Abstract
Centriole duplication is tightly controlled to occur once per cell cycle, and disruption of this synchrony causes centriole amplification, which is frequently observed in many cancers. Our previous work showed that nuclear distribution gene C (NudC)-like protein 2 (NudCL2) localizes to centrosomes; however, little is known about the role of NudCL2 in the regulation of centrosome function. Here, we find that NudCL2 is required for accurate centriole duplication by stabilizing the E3 ligase HECT domain and RCC1-like domain-containing protein 2 (HERC2). Knockout (KO) of NudCL2 using CRISPR/Cas9-based genome editing or depletion of NudCL2 using small interfering RNA causes significant centriole amplification. Overexpression of NudCL2 significantly suppresses hydroxyurea-induced centriole overduplication. Quantitative proteomic analysis reveals that HERC2 is downregulated in NudCL2 KO cells. NudCL2 is shown to interact with and stabilize HERC2. Depletion of HERC2 leads to the similar defects to that in NudCL2-downregulated cells, and ectopic expression of HERC2 effectively rescues the centriole amplification caused by the loss of NudCL2, whereas the defects induced by HERC2 depletion cannot be reversed by exogenous expression of NudCL2. Either loss of NudCL2 or depletion of HERC2 leads to the accumulation of ubiquitin-specific peptidase 33 (USP33), a centrosomal protein that positively regulates centriole duplication. Moreover, knockdown of USP33 reverses centriole amplification in both NudCL2 KO and HERC2-depleted cells. Taken together, our data suggest that NudCL2 plays an important role in maintaining the fidelity of centriole duplication by stabilizing HERC2 to control USP33 protein levels, providing a previously undescribed mechanism restraining centriole amplification.
Similar content being viewed by others
Introduction
Centrioles are cylindrical, microtubule-based structures that are essential for the formation of centrosomes, cilia, and flagella1,2. In general, a pair of centrioles recruits and organizes the pericentriolar material (PCM) to form a mature centrosome, which plays important roles in regulating cell shape, polarity, motility, mitosis, and cytokinesis1,3. In mammalian cells, centriole duplication occurs once per cell cycle, and is highly coordinated with DNA replication4. Centriole duplication consists of sequential steps that occur in a semiconservative and cell cycle-dependent manner. First, a pair of centrioles disengages during mitotic exit and early G1 phase. Then, a single new centriole, termed procentriole, begins to assemble adjacent to each pre-existing parental centriole in the G1/S transition, elongates through S and G2 phases, and finally becomes fully mature in the early phase of mitosis4,5. During this process, accurate control of the centriole number is crucial to maintain the number of centrosomes. Dysregulation of this process may cause centriole amplification, which leads to extranumerary centrosomes and results in genome instability2,4.
Nuclear distribution gene C (NudC)-like protein 2 (NudCL2) was cloned and characterized as a new homolog of NudC in mammalian cells6. In the filamentous fungus Aspergillus nidulans, NudC was first identified as an upstream regulator of NudF (a homolog of the human lissencephaly 1 gene product, LIS1) in the control of nuclear movement7,8. Mammalian NudC plays crucial roles in diverse cellular processes including cell division, neuronal migration, and ciliogenesis9. Our previous study has reported that NudCL2 plays an important role in regulating the LIS1/dynein pathway by enhancing the interaction between LIS1 and heat-shock protein 90 (Hsp90) to stabilize LIS16. Additional data demonstrated that NudCL2 functions as an Hsp90 cochaperone to stabilize subunits of the ring-shaped cohesin complex that mediates sister chromatid cohesion10. Interestingly, NudCL2 has been shown to be associated with centrosomes in human cells6; however, little is known about the potential role of NudCL2 in centrosomes.
In this study, we provide evidence that NudCL2 is crucial for accurate centriole duplication. NudCL2 interacts with and stabilizes the E3 ligase HECT domain and RCC1-like domain-containing protein 2 (HERC2). Either downregulation of NudCL2 or depletion of HERC2 causes centriole amplification. Ectopic expression of HERC2 reverses the centriole defects induced by NudCL2 deletion, but not vice versa. Thus, these data suggest that NudCL2 suppresses centriole amplification by stabilizing HERC2.
Results
NudCL2 is associated with centrosomes
Our previous work indicated that NudCL2 is localized to centrosomes in HeLa cells6. To confirm the centrosome localization of NudCL2 during cell cycle progression, we performed immunofluorescence staining with anti-NudCL2 and anti-γ-tubulin (a centrosomal marker) antibodies in U2OS and HeLa cells and found that NudCL2 was colocalized with γ-tubulin throughout the cell cycle (Fig. 1a, b). Subsequent results revealed that NudCL2 was also associated with green-fluorescent protein (GFP)-centrin (a centriolar marker) that was stably expressed in HeLa cells (Fig. 1c). Moreover, sucrose density gradient centrifugation experiments showed that NudCL2 was cosedimented with γ-tubulin and centrosomal protein of 110 kDa (CP110) in U2OS cells (Fig. 1d). These data indicate that NudCL2 is a centrosome-associated protein.
