Abstract
Zinc finger proteins (ZFPs) are the largest transcription factor family in mammals. About one-third of ZFPs are Krüppel-associated box domain (KRAB)-ZFPs and involved in the regulation of cell differentiation/proliferation/apoptosis and neoplastic transformation. We recently identified ZNF382 as a novel KRAB-ZFP epigenetically inactivated in multiple cancers due to frequent promoter CpG methylation. However, its epigenetic alterations, biological functions/mechanism and clinical significance in oesophageal squamous cell carcinoma (ESCC) are still unknown. Here, we demonstrate that ZNF382 expression was suppressed in ESCC due to aberrant promoter methylation, but highly expressed in normal oesophagus tissues. ZNF382 promoter methylation is correlated with ESCC differentiation levels. Restoration of ZNF382 expression in silenced ESCC cells suppressed tumour cell proliferation and metastasis through inducing cell apoptosis. Importantly, ZNF382 suppressed Wnt/β-catenin signalling and downstream target gene expression, likely through binding directly to FZD1 and DVL2 promoters. In summary, our findings demonstrate that ZNF382 functions as a bona fide tumour suppressor inhibiting ESCC pathogenesis through inhibiting the Wnt/β-catenin signalling pathway.
Similar content being viewed by others
Introduction
Oesophageal cancer is the eighth most common cancer with the sixth highest cancer mortality rate worldwide and a very low 5-year survival of 15%–25%1,2. Oesophageal squamous cell carcinoma (ESCC) comprises 90% of oesophageal cancer as the predominant type in China; the remaining cases are oesophageal adenocarcinoma (EAC)1,3. The incidence of oesophageal cancer varies geographically with the highest in a belt extending through central Asian to North-Central China1,2,4,5, reaching an incidence of >100/100,000 population annually2,6,7. The major risk factors for ESCC are poor nutrition, tobacco and alcohol consumption, whereas diet, obesity and gastroesophageal reflux disease are also EAC risk factors4,8,9. The incidence of ESCC has been decreasing a bit, but its 5-year survival remains poor; therefore a better understanding of ESCC pathogenesis is urgently needed3,6,10,11,12,13. Substantial evidences have shown that epigenetic modifications, primarily aberrant promoter CpG methylation of tumour suppressor genes (TSGs), consistently contribute to malignant transformation4,14,15. Alterations of promoter hypermethylation occur frequently in ESCC and involve multiple genes required for ESCC carcinogenesis14,16. Therefore, it is crucial to investigate the epigenetic abnormalities in ESCC.
Zinc finger proteins (ZFPs) comprise the largest group of transcription factors. Their zinc finger domains bind to gene promoters to activate or repress gene expression17. Nearly one-third of mammalian ZFPs contain a highly conserved Krüppel-associated box (KRAB) motif, which contributes to transcriptional repression by recruitment of histone deacetylase (HDAC) complexes18,19,20,21. We recently identified a novel ZFP ZNF382, the human homologue of rat tumour suppressor KS1 located at 19q13.12, that functions as a transcriptional repressor by binding to the KAP-1 co-repressor protein22. ZNF382 is widely expressed in normal tissues but reduced or silenced in multiple carcinomas due to aberrant promoter CpG methylation22,23,24. Although evidence has indicated that ectopic expression of ZNF382 suppresses tumour cell proliferation and promotes apoptosis, its biological functions and underlying mechanisms in ESCC pathogenesis remain to be investigated24.
Wnt signalling includes canonical Wnt/β-catenin pathway and noncanonical pathway and is an oncogenic activation event in many cancers, especially in gastrointestinal cancers. Numerous studies have reported that Wnt signalling is associated with the initiation and progression of human ESCC. Furthermore, comprehensive genomic analysis of ESCC revealed that altered genes in Wnt pathway were identified in ~86% of ESCC cases, adding to our understanding of pathogenenic role of Wnt pathway in ESCC tumorigenesis25,26. In this study, we investigated the methylation status of ZNF382 in primary ESCC and its biological functions in silenced ESCC cell lines. We further explored the mechanism of ZNF382 on tumour suppression of ESCC.
Results
Promoter methylation leads to ZNF382 downregulation in ESCC
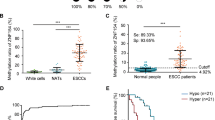
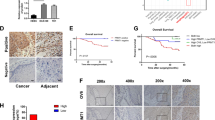
ZNF382 downregulation in some carcinomas has been previously reported24. We thus further analysed ZNF382 mRNA expression via qRT-PCR in 15 cases of ESCC and paired adjacent non-cancerous tissues, and we found that ZNF382 expression in the ESCC samples was significantly reduced compared with paired adjacent non-cancerous tissues (Fig. 1a). We then evaluated ZNF382 expression in a panel of ESCC cell lines and normal oesophagus tissues by real-time PCR (Fig. 1b) and qRT-PCR (Supplemental Fig. 1). ZNF382 was highly expressed in normal oesophagus tissues, but almost completely silenced in KYSE150, KYSE410 and KYSE510 cells. We also examined ZNF382 protein expression in ESCC and paired adjacent non-cancer tissues by immunohistochemistry (IHC). ZNF382 protein expression was significantly weaker in ESCC than in adjacent non-cancerous tissues and primarily localised in cell nuclei (Fig. 1c). In addition, ZNF382 expression was analysed using the online Gene Expression across Normal and Tumour (GENT) tissue database (http://medical-genome.kribb.re.kr/GENT/search/search.php), and ZNF382 was also found to be downregulated in ESCC tissues compared with normal oesophageal epithelial tissues (Fig. 1d, p= 0.0016). Furthermore, the correlation of ZNF382 expression and overall survival (OS) in oesophageal cancer patients was investigated according to the cBioPortal for Cancer Genomics (http://www.cbioportal.org/) in The Cancer Genome Atlas (TCGA) database. Higher levels of ZNF382 expression were associated with a better OS rate in oesophageal cancer patients (Fig. 1e, p= 0.028), suggesting that ZNF382 may be an independent prognostic factor in oesophageal cancer.
