Abstract
Gastric cancer ranks as the fifth most common human malignancy and the third leading cause of cancer related deaths. Depending on tumor stage, endoscopic or surgical resection supported by perioperative chemotherapy is the only curative option for patients. Due to late clinical manifestation and missing reliable biomarkers, early detection is challenging and overall survival remains poor. Organoids are cell aggregates cultured in three-dimensions that grow with similar characteristics as their tissue-of-origin. Due to their self-renewal and proliferative capacity, organoids can be maintained long term in culture and expanded in many cases in an unlimited fashion. Patient-derived organoid (PDO) libraries function as living biobanks, allowing the in depth analysis of tissue specific function, development and disease. The recent successful establishment of gastric cancer PDOs opens up new perspectives for multiple translational clinical applications. Here, we review different adult stem cell derived gastric organoid model systems and focus on their establishment, phenotypic and genotypic characterizations as well as their use in predicting therapy response.

Similar content being viewed by others
Facts
-
The stomach is an organ important for food processing, permanently in contact with nutrients and bacteria. To ensure a functional mucosa, a continuous self-renewal of the epithelium is required.
-
Organoids are a three-dimensional cell culture system showing self-renewal, differentiation, and proliferation capabilities.
-
Gastric organoids can be established from adult stem cells of primary tissue, embryonic stem cells or induced pluripotent stem cells.
-
Gastric cancer ranks as the fifth most common world’s malignancy and the third leading cause of cancer related deaths.
-
PDO biobanks of gastric cancer represent a useful tool in analyzing gastric cancer biology. PDOs allow individualized in vitro therapy and resistance testing.
Open questions
-
Is it feasible to generate and characterize cancer organoids between the patient’s cancer biopsy and the start of therapy in order to provide a therapy recommendation?
-
Adult stem cell derived organoid cultures are purely epithelial. To what extent does this situation recapitulate the biological behavior in vivo?
-
Is intra-tumoral heterogeneity present to various degrees in most cancers, a limiting factor for precision medicine, at least when organoids are derived from single biopsies with limited representation of the full genetic spectrum?
The stomach—organ for food storage and digestion
Anatomy, gland structure and function
The stomach is a muscular organ playing an important role in food storage and digestion. It consists of four main parts: cardia, fundus, corpus (body), and antrum (pylorus) (Fig. 1A) [1]. The corpus is the main part of the stomach secreting acid and digestive enzymes. The antrum plays an important role in hormone and mucus secretion. The murine stomach additionally possesses a squamous-epithelium lined forestomach important for storage and mechanical dissociation of food [2,3,4,5]. The stomach is in permanent contact to nutrients, toxins and bacteria that altogether generate a toxic environment [6]. To ensure an intact and functional mucosa, a continuous self-renewal of the epithelium is required.
The stomach mucosa is composed of an epithelial layer organized into glands. Four different regions are differentiated from bottom to top: base, neck, isthmus and pit. The corpus epithelium presents long glands with short pits containing mucus secreting pit and neck cells, acid secreting parietal cells, hormone secreting endocrine cells and digestive enzyme secreting chief cells (Fig. 1B) [7]. Antral glands are shorter, and have a relatively larger pit region. They also contain mucus secreting pit and neck cells as well as endocrine cells, but fewer basally located chief cells and no parietal cells [8]. The isthmus region contains proliferating cells and has long been considered the home of gastric epithelial stem cells [9]. As suggested by their names, mucus pit cells are found exclusively in the pit region and mucus neck cells in the neck region, whereas chief cells are located at the base region only. Hormone producing endocrine and acid secreting parietal cells scatter throughout the whole gland (Fig. 1B) [10,11,12,13].
Gastric stem cells
The presence of stem cells allows the life-long self-renewal capacity of the stomach. In contrast to the small intestine, where leucine-rich repeat-containing G-protein coupled receptor 5 (Lgr5) expressing cells at the crypt base constitute the undisputed stem cell population, the identity of the stomach stem cell is still under debate. The intestinal stem cell marker Lgr5 was found to be also expressed at the bottom of the antral gland (Fig. 1B). Lineage tracing proved the stem cell properties of this cell type, as Lgr5+ cells self-renewed and were able to differentiate into all cell types of the antral epithelium [3, 14]. For the corpus gland, radiolabeling electron microscopy suggested a granule free undifferentiated cell within the isthmus [13, 15]. Trefoil factor 2 (Tff2) mRNA transcript expressing cells, as opposed to TFF2 protein expressing neck cells, were shown to constitute short-lived progenitor cells of mucus neck, chief and parietal cells[16],. SRY (sex determining region Y)-box 2 (Sox2) expressing cells have also been proposed to constitute long-lived stem cells of the gastric epithelium [17]. Nevertheless, Sox2 is widely expressed throughout the gastric unit. A further study promoted basic helix-loop helix family member a15 (Bhlha15 or Mist1) expressing isthmus cells as stem cells of the stomach epithelium [18]. Of note, Mist1 is also expressed by chief cells at the bottom of the glands. Cells expressing the tumor necrosis factor receptor superfamily member 19 (Tnfrsf19 or Troy) gene located at the gland base were shown by lineage tracing to self-renew and generate all cell types of the gastric gland, albeit inefficiently [19, 20]. Surprisingly, Troy+ cells are fully differentiated chief cells. Due to their quiescent, slow proliferating nature, which can be activated by tissue damage, Troy+ cells were proposed to serve as reserve stem cells [19]. This phenomenon opposes classical textbook knowledge, which describes an uni-directional differentiation flow of cells coming from stem cells toward differentiated cells, but might be a back-up mechanism active in several tissues [21]. It therefore appears that the “stem cell state” at least in certain stem cell populations can be influenced by the local microenvironment, which is subject to change in case of disturbance of homeostasis. A more general definition for a stem cell might therefore be: “A stem cell has the ability to replace lost tissue through cell division” [22].
In order to clarify the role and potential interplay of the two stem cell populations in the isthmus and base, Han et al. applied a detailed lineage tracing approach [23]. The isthmus stem cells, residing in the narrow zone between the pit and neck region, are multipotent and maintain the pit-isthmus-neck region by stochastic self-renewal, while the Troy+ stem cell population maintains chief cell regeneration at the gland bottom. This unifying concept was confirmed by a recent publication by Burclaff et al [24].
Similar to other tissues, gastric stem cells are discussed as the origin of gastric cancer. As they are long-lived, they are prone to accumulate mutations over time, eventually leading to aberrant signaling and finally full-blown gastric cancer. For the intestine, the Lgr5+ stem cell population has already been proven to constitute an important origin of intestinal cancer [25, 26].
Gastric cancer
Gastric cancer ranks as the fifth most common world’s malignancy after cancers of the lung, breast, colorectum and prostate. The incidence rate decreased since the mid-20th century [27]. However, it remains the third leading cause of cancer related deaths worldwide [28, 29]. Adenocarcinoma of the esophagogastric junction (AEG) represents a cancer with clear histopathological and molecular overlap to gastric cancer. Of note, this entity shows a rising incidence rate over the last decades [30, 31]. Due to late clinical signs, diagnosis is delayed in three quarters of patients presenting with non-curable disease resulting in a poor overall prognosis [32]. Endoscopic or surgical resection is the only curative option, often supported by interdisciplinary approaches, i.e., perioperative chemotherapy. The most common conventional chemotherapeutic drugs used are fluoropyrimidines (i.e., 5-fluoruracil (5-FU), capecitabine, S-1), platinum compounds (i.e., cisplatin, oxaliplatin), docetaxel and epirubicin [33,34,35]. Besides the classical chemotherapy, genetic alterations represent molecular targets for potential targeted treatments. At the moment the only approved targeted therapies are trastuzumab, a monoclonal antibody against the human epidermal growth factor receptor 2 (HER2)/neu signaling, and the anti-vascular endothelial growth factor receptor (VEGFR) antibody ramucirumab [36, 37]. Other targeted therapies, like the anti-epidermal growth factor receptor (EGFR) antibodies cetuximab and panitumumab or the anti-VEGFR antibody bevacizumab, have so far failed to improve the survival rates. However, the latter studies were performed in mostly unselected patient cohorts due to missing biomarkers.