a, b U2OS and HeLa cells were fixed and subjected to immunofluorescence analyses with anti-NudCL2 (green) and anti-γ-tubulin (red) antibodies, respectively. c NudCL2 is colocalized with centrin. HeLa cells stably expressing green-fluorescent protein (GFP)-centrin were fixed and processed for immunofluorescence analysis with anti-NudCL2 (red) antibody. Higher magnifications of the boxed regions are displayed. d NudCL2 is cosedimented with centrosomes. U2OS cells were treated with 10 μg/ml nocodazole and 5 μg/ml cytochalasin B for 1.5 h. Lysates of the cells were fractionated by sucrose gradient centrifugation and applied for western blot analysis with the indicated antibodies. DNA was visualized with 4′,6-diamidino-2-phenylindole (DAPI) (blue). Scale bars, 10 μm
Downregulation of NudCL2 causes centriole amplification
To explore the role of NudCL2 in centrosome function, we employed CRISPR/Cas9-mediated genome editing to knock out NudCL2 in mammalian cells. A CRISPR/Cas9 plasmid with a short guide RNA (sgRNA) that recognizes the first exon of NudCL2 was constructed and transfected into U2OS cells (Fig. 2a). PCR amplification of genomic DNA followed by Sanger sequencing revealed indels that are predicted to cause frameshift mutations at the NudCL2 DNA locus (Fig. 2b). Immunoblotting confirmed that NudCL2 protein disappeared in the mutant cells (Fig. 2c). In NudCL2 knockout (KO) cells at interphase, the number of cells with more than four centrin, four CP110, or two γ-tubulin dots increased approximately three-fold compared to the wild-type (WT) cells (Fig. 2d–h), suggesting that loss of NudCL2 causes centriole amplification. The similar results were observed in NudCL2 KO DLD1 cells and NudCL2-depleted CAL51 cells (Supplementary Figs. 1 and 2). Moreover, the increase in centriole number observed in NudCL2 KO cells was significantly reversed by ectopic expression of NudCL2 (Fig. 2i–l). Given that cell cycle arrest may induce centriole amplification2,11, we determined whether centriole amplification induced by NudCL2 deletion resulted from a change in cell cycle progression in NudCL2 KO cells. Fluorescence-activated cell sorting (FACS) analysis showed that there was no significant difference between the WT and NudCL2 KO cells (Fig. 2m, n). Together, these data indicate that NudCL2 plays an important role in restraining centriole amplification.
a Schematic representation of NudCL2 gene targeting strategy. b Indel mutations of the NudCL2 DNA locus in two NudCL2 knockout cell lines. c Western blot analysis of NudCL2 protein in control and NudCL2 KO U2OS cells. β-actin, a loading control. d–f Control and NudCL2 KO U2OS cells were fixed and processed for immunofluorescence analysis with anti-centrin (green) and anti-CP110 (red) antibodies. Higher magnifications of the boxed regions are displayed. The frequencies of cells with more than four centrin and four CP110 dots were calculated, respectively. g, h Control and NudCL2 KO U2OS cells were fixed and stained with anti-γ-tubulin (green) and anti-CP110 (red) antibodies. Higher magnifications of the boxed regions are shown. The number of cells with more than two γ-tubulin dots was plotted. i–l Control and NudCL2 KO U2OS cells were transfected with green-fluorescent protein (GFP)-NudCL2 or GFP vector for 48 h and subjected to western blot and immunofluorescence analyses, respectively. β-actin, a loading control. The frequencies of cells with more than four centrin, four CP110, and two γ-tubulin dots were plotted, respectively. m, n The cell cycle distribution of control and NudCL2 KO U2OS cells was analyzed by flow cytometry. o, p Cells were fixed and immunostained with anti-α-tubulin (green) and anti-CP110 (red) antibodies. Representative images of mitotic cells with bipolar, pseudobipolar, multipolar, or monopolar spindles are shown. The percentages of cells with various mitotic phenotypes were calculated. DNA was visualized with 4′,6-diamidino-2-phenylindole (DAPI) (blue). Scale bars, 10 μm. Quantitative data are expressed as the mean ± SD (at least three independent experiments). More than 150 cells were counted in each experiment. *p < 0.05, **p < 0.01, ***p < 0.001, ns, no significance, Student’s t test
Centrosomes are essential for bipolar spindle assembly and accurate chromosome segregation in mammalian cells1. Centriole amplification leads to supernumerary centrosomes in the subsequent cell cycle, which cluster to generate pseudobipolar spindles after transient spindle multipolarity, promoting chromosome missegregation2,12,13,14,15. To investigate the effects of NudCL2 deletion on mitotic spindle formation, we performed immunostaining analysis with anti-α-tubulin and anti-CP110 antibodies. The data showed that the frequency of cells exhibiting pseudobipolar spindles was significantly increased in NudCL2 KO cells compared with that in WT cells (Fig. 2o, p), implying that loss of NudCL2 influences the formation of bipolar spindles.
Overexpression of NudCL2 suppresses centriole overduplication
Treatment with hydroxyurea (HU, a DNA synthesis inhibitor) uncouples centriole duplication from DNA replication and induces multiple rounds of centriole duplication in a prolonged S phase in U2OS cells, resulting in centriole amplification16,17. To confirm the role of NudCL2 in restraining centriole amplification, U2OS cells were transfected with Myc-NudCL2 followed by HU treatment. We found that NudCL2 overexpression did not obviously affect cell cycle progression as determined by FACS analysis (Fig. 3a, b). Immunostaining analyses with anti-centrin and anti-CP110 antibodies revealed that, upon HU treatment, the percentage of cells with more than four centrioles was significantly lower in Myc-NudCL2-expressing cells than in control cells (Fig. 3c–f), indicating that overexpression of NudCL2 suppresses centriole overduplication induced by HU treatment.
U2OS cells transfected with Myc-NudCL2 or Myc vector were treated with or without hydroxyurea (HU) for 48 h, and then subjected to the following analyses. a Western blot analysis of the expression of Myc-NudCL2. β-actin, a loading control. b Flow cytometry analysis of the cell cycle distribution. c–f Immunofluorescence analyses were carried out by using anti-centrin (green) or anti-CP110 (red) antibodies. Higher magnifications of the boxed regions are shown. The frequencies of cells with more than four centrin or four CP110 dots were plotted. DNA was visualized with 4′,6-diamidino-2-phenylindole (DAPI) (blue). Scale bars, 10 μm. Quantitative data are expressed as the mean ± SD (at least three independent experiments). More than 300 cells were counted in each experiment. **p < 0.01, ns, no significance, Student’s t test
Downregulation of NudCL2 induces HERC2 degradation
Previous work in our lab indicated that NudCL2 plays an important role in the regulation of protein stability6,10. To determine the mechanism of NudCL2 in centriole duplication, we performed isobaric tags for relative and absolute quantitation (iTRAQ)-based quantitative proteomic analysis using NudCL2 KO cells, and found that 61 proteins were differentially expressed (KO-1/WT fold change >1.2 or <0.83, p < 0.05) (Fig. 4a, b). A range of centrosomal proteins have been reported to regulate centriole duplication1. To identify the potential centrosomal proteins that may be involved in the function of NudCL2 in suppressing centriole amplification, we compared our quantitative proteomics data with centrosome-associated proteins from centrosome database (http://centrosome.dacya.ucm.es)18. We found that five centrosomal proteins overlapped in the two datasets, including HERC2, heat-shock cognate B, heat-shock protein family A member 2, mitogen-activated protein kinase kinase 1, and regulator of microtubule dynamics 1 (Fig. 4c, d). Among these proteins, only HERC2 was downregulated in NudCL2 KO cells, suggesting that HERC2 may be stabilized by NudCL2.
