Abstract
The recombinase RAD51 catalyzes the DNA strand exchange reaction in homologous recombination (HR) during both mitosis and meiosis. However, the physiological role of RAD51 during spermatogenesis remains unclear since RAD51 null mutation is embryonic lethal in mice. In this study, we generated a conditional knockout mouse model to study the role of RAD51 in spermatogenesis. Conditional disruption of RAD51 in germ cells by Vasa-Cre led to spermatogonial loss and Sertoli cell-only syndrome. Furthermore, tamoxifen-inducible RAD51 knockout by UBC-CreERT2 confirmed that RAD51 deletion led to early spermatogenic cells loss and apoptosis. Notably, inducible knockout of RAD51 in adult mice caused defects in meiosis, with accumulated meiotic double-strand breaks (DSBs), reduced numbers of pachytene spermatocytes and less crossover formation. Our study revealed an essential role for Rad51 in the maintenance of spermatogonia as well as meiotic progression in mice.
Similar content being viewed by others
Introduction
Spermatogenesis is a complex process involving mitotic cell division, meiosis, and the process of spermiogenesis [1]. In the mouse, the first wave of spermatogenesis begins shortly after birth when gonocytes resume mitotic division [2]. Gonocytes give rise to either undifferentiated spermatogonia or differentiating spermatogonia, which differentiate into meiotic spermatocytes [3]. Spermatocytes undergo two meiotic divisions to form haploid round spermatids, which are transformed into spermatozoa [4] Meiosis is an essential step of spermatogenesis, during prophase I of meiosis, homologous chromosomes pair, synapse, and exchange their genetic material through homologous recombination (HR) [5].
HR is essential for high-fidelity DNA repair during mitotic proliferation and meiosis [6]. Mitotic HR promotes genome stability through the precise repair of DNA double-strand breaks (DSBs) [7]. Meiotic HR is a specialized process that involves homologous chromosome pairing and strand exchange to guarantee proper chromosome segregation and genetic diversity [8]. Meiotic recombination is initiated by DSBs generated by the topoisomerase like DNA transesterase SPO11 [9]. The dynamics of γH2AX distribution is one of the major characteristics of meiosis and is commonly used as a marker for meiotic progression [10].
The repair of meiotic DSBs by HR assures the exchange of genetic material between parental haplotypes in the germline [11]. ATM kinase is needed to limit the number of meiotic DSBs and is required for complete DNA repair [12]. ATM is also essential for proper crossover formation in mouse spermatocytes [13]. After DSB formation in meiotic chromosomes, the ends surrounding the breaks are resected to create 3′ single-stranded DNA (ssDNA) overhangs [14]. The emerging ssDNA is bound and protected by heterotrimeric replication protein A (RPA) to prevent degradation and secondary structure formation [15]. Subsequently, RAD51 and DMC1 are recruited and promote the removal of RPA from the ssDNA and facilitate the invasion of 3′-extended strand into the duplex of the homologue for successful recombination and synapsis formation [16, 17].
RAD51 and DMC1 are two RecA-like recombinases in eukaryotes. DMC1 is detected in leptotene-to-zygotene spermatocytes in mice [17]. In Dmc1-deficient mice, germ cells arrest in the early zygotene stage of meiosis and then undergo apoptosis [18]. Rad51 is a conserved eukaryotic protein that catalyzes the HR repair of DNA DSBs that occur during mitosis and meiosis [19]. Human and mouse RAD51 are almost identical and are highly homologous (83%) with the yeast rad51 protein [20]. RAD51 is transcribed at a high level in the thymus, spleen, testis, and ovary and at a lower level in the brain [21]. Homozygous knockout of RAD51 in mice results in early embryonic lethality and heterozygous mice appear normal and are fertile [22]. Knockdown of RAD51 by siRNAs in the seminiferous tubules of mouse testis led to reduced numbers of spermatocytes and reduced crossover formation [23]. However, the physiological role of RAD51 during spermatogenesis remains unclear.
In this study, we generated a conditional knockout mouse model to study the role of RAD51 in spermatogenesis. Conditional disruption of RAD51 in germ cells by Vasa-Cre led to sterility due to complete germ cell loss. We further generated a tamoxifen-inducible RAD51 knockout model by UBC-CreERT2 in adult mice. We demonstrated that RAD51 deficient male mice led to impaired DSB repair and reduced formation of crossovers during meiosis.
