Abstract
Tumor progression and evolution are frequently associated with chromosomal instability (CIN). Tumor cells often express high levels of the mitotic checkpoint protein MAD2, leading to mitotic arrest and cell death. However, some tumor cells are capable of exiting mitosis and consequently increasing CIN. How cells escape the mitotic arrest induced by MAD2 and proliferate with CIN is not well understood. Here, we explored loss-of-function screens and drug sensitivity tests associated with MAD2 levels in aneuploid cells and identified that aneuploid cells with high MAD2 levels are more sensitive to FOXM1 depletion. Inhibition of FOXM1 promotes MAD2-mediated mitotic arrest and exacerbates CIN. Conversely, elevating FOXM1 expression in MAD2-overexpressing human cell lines reverts prolonged mitosis and rescues mitotic errors, cell death and proliferative disadvantages. Mechanistically, we found that FOXM1 facilitates mitotic exit by inhibiting the spindle assembly checkpoint (SAC) and the expression of Cyclin B. Notably, we observed that FOXM1 is upregulated upon aneuploid induction in cells with dysfunctional SAC and error-prone mitosis, and these cells are sensitive to FOXM1 knockdown, indicating a novel vulnerability of aneuploid cells.
Similar content being viewed by others
Introduction
Chromosomal instability (CIN), defined as the continuous loss or gain of chromosomes is the result of increased levels of mitotic errors and aneuploidy [1]. Although CIN promotes tumor evolution, drug resistance and tumor heterogeneity, excessive CIN leads to cell death [2,3,4]. The mechanisms conferring cancer cells tolerance to high levels of CIN are not fully understood [5]. The identification of specific targets against these bypass mechanisms could provide therapeutic strategies against CIN tumors [6, 7].
The spindle-assembly checkpoint (SAC) stalls the onset of anaphase until all kinetochores are properly bounded to microtubules during metaphase, preventing chromosome errors and CIN [8, 9]. Overexpression of SAC proteins is common among human cancers and defective SAC functioning facilitates ongoing CIN [10, 11]. Among these SAC proteins, MAD2 has been found to be upregulated in different types of cancer and its overexpression, in transgenic mouse models, induces mitotic arrest and increases the number of mitotic errors [12]. In addition, Mad2 overexpression in Kras-driven mouse breast tumors leads to increased somatic copy number alterations compared with Kras tumors [13]. However, how tumor cells overcome Mad2-induced mitotic arrest and tolerate Mad2-induced CIN is still unclear. Understanding the molecular mechanism behind could reveal novel therapeutic strategies for unstable MAD2-overexpressing cancers.
Microtubule (MT) poisons block mitosis by interfering with microtubule dynamics and therefore activating the SAC and have been widely applied in the treatment of solid cancers [14, 15]. Intriguingly, clinical trials indicate that the cytotoxic effect of microtubule-targeting drugs might not only rely on the induced mitotic arrest but also in the excessive CIN generation [16].
One key mediator of anti-mitotic therapeutic response is the Forkhead Box M1 (FOXM1) transcription factor, which is involved in mitotic progression, spindle assembly and chromosome segregation [17,18,19]. FOXM1 preserves mitotic spindle formation and prevents mitotic catastrophe induced by the MT poison paclitaxel [20] while repression of FOXM1 increases paclitaxel-induced mitotic cell death through modulation of the apoptotic pathway [21]. FOXM1 also improved age-associated mitotic defects in elderly human dermal fibroblasts, leading to decreased aneuploid levels [22].
Here, we identify FOXM1 to be essential for the survival of tumor cells with MAD2 overexpression (OE) in mouse and human cell lines. FOXM1 overexpression facilitates mitotic exit and maintains chromosome segregation fidelity of MAD2-overexpressing and nocodazole-treated cells by disrupting SAC signaling. Analysis of human tumors showed that high FOXM1 expression and increased aneuploidy were associated with poor prognosis, and cells with tetraploidization were more sensitive to depletion of FOXM1. Our results revealed that the upregulation of FOXM1 is a mechanism that allows cells to bypass mitotic arrest and tolerate CIN in MAD2-overexpressing cells.
Materials and methods
Human cancer cell lines data
Aneuploidy score (AS) and gene expression data sets were obtained from DepMap portal (https://depmap.org/portal/). Aneuploid and near euploid cell lines were split into two groups: the top and bottom quartiles of AS. Aneuploid cell lines were further separated as top and bottom sextile of MAD2 levels, according to the MAD2 expression levels to gain further insight into specific genes that are essential in the MAD2_high cell lines. Genetic dependency data sets were CRISPR (DepMap 22Q2 Public + Score, Chronos) and RNAi (Achilles + DRIVE + Marcotte, DEMETER2). Essential genes in high or low MAD2 aneuploid compared to near euploidy were determined by using R package Limma. The different dependency of each gene between groups was evaluated and considered as significant gene when the fold change of mean dependency was above 1 and the P-value was <0.01. P-values were derived from Wilcoxon t-test. To compare AS or gene expression among groups, we applied Prism9 to generate scatterplots. Drug sensitivity data sets were obtained from drug sensitivity (AUC) [23] and Drug Repurposing Secondary Screen 19Q4 [23]. The sensitivity of drug screens was compared among groups. P-values were determined by one-way ANOVA. EC50 was compared between euploidy and aneuploidy with high or low MAD2, respectively. P-values were determined by Wilcoxon t-test.
GEO and TCGA human cancer data analysis
Gene expression of human breast cancer data sets was obtained from Gene Expression Omnibus (GEO) datasets (http://www.ncbi.nlm.nih.gov/geo). Two groups of samples were determined as the top and bottom sextile of MAD2 levels.
Copy number alterations and gene expression (FPKM-UQ) of patient samples from TCGA were obtained from UCSC Xena functional genomics explorer. Data of mean absolute change in copy number segment were filtered to remove the bottom quartile across all segment lengths to better obtain ploidy changes. Top and bottom quartiles of mean absolute change were considered as aneuploid and euploid cancers. Comparison of genes between groups was done by Limma package. Genes with >2.5-fold change of expression and <0.01 of P-value were considered as significant. P-values were determined by Wilcoxon t-test.
Mouse models and Xenograft experiment
KH2-HA-Mad2, TetO-Mad2, TetO-KrasG12D, MMTV-rtTA and H2B-GFP mice were generated as described previously [13]. All animals were in FVB background and housed in specific pathogen-free conditions. Breeding and experiments were performed at DKFZ animal facilities under permit numbers G231/15 and G18/21 from Regierungspräsidium of Karlsruhe, Germany. To induce transgenes, 8-week-old female mice were administrated with doxycycline via impregnated food pellets (625 mg/kg; Harlan-Teklad). Tumor growth was regularly examined and tumors were collected when they reached 1.5 cm3. Xenografts of Balbc nude mice: 250,000 MCF-7 cells in 0.1 mL serum-free medium were injected subcutaneously in 6-week-old female mice. Nude mice were administrated with doxycycline food 3 days before cell injection. Tumors were measured every two days and harvested when size reached 1.5 cm3. Tumor size was calculated with formula length (L) × width (W)2/2. No statistical method was used to estimate the sample size of the mouse models or xenograft mice. The sample size was based on the previous experimental observations [13]. No data were excluded from the animal experiment. The researchers were blind to the allocation of the animal groups during the experiment.