While the World Health Organization (WHO) classified gastric cancer into four main subtypes, the widely used Lauren classification divides gastric cancer into the intestinal, diffuse and intermediate subtype [38]. The intestinal subtype shows a glandular morphology of tumor cells and often metastasizes to the liver and lung. In contrast, the diffuse subtype is characterized by a non-coherent diffuse growth of cancer cells, characteristic signet-ring cells and a frequent metastatic spread to the peritoneum. In addition, the Cancer Genome Atlas (TCGA) study described a robust classification system into four subtypes based on molecular alterations [39]. The first group of gastric tumors is associated with an Epstein Barr virus (EBV) infection and is therefore termed “EBV subtype”. Tumors often show phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit alpha (PIK3CA) and AT-rich interactive domain-containing protein 1 A (ARID1A) mutations, cyclin-dependent kinase inhibitor 2 A (CDKN2A) silencing and a widespread hypermethylation of promotor regions. The second group of gastric tumors shows microsatellite instability (MSI) through hypermutation, MutL homolog 1 (MLH1) silencing and mutations of the DNA damage repair system. Accordingly, this group is termed “MSI subtype”. The third genetically defined gastric cancer cohort is named “chromosomal instability” (CIN) subtype. Cancer cells characteristically show an intestinal glandular histology with frequent tumor protein 53 (TP53) mutations, receptor tyrosine kinase (RTK)-RAS pathway activation by receptor amplifications and a high amount of somatic copy number alterations (SCNA). The fourth molecular subtype shows a diffuse morphology of cancer cells due to the frequent loss of the cell adhesion molecule cadherin 1 (CDH1). In addition frequent alterations are found in Ras homolog family member A (RHOA) and ARID1A. Due to a relatively low amount of SCNA, it is named “genomically stable” (GS) subtype.
Organoids—a self-organizing and self-renewing three-dimensional cell culture model
Organoids are a recently developed three-dimensional (3D) cell cultivation system from adult stem cells (AdSCs) of primary tissue or embryonic stem cells (ESCs)/induced pluripotent stem cells (iPSCs) (together PSCs) [40, 41]. Cells are embedded in a laminin-rich extracellular matrix mimicking a native extracellular microenvironment. Some AdSC-derived organoids maintain or establish an intact stem cell niche, while others niche factors have to be supplemented in the medium. Stem cells in organoid cultures typically show a self-renewal and often an unlimited proliferative capacity. Depending on the amount of supplemented growth factors via the medium, cells within AdSC-derived organoids can differentiate into all or at least several differentiated cell types in the epithelium of the tissue they are derived from. PSC-derived organoids contain cells from different germ layers. Organoids typically exhibit physiological functions of the derived organs [41,42,43,44,45]. Therefore they constitute in many cases a near physiological cell culture model.
In 2009 the laboratory of Hans Clevers was the first to describe the AdSC-derived organoid system. In a seminal report, Sato et al. described a method to culture mouse intestinal organoids based on the knowledge of the specific growth factors required by intestinal stem cells [46]. The small intestinal epithelium is divided into crypts and villi as its main units [47]. The identification of the Lgr5+ stem cell located at the crypt base and their transcriptional profile formed the scientific basis of small intestinal organoid establishment [48, 49]. Mouse Lgr5+ stem cells were embedded in an extracellular matrix and overlaid with a medium containing the following niche factors: a WNT signaling agonist (Rspondin), epidermal growth factor (EGF) and Noggin. In general, WNT signaling is an essential requirement for crypt proliferation as well as for the maintenance of the intestinal stem cell pool [50,51,52]. WNT ligands are secreted by Paneth cells and stromal cells surrounding the intestinal crypt [53,54,55]. Within the intestinal murine organoid culture, which is a stroma-free cell culture system, Paneth cells constitute the essential WNT source for stem cells [56]. Rspondin is a potent WNT agonist enhancing WNT signaling via the LGR5/ring finger protein 43 (RNF43) axis [57]. Rspondin inhibits the action of RNF43/zinc and ring finger 3 (ZNRF3) by forming a complex with Rspondin and LGR4/5. This allows cells to express enough level of the WNT receptor Frizzled on the plasma membrane, which ensures constitutive WNT stimulation [58, 59]. Another important factor is EGF, which is important for epithelial proliferation and survival of undifferentiated cells [60,61,62]. EGF is secreted by Paneth cells [56], but still needs to be added to the intestinal organoid medium to allow long term growth. Bone morphogenetic proteins (BMPs) belong to the transforming growth factor-beta (TGF-β) family. TGF-β/BMP signaling inhibits epithelial proliferation and promotes differentiation in the intestinal crypt [63,64,65]. To prevent stem cell differentiation, Noggin as a BMP antagonist needs to be added to the growth medium. Within this culture setup, seeded single Lgr5+ cells are able to develop into mature organoids [46]. The intestinal organoid culture was later adapted to other mouse organs i.e., liver, lung, stomach, pancreas, prostate, endometrium, bladder, and salivary gland [14, 19, 66,67,68,69,70,71,72,73,74,75,76,77]. Organoids can also be grown from human tissue by adopting the growth requirements of murine culture protocols, allowing the establishment of large organoid collections [75, 77,78,79,80,81,82,83,84].
All those cultures have in common that they are grown in an extracellular matrix, which allows an outgrowth in 3D, and that they are cultured with a specific set of supplemented growth factors mimicking the niche microenvironment of each tissues and organs. Depending on the culture medium, differentiation of stem cells into organ-specific lineages is in part hampered by the presence of stem cell factors. For example, mouse colon organoids need a high concentration of WNT for their maintenance, which at the same time blocks differentiation of the stem cells [78]. Many cultures therefore consist of mainly stem and progenitor cells with rather immature differentiated cells, while fully differentiated cell types can only be observed after removing stem cell growth factors in the medium, resulting in the loss of stem cell driven longevity. Nevertheless, these stepwise culture conditions (expansion media and differentiation media) allows researchers to culture both stem/progenitor cells and differentiated cells. Further improvements of the culture conditions might resolve this problem in the future. For example, an improved medium condition by adding insulin-like-growth-factor-1 (IGF-1) and fibroblast growth factor-2 (FGF2), instead of adding EGF to the growth medium, preserved cellular diversity in normal human intestinal organoids, leading to the presence of stem and progenitor as well as differentiated goblet cells and a few Paneth cells [85].
The main advantage of the organoid culture system over most other primary cell cultures is the possibility to maintain the genomic stability of cells for a long period of time while retaining the characteristics of the tissue-of-origin [86, 87]. Organoids can be split, frozen and thawed like the traditional 2D cultures. All these advantageous features make organoids a useful technology for many laboratory applications. Organoid cultures are compatible to molecular characterization by genomic, (single cell) transcriptomic and (large-scale) proteome analyses. It is also possible to apply genetic manipulations using lentiviruses, bacterial artificial chromosomes (BAC) or the clustered regularly interspaced short palindromic repeats (CRISPR/Cas9) system [80, 83, 88,89,90,91,92,93,94,95,96]. Bacterial or viral infections into the lumen of organoids play an important role for studying infectious diseases [97,98,99].
In conclusion, the current organoid technology enables patient-derived organoid collections, genetic manipulations, and various omics studies, opening up unprecedented new possibilities for individualized treatment testing and in depth analysis of the underlying pathobiology. In particular, with the help of organoid biobanks, high-throughput drug screenings can be performed to identify novel treatment options [100].
Gastric organoids
Adult stem cells and pluripotent stem cells allow the establishment of gastric organoids
Gastric organoids may be initiated from stomach tissue derived AdSCs as well as PSCs. The main difference between the systems is the presence of mesenchymal cells within the PSC-derived organoid culture. AdCSs can only generate the specific cells from the tissue-of-origin, whereas PSCs have by nature the ability to differentiate into any cell types. Therefore, PSC-derived organoids require a stepwise differentiation protocol that guides PSC differentiation into the target tissue identity, while AdSC-derived gastric organoids from the start only require a single growth factor-enriched medium. Consequently, the time to differentiate PSCs into organoids takes ~30–60 days, while AdSC-derived organoids are established within 7–14 days.
