Abstract
Overcoming drug-resistance is a big challenge to improve the survival of patients with epithelial ovarian cancer (EOC). In this study, we investigated the effect of chloroquine (CQ) and its combination with cisplatin (CDDP) in drug-resistant EOC cells. We used the three EOC cell lines CDDP-resistant A2780-CP20, RMG-1 cells, and CDDP-sensitive A2780 cells. The CQ-CDDP combination significantly decreased cell proliferation and increased apoptosis in all cell lines. The combination induced expression of γH2AX, a DNA damage marker protein, and induced G2/M cell cycle arrest. Although the CQ-CDDP combination decreased protein expression of ATM and ATR, phosphorylation of ATM was increased and expression of p21WAF1/CIP1 was also increased in CQ-CDDP-treated cells. Knockdown of p21WAF1/CIP1 by shRNA reduced the expression of γH2AX and phosphorylated ATM and inhibited caspase-3 activity but induced ATM protein expression. Knockdown of p21WAF1/CIP1 partly inhibited CQ-CDDP-induced G2/M arrest, demonstrating that knockdown of p21WAF1/CIP1 overcame the cytotoxic effect of the CQ-CDDP combination. Ectopic expression of p21WAF1/CIP1 in CDDP-treated ATG5-shRNA/A2780-CP20 cells increased expression of γH2AX and caspase-3 activity, demonstrating increased DNA damage and cell death. The inhibition of autophagy by ATG5-shRNA demonstrated similar results upon CDDP treatment, except p21WAF1/CIP1 expression. In an in vivo efficacy study, the CQ-CDDP combination significantly decreased tumor weight and increased expression of γH2AX and p21WAF1/CIP1 in A2780-CP20 orthotopic xenografts and a drug-resistant patient-derived xenograft model of EOC compared with controls. These results demonstrated that CQ increases cytotoxicity in combination with CDDP by inducing lethal DNA damage by induction of p21WAF1/CIP1 expression and autophagy inhibition in CDDP-resistant EOC.
Similar content being viewed by others
Facts
-
Overcoming drug-resistance is a major challenge in improving the survival of patients with epithelial ovarian cancer (EOC).
-
Chloroquine is an anti-malaria drug and inhibits autophagy.
-
Chloroquine in combination with cisplatin increases cytotoxicity of drug-resistant EOC through induction of lethal DNA damage.
-
Induction of p21WAF1/CIP1 expression and inhibition of autophagy mediated by chloroquine lead to lethal DNA damage.
-
Combination of chloroquine with cisplatin represents a novel strategy to treat drug-resistant EOC.
Introduction
Platinum-based drugs are generally accepted main drugs for treatment of epithelial ovarian cancer (EOC). However, some patients with EOC have intrinsic resistance or develop acquired resistance due to repeated administration1,2. Overcoming drug resistance is thus a big challenge to improve the survival of EOC patients3,4. Chemotherapeutic drugs for EOC such as platinum-based cisplatin (CDDP) are known to cause DNA damage, and leads to cell death5,6. However, chemo-resistant cancer cells can survive from anti-cancer drug-induced DNA damage via DNA repair pathways or bypassing cell cycle checkpoints7,8. Therefore, DNA damage/repair-related signaling pathways are important targets in treatment of chemo-resistant cancer.
Protein kinases including ataxia telangiectasia mutated (ATM) and RAD3-related (ATR) recognize DNA damage and induce DNA damage response signals9,10. Upon DNA damage, ATM/ATR activate the checkpoint kinases Chk2 and Chk1 by phosphorylation and also activate p53, inducing p21WAF1/CIP1 (also called Cyclin-dependent kinase inhibitor 1) and inhibiting Cyclin-dependent kinases to arrest the cell cycle and repair DNA damage. ATM/ATR phosphorylate histone variant H2AX at serine 139 which is called γH2AX in the response of DNA damage and therefore γH2AX is considered as a DNA damage marker protein11. Cyclin-dependent kinase 1, Cdk1 (Cdc2), is involved in cell cycle progression through an interaction with cyclin B1. Phosphorylation of Cdc2 at tyrosine 15 (Cdc2-pY15), an inactive form, arrests the cell cycle at G2/M12. During cell cycle arrest, DNA can be repaired, allowing cell survival, or persistent DNA damage induces cell death. In DNA-damaging anti-cancer agents, repair and cell cycle checkpoints are considered as mechanisms for acquiring resistance13.
Autophagy can be a survival signal under hypoxic and limited nutrient conditions, which are hallmarks of cancers14,15. Drug-resistant cancers can survive through induction of autophagy by anti-cancer drugs; therefore, autophagy was considered to induce resistance in chemotherapy. Therefore, autophagy inhibitors have become candidates to treat drug-resistant cancers.
Drug-resistant cancers can survive through induction of autophagy14,15; therefore, autophagy was considered to induce resistance in chemotherapy. Therefore, autophagy inhibitors have become candidates to treat drug-resistant cancers.
Chloroquine (CQ) is an anti-malaria drug and inhibits autophagy through inhibition of autophagosome-lysosome fusion16. The anti-tumor effect of CQ has been reported in different types of cancer17,18. CQ induced cell death through activation of p53 pathway in glioblastoma19. Cancer cells can survive from oxidative stress by removing reactive oxygen species (ROS) by autophagy. CQ induced cancer cell death by inhibiting autophagy-mediated removal of ROS20. CQ was also studied as an anti-cancer drug in combination with other anti-cancer drugs. CDDP induced cell proliferation through activation of the Akt/mTOR signaling pathway by activation of autophagy, and CQ induced cell death by inhibition of CDDP-induced autophagy in ovarian cancer cell lines21. CQ combined with an mTOR inhibitor, enhanced radiosensitivity of colon cancer22. Although there are several results on the cytotoxic effect of combined CQ and anti-cancer drugs, the exact functional mechanism or targeting signals have not been identified.
p21WAF1/CIP1 is induced by p53 upon DNA damage and has critical roles in cell cycle distribution, apoptosis, cell growth, and senescence23. The function of p21WAF1/CIP1 in cancer cell growth has been reported that ectopic expression of p21WAF1/CIP1 induced apoptosis in several cancers24,25,26. Furthermore, anti-cancer drugs induced cell death by induction of p21WAF1/CIP1 expression27,28. In contrast, the anti-apoptotic effect of p21WAF1/CIP1 in response of anti-cancer drugs has been reported in prostate and colon cancers29,30. Conflicting function of p21WAF1/CIP1 on cell growth and apoptosis is dependent on subcellular localization, post-translational modifications, and p53 status31. p21WAF1/CIP1 inhibited Cdc2 and cyclin B1 expression, thereby induced G2/M arrest and also induced S phase arrest by inhibition of Cdk231. p21WAF1/CIP1 interacts with many proteins such as PCNA, Cdk2, and Cdk4, so the function of p21WAF1/CIP1 might be dependent on the cell context27,32.
In this study, we evaluated the effect of CQ on DNA damage/cell cycle in regard to cytotoxicity in drug-resistant EOC. We found that the CQ-CDDP combination increased cytotoxicity in drug-resistant EOC cell lines and induced γH2AX, demonstrating that the induced cytotoxicity might be caused by severe DNA damage. The CQ-CDDP combination sustained G2/M arrest and induced expression of p21WAF1/CIP1. The induced cytotoxicity of EOC by CQ was dependent on its autophagy inhibitor activity and induction of p21WAF1/CIP1. In an in vivo study, the CQ-CDDP combination significantly decreased tumor weights in drug-resistant cell line-derived xenograft and patient-derived xenograft (PDX) models. Taken together, these results demonstrated that CQ could increase cytotoxicity in combination with CDDP by inducing lethal DNA damage through induction of p21WAF1/CIP1 expression and autophagy inhibition in CDDP-resistant EOC.