a Schematic representation of the isobaric tags for relative and absolute quantitation (iTRAQ)-based quantitative proteomic analysis. b Volcano plot showing p values (−log 10) versus the protein ratio of KO-1/WT cells (log 2). Proteins exhibiting a fold change >1.2 and p <0.05 were defined as “upregulation” (red dots), and those with a fold change <0.83 and p <0.05 were shown as “downregulation” (blue dots). Others were defined as “no change” (gray dots). c Venn diagram showing the overlap between the differentially expressed proteins from the quantitative proteomic analysis and the centrosomal proteins from the centrosome database. d Five centrosomal proteins that overlap in Fig. 4c are shown. e Western blot analysis of HERC2 protein in control and NudCL2 KO U2OS cells using the indicated antibodies. β-Actin, a loading control. f Quantitative reverse transcription PCR (RT-PCR) analysis of HERC2 messenger RNA (mRNA) in control and NudCL2 KO U2OS cells. Quantitative data are expressed as the mean ± SD (at least three independent experiments). ns, no significance, Student’s t test. g Control and NudCL2 KO U2OS cells were transfected with Myc-NudCL2 or Myc vector for 48 h and subjected to western blot analysis with anti-HERC2 and anti-NudCL2 antibodies. β-actin, a loading control. h Control and NudCL2 KO U2OS cells were treated with MG132 or dimethyl sulfoxide (DMSO) for 2 h. Lysates of the cells were applied for western blot analysis with anti-HERC2 and anti-NudCL2 antibodies. β-actin, a loading control. i U2OS cells transfected with the indicated vectors for 48 h were harvested and lysed in TBSN (20 mM Tris [pH 8.0], 150 mM NaCl, 0.5% Nonidet P-40, 5 mM EGTA, 1.5 mM EDTA, 0.5 mM Na3VO4, 20 mM p-nitrophenyl phosphate) lysis buffer. Immunoprecipitation analysis was performed using the indicated antibodies. j HEK-293 cells were harvested and lysed. Immunoprecipitation analysis was carried out using the indicated antibodies
To confirm the downregulation of HERC2 induced by NudCL2 deletion, we performed western blot and reverse transcription PCR (RT-PCR) experiments. The results showed that the protein level of HERC2, but not its messenger RNA (mRNA) level, was substantially reduced in NudCL2 KO U2OS cells compared to WT cells (Fig. 4e, f). In addition, KO of NudCL2 in DLD1 cells or depletion of NudCL2 in CAL51 cells also resulted in an obvious decrease in HERC2 protein levels (Supplementary Fig. 3). The reduction in HERC2 protein levels was rescued by ectopic expression of NudCL2 in NudCL2 KO cells (Fig. 4g). In addition, cycloheximide (CHX) chase analysis revealed that the rate of HERC2 degradation was faster in NudCL2 KO cells than in WT cells (Supplementary Fig. 4), implying that NudCL2 may be involved in the regulation of HERC2 protein stability. Furthermore, we employed the proteasome inhibitor MG132 to treat NudCL2 KO cells and found that HERC2 degradation was decreased (Fig. 4h). Additional immunoprecipitation data showed that NudCL2 interacted with HERC2 (Fig. 4i, j). Together, these data strongly imply that NudCL2 plays an important role in stabilizing HERC2.
HERC2 is involved in the regulation of centriole duplication in NudCL2 KO cells
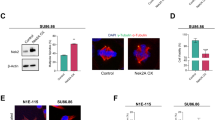
HERC2 has been reported to participate in many fundamental cellular processes, including DNA repair, DNA replication, and PCM morphology maintenance19. Nevertheless, whether HERC2 plays a role in centriole duplication is still unknown. To examine the role of HERC2 in centriole duplication, we employed two small interfering RNAs (siRNAs) targeting different HERC2 mRNA regions (siHERC2-1 and -2). Western blot analysis showed that the HERC2 proteins were efficiently downregulated at 72 h post transfection (Fig. 5a). Importantly, immunostaining with anti-centrin and anti-CP110 antibodies revealed an approximately three-fold increase in the number of cells with more than four centrioles in HERC2-depleted cells compared to the control (Fig. 5b–d). Flow cytometry data showed that knockdown of HERC2 had no significant effect on cell cycle progression (Fig. 5e, f). Furthermore, downregulation of HERC2 resulted in a significant increase in mitotic cells with pseudobipolar spindles (Fig. 5g, h). Thus, these results strongly suggest that HERC2 plays an important role in restraining centriole amplification.