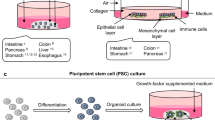
The first murine adult stem cell derived stomach organoid culture was established from antrum glands containing Lgr5+ stem cells. The protocol was developed on the basis of the intestinal organoid culture system by the addition of fibroblast growth factor 10 (FGF10) and the hormone gastrin (Fig. 2A) [14]. In addition, markers of chief cells (pepsinogen C (PGC)) and mucus neck cells (MUC6) could be observed. Reduction of the WNT concentration resulted in the generation of the differentiated lineages of mucous pit and endocrine cells, while parietal cells were not observed [14]. The same conditions were later used for murine corpus organoids originating from Troy+ stem cells (Fig. 2B) [19]. These organoids expressed markers of chief cells and mucus neck cells. Upon the withdrawal of WNT, Noggin and FGF10 differentiated pit cells could be observed, but no endocrine or parietal cells.
A Mouse or human antral organoids and (B) mouse or human corpus organoids are initiated by isolating stomach glands, embedding them in extracellular matrix and supporting them with growth media composed of EGF, WNT, Rspondin, Noggin, FGF10 and gastrin. The inhibition of the TGF-β signaling pathway increased the longevity of human corpus organoids. C Differentiation of PSC-derived human antral and corpus organoids. PSCs were obtained from blastocysts (embryonic stem cells (ESC)) or by reprogramming of differentiated cells (induced PSCs (iPSCs)). Cells were differentiated into endoderm by addition of activin A and BMP4. Posterior foregut formation was reached by supplementing with FGF4 and WNT or CHIR99021. Noggin was added for foregut differentiation. Embedding of these cells into an extracellular matrix led to the generation of 3D foregut spheroids. Antral differentiation was achieved by retinoic acid (RA) and EGF treatment. To direct foregut into corpus lineage the organoids were cultured with CHIR99021, EGF and FGF10. D Differentiation of PSC-derived mouse corpus organoids. PSCs were cultured as embryoid bodies and exposed to sonic hedgehog (SHH), the WNT antagonist dickkopf 1 (DKK1) as well as Noggin. Embedding of spheroids into extracellular matrix with addition of FGF10, Noggin, WNT and Rspondin led to mouse corpus gland formation.
Human antral organoids can be established using the mouse protocol (Fig. 2A) [101]. Human corpus organoids need the inhibition of the TGF-β signaling pathway by A83-01 (activin receptor-like kinase (ALK 5) inhibitor) for successful long term growth (Fig. 2B) [99].
The differentiation of PSCs into organoids allows the establishment of gastric organoids containing both epithelial and mesenchymal cells. McCracken et al. described the first differentiation protocol of human PSCs into gastric organoids (Fig. 2C) [102, 103]. Firstly, human PSCs were differentiated into endoderm by addition of activin A and BMP4. Activin A stimulates Nodal signaling, a highly conserved pathway important for foregut formation. Posterior foregut formation was achieved by addition of FGF4 and WNT or CHIR99021, a glycogen synthase kinase 3(GSK3)/β inhibitor stimulating the WNT pathway. To generate foregut, from which the stomach derives, Noggin was additionally applied and led to inhibition of BMP signaling. Embedding of these cells into an extracellular matrix led to the generation of 3D foregut spheroids. Antral differentiation was achieved by retinoic acid (RA) and EGF treatment. The complete differentiation required ~34 days resulting in antral organoids containing pit, mucus neck, and enteroendocrine cells [102, 104]. To direct foregut into corpus, the organoids were further supplemented with CHIR99021, EGF, and FGF10. To stimulate the production of parietal cells, the medium was supplemented subsequently with BMP4 and the MEK inhibitor PD0325901 (Fig. 2C) [103]. Differentiated corpus organoids contained pit, mucus neck, endocrine, chief and parietal cells [103, 104]. In a similar way, using a stepwise differentiation protocol, Noguchi et al. successfully generated organoids from murine PSCs (Fig. 2D) [105]. Here, the PSCs were cultured as embryoid bodies and exposed to sonic hedgehog (SHH), the WNT antagonist dickkopf 1 (DKK1) as well as Noggin. SHH activation as well as WNT signaling inhibition allowed tube-like structure formation and the generated spheroids resembled early stomach like structures. Embedding of spheroids into extracellular matrix with addition of FGF10, Noggin, WNT and Rspondin led to corpus gland formation after ~60 days. Similarly to the human PSC-derived corpus organoids, the murine PSC-derived organoids contained pit, mucus neck, endocrine, chief and parietal cells (Fig. 2D) [105].
In order to overcome the limitation of AdSC-derived organoids with its purely epithelial composition, a co-cultivation protocol of murine AdSC-derived organoids with mesenchymal cells was established [106]. The presence of mesenchymal niche cells resulted in the generation of all cells of the stomach epithelium including parietal cells (although only for a limited time span).
Overall, gastric normal organoids represent an excellent model system to answer questions of a wide range of topics, from basic science to translational clinical studies. Organoids can i.e., be used to recapitulate organ development [107]. They represent a useful tool to study Helicobacter pylori infection [108, 109]. Furthermore, organoids have been used successfully for disease modeling using CRISPR/Cas9 [110]. We will further focus on the establishment, characterization, and analysis of patient-derived gastric cancer organoid biobanks.
Patient-derived gastric cancer organoids
Cancer organoids as avatars of a patient’s tumor hold a great promise. The individual cancer organoid can be used to predict therapeutic responses to certain drugs, while the establishment of large PDO biobanks in combination with drug screens might be useful to delineate novel therapeutic strategies in gastric cancer in general.
Recently, four independent groups reported the generation of gastric PDOs [81,82,83,84] (Table 1). The protocol to culture cancer organoids was based on the described protocol for normal gastric tissue organoids [99]. Tissue samples from histologically confirmed (metastatic) gastric or esophagogastric junction adenocarcinoma were obtained from surgical resection specimens as well as endoscopic, ultrasound- and computed tomography (CT)-guided biopsies or ascites punctures (Table 1) [81,82,83,84]. Seidlitz et al. generated a biobank composed of 20 different human gastric cancer organoids with an in depth molecular analysis of four lines [81]. Vlachogiannis et al. generated a PDO biobank including cancers of different gastrointestinal origin, incl. four gastric cancers [82]. Of note, the organoids were generated from patients within clinical trials, allowing a correlation of patient to organoid response. Nanki et al. generated a biobank of 37 molecularly characterized gastric cancer organoids [83]. The study convincingly established a direct link between mutation pattern and growth factor independency in the culture medium, highlighting the different niche dependencies of individual cancers. The largest biobank was generated by Yan et al., consisting of 46 molecularly characterized organoid lines [84]. Interestingly, this study also compared organoids derived from multiple biopsies of the same patient, thus allowing the analysis of subclones within the primary cancer.
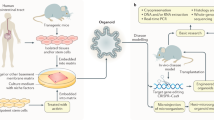
The general setup for gastric cancer organoid establishment followed the same lines in all studies. After obtaining the tissue, the specimen is enzymatically digested, extracted cells are embedded in extracellular matrix and overlaid with medium (Fig. 3A). Different protocols for the enzymatic digestion have been described: incubation with dispase II and collagenase XI [81], EDTA and TrypLe [82], Liberase TH and TrypLe Express [83] or collagenase and hyaluronidase [84]. Growth media composition also varied slightly within the studies (Table 1). A general problem in cancer organoid generation is the “contamination” of the culture with normal cell-derived organoids from the specimen. Different strategies have been used to enrich for cancer organoids [111]. In an elegant way, Nanki et al. enriched cancer organoids by blocking the frequently altered signaling pathways i.e., TP53, RHO, TGF-β and RAS-phosphoinositide 3-kinase (PI3K), which non-mutated normal organoids do not tolerate [83]. This resulted in an increase of cancer organoid generation efficiency from ~55 to 75% [83]. Yan et al. enriched tumor organoids by microscopical selection and, in case of TP53 mutation, by nutlin3a treatment (Table 1) [84].
A Scheme of PDO culture generation from a cancer biopsy: enzymatic digestion, embedding in extracellular matrix, addition of growth medium and cancer organoid enrichment by media compound withdrawal and/or addition of mutation related inhibitors. B Overview of PDO characterization based on morphology, molecular pattern, and drug response testing.
