Abstract
Background
Despite a clear link between aging and cancer, there has been inconclusive evidence on how biological age (BA) may be associated with cancer incidence.
Methods
We studied 308,156 UK Biobank participants with no history of cancer at enrolment. Using 18 age-associated clinical biomarkers, we computed three BA measures (Klemera-Doubal method [KDM], PhenoAge, homeostatic dysregulation [HD]) and assessed their associations with incidence of any cancer and five common cancers (breast, prostate, lung, colorectal, and melanoma) using Cox proportional-hazards models.
Results
A total of 35,426 incident cancers were documented during a median follow-up of 10.9 years. Adjusting for common cancer risk factors, 1-standard deviation (SD) increment in the age-adjusted KDM (hazard ratio = 1.04, 95% confidence interval = 1.03–1.05), age-adjusted PhenoAge (1.09, 1.07–1.10), and HD (1.02, 1.01–1.03) was significantly associated with a higher risk of any cancer. All BA measures were also associated with increased risks of lung and colorectal cancers, but only PhenoAge was associated with breast cancer risk. Furthermore, we observed an inverse association between BA measures and prostate cancer, although it was attenuated after removing glycated hemoglobin and serum glucose from the BA algorithms.
Conclusions
Advanced BA quantified by clinical biomarkers is associated with increased risks of any cancer, lung cancer, and colorectal cancer.
Similar content being viewed by others
Background
Aging is closely linked to cancer [1,2,3], in which some of the proposed hallmarks of aging, such as genomic instability, cellular senescence, and epigenetic alteration [4], also overlap with the hallmarks of cancer [5]. Although chronological age (CA) is the dominant risk factor for most cancers [6], it does not capture the heterogeneity between older individuals. Biological age (BA), on the other hand, combines information from biological markers and may better reflect an individual’s physiology and risks of age-related diseases and death [7].
In recent years, various BA measures have been proposed and validated, including telomere length, deficit-accumulation frailty indices, epigenetic clocks based on DNA methylation markers, and algorithms that combine information on multiple clinical biomarkers [7, 8]. Accumulating evidence from observational studies has suggested that epigenetic clocks may predict cancer risks [9,10,11]. Recently, a Mendelian randomization study provided further support on the potential causal relationship between GrimAge acceleration, a second-generation epigenetic clock that reflects not only CA but also mortality and smoking [12], and the risk of colorectal cancer [13]. On the other hand, various biomarkers from blood chemistries (e.g., total cholesterol [14], glucose [15], C-reactive protein [16]) and other clinical data (e.g., waist circumference [17], forced expiratory volume [FEV1] [18]) have been linked to cancer risks. However, there has been a lack of data on whether composite measures of BA based on these routinely collected clinical biomarkers may predict cancer risks. As different types of BA measures may capture slightly different aspects of the aging process [19, 20], deciphering the link between various BA measures and cancers is important for understanding the mechanisms underlying aging and cancer.
Therefore, the aim of this study was to investigate the relationships between BA quantified based on clinical biomarkers and the risk of any cancer and site-specific cancers (including breast, prostate, lung, colorectal, and melanoma skin cancer), using data from the large population-based UK Biobank cohort. We hypothesize that these composite biomarker measures, as proxies for biological aging capturing the overall aging process, would be associated with increased risks of cancers independent of CA and other risk factors.
Methods
Study population
During 2006–2010, the UK Biobank enrolled over 500,000 participants aged 37–73 years from the general population [21]. At baseline, participants completed a touch-screen questionnaire, provided biological samples, and had physical measurements taken in 22 assessment centers throughout England, Wales, and Scotland. The UK Biobank study was approved by the North West Multi-Centre Research Ethics Committee. All participants provided written informed consent.
In this analysis, we included 331,699 UK Biobank participants who had complete data on the 18 biomarkers used in our BA algorithms (Table 1). We also excluded n = 177 with outlier BA values (considered as ±5 standard deviations [SD] from mean) and n = 23,366 with any cancer diagnosis (except non-melanoma skin cancer) before baseline, yielding an analytical sample of n = 308,156.
