Abstract
Factors mediating mobilization of osteoblastic stem and progenitor cells from their bone marrow niche to be recruited to bone formation sites during bone remodeling are poorly known. We have studied secreted factors present in the bone marrow microenvironment and identified KIAA1199 (also known as CEMIP, cell migration inducing hyaluronan binding protein) in human bone biopsies as highly expressed in osteoprogenitor reversal cells (Rv.C) recruited to the eroded surfaces (ES), which are the future bone formation sites. In vitro, KIAA1199 did not affect the proliferation of human osteoblastic stem cells (also known as human bone marrow skeletal or stromal stem cells, hMSCs); but it enhanced cell migration as determined by scratch assay and trans-well migration assay. KIAA1199 deficient hMSCs (KIAA1199down) exhibited significant changes in cell size, cell length, ratio of cell width to length and cell roundness, together with reduction of polymerization actin (F-actin) and changes in phos-CFL1 (cofflin1), phos-LIMK1 (LIM domain kinase 1) and DSTN (destrin), key factors regulating actin cytoskeletal dynamics and cell motility. Moreover, KIAA1199down hMSC exhibited impaired Wnt signaling in TCF-reporter assay and decreased expression of Wnt target genes and these effects were rescued by KIAA1199 treatment. Finally, KIAA1199 regulated the activation of P38 kinase and its associated changes in Wnt-signaling. Thus, KIAA1199 is a mobilizing factor that interacts with P38 and Wnt signaling, and induces changes in actin cytoskeleton, as a mechanism mediating recruitment of hMSC to bone formation sites.
Similar content being viewed by others
Introduction
Human osteoprogenitor cells, also known as human skeletal stem cells, marrow stromal or mesenchymal stem cells (hMSCs), represent a population of non-hematopoietic cells that exist at different locations within the bone marrow near eroded surfaces and can differentiate into mature osteoblastic bone forming cells1,2. The initiation of in vivo bone formation during skeletal remodeling and bone regeneration during fracture healing depend on the mobilization of sufficient number of osteoprogenitor cells to future bone formation sites1. This critical recruitment is impaired during aging and in metabolic bone diseases, including osteoporosis1,3.
As bone remodeling takes place asynchronously in the skeleton, the coupling of bone formation to resorption is tightly orchestrated by local coupling factors. These coupling factors are believed to mobilize osteoprogenitor cells from their niche, and recruit them to eroded surface prior to initiation of bone formation1. However, the identity of these factors is under investigation and currently only few have been identified and shown to be produced by osteoclastic, osteoblastic cells or other cells in the hematopoietic microenvironment4.
From a translational perspective, hMSCs have been employed in an increasing number of clinical trials for enhancing bone formation and tissue regeneration2. However, systemically infused hMSCs exhibit poor homing to the injured tissues5,6 and the majority of the cells are trapped in the lungs with very few cells reaching and engrafting in the skeleton7,8. To achieve clinical goals of using hMSCs in therapy, there is a need for identifying molecules and factors that enhance hMSCs migration and motility9,10,11.
Several factors have been identified to ‘mobilize’ hematopoietic stem cells out of their niche as the first step for induction of differentiation12, but very few factors have been reported to enhance hMSCs mobilization from their bone marrow niche. Substance P has been reported to mobilize a subgroup of bone marrow stromal cells with MSC-like phenotype13. Also, following bone fracture, the number of circulating human MSC-like cells increased14 suggesting that changes in bone microenvironment following bone fracture, release osteoprogenitor cells mobilizing factors that are yet to be identified.
We have previously performed a global quantitative proteomic studies on hMSCs secretome, and identified a number of secreted factors which regulate MSCs lineage allocation, differentiation and functions15, e.g., Legumain (LGMN) and Collapsin Response Mediator Protein 4 (CRMP4)16,17. Among the identified factors, KIAA1199 was found to be highly expressed by hMSCs in vitro and in vivo but its function in hMSCs biology is not known.
KIAA1199, also named as CEMIP (cell migration inducing protein), is expressed from a gene located on chromosome 15q25.1 and encodes 150 kDa protein18 with N-terminal secretion signal peptide. KIAA119 has a PbH1 domain consisting of parallel beta-helix repeats, which is predicted to function in polysaccharide hydrolysis19, G8 domain containing eight conserved glycine residues and five repeated beta-strand pairs and one alpha-helix20, and two GG domains consisting of seven beta-strands and two alpha-helices21. Many G8-containing proteins are integral membrane proteins with signal peptides and/or transmembrane segments, suggesting that KIAA1199 is a secreted factor that plays a role in extracellular ligand binding and processing. The biological role of KIAA1199 has been studied in cancer biology and a number studies has demonstrated high expression levels in cancer cell lines and its association with invasive and metastatic disease22,23.
In the present study, we examined regulatory role of KIAA1199 in hMSCs migration, as well as its cellular and molecular mechanism of action. We observed that KIAA1199 is expressed in osteoprogenitor cells and enhances their migration abilities through regulation of cell shape, actin cytoskeletal dynamics and regulation actin depolymerizing factors Cofilin1 (CFL1), LIM domain kinase 1 (LIMK1) and Destrin (DSTN). Furthermore, KIAA1199 promotes cell migration by cooperative activation of canonical Wnt and p38/MAPK signaling.