The further parts of this review focus on the characterization of patient derived gastric cancer organoids concerning phenotypic and molecular characterization as well as drug response testing (Fig. 3B).
Phenotypic characterization
Individual PDOs show different growth patterns characteristic to each line. Seidlitz et al. documented cystic organoids with or without a thickened (multilayered) wall, a non-coherent grape-like growth pattern and organoids with a compact cell cluster with no lumen [81]. The studies of Nanki and Yan grouped the divergent morphologies into three subtypes and correlated them to the Lauren classification: a solid-subtype derived mainly from diffuse gastric cancer showing amorphous solid configurations and a discohesive growth pattern, a glandular-subtype derived mainly from intestinal cancer showing glandular structures with a single lumen, and the mixed-subtype [38, 83, 84]. Cancer organoids also varied in proliferation rate, in line with different growth characteristics seen in patients [81]. Xenotransplantation of PDOs confirmed the tumorigenic potential of generated organoids and additionally the recapitulation of primary cancer morphology and histological subtypes [81, 83]. The link between phenotype and genotype was elegantly demonstrated by Nanki et al. by performing a knockout of CDH1 using CRISPR/Cas9. The CDH1 knockout led to a phenotypical change of organoids from normal cystic structures to solid structures with a vigorous migratory activity resembling PDOs with CDH1 mutation [83]. This experiment nicely demonstrates the usefulness of the organoid system in combination with genetic engineering to gain insights into phenotypical and functional mechanisms of certain mutations.
Molecular characterization
The availability of high throughput sequencing modalities allows molecular profiling by genetic, transcriptomic, and epigenetic analyses outlining the (epi)genetic landscape of the individual tumor. Observed mutation patterns of gastric cancer PDOs recapitulate the described TCGA stomach cancer subtypes, and PDOs of all subtypes can be generated [39]. PDO cultures of the MSI, GS, CIN as well as EBV subtype could be established [81, 83, 84]. Furthermore, a 96% overlap in the mutational spectrum of organoid and parental tissue was found [82]. One potential pitfall when using PDOs as avatars of a patient’s tumor is its derivation from a biopsy or small piece of tissue from a resection specimen. Intra-tumoral heterogeneity might play an important role in therapy resistance development out of underrepresented but resistant subclones. The generation of organoid lines from different areas of resection specimens in colorectal cancer has unraveled such intra-tumoral heterogeneity [112]. It will be crucial to clarify the importance of intra-tumoral heterogeneity for therapy response testing in the future. As all subclones within one tumor often harbor a common mutational origin, the arising differences during cancer progression might not necessarily lead to a differential response among subclones. If intra-tumoral heterogeneity turns out to be crucial for therapy response prediction, taking several biopsies could be a solution, but might not be ethically feasible which could limit the potential clinical usefulness of PDOs. Yan et al. observed varying degrees of tumor heterogeneity in gastric cancer by comparing PDOs from primary tumor and lymph node metastasis [84]. A PDO library of different tumors (i.e., primary and several metastases) of an individual patient constitutes a novel research tool to study the consequence of intra-tumoral heterogeneity.
PDO survival under growth factor withdrawal of relevant media compounds can indicate an acquired independency of certain pathways due to genetic alterations (Fig. 4). Nanki et al. investigated phenotype-genotype correlations by focusing on niche factor dependencies and occurred genetic aberrations [83]. Ki-ras2 kirsten rat sarcoma viral oncogene homolog (KRAS) mutation and RTK amplifications like erb-b2 receptor tyrosine kinase 2 (ERBB2) or erb-b2 receptor tyrosine kinase 3 (ERBB3) led to the acquisition of EGF and FGF growth factor independency (Fig. 4A). The upregulation of epiregulin (EREG), a ligand of EGFR, mediated EGF and FGF independency suggesting an EREG autocrine loop for pathway activation (Fig. 4A) [83].
A EGFR/FGFR signaling pathway in normal and gastric cancer organoids. Normal gastric cancer organoids are dependent on EGF/FGF10. KRAS mutation or receptor tyrosine kinase (RTK) amplification lead to an EGF/FGF10 independency. The upregulation of the EGFR ligand epiregulin (EREG) also mediates EGF/FGF independency. B TGF-β and BMP signaling in normal and cancer gastric organoids. In normal gastric organoid cultures, TGF-β and BMP signaling has to be inhibited to avoid differentiation and ensure proliferation. Cancer organoids with mutations in these pathways tolerate TGF-β and BMP signaling. C WNT signaling in normal and gastric cancer organoids. WNT receptor stimulation is important for organoid proliferation. Cancer organoids with APC alterations develop WNT independency. Porcupine plays an important role in WNT ligand production. Inhibition of this protein revealed dependencies on autocrine WNT loops. D Rspondin/ZNFR3/RNF43 signaling in normal and gastric cancer organoids. Rspondin is essential for normal gastric organoid growth. Alterations of ZNFR3 and RNF43 or the single RNF43D300Y mutation results in Rspondin independence.
Gastric organoid growth is dependent on TGF-β and BMP signaling inhibition. However, TGF-β and BMP4 treatment of organoids carrying mutations in the transforming growth factor beta receptor 2 (TGFBR2) and the smad family member 4 (SMAD4) did not influence proliferation rate of organoids (Fig. 4B). Some gastric cancer organoids with no alterations in the aforementioned genes also tolerated stimulation with TGF-β and BMP4, hinting to additional nongenetic mechanisms that lead to the tolerance of PDOs toward TGF-β and BMP signaling [83, 113].
WNT and Rspondin are essential for successful normal gastric organoid growth (Fig. 4C, D). Therefore, gastric cancer organoids often acquire WNT pathway independency during tumorigenic progression e.g., by adenomatous polyposis coli (APC) gene mutations [81, 83]. Another mechanism constitutes the upregulation of WNT ligand production, leading to an autocrine self-stimulation of the tumor (Fig. 4C). This mechanism can be suppressed by treatment with the WNT ligand production inhibitor porcupine [114].
Among the WNT ligand dependent PDOs some represented an additionally unique Rspondin independency. Rspondin normally binds to LGR4/5 stabilizing the WNT receptors Frizzled and LRP (Fig. 4D). In the absence of Rspondin, RNF43 as well as ZNRF3 are ubiquitinating the WNT receptors [58, 59, 115]. RNF43 mutations are found in only 5% of microsatellite stable tumors, but have a high frequency of 55% in MSI subtype patients [116]. Interestingly, some stomach and intestinal cancer organoids with single RNF43 mutations were still Rspondin dependent [58, 83]. Subsequent genetic analyses revealed that double mutations, homozygous deletions as well as mRNA downregulation of RNF43 and the corresponding homolog ZNFR3 resulted in Rspondin independency. Interestingly, this was additionally seen for a RNF43D300Y single mutation (Fig. 4D).
Drug response testing
Treating gastric cancer PDOs with chemotherapeutic drugs frequently used in gastric cancer treatment resulted in varying degrees of response, comparable in its spectrum to clinical responses of patients [81, 84]. In addition, PDOs allow small to medium size drug screens that might identify additional vulnerabilities (Fig. 5A) [79]. Also gastric cancer PDOs have been exposed to such screens, resulting in interesting divergent response patterns between individual PDO lines [82, 84].
A General drug-screening setup with dose-response curves. B Targeted therapy options of PDOs depending on the present mutations. ERBB2 amplifications and/or mutations can be targeted with the monoclonal antibody trastuzumab. Aberrant PI3K signaling can be inhibited using the small molecules MK-2206 and GSK690693 targeting AKT1. Loss of the tumor suppressor CDKN2A results in an uncontrolled cell cycle. Resulting proliferation can be suppressed by treatment with palbociclib and abemaciclib. c Immunotherapeutic treatments can be tested in co-cultures of PDOs with reactive T-cells.