Biological age measures
We quantified BA based on three composite measures of blood chemistry and other clinical data: Klemera-Doubal method (KDM) [22], PhenoAge [23], and homeostatic dysregulation (HD) [24]. Details on the calculations and interpretations of the three measures have been summarized previously [25, 26]. Briefly, KDM is calculated from a series of regressions of biomarkers on CA and can be interpreted as the age at which the average physiology in the US National Health and Nutrition Examination Surveys (NHANES) III (i.e., the training sample) matches the physiology of the person. PhenoAge is calculated based on a mortality prediction score of biomarkers and CA and can be interpreted as the age at which the average mortality risk in NHANES III matches the predicted mortality risk. Different from KDM and PhenoAge, HD does not include CA in the calculation, but it is calculated based on the Mahalanobis distance [27] for a set of biomarkers relative to a reference sample and can be interpreted as the deviation of the person’s physiology from a healthy sample of NHANES III participants aged 20–30. All three measures have previously been validated for their abilities to predict diseases, disability, and mortality [25, 26].
In general, any age-related biomarkers can be used for constructing the BA algorithms. To facilitate comparison, we selected the same set of biomarkers in our KDM, PhenoAge, and HD algorithms, which were computed using the R package BioAge [25] in three steps:
-
1.
Training in NHANES III. We first identified 19 potential biomarkers covering a range of organ systems (e.g., cardiometabolic, inflammatory, kidney, lung functions) that are routinely collected in clinical practice and were available in NHANES III and UK Biobank (Supplementary Fig. 1). Only those with ≤20% missing data and correlated with CA ( |r | > 0.1, in accordance with prior work [19, 28]) in NHANES III were considered. Pulse was excluded due to its high correlation with systolic blood pressure (r = 0.84). Therefore, our new KDM, PhenoAge, and HD algorithms included 18 biomarkers (Table 1): FEV1, systolic blood pressure, blood urea nitrogen, glycated hemoglobin (HbA1c), total cholesterol, creatinine, serum glucose, waist circumference, red cell distribution width, albumin, alkaline phosphatase, triglyceride, mean cell volume, uric acid, lymphocyte percent, red blood cell count, C-reactive protein, and diastolic blood pressure. Following previous work [25], we selected non-pregnant participants who aged 30–75 years and had complete biomarker data as the reference population for KDM (n = 7694). The reference population for PhenoAge included those aged 20–84 years and with complete biomarker data (n = 12,998). The reference population for HD included participants aged 20–30 years who were not obese and whose biomarker values were within the age- and sex-specific normal range (n = 258). Only one measurement occasion was available per person in the training set.
-
2.
Testing the new BA algorithms in comparison to the published algorithms for their ability to predict mortality in an independent cohort of NHANES IV participants (n = 3849), who were recruited during 1999–2014 and followed up to 2015. Similar to the original version of KDM [28] and PhenoAge [23] (constructed using an alternative list of biomarkers as shown in Table 1), our new BA algorithms were statistically significantly associated with mortality during a median follow-up time of 7.4 years (interquartile range 4.1–11.5) (Supplementary Table 1). Besides, the new KDM and PhenoAge were strongly correlated with CA (r > 0.9), and all BA measures were moderately correlated with each other in NHANES IV (Supplementary Figs. 2 and 3).
-
3.
Projecting the newly trained algorithms onto UK Biobank. Correlations among the 18 biomarkers are shown in Supplementary Table 2, and the distributions of the BA measures in UK Biobank are presented in Supplementary Fig. 4.
To calculate the deviation between BA and CA, we regressed out CA (as 3 degrees-of-freedom natural spline) from KDM and PhenoAge in each cohort and considered them as “age residuals” (also known as “age acceleration”) [29]. Residuals were not calculated for HD as it was not an age measure by definition and it already quantifies deviation from a reference population [25, 26], but it was log-transformed due to the skewed distribution. Higher values of KDM residual, PhenoAge residual, and HD represent advanced BA. The KDM residual, PhenoAge residual, and HD were then standardized with mean = 0 and SD = 1 to allow comparison of effect sizes in subsequent analyses.
Cancer ascertainment
We studied five common cancers in Europe, including breast (for women), prostate (for men), and for both sexes, lung, colorectal, and melanoma skin cancer [30]. Incident cancers were ascertained from the cancer registries in England, Wales, and Scotland, where complete follow-up was available through February 29, 2020. We defined cancer diagnoses using the International Classification of Diseases, 10th revision (ICD-10) codes: any cancer (C00-97, excluding non-melanoma skin cancer C44), breast (C50), prostate (C61), lung (including trachea, C33-34), colorectal (C18-20), and melanoma (C43).
