Abstract
Iroquois homeobox transcription factor 5 (IRX5) plays a pivotal role in extramedullary adipogenesis, but little is known about the effects of IRX5 on adipogenesis of human bone marrow-derived mesenchymal stem cells (hMSCs). In this study, we aimed to determine the effect of IRX5 on hMSCs adipogenesis. By means of qPCR analysis, we determined that IRX5 expression was elevated during adipogenic commitment of hMSCs. The biologic role of IRX5 was further investigated by employing a gain/loss-of-function strategy using an in vitro lentivirus-based system. IRX5 overexpression promoted adipogenesis whereas IRX5 knockdown reduced the adipogenic phenotype. RNA-seq and metabolomics revealed that IRX5 overexpression repressed glycolysis. Dual-luciferase assay results showed that IRX5 overexpression transcriptionally activates peroxisome proliferator-activated receptor gamma coactivator (PGC-1α). Metformin and PGC-1α inhibitor reversed IRX5-induced adipogenesis and glycolytic inhibition. Collectively, IRX5 facilitates adipogenic differentiation of hMSCs by transcriptionally regulating PGC-1α and inhibiting glycolysis, revealing a potential target to control bone marrow-derived mesenchymal stem cells (BMSCs) fate decision and bone homeostasis.
Similar content being viewed by others
Introduction
Although bone marrow adipose tissue (BMAT) was discovered more than a century ago, little is known about its origin and function in bone marrow (BM) [1]. BMAT is distinct from white adipose tissues (WAT), brown adipose tissues (BAT) and beige adipose tissues. It plays significant roles in bone homeostasis and whole-body energy metabolism [2]. Postnatally, BMAT originates from progenitors that are distinct from peripheral adipose tissues. BMAT is thought to be derived from BMSCs located within BM stroma, which can give rise to osteoblasts and adipocytes [3,4,5,6,7]. Increasing evidence suggests that there is a theoretical inverse relationship between adipogenic and osteogenic differentiation of BMSCs both in vitro and in vivo [8, 9]. It has been reported that abnormal accumulation of adipocytes in the BM in osteoporosis, estrogen deficiency, anorexia nervosa, chronic glucocorticoid treatment, and Cushing disease leads to decreased bone mass and abnormal bone remodeling [10,11,12,13]. A clear understanding of the cellular and molecular mechanisms of BMSCs adipogenic commitment is profoundly significant for the purposes of illuminating the pathogenesis of bone and metabolic diseases and finding novel, effective therapeutic targets.
IRX5 is a family member of Iroquois homeobox transcription factors (IRX) which shares a highly conserved homeodomain from worm to vertebrates [14]. It plays an essential role in many physiological and pathological processes by binding specific sequence sites ACANNTGT. A causal cis-regulatory variant in the first intron of the FTO gene, which has been reported to be most associated with obesity, leads to activation of downstream targets IRX5 and IRX3 through long range enhancer promoter interactions. Manipulation of IRX3 and IRX5 expression in three different cellular models including mouse embryonic fibroblast–derived adipocytes, white 3T3-L1 preadipocytes, and β-adrenergic–stimulated beige ME3 preadipocytes induces adipocyte lipid accumulation and repressed thermogenesis [15]. IRX5 null mice had a significant anti-obesity phenotype associated with a dramatic loss of fat mass [16]. This evidence supports the notion that IRX5 is important for the identification and differentiation of extramedullary adipocytes. Considering the biological differences between peripheral adipose tissues and BMAT, the function of IRX5 on BMSC adipogenesis needs to be further addressed.
In this study, we showed that IRX5 overexpression promotes adipogenesis of hMSCs by increasing PGC-1α expression and inhibiting glycolysis. This effect is reversed by a glycolysis activator, metformin, and a PGC-1α inhibitor, SR-18292.
Results
The differentiation ability of hMSCs
We began the study by testing adipogenic and osteogenic differentiation of hMSCs. hMSCs were induced with an adipogenic medium or osteogenic medium.
