Abstract
Cancer immunotherapy has rapidly transformed cancer treatment, yet resistance remains a significant hurdle, limiting its efficacy in many patients. Circular RNAs (circRNAs), a novel class of non-coding RNAs, have emerged as pivotal regulators of gene expression and cellular processes. Increasing evidence indicates their involvement in modulating resistance to cancer immunotherapy. Notably, certain circRNAs function as miRNA sponges or interact with proteins, influencing the expression of immune-related genes, including crucial immune checkpoint molecules. This, in turn, shapes the tumor microenvironment and significantly impacts the response to immunotherapy. In this comprehensive review, we explore the evolving role of circRNAs in orchestrating resistance to cancer immunotherapy, with a specific focus on their mechanisms in influencing immune checkpoint gene expression. Additionally, we underscore the potential of circRNAs as promising therapeutic targets to augment the effectiveness of cancer immunotherapy. Understanding the role of circRNAs in cancer immunotherapy resistance could contribute to the development of new therapeutic strategies to overcome resistance and improve patient outcomes.
Similar content being viewed by others
Facts
-
CircRNAs play a significant role in modulating immune-related pathways and the tumor microenvironment in cancer immunotherapy resistance.
-
Dysregulation of circRNAs can profoundly impact tumor progression, making therapeutic strategies targeting circRNA highly promising for clinical applications.
-
CircRNA research is still in its early stages, and clinical applications, particularly in terms of personalized treatment, are yet to be fully realized.
Open questions
-
What are the precise molecular mechanisms through which circRNAs modulate immune checkpoints and tumor microenvironment factors in different cancer types?
-
What are the implications of circRNA dysregulation in specific pathological stages of cancer progression for immunotherapy efficacy?
-
How can advanced methods for the design, synthesis, purification, and delivery of circRNA-based therapies be developed to enhance their clinical translation?
Introduction
Cancer is a significant global health challenge, annually claiming millions of lives [1]. The emergence of immunotherapy has ushered in a transformative era in oncology treatment, complementing traditional methods like surgery, radiotherapy, chemotherapy, and targeted therapy. This paradigm shift leverages the innate capability of the immune system to identify and eradicate cancer cells, offering a ray of hope [2]. However, despite notable successes in treating various cancers types, such as lung cancer, melanoma, and hepatocellular carcinoma, resistance remains a significant challenge [3]. As the most promising immunotherapy, the efficacy of anti-programmed death-1/programmed death ligand 1 (anti-PD-1/L1) therapies is confined to a minority of patients, with sustained positive responses observed in only 10–30% of cases [4]. Thus, a comprehensive understanding of the intricate mechanisms underlying immunotherapy resistance is imperative for devising effective treatment strategies.
Recent studies have spotlighted the pivotal role of circular RNAs (circRNAs) in orchestrating resistance to cancer immunotherapy [5]. CircRNAs, a unique class of endogenous non-coding RNAs forming covalently closed circular structures, defy the linear structure of other RNA molecules [6]. Initially deemed splicing artifacts, recent studies have unveiled their critical role in regulating gene expression by acting as miRNA sponges, RNA-binding protein sequestering agents, or transcriptional regulators. Dysregulation of circRNAs has been noted in various cancer types, with several studies illustrating their involvement in cancer progression and metastasis [7, 8]. Emerging evidence suggests that circRNAs also play a crucial role in regulating resistance to cancer immunotherapy. They can modulate various aspects of tumor-immune interactions, including immune cell infiltration, immune checkpoint regulation, and cytokine signaling pathways, thereby influencing the response to immunotherapy.
In this review, we provide an overview of the emerging role of circRNAs in regulating resistance to cancer immunotherapy. We discuss the current understanding of the mechanisms by which circRNAs modulate tumor-immune interactions and the impact of these interactions on the response to immunotherapy. Additionally, we highlight the potential of circRNAs as biomarkers for predicting response to immunotherapy and as therapeutic targets for overcoming resistance. Overall, this review offers a comprehensive understanding of the role of circRNAs in regulating resistance to cancer immunotherapy and their potential as therapeutic targets and biomarkers.
CircRNAs: biogenesis and functions (Fig. 1)
Overview of circRNA biogenesis and features
CircRNAs constitute a subtype of non-coding RNA characterized by a covalently closed-loop structure. In 1976, Sanger et al. first identified circular RNA molecules in nature through electron microscopy [9]. Subsequent decades revealed the presence of circRNAs in yeast mitochondria [10], Hepatitis delta virus [11], and genes capable of producing circRNA, such as the human dystrophin gene [12] and the mouse sex-determining region Y (Sry gene) [13]. Formed through back-splicing events, circRNAs involve the joining of the 3’ end of a downstream exon with the 5’ end of an upstream exon, resulting in a circular molecule [14, 15]. Initially perceived as rare splicing by-products, recent studies demonstrate their abundance and widespread expression across various tissues and organisms [16].
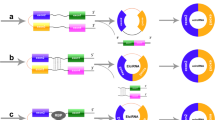
A Biogenesis of circRNAs. (a) conventional splicing. The conventional splicing process involves the formation of intronic lariats, escaping debranching and ligation, resulting in the creation of intronic circRNAs (CiRNAs). (b) exon skipping. (c) direct back-splicing. (d) nuclear export. The nuclear export of short (<400-nt) and long (>1200-nt) circRNAs is mediated by DDX39A and DDX39B, respectively. Another crucial regulator of nuclear export is the conserved Exportin 4 (XPO4). Additionally, m6A modification can facilitate the nuclear export of circRNAs. B Functions of circRNAs. (a) Transcriptional regulation: circRNAs can influence parental gene transcription by forming a three-stranded R-loop structure with its production site or by interacting with transcriptional complexes, such as U1 snRNP. (b) miRNA sponges: by competitively binding to miRNAs, circRNAs up-regulate downstream target mRNA and corresponding proteins, ultimately impacting cellular physiological processes. (c) RNA-binding protein decoys: acting as sponges, scaffolds, and recruiters, circRNAs interact with RNA-binding proteins, regulating various intracellular physiological processes. (d) Translating proteins: circRNAs exhibit two cap-independent translation modes: IRES-mediated and m6A-mediated. In the IRES-mediated process, IRES binds to the initiation factor eIF4G2, assembles with eIF4A and eIF4B, recruits the 40 s ribosome subunit, and forms a 43 s initiation complex that initiates translation upon encountering the initiation codon ATG. Another translation type is mediated by the m6A motif in circRNA, where m6A is recognized by the m6A reader YTH domain family protein 3 (YTHDF3), recruiting initiation factors and ribosome subunits to form translation-initiation complexes inducing translation.
The biogenesis of circRNAs relies on non-canonical spliceosome machinery, regulated by trans-acting proteins and cis-regulatory elements. The interplay between back splicing and canonical splicing, mediated by splicing factors, determines the equilibrium between the two processes [17]. Based on the sequence order of back splicing and canonical splicing, circRNA biosynthesis is categorized into two models: (1) Lariat-driven circularization (or the exon-skipping model) and direct back-splicing. In the Lariat-driven circularization model, canonical splicing precedes, producing a linear RNA with skipped exons and a lariat precursor. This precursor, containing introns and exons, undergoes back-splicing to generate a circRNA. (2) In the direct back-splicing model, back splicing occurs first, directly producing a circRNA and a linear exon-intron(s)-exon intermediate. This intermediate can undergo further splicing, resulting in a linear RNA with skipped exons [5, 18, 19].
CircRNAs are categorized into three main types based on their origin: exonic circRNAs (ecircRNAs), circular intronic RNAs (ciRNAs), and exon-intron circRNAs (EIciRNAs). EcircRNAs comprise one or more exons and are primarily located in the cytoplasm, while ciRNAs and EIciRNAs are situated in the nucleus. EcircRNAs predominate among the total circRNAs [20,21,22]. CircRNAs exhibit distinctive features, such as a covalently closed-loop structure conferring stability and resistance to Rnase degradation [23]. The longer half-lives of circRNAs (18.8–23.7 h) compared to their linear counterparts (4.0–7.4 h) contribute to their enhanced stability [24]. Additionally, circRNAs display tissue-specific and developmental stage-specific expression, with high levels in specific cell types, such as nerve cells in the brain. Many circRNAs are evolutionarily conserved, underscoring their potential functional roles [25, 26]. Furthermore, circRNAs vary in length, originate from different exons or introns, and exhibit diverse sequences, contributing to a wide range of potential functions. Serving as miRNA sponges, RNA-binding protein (RBP) decoys, or splicing modulators, circRNAs emerge as a diverse class of non-coding RNAs with crucial regulatory roles in eukaryotic gene expression.
Functions of circRNAs
CircRNAs have emerged as pivotal regulators of gene expression in eukaryotes, demonstrating diverse functions across various aspects:
MiRNA sponges: circRNAs function as competing endogenous RNAs (ceRNAs) that sequester miRNAs, thereby preventing them from targeting their mRNA counterparts. This sequestration leads to increased expression of mRNA targets, impacting cellular processes such as proliferation, differentiation, and apoptosis. MiRNA sponge circRNAs, usually exhibiting high expression, contain numerous microRNA response elements (MREs) [5, 6, 20]. A classic example is ciRS-7, which harbors over 70 selectively conserved miRNA target sites, inhibiting miR-7 activity and resulting in elevated levels of miR-7 targets [27]. Studies indicate that circRNAs act as miRNA sponges to either inhibit or promote tumor growth. For example, circCD44 directly binds to miR-502-5p, promoting the proliferation, migration, and invasion of triple-negative breast cancer [28]. Another instance is circMTO1, which inhibits liver cancer by sponging miR-9, subsequently up-regulating the expression of the tumor suppressor p21 [29]. Growing evidence supports the notion that circRNA-miRNA interactions are a universal regulatory mechanism.
RNA-binding protein decoys: circRNAs can act as decoys for RNA-binding proteins (RBPs), preventing their binding to target mRNAs. This interaction alters the stability or translation of mRNA targets, influencing cellular processes. CircRNAs interact with RBPs as protein sponges, scaffolds, and recruiters [6]. The initial example of circRNAs acting as protein sponges is circMbl, where the splicing factor muscleblind (MBL) and its flanking introns possess conserved MBL binding sites. Elevated MBL concentrations promote circMbl generation while reducing linear mRNA MBL production. Highly expressed circMbl also adsorbs MBL, impeding its neural function [6, 17, 20]. As protein scaffolds, circRNAs facilitate interactions between different proteins. For instance, circ-Foxo3 in breast cancer cells binds to p53 and E3 ubiquitin ligase mouse doubleminute 2 homolog (MDM2), forming a ternary complex and promoting p53 ubiquitination and degradation by MDM2 [30]. Additionally, circRNAs can recruit specific proteins to particular sites. For example, circRHOT1 induces liver cancer development by recruiting TIP60 (also known as histone acetyltransferase KAT5) to the promoter of nuclear receptor subfamily 2 group F member 6 (NR2F6), thereby inducing NR2F6 expression [6, 31]. In summary, circRNA-protein interactions can influence target gene expression, thereby affecting human disease development.
Transcriptional regulation: circRNAs can regulate gene expression by interacting with transcription factors or chromatin modifiers to impact target gene transcription. On one hand, circRNAs can influence transcription by forming a three-stranded R-loop structure with their production site. For instance, Ci-ankrd52 binds to the parental locus ANKRD52, forming an R-loop, activating RNase H1-mediated ci-ankrd52 digestion, disrupting the R-loop, and promoting transcriptional extension [32]. On the other hand, circRNAs can activate transcription factors (TFs). For instance, circPOK (encoded by the Zbtb7a gene) co-activates the ILF2/3 complex, binding to the proximal promoter of II6 [33]. Additionally, EIciRNAs enhance the transcription of parental genes by interacting with U1 small nuclear ribonucleoprotein (U1 snRNP), such as circEIF3J and circPAIP21922. Knocking down circEIF3J and circPAIP2 decreases the transcription levels of EIF3J and PAIP2, respectively [34].