Statistical analyses
Participants were followed from the date of baseline assessment to the date of cancer diagnosis, death, or end of follow-up, whichever occurred first. Hazard ratios (HRs) for cancer risks per 1-SD increase in each BA measure were estimated using multivariable Cox proportional-hazards models, where attained age was used as the underlying timescale. The models were first adjusted for birth year and sex, and were further adjusted for baseline assessment center, ethnic background, body mass index (BMI), smoking, alcohol, physical activity, education, and Townsend deprivation index [31] in analyses of all cancer sites. Additionally, we adjusted for cancer-specific covariates such as family history of cancers, women-specific factors (menopausal status, hormone replacement therapy use, oral contraceptive use, parity), cancer screening, diet, and sun exposure variables, as relevant for each cancer site based on the literature. Covariates used in each model are listed in the footnote of the corresponding tables, and their definitions and descriptive statistics are provided in Supplementary Table 3. Missing data on covariates were coded as indicator variables in the models. To assess the associations of individual clinical biomarkers—in comparison to composite BA measures—with cancer risks, we also calculated the HRs per 1-SD increase in each clinical biomarker from the fully-adjusted models.
We performed subgroup analyses to test whether the associations may differ by age at baseline (<60 vs. ≥60 years), sex (women vs. men), and ethnicity (white vs. non-white). For breast cancer and lung cancer, we additionally stratified the analyses by menopausal status (premenopausal vs. postmenopausal) and smoking (never-smokers vs. ever-smokers), respectively. The proportional-hazards assumption was formally tested using Schoenfeld residuals. When the proportional-hazards assumption did not hold in the exposure of interest (P < 0.05), we fitted a time-varying model by including interaction terms between the BA measure and age (split into 5-year intervals) to calculate HRs over different periods of follow-up. Besides, to examine potential non-linear relationships between BA measures and cancer risks, we compared model fit of a restricted cubic spline model with a linear model using likelihood ratio tests and plotted the models with evidence of non-linearity (P < 0.05).
Several sensitivity analyses were performed. First, we assessed the association between the original KDM [28] and PhenoAge [23] algorithms (Table 1) and cancer risks to analyze if the biomarker composition would affect the results. Second, as we found an unexpected protective effect of BA measures for prostate cancer and that HbA1c and serum glucose may be associated with reduced prostate cancer risk [32, 33], we repeated the analysis using modified versions of KDM, PhenoAge, and HD computed from 16 biomarkers (i.e., removing HbA1c and serum glucose) to further examine whether the observed inverse relationship may be influenced by these two biomarkers. Third, instead of using indicator variables for missing values, we performed a complete-case analysis using available data (i.e., excluding individuals with missing data on any covariates). Finally, as individuals with a cancer diagnosis during the first 2 years of follow-up might have undiagnosed or subclinical disease at baseline, we performed a sensitivity analysis by excluding the first 2 years of follow-up to minimize reverse causation.
All analyses were performed in R 4.1.3 and Stata 16. To account for multiple testing (3 BA measures × 5 cancers), we applied the Bonferroni correction and considered a two-sided P < 0.0033 (i.e., 0.05/15) as statistically significant.
Results
Sample characteristics
Of the 308,156 UK Biobank participants, the mean age at baseline was 56.2 years (SD 8.1) and 163,022 (52.9%) were women (Table 2). During a median follow-up of 10.9 years (interquartile range 10.1–11.6), a total of 16,933 (10.4%) and 18,493 (12.7%) incident cancers were reported in women and men, respectively.
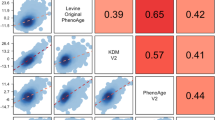
Correlations between the BA measures and CA in UK Biobank are shown in Fig. 1. As expected, CA was strongly correlated with KDM (i.e., an algorithm representing predicted CA; r = 0.85) and PhenoAge (i.e., representing predicted age-associated mortality; r = 0.84), while a weaker correlation was found between CA and HD (i.e., representing deviation of physiology from a healthy reference; r = 0.25). The residual (i.e., removing the effect of CA) of KDM was moderately correlated with the PhenoAge residual (r = 0.64) and HD (r = 0.60), but there was only weak correlation between the PhenoAge residual and HD (r = 0.29). Compared to men, women had a higher mean KDM residual (0.03 vs. −0.12) and HD (6.81 vs. 6.54 log units), but a lower mean PhenoAge residual (−0.65 vs. 0.67) (Table 2).