Results
RAD51 is highly expressed in the mouse testis, predominantly in germ cells
To understand the expression and physiological role of RAD51, we measured the expression of RAD51 in different tissues of adult mice by western blot. RAD51 protein was shown to be highly expressed in the testis, spleen, and ovary (Fig. 1A). We next analyzed Rad51 expression in testes at different stages of mouse spermatogenesis using publicly available RNA-seq data [24]. We found that Rad51 had a low level of expression between E15.5 and P0 and had significantly increased levels after birth (Fig. S1A). We further performed immunoblotting to determine RAD51 level in testis tissues and found RAD51 was expressed at a moderate level from postnatal day 6 (PD6) to PD12 and then significantly increased at PD14 (Fig. 1B). Subsequently, we isolated germ cells by the differential adhesion method as described previously [25]; the enrichment efficiency was confirmed by using germ cell and somatic cell markers detected by qPCR (Fig. S1B). Then, we compared RAD51 expression between germ cells and somatic cells by qPCR and western blotting and the results revealed that RAD51 was more highly expressed in germ cells than in somatic cells (Fig. 1C, D). We also performed immunohistochemistry staining in testes and found RAD51 was highly expressed in spermatogonia and less expressed in spermatocytes (Fig. 1E, and Fig. S1C). Further, we immunostained RAD51(green) and SYCP3 (red) in meiotic chromosome spreads and found that RAD51 was expressed in leptotene and zygotene spermatocytes, and the expression diminished in mid-pachytene spermatocytes (Fig. 1F). Thus, RAD51 is expressed in mitotic spermatogonia and early meiotic prophase spermatocytes.
A Western blotting was performed to detect RAD51 expression in different tissues of adult mice and GAPDH was used as the loading control. B RAD51 expression in different developmental stages of spermatogenesis was measured by Western blotting. C, D Primary isolation of spermatogenic cells and somatic cells from PD9 mouse testes. RT-qPCR (C) and western blotting (D) were conducted to test the expression of RAD51 and germ cell markers MVH and DAZL in the fraction of spermatogenic cells (FSPCs) and the fraction of somatic cells (FSCs). Data are presented as means ± S.D. (n = 3 biologically independent samples for RT-qPCR assay). ***p < 0.001. E Immunohistochemistry staining for RAD51 was analyzed in PD62 mouse testes. (red triangle: RAD51 positive cells; L: leptotene spermatocyte; P: pachytene spermatocyte; ES: elongating spermatid). Scale bar is 50 μm. F Immunofluorescence of RAD51 (green fluorescence) and SYCP3 (red fluorescence) on chromosome spreads in wild type. Scale bar, 5 μm.
Conditional disruption of RAD51 in germ cells results in sterility
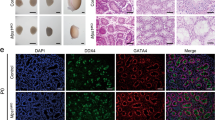
To explore the function of RAD51 in spermatogenesis, we generated a conditional Rad51 null allele by flanking exons 3–4 with loxP sites (Fig. 2A). Rad51 was specifically inactivated in germ cells by crossing Rad51-floxed mice with Vasa-Cre line, which expresses CRE recombinase in male germ cells from embryonic day 15.5 [26]. The genotype of the conditional knockout mice was confirmed by PCR of tail biopsy DNA (Fig. 2B). The knockout efficiency was determined by western blotting and IHC in testes (Fig. 2C and Fig. S2A), and the results showed that RAD51 protein was absent in Rad51-VKO (Rad51F/-; Vasa-Cre) compared to wild-type mouse germ cells. Fertility test revealed that no pups were obtained when adult Rad51-VKO male mice mated with wild-type fertile female mice for at least 6 months (Table S2). We then analyzed the PD40 testis size and found that Rad51-VKO mice had smaller testes than their littermate controls (Fig. 2D). To obtain more information about this phenotype, we measured the testes weight at different timepoints from PD8 to PD40 and found that homozygous Rad51-VKO males had dramatically reduced testis weight compared to their littermate controls from PD10 to PD40, whereas no significant differences were observed in PD8 mice (Fig. 2E). Immunohistochemical staining of mouse seminiferous tubules showed that RAD51 protein decreased in PD8 testes of Rad51-VKO (Fig. 2F). Consistently, Hematoxylin staining demonstrated germ cell loss in PD12 and PD60 Rad51-VKO mice compared to their littermate controls (Fig. 2G, H and Fig. S2B). In addition, co-immunofluorescence staining of the germ cell marker GCNA (green) and RAD51 revealed that GCNA-positive cells were found both in embryonic day (E)18.5 control and Rad51-VKO testes (Fig. S2C), indicating germ cells were still present in Rad51-VKO testes at E18.5. These data indicate that RAD51 is essential for normal spermatogenesis and male fertility.
A A schematic diagram illustrates the generation of the Rad51 conditional knockout mice. The strategy was to insert loxp sites to flank exons 3 and 4 of the mouse Rad51 gene. Cre recombinase mediated the removal of the floxed sequence to create a null allele. B Genotype identification of Rad51 conditional knockout mice was analyzed by PCR of tail biopsy DNA. C The knockout efficiency was determined by western blotting in PD40 testes of RAD51-VKO mice compared to their littermate controls. D The size of testes was compared in RAD51-VKO males compared to their littermate controls from PD40 mice. E The testes weights at PD8 to PD40 were measured in RAD51-VKO mice and their littermate controls (n = 5). Data are presented as means ± S.D. *p < 0.05, **p < 0.01, ***p < 0.001. F Immunohistochemical staining of mouse seminiferous tubules shows the efficiency of PD8 RAD51-VKO mice and their littermate control. Scale bars are 20 μm. G Haematoxylin staining was conducted in seminiferous tubules to analyze the number and type of testicular cells in PD12 RAD51-VKO mice and their littermate controls. Scale bar, 50 μm. (red triangles: Sertoli cell; green triangles: spermatogonia; yellow triangles: leptotene spermatocyte; gray triangles: zygotene spermatocytes). H Statistics of label the different types of germ and somatic cell types to A.