Materials and methods
Cell lines
Ovarian carcinoma cell lines A2780 (cisplatin-sensitive) and A2780-CP20 (cisplatin-resistant) were obtained from American Type Culture Collection (ATCC) and Dr. Anil K. Sood (M&D Anderson Cancer Center, Houston, TX, USA), respectively. Cells were maintained in RPMI (Gibco, CA, USA) containing 10% FBS and 100 units/ml penicillin/100 µg/ml streptomycin (Invitrogen, CA, USA) and were grown at 37 °C in a 5% CO2 incubator. RMG-1, an ovarian clear cell carcinoma cell line, was purchased from Japan Health Science Research Resources Bank (HSRRB, Osaka, JAPAN) and maintained in HAM’s F-12 media (Gibco) containing 10% FBS and penicillin/streptomycin. We obtained the authentications of the cell lines used in this study and checked the certificates of the cell lines using species verification and short tandem repeat DNA profiling assay. CDDP was purchased from Sigma (MO, USA) and suspended in distilled water. CQ purchased from Sigma was suspended in distilled water at 50 mg/ml concentration and used at 10–30 µM in all cell lines. Inhibitors for ATM (KU55933, Abcam, England) and ATR (AZD6738, AstraZeneca, England) were used at 15 µM and 1 µM, respectively. MG-132 (Z-L-Leu-D-Leu-L-Leu-al) and bafilomycin A1 were purchased from Sigma and used at 2 µM and 0.1 µM, respectively.
Preparation of lentivirus-expressing shRNA and stable cell lines
To prepare lentiviruses expressing shRNA, plasmids for lentivirus preparation, pMD.2G (Addgene, MA, USA) and psPAX2 (Addgene), and plasmid encoding either ATG5-, p21WAF1/CIP1-specific shRNA (Origene, MD, USA), or GFP-expressing scrambled shRNA (Origene) were transfected into 293T cells using Lipofectamine 2000 (Invitrogen). Lentivirus expressing shRNA was prepared as previously described33. Virus-containing media were stored in aliquots at −80 oC.
For preparing stable cells, A2780-CP20 cells were plated into 60 mm dishes one day before lentivirus infection. Cells were infected with lentivirus expressing ATG5-specific shRNA, p21WAF1/CIP1-specific shRNA or scrambled control shRNA in the presence of polybrene (8 µg/ml, Sigma) for 6 h and media was changed to complete growth media (RPMI media with 10% FBS and penicillin/streptomycin). At 48 h after virus infection, media were changed to complete growth media containing puromycin (Sigma). Cultures were monitored for surviving cells and surviving cells were either used for experiments or stored in RPMI media containing 20% FBS and 10% DMSO (Sigma) at −80 oC.
Western blot analysis
EOC cells were plated at 2 × 105 cells/well in a 6-well plate. Cells were treated with CDDP, CQ, or CQ-CDDP combination at the indicated concentrations for 24 h or 48 h. Cells were lysed in RIPA buffer containing 0.1 M sodium fluoride (NaF, Sigma), 0.1 M β-glycerophosphate (Sigma), protease inhibitor cocktail (Sigma), and 1 mmol/L phenylmethylsulfonyl fluoride (PMSF, Sigma). Western blot analysis was performed as previously described33. Primary antibodies against ATM (sc-23921), Cdc2 (sc-8395), Keap1 (sc-365626), and β-actin (sc-47778) were obtained from Santa Cruz Biotechnology (TX. USA); antibodies against p-ATR (#2853), ATR (#13934), p-Cdc2 (Tyr-15,#4539), p21WAF1/CIP1 (#2947), p-Akt (#9271), and Akt (#9272) were from Cell Signal Technology (MA, USA); antibodies against Cyclin B1 (ab32053), p-ATM (S1981, ab81292), and p62/SQSTM1 (ab56416) were from Abcam; and antibodies against γH2AX (NB100-384), LC3B (NB100-2220), and ATG5 (NB110-53818) were purchased from Novus (CO, USA). Antibody for Nrf2 was purchased from Thermo Fisher (PA5-27882).
Immunofluorescence staining
For immunofluorescence staining for γH2AX, cells were plated onto gelatin-coated glass coverslips in 6-well plates. Twenty-four or forty-eight hours after drug incubation, cells were fixed in 4% paraformaldehyde for 10 min and blocked with 3% BSA in 1X PBS. Cells were then incubated with anti-γH2AX antibody (NB100-384, Novus) for 1.5 h and examined after incubation with Alexa 568-conjugated anti-rabbit antibody for 1 h. Nuclear counterstaining was performed using 4′6-diamidino-2-phenylindole (DAPI, Sigma). Stained cells were examined under a confocal microscope (X400, LSM700, Carl Zeiss, Germany).
Active caspase-3 assay
To assess apoptotic cell death, cells were treated with CQ and CDDP for 48 h and caspase-3 activity in the cell lysates was measured using a caspase-3 (cleaved) human ELISA kit (Invitrogen). ELISA was performed according to the manufacturer’s protocol.
Cell cycle analysis by flow cytometry
Cells were plated at 3 × 105 cells into 60 mm-dishes and treated with drugs for 24 and 48 h. Cells were trypsinized, washed with cold PBS, and fixed in 70% ethanol overnight at −20 oC. Cells were washed with cold PBS and stained with propidium iodide (PI, Sigma) solution containing 50 µg/ml PI, 0.1 mg/ml RNase A (Sigma), and 0.05% Triton X-100 in PBS for 30 min at 37 oC. Cells were washed once with cold PBS and cell cycle analysis was performed using FACSVerseTM (BD FACS: Fluorescence-activated cell sorting, CA, USA).
Transfection of plasmid expressing HA-p21WAF1/CIP1
Plasmid expressing HA-p21WAF1/CIP1 was purchased from Addgene. Plasmid encoding HA-p21WAF1/CIP1 or empty vector was transfected into cells (2 × 105/well in 6-well plate) using Lipofectamine 2000 (Invitrogen). At 7 h after transfection, media was changed and drugs were treated to cells for an additional 36 h. After lysing cells in RIPA buffer, western blotting was performed.
Hoechst/propidium iodide staining
A2780-CP20 cells (1 × 106/dish) were plated in a 60 mm-dish and incubated with CDDP, CQ, or CQ combined with CDDP for 48 h. After incubation, culture media was collected in 15-ml falcon tubes and cells were washed with 1X cold PBS. Cells were trypsinized, were combined with culture media collected before trypsin treatment, and were washed twice with 1X cold PBS. Cells were incubated with Hoechst33342 and propidium iodide (Vybrant Apoptosis assay kit#5, ThermoFisher) for 10 min. Apoptosis and necrosis were measured with FACS Aria (BD Bioscience).