a U2OS cells were transfected with control or HERC2 small interfering RNAs (siRNAs) for 72 h and subjected to western blot analysis using anti-HERC2 antibody. β-actin, a loading control. b–d U2OS cells transfected with the indicated siRNAs were fixed and processed for immunofluorescence analysis with anti-centrin (green) and anti-CP110 (red) antibodies. Higher magnifications of the boxed regions are displayed. The frequencies of cells with more than four centrin and four CP110 dots were calculated, respectively. e, f Flow cytometry analysis of the cell cycle distribution in control and HERC2-depleted cells. g, h Cells were fixed and subjected to immunostaining analysis with anti-α-tubulin (green) and anti-CP110 (red) antibodies. Representative images of mitotic cells are shown. The frequencies of cells with various phenotypes were scored. i–k Control and NudCL2 KO U2OS cells transfected with Flag-HERC2 or Flag vector for 48 h were subjected to western blot and immunofluorescence analyses using the indicated antibodies. β-actin, a loading control. The frequencies of cells with more than 4 centrin or 4 CP110 dots were plotted. l–n U2OS cells transfected with vectors and siRNAs for 72 h were subjected to western blot and immunofluorescence analyses using the indicated antibodies. β-actin, a loading control. The frequencies of cells with more than four centrin and four CP110 dots were plotted, respectively. DNA was visualized with 4′,6-diamidino-2-phenylindole (DAPI). Scale bars, 10 μm. Quantitative data are expressed as the mean ± SD (at least three independent experiments). More than 150 cells were counted in each experiment. *p < 0.05, **p < 0.01, ***p < 0.001, ns, not significanct, Student’s t test
Since our data show that NudCL2 interacts with and stabilizes HERC2 and that both NudCL2 downregulation and HERC2 depletion lead to centriole amplification (Figs. 2, 4, and 5), we next asked whether NudCL2 negatively regulates centriole duplication through HERC2. Exogenous expression of HERC2 efficiently reversed centriole amplification in NudCL2 KO cells (Fig. 5i–k). By contrast, ectopic expression of NudCL2 failed to rescue the defects induced by HERC2 depletion (Fig. 5l–n). Collectively, these data suggest that HERC2 functions as downstream of NudCL2 to restrain centriole amplification.
USP33 is involved in the HERC2-mediated regulation of centriole duplication in NudCL2 KO cells
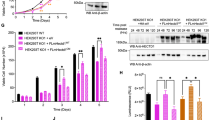
Given that the E3 ligase HERC2 target protein ubiquitin-specific peptidase 33 (USP33) plays an important role in the positive regulation of centriole duplication20,21, we asked whether USP33 acts as a downstream effector of the NudCL2/HERC2 axis to restrain centriole amplification during the cell cycle. First, we tested USP33 protein levels in HERC2-depleted cells and found that USP33 accumulated after HERC2 knockdown (Fig. 6a), consistent with a previous study20. Downregulation of USP33 reversed centriole amplification in HERC2-depleted cells (Fig. 6b–d), suggesting that HERC2 may control accurate centriole duplication through USP33. Next, we examined whether the protein stability of USP33 was regulated by NudCL2 and found that USP33 was obviously increased in NudCL2 KO cells compared to the control (Fig. 6e). Further analyses showed that NudCL2 KO decreased the degradation of USP33 (Supplementary Fig. 5). Importantly, ectopic expression of HERC2 reversed centriole amplification and USP33 accumulation induced by NudCL2 deletion (Figs. 5i–k, 6f). Knockdown of USP33 suppressed centriole amplification in NudCL2 KO cells (Fig. 6g–i). Moreover, immunoprecipitation experiments showed that HERC2 was associated with NudCL2 and USP33 (Fig. 6j). Taken together, our results indicate that USP33 functions as a downstream effector of the NudCL2/HERC2 pathway to restrain centriole overduplication (Fig. 6k).
a Western blot analysis of USP33 protein levels in control and HERC2-depleted U2OS cells. β-actin, a loading control. b–d U2OS cells transfected with the indicated small interfering (siRNAs) for 48 h were processed for western blot and immunofluorescence analyses with the indicated antibodies. β-actin, a loading control. The frequencies of cells with more than four centrin and four CP110 dots were calculated, respectively. e Western blot analysis of USP33 protein levels in control and NudCL2 KO U2OS cells. β-actin, a loading control. f Cells transfected with Flag-HERC2 or Flag vector for 48 h were subjected to western blot analysis with the indicated antibodies. β-actin, a loading control. g–i Control and NudCL2 KO U2OS cells transfected with control or USP33 siRNAs for 48 h were subjected to western blot and immunofluorescence analyses with the indicated antibodies. β-actin, a loading control. The frequencies of cells with more than four centrin and four CP110 dots were plotted, respectively. j U2OS cells transfected with Flag-HERC2 or Flag vector for 48 h were harvested and subjected to immunoprecipitation analysis using the indicated antibodies. Quantitative data are expressed as the mean ± SD (at least three independent experiments). More than 300 cells were counted in each experiment. *p < 0.05, **p < 0.01, Student’s t test. k Working model for the role of NudCL2 in centriole duplication. NudCL2 restrains centriole amplification by stabilizing HERC2. Knockout of NudCL2 leads to HERC2 degradation and USP33 accumulation, resulting in centriole amplification
Discussion
Centriole duplication is tightly controlled during cell cycle progression1. Accumulating studies have revealed that a number of proteins are involved in restraining centriole amplification, such as Krüppel-like factor 14, RNA-binding motif protein 14, cell division cycle 6, and so on22,23,24,25,26,27. However, the molecular mechanism of suppressing centriole amplification is still a mystery. In this report, we provide evidence that NudCL2 plays an essential role in restraining centriole amplification in mammalian cells. Downregulation of NudCL2 results in extranumerary centrioles (Fig. 2, Supplementary Figs. 1 and 2), and overexpression of NudCL2 inhibits HU-induced centriole overduplication (Fig. 3). NudCL2 is found to interact with and stabilize HERC2 (Fig. 4). Ectopic expression of HERC2 reverses centriole amplification induced by NudCL2 deletion (Fig. 5i–k). Either loss of NudCL2 or depletion of HERC2 increases USP33 protein levels (Fig. 6a, e, f). Moreover, downregulation of USP33 reverses centriole amplification induced by NudCL2 KO or HERC2 knockdown (Figs. 6b–d, g–i). Thus, these data indicate a hitherto undescribed mechanism in which NudCL2 functions as an important checkpoint protein to restrain centriole amplification by stabilizing HERC2 to decrease USP33 (Fig. 6k).