RAD51 deletion led to Sertoli cell-only phenotype in male mice
We then started to investigate the underlying mechanism for the lack of germ cells in RAD51 deficient mice. Transmission electron microscopy (TEM) was used to examine the detailed structure of the seminiferous tubules at PD8. Rad51-deficient testes exhibited a marked depletion of germ cells in the seminiferous tubules in comparison with their littermate controls (Fig. 3A, B). Next, we performed immunofluorescence experiments to examine the early spermatogonial stem cells (SSCs) marker PLZF, the synaptonemal complex (SC) marker SYCP3, and the DNA damage marker γH2AX. No PLZF-positive cells were observed in the seminiferous tubules of Rad51-VKO testes at PD10 (Fig. S3A). Meanwhile, no SYCP3 or γH2AX-positive cells were observed in the seminiferous tubules of RAD51-VKO testes at PD8 (Fig. S3B). In addition, immunofluorescence staining of the Sertoli cell marker SOX9 revealed that the seminiferous tubules of RAD51 VKO testes contained only Sertoli cells (Fig. 3C, D and Fig. S3C, S3D). Immunohistochemical staining of SOX9 further revealed that the expression level of SOX9 and the number of Sertoli cells were increased in RAD51VKO mice (Fig. 3E, F and Fig. S3E–G). Thus, ablation of RAD51 results in sertoli cell-only phenotypes, indicating a pivotal role of RAD51 in spermatogenesis.
A Transmission electron microscopy (TEM) was performed to examine the detailed structure of seminiferous tubules at PD8 in VKO mice and their littermate controls (red triangles: spermatogonia; red stars: Sertoli cells). Scale bar is 10 μm. B Statistics of the germ cells to (A). Data are presented as means ± S.D. ***p < 0.001. C Immunofluorescence staining of the Sertoli cell marker SOX9 (green fluorescence) and RAD51 (red fluorescence) was measured in the seminiferous tubules of RAD51-VKO and control testes at PD8. Scale bar is 10 μm. D Statistics of the RAD51 positive cells to (C). Data are presented as means ± S.D. ***p < 0.001. E, F Immunohistochemical staining of SOX9 was analyzed in the seminiferous tubules of VKO and control mice at PD8 and PD40. SOX9 positive cells were counted using ImageJ. Scale bar is 20 μm. Data are presented as means ± S.D. *p < 0.05, **p < 0.01, ***p < 0.001.
Inducible knockout of RAD51 causes spermatogonia loss and apoptosis
Cre-ERT2 encodes a Cre fused to a mutant estrogen ligand-binding domain (ERT2) that requires tamoxifen for activity [27]. UBC-CreERT2 mice express a Cre-ERT2 fusion gene under the control of the ubiquitin C (UBC) promoter. When UBC-Cre-ERT2 mice are crossed with mice containing a loxP-flanked alleles of interest genes. Tamoxifen treatment could induce global knockout in the adult mouse at any age [26]. UBC-Cre ERT2 mice have been used to study DDX5 in spermatogenesis [28]. To better characterize the effect of RAD51 on spermatogenesis, we crossed UBC-Cre ERT2 mice with Rad51flox/flox mice to generate a tamoxifen-inducible Rad51 knockout mouse model (Rad51F/F; UBC-Cre ERT2, Rad51-UKO). We then treated 8-week-olds male UKO (Rad51TAM-KO) and control mice with tamoxifen for five consecutive days and tissues were harvested at 8 days after the final injection. Subsequently, we measured the weight of the testes and found that the weight of Rad51-UKO testes was reduced to 80% of the weight of control testes (Fig. 4A, B). The knockout efficiency was then determined by qPCR, western blotting and IHC (Fig. 4C–E, and Fig. S4A). In addition, haematoxylin staining of seminiferous tubules showed early spermatogenic cell loss in the testes of RAD51-UKO mice at PD70 (Fig. 4F). In accordance with the results from VKO mice, immunohistochemical and immunofluorescence staining showed that PLZF-positive cells were lost in the seminiferous tubules of RAD51-UKO mice compared to controls at PD70 (Fig. 4G and Fig. S4B). To further investigate this phenotype, we treated PD11 RAD51-UKO and control mice with tamoxifen for 3 consecutive days and compared testis sections with haematoxylin staining at PD14. Unlike control testes with the normal progression of spermatocytes, in which pachytene spermatocytes could be observed, no meiotic cells were found in the testes of RAD51-UKO mice, only mitotic cells were observed (Fig. 4H). Furthermore, immunostaining of the meiotic markers SYCP3 and γH2AX confirmed the lack of SYCP3 and γH2AX expression in germ cells with RAD51 deletion (Fig. 4I, Fig. S4C). We speculated that the depletion of spermatogenic cells might be due to apoptosis. The TUNEL assay showed an increase in the number of apoptotic spermatocytes in the seminiferous tubules of RAD51-UKO mice compared to controls at PD14 (Fig. 4J, K) or PD70 (Fig. S4D, E). These data reveal that RAD51 deletion leads to early spermatogenic cell loss and apoptosis.