Development of in vivo cell line xenograft and PDX models
In vivo animal study was reviewed and approved by the Institutional Animal Care and Use Committee at Samsung Biomedical Research Institute (protocol no. H-A9-003), which is permissioned by the Association for the Assessment and Accreditation of Laboratory Animal Care International and compliant with the guidelines of the Institute of Laboratory Animal Resources. Female BALB/c nude mice were obtained from Orient Bio (Seongnam, South Korea). Orthotopic models were prepared with A2780-CP20 cells (1 × 106 cells/0.2 ml Hank’s balanced salt solution) by injection into the peritoneal cavities of BALB/c nude mice34. A PDX model of ovarian cancer was established using 6 to 8 weeks old mice as described in our previous report35. This study with PDX mouse model was approved by the Samsung Medical Center Institutional Review Board (file#201004004) and performed in accordance with approved guidelines and regulations. Orthotopic cell-line xenograft and PDX models were randomly divided into 4 groups (n = 10 per group) before administrating drugs. CDDP (4 mg/kg, once a week), CQ (50 mg/kg, three times a week), CQ and CDDP, or PBS were injected into model mice for the subsequent 3 weeks. Tumor development was monitored daily and mice were sacrificed at the end of treatment. Body weight and tumor weight were measured.
Immunohistochemistry
For immunohistochemical staining of γH2AX as a marker for DNA damage, p21WAF1/CIP1, and Ki-67 as a marker for cell proliferation, tissue sections (4 µm-thick) were incubated with anti-γH2AX rabbit polyclonal antibody (NB100-384, Novus), anti-p21WAF1/CIP1 antibody (#2947, Cell signaling), or anti-Ki67 rabbit polyclonal antibody (NB600-1252, Novus) overnight at 4 °C in a humidity chamber after heat-induced epitope retrieval (HIER) with citrate buffer (pH 6.0; Dako, CA, USA). The sections were then incubated with HRP-conjugated secondary antibody against rabbit IgG (P-0448, Dako) for 30 min at RT. The color reaction was developed using the ready-to-use DAB (3,3’-diaminobenzidine) substrate-chromogen solution (Dako) for 5 min and sections were counterstained with Mayer’s hematoxylin (Sigma) for 30 s before dehydration and mounting. Slides were scanned with Scanscope microscopy (Aperio, Germany).
Statistical analysis
Data were analyzed with Mann–Whitney U test and one-way analysis of variance (ANOVA) followed by the Newman–Keuls multiple comparison tests, as appropriate, using a statistical software package (Prism, GraphPad, CA, USA). P values less than 0.05 were considered statistically significant.
Results
CQ increases CDDP-induced cell death in EOC cells
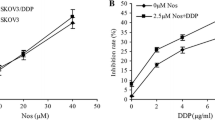
Lethal concentration 50 (LC50) of CQ or CDDP in the EOC cell lines A2780, A2780-CP20, and RMG-1 was investigated after 72 h treatment with each drug (Supplementary Fig. S1). In A2780 cells, 10 μM of CDDP induced cell death in more than 80% of cells, and the combination of CDDP with CQ further increased CDDP-induced cell death (Fig. 1a). However, in A2780-CP20 and RMG-1 cells, 10 μM of CDDP induced cell death in 20 and 46% of cells, respectively, and the CQ-CDDP combination induced cell death by 50 and 70% compared with controls (Fig. 1a and Supplementary Fig. S2a).
a CDDP-sensitive (A2780) and CDDP-resistant (A2780-CP20) EOC cells were treated with CDDP and CQ for 72 h, and cell viability was measured by MTT assay. Results are demonstrated by a bar graph. b Apoptotic cell death was measured by ELISA for detecting active caspase-3. A2780 and A2780-CP20 cells were treated with CDDP (1 µM and 5 µM, respectively) and CQ (20 µM and 30 µM, respectively) as indicated for 48 h, and cell lysates were used for caspase-3 assay. Results are shown as the mean ± SD of triplicate observations from three experiments (n = 3, *P < 0.05, **P < 0.01, ***P < 0.001). c A2780-CP20 cells were stained with Hoechst/propidium iodide to detect the contribution to apoptosis and necrotic cell death after treatment with CDDP and CQ for 48 h. The representative FACS data obtained from three experiments and a bar graph representing apoptotic cells are shown as the mean ± SD (n = 3, ***P < 0.001).
In apoptosis assays measuring active caspase-3, different amounts of CDDP were used for drug-sensitive A2780 and drug-resistant A2780-CP20 and RMG-1 cells (1 μM and 5 μM, respectively). CQ-CDDP combination significantly increased apoptosis in all cells (Fig. 1b and Supplementary Fig. S2b). Interestingly, in A2780-CP20 and RMG-1 cells, this combination more profoundly increased apoptosis compared with CQ or CDDP alone. To characterize the contribution of apoptosis/necrosis in this experimental condition, the apoptotic and necrotic cells were analyzed by Hoechst/propidium iodide staining (Fig. 1c). Although the proportion of necrosis was 0.1, 0.5, 0.2, and 0.7% of total cells in the control, CDDP, CQ, and CQ-CDDP treatments, respectively, apoptosis was increased in the CQ-CDDP combination compared with the control, consistent with the result of caspase-3 assay presented in Fig. 1b.
CQ increases CDDP-induced DNA damage
CDDP induces DNA damage, leading to DNA damage-mediated cancer cell death5. The effect of the CQ-CDDP combination on DNA damage was studied by measuring amount of γH2AX. Treatment of CDDP and the combination with CQ induced γH2AX in A2780 cells in a time-dependent manner. However, in A2780-CP20 and RMG-1 cells, the combination with CQ increased γH2AX compared with CDDP alone at 48 h (Fig. 2a and Supplementary Fig. S2c). Induction of γH2AX by the CQ-CDDP combination was also determined by immunofluorescence staining in A2780-CP20 cells at 24 h and 48 h and demonstrated that the combination further increased γH2AX foci in nuclei, representing increased DNA damage (Fig. 2b, c).
a EOC cells were treated with either CDDP, CQ, or the combination of CQ and CDDP for 24 h and 48 h. Phosphorylation of histone H2AX (γH2AX) was detected by western blot using anti-γH2AX antibody. b Nuclear foci formation of γH2AX was determined by immunofluorescent staining in A2780-CP20 cells treated with each drug as indicated (X400, scale bar represents 20 μm). DAPI was used for nuclear staining. c Nuclear foci formation of γH2AX obtained from b was quantified by a bar graph after calculating the number of nuclear foci of γH2AX and nuclei. The bar graph was generated with data obtained from three experiments (n = 3, *P < 0.05, **P < 0.01, ***P < 0.001).
The CQ-CDDP combination arrests the cell cycle at G2/M and regulates DNA damage-related signaling proteins
The cell cycle is arrested upon DNA damage to allow the cell to repair the damage. Therefore, we assessed the cell cycle of EOC cells by FACS analysis after drug treatment. CDDP treatment for 24 h arrested the cell cycle of A2780 cells at G2/M, and this cell cycle arrest persisted until 48 h (Fig. 3a). The combination of CQ-CDDP in A2780 cells induced G2/M arrest with increasing incubation time. Interestingly, CQ-CDDP combination also induced S phase arrest at 24 h and slightly decreased it at 48 h. In A2780-CP20 cells, although 24 h treatment of CDDP also arrested the cell cycle at G2/M, this arrest was reversed during 48 h treatment (Fig. 3b). In contrast, the CQ-CDDP combination induced G2/M arrest in A2780-CP20 cells at 24 h, and the prolonged incubation further increased the number of G2/M-arrested cells (from 37% of G2/M-arrested cells at 24 h to 52% at 48 h). RMG-1 cells showed similar responses to A2780-CP20 cells (Supplementary Fig. S3a).
a A2780 and b A2780-CP20 cells were incubated with each drug or the combination of drugs for 24 h and 48 h. Cells were stained with propidium iodide, and cell cycle distribution was measured by flow cytometry. Flow cytometry was performed three times, and representative data are presented. Cell cycle distribution was demonstrated by a bar graph and the statistical data represented for G2/M-arrested cell numbers compared with the control-treated cells. c Activation and expression of cell cycle-related proteins (Cdc2 and cyclin B1) were studied by western blot analysis in A2780 and A2780-CP20 cells using anti-phospho-specific antibody and antibodies recognizing the total proteins. The amount of phosphorylation of Cdc2 normalized by the amount of total Cdc2 expression in A2780 and A2780-CP20 cells is represented by a bar graph in the upper panel and lower panel, respectively. d DNA damage-recognizing proteins, ATM and ATR, were analyzed in A2780 and A2780-CP20 cells by western blot using phospho-specific antibodies and antibodies recognizing both the phosphorylated and unphosphorylated proteins. Arrow indicates ATR. Western blotting was performed at least three times, and a representative figure is presented. The amount of phosphorylation of ATM normalized by total ATM expression in A2780 and A2780-CP20 cells is represented by a bar graph in the upper panel and lower panel, respectively (n = 3, mean ± SD, *P < 0.05, **P < 0.01, ***P < 0.001).