a ZNF382 expression in primary ESCC (n = 15) and paired adjacent non-cancerous tissues (n = 15) by qRT-PCR. Mean ± SD, p < 0.001. b ZNF382 expression in representative ESCC cells compared with normal oesophageal tissues was assayed by RT-PCR. c Representative IHC for ZNF382 in ESCC tissues (right) and paired adjacent non-cancerous tissues (left). d ZNF382 expression in ESCC tissues (n = 40) and normal oesophageal tissues (n = 9) from online Gene Expression across Normal and Tumour Tissues (GENT) database (http://medical-genome.kribb.re.kr/GENT/search/search.php). e Kaplan–Meier cumulative survival curves of ZNF382 expression in oesophageal cancer patients are presented to explore the prognostic implication of ZNF382. Data were extracted from the Cancer Genome Analysis Database, and p value indicated statistical significance. f Restoration of ZNF382 expression with 5-aza-2′-deoxycytidine (Aza) and trichostatin A (TSA) treatments in ESCC cells. Demethylation was measured by MSP. M methylated, U unmethylated. g Methylation status of ZNF382 in oesophageal cancer tissues (n = 188) and normal oesophagus tissues (n = 10) from human disease methylation database (http://www.biobigdata.com/diseasemeth/). h–l ZNF382 methylation in primary ESCC tissues (n = 114) and normal tissues (n = 3), measured by methylation-specific PCR (MSP). M methylated, U unmethylated
Abnormal promoter methylation is a critical mechanism responsible for the silencing of TSGs in tumourigenesis27. Our previous research demonstrated that ZNF382 is frequently methylated in solid cancers. To observe whether ZNF382 expression is regulated by promoter methylation, we treated KYSE150, KYSE410 and KYSE510 cells with the demethylation drug Aza combined with the HDAC inhibitor trichostatin A (TSA), and the synergistic treatments significantly restored ZNF382 expression, together with increased unmethylated alleles and decreased methylated alleles (Fig. 1f). In addition, ZNF382 promoter methylation in primary oesophageal cancer tissues and normal oesophagus tissues from a human disease methylation database (http://www.biobigdata.com/diseasemeth/) was analysed. ZNF382 methylation was far more prevalent in oesophageal cancer tissues than in normal oesophagus tissues (Fig. 1g, p= 0.0023). Thus, the overall results indicate that promoter methylation may be a primary mechanism of ZNF382 silencing in ESCC cell lines.
ZNF382 methylation in ESCC is correlated with clinicopathological features
To further investigate ZNF382 promoter methylation status in ESCC, methylation-specific PCR (MSP) was performed in primary ESCC and the normal oesophagus tissues. The data showed that the ZNF382 promoter was hypermethylated in 87% (99/114) of ESCC tissues and in 33% (1/3) of normal oesophageal tissues, suggesting the frequent methylation of ZNF382 in ESCC (Fig. 1h–i and Table 1). Next, we analysed the relationship between ZNF382 methylation status and clinical features, and a significant association was found with tumour differentiation (Table 2) (χ2 test, p= 0.029).
Ectopic expression of ZNF382 suppresses ESCC cell proliferation
ZNF382 ectopic expression in ZNF382-stably transfected KYSE150, KYSE410 and KYSE510 cells was verified by RT-PCR and western blot, respectively (Fig. 2a, b). To investigate the effect of ZNF382 on ESCC cell proliferation, cell viability and colony formation assays were performed. ZNF382-transfected cells exhibited lower viability at 24, 48 and 72 h (Fig. 2c) and 60% to 80% less colony formation than controls (Fig. 2d). In addition, we also carried out Edu cell proliferation assays, and a significant decrease in the percentage of Edu-positive cells was observed after transfection of ESCC cells with ZNF382 (Fig. 2e and Supplemental Figure 2).
a, b Ectopic expression of ZNF382 in ESCC cells was measured by RT-PCR and western blot assays, respectively. c Cell viabilities were evaluated at 24, 48 and 72 h after transfection with ZNF382 in KYSE150, KYSE410 and KYSE510 cells. Mean ± SD, ***p < 0.001. d Representative images of showing the colony formation ability of in vector- and of ZNF382- transfected KYSE150, KYSE410 and KYSE510 cells (left panel). The colony formation rates are shown as the mean ± SD (right panel) from three independent experiments (***p< 0.001). e Edu cell proliferation assays of KYSE150, KYSE410 and KYSE510 cells. Scale bars: 800 μm. Data are from three independent assays and are shown as the mean ± SD, ***p < 0.001. f The cell cycle distribution was measured in vector- and ZNF382-stably transfected KYSE150, KYSE410 and KYSE510 cells by flow cytometry analysis. Representative cell cycle distribution plots and histograms of cell cycle alterations are shown (**p <0.01, *p< 0.05). g Western blot assay of Cdc25C, Cdc2 and CyclinB1 protein expression levels. h The percentages of apoptotic cells in KYSE150, KYSE410 and KYSE510 cells with ZNF382 ectopic expression were evaluated. Cell apoptosis alterations were revealed by histograms (***p < 0.001). i The expression of cleaved-caspase8, cleaved-caspase3 and cleaved PARP was measured by western blot assay in ZNF382-transfected ESCC cells. All experiments were repeated in triplicate and are shown as the mean ± SD
ZNF382 induces G2/M cell cycle arrest and apoptosis in ESCC cells
A cell cycle assay was performed to determine how ZNF382 affects cell proliferation. Compared with controls, ZNF382 significantly increased KYSE150, KYSE410 and KYSE510 cells in the G2/M phase after ZNF382 transfections (Fig. 2f). This effect was accompanied by Cdc25C, Cdc2 and CyclinB1 inhibitions (Fig. 2g). Flow cytometry analysis of cell apoptosis was performed to evaluate the mechanisms of ZNF382 suppression of tumour cell growth. As shown in Figure 2h, the percentage of apoptotic cells was remarkably increased after ZNF382- transfection in ESCC cells. Caspases were involved in the proteolytic cascade, which further induced apoptosis. Western blots showed that ZNF382 significantly increased the cleavage of caspase8, caspase3 and PARP in ESCC cells compared with vector controls (Fig. 2i).