A, B The size (A) and weight (B) of testes were compared in mice with tamoxifen-induced knockout of Rad51 (Rad51TAM-KO, UKO) and control mice at PD70 (n = 5). C, D RT-qPCR (C) and Western blotting (D) were conducted to determine the knockout efficiency of RAD51-UKO mice compared to control mice in testes. Tubulin was used as the loading control. Data are presented as means ± S.D. **p < 0.01. E Immunohistochemical staining of RAD51 in seminiferous tubules was used to confirm the knockout efficiency at PD70. (P: pachytene spermatocyte; RS: round spermatid; ES: elongating spermatid). Scale bar is 50 μm. F Haematoxylin staining was conducted in seminiferous tubules to analyze the number and type of testicular cells in RAD51-UKO and control testes at PD70. (P: pachytene spermatocyte; RS: round spermatid; ES: elongating spermatid). Scale bar is 50 μm. G Immunohistochemical staining of PLZF was applied to analyze in the seminiferous tubules of RAD51-UKO mice at PD70. (A: spermatogonia; P: pachytene spermatocyte; RS: round spermatid; ES: elongating spermatid). Scale bar is 50 μm. H Haematoxylin staining was carried out in the seminiferous tubules of RAD51-UKO and control mouse testes at PD14. (red triangle: spermatogonia; red star: pachytene spermatocytes; red arrow: type B spermatogonia; green arrow: mitotic cells). Scale bar, 50 μm. I Immunofluorescence staining of SYCP3 and γH2AX was applied to the seminiferous tubules of RAD51-UKO and control testes at PD14. Scale bar, 20 μm. J A TUNEL assay was applied to control and RAD51-UKO testes. Cells strained green are TUNEL-positive cells and DNA was stained with DAPI. Scale bar, 20 µm. K Statistics of a number of TUNEL-positive cells per tubule in control and RAD51-UKO testes at PD14 tubules were counted. Student’s t-test, error bars indicate SEM. ***P < 0.001.
Inducible knockout of Rad51 leads to meiosis defects
While RAD51 is believed to be essential for both mitotic and meiotic recombination. However, the role of RAD51 in meiosis remains poorly characterized in mice due to the embryonic lethality of the RAD51 homozygous knockout mice. We sought to investigate the functions of RAD51 in male meiosis using a tamoxifen-inducible Rad51 knockout mouse model treated with tamoxifen as described above. We immunostained the testis sections for the meiotic marker SYCP3 and performed a quantitative analysis of spermatocytes at different meiotic stages in RAD51-UKO and control mice. Interestingly, compared to the profile in control mice, we observed a significant increase in the proportion of diplotene spermatocytes (from 48.83% to 82.74%) and a decrease in the proportion of pachytene spermatocytes (from 41.37 to 12.57%) in RAD51-UKO mice (Fig. 5A, B and Fig. S5A). To further investigate the role of RAD51 in the repair of meiotic DSBs, we performed immunostaining of γH2AX, a marker of meiotic DSBs, on spread spermatocytes. We observed the accumulation of γH2AX foci on the axes of autosomal chromosomes in pachytene spermatocytes in RAD51-UKO mice (Fig. 5C, D and Fig. S5B). Similarly, we observed a significant increase of DMC1 foci in the pachytene stage in RAD51-UKO mice compared to control mice (Fig. 5E, F and Fig. S5C). These data indicate that depletion of RAD51 results in an increased incidence of unrepaired DNA breaks. Moreover, we immunostained MLH1 to evaluate crossover maturation in the pachytene stage and found a significant decrease in MLH1 foci in RAD51-UKO mice compared to control mice (Fig. 5G, H and Fig. S5D). These data reveal that RAD51 is critical for DSB repair and meiotic progression.
A Immunofluorescence of SYCP3 expression shown on chromosome spreads from control and RAD51-UKO testes at PD70. Scale bar, 20 μm. B Statistics of each meiotic period corresponding to (A). C, D Immunofluorescence of SYCP3 and γH2AX on chromosome spreads from control and RAD51-UKO testes at PD70 was shown and counted. Scale bar, 5 μm. E, F Immunofluorescence of SYCP3 and DMC1 on chromosome spreads in control and RAD51-UKO testes at PD70 was shown and counted. Scale bar, 5 μm. G, H Immunofluorescence of SYCP3 and MLH1 on chromosome spreads from control and RAD51-UKO testes at PD70 was shown and quantified. Scale bar, 5 μm. Data are presented as means ± S.D.*p < 0.05, **p < 0.01, ***p < 0.001.
Discussion
Rad51 mediates the HR repair both in mitosis and meiosis [29]. However, because of the early embryonic lethality of constitutive RAD51 knockout mice, the importance of RAD51 in spermatogenesis has not been fully elucidated. In this study, we explore the physiological role of RAD51 in spermatogenesis using a conditional knockout mouse model for the first time. We found that RAD51 was highly expressed in germ cells of mouse testis. Conditional disruption of RAD51 in germ cells with Vasa-Cre results in SSCs loss and Sertoli cell-only phenotype. In a tamoxifen-inducible Rad51 knockout mouse model, we observed that RAD51 deletion led to early spermatogenic cells loss and apoptosis. Moreover, we demonstrate that RAD51 ablation results in defective DSB repair and crossover formation of male meiosis.
