2 overexpression (d) or knockdown (e). Each experiment was independently repeated at least three times. f Luciferase activity of the reporter gene driven by serially truncated/mutated FGF1 promoters. Each experiment was independently repeated at least three times. g ChIP assay confirms the direct binding of LHX2 to the FGF1 promoters in HONE1 and SUNE1 cells. Each experiment was independently repeated at least three times. h Western blot assay reveals the p-STAT3, pERK1/2 and p-AKT were activated by LHX2. Each experiment was independently repeated at least three times. i GSEA enrichment plots demonstrates that β-catenin signalling is associated with upregulation of LHX2 in GSE53819. j Western blot assay of β-catenin and Ser9-GSK3β in HONE1 and SUNE1 cells stably overexpressing LHX2 or containing the empty vector. Each experiment was independently repeated at least three times. k RT-qPCR analysis of β-catenin response genes in HONE1 and SUNE1 cells stably overexpressing LHX2 or containing the empty vector. Each experiment was independently repeated at least three times. l Representative images of immunofluorescent staining of β-catenin in HONE1 and SUNE1 cells stably overexpressing LHX2 or containing the empty vector. Each experiment was independently repeated at least three times. Data shown as mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001.
Accumulating evidence has revealed that FGF1 can bind to FGFR and activate the STAT3, AKT and ERK signalling pathways [28]. Therefore, we detected these signalling pathways and found LHX2 overexpression activated p-STAT3, p-ERK and p-AKT (Fig. 4h). In light of the activation of p-AKT by LHX2, we hypothesised that LHX2 may facilitate β-catenin stability via downregulating glycogen synthase kinase-3β (GSK3β) activity by phosphorylation at Ser9 of GSK3β. GSEA demonstrated that LHX2 was associated with the β-catenin signalling pathway (Fig. 4i). LHX2 overexpression increased nuclear levels of β-catenin and total expression levels of β-catenin and p-Ser9-GSK3β in both HONE1 and SUNE1 cells (Fig. 4j). Moreover, the expression levels of β-catenin-targeted genes were substantially increased in LHX2-overexpressed NPC cells, including ZEB1 and TWST1 (Fig. 4k). Immunofluorescence analysis showed that LHX2 overexpression induced the accumulation of nuclear β-catenin (Fig. 4l). Importantly, FGF1 expression was positively associated with ZEB1 and TWIST1 expression in pan-cancers (Supplementary Figs. S6 and S7). These findings suggest that LHX2 activates pro-survival, β-catenin signalling pathways via transcriptionally regulating FGF1 expression in NPC.
FGF1 mediates LHX2-induced proliferation, migration and invasion in an autocrine or paracrine manner
To explore whether FGF1 contributes to LHX2-induced proliferation, migration and invasion in NPC cells. We inhibited FGF1 expression in NPC cells overexpressing LHX2 with siRNA. We found FGF1 inhibition restrained the proliferative, migratory and invasive abilities of NPC cells induced by ectopic expression of LHX2 (Fig. 5a–c and Supplementary Fig. S8A–C). Furthermore, inhibition of FGF1 significantly blocked the activation of ERK1/2, AKT, STAT3 and β-catenin signalling pathways stimulated by LHX2 overexpression (Fig. 5d, e). As cancer cells secrete FGFs, we transfected NPC cells with an FGF1 overexpression plasmid or empty vector to determine whether FGF1 promotes NPC progression in an autocrine/paracrine secretion manner. The conditioned medium (CM) derived from transfected HONE1 and SUNE1 cells was centrifuged and added to the indicated NPC cells (Supplementary Fig. S8D). Notably, FGF1-CM activated STAT3, ERK and AKT signalling as shown by the consistently elevated expression levels of p-STAT3, pERK1/2 and p-AKT when LHX2 was overexpressed (Fig. 5f). Moreover, Ser9-GSK3β/β-catenin signalling and EMT-related proteins were also stimulated by FGF1-CM (Fig. 5g). These observations reveal that FGF1 serves as a functional target of LHX2 in NPC cells and inhibition of FGF1 could block the LHX2-induced proliferation, migration and invasion of NPC cells.