ZNF382 inhibits ESCC cell migration and invasion
To investigate the effect of ZNF382 on cell migration, wound healing assays were carried out. ZNF382-stably transfected ESCC cells migrated into the wounded areas slower than the control cells at 24 h (Fig. 3a). Moreover, the effect of ZNF382 on cell migration and invasion were further analysed by transwell assay. The results showed that ectopic expression of ZNF382 significantly decreased the number of migrating (Fig. 3b, c) and invading cells (Fig. 3d, e) compared with controls. The effect of ZNF382 on ESCC cell metastasis was mediated by E-cadherin (CDH1) upregulation and vimentin (VIM), N-cadherin (CDH2), SNAIL1 and SLUG downregulation (Fig. 3f).
a The cell migration abilities of ESCC cells were evaluated by wound healing assays. Photographs were captured at 0, 12 and 24 h. The representative wound healing ratio is shown. Scale bars: 400 μm. Mean ± SD, **p < 0.01, *p < 0.05. The cellular migration (b, c) and invasion (d, e) abilities of ESCC cells upon ectopic expression of ZNF382 were measured by transwell assays with or without Matrigel. Representative images were photographed following fixation and staining. Assays were performed in triplicate, ***p< 0.001. Scale bars: 100 μm. f Expression of representative EMT markers by western blot assay
Knockdown of ZNF382 promotes cell growth and induces metastasis
To further validate the suppression function of ZNF382 in ESCC, we investigated the effect of ZNF382 through knockdown of ZNF382 in ZNF382-expressing ESCC cell line EC1 by siRNA transfection. Knockdown efficiency was detected by western blot (Fig. 4a). Knockdown of ZNF382 markedly increased EC1 cell viability, according to CCK8 assays compared with cells transfected with control siRNA (Fig. 4b). To evaluate the effect of ZNF382 on cell cycle regulation, flow cytometry assays were employed. The distribution of EC1 cells in G2/M phase was significantly reduced after knocking down ZNF382 (Fig. 4c). Moreover, we observed enhanced cell migration and invasion after ZNF382 knocking down. Compared with control, the number of migrated and invading EC1 cells were dramatically increased after knockdown of ZNF382 (Fig. 4d). These knockdown data confirmed that ZNF382 functions as a tumour suppressor in ESCC cells.
a Evaluation of ZNF382 knockdown in EC1 cells after transfection with ZNF382 siRNA and si-Scramble by western blot. b Knockdown of ZNF382 on cell viability was measured by cell viability assay. Each experiment was repeated three times. **p <0.01. c Cell cycle distribution after ZNF382 knockdown in EC1 cells. Each experiment was repeated three times. **p <0.01. d Transwell assay showed cell migration and invasion of EC1 cells after ZNF382 knockdown. Scale bars: 200 μm. Each experiment was repeated three times. *p< 0.05
Identification of differentially expressed genes regulated by ZNF382 in ESCC
To better investigate the underlying mechanism of ZNF382 in ESCC, RNA-Seq was performed in ZNF382-stably transfected KYSE150 cells. The entire distribution of differentially expressed genes was shown by a volcano plot, and a total of 3276 upregulated and 3557 downregulated genes were observed (Fig. 5a, *p < 0.05). We next performed the hierarchical cluster analysis of 42 differentially expressed genes selected on the basis of their expression patterns, and two major groups were classified (Fig. 5b). To further confirm the differential expression of genes identified by RNA-Seq analysis, qRT-PCR was performed to validate the gene expression levels, and 15 differentially expressed genes modulated by ZNF382 were verified in both KYSE150 and KYSE410 cells (Fig. 5c–e).
a The entire distribution of differentially expressed genes in ZNF382-stably transfected KYSE150 cell is shown by a volcano plot. b The hierarchical cluster analysis of selected differentially expressed genes is shown as a heat map. c Ectopic expression of ZNF382 in KYSE150 and KYSE410 cells. Mean ± SD, ***p< 0.001. d, e Validation of differentially expressed genes in KYSE150 and KYSE410 cells ectopically expressing ZNF382 by qRT-PCR. Assays were repeated in triplicate. Mean ± SD, ***p< 0.001, **p< 0.01. f QRT-PCR was used to evaluate FZD1 and DVL2 expression in KYSE150 and KYSE410 cells. Mean ± SD, **p< 0.01, *p< 0.05. Downstream target genes of β-catenin in KYSE150 and KYSE410 cells ectopically expressing ZNF382 were assayed by RT-PCR (g) and western blot (h). GAPDH and β-actin were used as internal controls, respectively. All assays were performed in triplicate. i The effect of ZNF382 on Wnt/β-catenin signalling was determined by TCF/LEF luciferase reporter activity assays. All the experiments were performed in triplicate. Mean ± SD, ***p< 0.001
Moreover, these significantly regulated genes identified in both KYSE150 and KYSE410 cells were involved in cell metastasis (SLUG, SNAIL and SMAD3), cell growth (CSF1 and VEGFA), stress-induced protector (SESN3) and stem cell regulation (SOX2 and STAT5B), adding to our understanding of the possible mechanisms of ZNF382 tumour suppression in ESCC.