Besides classical chemotherapeutics and time and cost consuming drug screens individual PDOs can also be treated with targeted drugs against identified molecular alterations in i.e., sequencing data. Amplifications of ERBB2 are found in 22% of gastric cancer patients and patients carrying such a amplification can be successfully treated with trastuzumab [37]. Accordingly, response could be documented for a gastric cancer PDO line with an ERBB2 amplification, as well as in an PDO with an ERBB2 pathway activating mutation [81]. Mutations in the AKT serine/threonine kinase 1 (AKT1) gene resulted in a strong response to treatment with MK-2206 and GSK690693, both inhibitors of AKT [82]. Alterations in the tumor suppressor CDKN2A, frequently observed in gastric cancer (34%), play a key role in tumorigenic cell cycle progression [39]. Loss of the gene results in a permanently active cell cycle and therefore aberrant proliferation. Treatment with palbociclib or abemaciclib, both cyclin dependent kinase 4/6 (CDK4/6) inhibitors, resulted in a suppression of proliferation (Fig. 5B) [81, 84].
The use of organoids in personalized medicine, i.e., to delineate treatment strategies for individual patients; depends on the rate of correct prediction of response. In order to ascertain the predictive value of PDOs, co-clinical trials are needed to document side-by-side the clinical effect in vivo and the effect in organoids in vitro. The first study to document such correlations was carried out by the group of Nicola Valeri [82]. Overall, a strikingly high sensitivity, specificity as well as positive and negative predictive value or organoids to predict response to targeted drugs could be documented.
Taken together, first data indicate the usefulness of patient derived organoids in therapy prediction. In our experience, cancer organoids can be established from biopsies within 2–3 weeks for ~50% of patients to an extend that allows to test responses to standard chemotherapeutics. This is similar to the time frame of the routine work (i.e., CT and laparoscopic staging, intravenous port implantation) that happens before starting the neoadjuvant chemotherapy. Patients in a palliative setting normally receive first line of chemotherapy according to established protocols, before further personalized approaches are applied. This usually gives enough time to expand cultures to perform drug screens, i.e., in the form of a high-throughput drug screen as established by Du et al., before suggestions for second/third line treatment regimens are needed [117]. In our view, it will therefore be possible in the future to integrate PDOs into the clinical decision-making process. Of note, Yan et al. reported a heterogeneous drug response between PDOs from different tumor regions of one tumor [84]. Further co-clinical and first prospective studies in larger patient cohorts are therefore needed to clearly define the role of PDOs in different therapy settings (i.e., neoadjuvant/palliative). In addition, for novel immunotherapeutic approaches, co-culture systems with immune cells also need to be validated concerning in vitro functionality of tumor-immune cell interaction [118]. First promising data hint toward the possibility for functional testing of e.g., checkpoint inhibition therapies using immune cell-organoid co-cultures (Fig. 5C) [119].
Finally, for further interest we would like to direct the readers to other excellent reviews on different aspects of the topic of organoids [40, 100, 120,121,122,123,124,125,126,127,128,129,130]. The generation of tumor organoid libraries from other entities besides gastric cancer is reviewed in several articles [131,132,133,134,135,136].
Summary and future perspectives
Organoids open up new possibilities in characterization and understanding development, tissue homeostasis, and diseases. Organoids from normal tissues are used in a wide range of laboratory applications like CRISPR/Cas9 gene editing, bacterial or viral infections as well as transplantation assays. Large human cancer organoid biobanks from different cancer entities have been established and characterized in detail. Organoid libraries provide a human cancer resource to analyze cancer biology in living cells. Individualizing cancer treatment based on drug response testing has become feasible. In the future, patient derived organoids can bridge the gap between molecular genetics, current biological understanding, and clinical therapy.
Change history
18 August 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41418-021-00835-7
References
Karam SM. Lineage commitment and maturation of epithelial cells in the gut. Front Biosci. 1999;4:286–98.
Roman AKS, Shivdasani RA. Boundaries, junctions and transitions in the gastrointestinal tract. Exp Cell Res. 2011;317:2711–8.
Kim T-H, Shivdasani RA. Stomach development, stem cells and disease. Development. 2016;143:554–65.
Willet SG, Mills JC. Stomach organ and cell lineage differentiation: from embryogenesis to adult homeostasis. Cmgh. 2016;2:546–59.
Bartfeld S, Koo B-K. Adult gastric stem cells and their niches. Wiley Interdiscip Rev Dev Biol. 2017;6:e261.
Koelz H. Gastric acids in vertebrates. Scand J Gastroenterol Suppl. 1992;193:2–6.
Karam SM, Leblond CP. Identifying and counting epithelial cell types in the “corpus” of the mouse stomach. Anat Rec. 1992;232:231–46.
Lee ER, Trasler J, Dwivedi S, Leblond CP. Division of the mouse gastric mucosa into zymogenic and mucous regions on the basis of gland features. Am J Anat. 1982;164:187–207.
Bjerknes M, Cheng H. Multipotential stem cells in adult mouse gastric epithelium. Am J Physiol Liver Physiol. 2002;283:G767–77.
Karam SM, Leblond CP. Dynamics of epithelial cells in the corpus of the mouse stomach. II. Outward migration of pit cells. Anat Rec. 1993;236:280–96.
Karam SM, Leblond CP. Dynamics of epithelial cells in the corpus of the mouse stomach. III. Inward migration of neck cells followed by progressive transformation into zymogenic cells. Anat Rec. 1993;236:297–313.
Karam SM, Leblond CP. Dynamics of epithelial cells in the corpus of the mouse stomach. V. Behavior of entero-endocrine and caveolated cells: General conclusions on cell kinetics in the oxyntic epithelium. Anat Rec. 1993;236:333–40.
Karam SM, Leblond CP. Dynamics of epithelial cells in the corpus of the mouse stomach. I. Identification of proliferative cell types and pinpointing of the stem cell. Anat Rec. 1993;236:259–79.
Barker N, Huch M, Kujala P, van de Wetering M, Snippert HJ, van Es JH, et al. Lgr5+ve Stem Cells Drive Self-Renewal in the Stomach and Build Long-Lived Gastric Units In Vitro. Cell Stem Cell. 2010;6:25–36.
Mills JC, Shivdasani RA. Gastric epithelial stem cells. Gastroenterology. 2011;140:412–24.
Quante M, Marrache F, Goldenring JR, Wang TC. TFF2 mRNA transcript expression marks a gland progenitor cell of the gastric oxyntic mucosa. Gastroenterology. 2010;139:2018–27.
Arnold K, Sarkar A, Yram MA, Polo JM, Bronson R, Sengupta S, et al. Sox2 + adult stem and progenitor cells are important for tissue regeneration and survival of mice. Cell Stem Cell. 2011;9:317–29.
Hayakawa Y, Ariyama H, Stancikova J, Sakitani K, Asfaha S, Renz BW, et al. Mist1 Expressing Gastric Stem Cells Maintain the Normal and Neoplastic Gastric Epithelium and Are Supported by a Perivascular Stem Cell Niche. Cancer Cell. 2015;28:800–14.
Stange DE, Koo BK, Huch M, Sibbel G, Basak O, Lyubimova A, et al. Differentiated Troy+ chief cells act as reserve stem cells to generate all lineages of the stomach epithelium. Cell. 2013;155:357–68.
Leushacke M, Tan SH, Wong A, Swathi Y, Hajamohideen A, Tan LT, et al. Lgr5-expressing chief cells drive epithelial regeneration and cancer in the oxyntic stomach. Nat Cell Biol. 2017;19:774–86.
Desai TJ, Krasnow MA. Differentiated cells in a back-up role. Nature. 2013;503:204–5.
Post Y, Clevers H. Defining adult stem cell function at its simplest: the ability to replace lost cells through mitosis. Cell Stem Cell. 2019;25:174–83.
Han S, Fink J, Jörg DJ, Lee E, Yum MK, Chatzeli L, et al. Defining the Identity and Dynamics of Adult Gastric Isthmus Stem Cells. Cell Stem Cell. 2019;25:342–56.e7.
Burclaff J, Willet SG, Sáenz JB, Mills JC. Proliferation and Differentiation of Gastric Mucous Neck and Chief Cells During Homeostasis and Injury-induced Metaplasia. Gastroenterology. 2020;158:598–609.
Barker N, Ridgway RA, van Es JH, van de Wetering M, Begthel H, van den Born M, et al. Crypt stem cells as the cells-of-origin of intestinal cancer. Nature. 2009;457:608–11.