Material and methods
In situ hybridization
Formalin-fixed, decalcified and paraffin-embedded bone specimens from eight human controls were included in situ hybridization analysis. Four of the human specimens were diagnostic iliac crest biopsies obtained from control patients formerly under investigation for a hematological disorder, as previously described24, while the remaining four human specimens were from the proximal femur of coxa valga patients, as previously described1. The study is approval by the Danish National Committee on Biomedical Research Ethics (S-20070121 and S-20120193). Paraffin sections (3.5-µm thick) were in situ hybridization using an enhanced version of the RNAScope 2.5 high definition procedure (310035, ACD Bioscience, Attikis, Greece). Sections were rehydrated, deparaffinized and pretreated as previously24, and hybridized overnight at 40°C with 20-ZZ-pair probes binding either the X-Y region of the human KIAA1199 mRNA diluted 1:1 in probe diluent (Cat No. 449819, ACD Bioscience). A negative control only hybridized with probe diluent was included. The amplification was conducted according to the instructions provided by the manufacturer. The horse radish peroxidase was further enhanced with digoxigenin (DIG)-labeled tyramide (NEL748001KT, PerkinElmer, Skovlunde, Denmark), which was labeled with alkaline-phosphatase-conjugated sheep anti-DIG FAB-fragments (11093274910, Roche, Hvidovre, Denmark) and visualized with Liquid Permanent Red (DAKO, Glostup, Denmark). Finally, the sections were counterstained with Mayer’s Haematoxylin and mounted with Aqua-Mount.
Cell culture
As a model for human osteoprogenitor cells, we employed the telomerized hMSCs (hMSC-TERT) line that was established in our laboratory by stable overexpression of human telomerase reverse transcriptase gene (hTERT)25. hMSC-TERT expresses all known markers of osteoprogenitor cells including the ability to form bone when implanted in vivo25. To simplify, hMSC-TERT will be referred to as hMSCs. Cells were cultured in Minimum Essential Medium (MEM) with 10% fetal bovine serum (FBS) and penicillin-streptomycin (P/S) (1%). All reagents were purchased from Life Technologies (Nærum, Denmark). Cells were incubated in 5% CO2 incubators, 37 °C and humidity of 95%.
Cell transfection and infection
hMSCs and HEK293T cells were cultured until 70–80% confluent and transfected by small interfering RNA (siRNA) or constructed plasmids using Lipofectamine 2000 (Thermo-Fisher Scientific, Roskilde, Denmark) according to the manufacturer’s instructions. siRNAs for specific genes or control non-targeting siRNA were all purchased from Silencer® Selector siRNA library from Thermo-Fisher Scientific. The KIAA1199 (Gene ID: NM_001293298.1) expressing plasmid construct was produced with pLVX-mCMV-ZsGreen-Puro vector (BioWit, Shenzhen, China). Plasmid DNA preparations were made from over-night cultures of individual colonies using the Plasmid MidiPrep System (Qiagen, Stockach, Germany) following the manufacturer’s protocol.
Lentiviruses were made by transfecting packaging cells (HEK293T) with a three-plasmid system: DNA for transfections was prepared by mixing plasmids for KIAA1199, pVSVg, and psPAX2 with Lipofectamine® 2000 Reagent according to the manufacturer’s instructions (Thermo-Fisher Scientific, Roskilde, Denmark). Lentiviral supernatants were collected at 36 h post-transfection and added polybrene to 8 μg/ml, then immediately used to infect target cells. Cells were incubated for 24 h and the media was changed to remove virus particles. Transduced cells were cultured in the presence of 1.0 μg/ml puromycin to select for positive cells for one month. A control cell line was generated by transducing hBMSC-TERT with virus containing the empty pBABE-puro vector.
Quantitative real-time PCR (qRT-PCR)
RNA from cells was isolated by TRIzol® according to the manufacturer’s instructions (Thermo-Fisher Scientific, Roskilde, Denmark). The first strand complementary DNA was synthesized from 1 µg total RNA by Revert aid cDNA kit (Sigma, Copenhagen, Denmark). RT-qPCR was performed by ABI StepOneTM Real-TIME PCR machine with SYBR green (Applied Biosystems, Roskilde, Denmark). The primers of genes used in the study were listed in supplemental Table 1. The data was normalized to geometric means of the several reference genes and analyzed by a comparative CT method where ΔCT is the difference between the CT values of the target and the geometric mean of the reference genes.
Collection of conditioned medium
Cells were cultured as required at the specified time point, and washed with PBS and incubated in serum-free medium for 8–12 h. The conditioned medium (CM) was then collected. Floating cells were spun down at 250×g for 5 min. The supernatant was kept in –80 °C. For Western blot analysis at protein in CM, CM was concentrated (1:20) by centrifugation at 3000×g for 30–45 min with Amicon Ultra Centricons (10 kDa NMWL) (Millipore, Hellerup, Denmark).
Western blot analysis
Western blot analysis was performed as described previously26. The cells were washed in cold PBS twice and lysed in RIPA buffer (Thermo-Fisher Scientific, Roskilde, Denmark) supplemented with protease inhibitors (Roche, Hvidovre, Denmark) for 30 min in cold room. Samples were centrifuged at 12,000 r.p.m., 4 °C for 10 min. Protein concentrations were determined with a BCA kit (Thermo-Fisher Scientific, Roskilde, Denmark), and equal amounts (30 µg) of proteins were loaded on a polyacrylamide gel (Thermo-Fisher Scientific, Roskilde, Denmark). Blotted nitrocellulose membranes were incubated overnight with primary antibody at 4 °C, and were developed after 1 h incubation with secondary anti-rabbit horseradish peroxidase-conjugated antibody (Santa Cruz Biotechnology, Heidelberg, Germany) using an ECL Western blotting kit (GE Healthcare, Brøndby, Denmark) and Kodak films. Antibodies for KIAA1199 was purchased from ProteinTech, Manchester, UK; Antibodies (total or phosphor) specific for CFL1, LIMK1, p38, GSK3β and beta-catanin were obtained from Cell Signaling (Herlev, Denmark); antibodies for DSTN and actin were bought from Sigma-Aldrich (Copenhagen, Denmark). All antibodies were used at 1:1000 dilutions except actin antibodies (1:2500).
Cell counting and cell viability
The cells were washed by PBS, trypsinized, and stained with 0.4% Trypan Blue (1:1 V/V) and counted using a hemocytometer. Cell viability was determined by Cell Titer-Blue® cell viability assay according to the manufacturer´s instructions (Promega Biotech AB, NACKA, Sweden). In briefly, cells were cultured in 96-well plates and 20 µl/well of CellTiter-Blue Reagent was added and incubated for 1 h at 37 °C. Fluorescence at 560/590 nm by FLUO Star Omega Plate Reader (BMG Laboratories, Ortenberg, Germany) was determined.