ZNF382 inhibits Wnt/β-catenin signalling through binding to DVL2 and FZD1 promoters
ChIP-Seq data of HEK293 cells stably expressing eGFP-ZNF382 fusion protein were downloaded from the Gene Expression Omnibus (GEO) database. Enriched peaks located in the transcriptional start sites of FZD1 and DVL2 genes were obtained, suggesting that FZD1 and DVL2 are the direct targets of ZNF382 (Supplemental Fig. 3). List of Chip-Seq peaks for ZNF382 targets were listed in Table S2.
Thus, we hypothesised that ZNF382 likely plays a role in Wnt/β-catenin signalling through directly binding to FZD1 and DVL2 promoters. Reduced FZD1 and DVL2 accompanied by downstream target genes such as C-MYC, CCND1, MMP2 and MMP3 expression levels were observed at both the transcriptional (Fig. 5f–g) and protein levels (Fig. 5h). Then, we examined the expression of active β-catenin via western blot. As shown in Figure 5h, the expression of total β-catenin remained unchanged, while the expression of active β-catenin decreased. In addition, inhibition of TOPflash luciferase activity more fully demonstrated that ZNF382 antagonised Wnt/β-catenin signalling by decreasing active β-catenin levels (Fig. 5i). These results conclusively suggest that ZNF382 exerts tumour suppression function through antagonising Wnt/β-catenin signalling by directly binding to DVL2 and FZD1 promoters.
Pathway enrichments and protein–protein interaction network
Gene Ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichments based on RNA-Seq data were further analysed and are shown in Figure 6a, b. The Wnt pathway was significantly enriched in both GO and KEGG pathways and other processes, such as apoptosis induction, negative regulation of RNA polymerase II transcription and regulation of stem cell pluripotency were also included. In addition, the protein–protein interaction (PPI) network showed interactions within the network of genes transcribed by ZNF382, which involved cell development, cell cycle regulation and metastasis (Fig. 6c).
a GO biological pathway analysis. * indicates significantly enriched pathways. b KEGG pathway categories of differentially expressed genes. The rich factor represents the proportion of differentially expressed genes within a specific term, and the size of the point represents the number of related differentially expressed genes. The q-value is the calibrated p value. c The interactions within the network of genes transcribed by ZNF382 are shown by the PPI network. Genes in the network are shown as nodes, and edges represent the interactions of genes. d A schematic diagram of the mechanism by which ZNF382 antagonises Wnt/β-catenin signalling in ESCC
Discussion
ZFPs comprising 742 members and involving 423 genes make up the largest transcription factor family in mammals28,29. The KRAB domain, located in the amino terminal of ZFP, exerts transcriptional repressor functions16,19,21,30 by sequence-specific DNA methylation29,31,32. Due to the potent repressor domain, ZFPs play an essential role in cell metabolism, differentiation and apoptosis21,33,34. We investigated the novel 19q13 KRAB-ZFP ZNF382, which is orthologous to rat KS1, and found that ZNF382 was highly expressed in normal tissues but frequently silenced in multiple carcinomas due to promoter CpG methylation22,24,29,35. ZNF382 was significantly reduced or silenced in ESCC tissues compared with normal oesophageal tissues. The MSP assay also revealed that ZNF382 promoter was methylated in 87% (99/114) of ESCC cases, suggesting a high frequency of ZNF382 promoter methylation in ESCC. In addition, data from TCGA database showed that higher level of ZNF382 heralded a better OS in ESCC patients, indicating that ZNF382 might be an independent prognostic factor for ESCC patients.
Here, we demonstrated the tumour suppressive effects of ZNF382 in ESCC cells. Our results indicate that inhibition of cell proliferation by ectopic expression of ZNF382 in ESCC cells might be through G2/M cell cycle arrest and apoptosis, accompanied by reduced of Cdc25C, Cdc2 and CyclinB1 checkpoint regulators and induced cleavage of apoptotic markers caspase8, caspase3 and PARP. The effect of ZNF382 on cell metastasis was further evaluated by scratch and transwell assays. The decreased migration and invasion abilities of ESCC cells, with upregulation of epithelial marker CDH1 and downregulation of VIM, CDH2, SNAIL1 and SLUG resulted from negative regulation by ZNF382, which was also reflected in the epithelial-mesenchymal transition (EMT) of ESCC cells.
RNA-Seq was additionally performed to explore the underlying mechanism of ZNF382 in ESCC, and a total of 3276 upregulated and 3557 downregulated genes were identified. Subsequently, 42 differentially expressed genes were selected among which 29 DEGs exhibited a fold change >1.5, for further hierarchical cluster analysis. Validation by qRT-PCR revealed 15 differentially expressed genes modulated by ZNF382 that were involved in cell metastasis (SLUG, SNAIL and SMAD3), cell growth (CSF1 and VEGFA), stress-induced protection (SESN3) and stem cell regulation (SOX2 and STAT5B) were observed in both KYSE150 and KYSE410 cells.
Wnt signalling pathway participates in the carcinogenesis of multiple types of tumours, especially gastrointestinal cancers. Numerous articles have reported that Wnt pathway alterations are involved in the development of oesophageal squamous carcinomas. Wnt signalling mainly includes canonical and noncanonical pathways. The canonical pathway (WnT/β-catenin signalling) is the most frequently reported pathway and has been associated with the regulation of tumour growth, metastasis, and transformation.
In the present investigation, we identified two target genes, FZD1 and DVL2, from ChIP-Seq of HEK293 cells stably transfected with ZNF382. Both of these genes are critical elements in the Wnt pathway. ZNF382 negatively regulates the Wnt/β-catenin signalling by directly binding to FZD1 and DVL2 promoters, inhibiting β-catenin activation and T cell factor/lymphoid-enhancer factor (TCF/LEF)-dependent luciferase reporter activity, subsequently downregulating the downstream target genes of β-catenin signalling including C-MYC, CYCLIND1, MMP2, MMP3 and MMP9 expressions (Fig. 6d).