To determine the mechanism involved in the G2/M cell cycle arrest induced by the CQ-CDDP combination, we examined the phosphorylation of Cdc2 at tyrosine 15 (Cdc2-pY15, the inactive form), which is known to lead to G2/M arrest. CDDP induced Cdc2 expression in both A2780 and A2780-CP20 cells. Amount of Cdc2-pY15 was increased during 48 h treatment of the CQ-CDDP combination in all cells (Fig. 3c and Supplementary Fig. S3b). The ratio of p-Cdc2/Cdc2 was increased in CQ-CDDP-treated A2780-CP20 cells with 48 h incubation. However, in A2780 cells, treatments of CDDP and CQ-CDDP combination induced Cdc2 protein expression but did not significantly increase its phosphorylation. The error bars in A2780 cells show lack of statistical significance (Fig. 3c upper panel). Expression of Cyclin B1 was increased by CDDP and had no effect by CQ-CDDP combination in either A2780 or A2780-CP20 cells. We also evaluated upstream signaling of Cdc2. ATM, a DNA damage-recognition protein, was activated by CDDP in EOC cells. Although the CQ-CDDP combination inhibited ATM protein expression, the combination induced phosphorylation of ATM in all EOC cells compared with CDDP alone (Fig. 3d and Supplementary Fig. S3c). Although CDDP slightly induced phosphorylation of ATR, the CQ-CDDP combination had no effect on phosphorylation of ATR at 48 h in all EOC cells (Fig. 3d and Supplementary Fig. 3c). However, ATR protein expression was highly decreased by the CQ-CDDP combination. Taken together, these results show that the CQ-CDDP combination might induce G2/M cell cycle arrest through activation of ATM and inhibition of Cdc2.
The CQ-CDDP combination has similar effects to ATM and ATR inhibitors
We compared the effects of ATM or ATR inhibitors with the CQ-CDDP combination on cell cycle and expression of signaling proteins. Inhibitors specific for ATM (ATMi) or ATR (ATRi) were incubated either alone or in combination with CDDP in A2780-CP20 cells. ATMi alone seemed to induce G2/M arrest by about 1.5-fold compared with the control at 48 h incubation but this induction was not statistically significant as calculated from three experiments (Supplementary Fig. S4). However, in combination with CDDP, these inhibitors significantly induced G2/M arrest (Fig. 4a). The pattern of G2/M arrest by the CQ-CDDP combination in A2780-CP20 cells was similar to that of the combination of ATMi or ATRi with CDDP, in that G2/M arrest lasted until 48 h. ATMi inhibited CDDP-induced ATM phosphorylation at 48 h, demonstrating that the inhibitor successfully inhibited its target protein (Fig. 4b). Combination of ATRi with CDDP highly increased γH2AX expression compared with CDDP or inhibitor alone. Combination of ATMi with CDDP increased Cdc2-pY15 at 48 h incubation, which was the same effect as with the CQ-CDDP combination (Fig. 4c).
a Cell cycle distribution was analyzed by flow cytometry in A2780-CP20 cells treated with CDDP alone or in combination with ATM or ATR inhibitor for 24 h and 48 h. Flow cytometry was performed three times, and representative data are presented. Cell cycle distribution was demonstrated by a bar graph and the statistical data represented for G2/M-arrested cell numbers compared with the control cells. b A2780-CP20 cells were incubated with CDDP, CQ, ATR inhibitor, ATM inhibitor, or the combination of CDDP with either CQ, ATRi, or ATMi for 24 h and 48 h. Western blot for ATM and ATR was performed using antibodies for phospho-specific and antibodies recognizing both the phosphorylated and unphosphorylated proteins as indicated. The amount of phosphorylation of ATM normalized by the amount of total ATM expression is represented by a bar graph in the right panel. Arrow indicates ATR. c Expression of cell cycle-related proteins was determined in A2780-CP20 cells after 24 h and 48 h treatment with the CQ-CDDP combination. Western blot analysis was performed three times, and the representative data are presented (n = 3, mean ± SD, *P < 0.05, **P < 0.01, ***P < 0.001).
CQ increases CDDP-induced cell death via inhibition of autophagy and induced p21WAF1/CIP1 protein expression
We next evaluated whether the inhibition of autophagy by CQ is involved in CDDP-induced cell death. To inhibit autophagy, lentivirus expressing ATG5-specific shRNA was generated and used to infect A2780-CP20 cells (ATG5-shRNA cells). Knockdown of ATG5 was analyzed by western blot, and expression of p62 was increased in ATG5-shRNA cells and CQ-treated A2780-CP20 cells, indicating autophagy inhibition (Fig. 5a). Conversion from LC3-I to LC3-II was inhibited in ATG5-shRNA cells compared with the control cells determined by increased amount of LC3-I. In contrast, since CQ inhibits fusion between autophagosome and lysosome16, LC3-II accumulated in cells treated with CQ. Cell viability of ATG5-shRNA or control-shRNA cells were measured by MTT assay after incubation with different amounts of CDDP. In contrast with control cells, CDDP at 4 µM resulted in 50% cell death in ATG5-shRNA cells, demonstrating that inhibition of autophagy sensitized A2780-CP20 cells to CDDP (Fig. 5b). In caspase-3 assay, CDDP in ATG5-shRNA cells induced caspase-3 activity compared with CDDP-treated control-shRNA cells, and this cell death increase was further induced by CQ-CDDP treatment in ATG5-deficient cells (Fig. 5c). Interestingly, CQ alone had no significant effect on caspase-3 activity.