TCF luciferase reporter assay
TCF-dependent transcriptional activity was studied in hMSC-TERT transduced with TCF promoter constructs along with Renilla luciferase reporter gene (50:1 ratio) as described previously27. After 48 h of siRNA transfection, firefly and Renilla luciferase activities were measured using the Dual-Glo Luciferase Assay System (Promega Biotech AB).
In vitro scratch assay
The confluent monolayer of cells was scratched with a 1000-μl pipette tip smoothly to induce a gap without cell detachment, and the cells were cultured for 8–48 h. Images were collected at the same positions of the plate at different time points post-scratching. Images were quantified using Image-J software that measured the migrated area.
Trans-well migration assay
Cells were cultured to 70–80% confluence and starved for 18–20 h in serum-free LG DMEM (Life Technologies, Tastrup, Denmark). Transwell migration assay was performed using 12 mm Millicell® inserts (Millipore). Briefly, adding 5% FBS MEM or specific conditional medium in the lower chamber of well, test cells were plated in the upper chamber in 0.2% FBS DMEM medium and incubated at 37 °C for 16 h. The cells on the surface of the upper chamber were removed by cotton swabs; the membranes in the inserts were removed and fixed by formalin buffer (Sigma-Aldrich) for 5 min, stained in Hemacolor staining solution (Merck, Germany) for 15 min. Transfer the membranes to glass slide (Menzelgläzer, Germany) and scan the whole membrane by a Lecica DM4500 microscope (Olympus, UK). The numbers of migrated cells or sizes of membrane area covered by the cells were determined by Surveyor Turboscan Mosaic acquisition imaging system (Objective Imaging, Cambridge, UK).
Single cell morphology analysis by Operetta high content imaging system
Single cell morphology analysis was performed using Operetta CLSTM High Content Imaging system (PerkinElmer, Skovlunde, Denmark) as we described in a previous study28. Briefly, the cells were trypsinized and seeded as 1000 cell/well into clear bottom 96-well CellCarrier™ microtiter plates (PerkinElmer, Denmark). The cells were fixed in 4% paraformaldehyde for 10 min, washed with PBS, stained with DAPI (Sigma-Aldrich, D8417) and Phalloidin-TRITC (Sigma-Aldrich®, P1951). Fluorescent images were analyzed at × 10 and × 40 magnification using Harmony High Content Imaging and Analysis Software (PerkinElmer, Denmark). All pictures were acquired with the same contrast and brightness parameters, and data analysis performed by same parameters setting.
Statistical analysis
Data are expressed as the mean and standard deviation (SD). Student’s t-test was used to assess differences between two groups, and one-way analysis of variance (ANOVA) was used for statistical testing involving more than two groups. P < 0.05 considered to be significant. n = number of independent experiment and each experiment was repeated as least 3 times.
Results
KIAA1199 is expressed in osteoprogenitor cells in human bone biopsies
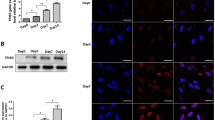
Our previous study had identified KIAA1199 as a secreted factor by human osteoblastic stem cells, and its secretion is enhanced during osteoblast differentiation in vitro15. To corroborate the cellular expression of KIAA1199 in the bone marrow microenvironment in vivo, we performed in situ hybridization analysis of adult human iliac crest bone biopsies that revealed KIAA1199 expression was localized to bone formation sites during bone remodeling and expressed specifically in immature and mature osteoblastic cells along bone surfaces in both cancellous and cortical bone (Fig. 1). KIAA1199 expression was abundant in the osteoblastic reversal cells (Rv.C) recruited to the eroded surfaces (ES), which are the future bone formation sites (Fig. 1). These cells have recently been reported to reflect motile osteoprogenitor cells which differentiate into bone-forming osteoblastic cells when reaching a critical cell density29. Moreover, KIAA1199 expression was present in osteoblastic bone lining cells (BLCs) and bone marrow envelope (BME) above quiescent surfaces, as well as osteoblastic canopy cells above eroded and osteoid surfaces (Fig. 1). These cells have been reported to be local source of osteoprogenitor cells (hMSCs) which are recruited to the eroded and osteoid surfaces during bone remodeling29. KIAA1199 expression was rich present in mature bone-forming osteoblast, but exhibited limited expressed in osteocytes and was absent in osteoclasts (Fig. 1).