It is worth mentioning that amplification of CYCLIND1, an essential downstream target gene of β-catenin, has been detected in ESCC samples1. Overexpression of CYCLIND1 is common in ESCC cell lines and is a poor prognostic marker of poor ESCC outcomes, due to its strong association with increased lymph nodes or distant metastasis, high proliferation rates and poor responses to chemotherapy1. Thus, evaluation of CYCLIND1 status is critical for ESCC patients. Here, we found that ZNF382 negatively modulates Wnt/β-catenin signalling, further inhibiting target gene CYCLIND1 expression. These findings indirectly illustrate that ZNF382, as a TSG, could be a promising prognostic predictor for ESCC patient outcomes.
Conclusions
In conclusion, ZNF382 was frequently inactivated by promoter CpG methylation, and suppressed ESCC carcinogenesis through directly regulating Wnt/β-catenin signalling.
Materials and methods
Primary tumour samples and cell lines
Multiple ESCC cell lines including KYSE150, KYSE410 and KYSE510 were used24. The cells were cultured in RPMI 1640 medium (Gibco-BRL, Karlsruhe, Germany), supplemented with 10% foetal bovine serum (Gibco-BRL), at 37 °C in 5% CO2 atmosphere. Primary ESCC tissues, adjacent non-cancerous tissues and normal oesophageal tissues were obtained from patients who underwent surgery at the Department of Cardiothoracic Surgery of the First Affiliated Hospital of Chongqing Medical University. All samples were stored at −80 °C until being evaluated by pathologists at the Chongqing Key Laboratory of Molecular Oncology and Epigenetic of the First Affiliated Hospital of Chongqing Medical University. All participants provided written consent before enrolment, and the study was approved by the Ethics Committees of the First Affiliated Hospital of Chongqing Medical University.
RNA and DNA extraction
Total RNA was isolated from cells and tissues with the TRIzol reagent (Invitrogen, Carlsbad, CA, USA). Genomic DNA was extracted from cells and tissues with a QIAamp DNA Mini Kit (Qiagen, Hilden, Germany) following the manufacturers’ protocols36. RNA and DNA concentrations were measured with a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, Waltham, MA), and sample quality was determined by gel electrophoresis.
Reverse transcription, semi-quantitative PCR (RT-PCR) and quantitative PCR (qRT-PCR)
Reverse transcription and semi-quantitative PCR (RT-PCR) were performed as previously described using Go-Taq polymerase (Promega, Madison, WI)37. RT-PCR was performed using a final volume of 10 μL reaction mixture containing 2 μL cDNA. GAPDH served as an internal control. The amplified PCR products were assayed on 2% agarose gels. A SYBR Green PCR Master Mix kit (Invitrogen) and an Applied Biosystems 7500 Real-Time PCR System (Applied Biosystems, Foster City, CA) were used for qRT-PCR36. The relative expression of ZNF382 was estimated with the 2(−ΔCt) method38 and all assays were performed in triplicate. The primers used are listed in Supplementary Table 1.
5-Aza-2′-deoxycytidine (Aza) and trichostatin A (TSA) treatment
Combined treatment of Aza, a DNA methyltransferase inhibitor and TSA, a HDAC inhibitor synergistically activated methylated genes. ESCC cell lines were treated with 10 mmol/L Aza (Sigma-Aldrich, Steinheim, Germany) for three days and 100 nmol/L TSA (Sigma-Aldrich) for an additional 24 h as previously described24. Cells were then used for RNA and DNA isolation.
Bisulfite treatment and methylation-specific PCR (MSP)
To evaluate the methylation status of ZNF382, bisulfite modification of DNA and MSP were performed as previously described24,39,40. In brief, bisulfite-treated DNA was amplified by MSP with methylation-specific primers. Methylated and non-methylated DNAs were used as positive and negative controls. MSP products were separated on 2% agarose gels (MBI Fermentas, Vilnius, Lithuania). The primers are listed in Supplementary Table 1.
Transfection and construction of stable cell lines
A ZNF382 expression vector (PcDNA3.1–ZNF382-FLAG) and a control vector (PcDNA3.1) were used24. Plasmids were transfected into ESCC cell lines using Lipofectamine 2000 (Invitrogen) at a final concentration of 4 μg. The transfected cells were harvested at 48 h. ZNF382-stably transfected ESCC cells were selected with G418 (Invitrogen, Gibco) for 2 weeks.
Knockdown of ZNF382
ZNF382 siRNA kits were purchased from GenePharma (Jiangsu, China). All transfections were performed using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instruction with a concentration of 50 nM siRNA. The cells were harvested for subsequent assays at 72 h after transfection.
Cell viability assay
Stably transfected KYSE150, KYSE410 and KYSE510 cells were seeded in 96-well plates and grown overnight. Cell viabilities were then evaluated with a Cell Counting Kit-8 (Beyotime Institute of Biotechnology, Jiangsu, China) at 24, 48 and 72 h, respectively. All experiments were independently repeated three times.
Colony formation assay
Stably transfected KYSE150, KYSE410 and KYSE510 cells were plated in six-well plates and cultured with various concentrations of G418 for 10–14 days. Surviving colonies with >50 cells were counted following fixation and stained with gentian violet (Beyotime). All experiments were repeated three times.
5-Ethynyl-2′-deoxyuridine (Edu) cell proliferation assay
Cell proliferation was evaluated with an Edu cell proliferation kit (RiboBio, Guangzhou, China). Edu is a thymine nucleoside analogue that replaces thymine during DNA replication, and DNA replication activity is assayed by labelling with specific Edu and Apollo fluorescent dyes. All assays were repeated three times.
Wound healing and transwell assay
Cell mobility was evaluated with a wound healing assay. Briefly, ZNF382-stably transfected KYSE150, KYSE410 and KYSE510 cells were seeded in six-well plates and grown until confluence. Linear scratch wounds were created with a pipette tip and cell migration distance was measured at various times by phase contrast microscopy (Leica DMI4000B, Milton Keynes, Bucks, UK).