a A2780-CP20 cells stably expressing control or ATG5 shRNA were treated with CDDP for 48 h, and the expression of ATG5 and autophagy marker proteins, LC3 and p62, were analyzed by western blot using specific antibodies. Knockdown efficiency of ATG5 was shown by western blot. Expression of LC3 and p62 was also analyzed by western blot in A2780-CP20 cells treated with CDDP, CQ, or the CQ-CDDP combination for 48 h. β-actin was used for a protein-loading control. shRNA/CP20 represents control-shRNA- or ATG5-shRNA-expressing A2780-CP20 cells. Western blot was performed three times, and representative data are presented (left panel). The amounts of p62 and LC3-I and -II normalized to the amount of β-actin are represented by a bar graph (middle and right panels, respectively). b Cell viability was measured by MTT assay in A2780-CP20 cells stably expressing control- (closed squares) or ATG5-shRNA (open triangles) in the presence of different concentrations of CDDP at 72 h. c Apoptotic cell death was measured by ELISA for detecting active caspase-3. Control-shRNA and ATG5-shRNA/A2780-CP20 cells were treated with CDDP and CQ as indicated for 48 h, and cell lysates were used for caspase-3 assay. Results are shown as the mean ± SD of duplicate observations from three experiments (n = 3, **P < 0.01, ***P < 0.001). d To analyze DNA damage recognition proteins and downstream cell cycle-related proteins, cell lysates were analyzed on a 4–12% gradient SDS-PAGE by western blot in shRNA/A2780-CP20 stable cells after incubation with CDDP, CQ, and CQ-CDDP combination for 48 h. Amounts of phosphorylated ATM and Cdc2 normalized by the amount of each total protein are represented by a bar graph in the lower panels. e Cell cycle distribution of A2780-CP20 stable cells expressing either control- or ATG5-shRNA was analyzed after incubation with CDDP, CQ or CQ combined with CDDP for 48 h (upper panel). Flow cytometry was performed for three experiments, and representative data are shown. Cell cycle distribution was represented by bar graphs and the statistical data demonstrated cell number in G2/M (lower panel) compared with the control. f Expression of p21WAF1/CIP1 was analyzed by western blot of A2780 and A2780-CP20 cells after incubation with CDDP, CQ, or CQ-CDDP combination. g A2780-CP20 cells were incubated with CDDP, CQ, ATR inhibitor, ATM inhibitor, or the combination of CDDP with either CQ, ATRi, or ATMi for 48 h. Expression of p21WAF1/CIP1 was analyzed by western blot. h Expression of p21WAF1/CIP1 was determined by western blot in shRNA/A2780-CP20 stable cells after incubation with CQ for 48 h. Expression of p21WAF1/CIP1 was determined by western blot analysis in control shRNA and ATG5 shRNA/A2780-CP20 cells treated with drugs as indicated. i A2780-CP20 cells were incubated with CDDP, CQ, bafilomycin A1 (Baf-A1), CQ-Baf-A1 combination, MG-132, or the combination of CDDP with CQ or MG-132 as indicated. CQ and CDDP were treated for 48 h and Baf-A1 and MG-132 were treated overnight. Western blot was performed using anti-p21WAF1/CIP1 antibody. Western blot analysis was performed at least three times, and representative data are presented in this study (mean ± SD, *P < 0.05, **P < 0.01, ***P < 0.001).
To study the effect of autophagy inhibition on DNA damage/cell cycle-related proteins, western blot analysis was performed. Total expression of ATM was inhibited by knockdown of ATG5. Treatment with CDDP in ATG5-shRNA cells induced phosphorylation of ATM compared with control-treated ATG5-shRNA cells (Fig. 5d). CQ-CDDP combination further induced phosphorylation of ATM. Although ATR expression was inhibited by CQ-CDDP combination in both control- and ATG5-shRNA cells, its phosphorylation was not affected. Knockdown of ATG5 induced expression of γH2AX, demonstrating that inhibition of autophagy enhanced DNA damage. Treatment of CDDP further induced γH2AX in ATG5-shRNA and control-shRNA cells. Interestingly, treatment of CQ-CDDP combination in ATG5-shRNA cells induced γH2AX to a lesser extent than the control-shRNA cells treated with CQ-CDDP combination. CQ alone had no effect on γH2AX expression. Cdc2-pY15 was increased by CQ-CDDP combination in ATG5-shRNA cells, demonstrating inhibition of Cdc2 by inhibition of autophagy. Cdc2 and cyclin B1 are involved in G2/M cell cycle progression, and the effect of inhibition of autophagy on cell cycle distribution was assessed. G2/M arrest was increased in ATG5-shRNA cells treated with CDDP and was further induced by CQ-CDDP combination (Fig. 5e).
p21WAF1/CIP1 is activated by DNA damage and induced G2/M arrest23; therefore, the effect of CQ-CDDP on p21WAF1/CIP1 expression was investigated. In A2780 and RMG-1 cells, protein expression of p21WAF1/CIP1 was highly induced by CDDP (Fig. 5f and Supplementary Fig. S5, respectively). However, p21WAF1/CIP1 expression was highly induced by CQ alone and the CQ-CDDP combination in A2780-CP20 cells. We examined whether ATMi or ATRi inhibitors could induce p21WAF1/CIP1. Induction of p21WAF1/CIP1 was hardly observed in cells treated with ATRi alone but ATRi combined with CDDP highly induced its expression (Fig. 5g). However, ATMi alone and ATMi-CDDP combination had no effect on induction of p21WAF1/CIP1. To determine whether autophagy inhibition induced p21WAF1/CIP1, expression of p21WAF1/CIP1 was assessed in ATG5-shRNA cells (Fig. 5h). In control-shRNA cells, single treatment of CDDP and CQ induced p21WAF1/CIP1, and its expression was further enhanced upon treatment with CQ-CDDP combination. However, although knockdown of ATG5 induced p21WAF1/CIP1 by CDDP, CQ, and CQ-CDDP combination, it was not as strong as in the control-shRNA cells, demonstrating that p21WAF1/CIP1 was not efficiently induced in ATG5-shRNA cells. Another autophagy inhibitor, Bafilomycin A1 (Baf-A1), was used to examine effect of autophagy on induction of p21WAF1/CIP1. Induction of p21WAF1/CIP1 was compared with that in cells treated with CQ-CDDP, Baf-A1, the combination of Baf-A1-CQ, the proteasome inhibitor MG-132, or co-treatment with MG-132 and CDDP (Fig. 5i). Baf-A1 alone slightly induced p21WAF1/CIP1 expression, while Baf-A1-CQ combination further induced its expression. However, treatment of MG-132 or its combination with CDDP strongly induced p21WAF1/CIP1. These findings suggest that enhanced p21WAF1/CIP1 expression by the CQ-CDDP combination may be due to inhibition of both autophagy and the proteasome by CQ.
CQ-CDDP-induced cell death was partially dependent on induction of p21WAF1/CIP1
The functional significance of CQ-induced p21WAF1/CIP1 was studied by knockdown of p21WAF1/CIP1 with its specific shRNA in A2780-CP20 cells and by its ectopic expression with HA-tagged p21WAF1/CIP1. Knockdown efficiency of p21WAF1/CIP1 was analyzed by western blot (Fig. 6a). Proteins involved in DNA damage and cell cycle arrest were assessed in p21WAF1/CIP1-shRNA cells compared with control-shRNA cells. Combination of CQ-CDDP as well as CDDP and CQ single treatment induced ATM protein expression in p21WAF1/CIP1-shRNA cells compared with control cells. However, phosphorylated ATM was reduced in p21WAF1/CIP1-shRNA cells. ATR protein expression was also increased in CQ-CDDP-treated p21WAF1/CIP1-shRNA cells compared with CQ-CDDP-treated control cells. However, phosphorylation of ATR was not affected in p21WAF1/CIP1-shRNA cells. Expression of γH2AX was highly reduced by CQ-CDDP combination in p21WAF1/CIP1-shRNA cells (Fig. 6b). Cdc2-pY15 was also decreased in p21WAF1/CIP1-shRNA cells treated with CQ-CDDP compared with CQ-CDDP-treated control cells, demonstrating reduced inhibition of Cdc2 by knockdown of p21WAF1/CIP1. Caspase-3 activity in CQ-CDDP combination-treated cells was inhibited by about 20% by knockdown of p21WAF1/CIP1 compared with that of CQ-CDDP-treated control cells (Fig. 6c). Although knockdown of p21WAF1/CIP1 also induced G2/M arrest by CDDP and CQ-CDDP combination, the number of G2/M-arrested cells were reduced from 51.29% to 47.62% and from 57.75% to 42.26% in CDDP- and CQ-CDDP-treated p21WAF1/CIP1-shRNA cells compared with the control cells, respectively (Fig. 6d). Ectopic expression of HA-p21WAF1/CIP1 induced phosphorylation of ATM but decreased ATM expression in CDDP-treated ATG5-shRNA cells compared with CDDP-treated vector-transfected cells (Fig. 6e). In control-shRNA cells, HA-p21WAF1/CIP1 in the presence of CDDP induced γH2AX compared with CDDP-treated vector-transfected control-shRNA cells (Fig. 6f). In ATG5-shRNA cells, γH2AX was increased compared with control-shRNA cells and the enhanced expression was further increased by HA-p21WAF1/CIP1 and CDDP treatment. Ratio of p-Cdc2/Cdc2 was increased in CDDP-treated HA-p21WAF1/CIP1-transfected cells, demonstrating inhibition of Cdc2 by HA-p21WAF1/CIP1. Caspase-3 activity was increased in CDDP-treated HA-p21WAF1/CIP1-transfected cells compared with the CDDP-treated vector-transfected cells (Fig. 6g). These results demonstrate that overexpressed p21WAF1/CIP1 may partially replace the function of CQ in the presence of CDDP.