Translating proteins: while circRNAs are generally considered non-coding due to the lack of a 5’ cap and 3’ poly(A) tail [35], existing studies suggest that circRNAs can be translated through cap-independent mechanisms mediated by internal ribosome entry sites (IRES) [36], as well as by N6-methyladenosine (m6A) modification [37]. In the IRES-mediated process, IRES binds to the initiation factor eIF4G2, assembling with eIF4A and eIF4B to recruit 40 s ribosomal subunits and form the 43 s initiation complex, initiating translation upon encountering the initiation codon ATG [38]. Another non-cap translation type is predominantly mediated by m6A located in the 5’-untranslated region. This m6A is recognized by the m6A reader YTH domain family protein 3, recruiting initiation factors and ribosomal subunits to form translation initiation complexes, inducing translation [39]. CircRNAs-mediated protein translation can regulate tumor growth. For instance, the study by Yang Y et al. demonstrates that circFBXW7 encodes the functional protein FBXW7-185aa, which competes with FBXW7α for deubiquitinase USP28, resulting in free FBXW7α-induced ubiquitination degradation of c-Myc and inhibiting glioblastoma development in the brain [40]. It is noteworthy that circRNAs translate proteins to maintain cell survival under stress conditions such as hypoxia, heat shock, or viral infection [41]. However, under non-stress conditions, circRNAs may not undergo translation due to the predominance of cap-dependent translation [42].
Based on these functions, circRNAs are implicated in various diseases, including cancer, cardiovascular disease, and neurological disorders. Recently, circRNAs have emerged as potential targets for cancer therapy, given their involvement in cellular processes related to cancer progression, such as cell proliferation, invasion, metastasis, and drug resistance. For instance, circHIPK3 is upregulated in various cancer types and promotes resistance to chemotherapy and radiotherapy by sponging multiple miRNAs [43, 44]. Targeting this circRNA could sensitize cancer cells to these treatments. CircVAPA is upregulated in non-small cell lung cancer, acting as a sponge for miR-377-3p and miR-494-3p, thereby accelerating tumor proliferation and differentiation through the IGF1R/AKT axis. Knockdown of circVAPA significantly enhances the effect of IGF1R kinase inhibitor (BMS-536924) in inhibiting tumor growth [45]. These studies underscore the potential of targeting circRNAs for cancer therapy. Nevertheless, further research is needed to comprehensively understand circRNA regulation mechanisms in cancer and develop effective circRNA-targeting therapies.
Cancer immunotherapy resistance
Cancer immunotherapy (Fig. 2)
The immune system plays a critical role in protecting the body against cancer. T-cells are one of the key components of the immune system, as they are responsible for recognizing and attacking abnormal or infected cells, including cancer cells. However, cancer cells can often evade detection by the immune system through two main mechanisms: either by producing signals that suppress the immune response or by presenting themselves as “self” to the immune system to avoid being recognized as foreign. Cancer immunotherapy utilizes the body’s immune system to fight cancer. It works by stimulating the immune system to recognize and attack cancer cells. There are several different types of cancer immunotherapy, including immune checkpoint inhibitors (ICIs), adoptive cell therapy (ACT), cancer vaccines, oncolytic virus therapy, and immune system modulators.
Immune checkpoint inhibitors bind to immunosuppressive proteins on the cell surface, restoring the antitumor function of T cells. Adoptive cell therapy: CAR-immune cells are engineered to specifically target antigens on the surface of tumor cells, enhancing the immune system’s ability to combat cancer. Therapeutic cancer vaccines: Therapeutic cancer vaccines deliver antigens to Antigen-Presenting Cells (APCs), activating and inducing cytotoxic T lymphocyte responses to fight against cancer cells. Oncolytic virus therapy: Oncolytic viruses cause oncolysis, releasing viral offspring, Pathogen-Associated Molecular Patterns (PAMPs), Damage-Associated Molecular Patterns (DAMPs), and Tumor-Associated Antigens (TAAs). This stimulates the immune system to target and destroy cancer cells. Small molecule immunomodulators: Illustrated by icaritin soft capsules, icaritin targets the MyD88/IL-6/JAK/STAT3 signaling pathway. This results in reduced cytokine production (e.g., TNF-α and IL-6) and a decrease in the expression of immune checkpoints (e.g., PD-L1), contributing to enhanced antitumor immunity.
Immune checkpoints block (ICB) therapy
ICIs can block specific proteins on immune cells that prevent them from attacking cancer cells. Immune checkpoints are cell surface receptors expressed by immune cells, a group of co-stimulatory signals, including stimulatory and inhibitory molecules, that regulate the activation and effector functions of immune cells, such as cytotoxic T lymphocyte-associated antigen-4 (CTLA-4), programmed death-1/programmed death ligand 1 (PD-1/PD-L1), and T cell immunoglobulin and mucin-3 (Tim-3) [46]. Under normal circumstances, immune checkpoints (ICPs) maintain self-tolerance and immune homeostasis, but when malignant tumors occur, ICPs are occupied, such as PD-L1 on tumor cells binding to PD-1 on T cells and thus preventing effector T cells from functioning, allowing tumor cells to achieve immune escape and promoting tumor growth [47]. Blockade of immune checkpoints can reactivate tumor immunity. Currently, the main ICIs commonly used in clinical practice target CTLA4, PD-1 and PD-L1, such as ipilimumab, nivolumab, and durvalumab [48]. For the remarkable results of ICIs in anti-tumor, James P. Allison and Tasuku Honjo, two scientists working on CTLA-4 and PD-1, respectively, were also awarded the 2018 Nobel Prize in Medicine and Physiology [49]. At present, ICIs are also approved for a growing number of indications, including metastatic melanoma, renal cell carcinoma, advanced non-small cell lung cancer, bladder cancer, lymphoma, and more [50]. The simultaneous use of different types of ICIs can also exert synergistic anti-tumor effects. For example, anti-PD-1/L1 therapy combined with anti-CTLA4 has become the standard for the treatment of melanoma, and the combination of nivolumab and ipilimumab in patients with advanced melanoma can improve the survival rate of patients of 5 years [51].
Adoptive cell therapy (ACT)
ACT involves removing immune cells from a patient, growing them in a lab, and then infusing them back into the patient to attack cancer cells [52]. ACT includes tumor infiltrating lymphocytes (TILs) therapy, endogenous T cells therapy, T cell receptor therapy, and chimeric antigen receptor (CAR) T cell therapy [53]. One of the most promising therapeutic methods is CAR-T therapy, which genetically modifies a patient’s T cells to target specific antigens on tumor cells. CAR-T cells do not rely on major histocompatibility complex (MHC) molecules to recognize antigens, which overcomes the problem of missing the expression of MHC class I molecules in tumor cells [54]. This method has shown good efficacy in patients with chemotherapy-refractory tumors of hematologic malignancies such as leukemia, myeloma, and non-Hodgkin B-cell lymphoma [55]. At present, in addition to CAR-T, CAR-natural killer cells (CAR-NK), and CAR-macrophages (CAR-M) have been introduced as a supplement or alternative to CAR-T cell therapy [52].
Therapeutic cancer vaccines
Therapeutic tumor vaccines are a safe way to boost T-cell responses and produce therapeutic effects at all stages of disease in tumor patients [56]. Tumor vaccines consist of tumor antigen, formulation, immune adjuvant, and delivery vehicle [57]. There are four main vaccine types: tumor whole-cell vaccines, genetically engineered vaccines, protein peptide vaccines, and dendritic cell vaccines [58]. Successful therapeutic cancer vaccines require high-quality antigens to be delivered to antigen-presenting cells and activated and induce cytotoxic T lymphocyte responses, maintain immune cell infiltration in tumor microenvironment (TME), and ultimately induce tumor regression, build durable anti-tumor memories, and avoid non-specific or adverse reactions [59]. Ideally, vaccines would target tumor-specific antigens to avoid an autoimmune response, but many vaccines target tumor-associated antigens (TAAs), which are often recognized by the immune system as ‘self’ [56]. Although flawed, this does not negate the therapeutic effectiveness of tumor vaccines. In a Phase I dose-escalation trial (NCT02410733), the mRNA lipid complex vaccine (BNT111) exhibited promising safety and efficacy in advanced melanoma patients, with an immune response against one or more tumor-associated antigens detected in over 75% of participants. Although therapeutic mRNA vaccines are not yet standard, their combination with ICIs in clinical trials has shown satisfactory results [60].
Oncolytic virus therapy
Oncolytic virus (OVs) immunotherapy exploits viruses to selectively infect and replicate within tumor cells, leading to tumor cell lysis, the release of tumor antigens, and the activation of the immune system through additional damage-associated molecular patterns and viral pathogen-associated molecular patterns (PAMPs) [61]. OVs, either naturally occurring or genetically modified, selectively eliminate tumor cells and induce host anti-tumor immunity [62]. Common OVs include newcastle disease viruses, herpes viruses, coxsackievirus, measles viruses, adenoviruses, polioviruses, poxviruses, and reoviruses [63]. Genetically engineered OVs enhance tumor selectivity, promote replication ability, and increase immunogenicity [64]. In clinical settings, OVs are often combined with other treatment methods to enhance treatment sensitivity. For instance, in a phase II clinical trial, the combination of talimogene laherparepvec and ipilimumab outperformed ipilimumab alone, increasing the objective response rate from 18 to 39% in patients with advanced melanoma [65].
Small molecule immunomodulators therapy
Most current immunotherapies rely on antibodies, which, while exhibiting specificity and efficacy in pharmacodynamics, face limitations in pharmacokinetics, such as extended half-life and poor tumor tissue permeability. To overcome these challenges, researchers have developed small molecule-based immunotherapy methods. The combined use of small molecule immunomodulators and antibody drugs produces synergistic effects, representing a complementary mode of tumor therapy [66]. The therapeutic strategy of small molecule immunomodulators involves targeting intracellular and extracellular molecules to impact various intracellular signaling pathways, thereby inhibiting tumor growth. Examples include small molecule PD-L1 inhibitors, PD-L1-targeting PROTAC degraders, chemokine receptor antagonists, RORγt agonists, small molecule TGF-β inhibitors, small molecule STING agonists, etc. Although many of these are in clinical trials, no small molecule-based cancer immunotherapies have yet gained approval from the US Food and Drug Administration (FDA) [66]. However, a Chinese herbal monomer preparation named icaritin soft capsules has been approved as an immunomodulatory agent for treating advanced hepatocellular carcinoma (HCC) in China [67]. Icaritin interacts with the MyD88/IкB kinase α protein complex, suppresses the IL-6/JAK/STAT3 signaling pathway, reduces cytokine generation, and decreases the expression of immune checkpoints. Furthermore, icaritin inhibits the bioactivity of myeloid-derived suppressor cells (MDSCs) by down-regulating tumor-associated splenic extramedullary hematopoiesis [67].
In instances where a single immunotherapy approach proves less effective, a combination of therapies, such as combining immunotherapy with targeted therapy, radiotherapy, or chemotherapy, can be chosen. Detailed combination strategies that maximize the benefits of immunotherapy are reviewed by Zhu et al. and Yap et al. [68, 69]. Ongoing research is directed toward enhancing the effectiveness and safety of immunotherapy and identifying patients most likely to benefit from this form of treatment [70].