RNA-seq analysis of mouse tumor samples
RNA-seq data was obtained from the European Nucleotide Archive (ENA; http://www.ebi.ac.uk/ena) under number PRJEB13611 [24]. The data matrix was processed with DEseq2 VST normalization. Comparison of gene expression between K and KM tumors was analyzed via Lima package. DEGs were identified if log2 fold changes >1.0 and P < 0.05. Metascape was used to perform pathway analysis. Significant pathways were identified if P < 0.05 and FDR < 0.5.
Cell culture and in vitro experiments
Mouse breast tumor cells were harvested from KH2-HA-Mad2, TetO-KrasG12D, MMTV-rtTA; TetO-Mad2, TetO-KrasG12D, MMTV-rtTA and from TetO-KrasG12D, MMTV-rtTA and cultured as previously published [13]. MEFs were prepared from KH2-HA-Mad2/Rosa26-rtTA and maintained using procedures described in [12]. MDA-MB-231 (RRID: CVCL_0062), MCF7 (RRID:CVCL_0031) and CAL51 (RRID: CVCL_1110) cells were cultured in DMEM (Life Technologies, 419650394) with 10% fetal bovine serum, Tetracycline free (VWR, S181T-500) and 1% penicillin-streptomycin (Life Technologies, 15140122). MCF10A (RRID: CVCL_0598) cells were maintained as described in [25]. All cell lines have been authenticated using STR profiling. All experiments were performed with mycoplasma-free cells. To generate MAD2 and FOXM1 inducible cell lines, cells were infected with rtTA or rtTA-GFP-expressing retrovirus. Selection was performed with puromycin (1 μg/ml) or FAC sorting, then infected with inducible Tet-ON lentiviruses carrying human MAD2 and FOXM1 cDNA (MAD2 from Addgene #136347; FOXM1b from Addgene #68811; FOXM1c from Addgene #68810) and selected with hygromycin (300 μg/ml) or puromycin (1 μg/ml). Doxycycline concentration in all cultured experiments was 1 μg/ml. To inhibit FOXM1, 200 μM siRNA and lipofectamine 2000 transfection reagent (Invitrogen) were prepared in Opti-MEM reduced-serum media (Gibco). Cells were treated with siRNA for the indicated number of days. The siRNA used were FOXM1 siRNA (human) (sc-37615), Foxm1 siRNA (mouse) (sc-44877), Control siRNA (sc-37007).
Cell cycle profile was performed by counterstaining DNA with propidium iodide (Life Technologies, BMS500PI) and analyzed by a FACS-Canto flow cytometry device (BD Biosciences). TUNEL staining was done following manufacture instructions (Roche, 12156792910). Cell proliferation was determined with the Promega, G3580 kit following manufacture instructions. For immunostaining, cells were first harvested in a dilute suspension and deposited onto a slide with a cytospin centrifuge, according to protocols (Thermo, TH-CYTO4). Then cells were fixed with 4% paraformaldehyde (VWR, J61899.AP). Blocking was done using 10% donkey or goat serum (Jackson Immuno) in PBS with 0.15% triton X. Primary antibodies were : FOXM1 (1:500, Abcam, ab207298), MAD2 (1: 500, BD biosciences, 610678), HA (1:1000, Covance, MMS-101R), FLAG (1:500, Sigma, F7425), pH3 (1:1000, Cell signaling, 9701 S), Aurora B (1:500, BD biosciences, 611082). Secondary antibodies were Alexa fluorophore-labeled donkey/goat IgG (1:500, Invitrogen). Images were taken in a Leica SP5 confocal microscope and Tissuegnostic TissueFAX system and analyzed using ImageJ. Time-lapse imaging was performed for at least 12 h using a time-lapse microscope 2 μm optical sectioning across 20 μm stack, every 5 min.
Concentrations of nocodazole (Sigma, 487928) were 200 ng/ml, as indicated in figure legends. Tetraploid MCF7 and CAL51 were generated by cytokinesis inhibition using 0.75 μM dihydrocytochalasin B (DCB, inhibitor of actin polymerization, Sigma-Aldrich D1641) for 18 h overnight. Afterwards, cells were washed 3 times with PBS and cultured in DMEM supplemented with 10% FBS and 1% penicillin-streptomycin for an additional 20 h.
Three-dimensional organotypic assays
Mammary epithelial cells were harvested from 8-week-old female mice. 3D cell culture and staining were performed according to [13]. Primary antibodies were HA (1:1000, Covance, MMS-101R), FOXM1 (1:500, HUABIO, ER 1706-62), Images were taken with a Leica TCS SP5 confocal microscope and analyzed using ImageJ.
Quantitative PCR
Frozen tissue from mice was ground with mortar and pestle on dry ice and RNA was purified using RNeasy Mini Kit (Qiagen). To synthesize cDNA, we used QuantiTect Reverse Transcription Kit (Qiagen). For real-time quantification, 8 ng cDNA was used as a starting material along with SYBR Green PCR Master Mix (2x) (Applied Biosystems) in a LightCycler II ® 480 (Roche). PCR program : 95 °C for 5', 45x [95 °C for 10'', 60 °C for 15'', 72 °C for 15''], [95 °C for 5', 65 °C for 1’]; Formulas for calculation of gene expression: ΔCt = Ct (gene of interest) – Ct (reference gene); ΔΔCt = ΔCt – ΔCt (reference sample); Primers were Actin F: GCTTCTTTGCAGCTCCTTCGT and Actin R: ACCAGCCGCAGCGATATCG; FOXM1 F: GCGTTAAGCAGGAACTGGAA and FOXM1 R: TCAGACACAGAGTCCTGCCA.
Western blot
40 μg of cell lysates obtained from ground-frozen tissue or cells were used for assessing protein expression. Primary antibodies used for detection were FOXM1 (1:1000, Abcam, ab207298), MAD2 (1:1000, BD biosciences, 610678), HA (1:1000, Covance, MMS-101R), Cyclin B (1:200, Santa Cruz, SC-245), Cleaved-Caspase 3 (1:1000, Cell signaling, 9661), p-Histone H2AX (1:250, Santa Cruz, SC-517348), CDC20 (1:250, Santa Cruz, SC-13162), Actin (1:3000, Sigma, A2066). Protein band quantification was carried out using ImageJ.
Statistical analysis
Prism9 was used for statistical analysis. Statistical analyses between two groups were carried out using unpaired t-test, between more than two groups were performed with one-way analysis of variance, followed by Tukey’s multiple comparisons test or two-way analysis of variance, followed by Sidak’s multiple comparisons test. P-values were indicated in figure legends. P-value < 0.05 was considered as significant. Scatterplots were shown as mean and SEM. Points and connecting lines were shown as mean and error with SEM. Cell number and animal number were presented as n.
Results
High levels of MAD2 confer aneuploid cells sensitivity to FOXM1 inhibition
To understand how the genetic landscape in human tumors is altered by MAD2 expression, we performed independent analysis from data obtained from a human cancer cell line dataset (DepMap, n = 1389) and human breast tumors (GSE102484, n = 683; GSE54002, n = 417; GSE76275, n = 265). Cancer cell lines and GSE tumors were divided into sextiles according to MAD2 expression levels. Differential expression analysis between the top and bottom sextiles revealed 62 differentially expressed genes (62 DEG) that were frequently upregulated in all MAD2_high cancer cell lines and MAD2_high tumors (P < 0.01; log2 fold change > 1.5) (Supplementary Fig. 1A).