Rad51 mutations in mice cause early embryo lethality, indicating the essential function of RAD51 in mitotic development [22]. In budding yeast, rad51 mutants show reduced spore formation and viability [30]. Loss of the RAD51 function in mice results in a defect in mitosis that is accompanied by chromosome fragmentation [31]. Disruption of the RAD51 gene in the chicken B-cell line DT40 led to G2/M phase arrest, and eventually dying [32]. However, loss-of-function mutant in the Arabidopsis homolog of RAD51 has no detectable abnormality in mitosis. Our findings showed that RAD51 deletion led to early spermatogenic cells loss and apoptosis, supporting the essential role of RAD51 in mitosis. A previous study reported that RAD51 contributes to G2/M transition in mouse embryonic stem cells [33]. RAD51 is responsible for recombinational repair of DNA damage during mitosis [34]. RAD51 also plays non-repair functions, which stimulates replication forks reversal and stabilizes stalled forks [35]. RAD51 promotes mitotic DNA synthesis and successful chromosome segregation in mitosis [36].
In meiosis, we found that inducible knockout of RAD51 at PD14 (three days post tamoxifen treatment), germ cells failed to enter meiosis as evidenced by histological analysis and the lack of SYCP3 and γH2AX expression (Fig. 4I). Nevertheless, a previous study shows microinjection of siRNAs into seminiferous tubules of mouse testis results in no significant reduction of EdU positive cells in the RAD51 knockdown tubules [23]. The discrepancy may be explained by the incomplete knockdown of RAD51 in mouse testis since heterozygous RAD51 knockout mice are viable and fertile [37]. Both RAD51 and DMC1 are essential for HR during meiosis and they might have distinct and overlapping roles in recombination. In yeast, Rad51 foci form normally in Dmc1 mutant strain whereas the formation of Dmc1 foci is greatly reduced in Rad51 mutant cells [38]. In budding yeast, Dmc1-deficient strain exhibits defects in meiotic recombination and SC formation and cells arrested at the pachytene stage [39]. Similar but not the same, spermatocytes of DMC1 knockout mice are arrested at the early zygotene stage, with the failure of homologous chromosomes to undergo synapsis [18]. DMC1 and RAD51 have distinct spatial localization on ssDNA [14]. In yeast, Rad51 is an accessory factor for the strand exchange activity of Dmc1 during meiosis [19]. However, our study revealed that the inducible knockout of RAD51 led to a significant decrease in the proportion of zygotene and pachytene spermatocytes relative to control mice (Fig. 5A, B). In addition, we found that DMC1 foci were increased in the pachytene stage in RAD51 deficient spermatocytes (Fig. 5E, F), suggesting that RAD51 and DMC1 have concerted functions during meiotic HR in male mice. Our study demonstrates that RAD51 is essential for meiotic recombination in mice. Further studies are needed to elucidate the role of RAD51 and DMC1 in mammalian meiosis.
In summary, we demonstrated that RAD51 is essential for spermatogonial mitosis as well as meiotic recombination in mice. Inactivation of Rad51 causes spermatogonia loss and Sertoli cell-only phenotype. Inducible knockout of Rad51 leads to leads to meiosis defects, with a significant decrease of zygotene and pachytene spermatocytes. Thus, RAD51 is indispensable for the survival of spermatogonia as well as meiotic progression in mice.
Materials and methods
Mice
Rad51-floxed line was generated by flanking exon 3 and 4 of Rad51 with loxP sites (Constructed at Nanjing Model Organism Center). The resulting founder male mice were mated to WT C57BL/6 J (B6) female mice to obtain heterozygous Rad51-floxed mice. Progeny was screened by PCR for germ line transmission of the targeted alleles. All mice described above were maintained in C57BL/6J (B6) background. Rad51- floxed mice (Rad51flox/flox) were then crossed to Vasa-Cre mouse line (Jackson laboratory) [26]. All animal experiments were conducted in accordance with the guidelines in the Animal Care and Use Committee at Shandong University. To generate tamoxifen-inducible Rad51 knockout mice, Rad51flox/flox-UBC-CreERT2 mice (which were obtained from the Jackson Laboratory) were subjected to tamoxifen treatment. Tamoxifen (Sigma, T5648) was dissolved in corn oil at a concentration of 20 mg/ml and injected into the abdominal cavity of 8-week-old male mice at a dose of 4 mg/30 g body weight for five consecutive days and tissues were harvested at 8 days after the final injection [28, 40]. The tamoxifen-inducible Rad51 knockout mice were referred to as Rad51-UKO mice. All the primers for PCR genotyping were listed in Supplementary information, Table S1.