Biological age and cancer incidence
After adjusting for sociodemographic and lifestyle factors, all BA measures were associated with an elevated risk of any cancer (KDM residual: HR per 1-SD increase=1.04, 95% confidence interval [CI] = 1.03–1.05; PhenoAge residual: 1.09, 1.07–1.10; HD: 1.02, 1.01–1.03) (Table 3 and Supplementary Fig. 5). Similarly, the BA measures were associated with increased risks of lung cancer and colorectal cancer in the full models adjusted for cancer-specific risk factors. Among the three measures, PhenoAge residual had the strongest effect estimate for lung cancer (HR = 1.35, 95% CI = 1.30–1.40), and HD had the strongest effect estimate for colorectal cancer (HR = 1.10, 95% CI = 1.07–1.15). Only PhenoAge residual (HR = 1.05, 95% CI = 1.02–1.08), but not KDM residual or HD, was associated with a higher risk of breast cancer. We also observed significant protective effects of KDM and PhenoAge residuals for prostate cancer, although the associations were slightly attenuated after adjusting for prostate cancer-specific factors. None of the BA measures were statistically significantly associated with melanoma after adjustment for sociodemographic and lifestyle factors (Table 3). Many of the 18 clinical biomarkers incorporated within BA measures were also associated with cancer risks individually (Supplementary Table 4 and Supplementary Fig. 5). For instance, higher FEV1 was associated with decreased risks of any cancer and lung cancer but increased risk of prostate cancer, and higher systolic blood pressure was associated with elevated risk of breast cancer.
Associations between BA measures and cancer risks were mostly consistent across subgroups split by age and sex; however, the associations of KDM and PhenoAge residuals with colorectal cancer appeared to be sexually dimorphic and were only significant among men (Supplementary Table 5). When stratifying by ethnicity, results in non-white participants were generally statistically non-significant, possibly due to the limited sample size. Moreover, the associations between BA measures and lung cancer were only significant among ever-smokers but not never-smokers (P for interaction < 0.001), and the association between PhenoAge and breast cancer was stronger in postmenopausal vs. premenopausal women (P for interaction = 0.002) (Supplementary Table 5).
We calculated age-varying HRs when the proportional-hazards assumption was not met. As shown in Fig. 2, the associations of PhenoAge residual with any cancer and breast cancer appeared to be stronger around 60–80 years, while the associations of PhenoAge and KDM residuals with colorectal cancer tended to be weaker with advancing age. There was some evidence of non-linear associations between the residuals of KDM and PhenoAge and the risk of any cancer, lung cancer, and colorectal cancer (P < 0.05), with a sharp increase in the risk of any cancer among individuals with a BA value above the mean (Fig. 3).
a PhenoAge residual and risk of any cancer; b PhenoAge residual and risk of breast cancer in women; c KDM residual and risk of colorectal cancer; d PhenoAge residual and risk of colorectal cancer. Only the models with evidence of non-proportional hazards (P < 0.05) are shown. Estimates were obtained by including interaction terms between the exposure and the time variable (i.e., attained age, split into 5-year intervals) in the Cox models. The shaded areas indicate 95% confidence intervals (HRs outside the boundaries between 0.6 and 1.6 are not shown). The proportional hazards assumption of the biological age measures fitted in the Cox models was tested using Schoenfeld residuals. All models were adjusted for age (time scale), birth year, sex, baseline assessment center, ethnic background, body mass index, smoking status, physical activity level, alcohol consumption, education level, deprivation index quintiles, and the cancer-specific covariates as detailed in the footnote of Table 3 and in Supplementary Table 3. HR hazard ratio, KDM Klemera-Doubal method, SD standard deviation.
a Any cancer; b Prostate cancer in men; c Lung cancer; d Colorectal cancer. Only the models with evidence of non-linearity (P < 0.05) are shown. The black solid lines represent hazard ratios and the corresponding 95% confidence intervals (shaded areas) estimated using restricted cubic spline Cox regression models with knots at the 25th, 50th, and 75th percentiles. The dashed lines represent the estimates obtained from models assuming linear relationships. The mean of each BA measure was used as the reference value. P values for non-linearity were from likelihood ratio tests comparing the spline models with linear models. All models were adjusted for age (time scale), birth year, sex, baseline assessment center, ethnic background, body mass index, smoking status, physical activity level, alcohol consumption, education level, deprivation index quintiles, and the cancer-specific covariates as detailed in the footnote of Table 3 and in Supplementary Table 3. HD homeostatic dysregulation, KDM Klemera-Doubal method, SD standard deviation.