Mechanisms underlying resistance to cancer immunotherapy (Fig. 3)
Resistance to cancer immunotherapy refers to the failure of a patient’s immune system to effectively target and eliminate cancer cells, despite treatment with immunotherapeutic agents. The mechanisms underlying resistance are complex and multifaceted, stemming from various factors. Here, we will introduce several resistance mechanisms prevalent in various types of tumor immunotherapy.
T cells, pivotal in recognizing and attacking tumor cells through TCR-mediated MHC-binding peptide antigens (center), encounter resistance mechanisms originating from intrinsic (right) and extrinsic factors (left). Internal factors, primarily arising from defective antigen presentation, antigen deletion, and signaling pathway alterations due to gene mutations, collectively result in compromised T-cell recognition and response against tumor cells. External factors encompass T-cell dysfunction, up-regulation of inhibitory immune checkpoints, and the impact of the immunosuppressive Tumor Microenvironment (TME) and host factors. T-cell dysfunction and heightened inhibitory immune checkpoints contribute to immune escape by tumor cells. The immunosuppressive TME, characterized by nutrient deficiency, hypoxia, acidity, and a plethora of immunosuppressive cells, fosters an environment detrimental to antitumor immune responses. The acidic microenvironment, emanating from lactic acid release by tumor cells, inhibits the cytotoxicity and proliferation of CD8+ T cells. Moreover, lactate and chemokines (e.g., CCL5, CCL7, CXCL12) secreted by tumor cells orchestrate the recruitment and induction of immunosuppressive cells like Treg cells, tumor-associated macrophages, and myeloid-derived suppressor cells into the TME. These recruited cells further secrete inhibitory cytokines (e.g., TGF-β, IL-10, IL-35), hindering the functionality of T cells. Host factors predominantly involve patient-specific elements such as gender, age, weight, and gastrointestinal flora, all contributing to the complex landscape of tumor immunotherapy resistance.
Tumor-intrinsic factors of immunotherapy resistance
Lack of tumor antigen and reduced immunogenicity
tumor immunogenicity, correlated with T cell recognition, hinges on the tumor’s ability to produce neoantigens—an integral factor determining immunotherapy response [71]. The loss of tumor antigens significantly impedes T-cell recognition and functionality, resulting in immunotherapy failure [72]. High immunogenic tumors, such as melanoma and kidney cancer, exhibit better immunotherapy efficacy, while tumors with low immunogenicity, like pancreatic and prostate cancer, are less responsive to ICI treatment [73]. Microsatellite instability (MSI) in tumors with mismatch repair deficiency (MMR) can lead to high tumor mutational burden (TMB), enhancing immunogenicity and ICI therapy response [72, 74]. MSI and TMB serve as biomarkers predicting immunotherapy efficacy [75].
Deficiency in antigen presentation
the antigen peptide-MHC I complex activation of CD8+ T cells is crucial for anti-tumor efficacy [76]. Mutations in genes involved in antigen processing and presentation, including MHC molecules, beta 2 microglobulin (B2M), large multifunctional protease (LMP), and transporter associated with antigen processing (TAP), contribute to ICI resistance [73]. Notably, B2M gene mutations, particularly homozygous truncation, impede MHC Class I molecule expression, affecting antigen presentation and ICI response [77]. Downregulation or loss of B2M expression has been observed in resistant cases, impacting MHC I or HLA I expression [78, 79]. For instance, CD19 deletion in B-cell acute lymphoblastic leukemia renders CART19 treatment ineffective [80].
Alterations in signaling pathways
various signaling pathway alterations, such as enhanced PI3K/AKT signaling and loss of interferon-gamma (IFN-γ) signaling, contribute to immunotherapy resistance. PTEN loss in melanoma activates PI3K/AKT, suppressing T cell infiltration and autophagy, leading to resistance [81]. IFN-γ, vital for anti-tumor effects, loses effectiveness when tumors acquire JAK1/2 loss-of-function mutations, hindering IFN-γ-induced PD-L1 expression and MHC-I molecules [79, 82, 83]. WNT/β-catenin and MAPK signaling also associate with immunotherapy resistance [84].
Tumor-extrinsic factors of immunotherapy resistance: tumor microenvironment
T cell dysfunction and upregulation of suppressive immune checkpoint expression
Prolonged antigenic stimulation induces abnormal T cell function, marked by reduced proliferative capacity, diminished effector function, and increased inhibitory receptor expression [85]. Disruptions in T cell immune function, including antigen recognition, activation, differentiation, and chemotaxis, may result in ineffectiveness of anti-PD therapy [86]. Studies have also shown that blockade of a single ICP leads to compensatory upregulation of other ICPs and affects the efficacy of drug therapy [87]. For example, in the anti-PD-1 treatment of head and neck tumors, TIL compensatoryly upregulates Tim-3 in a PI3K/Akt-dependent manner. Tim-3 inhibits T cell activation by inhibiting the phosphorylation of Akt/S6, thereby making anti-PD-1 therapy resistant [88]. It has been reported that the combination of anti-PD-1 and anti-Tim-3 will produce better therapeutic effects [89]. Upregulation of other immune checkpoints, PD-1/PD-L1, VISTA, TIGIT, LAG-3, also promotes immunotherapy resistance [90]. For example, upregulated expression of VISTA is one of the factors responsible for resistance to anti-PD-1 therapy in patients with metastatic melanoma[91].
Immunosuppressive tumor microenvironment
The TME, comprising tumor and stromal cells, immune cells, blood vessels, and signaling molecules, plays a pivotal role in immunotherapy resistance. An immunosuppressive TME, marked by nutritional deficiencies, hypoxia, acidity, and abundant immunosuppressive cells (Treg cells, tumor-associated macrophages, myeloid-derived suppressor cells), impedes anti-tumor responses and promotes resistance [92,93,94]. Tumor cells reshape the TME through metabolic processes like aerobic glycolysis (Warburg effect), creating an acidic environment that hampers CD8+ T cell cytotoxicity and proliferation, inhibiting immunotherapy sensitivity [95]. Tumor cells also influence the TME by secreting chemokines that recruit immunosuppressive cells, affecting effector T cell function. For instance, MDSCs inhibit the therapeutic effects of anti-CTLA-4 in head and neck tumors [96].
Tumor-extrinsic factors of immunotherapy resistance: host factors
Patient-specific factors, including gender, diet, obesity, and gut microbiota, may impact immunotherapy efficacy [83]. Among these, gut microbiota has garnered significant attention, influencing resistance to ICI therapy. Analysis of the gut microbiome in melanoma patients treated with anti-PD-1 revealed associations between specific bacterial abundances and prolonged progression-free survival [97]. Gut microbes have been identified as potential determinants or biomarkers of immunotherapy response in various cancers [98, 99].
Other factors contributing to resistance include patient-specific differences in immune function and suboptimal dosing or treatment regimens. In essence, immunotherapy resistance involves intricate interactions among tumor cells, immune cells, cytokines, and signaling pathways. A comprehensive understanding of these mechanisms is crucial for developing effective strategies to overcome resistance and enhance clinical outcomes.
CircRNAs in regulating response to cancer immunotherapy
The potential of circRNAs as biomarkers for predicting immunotherapy response
Beyond traditional protein biomarkers like PD-L1, TMB, MSI-H, and dMMR [58, 100,101,102], circRNAs have emerged as innovative biomarkers for tumor immunotherapy, offering potential targets due to their stability in body fluids and specificity across tissues and developmental stages [103]. Recent findings demonstrate that plasma hsa_circ_0000190 levels are inversely correlated with immunotherapy response in advanced lung cancer patients. Notably, hsa_circ_0000190 exhibits promise as a novel biomarker for immunotherapy effectiveness, independent of PD-L1 expression levels [104]. Zhou et al. identified the inaugural circRNA signature (hsa_circ_0006408, hsa_circ_0032116, hsa_circ_0003633, hsa_circ_0066874, and hsa_circ_0006508) capable of predicting survival benefits in advanced melanoma patients undergoing anti-PD-1 immunotherapy [105]. Another study identified has_CDR1 as a potential response biomarker for effectively predicting anti-PD-1 therapy outcomes in patients with cutaneous metastatic melanoma [106]. Additionally, Gao et al. demonstrated that hsa_circ_0066351, through the construction of risk assessment models, is associated with colorectal cancer prognosis and immunotherapy response [107]. These investigations underscore the potential of circRNAs as valuable biomarkers for predicting immunotherapy response, presenting an opportunity to inform treatment decisions and enhance patient outcomes. However, the clinical application of circRNAs as predictive biomarkers for immunotherapeutic response necessitates further exploration.
Role of circular RNAs in regulating resistance to cancer immunotherapy (Fig. 4)
Immunotherapy resistance can occur through various mechanisms, including alterations in antigen presentation, reduction of immunogenicity, and activation of negative immune checkpoints, etc. The unique closed-loop structure and biological functions of circRNAs lead to their ability to inhibit or promote tumor progression and mediate resistance to cancer therapy [108]. Here, we summarize the mechanisms by which circRNAs are involved in immunotherapy resistance in different tumors (Table 1).
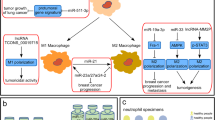
A CircRNAs in immunotherapy resistance in different tumors. The purple and pink circles represent down-regulated and up-regulated circRNAs, respectively. Orange and blue colors signify downstream target miRNAs and target proteins influenced by circRNAs, respectively. Sections of the figure were sourced from SMART—Servier Medical Art (https://smart.servier.com, last accessed September 17th, 2023). B Molecular mechanisms of circRNAs in ICI therapeutic resistance. Illustrated are three exemplar molecular mechanisms featuring CircIGF2BP3, CircIZNF451, and CircMGA contributing to Immune Checkpoint Inhibitor (ICI) therapeutic resistance.
Lung cancer (LC)
Lung cancer is a highly lethal tumor globally. Immunotherapy has improved long-term survival, about 40–50% of patients show resistance in the first cycle [109]. CircRNAs play a pivotal role in diminishing immunotherapy sensitivity through intricate molecular mechanisms across various lung cancer types. Notable examples include CircIGF2BP3 in NSCLC, associated with poor prognosis by reducing CD8+ T cell immune infiltration and upregulating PKP3 expression via sponging miR-328-3p and miR-3173-5p, ultimately preventing PD-L1 degradation and promoting immune escape [110]. CircCELF1 restricts T cell recruitment at the TME and inhibits anti-PD-1 therapy by sponging miR-491-5p, leading to upregulated EGRF expression [111]. CircASCC3 induces C5a activation, remodels the immunosuppressive TME, and induces anti-PD-1 therapeutic resistance [112]. Similarly, CircFGFR1 exerts immunosuppressive effects by sponging miR-381-3p, reducing CD8+ T cell infiltration [113], while hsa_circ_0003222, hsa_circ_0020714 and hsa_circ_0000190 promote proliferation, stemness, and anti-PD-1 therapeutic resistance in NSCLC [114,115,116]. Exosomal CircUSP7 inhibits CD8+ T cell function, contributing to immunotherapy resistance [117]. In lung adenocarcinoma (LUAD), circRUNX1 affects antigen presentation by sponging miR-4739 [118], and circ_0004140 reduces anti-PD-1 therapeutic effect by promoting CCL22 expression through sponging miR-1184, recruiting Treg cells [119]. Exosomal circZNF451 plays a crucial role in restructuring the tumor-immune microenvironment by influencing macrophage polarization through the FXR1-ELF4–IRF4 axis, serving as a promising biomarker to predict the responsiveness of PD-1 blockade in LUAD [120]. Additionally, circHMGB2 limits PD-1 blockade efficacy by upregulating CARM1 through interacting with miR-181a-5p, inhibiting the type 1 IFN response and desensitizing tumor cells to cytotoxic T-cell-mediated immune responses [121].