Next, to understand how cells with high levels of MAD2 are able to tolerate CIN, we investigated the vulnerabilities of aneuploid MAD2_high cell lines. Cancer cell lines from DepMap were divided into quartiles based on their aneuploidy score (AS) [26]. Aneuploid cell lines (Top AS quartiles) were further distributed into sextiles according to their MAD2 levels. MAD2_high (Top sextile) and MAD2_low (Bottom sextile) groups were considered for further analysis. The cell lines in the low AS quartile were considered the near euploid group. Based on this stratification, we performed a comparison of genetic dependency between MAD2_high aneuploid (aneuploid with high levels of MAD2) and euploid cell lines (Fig. 1A). Analysis of CRISPR-Cas9 datasets revealed that 115 genes were essential in MAD2_high aneuploid cells (Fig. 1B, Table 1) [26]. Three genes (RACGAP1, ASPM, FOXM1) were present in both DEGs and CRISPR datasets (Supplementary Fig. 1B). We then investigated RNAi (Achilles+DRIVE) datasets to look for genes whose depletion was more lethal to MAD2_high aneuploid cell lines than to euploid ones. We identified 39 differential dependencies of MAD2_high aneuploid cells (Fig. 1C, Table 1) and confirmed that FOXM1 was a common dependency for MAD2_high aneuploid cells in both RNAi and CRISPR datasets (Fig. 1D). Further analysis showed that MAD2_high aneuploid cells exhibited increased FOXM1 mRNA levels compared to euploid cells or MAD2_low aneuploid ones (Fig. 1E).
A Schematic of our comparison of genetic and chemical dependencies between MAD2 high and low cancer cell lines. Cell lines were assigned aneuploidy scores (AS), and the genetic and drug sensitivity landscapes were compared between the top and bottom AS quartiles. Cell lines in the top quartile of aneuploidy scores were divided into top and bottom sextiles according to their MAD2 expression (MAD2_high aneuploid vs near euploid cell lines and MAD2_low aneuploid vs near euploid cell lines). B Essential genes in MAD2_high aneuploid cells when compared to euploid cells, based on a CRISPR-Cas9 screen. The unique differential genetic dependencies in MAD2_high and euploid groups are shown in a volcano plot. The three genes present in both DEGs and CRISPR datasets are highlighted in red. C Essential genes for MAD2_high and euploid cell lines in RNAi datasets are shown as relative dependency. FOXM1 is highlighted in red. D The number of essential genes of MAD2_high aneuploid in RNAi and CRISPR datasets. E Comparison of aneuploidy scores and mRNA expression levels of MAD2 and FOXM1 between euploid and aneuploid cancer cell lines. ****P < 0.0001 and ***P = 0.0002; One-way ANOVA.
FOXM1 controls cell cycle-related gene expression and regulates chromosome stability [27, 28]. To understand if the requirement of FOXM1 is associated with aneuploidy or with MAD2 levels, we examined essential genes of MAD2_low aneuploid cells as well as those in MAD2_high euploid cells. In both cases, FOXM1 was not found in either CRISPR or RNAi datasets (Table 2). Moreover, CRISPR and RNAi datasets analyses indicated that MAD2_high euploid cells are not sensitive to FOXM1 inhibition (Supplementary Fig. 1D). Since FOXM1 mRNA expression is similar in MAD2_high euploid and aneuploid cells (Supplementary Fig. 1C), we hypothesized that FOXM1 essentiality might be related to MAD2 expression and aneuploidy status.
To further confirm if FOXM1 is crucial in MAD2_high aneuploid cells, we examined two different drug screening datasets between MAD2_high aneuploid cell lines and euploid cell lines (Fig. 1A). Drug sensitivity data (AUC) suggested that MAD2_high aneuploid cells were more sensitive than euploid cells to the FOXM1 inhibitor Thiostrepton (Supplementary Fig. 1E). Moreover, FOXM1 has been described to be a target for the proteasome inhibitors Bortezomib, Delanzomib and Oprozomib [29, 30]. Analysis of PRISM drug dose-level dataset indicated that MAD2_high aneuploid cells were more sensitive to proteasome inhibitors than euploid cells (Supplementary Fig. 1F). Altogether, these results suggest that aneuploid cells with high MAD2 levels are more vulnerable to FOXM1 inhibition.
FOXM1 inhibition decreases mitotic fidelity in short-term MAD2 overexpressing cells
Dysregulation of FOXM1 disrupts mitosis and increases CIN [31, 32]. To study the impact of FOXM1 inhibition on mitosis in MAD2-overexpressing cells (MAD2 OE), we generated doxycycline (Dox) inducible human MAD2-expressing breast cancer cell lines (MCF7, CAL51, MDA-MB-231, and MCF10A). MAD2 overexpression resulted in increased FOXM1 levels after 6 days (Fig. 2A and Supplementary Fig. 2A and 2B) and reduced cell viability in all cell lines over time (Fig. 2B and Supplementary Fig. 2C). We then used siRNAs against FOXM1 in these cell lines, infected with an empty vector or with the Dox inducible MAD2 vector and monitored the consequences of FOXM1-downregulation (FOXM1 KD) 6 days after induction. Cell viability was significantly reduced in MAD2 OE cells after FOXM1 KD compared to the inhibition of FOXM1 alone (Fig. 2C and Supplementary Fig. 2D).
A Western blots of FOXM1 and MAD2 in human breast cell lines infected with an empty vector (EV) or a Dox-inducible MAD2 expressing vector (MAD2 OE) after dox administration for 6 days. ACTIN was used as a loading control. Three biological replicates were analyzed. B Cell viability of each human cell line after dox treatment for 3 or 6 days. Each dox-treated cell line was normalized to the untreated one. MCF7: ***P < 0.0005; MDA-MB-231: ***P < 0.0001; CAL51: ****P < 0.0001, ***P = 0.0004, **P = 0.0027; MCF10A: **P < 0.008, *P = 0.0359, One-way ANOVA. Each dot is a biological replicate. C Cell viability of human cell lines after MAD2 overexpression and FOXM1 knockdown by siRNA for 6 days. Values of each cell line were normalized to those of each EV group. Each dot is a biological replicate. MCF7: *P = 0.0228; CAL51: *P = 0.0124; Two-way ANOVA. D Mitotic duration of MCF7 cells with or without MAD2 overexpression and siFOXM1 after 3 days. *P = 0.01, ****P < 00001; Two-way ANOVA. (EV, 100 cells; MAD2 OE, 92 cells; FOXM1 KD, 95 cells; MAD2 OE/FOXM1 KD, 119 cells in at least 3 independent experiments). E Cell fate of MCF7 cells represented as the incidence of each cell fate/total number of mitotic cells. (EV, 100 cells; MAD2 OE, 92 cells; FOXM1 KD, 95 cells; MAD2 OE/FOXM1 KD, 119 cells in at least 3 independent experiments). F Mitotic duration of CAL51 cells with or without MAD2 overexpression and siFOXM1 after 3 days. *P = 0.02, ****P < 00001; Two-way ANOVA. (EV, 133 cells; MAD2 OE, 237 cells; FOXM1 KD, 95 cells; MAD2 OE/FOXM1 KD, 234 cells, in at least 3 independent experiments). G Cell fate of CAL51cells represented as the incidence of each cell fate/total number of mitotic cells. (EV, 133 cells; MAD2 OE, 237 cells; FOXM1 KD, 95 cells; MAD2 OE/FOXM1 KD, 234 cells, in at least 3 independent experiments). H Representative western blot of KH2-HA-Mad2/Rosa26-rtTA MEFs and quantification in 5 MEFs lines, that were either not induced (CTRL) or on Dox for 30 h. Actin was used as loading control. FOXM1/ACTIN levels were normalized to control groups. *P = 0.0193; Unpaired t-test. 5 biological replicates were analyzed. I Mitotic duration of MEFs with or without MAD2 overexpression and siFOXM1 (CTRL, 38 cells; MAD2 OE, 64 cells; FOXM1 KD, 53 cells; MAD2 OE/FOXM1 KD, 66 cells in 3 independent movies). ***P = 0.0007; ****P = < 0.0001. One-way ANOVA. J Cell fate of MEFs with or without MAD2 overexpression and siFOXM1 (CTRL, 38 cells; MAD2 OE, 64 cells; FOXM1 KD, 53 cells; MAD2 OE/FOXM1 KD, 66 cells in 5 MEFs cell lines). K Representative western blot of FOXM1 and HA-Mad2 in EP cells from two TetO-Mad2/MMTV-rtTA Dox-inducible transgenic mice (M1 and M2) without DOX (CTRL) and with DOX. Actin was used as loading control. L Representative immunofluorescence images of HA, FOXM1 and DAPI on fixed mammary spheroid cultures after 36 h on Dox or/and FOXM1 inhibition by RCM-1. Scale bar: 10 μm. Percentage of apoptotic cells per acinus was calculated as apoptotic cells/total cells in each acinus (CTRL, 6 acini; MAD2 OE, 56 acini; FOXM1 KD, 24 acini; MAD2 OE/FOXM1 KD, 32 acini). ***P = 0.0004, *P = 0.0329 and 0.0178; One-way ANOVA. For original and additional wbs see Supplemental Material. Raw data for D,E,F,G, I J is included in Supplemental Tables.