a, b Proteins related to DNA damage and cell cycle progression were analyzed by western blot in control-shRNA and p21WAF1/CIP1-shRNA/A2780-CP20 cells. The amount of ATM normalized by β-actin is represented by a bar graph. Amounts of phosphorylated ATM and Cdc2 normalized by the amount of each total protein are represented by a bar graph in the right panel (upper and lower panels, respectively). c Active caspase-3 was measured in control-shRNA and p21WAF1/CIP1-shRNA cells after treatment of drugs for 48 h as indicated. Results are shown as the mean ± SD of triplicate observations from three experiments. d Cell cycle distribution of A2780-CP20 stable cells expressing either control- or p21WAF1/CIP1-shRNA was analyzed after incubation with CDDP, CQ, or CQ combined with CDDP for 48 h. Flow cytometry was performed for three experiments, and representative data are shown. Cell cycle distribution was represented by bar graphs and the statistical data demonstrated cell number in G2/M (right panel) compared with the control. e, f HA-tagged p21WAF1/CIP1 or the empty vector were transfected into control-shRNA and ATG5-shRNA/A2780-CP20 cells. After transfection, cells were incubated with CDDP for 48 h. Expression of HA-tagged p21WAF1/CIP1 was analyzed by western blot using anti-p21WAF1/CIP1 antibody. Expression of proteins involved in DNA damage and cell cycle progression as well as γH2AX was detected by western blot using their specific antibodies. β-actin served as a protein-loading control. Arrow indicates ATR expression. g Caspase-3 activity was measured by detecting active caspase-3 in cells transfected with HA-tagged p21WAF1/CIP1 plasmid or the control vector, followed by incubation with CDDP. Results are shown as the mean ± SD of duplicate observations from three experiments (n = 3, mean ± SD, *P < 0.05, **P < 0.01, ***P < 0.001, ns not significant).
The CQ-CDDP combination significantly decreases tumor weight in orthotopic and drug-resistant PDX mouse models for EOC
To test the efficacy of CQ-CDDP combination in vivo, we used an orthotopic mouse model established with A2780-CP20 cells (Fig. 7a–d) and a PDX model established with human EOC tissue35 (Fig. 7e–h). The human EOC tissue used in generation of a PDX model was from a 62-year-old woman with platinum resistance who was heavily pretreated with chemotherapy of paclitaxel-carboplatin, docetaxel, belotecan, and etoposide-ifosphamide and initially presented with FIGO stage IIIC, high-grade serous histology. The tumors obtained from orthotopic and PDX xenograft mice injected with the CQ-CDDP combination, were significantly smaller than those obtained from control, CDDP alone, and CQ alone groups (Fig. 7a and e, respectively). Immunohistochemical staining for Ki67, a proliferation marker protein, revealed that the numbers of proliferating cells in tumors obtained from the CQ-CDDP combination groups in orthotopic and PDX xenograft mice were reduced by 50 and 60% of that in controls, respectively (Fig. 7b and f). The combination group showed significantly increased immunostaining of γH2AX compared with the control groups (Fig. 7c and g). Expression of p21WAF1/CIP1 was increased in tumors from the CQ-CDDP combination group in orthotopic and PDX xenograft mice compared with the control groups (Fig. 7d and h, respectively).
a Drug-resistant EOC orthotopic mice were generated by i.p. injection of A2780-CP20 cells, and each drug was injected into the mice. After mice were sacrificed, tumors were isolated and weighed. Tumor weights are shown on a scatter plot. b–d Tumor tissues obtained from orthotopic mice were stained with antibodies against Ki67, a proliferation marker; γH2AX, a DNA damage marker protein; or p21WAF1/CIP1 (X200, left panels). Bar graphs (right panels) show numbers of Ki67-, γH2AX-, or p21WAF1/CIP1-stained nuclei relative to total hematoxylin-stained nuclei. Scale bar represents 200 μm. e EOC PDX mice were generated using subrenal implantation with a tumor from a patient with drug-resistant serous papillary adenocarcinoma; each drug or the combination of CQ and CDDP was injected into the peritoneal cavity of xenograft mice. Left panel shows tumor size compared with a kidney obtained from the same mouse, and right panel shows tumor weight on a scatter plot. f–h Tumor tissues obtained from PDX mice were stained with antibodies against Ki67, γH2AX, and p21WAF1/CIP1 (X200, left panels). Scale bar represents 200 μm. Bar graphs (right panels) show numbers of Ki67-, γH2AX-, or p21WAF1/CIP1-stained nuclei relative to total hematoxylin-stained nuclei. All data represent the mean ± SD (*P < 0.05, **P < 0.01, ***P < 0.001). i A proposed functional mechanism of CQ-CDDP in chemo-resistant EOC.
Discussion
In this study, we found that CQ increases cytotoxicity in combination with CDDP by inducing lethal DNA damage through induction of p21WAF1/CIP1 expression and autophagy inhibition in CDDP-resistant EOC.
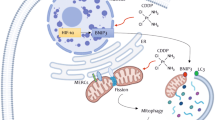
A proposed functional mechanism of CQ-CDDP combination in drug-resistant EOC is presented in Fig. 7i. CDDP induces DNA damage and activates a signaling cascade involved in DNA damage/cell cycle regulation. In chemo-sensitive cancer, CDDP activates ATM/ATR signaling cascades by triggering DNA damage, activates p53 to induce p21WAF1/CIP1, arrests cell cycle, inhibits DNA repair, and finally leads to cell death. However, in chemo-resistant cancer, cells can survive CDDP-induced DNA damage by overcoming cell cycle arrest. Treatment with CQ combined with CDDP in chemo-resistant cancer induces p21WAF1/CIP1. Although total ATM expression is decreased, the induced p21WAF1/CIP1 activates ATM and inhibits Cdc2, leading to G2/M arrest. Expression of p21WAF1/CIP1 highly induces γH2AX, demonstrating increased DNA damage and potential cell death. Autophagy inhibitory function of CQ inhibits ATM/ATR expression but slightly induces activation of ATM and inhibition of Cdc2 and thereby induces persistent G2/M arrest. Increased DNA damage (detected by γH2AX expression level) by CQ-CDDP finally leads to cell death. Therefore, induction of p21WAF1/CIP1 and inhibition of autophagy may additively function in cytotoxicity of CQ-CDDP combination in drug-resistant EOC.