Emerging studies have identified a number of centrosomal proteins such as polo-like kinase 1 (Plk1), polo-like kinase 4 (Plk4), spindle assembly abnormal protein 6 (SAS6), separase, and SCL/TAL1-interrupting locus (STIL) that play crucial roles in the centriole duplication cycle1. During centriole biogenesis, Plk4 is recruited to the base of centrioles by centrosomal protein of 152 kDa and centrosomal protein of 192 kDa28,29. Then, Plk4 phosphorylates STIL and triggers the recruitment of SAS6 to centrioles, which initiates centriole assembly30,31,32. Additionally, Plk1 and separase participate in centriole disengagement, a critical licensing step for centriole duplication in the next cell cycle33. Overexpression of these proteins induces centriole amplification34,35,36,37,38. Here, we found that the NudCL2/HERC2/USP33 axis is essential for the precision regulation of centriole duplication. Intriguingly, our data reveal that NudCL2 deletion has no obvious effect on the protein levels of the above regulators, including Plk1, Plk4, SAS6, separase, and STIL (Supplementary Fig. 6), suggesting an unknown mechanism underlying the role of NudCL2 in centriole duplication.
Accumulating data implicate that HERC2 plays important roles in many cellular processes. HERC2 is required for the proper maturation of double-strand break responses by promoting the retention of DNA repair factors on damaged chromosomes39. HERC2 promotes DNA replication by facilitating minichromosome maintenance complex component 2 phosphorylation40. In addition, HERC2 has been found to participate in cell cycle regulation by inhibiting G2–M checkpoint activity via destabilizing breast cancer type 1 susceptibility protein41. Recently, HERC2 has been reported to localize to centrosomes and maintain PCM morphology by interacting with and ubiquitinating neuralized E3 ubiquitin protein ligase 442; however, whether HERC2 plays a role in centriole duplication remains unknown. Here, our data show that downregulation of HERC2 leads to centriole amplification and pseudobipolar spindle formation in mammalian cells, suggesting a role of HERC2 in the accurate control of centriole duplication (Fig. 5). USP33 is a HERC2 target protein and participates in positively regulating centriole duplication20,21. In this study, we find that USP33 is accumulated in HERC2-depleted cells. Downregulation of USP33 is able to effectively reverse centriole amplification induced by HERC2 knockdown. These data suggest a previously uncharacterized function of HERC2 in restraining centriole amplification by destabilizing USP33.
In this study, we found that downregulation of NudCL2 causes HERC2 degradation (Fig. 4); however, the mechanism underlying the stabilization of HERC2 regulated by NudCL2 is still unknown. NudCL2 contains a core structure of p23 (p23 domain), which acts as an Hsp90 cochaperone to regulate the folding and maturation of client proteins9. Our recent work showed that NudCL2 functions as an Hsp90 cochaperone to stabilize cohesin subunits by modulating Hsp90 ATPase activity10. Based on these scenarios, further studies are clearly needed to determine whether Hsp90 is involved in the regulation of HERC2 stability.
Vertebrate NudC has three homologs: NudC, NudCL, and NudCL2, all of which have been reported to localize at centrosomes, an important organelle for cell cycle and ciliogenesis6,43,44,45. Depletion of NudC leads to multiple mitotic defects including the multipolar spindles and the lagging chromosomes, and causes ciliary defects in mammalian cells and zebrafish46,47. Downregulation of NudCL results in the similar phenotypes to that of NudC depletion during mitosis48. NudCL depletion also influences cilia assembly in interphase49. Here, our data indicate that NudCL2 acts as a centrosome duplication checkpoint protein to restrain centriole amplification by destabilizing HERC2. Taken together, these studies suggest that members of the NudC family function as important regulators of centrosome function.
Materials and methods
Plasmids, primers, and oligonucleotides
The GFP-NudCL2 and Myc-NudCL2 plasmids were constructed as described previously6. Plasmids expressing Flag-tagged full-length HERC2 were obtained from Addgene (55613)20, and the sequence of HERC2 was confirmed by DNA sequencing. All siRNAs were designed or synthesized by GenePharma (Shanghai, China). The sense sequences of the siRNA duplexes are as follows:
siHERC2-1: 5′-GGAAAGCACUGGAUUCGUUTT-3′30;
siHERC2-2: 5′-GAAGGUGGCUGUUCACUCATT-3′30;
siUSP33: 5′-GAUCAUGUGGCGAAGCAUATT-3′21;
siNudCL2-1: 5′-ACCUUGAGAAAUAACTGCUTT-3′10;
siNudCL2-2: 5′-GACUUCUCUACUAGAAUCUTT-3′.
Cell culture, transfection, and drug treatment
U2OS, HeLa, and HEK-293 cells were maintained in Dulbecco’s modified Eagle’s medium (Corning, Shanghai, China) supplemented with 10% fetal bovine serum (PAA Laboratories, Northbrook, IL, USA). The human colorectal adenocarcinoma cell line DLD1 and the human breast cancer cell line CAL51 were cultured in RPMI-1640 (Corning) supplemented with 10% fetal bovine serum (PAA Laboratories). Cells were cultured in humidified incubators at 37 °C with 5% CO2. Transfections of plasmids and siRNAs were carried out using polyjet (SignaGen Laboratories, Rockville, MD, USA) and Lipofectamine RNAiMAX (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions, respectively. In the centriole-overduplication assay, 4 mM HU (Sigma-Aldrich, St. Louis, MO, USA) was used to treat U2OS cells for 48 h. For CHX (Sigma-Aldrich, St. Louis, MO, USA) chase analysis, 100 µg/ml CHX was used for the indicated times as described in the text. To block the proteasome-dependent degradation pathway, 1 µM MG132 (Millipore, Billerica, MA, USA) was added to U2OS cells for 2 h.