We then selected two of these cell lines, one unstable breast cancer cell line, MCF7 and one diploid cell line, CAL51, and performed live cell imaging to study if mitotic defects or mitotic timing were contributing to the reduced viability. Time-lapse microscopy revealed that FOXM1 KD prolonged mitotic duration in MAD2 expressing MCF7 and CAL51 cells but not in the ones infected with an empty vector (Fig. 2D and F). In addition, we found a significant increase in the number of mitotic errors when FOXM1 was downregulated in both cell lines (Fig. 2E and G and Supplementary Fig. 2E, F). Strikingly, MAD2-expressing cell lines had the highest rates of mitotic errors after FOXM1 depletion, suggesting that FOXM1 depletion not only affected mitotic duration but also increased the incidence of lagging chromosomes and other mitotic errors when MAD2 was overexpressed.
We next generated mouse embryonic fibroblasts (MEFs) from KH2-Mad2/Rosa26-rtTA Dox inducible transgenic mice [13]. Induction of MAD2 after 30 h on Dox led to higher FOXM1 protein levels (Fig. 2H). We previously published that overexpression of MAD2 causes mitotic delay and CIN in these cells [12]. Inhibition of FOXM1 had no significant effect on mitotic duration in wild-type MEFs while it induced a mild mitotic delay in MAD2-overexpressing cells (Fig. 2I). In addition, we observed increased number of mitotic errors upon MAD2 OE or FOXM1 KD in cells that were further increased in the combination of both. Interestingly, an increase in cell death and mitotic errors was observed in MAD2 OE cells after FOXM1 KD (Fig. 2J). These results indicate a causal link between MAD2 induced CIN and FOXM1 dependency. Altogether, our results show that whereas normal cells tolerate the downregulation of FOXM1, it results in prolonged mitosis and increased CIN levels in MAD2 OE cells.
Finally, we used mammary epithelial (EP) cells from our TetO-HA-Mad2/MMTV-rtTA doxycycline-inducible transgenic mice (M) where short-term MAD2 overexpression has been shown to induce mitotic arrest and cell death [13]. Consistent with our observations in MAD2 OE MEFs, we observed high FOXM1 protein levels after MAD2 induction in EP cells (Fig. 2K) and immunofluorescence staining confirmed that tipically cells with HA-MAD2 overexpression showed FOXM1 upregulation (Supplementary Fig. 2G). We then measured the effect of FOXM1 downregulation using the RCM-1 inhibitor on 3D cultures grown from the EP of M mice [33]. MAD2 overexpression led to the accumulation of dying cells with no expression of FOXM1, while cells at the rim of these spheres retained expression of HA-MAD2 and FOXM1. Additional inhibition of FOXM1 in MAD2 overexpressing organoids led to an increase in the number of apoptotic cells and the shrinkage and condensation of the acinar structures, while FOXM1 KD in normal EP cells did not increase apoptotic cells (Fig. 2L). Thus, FOXM1 KD is detrimental in human breast cancer cells and mouse cells that express high levels of MAD2.
FOXM1 is required for the fitness of high MAD2 unstable cells
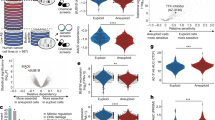
MAD2 overexpression in Kras-induced breast tumors leads to the formation of chromosomally unstable tumors that retained MAD2 expression [13], suggesting that these MAD2 tumor cells adapt over time to high MAD2 levels. To investigate, whether Foxm1 can play a role in this adaptation process, we asked if long-term induction of MAD2 also resulted in increased FOXM1 levels. We harvested breast tumor cells from TetO-Kras/MMTV-rtTA transgenic mice (K) and TetO-Kras/TetO-Mad2/MMTV-rtTA mice (KM) and observed higher Foxm1 mRNA and protein levels in KM compared to K tumor cells (Fig. 3A, B). To explore the transcription profiling of tumor cells, RNA sequencing data from K and KM tumors was analyzed [24]. Gene set enrichment analysis (GSEA) revealed downregulation in pathways related to APC/C:CDH1 function, chromosomal segregation and anti-apoptotic signaling in KM tumors when compared to K tumors (Supplementary Fig. 3A). These results suggest that long-term MAD2 overexpression might lead to a dysfunctional regulation of mitosis and apoptosis in mouse breast tumor cells. To understand the consequences of FOXM1 inhibition after long-term MAD2 expression, we used siRNA to knockdown Foxm1 in K and KM tumor cells. siFoxm1 for 6 days in KM cells resulted in a significantly reduced cell viability (Fig. 3C), the accumulation of G2/M cells (Fig. 3D) and increased TUNEL-positive cells (Fig. 3E) when compared to non-treated ones, while inhibition of Foxm1 in K cells had no effect. Moreover, treatment with the RCM-1 inhibitor or siFoxm1 led to increased levels of Cleaved Caspase-3 and Gamma-H2AX (γ-H2AX) in KM cells (Fig. 3F). Although total inhibition of FOXM1 was not achieved in KM cells, we still observed that KM unstable cells are significantly more vulnerable than K cells to the downregulation of FOXM1.