We also studied the effect of CQ-CDDP combination on signaling pathways related to autophagy and apoptosis. Akt regulates autophagy and apoptosis, and the effect of CQ-CDDP on Akt was determined (Supplementary Fig. S6). CDDP-induced Akt expression and activated Akt activity in A2780-CP20 cells as measured by phosphorylation of Akt using its phosphor-specific antibody. However, CQ inhibited CDDP-induced Akt activity and its expression. These results demonstrate that inhibition of Akt by CQ-CDDP combination may play a role in cytotoxicity of EOC.
The cytotoxic effect of CQ is related to mitochondrial dysfunction induced by increasing mitochondrial reactive oxygen species (ROS)36. Therefore, mitochondrial ROS was measured in A2780-CP20 cells treated with drugs using MitoSox kit (Supplementary Fig. S6b). CQ-induced mitochondrial ROS and this induction was further increased by CQ-CDDP combination. Nrf2 is an antioxidant protein, and its interacting protein, Keap1, is a negative regulatory protein of Nrf2 via induction of its degradation37. p62 (an autophagy marker protein) interacts with Keap1 and thereby inhibits degradation of Nrf2. Expression of Nrf2 and Keap1 was assessed (Supplementary Fig. S6c). CDDP induced Keap1 and CQ inhibited CDDP-induced Keap1 expression. Expression of Nrf2 was not significantly changed. These results suggest that CQ-induced mitochondrial ROS was not related to the expression level of Nrf2. Taken together, CQ-induced mitochondrial ROS may contribute to the cytotoxic effect of CQ when combined with CDDP.
Cellular senescence leads to permanent cell cycle arrest38. The effect of CQ on senescence was assessed by staining A2780 and A2780-CP20 cells with senescence-associated β-galactosidase after treatment with drugs as indicated (Supplementary Fig. S6d). In A2780 cells, CDDP induced cellular senescence, while CQ inhibited CDDP-induced senescence. In A2780-CP20 cells, CDDP did not significantly induce senescence. Thus, senescence may not be a critical factor for CQ-CDDP-mediated cytotoxicity in drug-resistant EOC.
Studies on inhibitors to target DNA damage/cell cycle checkpoint for therapeutic purposes in different type of cancers are ongoing39,40,41. Recently, inhibitors for ATM and ATR have been investigated in preclinical applications42,43. In this study, ATMi-CDDP combination increased inactive Cdc2 but did not induce γH2AX expression. H2AX is phosphorylated by ATM, ATR, and DNA-PK. CDDP activates ATM upon DNA damage44. Therefore, in the CDDP-induced DNA damage/repair signaling pathway, ATM is more likely to be important protein for γH2AX than other kinases. In contrast, ATRi-CDDP combination induced expression of γH2AX but inactivation of Cdc2 was not observed. The phosphorylated Cdc2 at Y15 is dephosphorylated by Cdc25C phosphatase and thereby is degraded. Cdc25C is inhibited by the ATR/Chk1 signaling pathway, and inhibition of ATR by its inhibitor may result in active Cdc25C and thereby enhanced Cdc2 degradation45. These results demonstrated that, although CQ, ATRi, and ATMi presented similar results in DNA damage/cell cycle signaling pathway in the presence of CDDP, the detailed functional mechanisms may differ in these drugs.
In this study, CQ and CQ-CDDP combination induced p21WAF1/CIP1 expression in drug-resistant EOC, and this induction was not sufficient in ATG5-shRNA cells compared with control-shRNA cells. It seems that p21WAF1/CIP1 may be degraded via a proteasome degradation pathway in drug-resistant EOCs since p21WAF1/CIP1 was highly increased in the presence of MG-132. In support of our data, proteasomal degradation of p21WAF1/CIP1 has been reported in several cell lines46,47,48. Furthermore, inhibition of lysosomal degradation by CQ led to inhibition of proteasomal degradation, but knockdown of ATG5 could not inhibit proteasomal activity49,50, consistent with our result that induction of p21WAF1/CIP1 was not sufficient in ATG5-shRNA cells. These study findings indicate that CQ may enhance p21WAF1/CIP1 expression by inhibition of proteasomal degradation and by partly its autophagy inhibition.
In conclusion, we found that CQ in combination with CDDP increased the cytotoxicity of drug-resistant EOC cells mainly by inducing lethal DNA damage through induction of p21WAF1/CIP1 expression and autophagy inhibition. This study suggests that CQ may be a promising drug for combination therapy with anti-cancer drugs to treat chemo-resistant EOCs.
References
Kim, S. et al. Tumor evolution and chemoresistance in ovarian cancer. NPJ Precis Oncol. 2, 20 (2018).
Agarwal, R. & Kaye, S. B. Ovarian cancer: strategies for overcoming resistance to chemotherapy. Nat. Rev. Cancer 3, 502–516 (2003).
Chen, X. et al. Sequential combination therapy of ovarian cancer with cisplatin and gamma-secretase inhibitor MK-0752. Gynecol. Oncol. 140, 537–544 (2016).
Vetter, M. H. & Hays, J. L. Use of targeted therapeutics in epithelial ovarian cancer: a review of current literature and future directions. Clin. Ther. 40, 361–371 (2018).
Dasari, S. & Tchounwou, P. B. Cisplatin in cancer therapy: molecular mechanisms of action. Eur. J. Pharm. 740, 364–378 (2014).
Roos, W. P. & Kaina, B. DNA damage-induced cell death: from specific DNA lesions to the DNA damage response and apoptosis. Cancer Lett. 332, 237–248 (2013).
Sakthivel, K. M. & Hariharan, S. Regulatory players of DNA damage repair mechanisms: role in cancer chemoresistance. Biomed. Pharmacother. 93, 1238–1245 (2017).
Nagel, Z. D. et al. DNA repair capacity in multiple pathways predicts chemoresistance in glioblastoma multiforme. Cancer Res. 77, 198–206 (2017).
Ashwell, S. & Zabludoff, S. DNA damage detection and repair pathways–recent advances with inhibitors of checkpoint kinases in cancer therapy. Clin. Cancer Res. 14, 4032–4037 (2008).
Marechal, A. & Zou L. DNA damage sensing by the ATM and ATR kinases. Cold Spring Harb. Perspect. Biol. 5, a012716 (2013).
Mah, L. J., El-Osta, A. & Karagiannis, T. C. gammaH2AX: a sensitive molecular marker of DNA damage and repair. Leukemia 24, 679–686 (2010).
Sanchez, Y. et al. Conservation of the Chk1 checkpoint pathway in mammals: linkage of DNA damage to Cdk regulation through Cdc25. Science 277, 1497–1501 (1997).
Abraham, R. T. Cell cycle checkpoint signaling through the ATM and ATR kinases. Genes Dev. 15, 2177–2196 (2001).
Levine, B. & Kroemer, G. Autophagy in the pathogenesis of disease. Cell 132, 27–42 (2008).
Levy, J. M. M., Towers, C. G. & Thorburn, A. Targeting autophagy in cancer. Nat. Rev. Cancer 17, 528–542 (2017).
Mauthe, M. et al. Chloroquine inhibits autophagic flux by decreasing autophagosome-lysosome fusion. Autophagy 14, 1435–1455 (2018).
Draz, H. et al. Autophagy inhibition improves the chemotherapeutic efficacy of cruciferous vegetable-derived diindolymethane in a murine prostate cancer xenograft model. Invest N. Drugs 36, 718–725 (2018).
Xu, R., Ji, Z., Xu, C. & Zhu, J. The clinical value of using chloroquine or hydroxychloroquine as autophagy inhibitors in the treatment of cancers: a systematic review and meta-analysis. Medicine 97, e12912 (2018).