Transwell chambers with an 8 μm pores (Corning Life Sciences, Bedford, MA) were used with or without Matrigel (BD Biosciences, San Jose, CA) or not to measure cell migration and invasion, respectively. Cells on the lower membrane surface were counted following fixation and staining. All assays were performed in triplicate.
Flow cytometry assay
Cell cycle arrest and apoptosis were assayed via flow cytometry as previously described37. In brief, the cells were harvested with trypsin, fixed with ice-cold 70% ethanol and stained with propidium iodide (PI) to assay the cell cycle distribution. Apoptosis was assayed with annexin V–fluorescein isothiocyanate and PI following the kit manufacturer’s protocol. All data were measured using a Cell Quest kit (BD Biosciences) and experiments were independently repeated in triplicate.
Western blot assay
Western blot assays were performed as previously described36. Briefly, whole cells lysates were prepared with lysis buffer (Beyotime) containing a protease inhibitor cocktail (Sigma-Aldrich, St. Louis, MO). Lysates were sonicated, and the protein concentration was determined using a bicinchoninic acid (BCA) assay. Aliquots of 50 µg protein were separated with 10–12% sodium dodecyl sulphate polyacrylamide gel electrophoresis and then transferred to polyvinylidene fluoride membranes (Merck Millipore, Billerica, MA). After blocking with 5% nonfat milk, the membranes were incubated at 4 °C overnight with anti-DDK (FLAG) monoclonal antibody (TA50011-100; OriGene, Rockville, MD), anti-FLAG M2 antibody (#14793; Cell Signaling Technology, Danvers, MA), cleaved caspase 3 (#9661, Cell Signaling Technology), cleaved caspase 8 (#8592; Cell Signaling Technology), cleaved PARP (#5625, Cell Signaling Technology), cyclin D1 (#1677; Epitomics, Burlingame, CA), c-Myc (#1472-1, Epitomics), active β-catenin (#05-665,Merck Millipore), total β-catenin (#2677, Cell Signaling Technology), β-actin (sc-47778; Santa Cruz Biotechnology), E-cadherin (#1702-1, Epitomics), vimentin (#2707-1, Epitomics), N-cadherin (ab98952; Abcam), Snail1 (#3897;Cell Signaling Technology), SLUG (#9585, Cell Signaling Technology), anti-Frizzled homologue 1 antibody (ab71342, Abcam), Dvl2 (sc-420081, Santa Cruz), Cdc2 (sc-8395, Santa Cruz), Cdc25C (sc-24540, Santa Cruz), or cyclinB1 (sc-4073, Santa Cruz) primary monoclonal antibodies. Membranes were visualised with Electrochemiluminescence Plus Detection Reagents (RPN2132; GE Healthcare Life Science, Buckinghamshire, UK). All assays were independently repeated three times.
Immunohistochemistry (IHC)
IHC was performed with an UltraSensitive SP Kit (Maixin-Bio, Fujian, China) following the manufacturer’s instructions as previously described41,42. Cells were incubated with a primary antibody against ZNF382 (1:50 dilution, Sigma-ldrich) overnight at 4 °C and then incubated with a secondary antibody at 37 °C for 30 min. All IHC images were captured under a microscope at ×400 magnification.
Dual-luciferase reporter assay
Briefly, cells were co-transfected with pcDNA3.1–ZNF382-FLAG or pcDNA3.1 and TCF-responsive luciferase construct TOPflash or FOPflash reporter plasmids plus Renilla luciferase reporter pRL-TK in 24-well plates. Luciferase activities were read 48 h later using a dual-luciferase reporter assay kit (Promega). All assays were independently performed three times.
RNA sequencing (RNA-Seq)
RNA-Seq was performed by the Novogene Bioinformatics Technology Company (Beijing, China). RNA integrity was assessed with an RNA Nano 6000 Assay Kit included with the Bioanalyzer 2100 system (Agilent Technologies, CA). Sequencing libraries were prepared using a NEBNext Ultra RNA LibraryPrep Kit for Illumina (New England Biolabs) following the manufacturer’s recommendations. RNA-Seq was performed with an Illumina Hiseq 2000/2500 platform. Bowtie v. 2.0.643,44,45 and TopHat v. 2.0.946,47,48 were used for reads mapping to the reference genome. Genes with an adjusted p or0.05 and identified with DESeq49,50 were considered to be differentially expressed.
Peaks calling for chromatin immunoprecipitation sequencing (ChIP-Seq)
ChIP-Seq of ZNF382 from human HEK293 cells were obtained from the Gene Expression Omnibus (GEO, https://www.ncbi.nlm.nih.gov/geo) database. The GSM2424064_ENCFF191YCR_fold_change_over_control_GRCh38.bigWig file was downloaded. We annotated those peaks by using R package “CHIPseeker”51. Subsequently, the gene region were intercepted with BEDTools52 and exported to Integrated Genomics Viewer software to map the peaks53,54.
Pathway enrichment analysis
The function enrichments of differentially expressed genes were analysed with GO classification55 and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways56 from the Database for Annotation, Visualization and Integrated Discovery (DAVID, http://david.abcc.ncifcrf.gov/)57. The level of significance was p< 0.05.
PPI analysis
The PPI analysis of target genes was based on the STRING database (https://string-db.org/) and the networks were constructed by Cytoscape58.
Statistical analysis
Data were reported as the means ± standard deviation (SD). The significance of differences between the experimental and control values were tested the with two-tailed Student's t-test. Correlations of methylation status and clinicopathological features were evaluated using the χ2 and Fisher’s exact tests. For all tests, p< 0.05 was considered statistically significant.