Hepatocellular carcinoma (HCC)
Liver cancer, a prevalent global malignancy, is predominantly represented by HCC, constituting around 90% of all liver cancer cases. Detection of HCC typically occurs in advanced stages, and despite the incremental survival benefit offered by current therapies, the challenge of immunotherapy resistance persists due to the immunosuppressive TME [122]. Elevated expression of circMET in HCC cells correlates with increased invasion, metastasis, and resistance to immunotherapy. Mechanistically, circMET upregulates snail and its downstream DPP4 by sponging miR-30-5p, fostering immunosuppression through the miR-30-5p/snail/DPP4 axis, leading to CXCL10 degradation. The DPP4 inhibitor Sitagliptin enhances the effectiveness of anti-PD-1 therapy [123]. In HCC, resistance to anti-PD-1 treatment can be induced through tumor-macrophage interactions. Exosome circTMEM181 in the TME upregulates CD39 expression in macrophages, activating the ATP-adenosine pathway and inducing immunosuppression. Targeting the adenosine pathway can enhance the efficacy of ICB therapy [124]. Similarly, exosome-delivered circUHRF1 induces immunosuppressive effects by upregulating TIM-3 expression in NK cells, leading to NK cell exhaustion and inhibiting IFN-γ and TNF-α secretion, ultimately mediating HCC resistance to anti-PD-1 therapy. The underlying molecular mechanism involves circUHRF1-mediated degradation of miR-449c-5p, leading to heightened TIM-3 expression in NK cells and subsequent immunosuppression [125]. Exosome-derived circCCAR1 targets miR-127-5p to upregulate WTAP in activated T cells, causing CD8+ T cell dysfunction and promoting HCC resistance to anti-PD-1 therapy [126]. High levels of CircRHBDD1 are associated with poor patient prognosis and reduced sensitivity to anti-PD-1 therapy, and its inhibition enhances the therapeutic effect of anti-PD-1 by interacting with YTHDF1 to enhance PIK3R1 translation [127]. CircSOD2, highly expressed in HCC tissues, is linked to decreased CD8+ T cell activity in TME, inducing anti-PD-1 therapeutic resistance by binding to miR-497-5p to upregulate ANXA11 [128].
Colorectal cancer (CRC)
CRC holds the third position in global cancer incidence and ranks second in cancer-related deaths [129]. Traditional treatments face challenges due to chemotherapy drug side effects and the unique biological characteristics of tumor cells. Although ICB therapy has introduced a new approach, it proves effective in only ~15% of CRC patients with high microsatellite instability (MSI-H). Furthermore, some initially responsive patients develop acquired resistance [130]. In CRC patients, the upregulation of circQSOX1 contributes to resistance to anti-CTLA-4 treatment. Mechanistically, circQSOX1 sponges miR-326 and miR-330-5p, subsequently upregulating the expression of phosphoglycerate mutase 1 (PGAM1), promoting glycolysis in CRC cells. Additional investigations reveal that the knockdown of circQSOX1 reduces the infiltration of immunosuppressive Treg cells, thereby leading to the development of immunotherapy resistance by remodeling the immunosuppressive TME [131].
Bladder cancer(Bca)
ICB therapy has demonstrated significant clinical efficacy in BCa, ranking as the ninth most common tumor globally [132]. Despite these advances, the majority of patients do not respond to immunotherapy. A recent study reveals that the downregulation of circFAM13B expression in BCa correlates with a poorer prognosis and resistance to anti-PD-1 treatment. Mechanistically, circFAM13B inhibits the stability of PKM2 mRNA by interacting with IGF2BP1 via the K homology 3–4 (KH3-4) domains, consequently suppressing glycolysis in which PKM2 is involved. Consequently, the decreased circFAM13B fails to inhibit glycolysis, contributing to an immunosuppressive TME. In HuNOG mouse models bearing BCa tumors, the overexpression of circFAM13B enhances the effectiveness of CD8+ T cells and sensitivity to anti-PD-1 therapy [133]. Similarly, the downregulation of circMGA reduces the therapeutic sensitivity of anti-PD-1. CircMGA forms an RNA-protein complex by binding with HNRNPL, leading to the upregulation of CCL5 and increased infiltration of CD8+ T cells, thereby enhancing the immunotherapeutic effect of anti-PD-1 [134].
Melanoma
In the nearly decade since the approval of ipilimumab for metastatic melanoma treatment in 2011, despite the initial effectiveness of ICB therapy in 40–45% of patients, the majority eventually develop acquired drug resistance [135, 136]. A study by Wei et al. indicates a positive correlation between circ_0020710 expression and poor prognosis, with upregulated circ_0020710 reducing sensitivity to anti-PD-1 treatment. Mechanistically, circ_0020710 upregulates CXCL12 expression by sponging miR-370-3p. Elevated levels of CXCL12 contribute to the recruitment of immunosuppressive cells, fostering an immunosuppressive microenvironment and resulting in decreased infiltration of cytotoxic T lymphocytes. This ultimately leads to the development of immunotherapeutic resistance [137].
Head and neck squamous cell carcinoma (HNSCC)
Preclinical evidence suggests that HNSCC may elude immune surveillance and induce immunosuppression, resulting in a limited response to anti-PD-1 antibodies in most patients [138]. Cancer stem cells, a small subset of tumor cells crucial for self-renewal and promoting malignant tumor development, also contribute to resistance against radiotherapy and chemotherapy [139]. A recent study has unveiled that circRNA can diminish immunotherapy sensitivity by regulating HNSCC cell stemness. In the investigation conducted by Jia et al., circFAT1 expression was found to be upregulated in head and neck squamous cell carcinoma, correlating with poor prognosis and reduced efficacy of immunotherapy. CircFAT1 binds to STAT3 in the cytoplasm, hindering SHP1-mediated dephosphorylation and activating STAT3. Knocking down circFAT1 attenuates tumor stemness and enhances CD8+ T cell infiltration at tumor sites, thereby boosting the effectiveness of anti-PD-1 therapy [140].
Gastric cancer
Gastric cancer poses a significant clinical challenge, and despite the introduction of anti-PD-1 therapy, the majority of patients do not achieve favorable outcomes. In a study by Chen et al., the upregulation of circDLG1 was found to be significantly associated with an aggressive tumor phenotype and poor prognosis in gastric cancer patients undergoing anti-PD-1 therapy. Mechanistically, circDLG1 upregulates the expression of CXCL12 by sponging miR-141-3p. The elevated CXCL12 levels further contribute to increased MDSCs and decreased infiltration of CD8+ T cells in TME. Notably, the knockdown of CXCL12 demonstrates the potential to enhance the sensitivity of gastric cancer to anti-PD treatment, suggesting a critical role for circDLG1 in mediating immunotherapeutic responses [141].
Future directions and discussion
CircRNAs, characterized by a stable covalent closed-loop structure, exhibit tissue-specific and developmental stage-specific distribution. Over the past decade, extensive research has deepened our understanding of the pivotal role of circRNAs in human diseases and physiological processes [142]. In oncology, circRNAs act as miRNA sponges, protein decoys, or translated proteins, exerting regulatory control over tumor growth. Recent explorations have extended their involvement in chemotherapy, radiation therapy, targeted therapy, and immunotherapy resistance. The current focus of circRNA research in immunotherapy resistance is mainly on lung cancer and HCC, particularly in the context of ICB therapy. Mechanistically, circRNAs modulate immunotherapy resistance by binding to miRNAs or serving as protein scaffolds. However, the potential involvement of circRNAs in immunotherapy resistance through other functions remains an area requiring further investigation.
Recently, immunotherapy has garnered significant attention as a highly effective treatment compared to traditional radiotherapy and chemotherapy. Despite achieving high response rates in clinical treatment, the emergence of drug resistance remains a substantial challenge [53]. Research on circRNAs has sparked a new wave in the field of immunotherapy. As scientists delve into understanding the role of circRNAs in regulating resistance to cancer immunotherapy, the identification of novel circRNAs involved in this process could unveil new targets for therapeutic intervention. With over 100,000 different human circRNAs discovered to date, their dysregulation is linked to tumor development across various cancers such as breast, liver, lung, prostate, neuroblastoma, and stomach cancers [143,144,145,146,147]. This dysregulation may contribute to tumor progression or treatment resistance [148, 149]. However, further studies are needed to determine the functionality of differentially expressed circRNAs in normal and tumor tissues, their role in tumorigenesis, development, regulation of immunotherapy resistance, and their potential as drug targets.
PD-L1 and PD-1 are pivotal immune checkpoint proteins targeted in cancer immunotherapy. They regulate immune responses and are significant therapeutic targets. PD-L1, often upregulated in tumors, enables immune evasion by binding to PD-1 on T cells, leading to T cell dysfunction and apoptosis. ICIs that block the PD-1/PD-L1 axis have transformed cancer treatment by reactivating T cell anti-tumor responses. However, resistance to ICIs remains a challenge. Recent studies have focused on circRNAs’ role in modulating the PD-1/PD-L1 pathway, shedding light on novel mechanisms of immunotherapy resistance. For instance, circRNAs like circUHRF1 have been found to be significantly upregulated in oral squamous cell carcinoma (OSCC), and acts as a sponge for miR-526b-5p, leading to increased PD-L1 expression and immune evasion [150]. Similarly, in OSCC, m6A-circRNAs exhibit distinct modification patterns, providing insights into their roles in cancer progression [151]. These findings underscore the intricate regulatory network involving circRNAs, miRNAs, and immune checkpoint proteins in tumor immunotherapy resistance. Understanding this crosstalk is crucial for developing effective strategies to overcome immunotherapy resistance and improve patient outcomes in cancer treatment.
The low cellular content of circRNAs, coupled with the presence of linear mRNA sharing the same sequence, poses a significant obstacle to accurate circRNA identification and detection. The development of more precise detection methods is essential. Current circRNA detection methods include real-time quantitative polymerase chain reaction, droplet digital PCR, circRNA fluorescence in situ hybridization, high-throughput sequencing technology, nanopore sequencing technology, and emerging methods like Duplex-specific nuclease, and loop-mediated isothermal amplification (LAMP) [152]. These varied analytical methods offer scientists more choices, and emerging techniques promise to provide a more robust tool for the clinical application of circRNA.
The Cancer CircRNA Immunome Atlas (TCCIA), a recently launched website available at http://biotrainee.vip:18888/TCCIA/ or https://shiny.hiplot.cn/TCCIA, stands out as an innovative online platform facilitating the study of circRNA expression and analysis across 25 patient cohorts undergoing immunotherapy targeting CTLA4, PD-1, or PDL-1. This platform serves as a unique avenue to delve into circRNAs, evaluating their potential as biomarkers to predict immunotherapy responses and unraveling their broader implications in cancer research [153]. Volcano plots depicting the differential expression of circRNAs between non-responsive and responsive immunotherapy patients in various tumors were acquired from TCCIA (Fig. 5). The circRNAs, identified with circBase IDs, ranking among the top 5 up-or down-regulated in each tumor were highlighted in green (Log2FC ≤ −1 or Log2FC ≥ 1; P.adj < 0.05; method, DEseq2). Notably, our analysis revealed a downregulation of several circRNAs transcribed by the EYA1 gene in SKCM. Likewise, in SCLC, multiple circRNAs derived from NEB and ANKRD36C genes exhibited a significant down-regulation trend. We hypothesize that these circRNAs might play a role in tumor immunotherapy resistance or potentially serve as biomarkers for immunotherapy efficacy, and thorough validation through further studies is imperative.
A–G Differential expression of circRNAs in patients who responded and did not respond to immunotherapy in SKCM, RCC, STAD, NSCLC, SCLC, SGC and NHSC, respectively. The data were sourced from the TCCIA website (http://biotrainee.vip:18888/TCCIA/, last accessed on January 7th, 2024). CircRNAs lacking circBase ID are denoted in the format of circ + parent genes. Abbreviations: SKCM skin cutaneous melanoma, RCC renal cell carcinoma, STAD stomach adenocarcinoma, NSCLC non-small cell lung cancer; SCLC small cell lung cancer, SGC salivary gland cancer, NHSC head and neck squamous cell carcinoma.