a–e FGF1 siRNA was transfected into HONE1 cells stably overexpressing LHX2 or the empty vector. Colony-formation assay (a), wound-healing assay (b) and transwell migration and invasion assays (c). Western blot analysis of LHX2, FGF1 and pro-survival signalling pathways (d). Western blot analysis of β-catenin signalling pathway and EMT-related proteins (e). Each experiment was independently repeated at least three times. f, g HONE1 and SUNE1 cells were treated with FGF1-CM or EV-CM for the indicated time. Western blot analysis of pro-survival signalling pathways (f). Western blot analysis of β-catenin signalling pathway and EMT-related proteins (g). Each experiment was independently repeated at least three times. Data shown as mean ± SD. *P < 0.05, **P < 0.01. FGF1-CM FGF1-condition medium, EV empty vector.
Blocking FGF1/FGFR signalling suppresses the growth and metastasis of NPC cells
FGF1 commonly stimulates downstream signalling cascades by phosphorylating the FGFR [23]. To assess the impact exerted by inhibiting the FGF/FGFR axis on NPC cell proliferation, migration and invasion, NPC cells were treated with AZD4547, an FGFR-specific inhibitor. We observed extracellular recombinant FGF1 caused the activation of FGFR, and thus stimulated the cascade of phosphorylation of downstream signalling pathways, which could be abrogated by AZD4547 (Supplementary Fig. S9A, B). Consistently, administration of FGF1 dramatically increased the cell proliferation as well as migration and invasion of HONE1 cells, which could be attenuated by AZD4547 (Supplementary Fig. S9C–F). To get further insights into the impact of FGF/FGFR signalling blockage on LHX2-induced proliferation, migration and invasion in NPC, we administrated NPC cells with AZD4547. The inhibition of cellular FGFR phosphorylation was observed in a dose-dependent manner by AZD4547 (Fig. 6a and Supplementary Fig. S10A). In addition, the phosphorylation of FRS2, downstream marker of FGFR signalling, LHX2-mediated induction of STAT3, ERK, AKT and β-catenin signalling as well as EMT were also significantly abrogated by AZD4547 (Fig. 6b, c). Consistently, administration of AZD4547 dramatically blocked the positive effect of LHX2 on HONE1 cell proliferation, migration and invasion in vitro (Fig. 6d–f and Supplementary Fig. S10B). Furthermore, AZD4547 treatment significantly revised the accelerated tumour growth and metastasis by LHX2 overexpression in vivo (Fig. 6g–m). Together, these findings revealed an oncogenic role of the LHX2-induced FGF/FGFR signalling pathway in NPC tumorigenesis and the FGF/FGFR trapping by AZD4547 could reverse these effects.
a HONE1 cells were incubated for 4 h of AZD4547 at different concentrations and then lysed and immunoblotted for the indicated proteins. b, c HONE1 cells stably overexpressing LHX2 or the empty vector were treated with AZD4547 (100 nM) or DMSO. Whole-cell lysates were subjected to western blot analysis. Each experiment was independently repeated at least three times. d–f HONE1 cells stably overexpressing LHX2 or the empty vector were treated with AZD4547 (100 nM) or DMSO for 48 h. The cells were harvested and subjected to (d) colony-formation assay, (e) wound-healing assay and (f) transwell migration and invasion assays. Each experiment was independently repeated at least three times. g–j LHX2-overexpressed or vector HONE1 cells (2 × 106) were injected into the dorsal flank of mice. One week after injection, the mice were treated with AZD4547 (12.5 mg/kg/d) or vehicle orally for 3 weeks. Images of Xenograft tumours (g). Tumour volume growth curves (h). The volume of xenograft tumours (i). Average xenograft tumour weights (j). LHX2-overexpressed or vector HONE1 cells (1 × 106) were injected into the tail vein of nude mice and the mice were treated with AZD4547 (12.5 mg/kg/d) or vehicle orally for 3 weeks. k–m Representative images of metastatic lungs (k), representative HE images (l) and numbers of metastatic foci per lung (m) (n = 5 in each group). Data are shown as means ± SD. *P < 0.05, **P < 0.01, ***P < 0.001.