Enrichment of spermatogenic cells
The spermatogenic cells and somatic cells were enriched using a two-step enzymatic digestion process followed by a differential adhesion method as previously described with some modifications [41]. Briefly, after the tunica albuginea was disrupted, testes from postnatal day (PD) nine wild-type mice were transferred into 120 U/ml collagenase type I (Gibco, 17100-017) and incubated at 37 °C for 20 min with gentle shaking every 3–5 min to accelerate testis dissociation. Then, the cell suspension was centrifuged at 1000 rpm. for 5 min, and the pellet was digested with 0.25% trypsin-EDTA (Gibco, 25200-072) at 37 °C for 10 min to dissociate the seminiferous tubules into single cells. The suspension was neutralized with 5 ml DMEM supplemented with 10% FBS and centrifuged at 1000 rpm. for 5 min. The cell pellet was suspended in 8 ml of DMEM medium and seeded in a 10 cm culture dish. After 2–3 h of incubation at 37 °C, the floating and weakly adhering cells were transferred to a new 10 cm dish. The fraction of spermatogenic cells (FSPCs) comprised the floating and weakly adhering cells. The attached cells on the bottom of the dish were collected as the fraction of somatic cells (FSCs). The efficiency of separation was examined using RT-PCR and western blotting analysis.
Histological analysis, immunostaining, and imaging
Testes from control and RAD51-VKO/UKO male mice were isolated and fixed in 4% paraformaldehyde (PFA) overnight at 4 °C for immunostaining. The samples were dehydrated stepwise through an ethanol series (25, 50, 75, 85, 95, and 100% ethanol), embedded in paraffin and sectioned (4 μm). After dewaxing and hydration, the sections were stained with haematoxylin and imaged with an Olympus microscope. For immunostaining, after dewaxing and hydration, the section was placed in boiling EDTA and heated for 15 min for antigen retrieval. After washing with PBS three times, the sections were permeabilized with 0.2% Triton X-100 for 15 min, washed with PBS three times, blocked with 5% BSA for 1 h at room temperature and incubated overnight at 4 °C with primary antibodies diluted in western primary antibody diluent (Beyotime, P0023A). The next day, following three washes with PBS, secondary antibodies were added to the sections and incubated for 1 h at room temperature. The sections were then washed in PBS three times, washed with water several times, and incubated with DAPI (Abcam, ab104139) and sealed with nail polish. The primary antibodies used were as follows: rabbit anti-RAD51 polyclonal antibody (#PA5-27195, Thermo Fisher Scientific, 1:200); mouse anti-PLZF monoclonal antibody (sc-28319, Santa Cruz, 1:200); mouse anti-phospho-histone H2AX (Ser139/Tyr142) antibody (#05-636, Millipore, 1:300); mouse anti-SCP3 monoclonal antibody (sc-74569, Santa Cruz, 1:200); rabbit anti-SCP3 monoclonal antibody (sc-33195, Santa Cruz, 1:200); and rabbit anti-SOX9 monoclonal antibody (ab185966, Abcam, 1:250).
RNA extraction and real-time qPCR
Total RNA was extracted from whole testes using TRIzol reagent (Invitrogen, 15596-026) following the manufacturer’s instructions. After removing the residual genomic DNA, 1 μg of total RNA was reverse-transcribed into cDNA using the HiScript III RT SuperMix for qPCR (+gDNA wiper) (Vazyme, R223-01) according to the manufacturer’s protocol. Real-time RT-PCR was performed using ChamQ SYBR Color qPCR Master Mix (Vazyme, Q411-02) on a Quant Studio 3 (Thermo Fisher). The primers for qPCR assay are listed in Table S1.
Western blotting
Protein samples were prepared using cell lysis buffer for western blotting and IP without inhibitors mixed with a protease inhibitor cocktail (Beyotime, P011) and quantified using a BCA reagent kit (Beyotime, P0012-1). Equal amounts of total protein were separated in a 12% SDS–PAGE gel and transferred onto PVDF membranes. After blocking with 5% nonfat milk for 1 h at room temperature, the membranes were incubated with diluted primary antibodies at 4 °C overnight. After three washes with TBST, the membranes were incubated with horseradish peroxidase-conjugated secondary antibodies (1:10,000, Jackson ImmunoResearch) at room temperature for 1 h. The signals were developed with Pierce ECL Substrate (Thermo Fisher Scientific, #34080). The primary antibodies were as follows: rabbit anti-RAD51 polyclonal antibody (#PA5-27195, Thermo Fisher Scientific, 1:10,000); rabbit anti-DAZL polyclonal antibody (ab34139, Abcam, 1:5000); rabbit anti-DDX4/MVH polyclonal antibody (ab13840, Abcam, 1:5000); mouse anti-GAPDH monoclonal antibody (60004-1-Ig, Proteintech, 1:10,000); and mouse anti-tubulin monoclonal antibody (66031-1-Ig, Proteintech, 1:10,000).
Statistical analysis
The results of all quantitative experiments were based on at least three independent biological samples. All data are presented as means ± SEM. The statistical significance of the differences between the mean values for the different genotypes was measured by Student’s two-tailed t-test with a paired t-test. The data were considered significant when the p value was <0.05.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
de Kretser DM, Loveland KL, Meinhardt A, Simorangkir D, Wreford N. Spermatogenesis. Hum. Reprod. 1998;13:1–8.