Generation of NudCL2 KO cell lines by CRISPR/Cas9-mediated genome editing
The sgRNA, 5′-GAAGTTCAGGTGCCGCC-3′, targeting the first exon of the NudCL2 gene was designed and synthesized by Nanjing YSY Biotech Ltd (Nanjing, China). Then, the CRISPR/Cas9 plasmid was constructed by cloning the sgRNA into its backbone. U2OS, HEK-293, and DLD1 cells were transfected with this plasmid for 48 h followed by treatment with 1 μg/ml puromycin for 48 h. After selection, the cells were counted and diluted to a density of 1 cell per 200 µl of medium and seeded into 96-well plates to obtain single colonies. NudCL2 KO colonies were identified by western blot and genomic DNA sequencing analyses. The primers used to amplify the target region are as follows:
forward: 5′-AGGCGTAGCCTAAGCGTGGGATTC-3′;
reverse: 5′-ACCCAACAGTCGTTCAGGGAAACG-3′.
Antibodies
An anti-NudCL2 antibody was generated as described previously6. Antibodies against centrin 1 (Millipore, Billerica, MA, USA), CP110 (Proteintech, Wuhan, China), γ-tubulin (Sigma-Aldrich, St. Louis, MO, USA), α-tubulin (Sigma-Aldrich, St. Louis, MO, USA), β-actin (Sigma-Aldrich, St. Louis, MO, USA), Plk1 (Sigma-Aldrich, St. Louis, MO, USA), Flag (Beyotime Biotechnology, Shanghai, China), c-Myc (Santa Cruz Biotechnology, CA, USA), SAS6 (Santa Cruz Biotechnology, CA, USA), separase (Santa Cruz Biotechnology, CA, USA), CDK2 (Santa Cruz Biotechnology, CA, USA), cyclin A (Santa Cruz Biotechnology, CA, USA), cyclin E (Santa Cruz Biotechnology, CA, USA), STIL (Abcam, Cambridge, MA, USA), HERC2 (BD Biosciences, San Jose, CA, USA; Bethyl Laboratories, Montgomery, TX, USA), Plk4 (Proteintech, Wuhan, China), and USP33 (Proteintech, Wuhan, China) were acquired commercially.
Immunoprecipitation and western blot
Immunoprecipitation was performed as previously described6,50. Briefly, cells were lysed in TBSN buffer (20 mM Tris [pH 8.0], 150 mM NaCl, 0.5% Nonidet P-40, 5 mM EGTA, 1.5 mM EDTA, 0.5 mM Na3VO4, 20 mM p-nitrophenyl phosphate) containing a cocktail of protease inhibitors (Roche, Basel, Switzerland) and then subjected to immunoprecipitation with the indicated antibodies or anti-Flag antibody-coupled beads (Sigma-Aldrich, St. Louis, MO, USA). The proteins were separated in a sodium dodecyl sulfate (SDS)-polyacrylamide gel electrophoresis gel and transferred to a polyvinylidene fluoride membrane (Millipore, Billerica, MA, USA). The membranes were blocked with 5% skim milk at room temperature for 1 h, then incubated with the indicated primary antibodies and corresponding horse radish peroxidase-conjugated secondary antibodies (Cell Signaling Technology, Beverly, MA, USA), and, finally, detected by enhanced chemiluminescence (Fude biological technology, Hangzhou, China) according to the manufacturer’s instructions.
Immunofluorescence
Cells grown on coverslips were fixed with cold methanol at −20 °C for 5 min, permeabilized in 0.1% Triton X-100/phosphate-buffered saline (PBS) for 15 min, blocked with 3% bovine serum albumin/PBS for 30 min, and then incubated with the primary antibodies indicated in the text for 2 h at room temperature. After washing with 0.1% Triton X-100/PBS for 15 min, cells were incubated with secondary antibodies (Alexa Fluor 488- or 568-conjugated anti-rabbit or mouse IgG, Invitrogen) for 1 h at room temperature. DNA was stained with 4′,6-diamidino-2-phenylindole (DAPI) (Beyotime Technology, Shanghai, China). Finally, the mounted coverslips were analyzed by confocal fluorescence microscopy (IX81-FV1000, Olympus, Japan).
Fluorescence-activated cell sorting
For cell cycle analysis, cells were collected and washed with PBS and fixed in cold 70% ethanol. The samples were washed twice with PBS and incubated with a staining solution containing 20 µg/ml propidium iodide (Beyotime Technology, Shanghai, China) and 20 μg/ml RNase A (Beyotime Technology, Shanghai, China) at 37 °C for 30 min. The samples were analyzed using a flow cytometer (Cytomic FC 500 MCL, Beckman Coulter, Brea, CA, USA).
Quantitative RT-PCR
Total RNA was extracted with Trizol (Invitrogen) and reverse transcribed to obtain complementary DNA with HiScript Q RT SuperMix (Vazyme, Nanjing, China). Quantitative RT-PCR analysis for HERC2 mRNA was performed using a Bio-Rad CFX-Totch System (Bio-Rad Laboratories, Hercules, CA, USA). Primers used to amplify the target region are as follows:
forward: 5′-TGAAGAAGAAACTCCTGCACCT-3′;
reverse: 5′-GGTGGTGGCTGACTGGAC-3′.