The expression of LHX2 and FGF1 are positively correlated in NPC tissues
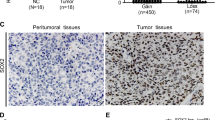
To extend our findings in human NPC tissues, we investigated the relationship between LHX2 and FGF1 expression in NPC tissues. Pearson correlation analysis showed that FGF1 and LHX2 mRNA expression were positively correlated in NPC clinical specimens (Fig. 7a). Consistently, TCGA datasets from other types of cancers further verified the positive correlation between FGF1 and LHX2 expression (Supplementary Fig. S11). Moreover, a positive correlation between FGF1 and LHX2 protein expression was confirmed in NPC tissue (Fig. 7b). In addition, high expression of FGF1, p-STAT3, p-ERK and p-AKT were observed in LHX2-overexpressing xenografts, as shown by IHC analysis (Fig. 7c).
Discussion
This study identified LHX2 as a novel transcription factor related to NPC progression. LHX2 transcriptionally regulated FGF1 expression and promoted the growth and metastasis of NPC in an FGF1/FGFR-dependent manner. Furthermore, FGF/FGFR trapping by AZD4547 could block the LHX2/FGF1-induced promotion of tumour growth and metastasis. Our study suggests LHX2 as a novel biomarker for NPC diagnosis and prognosis evaluation, as well as targets for therapeutic treatment (Fig. 8).
LHX2 transcriptionally activates FGF1 expression. Tumour cells produce FGF1, which binds to FGFR, and the subsequent downstream signalling occurs through the intracellular receptor substrates FGFR substrate 2 (FRS2), resulting in the phosphorylation and activation of MAPK/ERK, JAK2/STAT3, and PI3K/AKT signalling pathways. (i) These pro-survival pathways are responsible for the promotion of tumour proliferation and growth. (ii) Phosphorylated AKT stimulates the Ser9-GSK3β/β-catenin signalling, leading to the EMT of NPC via the β-catenin targeted ZEB1 and TWIST1 genes and promotes tumour cell migration and invasion. (iii) FGF/FGFR trapping by AZD4547 could block the LHX2/FGF1-induced promotion on tumour growth and metastasis.
A precise method for measuring the risk of recurrence or distant metastasis in NPC patients is needed in the clinic. By applying bioinformatic analysis of the TCGA and GEO databases, we found that LHX2 expression was dramatically increased in some solid tumours, including NPC. The carcinogenesis of LHX2 has been confirmed in osteosarcoma and prostate cancer [29, 30]. However, the role and mechanism of LHX2 in NPC were largely unknown. In this study, we verified that LHX2 was upregulated in NPC and high expression of LHX2 in NPC patients had worse clinical outcomes. These results suggest LHX2 is a promising prognostic biomarker for NPC detection and therapy.
FGFs activate MAPK-ERK, JAK2/STAT3 and PI3K/AKT signalling pathways, which has been identified as key mediators in tumour progression promotion [31, 32]. FGF1 was reported to promote the proliferation and invasion of LCSCs dependent on the MAPK/ERK signalling pathway [21] SH2B1β plays an important role in neurite outgrowth via enhancing and prolonging FGF1-induced MEK-ERK1/2 and PI3K-AKT pathways [33]. FGFR activation induces the accumulation of hyaluronan (HA) and increases the proliferation, migration and therapeutic resistance of breast cancer cells. Importantly, FGFR-mediated HA accumulation requires downstream activation of the STAT3 signalling pathway [34]. Consistent with these observations, we found FGF1 promote the growth of NPC cells primarily via activating phosphorylation of STAT3, ERK1/2 and AKT, which could be blocked by siFGF1 or FGFR inhibitor.
EMT is a physiological phenomenon, during this process, epithelial cells possess the characteristics of mesenchymal cells, including enhanced motility and invasive ability [35]. As the key initial step in the EMT, the downregulation of E-cadherin expression was observed in metastatic tumours, the transcriptional level of E-cadherin is repressed by several factors including, ZEB1 and Twist1, which have been identified as the effectors of β-catenin/TCF4 signalling in tumour invasiveness [36,37,38]. LHX2 has been reported to play an important role in epithelial–mesenchymal interactions, as identified in hair, liver and brain [10, 39, 40]. In this study, we demonstrated that LHX2 promotes EMT of NPC cells via transcriptionally activating FGF1, upregulating ZEB1 and TWIST1 through the β-catenin signalling pathway. In the absence of Wnt signal, APC/Axin/GSK3β could target β-catenin leading to its ubiquitination and proteasomal degradation [41]. The activation of GSK3β requires phosphorylation at Tyr216, but many signalling pathways are reported to downregulate GSK3β activity through phosphorylation at Ser9 [42, 43]. We found FGF1 binds to the FGFR, resulting in the phosphorylation of Akt, which in turn phosphorylates GSK3β at Ser9, rendering the kinase-inactive and resulting in decreased phosphorylation of downstream substrates, including β-catenin.