A Representative western blot and quantification of FOXM1 and HA-Mad2 in Kras (K1, K2, K3) and Kras/Mad2 (KM1, KM2, KM3) breast tumor cells. **P = 0.0043; Unpaired t-test. (n = 9 K and 9 KM tumors were analyzed and measured). B Quantitative RT-PCR of mouse Foxm1 expression in mammary tumors from K and KM animals (n = 8 K and n = 11 KM tumors). *P = 0.0338; Unpaired t-test. C Representative pictures (left) and relative cell viability (right) of K and KM tumor cells after siFoxm1 for 6 days (n = 6 K and n = 7 KM tumors with siFoxm1). Cell viability was normalized to untreated cells. *P = 0.0313; Unpaired t-test. D DNA content analysis of K and KM breast tumor cells after siFoxm1 for 6 days represented as the percentage of cells in each cell cycle phase. (n = 4 K and n = 6 KM tumors were analyzed). **P = 0.001, *P = 0.0304; Two-way ANOVA. E Percentage of TUNEL positive cells in K and KM tumor cells 6 days after Foxm1 knockdown (KD). (n = 4 K and 4KM tumors were analyzed). ***P = 0.0006; Two-way ANOVA. F Western blots of FOXM1, HA-Mad2, Cleaved-caspase3 (C-Casp3) and gama-H2AX in K and KM tumor cells after RCM-1 treatment or siFoxm1 for 6 days, performed in two biological replicates. ACTIN was used as a loading control. For original and additional wbs see Supplemental Material.
Finally, we overexpressed MAD2 in MCF7 cells for a long time (20 days) and analyzed the effect of inhibiting FOXM1. siFOXM1 led to severe mitotic arrest in MAD2 OE cells (Supplementary Fig. 3B), which further suggests an essential role of FOXM1 in mitotic segregation. We also noticed higher γ-H2AX in both parental and MAD2 OE MCF7 cell lines after FOXM1 inhibition, indicating a role of FOXM1 in regulating DNA damage [34]. Increased C-caspase3 levels in MAD2 OE cells with siFOXM1 illustrated that MAD2-expressing cells still rely on FOXM1 for mitotic exit and survival (Supplementary Fig. 3C). Collectively, these data suggest that FOXM1 is critical for the viability of CIN cells induced by MAD2 overexpression.
FOXM1 overexpression preserves mitotic fidelity in MAD2 overexpressing human breast cancer cell lines
To test whether FOXM1 plays a role in MAD2 tolerance in human cell, we infected MAD2-expressing human breast cancer cell lines with either an empty vector (EV) or with doxycycline-inducible human FOXM1 lentiviral vector (expressing FOXM1b or FOXM1c isoforms). Overexpression of FOXM1 in these cell lines showed similar viability compared to cells infected with the EV six days after induction. However, overexpression of FOXM1 significantly decreased the lethality of MAD2-expressing cells (Fig. 4A) and decreased the percentage of TUNEL-positive MCF7 cells (Supplemental Fig. 4A), suggesting that high FOXM1 levels can rescue MAD2-induced defective cell fitness. Since FOXM1 regulates cell cycle progression as well as chromosome segregation [35], we next sought to understand whether FOXM1 upregulation interferes with MAD2-mediated mitotic arrest and CIN in these cell lines. We performed live-cell imaging of MCF7 and CAL51 cells with or without MAD2 OE and FOXM1 OE. The overexpression of FOXM1 alone did not affect mitotic duration or mitotic errors when compared to EV cells. However, cells overexpressing MAD2 showed a significant decrease in mitotic time and reduced mitotic errors when FOXM1 was overexpressed at the same time (Fig. 4B, C). These results indicate that upregulation of FOXM1 facilitates mitotic exit in MAD2-expressing cells, allowing proper chromosome segregation and consequently, reducing MAD2-induced CIN.
A Cell viability of MCF7 and CAL51 cells after MAD2 or/and FOXM1 overexpression for 6 days. Values were normalized to that of each EV group (n = 3 biological replicates). MCF7: ***P = 0.0002, *P = 0.011; CAL51: *P = 0.0452, *P = 0.0225; Ordinary one-way ANOVA. B Mitotic duration and cell fate of MCF7 overexpressing MAD2 or/and FOXM1 for 3 days. Mitotic duration was considered from nuclear envelope breakdown until anaphase. ****P < 0001; Ordinary one-way ANOVA. Percentage of cell fate was calculated as the incidence of each cell fate/total number of mitotic cells. (EV, 112 cells; MAD2 OE, 128 cells; FOXM1c OE, 147 cells; MAD2 OE/FOXM1c OE, 130 cells). C Mitotic duration and cell fate in CAL51 cells with MAD2 or/and FOXM1 overexpression after 3 days. ****P < 0001; Ordinary one-way ANOVA. Percentage of cell fate was calculated as the incidence of each cell fate/total number of mitotic cells. (EV, 152 cells; MAD2 OE, 127 cells; FOXM1c OE, 123 cells; MAD2 OE/FOXM1c OE, 152 cells). Time lapse microscopy was performed in a minimum of 3 biological replicates. D Schematic showing MCF7 cells injected into nude mice that were fed with doxycycline 3 days before injection (n = 5 mice per group). E Tumor onset recorded as the first day after which a palpable tumor was detected. ***P < 0.001; Ordinary one-way ANOVA. Each dot represents a tumor. F Tumor volumes at day 56 after cell injection on the left. ***P < 0.001, **P = 0.0033, *P = 0.0182; Ordinary one-way ANOVA. Tumor volumes overtime on the right. Unpaired t-test was applied to compare tumor volumes at day 56. Raw data for B,C is included in Supplemental Tables.
MAD2 overexpression suppresses Aurora B activity, resulting in hyperstable microtubule-kinetochore attachments [36]. We observed that AURORA B expression was suppressed during mitosis in MAD2 OE cells, whereas the levels were maintained when FOXM1 and MAD2 were overexpressed in MCF7 cells (Supplementary Fig. 4B). Thus, this result suggests that FOXM1 regulates AURORA B allowing MAD2-overexpressing cells to exit mitosis after the induced cell cycle arrest.
To test if FOXM1 overexpression can rescue the detrimental effects of MAD2 OE in vivo, MCF7 cells with empty vector, MAD2 OE or MAD2 together with FOXM1 OE were injected as xenografts in nude mice (Fig. 4D). In line with our in vitro results, persistent high MAD2 levels resulted in decreased fitness and a delayed tumor onset while the combined overexpression of MAD2 and FOXM1 reverted this effect (Fig. 4E). Ten weeks after injection, 2 out of 5 animals injected with MAD2 OE cells developed tumors while the additional overexpression of FOXM1 allowed for 4 out of 5 animals to grow tumors (Fig. 4E). However, FOXM1 did not affect tumor growth at early (day 40) or late (day 56) time points throughout tumor progression (Fig. 4F). Thus, FOXM1 overexpression was not sufficient to accelerate the growth of MAD2 tumors, but it contributed to the tolerance of MAD2-induced detrimental effects on tumor cells.
To corroborate whether FOXM1 endows cells with tolerance to aneuploidy and CIN during chronic MAD2 OE, we analyzed the outcome of cell division in human breast cell lines 20 days after MAD2 induction. MCF7 cells after a long expression of MAD2 continue to present prolonged arrest in mitosis. While long-term MAD2 upregulation resulted in an average mitotic time of 2.67 h, further FOXM1 overexpression was not able to shorten this time (average 2.23 h) (Supplementary Fig. 4C). We next compared CIN levels between MAD2 OE and MAD2/FOXM1 OE by monitoring mitotic errors. A lower percentage of mitotic errors was observed in MAD2 OE cells with constitutive overexpression of FOXM1 indicating a protective role of FOXM1 against CIN during long-term MAD2 OE. (Supplementary Fig. 4D, E). Interestingly, once cells adapted to high MAD2 levels, no cell death was observed in either MAD2 OE cells alone or in the combination with FOXM1 upregulation and the spectrum of mitotic errors was different from the ones seen after a short exposure to MAD2.