As the individual biomarkers included in the BA measures had different effects on cancer risks (Supplementary Fig. 5), we performed sensitivity analysis using alternative BA algorithms incorporating other sets of biomarkers. Results were essentially unchanged when using the original KDM and PhenoAge algorithms (Supplementary Table 6), as well as the modified algorithms of KDM, PhenoAge, and HD (excluding HbA1c and serum glucose) as exposure variables (Supplementary Table 7), although these alternative algorithms were no longer statistically significantly associated with prostate cancer in the fully-adjusted models. The associations were also largely similar when performing the complete-case analysis rather than using indicator variables for missing values of covariates (Supplementary Table 8), and when excluding diagnoses occurring in the first 2 years of follow-up (Supplementary Table 9).
Discussion
In this large cohort of UK Biobank participants, we quantified BA based on clinical biomarkers (e.g., blood chemistries, blood pressure, lung function) and showed that they were associated with increased risks of any cancer, lung cancer, and colorectal cancer, independent of CA, sex, sociodemographic and lifestyle factors, and other cancer-specific risk factors. These associations were largely consistent over the follow-up period and across subgroups split by age and sex. We also found a positive association between the age-adjusted PhenoAge and breast cancer. Furthermore, a protective effect of an advanced BA on prostate cancer was seen in the main analysis, although this seemed to be primarily driven by HbA1c and serum glucose included in the BA algorithms.
There is no gold standard for measuring BA. Different BA measures (e.g., telomere length, epigenetic clocks, biomarker-based BA) are weakly correlated with each other, hence, they may explain different aspects of biological aging [19, 20]. To the best of our knowledge, this is the first study that has used clinical biomarker-based BA algorithms to predict cancer risks. Similar to previous studies [14,15,16,17,18, 32, 33], we found that individual clinical biomarkers had different effects on various cancer types. Importantly, when combining information from multiple biomarkers as the proxies for biological aging (i.e., reflecting the overall physiological status of a person), these composite measures of BA had modest, but significant associations with any cancer, lung cancer, and colorectal cancer. Our results are also consistent with the existing literature, which has shown an association between higher epigenetic ages and an increased risk of any cancer [9,10,11], lung cancer [11, 34, 35], and colorectal cancer [13, 34]. Taken together, these findings may indicate that multiple aging processes captured by BA measures (e.g., epigenetic alterations, inflammation, metabolic changes) could play a role in cancer development. Although our BA algorithms included FEV1, which is a lung function biomarker strongly associated with lung cancer [18], we also observed a strong association between the original Levine PhenoAge algorithm (without FEV1 included) and lung cancer, thus suggesting that the association between BA measures and lung cancer was not entirely due to reduced lung function. Meanwhile, the association between BA measures and lung cancer was only significant among ever-smokers, but not never-smokers, indicating that it could partly be confounded by smoking status, where smoking could lead to both advanced BA and lung cancer risk [36]. Further research is warranted to study the mechanisms underlying biological aging, lung cancer and colorectal cancer.
We studied three algorithms, the KDM, PhenoAge, and HD, which were trained using different methods and may therefore have slightly different implications on biological aging. While the associations of the three BA measures and most cancers were comparable, we only observed a statistically significant association of PhenoAge, but not KDM or HD, with breast cancer. This is similar to prior research showing a statistically significant association between the PhenoAge clock (i.e., a DNA methylation-based algorithm trained to predict the PhenoAge used in our study) and increased risk of breast cancer [37], but a weaker association when using Horvath and Hannum clocks (i.e., algorithms to predict CA) [37, 38]. As we incorporated the same set of biomarkers into our three BA measures, the inconsistent result is probably explained by the fact that PhenoAge captures not merely CA, but also the mortality risk predicted by the biomarkers, whereas KDM simply reflects CA and HD reflects physiological deviations from a healthy reference.
The inverse association between BA and prostate cancer found in our primary analysis is somewhat in line with previous studies which showed indications of a protective effect of the GrimAge clock for prostate cancer [13, 34]. Interestingly, in our study, this association was slightly attenuated after adjusting for prostate cancer-specific factors such as diabetes and having prostate-specific antigen (PSA) test, and was further attenuated to non-statistically significance after removing HbA1c and serum glucose from the BA algorithms. Diabetes [39], as well as HbA1c [32] and serum glucose levels [33], have been linked to a lower incidence of prostate cancer, partly because of the lower level of insulin-like growth factor-1 levels in diabetic patients, and partly because of the potential detection bias due to the lower PSA level in diabetic men [32, 39]. We therefore speculate that the apparent protective effect of higher BA for prostate cancer could be confounded by diabetes and altered glucose metabolism, which are also closely related to aging [40]. Finally, no significant association was found between BA measures and melanoma. As melanoma is a cancer that is relatively common in young adults [30], it may be less driven by systemic aging and more by other factors such as skin sun exposure.