Centrosome isolation
Centrosome purification was performed as previously described with some modifications51. U2OS cells that had been treated with a solution containing 10 μg/ml nocodazole (Santa Cruz Biotechnology, CA, USA) and 5 μg/ml cytochalasin B (Sigma-Aldrich, St. Louis, MO, USA) for 1.5 h were lysed with lysis buffer (1 mM HEPES [pH 7.2], 0.5% NP-40, 0.5 mM MgC12, 0.1% β-mercaptoethanol and protease inhibitors) on ice for 20 min. Swollen nuclei and chromatin aggregates were removed by centrifugation (2500 × g, 10 min). The supernatant was supplemented with HEPES buffer to a final concentration of 10 mM and incubated with 2 Units/ml of DNase I (Sigma-Aldrich, St. Louis, MO, USA) on ice for 30 min. The lysate was underlaid with 1 ml of 60% sucrose solution and centrifuged at 25,000 × g for 30 min at 4 °C. The crude centrosome preparation was diluted with lysis buffer and layered onto a discontinuous sucrose gradient (from bottom to top, containing 0.5, 0.3, and 0.3 ml of 70%, 50%, and 40% sucrose solutions, respectively) in a 5 ml tube, followed by centrifugation at 97,000 × g for 1.5 h at 4 °C. Subsequently, fractions were collected, diluted with 10 mM PIPES (pH 7.2), and centrifuged at 14,000 rpm for 10 min to pellet the centrosomes. The centrosome pellets were resuspended in 40 µl of SDS sample buffer.
Statistical analysis
All experiments were repeated at least three times. Two-tailed Student’s t tests were used for comparisons between two groups (GraphPad Prism 5).
References
Nigg, E. A. & Holland, A. J. Once and only once: mechanisms of centriole duplication and their deregulation in disease. Nat. Rev. Mol. Cell. Biol. 19, 297–312 (2018).
Nigg, E. A. & Raff, J. W. Centrioles, centrosomes, and cilia in health and disease. Cell 139, 663–678 (2009).
Azimzadeh, J. & Marshall, W. F. Building the centriole. Curr. Biol. 20, 816–825 (2010).
Holland, A. J., Lan, W. & Cleveland, D. W. Centriole duplication: a lesson in self-control. Cell Cycle 9, 2731–2736 (2010).
Nigg, E. A. & Stearns, T. The centrosome cycle: centriole biogenesis, duplication and inherent asymmetries. Nat. Cell Biol. 13, 1154–1160 (2011).
Yang, Y. et al. NudC-like protein 2 regulates the LIS1/dynein pathway by stabilizing LIS1 with Hsp90. Proc. Natl. Acad. Sci. USA 107, 3499–3504 (2010).
Osmani, A. H., Osmani, S. A. & Morris, N. R. The molecular cloning and identification of a gene product specifically required for nuclear movement in Aspergillus nidulans. J. Cell Biol. 111, 543–551 (1990).
Xiang, X., Osmani, A. H., Osmani, S. A., Xin, M. & Morris, N. R. NudF, a nuclear migration gene in Aspergillus nidulans, is similar to the human LIS-1 gene required for neuronal migration. Mol. Biol. Cell 6, 297–310 (1995).
Fu, Q., Wang, W., Zhou, T. & Yang, Y. Emerging roles of NudC family: from molecular regulation to clinical implications. Sci. China Life Sci. 59, 455–462 (2016).
Yang, Y. et al. NudCL2 is an Hsp90 cochaperone to regulate sister chromatid cohesion by stabilizing cohesin subunits. Cell Mol. Life Sci. 76, 381–395 (2019).
Anderhub, S. J., Krämer, A. & Maier, B. Centrosome amplification in tumorigenesis. Cancer Lett. 322, 8–17 (2012).
Kwon, M. et al. Mechanisms to suppress multipolar divisions in cancer cells with extra centrosomes. Genes Dev. 22, 2189–2203 (2008).
Leber, B. et al. Proteins required for centrosome clustering in cancer cells. Sci. Transl. Med. 2, 33ra38 (2010).
Ganem, N. J., Godinho, S. A. & Pellman, D. A mechanism linking extra centrosomes to chromosomal instability. Nature 460, 278–282 (2009).
Cosenza, M. R. et al. Asymmetric centriole numbers at spindle poles cause chromosome missegregation in cancer. Cell Rep. 20, 1906–1920 (2017).
Warnke, S. et al. Polo-like kinase-2 is required for centriole duplication in mammalian cells. Curr. Biol. 14, 1200–1207 (2004).
Zhu, F. et al. The mammalian SPD-2 ortholog Cep192 regulates centrosome biogenesis. Curr. Biol. 18, 136–141 (2008).
Alves-Cruzeiro, J. M., Nogales-Cadenas, R. & Pascual-Montano, A. D. CentrosomeDB: a new generation of the centrosomal proteins database for human and Drosophila melanogaster. Nucleic Acids Res. 42, 430–436 (2014).
Sánchez-Tena, S., Cubillos-Rojas, M., Schneider, T. & Rosa, J. L. Functional and pathological relevance of HERC family proteins: a decade later. Cell Mol. Life Sci. 73, 1955–1968 (2016).
Chan, N. C. et al. Degradation of the deubiquitinating enzyme USP33 is mediated by p97 and the ubiquitin ligase HERC2. J. Biol. Chem. 289, 19789–19798 (2014).
Li, J. et al. USP33 regulates centrosome biogenesis via deubiquitination of the centriolar protein CP110. Nature 495, 255–259 (2013).
Fan, G. et al. Loss of KLF14 triggers centrosome amplification and tumorigenesis. Nat. Commun. 6, 8450 (2015).
Shiratsuchi, G., Takaoka, K., Ashikawa, T., Hamada, H. & Kitagawa, D. RBM14 prevents assembly of centriolar protein complexes and maintains mitotic spindle integrity. EMBO J. 34, 97–114 (2015).
Xu, X. et al. DNA replication licensing factor Cdc6 and Plk4 kinase antagonistically regulate centrosome duplication via Sas-6. Nat. Commun. 8, 15164 (2017).
Guderian, G., Westendorf, J., Uldschmid, A. & Nigg, E. A. Plk4 trans-autophosphorylation regulates centriole number by controlling β-TrCP-mediated degradation. J. Cell Sci. 123, 2163–2169 (2010).
Puklowski, A. et al. The SCF–FBXW5 E3-ubiquitin ligase is regulated by PLK4 and targets HsSAS-6 to control centrosome duplication. Nat. Cell Biol. 13, 1004–1009 (2011).
D’Angiolella, V. et al. SCF (CyclinF) controls centrosome homeostasis and mitotic fidelity through CP110 degradation. Nature 466, 138–142 (2010).