Elevated FOXM1 and aneuploidy are associated with poor prognosis in BRCA patients
Previous data indicated that upregulation of FOXM1 facilitates mitotic exit of MAD2-overexpressing cells in the presence of CIN. We reasoned that aneuploid cancers might also require FOXM1 to compensate for deleterious CIN and hypothesized that cancers that allow propagation of CIN could be associated with a worse survival rate. These results were validated in human breast cell lines and mouse breast tumors, however, to further test this hypothesis, we analyzed TCGA data of BRCA patients (n = 1064) and divided them into two groups (FOXM1_low and FOXM1_high) based on FOXM1 mRNA levels. We then calculated the aneuploidy levels of these samples by determining the mean absolute changes in the copy number segment. Aneuploidy was considered as a deviation from 0, which represents euploidy. FOXM1_low samples showed lower copy number alteration (SCNA) counts while FOXM1_high samples had higher SCNA counts (Fig. 5A). However, no difference in overall survival was observed between these two groups (Fig. 5B). To assess the survival of patients with different aneuploidy levels related to FOXM1 expression, we divided patients into two groups based on SCNA counts (SCNA_low and SCNA_high). Patients with higher SCNA counts presented poor prognoses in FOXM1_high cancers, whereas changes in SCNA had no effect on the prognosis of FOXM1_low cancers (Fig. 5C). Patients with high FOXM1 expression and high SCNA had the worse survival compared to the other groups (Fig. 5D). In summary, these data suggest that survival of BRCA patients is not directly associated with FOXM1 levels. Instead, their outcomes are associated with the upregulation of FOXM1 and increased genomic alterations in the cancers. Thus, FOXM1 might facilitate tolerance of high aneuploid tumors and provide advantages for aneuploid cancer development.
A Mean of copy number changes in FOXM1_low and FOXM1_high BRCA cancer samples from TCGA; n = 1064. ****P < 0.0001; Unpaired t-test. B Survival analysis of FOXM1_low and FOXM1_high BRCA patients. P = 0.0763; Long-rank (Mantel-Cox) test. C Survival analysis of BRCA patients with low and high SCNA. FOXM1_high cancers (left): *P = 0.0025; FOXM1_low cancers (right): P = 0.0893; Long-rank (Mantel-Cox) test. D Survival analysis of BRCA patients with low SCNA compared to those with high SCNA and different levels of FOXM1. SCNA_high, FOXM1_high cancers vs SCNA_high, FOXM1_low cancers: *P = 0.0493; SCNA_high, FOXM1_high cancers vs SCNA_low cancers: ****P < 0.0001; Long-rank (Mantel-Cox) test.
FOXM1 facilitates mitotic exit after nocodazole-induced arrest
We speculated that increased FOXM1 levels antagonize mitotic checkpoint signaling and release cells from mitotic arrest. To test this hypothesis, we forced mitotic checkpoint activation by treating cells with the spindle poison nocodazole (Noco). A high dose of Noco treatment blocked cells in mitosis and in line with our previous results, FOXM1 overexpression released these cells from prolonged mitosis (Fig. 6A). In addition, high levels of FOXM1 partially rescued the mitotic slippage induced after Noco treatment (75% of mitotic slippage in Noco treated cells vs 68% in Noco/FOXM1overexpression), as well as cell death in mitosis (11% vs 2%) (Fig. 6B). SAC effectors including MAD2 and CDC20 are all involved in chromosome segregation [37]. Cyclin B is known to block mitotic exit regardless of SAC activity and is degraded by the CDC20-APC/C complex during mitosis [35]. We confirmed MAD2 and Cyclin B protein levels to be downregulated in Noco-treated FOXM1 OE cells compared to control cells after Noco treatment (Fig. 6C). This suggests that FOXM1 OE compromises SAC signaling and reduces Cyclin B in MCF7 cells, leading to chromosome segregation and exit from mitosis.
A Mitotic duration of MCF7 cells after nocodazole (200 μM) treatment, FOMX1 overexpression, and the combination of both for 24 h. ****P < 0.0001; Ordinary one-way ANOVA. (EV, 88 cells; NOCO, 94 cells; FOXM1 OE, 73 cells; NOCO/FOXM1 OE, 124 cells). B Cell fate of MCF7 cells with nocodazole (200 μM) treatment or/and FOXM1 overexpression. Data are represented as the incidence of each cell fate/total number of mitotic cells in at least 3 independent movies. (EV, 88 cells; NOCO, 94 cells; FOXM1 OE, 73 cells; NOCO/FOXM1 OE, 124 cells). C Western blots showing FOXM1, CDC20, CYLIN B and MAD2 protein levels in MCF7 cells after nocodazole treatment or/and FOXM1 overexpression for 24 h. ACTIN was used as a loading control. Two biological replicates. D Mitotic duration of CAL51 cells after Reversine, Nocodazole or MAD2 OE for 3 days and their combination together with siFOXM1 for 3 days. *P < 0.02, **P = 0.0029, ***P = 0.0002, ****P < 0.0001; Ordinary one-way ANOVA. (CTRL, 69 cells; Rev, 87 cells; Noco, 65 cells; MAD2 OE, 127; Rev/Noco, 139 cells; Rev/MAD2 OE, 78 cells; Rev/Noco/FOXM1 KD, 131 cells; Rev/MAD2 OE/FOXM1 KD, 70 cells). Time lapse microscopy was performed in a minimum of 3 biological replicates. E Percentage of mitotic errors in MCF7 cells (CTRL) and after treatment with DCB. ****P < 0.0001 Unpaired t-test. At least three biological replicates were analyzed. F Cell fate of MCF7 cells (CTRL) or treated with DCB. (CTRL, 112 cells, DCB, 88 cells) three independent movies were analyzed. G Representative western blot of FOXM1, MAD2 and ACTIN in human cell lines after treatment with dihydrocytochalasin B (DCB). 3 biological replicates. H Cell viability of MCF7 and CAL51 with or without DCB treatment and FOXM1 inhibition with siRNA for 3 or 6 days. Cell viability of each cell line was normalized to that of each untreated group (n = 3 independent experiments). ****P < 0001, ***P = 0.0028, *P = 0.0335; Unpaired t-test. For original and additional wbs see Supplemental Material. Raw data for A,B,D,E,F is included in Supplemental Tables.
Next, to test whether FOXM1 is responsible for allowing aneuploid cells to exit mitosis, we challenged CAL51 cells with Reversine, an MPS1 inhibitor that abrogates SAC signaling and promotes premature exit from mitosis [38]. While Reversine treatment in control cells or FOXM1 KD cells did not alter mitotic duration, mitotic arrest induced by MAD2 or Noco- was reverted after Reversine treatment in CAL51 cells (Fig. 6D). Importantly, Reversine was not able to rescue the prolonged mitosis in CAL51 cells when siFOXM1 was applied (Fig. 6D). These data suggest that FOXM1 is crucial to restoring mitotic exit in aneuploid cells with a hyperactivated SAC.