The primary strength of this study is the large sample size and the relatively long follow-up of ~10 years, which have allowed us to assess several cancers. Using the BioAge R package [25], we were able to assess the impact of multiple BA algorithms and compositions on cancer risks. Nevertheless, one limitation is the lack of data on tumor stage and grade. We therefore could not account for the severity of cancers. As we only have information on the clinical biomarkers at baseline, we were also unable to analyze if changes in BA over time may influence cancer risks. Moreover, the UK Biobank sample was mostly white participants (>94%), which may limit the generalizability of our results to other populations. Finally, as in other observational studies, although we have carefully adjusted for several cancer risk factors, the possibility of residual and unmeasured confounding cannot be ruled out.
In conclusion, our findings suggest that advanced biological aging may lead to increased risks of any cancer, lung cancer, and colorectal cancer, independent of age, sex, and common cancer risk factors. However, for other cancers such as breast cancer and prostate cancer, the associations may be influenced by the algorithm and composition of the BA being used. This work provides the basis for our further understanding on the biology underlying aging and cancer, and suggests that slowing down biological aging may be beneficial to mitigate cancer risks.
Data availability
UK Biobank is an open access resource. All bona fide researchers can apply to use its data for health-related research that is in the public interest (http://www.ukbiobank.ac.uk/register-apply).
Code availability
All codes used for generating the biological age measures and for conducting statistical analyses are available at https://github.com/jonathanklmak/UKB_bioage_cancer.
References
Aunan JR, Cho WC, Søreide K. The biology of aging and cancer: a brief overview of shared and divergent molecular hallmarks. Aging Dis. 2017;8:628–42. https://doi.org/10.14336/AD.2017.0103
Berben L, Floris G, Wildiers H, Hatse S. Cancer and aging: two tightly interconnected biological processes. Cancers. 2021;13. https://doi.org/10.3390/cancers13061400
Campisi J. Aging, cellular senescence, and cancer. Annu Rev Physiol. 2013;75:685–705. https://doi.org/10.1146/annurev-physiol-030212-183653.
López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. Hallmarks of aging: an expanding universe. Cell. 2023;186:243–78. https://doi.org/10.1016/j.cell.2022.11.001
López-Otín C, Pietrocola F, Roiz-Valle D, Galluzzi L, Kroemer G. Meta-hallmarks of aging and cancer. Cell Metab. 2023;35:12–35. https://doi.org/10.1016/j.cmet.2022.11.001
de Magalhães JP. How ageing processes influence cancer. Nat Rev Cancer. 2013;13:357–65. https://doi.org/10.1038/nrc3497
Jylhävä J, Pedersen NL, Hägg S. Biological age predictors. EBioMedicine. 2017;21:29–36. https://doi.org/10.1016/j.ebiom.2017.03.046.
Diebel LWM, Rockwood K. Determination of biological age: geriatric assessment vs biological biomarkers. Curr Oncol Rep. 2021;23:104. https://doi.org/10.1007/s11912-021-01097-9
Zheng Y, Joyce BT, Colicino E, Liu L, Zhang W, Dai Q, et al. Blood epigenetic age may predict cancer incidence and mortality. EBioMedicine. 2016;5:68–73. https://doi.org/10.1016/j.ebiom.2016.02.008
Dugué P-A, Bassett JK, Joo JE, Jung C-H, Ming Wong E, Moreno-Betancur M, et al. DNA methylation-based biological aging and cancer risk and survival: pooled analysis of seven prospective studies. Int J Cancer. 2018;142:1611–9. https://doi.org/10.1002/ijc.31189
Dugué P-A, Bassett JK, Wong EM, Joo JE, Li S, Yu C, et al. Biological aging measures based on blood DNA methylation and risk of cancer: a prospective study. JNCI Cancer Spectr. 2021;5:pkaa109. https://doi.org/10.1093/jncics/pkaa109
Lu AT, Quach A, Wilson JG, Reiner AP, Aviv A, Raj K, et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging. 2019;11:303–27. https://doi.org/10.18632/aging.101684