Although Tian-Song Liang has found that MicroRNA-506 can inhibit NPC tumour growth and metastasis by downregulating LHX2 [44], our study further extended the downstream targets and signalling pathway by which LHX2 serve as the oncogene in NPC, providing potential therapeutic strategies, such as FGF1/FGFR inhibitors. Treatment of FGF1 siRNA or FGFR inhibitor, efficiently decreased the growth and metastasis in LHX2-overexpressing NPC cells. The relevance of autocrine/paracrine FGF signalling has been shown in the development of various tumours, including multiple myeloma [45], breast cancer [46] and uveal melanoma [47], suggesting a promising target for tumour therapy. NSC12, a TX3-derived small molecule, is able to suppress FGFR activation and tumour growth in various FGF-dependent cancer models [48, 49]. In a Phase II study, erdafitinib, a tyrosine kinase inhibitor of FGFR1–4, exhibits antitumor activity in advanced urothelial carcinoma with FGFR alterations [50]. AZD4547 is a novel selective inhibitor of the FGFR1/2 tyrosine kinases. Administration of AZD4547 resulted in potent dose-dependent antitumor activity in preclinical models and has reached Phase II clinical investigations in malignancies with FGFR alterations, including lung cancer, endometrial cancer, multiple myeloma and breast cancer [51, 52]. Although AZD4547 did not meet the primary endpoint of the study, it did show a modest benefit in patients with FGFR mutations and fusions. Moreover, different approaches to targeting the FGF/FGFR system have been verified [47] and various selective TKI FGFR inhibitors have reached clinical trials on different kinds of tumours [53]. These studies highlight the potential of FGFR inhibitors as cancer therapeutics.
In summary, this study demonstrated that LHX2 is frequently upregulated in NPC and promotes NPC cell growth and metastasis both in vitro and in vivo. Furthermore, the role of LHX2 in NPC may depend on its transcriptional promotion of FGF1 and activated FGF1/FGFR signalling. Overall, our findings provide a new insight into the potential mechanism by which LHX2 regulates FGF1 expression in NPC progression.
Data availability
The obtained results of the research are available on reasonable request.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Lee AW, Ng WT, Chan LL, Hung WM, Chan CC, Sze HC, et al. Evolution of treatment for nasopharyngeal cancer-success and setback in the intensity-modulated radiotherapy era. Radiother Oncol. 2014;110:377–84.
Sun X, Su S, Chen C, Han F, Zhao C, Xiao W, et al. Long-term outcomes of intensity-modulated radiotherapy for 868 patients with nasopharyngeal carcinoma: an analysis of survival and treatment toxicities. Radiother Oncol. 2014;110:398–403.
Li Y, Tang LQ, Liu LT, Guo SS, Liang YJ, Sun XS, et al. Induction chemotherapy plus concurrent chemoradiotherapy versus concurrent chemoradiotherapy alone in locoregionally advanced nasopharyngeal carcinoma in children and adolescents: a matched cohort analysis. Cancer Res Treat. 2018;50:1304–15.
Peter IS, Davidson EH. Evolution of gene regulatory networks controlling body plan development. Cell. 2011;144:970–85.
Zheng LS, Yang JP, Cao Y, Peng LX, Sun R, Xie P, et al. SPINK6 promotes metastasis of nasopharyngeal carcinoma via binding and activation of epithelial growth factor receptor. Cancer Res. 2017;77:579–89.
Ren X, Yang X, Cheng B, Chen X, Zhang T, He Q, et al. HOPX hypermethylation promotes metastasis via activating SNAIL transcription in nasopharyngeal carcinoma. Nat Commun. 2017;8:14053.