Availability of data and materials
Data of prognostic significance is available from Cancer Genome Atlas database (http://www.cbioportal.org/). CHIP-Seq data analysed during the present study is available from Gene Expression Omnibus database (GEO, https://www.ncbi.nlm.nih.gov/geo). PPI analysis is available from the STRING database (https://string-db.org/). Schematic model drafted with the tool scienceslides software is available at http://www.scienceslides.com/.
References
Lam, A. K. Molecular biology of esophageal squamous cell carcinoma. Crit. Rev. Oncol. Hematol. 33, 71–90 (2000).
Pennathur, A., Gibson, M. K., Jobe, B. A. & Luketich, J. D. Oesophageal carcinoma. Lancet 381, 400–412 (2013).
Rustgi, A. K. & El-Serag, H. B. Esophageal carcinoma. New Engl. J. Med. 371, 2499–2509 (2014).
Kaz, A. M. & Grady, W. M. Epigenetic biomarkers in esophageal cancer. Cancer Lett. 342, 193–199 (2014).
Kamangar, F., Dores, G. M. & Anderson, W. F. Patterns of cancer incidence, mortality, and prevalence across five continents: defining priorities to reduce cancer disparities in different geographic regions of the world. J. Clin. Oncol. 24, 2137–2150 (2006).
Zhang, M. et al. Methylation of DACT2 accelerates esophageal cancer development by activating Wnt signaling. Oncotarget 7, 17957–17969 (2016).
Chen, W. et al. Cancer statistics in China, 2015. CA: Cancer J. Clin. 66, 115–132 (2016).
Ma, K., Cao, B. & Guo, M. The detective, prognostic, and predictive value of DNA methylation in human esophageal squamous cell carcinoma. Clin. Epigenetics. 8, 43 (2016).
Garcia-Rodriguez, L. A., Lagergren, J. & Lindblad, M. Gastric acid suppression and risk of oesophageal and gastric adenocarcinoma: a nested case control study in the UK. Gut 55, 1538–1544 (2006).
Chen, X. X. et al. Genomic comparison of esophageal squamous cell carcinoma and its precursor lesions by multi-region whole-exome sequencing. Nat. Commun. 8, 524 (2017).
Castro, C., Peleteiro, B. & Lunet, N. Modifiable factors and esophageal cancer: a systematic review of published meta-analyses. J. Gastroenterol. 53, 37–51 (2017).
Song, Q. et al. Chromosomal and genomic variations in esophageal squamous cell carcinoma: a review of technologies, applications, and prospections. J. Cancer 8, 2492–2500 (2017).
Alder, H., Taccioli, C., Chen, H., Jiang, Y. & Smalley, K. J. Dysregulation of miR-31 and miR-21 induced by zinc deficiency promotes esophageal cancer. Carcinogenesis 33, 1736–1744 (2012).
Tanaka, K. et al. Frequent methylation-associated silencing of a candidate tumor-suppressor, CRABP1, in esophageal squamous-cell carcinoma. Oncogene 26, 6456–6468 (2007).
Shen, H. & Laird, P. W. Interplay between the cancer genome and epigenome. Cell 153, 38–55 (2013).
Toh, Y., Egashira, A. & Yamamoto, M. Epigenetic alterations and their clinical implications in esophageal squamous cell carcinoma. Gen. Thorac. Cardiovasc. Surg. 61, 262–269 (2013).
Cheng, Y. et al. A novel 19q13 nucleolar zinc finger protein suppresses tumor cell growth through inhibiting ribosome biogenesis and inducing apoptosis but is frequently silenced in multiple carcinomas. Mol. Cancer Res. 10, 925–936 (2012).
Huntley, S. et al. A comprehensive catalog of human KRAB-associated zinc finger genes: insights into the evolutionary history of a large family of transcriptional repressors. Genome Res. 16, 669–677 (2006).
Margolin, J. F. et al. Kruppel-associated boxes are potent transcriptional repression domains. Proc. Natl Acad. Sci. USA 91, 4509–4513 (1994).
Sripathy, S. P., Stevens, J. & Schultz, D. C. The KAP1 corepressor functions to coordinate the assembly of de novo HP1-demarcated microenvironments of heterochromatin required for KRAB zinc finger protein-mediated transcriptional repression. Mol. Cell. Biol. 26, 8623–8638 (2006).
Friedman, J. R. et al. KAP-1, a novel corepressor for the highly conserved KRAB repression domain. Genes. Dev. 10, 2067–2078 (1996).
Gebelein, B., Fernandez-Zapico, M., Imoto, M. & Urrutia, R. KRAB-independent suppression of neoplastic cell growth by the novel zinc finger transcription factor KS1. J. Clin. Investig. 102, 1911–1919 (1998).
Gebelein, B. & Urrutia, R. Sequence-specific transcriptional repression by KS1, a multiple-zinc-finger-Kruppel-associated box protein. Mol. Cell. Biol. 21, 928–939 (2001).
Cheng, Y. et al. KRAB zinc finger protein ZNF382 is a proapoptotic tumor suppressor that represses multiple oncogenes and is commonly silenced in multiple carcinomas. Cancer Res. 70, 6516–6526 (2010).
Sawada, G. et al. Genomic landscape of esophageal squamous cell carcinoma in a japanese population. Gastroenterology 150, 1171–1182 (2016).
Song, Y. et al. Identification of genomic alterations in oesophageal squamous cell cancer. Nature 509, 91–95 (2014).
Jones, P. A. Epigenetics in carcinogenesis and cancer prevention. Ann. N.Y. Acad. Sci. 983, 213–219 (2003).
Urrutia, R. KRAB-containing zinc-finger repressor proteins. Genome Biol. 4, 231 (2003).
Lupo, A. et al. KRAB-Zinc finger proteins: a repressor family displaying multiple biological functions. Curr. Genom. 14, 268–278 (2013).
Witzgall, R., O’Leary, E., Leaf, A., Onaldi, D. & Bonventre, J. V. The Kruppel-associated box-A (KRAB-A) domain of zinc finger proteins mediates transcriptional repression. Proc. Natl. Acad. Sci. USA 91, 4514–4518 (1994).