Four human iliac crest bone specimens were in situ-hybridized for KIAA1199. KIAA199 expresses in reversal cells (Rv.C–light blue arrowheads), i.e., progenitor cells, colonizing the eroded surfaces (ES-future bone formation sites), the bone lining cells (BLC), bone marrow envelope (BME) (gray arrowheads) above quiescent surfaces (QS), as well as canopy cells (green arrowheads) above eroded surfaces (ES) that reported to be a local source of osteoprogenitor cells. KIAA1199 expression was also clear present in mature osteoblasts (OB–dark blue arrowheads) on osteoid surfaces (OS) and in some osteocytes, and absent in osteoclasts (OC). The overall expression level in each cell type is indicated below their illustrations: - no expression; + low expression, + + medium expression. Scale bar: 40 µm
KIAA1199 regulates hMSCs cell migration
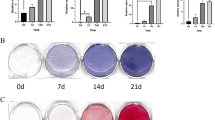
Based on the in vivo localization of KIAA1199, we studied its role in hMSCs migration and motility. hMSCs represent a population of stem and osteoprogenitor cells that express osteoblastic markers and can form mineralized matrix in vitro and form bone when implanted in vivo25. We established two hMSCs cell lines: KIAA1199 deficient (KIAA1199down) and KIAA1199 overexpressing (KIAA1199over) hMSCs. Inhibition of KIAA1199 expression was performed by specific siRNA and resulted in 90% reduction of expression evidenced by qRT-PCR and Western blot analysis (Fig. 2a), and the reduced levels of KIAA1199 lasted for more than 12 days (data not shown). KIAA1199over cells were created by lentivirus-mediated gene transfer that resulted in 8 fold higher stable expression of KIAA1199 compared to vector infected control (Fig. 2b). Changes of KIAA1199 expression in hMSCs did not affect cell proliferation (Fig. 2c&d). To study cell migration and motility, we employed two independent assays: in vitro scratch assay that examines cell migration in presence of intact cell–cell interactions and thus mimics migration of cells in vivo30 and Boyden chamber transwell migration assay testing the cell migration in suspension31. In scratch assay, KIAA1199down exhibited reduced cell motility, as well as reduction of the area covered by the cells, compared with non-target siRNA control cells (siR-Ctrl) (Fig. 3a&c). On the other hand, KIAA1199over showed enhanced cell motility and complete cell coverage of the scratch area when compared to vehicle control hMSCs (V-Ctrl) (Fig. 3b&d). In Boyden Chamber Trans-well migration assay, KIAA1199down exhibited a significant decrease in transwell cell migration (Fig. 3e) and the opposite effect was detected for KIAA1199over (Fig. 3f). Similar result also got in human primary culturing human bone marrow cells: knocking down of KIAA1199 by its specific siRNA inhibited cell migration as the hMSC-KIAA1199down cell line (Fig. S1). Since KIAA1199 is detected as a secreted factor in colorectal epithelial cells32 and it has also been reported to be present within the endoplasmic reticulum (ER)33. We detected KIAA1199 is expressed and secreted in hMSCs (Fig. S2), and we further studied whether the secreted form of KIAA1199 enhances cell migration. Adding conditioned medium from KIAA1199over (KIAA1199-CM) enhanced cell migration compared to control conditional medium (Ctrl-CM) (Fig. 3g) and also rescued the reduced migration in KIAA1199down hMSCs (Fig. 3h).
a KIAA1199 deficient hMSCs (KIAA1199down) were obtained by transfecting the cells with a specific siRNA for KIAA1199 (siR-KIAA1199) or with non-target siRNA control (siR-Ctrl); (b) KIAA1199-overexpression hMSCs (KIAA1199over) were established using a lentiviral transduction and, corresponding control was established by parallel infected by lentiviral vehicle (V-Ctrl). Expression levels of KIAA1199 in cells were determined using qRT-PCR and Western blot analysis (a, b). c–d Cell proliferation was determined by cell counting and cell viability assay
a–b In vitro scratch assay was performed in hMSC-KIAA1199down or hMSC-KIAA1199over cells. Photomicrographs were obtained at the same position of culture dish after 20 h incubation in 0.2% fetal bovine serum (FBS) cultured medium, Scale bar: 500 µm. c–d The ability of the cells to ‘heal the wound’ (migrated area) was determined as a ratio of cell covered area per total area by Image-J® program (n ≥ 3). e–f Boyden Chamber trans-well cell migration assay were performed in hMSC-KIAA1199down or hMSC-KIAA1199over and cell migration was measured after 16 h. The migrated cells were calculated by Image-J program (n ≥ 3). g hMSCs were placed in the upper chamber of Boyden chamber system and Lower chambers were filled with conditional mediums (CM) from KIAA1199 overexpression cells (KIAA1199-CM) or its corresponding vehicle control cells (Ctrl-CM). Cell migration was measured after 16 h. h KIAA1199 deficient hMSCs (KIAA1199down) and control cells (siR-Ctrl) were placed in upper chamber of Boyden chamber system, Ctrl-CM or KIAA1199-CM were added into lower chamber. Cell migration was measured after 16 h. The migrated cells were calculated by Image-J program (n ≥ 3). Data represent mean ± SD. *P < 0.05, **P < 0.01, ***P < 0.001. Scale bar for pictures: 500 µm
KIAA1199 regulates hMSCs cell morphology and actin cytoskeleton organization
Cell motility is associated with changes in cell morphology34. Thus, we examined the effect of KIAA1199 deficiency on hMSCs morphology employing high content imaging by the OperettaTM high-content image system. As shown in Fig. 4a, hMSCs-KIAA1199down exhibited changes in cell morphology from fibroblast-like to rounded morphology with reduced cell size and cell length, increased cell roundness, altered the ratio of cell width to length (Fig. 4b). Previous studies have shown that cytoskeletal changes in terms of polymerization filamentous actin (F-actin) and depolymerization to globular actin (G-actin) mediates morphological changes affecting cell motility and migration35,36. We observed that F-actin staining intensity was significantly reduced in KIAA1199down (Fig. 4c) and this was associated with increased levels of DSTN, decreased the phosphorylation of CFL1 and LIM domain kinase 1 (LIMK1) that are factors controlling actin polymerization and depolymerization (Fig. 4d). The opposite changes were observed in KIAA1199over that exhibited down-regulation of DSTN and up-regulation of P-CFL1 and P-LIMK1 (Fig. 4d).