Schepers AG, Snippert HJ, Stange DE, van den Born M, van Es JH, van de Wetering M, et al. Lineage tracing reveals Lgr5+ stem cell activity in mouse intestinal adenomas. Science (80-). 2012;337:730–5.
Malvezzi M, Bonifazi M, Bertuccio P, Levi F, Vecchia] C[La, Decarli A, et al. An Age-Period-Cohort Analysis of Gastric Cancer Mortality from 1950 to 2007 in Europe. Ann Epidemiol. 2010;20:898–905.
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Ferlay J, Colombet M, Soerjomataram I, Mathers C, Parkin DM, Piñeros M, et al. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int J Cancer. 2019;144:1941–53.
McColl KEL, Going JJ. Aetiology and classification of adenocarcinoma of the gastro-oesophageal junction/cardia. Gut. 2010;59:282–4.
Arnold M, Soerjomataram I, Ferlay J, Forman D. Global incidence of oesophageal cancer by histological subtype in 2012. Gut. 2015;64:381–7.
Hunt RH, Camilleri M, Crowe SE, El-Omar EM, Fox JG, Kuipers EJ, et al. The stomach in health and disease. Gut. 2015;64:1650–68.
Ychou M, Boige V, Pignon JP, Conroy T, Bouch O, Lebreton G, et al. Perioperative chemotherapy compared with surgery alone for resectable gastroesophageal adenocarcinoma: An FNCLCC and FFCD multicenter phase III trial. J Clin Oncol. 2011;29:1715–21.
Cunningham D, Allum WH, Stenning SP, Thompson JN, Van de Velde CJ, Nicolson M, et al. Perioperative chemotherapy versus surgery alone for resectable gastroesophageal cancer. N Engl J Med. 2006;355:11–20.
Al-Batran S-E, Hofheinz RD, Pauligk C, Kopp H-G, Haag GM, Luley KB, et al. Histopathological regression after neoadjuvant docetaxel, oxaliplatin, fluorouracil, and leucovorin versus epirubicin, cisplatin, and fluorouracil or capecitabine in patients with resectable gastric or gastro-oesophageal junction adenocarcinoma (FLOT4-AIO. Lancet Oncol. 2016;17:1697–708..
Fuchs CS, Tomasek J, Yong CJ, Dumitru F, Passalacqua R, Goswami C, et al. Ramucirumab monotherapy for previously treated advanced gastric or gastro-oesophageal junction adenocarcinoma (REGARD): an international, randomised, multicentre, placebo-controlled, phase 3 trial. Lancet. 2014;383:31–39.
Bang Y-J, Van Cutsem E, Feyereislova A, Chung HC, Shen L, Sawaki A, et al. Trastuzumab in combination with chemotherapy versus chemotherapy alone for treatment of HER2-positive advanced gastric or gastro-oesophageal junction cancer (ToGA): a phase 3, open-label, randomised controlled trial. Lancet. 2010;376:687–97.
Lauren P. The two histological main types of gastric carcinoma: diffuse and so-called intestinal-type carcinoma. Acta Pathol Microbiol Scand. 1965;64:31–49.
The Cancer Genome Atlas Research Network. Comprehensive molecular characterization of gastric adenocarcinoma. Nature. 2014;513:202–9.
Huch M, Koo B-K. Modeling mouse and human development using organoid cultures. Development. 2015;142:3113–25.
Shamir ER, Ewald AJ. Three-dimensional organotypic culture: experimental models of mammalian biology and disease. Nat Rev Mol Cell Biol. 2014;15:647–64.
Date S, Sato T. Mini-Gut Organoids: reconstitution of the Stem Cell Niche. Annu Rev Cell Dev Biol. 2015;31:269–89.
Kretzschmar K, Clevers H. Organoids: modeling Development and the Stem Cell Niche in a Dish. Dev Cell. 2016;38:590–600.
min S, Kim S, Cho S-W. Gastrointestinal tract modeling using organoids engineered with cellular and microbiota niches. Exp Mol Med. 2020;52:227–37.
Nakamura T, Sato T. Advancing intestinal organoid technology toward regenerative medicine. Cell Mol Gastroenterol Hepatol. 2018;5:51–60.
Sato T, Vries RG, Snippert HJ, van de Wetering M, Barker N, Stange DE, et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature. 2009;459:262–5.
Clevers H. The Intestinal Crypt, A Prototype Stem Cell Compartment. Cell. 2013;154:274–84.
Barker N, van Es JH, Kuipers J, Kujala P, van den Born M, Cozijnsen M, et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature. 2007;449:1003–7.
Muñoz J, Stange DE, Schepers AG, van de Wetering M, Koo B-K, Itzkovitz S, et al. The Lgr5 intestinal stem cell signature: robust expression of proposed quiescent ‘+4’ cell markers. EMBO J. 2012;31:3079–91.
Pinto D, Gregorieff A, Begthel H, Clevers H. Canonical Wnt signals are essential for homeostasis of the intestinal epithelium. Genes Dev. 2003;17:1709–13.
Korinek V, Barker N, Moerer P, van Donselaar E, Huls G, Peters PJ, et al. Depletion of epithelial stem-cell compartments in the small intestine of mice lacking Tcf-4. Nat Genet. 1998;19:379–83.
Kuhnert F, Davis CR, Wang H-T, Chu P, Lee M, Yuan J, et al. Essential requirement for Wnt signaling in proliferation of adult small intestine and colon revealed by adenoviral expression of Dickkopf-1. Proc Natl Acad Sci. 2004;101:266–71.
Farin HF, Van EsJH, Clevers H. Redundant Sources of Wnt Regulate Intestinal Stem Cells and Promote Formation of Paneth Cells. Gastroenterology. 2012;143:1518–29.
Kabiri Z, Greicius G, Madan B, Biechele S, Zhong Z, Zaribafzadeh H, et al. Stroma provides an intestinal stem cell niche in the absence of epithelial Wnts. Development. 2014;141:2206–15.
Durand A, Donahue B, Peignon G, Letourneur F, Cagnard N, Slomianny C, et al. Functional intestinal stem cells after Paneth cell ablation induced by the loss of transcription factor Math1 (Atoh1). Proc Natl Acad Sci. 2012;109:8965–70.
Sato T, van Es JH, Snippert HJ, Stange DE, Vries RG, van den Born M, et al. Paneth cells constitute the niche for Lgr5 stem cells in intestinal crypts. Nature. 2011;469:415–8.
de Lau W, Peng WC, Gros P, Clevers H. The R-spondin/Lgr5/Rnf43 module: regulator of Wnt signal strength. Genes Dev. 2014;28:305–16.
Koo B-K, Spit M, Jordens I, Low TY, Stange DE, van de Wetering M, et al. Tumour suppressor RNF43 is a stem-cell E3 ligase that induces endocytosis of Wnt receptors. Nature. 2012;488:665–9.
Hao H-X, Xie Y, Zhang Y, Charlat O, Oster E, Avello M, et al. ZNRF3 promotes Wnt receptor turnover in an R-spondin-sensitive manner. Nature. 2012;485:195–200.
Dignass A, Sturm A. Peptide growth factors in the intestine. Eur J Gastroenterol Hepatol. 2001;13:763–70.
Suzuki A, Sekiya S, Gunshima E, Fujii S, Taniguchi H. EGF signaling activates proliferation and blocks apoptosis of mouse and human intestinal stem/progenitor cells in long-term monolayer cell culture. Lab Investig. 2010;90:1425–36.
Abud HE, Watson N, Heath JK. Growth of intestinal epithelium in organ culture is dependent on EGF signalling. Exp Cell Res. 2005;303:252–62.
Qi Z, Li Y, Zhao B, Xu C, Liu Y, Li H, et al. BMP restricts stemness of intestinal Lgr5+ stem cells by directly suppressing their signature genes. Nat Commun. 2017;8:13824.
He XC, Zhang J, Tong W-G, Tawfik O, Ross J, Scoville DH, et al. BMP signaling inhibits intestinal stem cell self-renewal through suppression of Wnt–β-catenin signaling. Nat Genet. 2004;36:1117–21.