Zogbi C, Tesser RB, Encinas G, Miraglia SM, Stumpp T. Gonocyte development in rats: proliferation, distribution and death revisited. Histochem Cell Biol. 2012;138:305–22.
Yoshida S, Sukeno M, Nakagawa T, Ohbo K, Nagamatsu G, Suda T, et al. The first round of mouse spermatogenesis is a distinctive program that lacks the self-renewing spermatogonia stage. Development. 2006;133:1495–505.
Hermo L, Pelletier RM, Cyr DG, Smith CE. Surfing the wave, cycle, life history, and genes/proteins expressed by testicular germ cells. Part 1: background to spermatogenesis, spermatogonia, and spermatocytes. Microsc Res Tech. 2010;73:241–78.
Sepsi A, Schwarzacher T. Chromosome-nuclear envelope tethering - a process that orchestrates homologue pairing during plant meiosis? J Cell Sci. 2020;133:jcs243667
Wild P, Susperregui A, Piazza I, Dorig C, Oke A, Arter M, et al. Network rewiring of homologous recombination enzymes during mitotic proliferation and meiosis. Mol Cell. 2019;75:859–74. e854.
Moynahan ME, Jasin M. Mitotic homologous recombination maintains genomic stability and suppresses tumorigenesis. Nat Rev Mol Cell Biol. 2010;11:196–207.
Ribeiro J, Abby E, Livera G, Martini E. RPA homologs and ssDNA processing during meiotic recombination. Chromosoma. 2016;125:265–76.
Cole F, Keeney S, Jasin M. Evolutionary conservation of meiotic DSB proteins: more than just Spo11. Genes Dev. 2010;24:1201–7.
Jiang H, Gao Q, Zheng W, Yin S, Wang L, Zhong L, et al. MOF influences meiotic expansion of H2AX phosphorylation and spermatogenesis in mice. PLoS Genet. 2018;14:e1007300.
Brick K, Pratto F, Camerini-Otero RD. After the break: DSB end processing in mouse meiosis. Genes Dev. 2020;34:731–2.
Lange J, Pan J, Cole F, Thelen MP, Jasin M, Keeney S. ATM controls meiotic double-strand-break formation. Nature. 2011;479:237–40.
Barchi M, Roig I, Di Giacomo M, de Rooij DG, Keeney S, Jasin M. ATM promotes the obligate XY crossover and both crossover control and chromosome axis integrity on autosomes. PLoS Genet. 2008;4:e1000076.
Hinch AG, Becker PW, Li T, Moralli D, Zhang G, Bycroft C, et al. The Configuration of RPA, RAD51, and DMC1 binding in meiosis reveals the nature of critical recombination intermediates. Mol Cell. 2020;79:689–701. e610
Fanning E, Klimovich V, Nager AR. A dynamic model for replication protein A (RPA) function in DNA processing pathways. Nucleic Acids Res. 2006;34:4126–37.
Shang Y, Huang T, Liu H, Liu Y, Liang H, Yu X, et al. MEIOK21: a new component of meiotic recombination bridges required for spermatogenesis. Nucleic Acids Res. 2020;48:6624–39.
Shinohara A, Shinohara M. Roles of RecA homologues Rad51 and Dmc1 during meiotic recombination. Cytogenet Genome Res. 2004;107:201–7.
Yoshida K, Kondoh G, Matsuda Y, Habu T, Nishimune Y, Morita T. The mouse RecA-like gene Dmc1 is required for homologous chromosome synapsis during meiosis. Mol Cell. 1998;1:707–18.
Cloud V, Chan YL, Grubb J, Budke B, Bishop DK. Rad51 is an accessory factor for Dmc1-mediated joint molecule formation during meiosis. Science. 2012;337:1222–5.
Shinohara A, Ogawa H, Matsuda Y, Ushio N, Ikeo K, Ogawa T. Cloning of human, mouse and fission yeast recombination genes homologous to RAD51 and recA. Nat Genet. 1993;4:239–43.
Pires E, Sung P, Wiese C. Role of RAD51AP1 in homologous recombination DNA repair and carcinogenesis. DNA Repair. 2017;59:76–81.
Tsuzuki T, Fujii Y, Sakumi K, Tominaga Y, Nakao K, Sekiguchi M, et al. Targeted disruption of the Rad51 gene leads to lethality in embryonic mice. Proc Natl Acad Sci USA. 1996;93:6236–40.
Dai J, Voloshin O, Potapova S, Camerini-Otero RD. Meiotic knockdown and complementation reveals essential role of RAD51 in mouse spermatogenesis. Cell Rep. 2017;18:1383–94.
Cardoso-Moreira M, Halbert J, Valloton D, Velten B, Chen C, Shao Y, et al. Gene expression across mammalian organ development. Nature. 2019;571:505–9.
Liu W, Wang F, Xu Q, Shi J, Zhang X, Lu X, et al. BCAS2 is involved in alternative mRNA splicing in spermatogonia and the transition to meiosis. Nat Commun. 2017;8:14182.