Human bone marrow skeletal (stromal) stem cells (hMSCs) were cultured and transfected with siRNA specific for KIAA1199 or non-target control siRNAs to obtain KIAA1199 deficient hMSCs (KIAA1199down) and the corresponding control cells (siR-Ctrl). a The cells were transferred and seeded for 16 h, stained with TRITC-phalloidin and DAPI. Images were acquired on the Operetta® high content image system using ×10 or ×40 objective. b Parameters of cell morphology, including cell size (area) (µm2), cell length (µm), cell width (µm), cell roundness, and cell ratio (width to length) were analyzed by the Harmony® software. c The staining intensity of F-actin was analyzed by the Harmony® software for phalloidin stained F-actin. N ≥ 3, *P < 0.05, **P < 0.01. d Actin depolymerizing factors involved in actin cytoskeletal dynamics: CFL1, LIMK1 and DSTN, were measured by Western-blot analysis in KIAA1199down or KIAA1199over hMSCs
Wnt and p38 signaling mediate KIAA1199-enhanced cell motility
Previous studies suggested that Wnt signaling is important in regulation of stem cell migration37,38. To examine the interaction of Wnt/β-catenin signaling with KIAA1199, we employed the Wnt/TCF-luciferase reporter assay. As shown in Fig. 5a, hMSC-KIAA1199down exhibited significant decrease in Wnt/TCF activity compared with non-target siRNA controls. hMSC-KIAA1199over exhibited opposite changes with enhanced Wnt/TCF-luciferase reporter activity (Fig. 5b). Consistent with the results, mRNA gene expression levels of Wnt/β-catenin responsive genes, such as DKK2, Axin2, TNFRSF19–2, and NKD1 were all significantly inhibited in KIAA1199down cell while enhanced in KIAA1199over (Fig. 5c). Moreover, adding KIAA1199-CM rescued the inhibition of Wnt/TCF reporter activity in KIAA1199down hMSCs (Fig. 5d). We further examined several signaling pathways known to be important in cell motility39. Western-blot analysis showed that phosphorylation of P38 and active β-catenin were decreased in KIAA1199down hMSCs, while were enhanced in KIAA1199over hMSCs (Fig. 6a). Similar changes were observed in the phosphorylation of GSK3β (Thr390) (Fig. 6a). To confirm the association between P38 changes and hMSCs migration, inhibition of p38 signaling by P38 small molecule inhibitor SB203580 reduced the KIAA1199-induced cell migration (Fig. 6b). Furthermore, to corroborate the role of Wnt signaling, we treated the cells with Wnt signaling inhibitor WiKi4 that resulted in reduced the KIAA1199-induced cell migration in a dose dependent fashion (Fig. 6c).
a–b Wnt/TCF-luciferase-reporter was transduced into hMSCs deficient in KIAA1199 (KIAA1199down) that were created by siRNA-mediated knock-down and control cells transfected with non-target siRNA (siR-Ctrl) (a); or transduced into hMSCs overexpressing KIAA1199 (KIAA1199over) that was created by lentiviral transduction and corresponding control (V-Ctrl) was established by parallel infected by lentiviral vehicle (b). In presence of 50% Wnt3a condition medium induction, TCF-Luciferase activity was measured after 24 h incubation, firefly luciferase was normalized to parallel renilla luciferase activities in the same sample using the Dual-Glo Luciferase Assay System. c The mRNA expression levels of canonical Wnt signaling target genes NKD1, DKK2, TNFRSF19-2, and Axin2 were determined by qRT-PCR and normalized to internal controls. d KIAA1199down and control cells were cultured in control conditional medium (Ctrl-CM) or KIAA1199 conditioned medium (KIAA1199-CM), with Wnt3a stimulation, TCF-Luciferase activity was measured after 24 h incubation, firefly luciferase was normalized to parallel renilla luciferase activities in the same sample using the Dual-Glo Luciferase Assay System. At least three independent experiments were preformed, data presented as mean ± SD, *P < 0.05, **P < 0.01, ***P < 0.001
a KIAA1199down or KIAA1199over were starved for 6 h, and stimulated with 5% FBS medium for 15 min. Signaling pathways were analyzed by Western-blot analysis. b hMSCs were pretreated with P38 inhibitor (SB203580, 5 µM or 10 µM) for 2 h and trans-well migration assay was performed in the absence or presence of KIAA1199 conditioned medium (CM). c hMSCs were pretreated with Wnt signaling inhibitor (Wiki4, 0.1 µM or 1 µM) for 2 h and trans-well migration assay was performed in the absence or presence of KIAA1199 conditioned medium (CM). At least three independent experiments were preformed, data presented as mean ± SD, **P < 0.01, ***P < 0.001. Scale bar for pictures: 200 µm. d Mechanisms model for the regulation of KIAA1199 in hMSCs mobility and migration
Discussion
Identifying factors mediating osteoprogenitor cell mobilization to future bone formation sites during fracture healing is relevant for understanding the biology of bone remodeling and bone formation dynamics, as well as for clinical use of the cells for enhancing bone regeneration. In the current study, we identified a secreted factor KIAA1199 which is expressed in the human bone microenvironment by the motile osteoprogenitor cells and it enhances stem and osteoprogenitor cell motility and migration. We also demonstrated that KIAA1199 exerts significant changes in cell morphology, actin cytoskeletal dynamics and mediates its effects through cooperative regulation of the canonical Wnt and P38/MAPK signaling (Fig. 6d).
Bone formation during bone remodeling depends on migration of a cohort of osteoprogenitor cells to future bone formation sites. Recent histomorphometric studies that have employed a number of markers to identify the osteoprogenitor cell populations, demonstrated that osteoprogenitor cells correspond to the cells that are defined as ‘reversal cells’ in classical histomorphometries studies40. We observed that KIAA1199 is enriched in ‘reversal cells’ in human bone biopsies that are colonizing future bone formation sites. In addition, KIAA1199 is also expressed in their local reservoir of osteoprogenitor cells (canopy cells, bone marrow envelope cells, and bone lining cells)41. This pattern of localization support the hypothesis that mobilizing factors are needed to allow osteoprogenitor cells to reach the critical threshold cell density needed for initiating bone formation on eroded surfaces.
We have employed hMSCs as an in vitro model for osteoprogentior cells and the in vitro counterpart of the ‘reversal cells’. In support of this choice is that the in vivo phenotype of reversal cells is characterized by the expression of a number markers e.g., SMA, Runx2, Collagen type III40 that are highly expressed in cultured hMSCs42. In addition, cultured hMSCs can form mineralized matrix in vitro and form normal lamellar bone in vivo25 corroborating that the in vitro cultured hMSC are enriched in osteoprogenitor cells.