Haramis A-PG, Begthel H, van den Born M, van Es J, Jonkheer S, Offerhaus GJA, et al. De Novo Crypt Formation and Juvenile Polyposis on BMP Inhibition in Mouse Intestine. Science (80-). 2004;303:1684–6.
Hu H, Gehart H, Artegiani B, LÖpez-Iglesias C, Dekkers F, Basak O, et al. Long-Term Expansion of Functional Mouse and Human Hepatocytes as 3D Organoids. Cell. 2018;175:1591–606.
Peng WC, Logan CY, Fish M, Anbarchian T, Aguisanda F, Álvarez-Varela A, et al. Inflammatory Cytokine TNFα Promotes the Long-Term Expansion of Primary Hepatocytes in 3D Culture. Cell. 2018;175:1607–19.e15.
Cao W, Liu J, Wang L, Li M, Verstegen MMA, Yin Y, et al. Modeling liver cancer and therapy responsiveness using organoids derived from primary mouse liver tumors. Carcinogenesis. 2018;40:145–54.
Mullenders J, de Jongh E, Brousali A, Roosen M, Blom JPA, Begthel H, et al. Mouse and human urothelial cancer organoids: a tool for bladder cancer research. Proc Natl Acad Sci. 2019;116:4567–74.
Xin L, Lukacs RU, Lawson DA, Cheng D, Witte ON. Self-Renewal and Multilineage Differentiation In Vitro from Murine Prostate Stem Cells. Stem Cells. 2007;25:2760–9.
Huch M, Boj SF, Clevers H. Lgr5+ liver stem cells, hepatic organoids and regenerative medicine. Regen Med. 2013;8:385–7.
Huch M, Bonfanti P, Boj SF, Sato T, Loomans CJM, van de Wetering M, et al. Unlimited in vitro expansion of adult bi-potent pancreas progenitors through the Lgr5/R-spondin axis. EMBO J. 2013;32:2708–21.
Huch M, Dorrell C, Boj SF, van Es JH, Li VSW, van de Wetering M, et al. In vitro expansion of single Lgr5+ liver stem cells induced by Wnt-driven regeneration. Nature. 2013;494:247.
Lee J-H, Bhang DH, Beede A, Huang TL, Stripp BR, Bloch KD, et al. Lung stem cell differentiation in mice directed by endothelial cells via a BMP4-NFATc1-thrombospondin-1 axis. Cell. 2014;156:440–55.
Gao D, Vela I, Sboner A, Iaquinta PJ, Karthaus WR, Gopalan A, et al. Organoid cultures derived from patients with advanced prostate cancer. Cell. 2014;159:176–87.
Maimets M, Rocchi C, Bron R, Pringle S, Kuipers J, Giepmans BNG, et al. Long-Term In Vitro Expansion of Salivary Gland Stem Cells Driven by Wnt Signals. Stem Cell Rep. 2016;6:150–62.
Boretto M, Cox B, Noben M, Hendriks N, Fassbender A, Roose H, et al. Development of organoids from mouse and human endometrium showing endometrial epithelium physiology and long-term expandability. Development. 2017;144:1775–86.
Sato T, Stange DE, Ferrante M, Vries RGJ, Van Es JH, Van Den Brink S, et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology. 2011;141:1762–72.
Van De Wetering M, Francies HE, Francis JM, Bounova G, Iorio F, Pronk A, et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell. 2015;161:933–45.
Boj SF, Hwang CIl, Baker LA, Chio IIC, Engle DD, Corbo V, et al. Organoid models of human and mouse ductal pancreatic cancer. Cell. 2015;160:324–38.
Seidlitz T, Merker SR, Rothe A, Zakrzewski F, von Neubeck C, Grützmann K, et al. Human gastric cancer modelling using organoids. Gut. 2019;68:207–17.
Vlachogiannis G, Hedayat S, Vatsiou A, Jamin Y, Fernández-mateos J, Khan K et al. Patient-derived organoids model treatment response of metastatic gastrointestinal cancers. Science. 2018;926:920–6.
Nanki K, Toshimitsu K, Takano A, Fujii M, Shimokawa M, Ohta Y, et al. Divergent Routes toward Wnt and R-spondin Niche Independency during Human Gastric Carcinogenesis. Cell. 2018;174:856–69.
Yan HHN, Siu HC, Law S, Ho SL, Yue SSK, Tsui WY, et al. A Comprehensive Human Gastric Cancer Organoid Biobank Captures Tumor Subtype Heterogeneity and Enables Therapeutic Screening. Cell Stem Cell. 2018;23:882–97.
Fujii M, Matano M, Toshimitsu K, Takano A, Mikami Y, Nishikori S, et al. Human Intestinal Organoids Maintain Self-Renewal Capacity and Cellular Diversity in Niche-Inspired Culture Condition. Cell Stem Cell. 2018;23:787–93.e6.
Huch M, Gehart H, van Boxtel R, Hamer K, Blokzijl F, Verstegen MMA, et al. Long-Term Culture of Genome-Stable Bipotent Stem Cells from Adult Human Liver. Cell. 2015;160:299–312.
Georgakopoulos N, Prior N, Angres B, Mastrogiovanni G, Cagan A, Harrison D, et al. Long-term expansion, genomic stability and in vivo safety of adult human pancreas organoids. BMC Dev Biol. 2020;20:4.
Schwank G, Andersson-Rolf A, Koo B-K, Sasaki N, Clevers H. Generation of BAC Transgenic Epithelial Organoids. PLoS ONE. 2013;8:1–6.
Matano M, Date S, Shimokawa M, Takano A, Fujii M, Ohta Y, et al. Modeling colorectal cancer using CRISPR-Cas9–mediated engineering of human intestinal organoids. Nat Med. 2015;21:256–62.
Li X, Nadauld L, Ootani A, Corney DC, Pai RK, Gevaert O, et al. Oncogenic transformation of diverse gastrointestinal tissues in primary organoid culture. Nat Med. 2014;20:769–77.
Andersson-Rolf A, Mustata RC, Merenda A, Kim J, Perera S, Grego T, et al. One-step generation of conditional and reversible gene knockouts. Nat Methods. 2017;14::287–9. https://doi.org/10.1038/nmeth.4156.
Schwank G, Koo B-K, Sasselli V, Dekkers JF, Heo I, Demircan T, et al. Functional Repair of CFTR by CRISPR/Cas9 in Intestinal Stem Cell Organoids of Cystic Fibrosis Patients. Cell Stem Cell. 2013;13:653–8.
Drost J, van Jaarsveld RH, Ponsioen B, Zimberlin C, van Boxtel R, Buijs A, et al. Sequential cancer mutations in cultured human intestinal stem cells. Nature. 2015;521:43–47.
Andersson-Rolf A, Fink J, Mustata RC, Koo B-K. A video protocol of retroviral infection in primary intestinal organoid culture. J Vis Exp. 2014;90:e51765.
Koo B-K, Stange DE, Sato T, Karthaus W, Farin HF, Huch M, et al. Controlled gene expression in primary Lgr5 organoid cultures. Nat Methods. 2012;9:81–3.
Li VSW, Ng SS, Boersema PJ, Low TY, Karthaus WR, Gerlach JP, et al. Wnt signaling through inhibition of β-catenin degradation in an intact Axin1 complex. Cell. 2012;149:1245–56.
Wroblewski LE, Piazuelo MB, Chaturvedi R, Schumacher M, Aihara E, Feng R, et al. Helicobacter pylori targets cancer-associated apical-junctional constituents in gastroids and gastric epithelial cells. Gut. 2015;64:720–30.
Schlaermann P, Toelle B, Berger H, Schmidt SC, Glanemann M, Ordemann J, et al. A novel human gastric primary cell culture system for modelling Helicobacter pylori infection in vitro. Gut. 2016;65:202–13.
Bartfeld S, Bayram T, Van De Wetering M, Huch M, Begthel H, Kujala P, et al. In vitro expansion of human gastric epithelial stem cells and their responses to bacterial infection. Gastroenterology. 2015;148:126–36.
Fatehullah A, Tan SH, Barker N. Organoids as an in vitro model of human development and disease. Nat Cell Biol. 2016;18:246–54.