Morales Berstein F, McCartney DL, Lu AT, Tsilidis KK, Bouras E, Haycock P, et al. Assessing the causal role of epigenetic clocks in the development of multiple cancers: a Mendelian randomization study. eLife. 2022;11:e75374. https://doi.org/10.7554/eLife.75374
Kitahara CM, Berrington de González A, Freedman ND, Huxley R, Mok Y, Jee SH, et al. Total cholesterol and cancer risk in a large prospective study in Korea. J Clin Oncol. 2011;29:1592–8. https://doi.org/10.1200/JCO.2010.31.5200
Jee SH, Ohrr H, Sull JW, Yun JE, Ji M, Samet JM. Fasting serum glucose level and cancer risk in Korean men and women. J Am Med Assoc. 2005;293:194–202. https://doi.org/10.1001/jama.293.2.194
Allin KH, Nordestgaard BG. Elevated C-reactive protein in the diagnosis, prognosis, and cause of cancer. Crit Rev Clin Lab Sci. 2011;48:155–70. https://doi.org/10.3109/10408363.2011.599831
Kabat GC, Kim M, Hunt JR, Chlebowski RT, Rohan TE. Body mass index and waist circumference in relation to lung cancer risk in the women’s health initiative. Am J Epidemiol. 2008;168:158–69. https://doi.org/10.1093/aje/kwn109
Wasswa-Kintu S, Gan WQ, Man SFP, Pare PD, Sin DD. Relationship between reduced forced expiratory volume in one second and the risk of lung cancer: a systematic review and meta-analysis. Thorax. 2005;60:570 LP–575. https://doi.org/10.1136/thx.2004.037135
Li X, Ploner A, Wang Y, Magnusson PK, Reynolds C, Finkel D, et al. Longitudinal trajectories, correlations and mortality associations of nine biological ages across 20-years follow-up. eLife. 2020;9. https://doi.org/10.7554/eLife.51507
Belsky DW, Moffitt TE, Cohen AA, Corcoran DL, Levine ME, Prinz JA, et al. Eleven telomere, epigenetic clock, and biomarker-composite quantifications of biological aging: do they measure the same thing. Am J Epidemiol. 2018;187:1220–30. https://doi.org/10.1093/aje/kwx346
Sudlow C, Gallacher J, Allen N, Beral V, Burton P, Danesh J, et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 2015;12:e1001779. https://doi.org/10.1371/journal.pmed.1001779
Klemera P, Doubal S. A new approach to the concept and computation of biological age. Mech Ageing Dev. 2006;127:240–8. https://doi.org/10.1016/j.mad.2005.10.004
Levine ME, Lu AT, Quach A, Chen BH, Assimes TL, Bandinelli S, et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging. 2018;10:573–91. https://doi.org/10.18632/aging.101414
Cohen AA, Milot E, Yong J, Seplaki CL, Fülöp T, Bandeen-Roche K, et al. A novel statistical approach shows evidence for multi-system physiological dysregulation during aging. Mech Ageing Dev. 2013;134:110–7. https://doi.org/10.1016/j.mad.2013.01.004
Kwon D, Belsky DW. A toolkit for quantification of biological age from blood chemistry and organ function test data: BioAge. GeroScience. 2021;43:2795–808. https://doi.org/10.1007/s11357-021-00480-5
Graf GH, Crowe CL, Kothari M, Kwon D, Manly JJ, Turney IC, et al. Testing black-white disparities in biological aging among older adults in the United States: analysis of DNA-methylation and blood-chemistry methods. Am J Epidemiol. 2022;191:613–25. https://doi.org/10.1093/aje/kwab281
Mahalanobis PC. Mahalanobis distance. Proceedings of the National Academy of Sciences of India. 1936;49:234–256.
Levine ME. Modeling the rate of senescence: can estimated biological age predict mortality more accurately than chronological age? Journals Gerontol Ser A. 2013;68:667–74. https://doi.org/10.1093/gerona/gls233
McEwen LM, Jones MJ, Lin DTS, Edgar RD, Husquin LT, MacIsaac JL, et al. Systematic evaluation of DNA methylation age estimation with common preprocessing methods and the Infinium MethylationEPIC BeadChip array. Clin Epigenet. 2018;10:123. https://doi.org/10.1186/s13148-018-0556-2
Dyba T, Randi G, Bray F, Martos C, Giusti F, Nicholson N, et al. The European cancer burden in 2020: Incidence and mortality estimates for 40 countries and 25 major cancers. Eur J Cancer. 2021;157:308–47. https://doi.org/10.1016/j.ejca.2021.07.039
Townsend P, Phillimore P, Beattie A. Health and deprivation: inequality and the North. London: Croom Helm. Health Policy. 1988;10:207-6.