The identification of circRNAs as regulators of immune checkpoint gene expression presents a promising avenue for developing novel therapeutic strategies to enhance the effectiveness of cancer immunotherapy. Techniques such as gene knockout, RNA interference (RNAi), and antisense technologies are currently employed to inhibit the function of oncogenic circRNAs. In a recent study published in Nature Methods, the clustered regularly interspaced short palindromic repeats and associated proteins (CRISPR-Cas)13 system was demonstrated to specifically knock down circRNAs by targeting the back-splice junction (BSJ) instead of intronic complementary sequences [154]. Moreover, targeting signaling pathways influenced by circRNAs, as demonstrated in studies on breast cancer and glioblastoma [155, 156], can also exhibit anti-tumor effects when direct circRNA targeting is challenging.
Additionally, circRNAs can be utilized as tumor antigens or vaccine adjuvants in anti-tumor therapy [157]. Although clinical application of circRNA tumor vaccines is not yet realized, preclinical studies show promising results, with engineered circRNAs expected to be the next generation of RNA vaccines. Currently, three in vivo delivery vectors for circRNAs-Lipid nanoparticles (LNPs), Exosomes, and Virus-like particles (VLPs)-are being explored [158, 159]. LNPs offer advantages such as nuclease resistance, kidney clearance avoidance, and increased cellular uptake. Exosomes, natural endogenous carriers, provide low toxicity, no immunogenicity, and good permeability. VLPs, self-assembled nanocarriers, trigger a strong immune response without containing viral nucleic acids, demonstrating effective tissue targeting and deliverability. Several therapeutic studies have demonstrated the potential of these delivery methods. For example, PEG-Au-loaded circ-Foxo3 expression plasmids enhance the sensitivity of prostate cancer-bearing mice to docetaxel [160]. RGD-modified exosomes carrying circDIDO1 inhibit gastric cancer development [161], and oxaliplatin-resistant cells deliver circRNA ciRS-122 to sensitive cells via exosomes in colorectal cancer, reversing drug resistance in tumor-bearing mice [162]. While these delivery methods hold promise, challenges such as off-target effects, toxicity, standardized manufacturing, quality control, and the need for novel, precise drug delivery systems responsive to enzymes, pH, or hypoxia must be addressed. Comprehensive consideration based on target circRNA, tumor type, and clinical pathology is essential for successful implementation.
The discovery of circRNAs as regulators of immune checkpoint gene expression suggests their potential integration into existing cancer diagnostic and therapeutic workflows, aiming to improve the accuracy and efficacy of cancer treatment and enhance patient outcomes. Early cancer diagnosis remains a significant challenge, emphasizing the need for reliable biomarkers. CircRNAs show promise as biomarkers, capable of differentiating cancer subtypes and guiding treatment decisions. For instance, Tan et al. discovered F-circEA, a novel fusion circRNA specific to EML4-ALK-positive NSCLC patients, serving as a potential diagnostic biomarker and guiding targeted therapy with ALK inhibitors [6, 163]. CircRNAs may aid in immunotherapy selection by detecting their expression in tumor tissues. The expression of PD-L1, considered a biomarker for immunotherapy sensitivity, is intricately regulated by circRNAs, as illustrated in Table 2. However, it is essential to note that elevated PD-L1 expression does not universally guarantee better outcomes in ICB treatment. “Role of circular RNAs in regulating resistance to cancer immunotherapy” highlights the role of circIGF2BP3, which upregulates tumor cell PD-L1 expression, potentially leading to T cell dysfunction-mediated immune escape and immunotherapy resistance. Despite these complexities, circRNAs continue to play a crucial role in tumor diagnosis and treatment.
Conclusions
In this review, we introduce the biogenesis and biological functions of circRNAs, various immunotherapy methods, and factors linked to tumor immunotherapy resistance. We also explore the potential of circRNAs as biomarkers for predicting immune responses and summarize their current regulatory mechanisms in different tumor immunotherapy resistance scenarios. In current studies on immunotherapeutic resistance, circRNAs predominantly either promote or alleviate drug resistance by serving as miRNA sponges or protein decoys. However, the regulation of immunotherapy resistance by circRNAs might involve other mechanisms beyond these, warranting exploration. Uncovering additional mechanisms of circRNAs in immunotherapy resistance could lead to the identification of new therapeutic targets or biomarkers, enhancing tumor diagnosis and prognosis accuracy, enabling precise personalized treatment, and improving overall patient outcomes. The substantial potential of circRNAs in tumor treatment is promising, but the translation of preclinical research into clinical practice requires careful consideration.
References
Chhikara BS, Parang K. Global Cancer Statistics 2022: the trends projection analysis. Chem. Biol. Lett. 2023;10:451.
Farkona S, Diamandis EP, Blasutig IM. Cancer immunotherapy: the beginning of the end of cancer? BMC Med. 2016;14:73.
Chen S, Zhang Z, Zheng X, Tao H, Zhang S, Ma J, et al. Response efficacy of PD-1 and PD-L1 inhibitors in clinical trials: a systematic review and meta-analysis. Front Oncol. 2021;11:562315.
Andrews LP, Yano H, Vignali DAA. Inhibitory receptors and ligands beyond PD-1, PD-L1 and CTLA-4: breakthroughs or backups. Nat Immunol. 2019;20:1425–34.
Chen L-L. The expanding regulatory mechanisms and cellular functions of circular RNAs. Nat Rev Mol Cell Biol. 2020;21:475–90.
Kristensen LS, Jakobsen T, Hager H, Kjems J. The emerging roles of circRNAs in cancer and oncology. Nat Rev Clin Oncol. 2022;19:188–206.
Guo L, Jia L, Luo L, Xu X, Xiang Y, Ren Y, et al. Critical roles of circular RNA in tumor metastasis via acting as a sponge of miRNA/isomiR. Int J Mol Sci. 2022;23:7024. Available from: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC9267010/.
Papatsirou M, Artemaki PI, Karousi P, Scorilas A, Kontos CK. Circular RNAs: emerging regulators of the major signaling pathways involved in cancer progression. Cancers. 2021;13:2744.
Sanger HL, Klotz G, Riesner D, Gross HJ, Kleinschmidt AK. Viroids are single-stranded covalently closed circular RNA molecules existing as highly base-paired rod-like structures. Proc Natl Acad Sci. 1976;73:3852–6.
Arnberg AC, Van Ommen G-JB, Grivell LA, Van Bruggen EFJ, Borst P. Some yeast mitochondrial RNAs are circular. Cell. 1980;19:313–9.
Kos A, Dijkema R, Arnberg AC, Van Der Meide PH, Schellekens H. The hepatitis delta (δ) virus possesses a circular RNA. Nature. 1986;323:558–60.
Saad FA, Vitiello L, Merlini L, Mostacciuolo ML, Oliviero S, Danieli GA. A 3′ consensus splice mutation in the human dystrophin gene detected by a screening for intra-exonic deletions. Hum Mol Genet. 1992;1:345–6.
Capel B, Swain A, Nicolis S, Hacker A, Walter M, Koopman P, et al. Circular transcripts of the testis-determining gene Sry in adult mouse testis. Cell. 1993;73:1019–30.
Salzman J, Gawad C, Wang PL, Lacayo N, Brown PO. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS One. 2012;7:e30733.
Zhou W-Y, Cai Z-R, Liu J, Wang D-S, Ju H-Q, Xu R-H. Circular RNA: metabolism, functions and interactions with proteins. Mol Cancer. 2020;19:172.
Salzman J. Circular RNA expression: its potential regulation and function. Trends Genet. 2016;32:309–16.
Ashwal-Fluss R, Meyer M, Pamudurti NR, Ivanov A, Bartok O, Hanan M, et al. circRNA biogenesis competes with Pre-mRNA splicing. Mol Cell. 2014;56:55–66.
Chen L-L, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015;12:381–8.
Huang Y, Zhu Q. Mechanisms regulating abnormal circular RNA biogenesis in cancer. Cancers. 2021;13:4185.
Kristensen LS, Andersen MS, Stagsted LVW, Ebbesen KK, Hansen TB, Kjems J. The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet. 2019;20:675–91.
Chen L, Wang Y, Lin J, Song Z, Wang Q, Zhao W, et al. Exportin 4 depletion leads to nuclear accumulation of a subset of circular RNAs. Nat Commun. 2022;13:5769.
Li Z, Huang C, Bao C, Chen L, Lin M, Wang X, et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat Struct Mol Biol. 2015;22:256–64.
Chen L-L. The biogenesis and emerging roles of circular RNAs. Nat Rev Mol Cell Biol. 2016;17:205–11.
Enuka Y, Lauriola M, Feldman ME, Sas-Chen A, Ulitsky I, Yarden Y. Circular RNAs are long-lived and display only minimal early alterations in response to a growth factor. Nucleic Acids Res. 2016;44:1370–83.
Sang C, Rao D, Wu C, Xia Y, Si M, Tang Z. Role of circular RNAs in the diagnosis, regulation of drug resistance and prognosis of lung cancer. Oncol Lett. 2022;24:302.
Huang A, Zheng H, Wu Z, Chen M, Huang Y. Circular RNA-protein interactions: functions, mechanisms, and identification. Theranostics. 2020;10:3503–17.
Hansen TB, Jensen TI, Clausen BH, Bramsen JB, Finsen B, Damgaard CK, et al. Natural RNA circles function as efficient microRNA sponges. Nature. 2013;495:384–8.
Li J, Gao X, Zhang Z, Lai Y, Lin X, Lin B, et al. CircCD44 plays oncogenic roles in triple-negative breast cancer by modulating the miR-502–5p/KRAS and IGF2BP2/Myc axes. Mol Cancer. 2021;20:138.
Han D, Li J, Wang H, Su X, Hou J, Gu Y, et al. Circular RNA circMTO1 acts as the sponge of microRNA-9 to suppress hepatocellular carcinoma progression. Hepatology. 2017;66:1151–64.
Du WW, Fang L, Yang W, Wu N, Awan FM, Yang Z, et al. Induction of tumor apoptosis through a circular RNA enhancing Foxo3 activity. Cell Death Differ. 2017;24:357–70.
Wang L, Long H, Zheng Q, Bo X, Xiao X, Li B. Circular RNA circRHOT1 promotes hepatocellular carcinoma progression by initiation of NR2F6 expression. Mol Cancer. 2019;18:119.
Li X, Zhang J-L, Lei Y-N, Liu X-Q, Xue W, Zhang Y, et al. Linking circular intronic RNA degradation and function in transcription by RNase H1. Sci China Life Sci. 2021;64:1795–809.
Guarnerio J, Zhang Y, Cheloni G, Panella R, Mae Katon J, Simpson M, et al. Intragenic antagonistic roles of protein and circRNA in tumorigenesis. Cell Res. 2019;29:628–40.
Su Y, Feng W, Shi J, Chen L, Huang J, Lin T. circRIP2 accelerates bladder cancer progression via miR-1305/Tgf-β2/smad3 pathway. Mol Cancer. 2020;19:23.
Ma S, Kong S, Wang F, Ju S. CircRNAs: biogenesis, functions, and role in drug-resistant Tumours. Mol Cancer. 2020;19:119.
AbouHaidar MG, Venkataraman S, Golshani A, Liu B, Ahmad T. Novel coding, translation, and gene expression of a replicating covalently closed circular RNA of 220 nt. Proc Natl Acad Sci USA 2014;111:14542–7.