To further verify if FOXM1 is essential for aneuploid cells, we attempted to generate aneuploid cells by treating breast cancer cells with dihydrocytochalasin B (DCB), to block cytokinesis and induce tetraploidy [39]. Consistent with the previous reports [40, 41], an increase of mitotic errors was observed after DCB treatment, suggesting that cells became chromosomally unstable (Fig. 6E, F). MCF7 and CAL51 cell lines exhibited increased FOXM1 protein levels after DCB treatment. In addition, CAL51 cells treated with DCB also had higher MAD2 levels compared to non-treated cells (Fig. 6G). We next inhibited FOXM1 in these unstable cells. Long-term inhibition of FOXM1 impaired proliferation, especially in the DCB-treated cells (Fig. 6H) suggesting that chromosomally unstable cells have increased sensitivity to FOXM1 inhibition.
Discussion
In our current study, we report that depletion of Foxm1 in various cells that overexpress MAD2 (including human breast tumor cells, MEFs, normal mouse mammary epithelial cells and mouse tumor cells), leads to an extension of mitosis and an increase in mitotic errors. Tumor cells with high levels of MAD2 can adapt over time by accumulating FOXM1, which renders them vulnerable to FOXM1 inhibition. Our investigation has revealed that FOXM1 upregulation facilitates chromosome segregation and mitotic exit in human cell lines with high MAD2 levels by suppressing SAC signaling. As a result, these cells are rescued from their defective proliferation. Additionally, we have found that FOXM1 can reverse mitotic arrest and correct mitotic errors induced by other mechanisms such as nocodazole or cytokinesis failure.
Tumors can overcome the mitotic checkpoint induced by Mad2 overexpression and become aneuploid [12, 13]. The increasing number of mitotic errors suggest that cells force mitotic exit and regain cell fitness despite acquiring CIN. Interestingly, high levels of CIN can actually be tumor-suppressive, making high CIN tumors a potential therapeutic opportunity to halt CIN accumulation in unstable cells [5].
MAD2 overexpression in cells is detrimental, but the mechanisms by which cells overcome this effect are not well understood. By analyzing the genetic dependencies in high MAD2 aneuploid cell lines we found that FOXM1 is an essential gene in these cells. FOXM1 can activate various cell cycle-related proteins and is closely associated with CIN, suggesting that FOXM1 might be relevant to tolerate chromosomal instability. Our findings support the idea that high MAD2 CIN cells might require FOXM1 for maintaining proliferative fitness. However, once MAD2 is overexpressed for an extended period of time, chromosome errors persist, and the number of polyploid cells increases, suggesting that cells can escape the mitotic arrest via slippage. Although multiple mechanisms can allow cells to escape from mitotic arrest [15, 42], we found that the level of SAC proteins decreases in MAD2 overexpressing cells with FOXM1 upregulation, providing insights into the role of FOXM1 in the maintenance of chromosomal stability.
Similarly, inhibition of FOXM1 increases the levels of these SAC proteins, strengthening the checkpoint and inducing mitotic arrest and cell death. We conclude that FOXM1 is required for proliferative fitness in high MAD2 cells with CIN. Nevertheless, FOXM1 prevents mitotic cell death in cells with paclitaxel treatment [21], our results cannot exclude the possibility of cell death prevention of FOXM1during MAD2 overexpression.
FOXM1 insufficiency causes centrosome abnormalities and disrupts the spindle formation [43, 44]. Thus, FOXM1 is a valuable target to increase mitotic stress in CIN cells. Consistent with this, we find that short-term FOXM1 inhibition is insufficient to arrest cells in mitosis but leads to lagging chromosomes and chromosome misalignments in human cell lines. As mitotic errors accumulate, several cellular outcomes are expected. First, the mitotic checkpoint is activated leading to mitotic arrest [11]. Second, DNA damage and replication stress block cell cycle entry, leading to quiescence and senescence [45]. Finally, overwhelmingly high CIN levels activate apoptotic pathways and induce cell death [46]. In agreement, our results demonstrate that high levels of MAD2 confer aneuploid cell lines sensitivity to FOXM1 inhibition, reducing their proliferative ability as long-term FOXM1 inhibition induces high levels of mitotic aberrations. Thus, FOXM1 represents a critical target to tumor cells exhibiting CIN or aneuploidy.
Data availability
All data generated and analyzed during the study are available from the corresponding author upon reasonable request.
References
Schvartzman JM, Sotillo R, Benezra R. Mitotic chromosomal instability and cancer: mouse modelling of the human disease. Nat Rev Cancer. 2010;10:102–15.
Bakhoum SF, Landau DA. Chromosomal instability as a driver of tumor heterogeneity and evolution. Cold Spring Harb Perspect Med. 2017;7:a029611.
Ippolito MR, Martis V, Martin S, Tijhuis AE, Hong C, Wardenaar R, et al. Gene copy-number changes and chromosomal instability induced by aneuploidy confer resistance to chemotherapy. Dev Cell. 2021;56:2440–54.e6.
Lukow DA, Sausville EL, Suri P, Chunduri NK, Wieland A, Leu J, et al. Chromosomal instability accelerates the evolution of resistance to anti-cancer therapies. Dev Cell. 2021;56:2427–39.e4.
Sansregret L, Vanhaesebroeck B, Swanton C. Determinants and clinical implications of chromosomal instability in cancer. Nat Rev Clin Oncol. 2018;15:139–50.
Lee AJ, Endesfelder D, Rowan AJ, Walther A, Birkbak NJ, Futreal PA, et al. Chromosomal instability confers intrinsic multidrug resistance. Cancer Res. 2011;71:1858–70.
Bakhoum SF, Ngo B, Laughney AM, Cavallo JA, Murphy CJ, Ly P, et al. Chromosomal instability drives metastasis through a cytosolic DNA response. Nature 2018;553:467–72.
Musacchio A, Hardwick KG. The spindle checkpoint: structural insights into dynamic signalling. Nat Rev Mol Cell Biol. 2002;3:731–41.
Taylor SS, Scott MI, Holland AJ. The spindle checkpoint: a quality control mechanism which ensures accurate chromosome segregation. Chromosome Res. 2004;12:599–616.
Bates M, Furlong F, Gallagher MF, Spillane CD, McCann A, O'Toole S, et al. Too MAD or not MAD enough: The duplicitous role of the spindle assembly checkpoint protein MAD2 in cancer. Cancer Lett. 2020;469:11–21.
Yuan B, Xu Y, Woo JH, Wang Y, Bae YK, Yoon DS, et al. Increased expression of mitotic checkpoint genes in breast cancer cells with chromosomal instability. Clin Cancer Res. 2006;12:405–10.
Sotillo R, Hernando E, Diaz-Rodriguez E, Teruya-Feldstein J, Cordon-Cardo C, Lowe SW, et al. Mad2 overexpression promotes aneuploidy and tumorigenesis in mice. Cancer Cell. 2007;11:9–23.
Rowald K, Mantovan M, Passos J, Buccitelli C, Mardin BR, Korbel JO, et al. Negative selection and chromosome instability induced by Mad2 overexpression delay breast cancer but facilitate oncogene-independent outgrowth. Cell Rep. 2016;15:2679–91.
Weaver BA. How Taxol/paclitaxel kills cancer cells. Mol Biol Cell. 2014;25:2677–81.
Tischer J, Gergely F. Anti-mitotic therapies in cancer. J Cell Biol. 2019;218:10–1.
Zasadil LM, Andersen KA, Yeum D, Rocque GB, Wilke LG, Tevaarwerk AJ, et al. Cytotoxicity of paclitaxel in breast cancer is due to chromosome missegregation on multipolar spindles. Sci Transl Med. 2014;6:229ra43.