Gifford GB, Demitrack ES, Keeley TM, Tam A, La Cunza N, Dedhia PH, et al. Notch1 and Notch2 receptors regulate mouse and human gastric antral epithelial cell homoeostasis. Gut. 2017;66:1001–11.
McCracken KW, Catá EM, Crawford CM, Sinagoga KL, Schumacher M, Rockich BE, et al. Modelling human development and disease in pluripotent stem-cell-derived gastric organoids. Nature. 2014;516:400–4.
McCracken KW, Aihara E, Martin B, Crawford CM, Broda T, Treguier J, et al. Wnt/β-catenin promotes gastric fundus specification in mice and humans. Nature. 2017;541:182–7.
Broda TR, McCracken KW, Wells JM. Generation of human antral and fundic gastric organoids from pluripotent stem cells. Nat Protoc. 2019;14:28–50.
Noguchi TK, Ninomiya N, Sekine M, Komazaki S, Wang P-C, Asashima M, et al. Generation of stomach tissue from mouse embryonic stem cells. Nat Cell Biol. 2015;17:984–93.
Schumacher MA, Aihara E, Feng R, Engevik A, Shroyer NF, Ottemann KM, et al. The use of murine-derived fundic organoids in studies of gastric physiology. J Physiol. 2015;593:1809–27.
Eicher AK, Berns HM, Wells JM. Translating Developmental Principles to Generate Human Gastric Organoids. Cmgh. 2018;5:353–63.
Pompaiah M, Bartfeld S. Molecular Pathogenesis and Signal Transduction by Helicobacter pylori. Cell Mic. 2017;400:149–68.
Pompaiah M, Bartfeld S. Gastric Organoids: an Emerging Model System to Study Helicobacter pylori Pathogenesis. In: Tegtmeyer N, Backert S, (eds). Molecular Pathogenesis and Signal Transduction by Helicobacter pylori. Cham: Springer International Publishing; 2017. p. 149–68.
Fujii M, Clevers H, Sato T. Modeling Human Digestive Diseases With CRISPR-Cas9–Modified Organoids. Gastroenterology. 2019;156:562–76.
Wallaschek N, Niklas C, Pompaiah M, Wiegering A, Germer C-T, Kircher S, et al. Establishing Pure Cancer Organoid Cultures: Identification, Selection and Verification of Cancer Phenotypes and Genotypes. J Mol Biol. 2019;431:2884–93.
Roerink SF, Sasaki N, Lee-Six H, Young MD, Alexandrov LB, Behjati S, et al. Intra-tumour diversification in colorectal cancer at the single-cell level. Nature. 2018;556:457–62.
Fujii M, Shimokawa M, Date S, Takano A, Matano M, Nanki K, et al. A colorectal tumor organoid library demonstrates progressive loss of niche factor requirements during tumorigenesis. Cell Stem Cell. 2016;18:827–38.
Proffitt KD, Madan B, Ke Z, Pendharkar V, Ding L, Lee MA, et al. Pharmacological Inhibition of the Wnt Acyltransferase PORCN Prevents Growth of WNT-Driven Mammary Cancer. Cancer Res. 2013;73:502–7.
de Lau W, Barker N, Low TY, Koo B-K, Li VSW, Teunissen H, et al. Lgr5 homologues associate with Wnt receptors and mediate R-spondin signalling. Nature. 2011;476:293–7.
Wang K, Yuen ST, Xu J, Lee SP, Yan HHN, Shi ST, et al. Whole-genome sequencing and comprehensive molecular profiling identify new driver mutations in gastric cancer. Nat Genet. 2014;46:573–82.
Du Y, Li X, Niu Q, Mo X, Qui M, Ma T et al. Development of a miniaturized 3D organoid culture platform for ultra-high throughput screening. J Mol Cell Biol. 2020. https://doi.org/10.1093/jmcb/mjaa036.
Schnalzger TE, de Groot MHP, Zhang C, Mosa MH, Michels BE, Röder J, et al. 3D model for CAR-mediated cytotoxicity using patient-derived colorectal cancer organoids. EMBO J. 2019;38:e100928.
Dijkstra KK, Cattaneo CM, Weeber F, Chalabi M, van de Haar J, Fanchi LF, et al. Generation of Tumor-Reactive T Cells by Co-culture of Peripheral Blood Lymphocytes and Tumor Organoids. Cell. 2018;174:1586–98.e12.
Werner K, Weitz J, Stange DE. Organoids as model systems for gastrointestinal diseases: tissue engineering meets genetic engineering. Curr Pathobiol Rep. 2016;4:1–9.
Drost J, Clevers H. Translational applications of adult stem cell-derived organoids. Development. 2017;144:968–75.
Jin M-Z, Han R-R, Qiu G-Z, Ju X-C, Lou G, Jin W-L. Organoids: an intermediate modeling platform in precision oncology. Cancer Lett. 2018;414:174–80.
Artegiani B, Clevers H. Use and application of 3D-organoid technology. Hum Mol Genet. 2018;27:R99–107.
Merker SR, Weitz J, Stange DE. Gastrointestinal organoids: how they gut it out. Dev Biol. 2016;420:239–50.
Sato T, Clevers H. Growing self-organizing mini-guts from a single intestinal stem cell: mechanism and applications. Science (80-). 2013;340:1190–4.
Dedhia PH, Bertaux-Skeirik N, Zavros Y, Spence JR. Organoid models of human gastrointestinal development and disease. Gastroenterology. 2016;150:1098–112.
Lancaster MA, Knoblich JA. Organogenesis in a dish: modeling development and disease using organoid technologies. Science (80-). 2014;345:1247125. https://doi.org/10.1126/science.1247125
Wang X. Stem cells in tissues, organoids, and cancers. Cell Mol Life Sci. 2019;76:4043–70.
Schutgens F, Clevers H. Human organoids: tools for understanding biology and treating diseases. Annu Rev Pathol Mech Dis. 2020;15:211–34.
Dutta D, Heo I, Clevers H. Disease Modeling in Stem Cell-Derived 3D Organoid Systems. Trends Mol Med. 2017;23:393–410.
Lin M, Gao M, Cavnar MJ, Kim J. Utilizing gastric cancer organoids to assess tumor biology and personalize medicine. World J Gastrointest Oncol. 2019;11:509–17.
Aberle MR, Burkhart RA, Tiriac H, Olde Damink SWM, Dejong CHC, Tuveson DA, et al. Patient-derived organoid models help define personalized management of gastrointestinal cancer. Br J Surg. 2018;105:e48–60.
Weeber F, Ooft SN, Dijkstra KK, Voest EE. Tumor Organoids as a Pre-clinical Cancer Model for Drug Discovery. Cell Chem Biol. 2017;24:1092–100.
Tuveson D, Clevers H. Cancer modeling meets human organoid technology. Science (80-). 2019;364:952–5.
Bleijs M, van de Wetering M, Clevers H, Drost J. Xenograft and organoid model systems in cancer research. EMBO J. 2019;38:e101654.
Drost J, Clevers H. Organoids in cancer research. Nat Rev Cancer. 2018;18:407–18.
Acknowledgements
This work was supported by the Deutsche Krebshilfe (#111350), the Sander Stiftung (#2014.104.1), the Hector Stiftung (#M65.2) and the European Union (ERC #639050) to DES.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: Due to a retrospective open access order.
Edited by F Pentimalli
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Seidlitz, T., Koo, BK. & Stange, D.E. Gastric organoids—an in vitro model system for the study of gastric development and road to personalized medicine. Cell Death Differ 28, 68–83 (2021). https://doi.org/10.1038/s41418-020-00662-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-020-00662-2
- Springer Nature Limited
This article is cited by
-
The role of organoids in cancer research
Experimental Hematology & Oncology (2023)
-
Advances towards the use of gastrointestinal tumor patient-derived organoids as a therapeutic decision-making tool
Biological Research (2023)
-
Exploring the promising potential of induced pluripotent stem cells in cancer research and therapy
Molecular Cancer (2023)
-
Single-cell sequencing of ascites fluid illustrates heterogeneity and therapy-induced evolution during gastric cancer peritoneal metastasis
Nature Communications (2023)
-
Deterministic evolution and stringent selection during preneoplasia
Nature (2023)