KIAA1199 enhanced cell migration of hMSCs in two independent assays. The in vitro scratch assay that tests the effects on cell migration in presence of intact cell-extracellular matrix (ECM) and cell–cell interactions, thus mimicking the migration of cells in vivo30. In addition, Boyden chamber transwell migration assay was employed to test cell migration in suspension. The observed enhanced motility and migration of hMSCs by KIAA1199 corroborates previous findings of reported for its role in cancer cell biology. For example, in gastric cancer cell lines, KIAA1199 supported cancer cell migration and invasion and its expression was upregulated in invasive gastric cancer tissues associated with poor prognosis43. Similar results were obtained in breast cancer33, where KIAA1199 was associated with enhanced breast cancer cell motility and its expression was upregulated in invasive breast cancer associated with in vivo metastases and poor prognosis33.
Migration-stimulating signals induce morphologic and cytoskeletal changes34. We observed that the presence of KIAA1199 was associated with fibroblast-like cellular morphology, whereas its absence led to cell rounding and reduced F-actin levels. Similar to our findings, overexpressing KIAA1199 in a human SW480 colon cancer cells was associated with changes in cellular morphology from round cell to a flatter epithelial-like morphology32. Also, deficiency of KIAA1199 in human breast cancer cell lines MDA-MB-435 or MDA-MB-231 led to changed cell morphology from fibroblast-like to a polarized epithelial-like morphology. This was accompanied by a reorganization of F-actin from stress fibers to a cortical ring-like structure33. We have extended these results by providing a possible molecular explanation for the changes in actin cytoskeleton by demonstrating significant changes in the levels of actin depolymerizing factors, including Cofilin1 (CFL1) and Destrin (DSTN)28,44. Interestingly, we have previously reported that these factors to mediate significant changes in actin cytoskeleton during osteoblast differentiation of hMSCs and that the increase in p-CFL1, p-LIMK1, and the decreased DSNT were associated with enhanced osteoblast differentiation28, which suggests that the observed changes in cell morphology are needed for the bone forming functions of osteoprogenitor cells.
How does KIAA1199 exert its effects on cell motility? We have tested a number of signaling pathways regulating cell motility and migration. Wnt signaling has been reported to play a role in cell migration45,46 and it is an important regulator of bone formation in bone microenvironment47. We observed that KIAA1199 enhanced Wnt/β-catenin signaling pathway. Similar to our results, KIAA1199 upregulates Wnt/β-catenin signaling in gastric cancer cells and its deficiency reduces β-catenin expression in NCI-N87 and AGS cancer cells43. In addition to Wnt signaling, we observed that high levels of p38 were associated with KIAA1199-mediated increased in hMSCs migration. Interaction between Wnt and p38 has been described as regulator for MSCs biology48. Wnts are capable of activating p38 MAPK: positive effects of Wnt4 on osteoblastic cell differentiation of MSCs are mediated through cross-talk between Wnt and p38 MAPK signaling49; Wnt3a increases active p38 which in turn enhances osteoblast differentiation in C3H10T1/2 mesenchymal cells50. On the other hand, P38 MAPK regulated Wnt/beta-catenin through inactivation of GSK or LRP651,52. However, whether a physical interaction exists between Wnt, p38 signaling, and KIAA1199 remain to be determined.
We observed the expression of KIAA1199 in osteoprogenitor cells colonizing eroded surfaces, reflecting the reversal-resorption phase; corroborate a possible role in osteoprogenitor cell migration in vivo. Two previous findings support this hypothesis. First, KIAA1199 has recently been report to depolymerize hyaluronic acid (HA)53 and immunohistochemical studies have demonstrated the presence of HA in the endothelium lining of the sinusoids and endostium54 similar to the location where we identified KIAA1199-expressing cells. Second, KIAA1199 degrades HA that is induced by inflammatory mediators present in bone microenvironment and upregulated during bone remodeling55,56. Our data suggest a model where osteoprogenitor cells are retained in their niche through cell anchorage to matrix HA, probably through CD44 receptor which is highly expressed in osteoprogenitor cells57. During the reversal-resorption phase of bone remodeling, high levels of KIAA1199 likely degrades HA, leading to mobilization of osteoprogentior cells. This model suggest a new role for KIAA1199 as an ECM remodeling factor during the reversal-resorption phase that regulates adhesion and migration properties of osteoprogenitor cells. Our data further support the biological role of KIAA1199 in bone biology, as it interacts with Wnt and P38/MAPK singling, important regulators of bone formation and bone remodeling.
References
Lassen, N. E. et al. Coupling of bone resorption and formation in real time: new knowledge gained from human haversian BMUs. J. Bone Miner. Res. 32, 1395–1405 (2017).
Zaher, W., Harkness, L., Jafari, A. & Kassem, M. An update of human mesenchymal stem cell biology and their clinical uses. Arch. Toxicol. 88, 1069–1082 (2014).
Parfitt, A. M., Villanueva, A. R., Foldes, J. & Rao, D. S. Relations between histologic indices of bone formation: implications for the pathogenesis of spinal osteoporosis. J. Bone Miner. Res. 10, 466–473 (1995).
Sims, N. A. & Martin, T. J. Coupling the activities of bone formation and resorption: a multitude of signals within the basic multicellular unit. Bone. Rep. 3, 481 (2014).
Devine, S. M. et al. Mesenchymal stem cells are capable of homing to the bone marrow of non-human primates following systemic infusion. Exp. Hematol. 29, 244–255 (2001). [pii].
Horwitz, E. M. et al. Isolated allogeneic bone marrow-derived mesenchymal cells engraft and stimulate growth in children with osteogenesis imperfecta: Implications for cell therapy of bone. Proc. Natl Acad. Sci. USA 99, 8932–8937 (2002). 99/13/8932 [pii].
Karp, J. M. & Leng Teo, G. S. Mesenchymal stem cell homing: the devil is in the details. Cell Stem Cell 4, 206–216 (2009). S1934-5909(09)00056-3 [pii].
Bentzon, J. F. et al. Tissue distribution and engraftment of human mesenchymal stem cells immortalized by human telomerase reverse transcriptase gene. Biochem. Biophys. Res. Commun. 330, 633–640 (2005). [pii] 10.1016/j.bbrc.2005.03.072.