Quenneville, S. et al. The KRAB-ZFP/KAP1 system contributes to the early embryonic establishment of site-specific DNA methylation patterns maintained during development. Cell Rep. 2, 766–773 (2012).
Wiznerowicz, M. et al. The Kruppel-associated box repressor domain can trigger de novo promoter methylation during mouse early embryogenesis. J. Biol. Chem. 282, 34535–34541 (2007).
Schmitz, G., Heimerl, S. & Langmann, T. Zinc finger protein ZNF202 structure and function in transcriptional control of HDL metabolism. Curr. Opin. Lipidol. 15, 199–208 (2004).
Florio, F. et al. Biochemical and functional interaction between ZNF224 and ZNF255, two members of the Kruppel-like zinc-finger protein family and WT1 protein isoforms. Human. Mol. Genet. 19, 3544–3556 (2010).
Luo, K. et al. Expression of a novel Krupple-like zinc-finger gene, ZNF382, in human heart. Biochem. Biophys. Res. Commun. 299, 606–612 (2002).
Luo, X. et al. The tumor suppressor interferon regulatory factor 8 inhibits beta-catenin signaling in breast cancers, but is frequently silenced by promoter methylation. Oncotarget 8, 48875–48888 (2017).
Xiang, T. et al. The ubiquitin peptidase UCHL1 induces G0/G1 cell cycle arrest and apoptosis through stabilizing p53 and is frequently silenced in breast cancer. PloS ONE 7, e29783 (2012).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408 (2001).
Tao, Q. et al. Defective de novo methylation of viral and cellular DNA sequences in ICF syndrome cells. Human. Mol. Genet. 11, 2091–2102 (2002).
Tao, Q. et al. Methylation status of the Epstein-Barr virus major latent promoter C in iatrogenic B cell lymphoproliferative disease. Application of PCR-based analysis. Am. J. Pathol. 155, 619–625 (1999).
Mu, H. et al. Methylation of PLCD1 and adenovirus-mediated PLCD1 overexpression elicits a gene therapy effect on human breast cancer. Exp. Cell Res. 332, 179–189 (2015).
Mu, J. et al. Dickkopf-related protein 2 induces G0/G1 arrest and apoptosis through suppressing Wnt/beta-catenin signaling and is frequently methylated in breast cancer. Oncotarget 8, 39443–39459 (2017).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Marioni, J. C., Mason, C. E., Mane, S. M., Stephens, M. & Gilad, Y. RNA-seq: an assessment of technical reproducibility and comparison with gene expression arrays. Genome Res. 18, 1509–1517 (2008).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Wang, L. et al. DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 26, 136–138 (2010).
Anders, S. & Huber, W. Differential expression of RNA-Seq data at the gene level–the DESeq package. Heidelberg, Germany: European Molecular Biology Laboratory (EMBL). (2012) http://watson.nci.nih.gov/bioc_mirror/packages/2.11/bioc/vignettes/DESeq/inst/doc/DESeq.pdf.
Yu, G., Wang, L. G. & He, Q. Y. ChIPseeker: an R/Bioconductor package for ChIP peak annotation, comparison and visualization. Bioinformatics 31, 2382–2383 (2015).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinforma. 14, 178–192 (2013).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 11, R14 (2010).
Kanehisa, M. et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 36, D480–D484 (2008).
Dennis, G. et al. DAVID: Database for Annotation, Visualization, and Integrated Discovery. Genome biology. 4, P3 (2003).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Acknowledgements
This study was supported by National Natural Science Foundation of China (NSFC) (#81572769, #31420103915, #81772869, #81572327), National Key Research and Development Program of China (#2017YFE0191700) and the VC special research fund from The Chinese University of Hong Kong.
Authors’ contributions
C.Z., T.X., and Q.T.: conception and design of the study. C.Z., S.L., L.Y., R.S., Y.F., L.P., L.L., and X.W.: performed experiments and analysed data. C.Z., and L.Y.: collected samples. C.Z.: prepared figures and drafted the initial manuscript. G.R.: reviewed the manuscript. T.X., L.L., and Q.T.: revised and finalised the manuscript. All authors reviewed and approved the final manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All participants provided written consent before enrolment. This study was approved by the Ethics Committees of the First Affiliated Hospital of Chongqing Medical University and conformed to the tenets of the Declaration of Helsinki.
Consent for publication
I confirm this manuscript is original. If accepted, the article will not be published elsewhere without the written consent of the publisher.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
These authors are co-first authors: Chong Zhang, Tingxiu Xiang.
Edited by A. Oberst
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, C., Xiang, T., Li, S. et al. The novel 19q13 KRAB zinc-finger tumour suppressor ZNF382 is frequently methylated in oesophageal squamous cell carcinoma and antagonises Wnt/β-catenin signalling. Cell Death Dis 9, 573 (2018). https://doi.org/10.1038/s41419-018-0604-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-018-0604-z
- Springer Nature Limited
This article is cited by
-
The role and application of transcriptional repressors in cancer treatment
Archives of Pharmacal Research (2023)
-
Targeting epigenetic deregulations for the management of esophageal carcinoma: recent advances and emerging approaches
Cell Biology and Toxicology (2023)
-
Multi-omic characterization of genome-wide abnormal DNA methylation reveals diagnostic and prognostic markers for esophageal squamous-cell carcinoma
Signal Transduction and Targeted Therapy (2022)
-
Disruption of ZNF334 promotes triple-negative breast carcinoma malignancy through the SFRP1/ Wnt/β-catenin signaling axis
Cellular and Molecular Life Sciences (2022)
-
The tumor suppressor Zinc finger protein 471 suppresses breast cancer growth and metastasis through inhibiting AKT and Wnt/β-catenin signaling
Clinical Epigenetics (2020)