Meyer KD, Patil DP, Zhou J, Zinoviev A, Skabkin MA, Elemento O, et al. 5′ UTR m6A promotes cap-independent translation. Cell. 2015;163:999–1010.
Wang Y, Wu C, Du Y, Li Z, Li M, Hou P, et al. Expanding uncapped translation and emerging function of circular RNA in carcinomas and noncarcinomas. Mol Cancer. 2022;21:13.
Yang Y, Fan X, Mao M, Song X, Wu P, Zhang Y, et al. Extensive translation of circular RNAs driven by N6-methyladenosine. Cell Res. 2017;27:626–41.
Yang Y, Gao X, Zhang M, Yan S, Sun C, Xiao F, et al. Novel role of FBXW7 circular RNA in repressing glioma tumorigenesis. JNCI J Natl Cancer Inst. 2017;110:304–15.
Kong S, Tao M, Shen X, Ju S. Translatable circRNAs and lncRNAs: driving mechanisms and functions of their translation products. Cancer Lett. 2020;483:59–65.
Diallo LH, Tatin F, David F, Godet A-C, Zamora A, Prats A-C, et al. How are circRNAs translated by non-canonical initiation mechanisms? Biochimie. 2019;164:45–52.
Zhang Y, Li C, Liu X, Wang Y, Zhao R, Yang Y, et al. circHIPK3 promotes oxaliplatin-resistance in colorectal cancer through autophagy by sponging miR-637. EBioMedicine. 2019;48:277–88.
Liu Y, Xia L, Dong L, Wang J, Xiao Q, Yu X, et al. CircHIPK3 promotes gemcitabine (GEM) resistance in pancreatic cancer cells by sponging miR-330-5p and targets RASSF1. Cancer Manag Res. 2020;12:921–9.
Hua J, Wang X, Ma L, Li J, Cao G, Zhang S, et al. CircVAPA promotes small cell lung cancer progression by modulating the miR-377-3p and miR-494-3p/IGF1R/AKT axis. Mol Cancer. 2022;21:123.
Yu L, Sun M, Zhang Q, Zhou Q, Wang Y. Harnessing the immune system by targeting immune checkpoints: Providing new hope for Oncotherapy. Front Immunol. 2022;13:982026.
Wang J, Zheng Y, Tu C, Zhang H, Vanderkerken K, Menu E, et al. Identification of the immune checkpoint signature of multiple myeloma using mass cytometry‐based single‐cell analysis. Clin Transl Immunol. 2020;9:e1132. Available from: https://doi.org/10.1002/cti2.1132.
Wang Y, Wang M, Wu H-X, Xu R-H. Advancing to the era of cancer immunotherapy. Cancer Commun. 2021;41:803–29.
Wolchok J. Putting the immunologic brakes on cancer. Cell. 2018;175:1452–4.
Li X, Shao C, Shi Y, Han W. Lessons learned from the blockade of immune checkpoints in cancer immunotherapy. J Hematol Oncol. 2018;11:31.
Larkin J, Chiarion-Sileni V, Gonzalez R, Grob J-J, Rutkowski P, Lao CD, et al. Five-year survival with combined nivolumab and ipilimumab in advanced melanoma. N. Engl J Med. 2019;381:1535–46.
Liu Q, Li J, Zheng H, Yang S, Hua Y, Huang N, et al. Adoptive cellular immunotherapy for solid neoplasms beyond CAR-T. Mol Cancer. 2023;22:28.
Taefehshokr S, Parhizkar A, Hayati S, Mousapour M, Mahmoudpour A, Eleid L, et al. Cancer immunotherapy: challenges and limitations. Pathol - Res Pr. 2022;229:153723.
Gun SY, Lee SWL, Sieow JL, Wong SC. Targeting immune cells for cancer therapy. Redox Biol. 2019;25:101174.
Maalej KM, Merhi M, Inchakalody VP, Mestiri S, Alam M, Maccalli C, et al. CAR-cell therapy in the era of solid tumor treatment: current challenges and emerging therapeutic advances. Mol Cancer. 2023;22:20.
Adamik J, Butterfield LH. What’s next for cancer vaccines. Sci Transl Med. 2022;14:eabo4632.
Hu Z, Ott PA, Wu CJ. Towards personalized, tumour-specific, therapeutic vaccines for cancer. Nat Rev Immunol. 2018;18:168–82.
Wang D-R, Wu X-L, Sun Y-L. Therapeutic targets and biomarkers of tumor immunotherapy: response versus non-response. Signal Transduct Target Ther. 2022;7:331.
Saxena M, van der Burg SH, Melief CJM, Bhardwaj N. Therapeutic cancer vaccines. Nat Rev Cancer. 2021;21:360–78.
Lorentzen CL, Haanen JB, Met Ö, Svane IM. Clinical advances and ongoing trials of mRNA vaccines for cancer treatment. Lancet Oncol. 2022;23:e450–8.
Kaufman HL, Kohlhapp FJ, Zloza A. Oncolytic viruses: a new class of immunotherapy drugs. Nat Rev Drug Discov. 2015;14:642–62.
Malhotra J, Kim ES. Oncolytic viruses and cancer immunotherapy. Curr Oncol Rep. 2023;25:19–28.
Zou H, Mou X, Zhu B. Combining of oncolytic virotherapy and other immunotherapeutic approaches in cancer: a powerful functionalization tactic. Glob Chall. 2023;7:2200094.
Shalhout SZ, Miller DM, Emerick KS, Kaufman HL. Therapy with oncolytic viruses: progress and challenges. Nat Rev Clin Oncol. 2023;20:160–77.
Chesney J, Puzanov I, Collichio F, Singh P, Milhem MM, Glaspy J, et al. Randomized, open-label phase ii study evaluating the efficacy and safety of talimogene laherparepvec in combination with ipilimumab versus ipilimumab alone in patients with advanced, unresectable melanoma. J Clin Oncol. 2018;36:1658–67.
Wu Y, Yang Z, Cheng K, Bi H, Chen J. Small molecule-based immunomodulators for cancer therapy. Acta Pharm Sin B. 2022;12:4287–308.
Lu Y, Gao Y, Yang H, Hu Y, Li X. Nanomedicine‐boosting icaritin-based immunotherapy of advanced hepatocellular carcinoma. Mil Med Res. 2022;9:69.
Zhu S, Zhang T, Zheng L, Liu H, Song W, Liu D, et al. Combination strategies to maximize the benefits of cancer immunotherapy. J Hematol Oncol. 2021;14:156.
Yap TA, Parkes EE, Peng W, Moyers JT, Curran MA, Tawbi HA. Development of immunotherapy combination strategies in cancer. Cancer Discov. 2021;11:1368–97.
Kennedy LB, Salama AKS. A review of cancer immunotherapy toxicity. CA Cancer J Clin. 2020;70:86–104.
Bashash D, Zandi Z, Kashani B, Pourbagheri‐Sigaroodi A, Salari S, Ghaffari SH. Resistance to immunotherapy in human malignancies: Mechanisms, research progresses, challenges, and opportunities. J Cell Physiol. 2022;237:346–72.
Sun J-Y, Zhang D, Wu S, Xu M, Zhou X, Lu X-J, et al. Resistance to PD-1/PD-L1 blockade cancer immunotherapy: mechanisms, predictive factors, and future perspectives. Biomark Res. 2020;8:35.
Wang Z, Wu X. Study and analysis of antitumor resistance mechanism of PD1/PD‐L1 immune checkpoint blocker. Cancer Med. 2020;9:8086–121.
Baretti M, Le DT. DNA mismatch repair in cancer. Pharm Ther. 2018;189:45–62.
Duffy MJ, Crown J. Biomarkers for predicting response to immunotherapy with immune checkpoint inhibitors in cancer patients. Clin Chem. 2019;65:1228–38.
Pang K, Shi Z-D, Wei L-Y, Dong Y, Ma Y-Y, Wang W, et al. Research progress of therapeutic effects and drug resistance of immunotherapy based on PD-1/PD-L1 blockade. Drug Resist Updat. 2023;66:100907.
Wang H, Liu B, Wei J. Beta2-microglobulin(B2M) in cancer immunotherapies: biological function, resistance and remedy. Cancer Lett. 2021;517:96–104.
Gettinger S, Choi J, Hastings K, Truini A, Datar I, Sowell R, et al. Impaired HLA class I antigen processing and presentation as a mechanism of acquired resistance to immune checkpoint inhibitors in lung cancer. Cancer Discov. 2017;7:1420–35.
Zaretsky JM, Garcia-Diaz A, Shin DS, Escuin-Ordinas H, Hugo W, Hu-Lieskovan S, et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. N. Engl J Med. 2016;375:819–29.
Sotillo E, Barrett DM, Black KL, Bagashev A, Oldridge D, Wu G, et al. Convergence of acquired mutations and alternative splicing of CD19 enables resistance to CART-19 immunotherapy. Cancer Discov. 2015;5:1282–95.
Peng W, Chen JQ, Liu C, Malu S, Creasy C, Tetzlaff MT, et al. Loss of PTEN promotes resistance to T cell-mediated immunotherapy. Cancer Discov. 2016;6:202–16.
Vesely MD, Zhang T, Chen L. Resistance mechanisms to anti-PD cancer immunotherapy. Annu Rev Immunol. 2022;40:45–74.
Bai R, Chen N, Li L, Du N, Bai L, Lv Z, et al. Mechanisms of cancer resistance to immunotherapy. Front Oncol. 2020;10:1290.
Kalbasi A, Ribas A. Tumour-intrinsic resistance to immune checkpoint blockade. Nat Rev Immunol. 2020;20:25–39.
Xia A, Zhang Y, Xu J, Yin T, Lu X-J. T Cell Dysfunction in Cancer Immunity and Immunotherapy. Front Immunol. 2019;10:1719.
Ren D, Hua Y, Yu B, Ye X, He Z, Li C, et al. Predictive biomarkers and mechanisms underlying resistance to PD1/PD-L1 blockade cancer immunotherapy. Mol Cancer. 2020;19:19.
Huang R-Y, Francois A, McGray AR, Miliotto A, Odunsi K. Compensatory upregulation of PD-1, LAG-3, and CTLA-4 limits the efficacy of single-agent checkpoint blockade in metastatic ovarian cancer. Oncoimmunology. 2016;6:e1249561.
Shayan G, Srivastava R, Li J, Schmitt N, Kane LP, Ferris RL. Adaptive resistance to anti-PD1 therapy by Tim-3 upregulation is mediated by the PI3K-Akt pathway in head and neck cancer. Oncoimmunology. 2016;6:e1261779.
Koyama S, Akbay EA, Li YY, Herter-Sprie GS, Buczkowski KA, Richards WG, et al. Adaptive resistance to therapeutic PD-1 blockade is associated with upregulation of alternative immune checkpoints. Nat Commun. 2016;7:10501.
Chen X, Zhang W, Yang W, Zhou M, Liu F. Acquired resistance for immune checkpoint inhibitors in cancer immunotherapy: challenges and prospects. Aging. 2022;14:1048–64.
Kakavand H, Jackett LA, Menzies AM, Gide TN, Carlino MS, Saw RPM, et al. Negative immune checkpoint regulation by VISTA: a mechanism of acquired resistance to anti-PD-1 therapy in metastatic melanoma patients. Mod Pathol. 2017;30:1666–76.
Kim HJ, Ji YR, Lee YM. Crosstalk between angiogenesis and immune regulation in the tumor microenvironment. Arch Pharm Res. 2022;45:401–16.
Jiang T, Chen X, Ren X, Yang J-M, Cheng Y. Emerging role of autophagy in anti-tumor immunity: Implications for the modulation of immunotherapy resistance. Drug Resist Updat. 2021;56:100752.