Sonnen, K. F., Gabryjonczyk, A. M., Anselm, E., Stierhof, Y. D. & Nigg, E. A. Human Cep192 and Cep152 cooperate in Plk4 recruitment and centriole duplication. J. Cell Sci. 126, 3223–3233 (2013).
Kim, T. S. et al. Hierarchical recruitment of Plk4 and regulation of centriole biogenesis by two centrosomal scaffolds, Cep192 and Cep152. Proc. Natl. Acad. Sci. USA 110, 4849–4857 (2013).
Moyer, T. C., Clutario, K. M., Lambrus, B. G., Daggubati, V. & Holland, A. J. Binding of STIL to Plk4 activates kinase activity to promote centriole assembly. J. Cell Biol. 209, 863–878 (2015).
Dzhindzhev, N. S. et al. Two-step phosphorylation of Ana2 by Plk4 is required for the sequential loading of Ana2 and Sas6 to initiate procentriole formation. Open Biol. 7, 170247 (2017).
Ohta, M. et al. Direct interaction of Plk4 with STIL ensures formation of a single procentriole per parental centriole. Nat. Commun. 5, 5267 (2014).
Tsou, M. F. et al. Polo kinase and separase regulate the mitotic licensing of centriole duplication in human cells. Dev. Cell 17, 344–354 (2009).
Kleylein-Sohn, J. et al. Plk4-induced centriole biogenesis in human cells. Dev. Cell 13, 190–202 (2007).
Tang, C. J. et al. The human microcephaly protein STIL interacts with CPAP and is required for procentriole formation. EMBO J. 30, 4790–4804 (2011).
Peel, N., Stevens, N. R., Basto, R. & Raff, J. W. Overexpressing centriole-replication proteins in vivo induces centriole overduplication and de novo formation. Curr. Biol. 17, 834–843 (2007).
Izumi, H. et al. BubR1 localizes to centrosomes and suppresses centrosome amplification via regulating Plk1 activity in interphase cells. Oncogene 28, 2806–2820 (2009).
Haaß, W. et al. Measurement of separase proteolytic activity in single living cells by a fluorogenic flow cytometry assay. PLoS ONE 10, e0133769 (2015).
Bekker-Jensen, S. et al. HERC2 coordinates ubiquitin-dependent assembly of DNA repair factors on damaged chromosomes. Nat. Cell Biol. 12, 80–e0133786 (2010).
Izawa, N. et al. HERC2 interacts with Claspin and regulates DNA origin firing and replication fork progression. Cancer Res. 71, 5621–5625 (2011).
Wu, W. et al. HERC2 is an E3 ligase that targets BRCA1 for degradation. Cancer Res. 70, 6384–6392 (2010).
Al-Hakim, A. K., Bashkurov, M., Gingras, A. C., Durocher, D. & Pelletier, L. Interaction proteomics identify NEURL4 and the HECT E3 ligase HERC2 as novel modulators of centrosome architecture. Mol. Cell Proteom. 11, M111.014233 (2012).
Aumais, J. P. et al. NudC associates with Lis1 and the dynein motor at the leading pole of neurons. J. Neurosci. 21, RC187 (2001).
Aumais, J. P. et al. Role for NudC, a dynein-associated nuclear movement protein, in mitosis and cytokinesis. J. Cell Sci. 116, 1991–2003 (2003).
Cai, Y., Yang, Y., Shen, M. & Zhou, T. Inhibition of cytokinesis by overexpression of NudCL that is localized to the centrosome and midbody. Cell Res. 19, 1305–1308 (2009).
Zhou, T., Aumais, J. P., Liu, X., Yu-Lee, L. Y. & Erikson, R. L. A role for Plk1 phosphorylation of NudC in cytokinesis. Dev. Cell 5, 127–138 (2003).
Zhang, C. et al. NudC regulates actin dynamics and ciliogenesis by stabilizing cofilin 1. Cell Res. 26, 239–253 (2015).
Zhou, T., Zimmerman, W., Liu, X. & Erikson, R. L. A mammalian NudC-like protein essential for dynein stability and cell viability. Proc. Natl. Acad. Sci. USA 103, 9039–9044 (2006).
Asante, D., Stevenson, N. L. & Stephens, D. J. Subunit composition of the human cytoplasmic dynein-2 complex. J. Cell Sci. 127, 4774–4787 (2014).
Lu, Y. et al. Twa1/Gid8 is a β-catenin nuclear retention factor in Wnt signaling and colorectal tumorigenesis. Cell Res. 27, 1422–1440 (2017).
Shen, M. et al. Centrosomal protein FOR20 is essential for S-phase progression by recruiting Plk1 to centrosomes. Cell Res. 23, 1284–1295 (2013).
Acknowledgements
We are grateful to Wei Zhuo and Dante Neculai for providing the good suggestions about manuscript writing and to Guifeng Xiao and Wei Yin for help with the confocal imaging. We thank all the members of Zhou’s lab for the helpful comments and suggestions during the work. This work was supported by the National Natural Scientific Foundation of China (31671394, 31471259, 91740205, 31620103911, and 31571446) and the National Key Research and Development Program of China (2016YFA0100301).
Author information
Authors and Affiliations
Contributions
T.Z., M.L., and Y.Y. designed the experiments. M.L., X.X., W.W., and Y.G. performed the experiments. Q.S. and Y.L. provided assistance for quantitative proteomic analysis. Q.S., Y.L., M.L., and J.Z. discussed the results and commented on the manuscript. M.L. and Y.Y. wrote the original draft. Y.Y., F.W., W.L., and T.Z. edited the manuscripts. T.Z. supervised the project.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by G. Liccardi
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, M., Xu, X., Zhang, J. et al. NudC-like protein 2 restrains centriole amplification by stabilizing HERC2. Cell Death Dis 10, 628 (2019). https://doi.org/10.1038/s41419-019-1843-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-019-1843-3
- Springer Nature Limited