Kim, E. L. et al. Chloroquine activates the p53 pathway and induces apoptosis in human glioma cells. Neuro Oncol. 12, 389–400 (2010).
Liang, D. H. et al. The autophagy inhibitor chloroquine targets cancer stem cells in triple negative breast cancer by inducing mitochondrial damage and impairing DNA break repair. Cancer Lett. 376, 249–258 (2016).
Zhu, J. et al. Low concentration of chloroquine enhanced efficacy of cisplatin in the treatment of human ovarian cancer dependent on autophagy. Am. J. Transl. Res. 9, 4046–4058 (2017).
Shiratori, H. et al. The combination of temsirolimus and chloroquine increases radiosensitivity in colorectal cancer cells. Oncol. Rep. 42, 377–385 (2019).
Li, R. et al. Differential effects by the p21 CDK inhibitor on PCNA-dependent DNA replication and repair. Nature 371, 534–537 (1994).
Ramondetta, L., Mills, G. B., Burke, T. W. & Wolf, J. K. Adenovirus-mediated expression of p53 or p21 in a papillary serous endometrial carcinoma cell line (SPEC-2) results in both growth inhibition and apoptotic cell death: potential application of gene therapy to endometrial cancer. Clin. Cancer Res. 6, 278–284 (2000).
Tsao, Y. P. et al. Adenovirus-mediated p21((WAF1/SDII/CIP1)) gene transfer induces apoptosis of human cervical cancer cell lines. J. Virol. 73, 4983–4990 (1999).
Lincet, H. et al. The p21(cip1/waf1) cyclin-dependent kinase inhibitor enhances the cytotoxic effect of cisplatin in human ovarian carcinoma cells. Cancer Lett. 161, 17–26 (2000).
Liu, S., Bishop, W. R. & Liu, M. Differential effects of cell cycle regulatory protein p21(WAF1/Cip1) on apoptosis and sensitivity to cancer chemotherapy. Drug Resist. Updat. 6, 183–195 (2003).
Yang, H. L., Pan, J. X., Sun, L. & Yeung, S. C. p21 Waf-1 (Cip-1) enhances apoptosis induced by manumycin and paclitaxel in anaplastic thyroid cancer cells. J. Clin. Endocrinol. Metab. 88, 763–772 (2003).
Fizazi, K. et al. The association of p21((WAF-1/CIP1)) with progression to androgen-independent prostate cancer. Clin. Cancer Res. 8, 775–781 (2002).
Javelaud, D. & Besancon, F. Inactivation of p21WAF1 sensitizes cells to apoptosis via an increase of both p14ARF and p53 levels and an alteration of the Bax/Bcl-2 ratio. J. Biol. Chem. 277, 37949–37954 (2002).
Abbas, T. & Dutta, A. p21 in cancer: intricate networks and multiple activities. Nat. Rev. Cancer 9, 400–414 (2009).
Waga, S., Hannon, G. J., Beach, D. & Stillman, B. The p21 inhibitor of cyclin-dependent kinases controls DNA replication by interaction with PCNA. Nature 369, 574–578 (1994).
Oh, S. Y. et al. Autophagy regulates trophoblast invasion by targeting NF-kappaB activity. Sci. Rep. 10, 14033 (2020).
Mangala, L. S. et al. Therapeutic targeting of ATP7B in ovarian carcinoma. Clin. Cancer Res. 15, 3770–3780 (2009).
Heo, E. J. et al. Patient-derived xenograft models of epithelial ovarian cancer for preclinical studies. Cancer Res. Treat. 49, 915–926 (2017).
Qu, X. et al. Autophagy inhibitor chloroquine increases sensitivity to cisplatin in QBC939 cholangiocarcinoma cells by mitochondrial ROS. PLoS ONE 12, e0173712 (2017).
Jaramillo, M. C. & Zhang, D. D. The emerging role of the Nrf2-Keap1 signaling pathway in cancer. Genes Dev. 27, 2179–2191 (2013).
Rajendran, P. et al. Autophagy and senescence: a new insight in selected human diseases. J. Cell Physiol. 234, 21485–21492 (2019).
Mills, C. C., Kolb, E. A. & Sampson, V. B. Recent advances of cell-cycle inhibitor therapies for pediatric cancer. Cancer Res. 77, 6489–6498 (2017).
Wang, F. Z. et al. The checkpoint 1 kinase inhibitor LY2603618 induces cell cycle arrest, DNA damage response and autophagy in cancer cells. Apoptosis 19, 1389–1398 (2014).
Zhang, T. et al. The ATM inhibitor KU55933 sensitizes radioresistant bladder cancer cells with DAB2IP gene defect. Int. J. Radiat. Biol. 91, 368–378 (2015).
Min, A. et al. AZD6738, a novel oral inhibitor of ATR, induces synthetic lethality with ATM deficiency in gastric cancer cells. Mol. Cancer Ther. 16, 566–577 (2017).
Batey, M. A. et al. Preclinical evaluation of a novel ATM inhibitor, KU59403, in vitro and in vivo in p53 functional and dysfunctional models of human cancer. Mol. Cancer Ther. 12, 959–967 (2013).
Yimit, A., Adebali, O., Sancar, A. & Jiang, Y. Differential damage and repair of DNA-adducts induced by anti-cancer drug cisplatin across mouse organs. Nat. Commun. 10, 309 (2019).
Aleem, E. & Arceci, R. J. Targeting cell cycle regulators in hematologic malignancies. Front Cell Dev. Biol. 3, 16 (2015).
Blagosklonny, M. V., Wu, G. S., Omura, S. & el-Deiry, W. S. Proteasome-dependent regulation of p21WAF1/CIP1 expression. Biochem Biophys. Res. Commun. 227, 564–569 (1996).
Naujokat, C. et al. Proteasome inhibitors induced caspase-dependent apoptosis and accumulation of p21WAF1/Cip1 in human immature leukemic cells. Eur. J. Haematol. 65, 221–236 (2000).
Kim, H. J. et al. SPSB1 enhances ovarian cancer cell survival by destabilizing p21. Biochem Biophys. Res. Commun. 510, 364–369 (2019).
Myeku, N. & Figueiredo-Pereira, M. E. Dynamics of the degradation of ubiquitinated proteins by proteasomes and autophagy: association with sequestosome 1/p62. J. Biol. Chem. 286, 22426–22440 (2011).
Qiao, L. & Zhang, J. Inhibition of lysosomal functions reduces proteasomal activity. Neurosci. Lett. 456, 15–19 (2009).
Acknowledgements
We thank Dr. Anil K Sood (Department of Gynecologic Oncology and Department of Cancer Biology, University of Texas MD Anderson Cancer Center, Houston, TX, USA) for providing A2780-CP20 cells. This research was supported by the Basic Science Research Program through the National Research Foundation (NRF) funded by the Ministry of Science, ICT, and Future Planning (grant number: NRF-2017R1A2B4003434) and by grants from the National R&D Program for Cancer Control, Ministry for Health, Welfare and Family Affairs, Republic of Korea (1520100) and the National Research Foundation of Korea (NRF) funded by the Korean government (MEST) (No. 2016R1A2B3006644).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by B. Zhivotovsky
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hwang, J.R., Kim, W.Y., Cho, YJ. et al. Chloroquine reverses chemoresistance via upregulation of p21WAF1/CIP1 and autophagy inhibition in ovarian cancer. Cell Death Dis 11, 1034 (2020). https://doi.org/10.1038/s41419-020-03242-x
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-020-03242-x
- Springer Nature Limited