Zhang J, Li YQ, Guo R, Wang YQ, Zhang PP, Tang XR, et al. Hypermethylation of SHISA3 promotes nasopharyngeal carcinoma metastasis by reducing SGSM1 stability. Cancer Res. 2019;79:747–59.
Tetreault N, Champagne MP, Bernier G. The LIM homeobox transcription factor Lhx2 is required to specify the retina field and synergistically cooperates with Pax6 for Six6 trans-activation. Dev Biol. 2009;327:541–50.
Tornqvist G, Sandberg A, Hagglund AC, Carlsson L. Cyclic expression of lhx2 regulates hair formation. PLoS Genet. 2010;6:e1000904.
Hirota J, Mombaerts P. The LIM-homeodomain protein Lhx2 is required for complete development of mouse olfactory sensory neurons. Proc Natl Acad Sci USA. 2004;101:8751–5.
Kim JH, Youn BU, Kim K, Moon JB, Lee J, Nam KI, et al. Lhx2 regulates bone remodeling in mice by modulating RANKL signaling in osteoclasts. Cell Death Differ. 2014;21:1613–21.
Wu HK, Heng HH, Siderovski DP, Dong WF, Okuno Y, Shi XMK, et al. Identification of a human LIM-Hox gene, hLH-2, aberrantly expressed in chronic myelogenous leukaemia and located on 9q33-34.1. Oncogene. 1996;12:1205–12.
Shi X, Zhan L, Xiao C, Lei Z, Yang H, Wang L, et al. miR-1238 inhibits cell proliferation by targeting LHX2 in non-small cell lung cancer. Oncotarget. 2015;6:19043–54.
Gorantla B, Asuthkar S, Rao JS, Patel J, Gondi CS. Suppression of the uPAR-uPA system retards angiogenesis, invasion, and in vivo tumor development in pancreatic cancer cells. Mol Cancer Res. 2011;9:377–89.
Li J, Wang W, Chen S, Cai J, Ban Y, Peng Q, et al. FOXA1 reprograms the TGF-beta-stimulated transcriptional program from a metastasis promoter to a tumor suppressor in nasopharyngeal carcinoma. Cancer Lett. 2019;442:1–14.
Yip WK, Seow HF. Activation of phosphatidylinositol 3-kinase/Akt signaling by EGF downregulates membranous E-cadherin and beta-catenin and enhances invasion in nasopharyngeal carcinoma cells. Cancer Lett. 2012;318:162–72.
Zeng ZY, Zhou YH, Zhang WL, Xiong W, Fan SQ, Li XL, et al. Gene expression profiling of nasopharyngeal carcinoma reveals the abnormally regulated Wnt signaling pathway. Hum Pathol. 2007;38:120–33.
Zhou Y, Wu C, Lu G, Hu Z, Chen Q, Du X. FGF/FGFR signaling pathway involved resistance in various cancer types. J Cancer. 2020;11:2000–7.
Bouleau S, Grimal H, Rincheval V, Godefroy N, Mignotte B, Vayssiere JL, et al. FGF1 inhibits p53-dependent apoptosis and cell cycle arrest via an intracrine pathway. Oncogene. 2005;24:7839–49.
Liu P, Zhang R, Yu W, Ye Y, Cheng Y, Han L, et al. FGF1 and IGF1-conditioned 3D culture system promoted the amplification and cancer stemness of lung cancer cells. Biomaterials. 2017;149:63–76.
Ornitz DM, Itoh N. Fibroblast growth factors. Genome Biol. 2001;2:REVIEWS3005.
Zhang L, Kharbanda S, McLeskey SW, Kern FG. Overexpression of fibroblast growth factor 1 in MCF-7 breast cancer cells facilitates tumor cell dissemination but does not support the development of macrometastases in the lungs or lymph nodes. Cancer Res. 1999;59:5023–9.
Kin M, Sata M, Ueno T, Torimura T, Inuzuka S, Tsuji R, et al. Basic fibroblast growth factor regulates proliferation and motility of human hepatoma cells by an autocrine mechanism. J Hepatol. 1997;27:677–87.
Tomlinson DC, L’Hote CG, Kennedy W, Pitt E, Knowles MA. Alternative splicing of fibroblast growth factor receptor 3 produces a secreted isoform that inhibits fibroblast growth factor-induced proliferation and is repressed in urothelial carcinoma cell lines. Cancer Res. 2005;65:10441–9.