De Becker, A. & Riet, I. V. Homing and migration of mesenchymal stromal cells: how to improve the efficacy of cell therapy? World J. Stem Cells 8, 73–87 (2016).
Andersen, R. K. et al. Association between in vivo bone formation and ex vivo migratory capacity of human bone marrow stromal cells. Stem Cell Res. Ther. 6, 196 (2015). 10.1186/s13287-015-0188-9.
Levy, O. et al. A small-molecule screen for enhanced homing of systemically infused cells. Cell Rep. 10, 1261–1268 (2015). .
Bonig, H., Chudziak, D., Priestley, G. & Papayannopoulou, T. Insights into the biology of mobilized hematopoietic stem/progenitor cells through innovative treatment schedules of the CXCR4 antagonist AMD3100. Exp. Hematol. 37, 402–415 (2009).
Hong, H. S. et al. A new role of substance P as an injury-inducible messenger for mobilization of CD29(+) stromal-like cells. Nat. Med. 15, 425–435 (2009).
Eghbali-Fatourechi, G. Z. et al. Circulating osteoblast-lineage cells in humans. N. Engl. J. Med. 352, 1959–1966 (2005). 10.1056/NEJMoa044264.
Kristensen, L. P. et al. Temporal profiling and pulsed SILAC labeling identify novel secreted proteins during ex vivo osteoblast differentiation of human stromal stem cells. Mol. Cell. Proteom. 11, 989–1007 (2012).
Jafari, A. et al. Legumain regulates differentiation fate of human bone marrow stromal cells and is altered in postmenopausal osteoporosis. Stem Cell Rep. 8, 373–386 (2017). 10.1016/j.stemcr.2017.01.003.
Abdallah, B. M. et al. CRMP4 inhibits bone formation by negatively regulating BMP and RhoA signaling. J. Bone Miner. Res. 32, 913–926 (2017).
Abe, S., Usami, S. & Nakamura, Y. Mutations in the gene encoding KIAA1199 protein, an inner-ear protein expressed in Deiters’ cells and the fibrocytes, as the cause of nonsyndromic hearing loss. J. Hum. Genet. 48, 564–570 (2003).
Jenkins, J., Mayans, O. & Pickersgill, R. Structure and evolution of parallel beta-helix proteins. J. Struct. Biol. 122, 236–246 (1998).
He, Q. Y. et al. G8: a novel domain associated with polycystic kidney disease and non-syndromic hearing loss. Bioinformatics 22, 2189–2191 (2006).
Guo, J., Cheng, H., Zhao, S. & Yu, L. GG: a domain involved in phage LTF apparatus and implicated in human MEB and non-syndromic hearing loss diseases. FEBS Lett. 580, 581–584 (2006).
Kuscu, C. et al. Transcriptional and epigenetic regulation of KIAA1199 gene expression in human breast cancer. PLoS ONE 7, e44661 (2012).
Terashima, M. et al. KIAA1199 interacts with glycogen phosphorylase kinase beta-subunit (PHKB) to promote glycogen breakdown and cancer cell survival. Oncotarget 5, 7040–7050 (2014).
Abdelgawad, M. E. et al. Early reversal cells in adult human bone remodeling: osteoblastic nature, catabolic functions and interactions with osteoclasts. Histochem. Cell. Biol. 145, 603–615 (2016).
Simonsen, J. L. et al. Telomerase expression extends the proliferative life-span and maintains the osteogenic potential of human bone marrow stromal cells. Nat. Biotechnol. 20, 592–596 (2002).
Chen, L. et al. Delta-like 1/fetal antigen-1 (Dlk1/FA1) is a novel regulator of chondrogenic cell differentiation via inhibition of the Akt kinase-dependent pathway. J. Biol. Chem. 286, 32140–32149 (2011).
Chen, L. et al. MicroRNA-34a inhibits osteoblast differentiation and in vivo bone formation of human stromal stem cells. Stem Cells 32, 902–912 (2014).
Chen, L. et al. Inhibiting actin depolymerization enhances osteoblast differentiation and bone formation in human stromal stem cells. Stem Cell Res 15, 281–289 (2015).
Kristensen, H. B., Andersen, T. L., Marcussen, N., Rolighed, L. & Delaisse, J. M. Osteoblast recruitment routes in human cancellous bone remodeling. Am. J. Pathol. 184, 778–789 (2014).
Liang, C. C., Park, A. Y. & Guan, J. L. In vitro scratch assay: a convenient and inexpensive method for analysis of cell migration in vitro. Nat. Protoc. 2, 329–333 (2007).
Justus, C. R., Leffler, N., Ruiz-Echevarria, M. & Yang, L. V. In vitro cell migration and invasion assays. J. Vis. Exp. https://doi.org/10.3791/51046 (2014).
Tiwari, A. et al. Early insights into the function of KIAA1199, a markedly overexpressed protein in human colorectal tumors. PLoS ONE 8, e69473 (2013).
Evensen, N. A. et al. Unraveling the role of KIAA1199, a novel endoplasmic reticulum protein, in cancer cell migration. J. Natl Cancer Inst. 105, 1402–1416 (2013).
Seton-Rogers, S. Cell migration: changing shape. Nat. Rev. Cancer 15, 70 (2015). nrc3902 [pii].
Kasza, K. E. & Zallen, J. A. Dynamics and regulation of contractile actin-myosin networks in morphogenesis. Curr. Opin. Cell Biol. 23, 30–38 (2011).
Doherty, G. J. & McMahon, H. T. Mediation, modulation, and consequences of membrane-cytoskeleton interactions. Annu Rev. Biophys. 37, 65–95 (2008).
Neth, P. et al. The Wnt signal transduction pathway in stem cells and cancer cells: influence on cellular invasion. Stem Cell Rev. 3, 18–29 (2007).
Neth, P. et al. Wnt signaling regulates the invasion capacity of human mesenchymal stem cells. Stem Cells 24, 1892–1903 (2006). 10.1634/stemcells.2005-0503.