Jiang Z, Hsu JL, Li Y, Hortobagyi GN, Hung M-C. Cancer cell metabolism bolsters immunotherapy resistance by promoting an immunosuppressive tumor microenvironment. Front Oncol. 2020;10:1197.
Hayes C, Donohoe CL, Davern M, Donlon NE. The oncogenic and clinical implications of lactate induced immunosuppression in the tumour microenvironment. Cancer Lett. 2021;500:75–86.
Clavijo PE, Moore EC, Chen J, Davis RJ, Friedman J, Kim Y, et al. Resistance to CTLA-4 checkpoint inhibition reversed through selective elimination of granulocytic myeloid cells. Oncotarget. 2017;8:55804–20.
Gopalakrishnan V, Spencer CN, Nezi L, Reuben A, Andrews MC, Karpinets TV, et al. Gut microbiome modulates response to anti–PD-1 immunotherapy in melanoma patients. Science. 2018;359:97–103.
Lee KA, Thomas AM, Bolte LA, Björk JR, de Ruijter LK, Armanini F, et al. Cross-cohort gut microbiome associations with immune checkpoint inhibitor response in advanced melanoma. Nat Med. 2022;28:535–44.
Matson V, Fessler J, Bao R, Chongsuwat T, Zha Y, Alegre M-L, et al. The commensal microbiome is associated with anti-PD-1 efficacy in metastatic melanoma patients. Science. 2018;359:104–8.
Davis AA, Patel VG. The role of PD-L1 expression as a predictive biomarker: an analysis of all US Food and Drug Administration (FDA) approvals of immune checkpoint inhibitors. J Immunother Cancer. 2019;7:278.
Chan TA, Yarchoan M, Jaffee E, Swanton C, Quezada SA, Stenzinger A, et al. Development of tumor mutation burden as an immunotherapy biomarker: utility for the oncology clinic. Ann Oncol. 2019;30:44–56.
Marcus L, Lemery SJ, Keegan P, Pazdur R. FDA approval summary: pembrolizumab for the treatment of microsatellite instability-high solid tumors. Clin Cancer Res. 2019;25:3753–8.
Fontemaggi G, Turco C, Esposito G, Di Agostino S. New molecular mechanisms and clinical impact of circRNAs in human cancer. Cancers. 2021;13:3154.
Luo Y-H, Yang Y-P, Chien C-S, Yarmishyn AA, Ishola AA, Chien Y, et al. Plasma level of circular RNA hsa_circ_0000190 correlates with tumor progression and poor treatment response in advanced lung cancers. Cancers. 2020;12:1740.
Zhou J-G, Liang R, Wang H-T, Jin S-H, Hu W, Frey B, et al. Identification and characterization of circular RNAs as novel putative biomarkers to predict anti-PD-1 monotherapy response in metastatic melanoma patients—Knowledge from two independent international studies. Neoplasia N. Y N. 2023;37:100877.
Oliver J, Onieva JL, Garrido-Barros M, Berciano-Guerrero M-Á, Sánchez-Muñoz A, José Lozano M, et al. Association of circular RNA and long non-coding RNA dysregulation with the clinical response to immune checkpoint blockade in cutaneous metastatic melanoma. Biomedicines. 2022;10:2419.
Gao Y, Zhou Y, Wei L, Feng Z, Chen Y, Liu P, et al. Hsa_Circ_0066351 acts as a prognostic and immunotherapeutic biomarker in colorectal cancer. Front Immunol. 2022;13:927811.
Cui C, Yang J, Li X, Liu D, Fu L, Wang X. Functions and mechanisms of circular RNAs in cancer radiotherapy and chemotherapy resistance. Mol Cancer. 2020;19:58.
Błach J, Wojas-Krawczyk K, Nicoś M, Krawczyk P. Failure of immunotherapy—the molecular and immunological origin of immunotherapy resistance in lung cancer. Int J Mol Sci. 2021;22:9030.
Liu Z, Wang T, She Y, Wu K, Gu S, Li L, et al. N6-methyladenosine-modified circIGF2BP3 inhibits CD8+ T-cell responses to facilitate tumor immune evasion by promoting the deubiquitination of PD-L1 in non-small cell lung cancer. Mol Cancer. 2021;20:105.
Ge W, Chi H, Tang H, Xu J, Wang J, Cai W, et al. Circular RNA CELF1 drives immunosuppression and anti-PD1 therapy resistance in non-small cell lung cancer via the miR-491-5p/EGFR axis. Aging. 2021;13:24560–79.
Gao J, Zhang L-X, Ao Y-Q, Jin C, Zhang P-F, Wang H, et al. Elevated circASCC3 limits antitumor immunity by sponging miR-432–5p to upregulate C5a in non-small cell lung cancer. Cancer Lett. 2022;543:215774.
Zhang P-F, Pei X, Li K-S, Jin L-N, Wang F, Wu J, et al. Circular RNA circFGFR1 promotes progression and anti-PD-1 resistance by sponging miR-381-3p in non-small cell lung cancer cells. Mol Cancer. 2019;18:179.
Luo Y-H, Yang Y-P, Chien C-S, Yarmishyn AA, Adekunle Ishola A, Chien Y, et al. Circular RNA hsa_circ_0000190 facilitates the tumorigenesis and immune evasion by upregulating the expression of soluble PD-L1 in non-small-cell lung cancer. Int J Mol Sci. 2021;23:64.
Li C, Zhang J, Yang X, Hu C, Chu T, Zhong R, et al. hsa_circ_0003222 accelerates stemness and progression of non-small cell lung cancer by sponging miR-527. Cell Death Dis. 2021;12:807.
Wu J, Zhu M-X, Li K-S, Peng L, Zhang P-F. Circular RNA drives resistance to anti-PD-1 immunotherapy by regulating the miR-30a-5p/SOX4 axis in non-small cell lung cancer. Cancer Drug Resist. 2022;5:261–70. Available from: https://cdrjournal.com/article/view/4594.
Chen S-W, Zhu S-Q, Pei X, Qiu B-Q, Xiong D, Long X, et al. Cancer cell-derived exosomal circUSP7 induces CD8+ T cell dysfunction and anti-PD1 resistance by regulating the miR-934/SHP2 axis in NSCLC. Mol Cancer. 2021;20:144.
Zhang P-F, Xu Y-F, Zhang L-X, Wu J, Fan Y. CircRUNX1 Drives Immune Evasion and anti-PD1 Immunotherapy Resistance in Lung Adenocarcinoma by the miR-4739/PCSK9/MHC I Axis [Internet]. In Review; 2021 Aug. Available from: https://www.researchsquare.com/article/rs-835938/v1.
Liu Y, Zhang H, Zhang W, Xiang L, Yin Z, Xu H, et al. circ_0004140 promotes LUAD tumor progression and immune resistance through circ_0004140/miR-1184/CCL22 axis. Cell Death Discov. 2022;8:181.
Gao J, Ao Y-Q, Zhang L-X, Deng J, Wang S, Wang H-K, et al. Exosomal circZNF451 restrains anti-PD1 treatment in lung adenocarcinoma via polarizing macrophages by complexing with TRIM56 and FXR1. J Exp Clin Cancer Res CR. 2022;41:295.
Zhang L-X, Gao J, Long X, Zhang P-F, Yang X, Zhu S-Q, et al. The circular RNA circHMGB2 drives immunosuppression and anti-PD-1 resistance in lung adenocarcinomas and squamous cell carcinomas via the miR-181a-5p/CARM1 axis. Mol Cancer. 2022;21:110.
Shen W, Chen Y, Lei P, Sheldon M, Sun Y, Yao F, et al. Immunotherapeutic approaches for treating hepatocellular carcinoma. Cancers. 2022;14:5013.
Huang X-Y, Zhang P-F, Wei C-Y, Peng R, Lu J-C, Gao C, et al. Circular RNA circMET drives immunosuppression and anti-PD1 therapy resistance in hepatocellular carcinoma via the miR-30-5p/snail/DPP4 axis. Mol Cancer. 2020;19:92.
Lu J-C, Zhang P-F, Huang X-Y, Guo X-J, Gao C, Zeng H-Y, et al. Amplification of spatially isolated adenosine pathway by tumor–macrophage interaction induces anti-PD1 resistance in hepatocellular carcinoma. J Hematol Oncol. 2021;14:200.
Zhang P-F, Gao C, Huang X-Y, Lu J-C, Guo X-J, Shi G-M, et al. Cancer cell-derived exosomal circUHRF1 induces natural killer cell exhaustion and may cause resistance to anti-PD1 therapy in hepatocellular carcinoma. Mol Cancer. 2020;19:110.
Hu Z, Chen G, Zhao Y, Gao H, Li L, Yin Y, et al. Exosome-derived circCCAR1 promotes CD8+ T-cell dysfunction and anti-PD1 resistance in hepatocellular carcinoma. Mol Cancer. 2023;22:55.
Cai J, Chen Z, Zhang Y, Wang J, Zhang Z, Wu J, et al. CircRHBDD1 augments metabolic rewiring and restricts immunotherapy efficacy via m6A modification in hepatocellular carcinoma. Mol Ther Oncolytics. 2022;24:755–71.
Ye R, Lu X, Liu J, Duan Q, Xiao J, Duan X, et al. CircSOD2 contributes to tumor progression, immune evasion and anti-PD-1 resistance in hepatocellular carcinoma by targeting miR-497-5p/ANXA11 axis. Biochem Genet. 2023;61:597–614.
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2022. CA Cancer J Clin. 2022;72:7–33.
Weng J, Li S, Zhu Z, Liu Q, Zhang R, Yang Y, et al. Exploring immunotherapy in colorectal cancer. J Hematol Oncol. 2022;15:95.
Liu Z, Zheng N, Li J, Li C, Zheng D, Jiang X, et al. N6-methyladenosine-modified circular RNA QSOX1 promotes colorectal cancer resistance to anti-CTLA-4 therapy through induction of intratumoral regulatory T cells. Drug Resist Updat. 2022;65:100886.
Crispen PL, Kusmartsev S. Mechanisms of immune evasion in bladder cancer. Cancer Immunol Immunother. 2020;69:3–14.
Lv J, Li K, Yu H, Han J, Zhuang J, Yu R, et al. HNRNPL induced circFAM13B increased bladder cancer immunotherapy sensitivity via inhibiting glycolysis through IGF2BP1/PKM2 pathway. J Exp Clin Cancer Res CR. 2023;42:41.
Sun J, Zhang H, Wei W, Xiao X, Huang C, Wang L, et al. Regulation of CD8+ T cells infiltration and immunotherapy by circMGA/HNRNPL complex in bladder cancer. Oncogene. 2023;42:1247–62. Available from: https://www.nature.com/articles/s41388-023-02637-2.
Liu D, Lin J-R, Robitschek EJ, Kasumova GG, Heyde A, Shi A, et al. Evolution of delayed resistance to immunotherapy in a melanoma responder. Nat Med. 2021;27:985–92.
Huang AC, Zappasodi R. A decade of checkpoint blockade immunotherapy in melanoma: understanding the molecular basis for immune sensitivity and resistance. Nat Immunol. 2022;23:660–70.
Wei C-Y, Zhu M-X, Lu N-H, Liu J-Q, Yang Y-W, Zhang Y, et al. Circular RNA circ_0020710 drives tumor progression and immune evasion by regulating the miR-370-3p/CXCL12 axis in melanoma. Mol Cancer. 2020;19:84.
dos Santos LV, Abrahão CM, William WN. Overcoming resistance to immune checkpoint inhibitors in head and neck squamous cell carcinomas. Front Oncol. 2021;11:596290.
Nallasamy P, Nimmakayala RK, Parte S, Are AC, Batra SK, Ponnusamy MP. Tumor microenvironment enriches the stemness features: the architectural event of therapy resistance and metastasis. Mol Cancer. 2022;21:225.