Fischer M, Muller GA. Cell cycle transcription control: DREAM/MuvB and RB-E2F complexes. Crit Rev Biochem Mol Biol. 2017;52:638–62.
Carr JR, Park HJ, Wang Z, Kiefer MM, Raychaudhuri P. FoxM1 mediates resistance to herceptin and paclitaxel. Cancer Res. 2010;70:5054–63.
Arceci A, Bonacci T, Wang X, Stewart K, Damrauer JS, Hoadley KA, et al. FOXM1 deubiquitination by USP21 regulates cell cycle progression and paclitaxel sensitivity in basal-like breast cancer. Cell Rep. 2019;26:3076–86.e6.
Khongkow P, Gomes AR, Gong C, Man EP, Tsang JW, Zhao F, et al. Paclitaxel targets FOXM1 to regulate KIF20A in mitotic catastrophe and breast cancer paclitaxel resistance. Oncogene 2016;35:990–1002.
Vaz S, Ferreira FJ, Macedo JC, Leor G, Ben-David U, Bessa J, et al. FOXM1 repression increases mitotic death upon antimitotic chemotherapy through BMF upregulation. Cell Death Dis. 2021;12:542.
Macedo JC, Vaz S, Bakker B, Ribeiro R, Bakker PL, Escandell JM, et al. FoxM1 repression during human aging leads to mitotic decline and aneuploidy-driven full senescence. Nat Commun. 2018;9:2834.
Corsello SM, Nagari RT, Spangler RD, Rossen J, Kocak M, Bryan JG, et al. Discovering the anti-cancer potential of non-oncology drugs by systematic viability profiling. Nat Cancer. 2020;1:235–48.
Buccitelli C, Salgueiro L, Rowald K, Sotillo R, Mardin BR, Korbel JO. Pan-cancer analysis distinguishes transcriptional changes of aneuploidy from proliferation. Genome Res. 2017;27:501–11.
de Carcer G, Venkateswaran SV, Salgueiro L, El Bakkali A, Somogyi K, Rowald K, et al. Plk1 overexpression induces chromosomal instability and suppresses tumor development. Nat Commun. 2018;9:3012.
Cohen-Sharir Y, McFarland JM, Abdusamad M, Marquis C, Bernhard SV, Kazachkova M, et al. Aneuploidy renders cancer cells vulnerable to mitotic checkpoint inhibition. Nature 2021;590:486–91.
Gartel AL. FOXM1 in cancer: interactions and vulnerabilities. Cancer Res. 2017;77:3135–9.
Weiler SME, Pinna F, Wolf T, Lutz T, Geldiyev A, Sticht C, et al. Induction of chromosome instability by activation of Yes-associated protein and forkhead box M1 in liver cancer. Gastroenterology 2017;152:2037–51.e22.
Bhat UG, Halasi M, Gartel AL. FoxM1 is a general target for proteasome inhibitors. PLoS One. 2009;4:e6593.
Gartel AL. Thiostrepton, proteasome inhibitors and FOXM1. Cell Cycle. 2011;10:4341–2.
Jeffery JM, Kalimutho M, Johansson P, Cardenas DG, Kumar R, Khanna KK. FBXO31 protects against genomic instability by capping FOXM1 levels at the G2/M transition. Oncogene 2017;36:1012–22.
Barger CJ, Chee L, Albahrani M, Munoz-Trujillo C, Boghean L, Branick C, et al. Co-regulation and function of FOXM1/RHNO1 bidirectional genes in cancer. Elife. 2021;10:e55070.
Sun L, Ren X, Wang IC, Pradhan A, Zhang Y, Flood HM, et al. The FOXM1 inhibitor RCM-1 suppresses goblet cell metaplasia and prevents IL-13 and STAT6 signaling in allergen-exposed mice. Sci Signal. 2017;10:eaai8583.
Zona S, Bella L, Burton MJ, Nestal de Moraes G, Lam EW. FOXM1: an emerging master regulator of DNA damage response and genotoxic agent resistance. Biochim Biophys Acta. 2014;1839:1316–22.
Costa RH. FoxM1 dances with mitosis. Nat Cell Biol. 2005;7:108–10.
Kabeche L, Compton DA. Checkpoint-independent stabilization of kinetochore-microtubule attachments by Mad2 in human cells. Curr Biol. 2012;22:638–44.
Matsumura S, Toyoshima F, Nishida E. Polo-like kinase 1 facilitates chromosome alignment during prometaphase through BubR1. J Biol Chem. 2007;282:15217–27.
Maia ARR, Linder S, Song JY, Vaarting C, Boon U, Pritchard CEJ, et al. Mps1 inhibitors synergise with low doses of taxanes in promoting tumour cell death by enhancement of errors in cell division. Br J Cancer. 2018;118:1586–95.
Fujiwara T, Bandi M, Nitta M, Ivanova EV, Bronson RT, Pellman D. Cytokinesis failure generating tetraploids promotes tumorigenesis in p53-null cells. Nature 2005;437:1043–7.
Kuznetsova AY, Seget K, Moeller GK, de Pagter MS, de Roos JA, Durrbaum M, et al. Chromosomal instability, tolerance of mitotic errors and multidrug resistance are promoted by tetraploidization in human cells. Cell Cycle. 2015;14:2810–20.
Storchova Z, Kuffer C. The consequences of tetraploidy and aneuploidy. J Cell Sci. 2008;121:3859–66.
Cheng B, Crasta K. Consequences of mitotic slippage for antimicrotubule drug therapy. Endocr Relat Cancer. 2017;24:T97–T106.
Wonsey DR, Follettie MT. Loss of the forkhead transcription factor FoxM1 causes centrosome amplification and mitotic catastrophe. Cancer Res. 2005;65:5181–9.
Limzerwala JF, Jeganathan KB, Kloeber JA, Davies BA, Zhang C, Sturmlechner I, et al. FoxM1 insufficiency hyperactivates Ect2-RhoA-mDia1 signaling to drive cancer. Nat Cancer. 2020;1:1010–24.
Matthews HK, Bertoli C, de Bruin RAM. Cell cycle control in cancer. Nat Rev Mol Cell Biol. 2022;23:74–88.
Zhivotovsky B, Kroemer G. Apoptosis and genomic instability. Nat Rev Mol Cell Biol. 2004;5:752–62.
Acknowledgements
We thank the DKFZ central animal laboratory for animal husbandry; DKFZ core facilities of Light Microscopy and Flow Cytometry for technical support. Schemes were generated with BioRender.com. This study was supported by the Helmholtz Foundation and China Scholarship Council (CSC) to FP. We are grateful to Alberto Diaz and Sridhar Kandala for suggestions on this study and to Jimmy Monta, Simone Kraut, Justin Guddat for assistance with laboratory work.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
FP, SC and YC performed the experiments. MR and AAdIV generated CAL51-rtTA cell line. KS generated all plasmids used in this work. FP and RS designed the experiments, analyzed the data and wrote the manuscript. RS supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All animal experiments were performed under approval from protocol numbers G231/15 and G18/21 from Regierungspräsidium of Karlsruhe, Germany.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Massimiliano Agostini
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pan, F., Chocarro, S., Ramos, M. et al. FOXM1 is critical for the fitness recovery of chromosomally unstable cells. Cell Death Dis 14, 430 (2023). https://doi.org/10.1038/s41419-023-05946-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-023-05946-2
- Springer Nature Limited