Devreotes, P. & Horwitz, A. R. Signaling networks that regulate cell migration. Cold Spring Harb. Perspect. Biol. 7, a005959 (2015). a005959 [pii] 7/8/a005959 [pii].
Andersen, T. L. et al. Understanding coupling between bone resorption and formation: are reversal cells the missing link? Am. J. Pathol. 183, 235–246 (2013).
Kristensen, H. B., Andersen, T. L., Marcussen, N., Rolighed, L. & Delaisse, J. M. Increased presence of capillaries next to remodeling sites in adult human cancellous bone. J. Bone Miner. Res. 28, 574–585 (2013).
Larsen, K. H., Frederiksen, C. M., Burns, J. S., Abdallah, B. M. & Kassem, M. Identifying a molecular phenotype for bone marrow stromal cells with in vivo bone-forming capacity. J. Bone Miner. Res. 25, 796–808 (2010).
Jia, S. et al. KIAA1199 promotes migration and invasion by Wnt/beta-catenin pathway and MMPs mediated EMT progression and serves as a poor prognosis marker in gastric cancer. PLoS ONE 12, e0175058 (2017).
Kuure, S. et al. Actin depolymerizing factors cofilin1 and destrin are required for ureteric bud branching morphogenesis. PLoS Genet. 6, e1001176 (2010).
Vlad-Fiegen, A., Langerak, A., Eberth, S. & Muller, O. The Wnt pathway destabilizes adherens junctions and promotes cell migration via beta-catenin and its target gene cyclin D1. FEBS Open Bio 2, 26–31 (2012).
Aman, A. & Piotrowski, T. Wnt/beta-catenin and Fgf signaling control collective cell migration by restricting chemokine receptor expression. Dev. Cell. 15, 749–761 (2008).
Abdallah, B. M. & Kassem, M. New factors controlling the balance between osteoblastogenesis and adipogenesis. Bone 50, 540–545 (2012).
Bikkavilli, R. K. & Malbon, C. C. Mitogen-activated protein kinases and Wnt/beta-catenin signaling: Molecular conversations among signaling pathways. Commun. Integr. Biol. 2, 46–49 (2009).
Chang, J. et al. Noncanonical Wnt-4 signaling enhances bone regeneration of mesenchymal stem cells in craniofacial defects through activation of p38 MAPK. J. Biol. Chem. 282, 30938–30948 (2007).
Caverzasio, J. & Manen, D. Essential role of Wnt3a-mediated activation of mitogen-activated protein kinase p38 for the stimulation of alkaline phosphatase activity and matrix mineralization in C3H10T1/2 mesenchymal cells. Endocrinology 148, 5323–5330 (2007).
Bikkavilli, R. K., Feigin, M. E. & Malbon, C. C. p38 mitogen-activated protein kinase regulates canonical Wnt-beta-catenin signaling by inactivation of GSK3beta. J. Cell. Sci. 121, 3598–3607 (2008).
Cervenka, I. et al. Mitogen-activated protein kinases promote WNT/beta-catenin signaling via phosphorylation of LRP6. Mol. Cell. Biol. 31, 179–189 (2011).
Yoshida, M. et al. Expression analysis of three isoforms of hyaluronan synthase and hyaluronidase in the synovium of knees in osteoarthritis and rheumatoid arthritis by quantitative real-time reverse transcriptase polymerase chain reaction. Arthritis Res. Ther. 6, R514–R520 (2004).
Avigdor, A. et al. CD44 and hyaluronic acid cooperate with SDF-1 in the trafficking of human CD34+stem/progenitor cells to bone marrow. Blood 103, 2981–2989 (2004).
Nagaoka, A. et al. Regulation of Hyaluronan (HA) Metabolism Mediated by HYBID (Hyaluronan-binding Protein Involved in HA Depolymerization, KIAA1199) and HA Synthases in Growth Factor-stimulated Fibroblasts. J. Biol. Chem. 290, 30910–30923 (2015).
Shimoda, M. et al. Hyaluronan-binding protein involved in hyaluronan depolymerization controls endochondral ossification through hyaluronan metabolism. Am. J. Pathol. 187, 1162–1176 (2017). 10.1016/j.ajpath.2017.01.005.
Zhu, H. et al. The role of the hyaluronan receptor CD44 in mesenchymal stem cell migration in the extracellular matrix. Stem Cells 24, 928–935 (2006).
Acknowledgements
We thanks for Adiba Isa and Lifang Hu at the suggestion for the protocols to establish the cell migration assays, as well as Lone Christiansen, Kaja Laursen, and Birgit MacDonald for technical assistance. The project was supported by the NovoNordisk foundation (NNF15OC0016284), the Lundbeck foundation (R266–2017–4250), and a scholarship from Odense University Hospital PhD-grant and the University of Southern Denmark.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
All the authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, L., Shi, K., Andersen, T.L. et al. KIAA1199 is a secreted molecule that enhances osteoblastic stem cell migration and recruitment. Cell Death Dis 10, 126 (2019). https://doi.org/10.1038/s41419-018-1202-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-018-1202-9
- Springer Nature Limited
This article is cited by
-
Genome-wide investigation of lncRNAs revealed their tight association with gastric cancer
Journal of Cancer Research and Clinical Oncology (2024)
-
A Prognostic Methylation-Driven Two-Gene Signature in Medulloblastoma
Journal of Molecular Neuroscience (2024)
-
Combining single-cell transcriptomics and CellTagging to identify differentiation trajectories of human adipose-derived mesenchymal stem cells
Stem Cell Research & Therapy (2023)
-
KIAA1199 deficiency enhances skeletal stem cell differentiation to osteoblasts and promotes bone regeneration
Nature Communications (2023)
-
Secreted KIAA1199 promotes the progression of rheumatoid arthritis by mediating hyaluronic acid degradation in an ANXA1-dependent manner
Cell Death & Disease (2021)