Gallardo T, Shirley L, John GB, Castrillon DH. Generation of a germ cell-specific mouse transgenic Cre line, Vasa-Cre. Genesis. 2007;45:413–7.
Zhao J, Nassar MA, Gavazzi I, Wood JN. Tamoxifen-inducible NaV1.8-CreERT2 recombinase activity in nociceptive neurons of dorsal root ganglia. Genesis. 2006;44:364–71.
Legrand JMD, Chan AL, La HM, Rossello FJ, Anko ML, Fuller-Pace FV, et al. DDX5 plays essential transcriptional and post-transcriptional roles in the maintenance and function of spermatogonia. Nat. Commun. 2019;10:2278.
Woo TT, Chuang CN, Wang TF. Correction to: Budding yeast Rad51: a paradigm for how phosphorylation and intrinsic structural disorder regulate homologous recombination and protein homeostasis. Curr Genet. 2021;67:397–8.
Shinohara A, Gasior S, Ogawa T, Kleckner N, Bishop DK. Saccharomyces cerevisiae recA homologues RAD51 and DMC1 have both distinct and overlapping roles in meiotic recombination. Genes Cells. 1997;2:615–29.
Li W, Chen C, Markmann-Mulisch U, Timofejeva L, Schmelzer E, Ma H, et al. The Arabidopsis AtRAD51 gene is dispensable for vegetative development but required for meiosis. Proc Natl Acad Sci USA. 2004;101:10596–601.
Sonoda E, Sasaki MS, Buerstedde JM, Bezzubova O, Shinohara A, Ogawa H, et al. Rad51-deficient vertebrate cells accumulate chromosomal breaks prior to cell death. EMBO J. 1998;17:598–608.
Yoon SW, Kim DK, Kim KP, Park KS. Rad51 regulates cell cycle progression by preserving G2/M transition in mouse embryonic stem cells. Stem Cells Dev. 2014;23:2700–11.
Wassing IE, Esashi F. RAD51: Beyond the break. Semin Cell Dev Biol. 2021;113:38–46.
Wang AT, Kim T, Wagner JE, Conti BA, Lach FP, Huang AL, et al. A dominant mutation in human RAD51 reveals its function in DNA interstrand crosslink repair independent of homologous recombination. Mol Cell. 2015;59:478–90.
Wassing IE, Graham E, Saayman X, Rampazzo L, Ralf C, Bassett A, et al. The RAD51 recombinase protects mitotic chromatin in human cells. Nat Commun. 2021;12:5380.
Depienne C, Bouteiller D, Meneret A, Billot S, Groppa S, Klebe S, et al. RAD51 haploinsufficiency causes congenital mirror movements in humans. Am J Hum Genet. 2012;90:301–7.
Shinohara M, Gasior SL, Bishop DK, Shinohara A. Tid1/Rdh54 promotes colocalization of rad51 and dmc1 during meiotic recombination. Proc Natl Acad Sci USA. 2000;97:10814–9.
Sansam CL, Pezza RJ. Connecting by breaking and repairing: mechanisms of DNA strand exchange in meiotic recombination. FEBS J. 2015;282:2444–57.
Shi B, Xue J, Yin H, Guo R, Luo M, Ye L, et al. Dual functions for the ssDNA-binding protein RPA in meiotic recombination. PLoS Genet. 2019;15:e1007952.
Aponte PM. Isolation, culture, cryopreservation, and identification of bovine, murine, and human spermatogonial stem cells. Methods Mol Biol. 2020;2155:151–64.
Acknowledgements
We thank American Journal Experts (AJE) for English language editing on this paper. We also thank the Translational Medicine Core Facility of Shandong University for consultation and instrument availability that supported this work. This work was supported by the National Natural Science Foundation of China (81972437, 81672578), State Key Laboratory of Radiation Medicine and Protection (GZN1201804), and Key Research and Development Program of Shandong Province (2017GSF18188), S&T Program of Hebei (21377721D).
Author information
Authors and Affiliations
Contributions
Conception and design: JQ; ZL; CS; XZ. Development and methodology: JQ; LX; TH; Acquisition of data: JQ; JW; LX; HL; QD; XX. Analysis and interpretation of data: JQ; LX; TH; XZ. Administrative, technical, or material support: XZ; CS; HL; ZL. Study supervision: XZ; CS. Writing, review, and/or revision of the manuscript: All authors. Final approval: All authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Qin, J., Huang, T., Wang, J. et al. RAD51 is essential for spermatogenesis and male fertility in mice. Cell Death Discov. 8, 118 (2022). https://doi.org/10.1038/s41420-022-00921-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-022-00921-w
- Springer Nature Limited
This article is cited by
-
Fance deficiency impaired DNA damage repair of prospermatogonia and altered the repair dynamics of spermatocytes
Reproductive Biology and Endocrinology (2024)
-
RAD51 and Infertility: A Review and Case-Control Study
Biochemical Genetics (2024)
-
Sertoli cell-only syndrome: advances, challenges, and perspectives in genetics and mechanisms
Cellular and Molecular Life Sciences (2023)