King ML, Lindberg ME, Stodden GR, Okuda H, Ebers SD, Johnson A, et al. WNT7A/beta-catenin signaling induces FGF1 and influences sensitivity to niclosamide in ovarian cancer. Oncogene. 2015;34:3452–62.
El-Hariry I, Pignatelli M, Lemoine NR. FGF-1 and FGF-2 regulate the expression of E-cadherin and catenins in pancreatic adenocarcinoma. Int J Cancer. 2001;94:652–61.
Chandarlapaty S. Negative feedback and adaptive resistance to the targeted therapy of cancer. Cancer Discov. 2012;2:311–9.
Song H, Liu J, Wu X, Zhou Y, Chen X, Chen J, et al. LHX2 promotes malignancy and inhibits autophagy via mTOR in osteosarcoma and is negatively regulated by miR-129-5p. Aging. 2019;11:9794–810.
Shan G, Shao B, Liu Q, Zeng Y, Fu C, Chen A, et al. circFMN2 sponges miR-1238 to promote the expression of LIM-homeobox gene 2 in prostate cancer cells. Mol Ther Nucleic Acids. 2020;21:133–46.
Turner N, Grose R. Fibroblast growth factor signalling: from development to cancer. Nat Rev Cancer. 2010;10:116–29.
Lamothe B, et al. The docking protein Gab1 is an essential component of an indirect mechanism for fibroblast growth factor stimulation of the phosphatidylinositol 3-kinase/Akt antiapoptotic pathway. Mol Cell Biol. 2004;24:5657–66.
Lamothe B, Yamada M, Schaeper U, Birchmeier W, Lax I, Schlessinger J. SH2B1beta enhances fibroblast growth factor 1 (FGF1)-induced neurite outgrowth through MEK-ERK1/2-STAT3-Egr1 pathway. Cell Signal. 2009;21:1060–72.
Bohrer LR, Chuntova P, Bade LK, Beadnell TC, Leon RP, Brady NJ, et al. Activation of the FGFR-STAT3 pathway in breast cancer cells induces a hyaluronan-rich microenvironment that licenses tumor formation. Cancer Res. 2014;74:374–86.
Pastushenko I, Blanpain C. EMT transition states during tumor progression and metastasis. Trends Cell Biol. 2019;29:212–26.
Polyak K, Weinberg RA. Transitions between epithelial and mesenchymal states: acquisition of malignant and stem cell traits. Nat Rev Cancer. 2009;9:265–73.
Chang YW, Su YJ, Hsiao M, Wei KC, Lin WH, Liang CL, et al. Diverse targets of beta-catenin during the epithelial-mesenchymal transition define cancer stem cells and predict disease relapse. Cancer Res. 2015;75:3398–410.
Sanchez-Tillo E, de Barrios O, Siles L, Cuatrecasas M, Castells A, Postigo A. beta-catenin/TCF4 complex induces the epithelial-to-mesenchymal transition (EMT)-activator ZEB1 to regulate tumor invasiveness. Proc Natl Acad Sci USA. 2011;108:19204–9.
Pinto do OP, Richter K, Carlsson L. Hematopoietic progenitor/stem cells immortalized by Lhx2 generate functional hematopoietic cells in vivo. Blood. 2002;99:3939–46.
Porter FD, Drago J, Xu Y, Cheema SS, Wassif C, Huang SP, et al. Lhx2, a LIM homeobox gene, is required for eye, forebrain, and definitive erythrocyte development. Development. 1997;124:2935–44.
MacDonald BT, Tamai K, He X. Wnt/beta-catenin signaling: components, mechanisms, and diseases. Dev Cell. 2009;17:9–26.
Jensen J, Brennesvik EO, Lai YC, Shepherd PR. GSK-3beta regulation in skeletal muscles by adrenaline and insulin: evidence that PKA and PKB regulate different pools of GSK-3. Cell Signal. 2007;19:204–10.
Sutherland C, Leighton IA, Cohen P. Inactivation of glycogen synthase kinase-3 beta by phosphorylation: new kinase connections in insulin and growth-factor signalling. Biochem J. 1993;296:15–19.