Jia L, Wang Y, Wang C. circFAT1 promotes cancer stemness and immune evasion by promoting STAT3 activation. Adv Sci. 2021;8:2003376.
Chen D-L, Sheng H, Zhang D-S, Jin Y, Zhao B-T, Chen N, et al. The circular RNA circDLG1 promotes gastric cancer progression and anti-PD-1 resistance through the regulation of CXCL12 by sponging miR-141-3p. Mol Cancer. 2021;20:166.
Misir S, Wu N, Yang BB. Specific expression and functions of circular RNAs. Cell Death Differ. 2022;29:481–91.
Zhang X, Zhang J, Liu Q, Zhao Y, Zhang W, Yang H. Circ-CUX1 accelerates the progression of neuroblastoma via miR-16-5p/DMRT2 axis. Neurochem Res. 2020;45:2840–55.
Chen L, Nan A, Zhang N, Jia Y, Li X, Ling Y, et al. Circular RNA 100146 functions as an oncogene through direct binding to miR-361-3p and miR-615-5p in non-small cell lung cancer. Mol Cancer. 2019;18:13.
Zhang L, Song X, Chen X, Wang Q, Zheng X, Wu C, et al. Circular RNA CircCACTIN promotes gastric cancer progression by sponging MiR-331-3p and regulating TGFBR1 expression. Int J Biol Sci. 2019;15:1091–103.
Zhang P-F, Wei C-Y, Huang X-Y, Peng R, Yang X, Lu J-C, et al. Circular RNA circTRIM33–12 acts as the sponge of MicroRNA-191 to suppress hepatocellular carcinoma progression. Mol Cancer. 2019;18:105.
Vo JN, Cieslik M, Zhang Y, Shukla S, Xiao L, Zhang Y, et al. The landscape of circular RNA in cancer. Cell. 2019;176:869–881.e13.
Liang W-C, Wong C-W, Liang P-P, Shi M, Cao Y, Rao S-T, et al. Translation of the circular RNA circβ-catenin promotes liver cancer cell growth through activation of the Wnt pathway. Genome Biol. 2019;20:84.
Wang T, Liu Z, She Y, Deng J, Zhong Y, Zhao M, et al. A novel protein encoded by circASK1 ameliorates gefitinib resistance in lung adenocarcinoma by competitively activating ASK1-dependent apoptosis. Cancer Lett. 2021;520:321–31.
Zhao W, Cui Y, Liu L, Qi X, Liu J, Ma S, et al. Splicing factor derived circular RNA circUHRF1 accelerates oral squamous cell carcinoma tumorigenesis via feedback loop. Cell Death Differ. 2020;27:919–33.
Zhao W, Liu J, Wu J, Ma X, Wang X, Zhang L, et al. High-throughput microarray reveals the epitranscriptome-wide landscape of m6A-modified circRNA in oral squamous cell carcinoma. BMC Genom. 2022;23:611.
Mi Z, Zhongqiang C, Caiyun J, Yanan L, Jianhua W, Liang L. Circular RNA detection methods: a minireview. Talanta. 2022;238:123066.
Wang S, Xiong Y, Zhang Y, Wang H, Chen M, Li J, et al. TCCIA: a comprehensive resource for exploring CircRNA in cancer immunotherapy. J Immunother Cancer. 2024;12:e008040.
Li S, Li X, Xue W, Zhang L, Yang L-Z, Cao S-M, et al. Screening for functional circular RNAs using the CRISPR–Cas13 system. Nat Methods. 2021;18:51–9.
Zhong J, Yang X, Chen J, He K, Gao X, Wu X, et al. Circular EZH2-encoded EZH2-92aa mediates immune evasion in glioblastoma via inhibition of surface NKG2D ligands. Nat Commun. 2022;13:4795.
Xu Y, Zhang S, Liao X, Li M, Chen S, Li X, et al. Circular RNA circIKBKB promotes breast cancer bone metastasis through sustaining NF-κB/bone remodeling factors signaling. Mol Cancer. 2021;20:98.
Pandey PR, Young KH, Kumar D, Jain N. RNA-mediated immunotherapy regulating tumor immune microenvironment: next wave of cancer therapeutics. Mol Cancer. 2022;21:58.
Loan Young T, Chang Wang K, James Varley A, Li B. Clinical delivery of circular RNA: Lessons learned from RNA drug development. Adv Drug Deliv Rev. 2023;197:114826.
Li H, Peng K, Yang K, Ma W, Qi S, Yu X, et al. Circular RNA cancer vaccines drive immunity in hard-to-treat malignancies. Theranostics. 2022;12:6422–36.
Shen Z, Zhou L, Zhang C, Xu J. Reduction of circular RNA Foxo3 promotes prostate cancer progression and chemoresistance to docetaxel. Cancer Lett. 2020;468:88–101.
Guo Z, Zhang Y, Xu W, Zhang X, Jiang J. Engineered exosome-mediated delivery of circDIDO1 inhibits gastric cancer progression via regulation of MiR-1307-3p/SOCS2 Axis. J Transl Med. 2022;20:326.
Wang X, Zhang H, Yang H, Bai M, Ning T, Deng T, et al. Exosome-delivered circRNA promotes glycolysis to induce chemoresistance through the miR-122-PKM2 axis in colorectal cancer. Mol Oncol. 2020;14:539–55.
Tan S, Gou Q, Pu W, Guo C, Yang Y, Wu K, et al. Circular RNA F-circEA produced from EML4-ALK fusion gene as a novel liquid biopsy biomarker for non-small cell lung cancer. Cell Res. 2018;28:693–5.
Wang J, Zhao X, Wang Y, Ren F, Sun D, Yan Y, et al. circRNA-002178 act as a ceRNA to promote PDL1/PD1 expression in lung adenocarcinoma. Cell Death Dis. 2020;11:32.
Li L, Zhang Q, Lian K. Circular RNA circ_0000284 plays an oncogenic role in the progression of non-small cell lung cancer through the miR-377-3p-mediated PD-L1 promotion. Cancer Cell Int. 2020;20:247.
Hong W, Xue M, Jiang J, Zhang Y, Gao X. Circular RNA circ-CPA4/ let-7 miRNA/PD-L1 axis regulates cell growth, stemness, drug resistance and immune evasion in non-small cell lung cancer (NSCLC). J Exp Clin Cancer Res CR. 2020;39:149.
Lei J, Zhu J, Hui B, Jia C, Yan X, Jiang T, et al. Circ-HSP90A expedites cell growth, stemness, and immune evasion in non-small cell lung cancer by regulating STAT3 signaling and PD-1/PD-L1 checkpoint. Cancer Immunol Immunother. 2023;72:101–24.
Li J, Xu J, Wu G, Ren Y, Wang X, Zhang Q. Circular RNA hsa_circ_0068252 functions in cisplatin resistance and immune response via miR-1304-5p/PD-L1 axis in non-small cell lung cancer. Chemotherapy. 2022;67:223–33.
Yang J, Jia Y, Wang B, Yang S, Du K, Luo Y, et al. Circular RNA CHST15 sponges miR-155-5p and miR-194-5p to promote the immune escape of lung cancer cells mediated by PD-L1. Front Oncol. 2021;11:595609.
Tian Q, Wu T, Zhang X, Xu K, Yin X, Wang X, et al. Immunomodulatory functions of the circ_001678/miRNA-326/ZEB1 axis in non-small cell lung cancer via the regulation of PD-1/PD-L1 pathway. Hum Mol Genet. 2022;31:4094–106.
Niu R, Li D, Chen J, Zhao W. Circ_0014235 confers Gefitinib resistance and malignant behaviors in non-small cell lung cancer resistant to Gefitinib by governing the miR-146b-5p/YAP/PD-L1 pathway. Cell Cycle. 2022;21:86–100.
Zhang X, Xu L, Wang F. Hsa_circ_0020397 regulates colorectal cancer cell viability, apoptosis and invasion by promoting the expression of the miR-138 targets TERT and PD-L1: Circ_0020397 regulates growth in colorectal cancer cells by sequestering miR-138. Cell Biol Int. 2017;41:1056–64.
Tanaka E, Miyakawa Y, Kishikawa T, Seimiya T, Iwata T, Funato K, et al. Expression of circular RNA CDR1‑AS in colon cancer cells increases cell surface PD‑L1 protein levels. Oncol Rep. 2019;42:1459–66. Available from: http://www.spandidos-publications.com/10.3892/or.2019.7244.
Xu Y-J, Zhao J-M, Gao C, Ni X-F, Wang W, Hu W-W, et al. Hsa_circ_0136666 activates Treg-mediated immune escape of colorectal cancer via miR-497/PD-L1 pathway. Cell Signal. 2021;86:110095.
Wu F, Sun G, Zheng W, Tang W, Cheng Y, Wu L, et al. circCORO1C promotes the proliferation and metastasis of hepatocellular carcinoma by enhancing the expression of PD‐L1 through NF‐κB pathway. J Clin Lab Anal. 2021;35:e24003.
Chen Z-Q, Zuo X-L, Cai J, Zhang Y, Han G-Y, Zhang L, et al. Hypoxia-associated circPRDM4 promotes immune escape via HIF-1α regulation of PD-L1 in hepatocellular carcinoma. Exp Hematol Oncol. 2023;12:17.
Zhang D, Fu Z, Guo Y, Guo F, Wan Y, Guan G. Circ_0000052/miR‐382‐3p axis induces PD‐L1 expression and regulates cell proliferation and immune evasion in head and neck squamous cell carcinoma. J Cell Mol Med. 2023;27:113–26.
Ge J, Wang J, Xiong F, Jiang X, Zhu K, Wang Y, et al. Epstein–Barr virus-encoded circular RNA CircBART2.2 promotes immune escape of nasopharyngeal carcinoma by regulating PD-L1. Cancer Res. 2021;81:5074–88.
Huang L, Ma J, Cui M. Circular RNA hsa_circ_0001598 promotes programmed death-ligand-1-mediated immune escape and trastuzumab resistance via sponging miR-1184 in breast cancer cells. Immunol Res. 2021;69:558–67.
Li J, Dong X, Kong X, Wang Y, Li Y, Tong Y, et al. Circular RNA hsa_circ_0067842 facilitates tumor metastasis and immune escape in breast cancer through HuR/CMTM6/PD-L1 axis. Biol Direct. 2023;18:48.
Dong L, Huang J, Gao X, Du J, Wang Y, Zhao L. CircPCBP2 promotes the stemness and chemoresistance of DLBCL via targeting miR‐33a/b to disinhibit PD‐L1. Cancer Sci. 2022;113:2888–903.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (82274142 and 82374088), and the Sichuan Science and Technology Program (2023NSFSC0681 and 2022ZYD0092).
Author information
Authors and Affiliations
Contributions
Fangyi Long designed and guided the review. Yu Ma and Ting Wang wrote and edited the manuscript. Xudong Zhang and Pinghan Wang helped with reference collection and draw the figures. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by: George Calin
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ma, Y., Wang, T., Zhang, X. et al. The role of circular RNAs in regulating resistance to cancer immunotherapy: mechanisms and implications. Cell Death Dis 15, 312 (2024). https://doi.org/10.1038/s41419-024-06698-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-024-06698-3
- Springer Nature Limited
This article is cited by
-
Circular RNAs in tumor immunity and immunotherapy
Molecular Cancer (2024)
-
Epigenetic modification of ferroptosis by non-coding RNAs in cancer drug resistance
Molecular Cancer (2024)
-
Circular RNA in cancer
Nature Reviews Cancer (2024)
-
Unraveling the noncoding RNA landscape in glioblastoma: from pathogenesis to precision therapeutics
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)