Monroy-Iglesias MJ, Russell B, Crawley D, Allen NE, Travis RC, Perez-Cornago A, et al. Metabolic syndrome biomarkers and prostate cancer risk in the UK Biobank. Int J Cancer. 2021;148:825–34. https://doi.org/10.1002/ijc.33255
Perez-Cornago A, Fensom GK, Andrews C, Watts EL, Allen NE, Martin RM, et al. Examination of potential novel biochemical factors in relation to prostate cancer incidence and mortality in UK Biobank. Br J Cancer. 2020;123:1808–17. https://doi.org/10.1038/s41416-020-01081-3
Levine ME, Hosgood HD, Chen B, Absher D, Assimes T, Horvath S. DNA methylation age of blood predicts future onset of lung cancer in the women’s health initiative. Aging. 2015;7:690–700. https://doi.org/10.18632/aging.100809
Hillary RF, Stevenson AJ, McCartney DL, Campbell A, Walker RM, Howard DM, et al. Epigenetic measures of ageing predict the prevalence and incidence of leading causes of death and disease burden. Clin Epigenet. 2020;12:115. https://doi.org/10.1186/s13148-020-00905-6
Mamoshina P, Kochetov K, Cortese F, Kovalchuk A, Aliper A, Putin E, et al. Blood biochemistry analysis to detect smoking status and quantify accelerated aging in smokers. Sci Rep. 2019;9:142. https://doi.org/10.1038/s41598-018-35704-w
Kresovich JK, Xu Z, O’Brien KM, Weinberg CR, Sandler DP, Taylor JA. Methylation-based biological age and breast cancer risk. J Natl Cancer Inst. 2019;111:1051–8. https://doi.org/10.1093/jnci/djz020
Ambatipudi S, Horvath S, Perrier F, Cuenin C, Hernandez-Vargas H, Le Calvez-Kelm F, et al. DNA methylome analysis identifies accelerated epigenetic ageing associated with postmenopausal breast cancer susceptibility. Eur J Cancer. 2017;75:299–307. https://doi.org/10.1016/j.ejca.2017.01.014
Feng X, Song M, Preston MA, Ma W, Hu Y, Pernar CH, et al. The association of diabetes with risk of prostate cancer defined by clinical and molecular features. Br J Cancer. 2020;123:657–65. https://doi.org/10.1038/s41416-020-0910-y
Kalyani RR, Egan JM. Diabetes and altered glucose metabolism with aging. Endocrinol Metab Clin North Am. 2013;42:333–47. https://doi.org/10.1016/j.ecl.2013.02.010
Acknowledgements
This research was conducted using the UK Biobank resource, as part of the registered project 22224. This study was accomplished within the context of the Swedish National Graduate School on Ageing and Health (SWEAH).
Funding
This study was supported by the Swedish Cancer Society (21 1571 Pj), the Swedish Research Council (2018-02077, 2019-01272, 2020-06101), the Academy of Finland (grant no. 349335), the Sigrid Jusélius Foundation, the Loo & Hans Osterman Foundation, the Strategic Research Program in Epidemiology at Karolinska Institutet, the Karolinska Institutet Foundations, and the King Gustaf V and Queen Victoria’s Foundation of Freemasons. CEM was supported by a NIHR Academic Clinical Fellowship. Open access funding provided by Karolinska Institute.
Author information
Authors and Affiliations
Contributions
Conceptualization: JKLM and SH. Data curation: JKLM. Formal analysis: JKLM and CEM. Methodology: JKLM, CEM and RK-H. Supervision: RK-H, JJ and SH. Interpretation of results: all authors. Writing—original draft: JKLM. Writing—review & editing: all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The UK Biobank study was performed in accordance with the Declaration of Helsinki and had obtained ethical approval from the North West Multi-centre Research Ethics Committee (approval number: 11/NW/03820). All participants have provided their informed consent to participate.
Consent for publication
Not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mak, J.K.L., McMurran, C.E., Kuja-Halkola, R. et al. Clinical biomarker-based biological aging and risk of cancer in the UK Biobank. Br J Cancer 129, 94–103 (2023). https://doi.org/10.1038/s41416-023-02288-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-023-02288-w
- Springer Nature Limited
This article is cited by
-
Associations of daily eating frequency and nighttime fasting duration with biological aging in National Health and Nutrition Examination Survey (NHANES) 2003–2010 and 2015–2018
International Journal of Behavioral Nutrition and Physical Activity (2024)
-
Association between biological aging and periodontitis using NHANES 2009–2014 and mendelian randomization
Scientific Reports (2024)
-
Assessing Frailty in Gastrointestinal Cancer: Two Diseases in One?
Current Oncology Reports (2024)
-
Association between plant and animal protein and biological aging: findings from the UK Biobank
European Journal of Nutrition (2024)