Liang TS, Zheng YJ, Wang J, Zhao JY, Yang DK, Liu ZS. MicroRNA-506 inhibits tumor growth and metastasis in nasopharyngeal carcinoma through the inactivation of the Wnt/beta-catenin signaling pathway by down-regulating LHX2. J Exp Clin Cancer Res. 2019;38:97.
Ronca R, Ghedini GC, Maccarinelli F, Sacco A, Locatelli SL, Foglio E, et al. FGF trapping inhibits multiple myeloma growth through c-Myc degradation-induced mitochondrial oxidative stress. Cancer Res. 2020;80:2340–54.
Santolla MF, Maggiolini M. The FGF/FGFR system in breast cancer: oncogenic features and therapeutic perspectives. Cancers. 2020;12:3029.
Rezzola S, Ronca R, Loda A, Nawaz MI, Tobia C, Paganini G, et al. The autocrine FGF/FGFR system in both skin and uveal melanoma: FGF trapping as a possible therapeutic approach. Cancers. 2019;11:1035.
Castelli R, Giacomini A, Anselmi M, Bozza N, Vacondio F, Rivara S, et al. Synthesis, structural elucidation, and biological evaluation of NSC12, an orally available fibroblast growth factor (FGF) ligand trap for the treatment of FGF-dependent lung tumors. J Med Chem. 2016;59:4651–63.
Ronca R, Giacomini A, Di Salle E, Coltrini D, Pagano K, Ragona L, et al. Long-pentraxin 3 derivative as a small-molecule FGF trap for cancer therapy. Cancer Cell. 2015;28:225–39.
Loriot Y, Necchi A, Park SH, Garcia-Donas J, Huddart R, Burgess E, et al. Erdafitinib in locally advanced or metastatic urothelial carcinoma. N. Engl J Med. 2019;381:338–48.
Gavine PR, Mooney L, Kilgour E, Thomas AP, Al-Kadhimi K, Beck S, et al. AZD4547: an orally bioavailable, potent, and selective inhibitor of the fibroblast growth factor receptor tyrosine kinase family. Cancer Res. 2012;72:2045–56.
Chae YK, Hong F, Vaklavas C, Cheng HH, Hammerman P, Mitchell EP, et al. Phase II study of AZD4547 in patients with tumors harboring aberrations in the FGFR pathway: results from the NCI-MATCH Trial (EAY131) Subprotocol W. J Clin Oncol. 2020;38:2407–17.
Botrus G, Raman P, Oliver T, Bekaii-Saab T. Infigratinib (BGJ398): an investigational agent for the treatment of FGFR-altered intrahepatic cholangiocarcinoma. Expert Opin Investig Drugs. 2021;30:309–16.
Acknowledgements
We thank Jun Ma (Department of Radiation Oncology, State Key Laboratory of Oncology in Southern China, Collaborative Innovation Center for Cancer Medicine, Guangdong Key Laboratory of Nasopharyngeal Carcinoma Diagnosis and Therapy, Sun Yat-sen University Cancer Center) for his helpful discussion and his laboratory for kindly providing the NPC cell lines.
Funding
This work was supported by the National Natural Science Foundation of China (grant numbers: 81773354), the National Natural Science Foundation of China (grant numbers: 81872195), the National Natural Science Foundation of China (grant numbers: 82003212) and Guangzhou Key Medical Discipline Construction Project Fund.
Author information
Authors and Affiliations
Contributions
TX performed the majority of the experiments; JL provided clinical samples; WL, KD and TX performed the in vivo experiments; CL, BW, YT, RL, HJ and XH provided technical support; YY, BW and JZ contributed to study concept and design; TX drafted the manuscript; BW and YY contributed to study supervision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study was performed in accordance with the ethical standards of the Declaration of Helsinki and according to national and international guidelines. Our study has been approved by the Institutional Ethical Review Board of the Affiliated Cancer Hospital and Institute of Guangzhou Medical University.
Consent to publish
None.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Xie, T., Du, K., Liu, W. et al. LHX2 facilitates the progression of nasopharyngeal carcinoma via activation of the FGF1/FGFR axis. Br J Cancer 127, 1239–1253 (2022). https://doi.org/10.1038/s41416-022-01902-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-022-01902-7
- Springer Nature Limited