As expected, adipogenesis-associated genes such as PPAR-γ, CEBP-α, LPL, FABP4, CD36 and PGC-1α were significantly up-regulated at D4 and D8 under adipogenic differentiation (Fig. 1A). After 16 days of adipogenic induction, Oil red O staining indicated that a substantial number of oil droplets had accumulated in cells (Fig. 1C). For osteogenic differentiation, RUNX2, COL1A and SPP1 were up-regulated during the early stage of osteogenesis while IBSP was up-regulated during the later period (Fig. 1B). ALP activity assay, ALP staining, and alizarin red staining showed high ALP activity and mineralization levels in differentiated cells (Fig. 1D, E). The results therefore suggest that hMSCs is a valid cell type that can be used to study lineage differentiation.
A mRNA levels of CEBP-α, PPAR-γ, LPL, FABP4, CD36 and PGC-1α during hMSCs adipogenic differentiation at D0, D4 and D8 (n = 3). B mRNA levels of RUNX2, ALPL, COL1A, SPP1 and IBSP during hMSCs osteogenic differentiation at D0, D7, D14 and D21 (n = 3). C Oil red O staining was performed at D16 after adipogenic induction. D, E ALP activity assay (n = 6) and ALP staining were performed at D7 (D, E), alizarin red staining was performed at D21 after osteo genic induction (E). Results are shown as mean ± SD. * indicates a significant difference from the control group, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
IRX5 overexpression and knockdown play opposite effects on adipogenic differentiation of hMSCs
Endogenous IRX5 expression increased during adipogenic differentiation, indicating the potential role of IRX5 in regulating adipogenic differentiation (Fig. 2A).
A Endogenous mRNA levels of IRX5 during adipogenic differentiation at D0, D4 and D8. B, C IRX5 overexpression efficiency were determined by qRT-PCR (B) and Western blot (C). D CCK-8 assay. E Protein levels of PPAR-γ and CEBP-α in Plvx/CTRL and Plvx/IRX5 groups by western blot assay. F Oil red O staining was performed at D16 after adipogenic induction. G mRNA levels of IRX5 during adipogenic differentiation at D0 and D4 in Plvx/CTRL and Plvx/IRX5 groups. H mRNA levels of CEBP-α, PPAR-γ, LPL, FABP4, CD36 and PGC-1α during adipogenic differentiation at D0 and D4. Scale bar, 500 μm. Results are shown as mean ± SD. n = 3. *p < 0.05, ** p < 0.01, ***p < 0.001, ****p < 0.0001.
Overexpression efficiency was detected by qRT-PCR and western blot in hMSCs (Fig. 2B, C and Fig. S3A), in ST2 cells (Figs. S1A, B and S8A) and in HELA cell line (Figs. S2A, B and S9A). CCK-8 assay was performed to exclude the effects of IRX5 in cell proliferation. These results indicated that proliferation was not affected by infection or IRX5 overexpression (Fig. 2D). Two key transcription factors of adipogenic differentiation, PPAR-γ and CEBP-α, were upregulated in Plvx/IRX5 hMSCs as shown in western blots (Fig. 2E and Fig. S3B). Oil red O staining for lipid accumulation at day 16 showed that more lipid droplets had accumulated in the Plvx/IRX5 group than the Plvx/CTRL group (Fig. 2F). IRX5 overexpression also increased the expression of important adipogenic associated genes, such as PPAR-γ, CEBP-α, LPL, FABP4 and CD36 (Fig. 2H and Fig. S1G). Consistent with endogenous expression levels of IRX5, both Plvx/CTRL and Plvx/IRX5 groups exhibited growing IRX5 levels during adipogenesis (Fig. 2G). On the other hand, knockdown of IRX5 was verified by qRT-PCR and western blot (Fig. 3A, B and Fig. S4). CEBP-α protein levels were down-regulated (Fig. 3B and Fig. S4). Oil red O staining confirmed a clear reduction for the accumulation of lipid droplets in IRX5 knockdown hMSCs (Fig. 3C); qRT-PCR analysis indicated that IRX5 knockdown inhibited the expression of PPAR-γ, CEBP-α, CD36 and FABP4 (Fig. 3D) after induction for 8 days. Thus, IRX5 overexpression promotes adipogenic differentiation of hMSCs, while IRX5 knockdown inhibits adipogenesis.
A, B IRX5 knockdown efficiency were verified by qRT-PCR (A) and western blot (B). B IRX5 and CEBP-α protein expression. C Oil red O staining analysis of lipid droplets at D16 after adipogenic differentiation. D qRT-PCR analysis of PPAR-γ, CEBP-α, CD36 and FABP4 at D8 after adipogenic differentiation. Scale bar, 500 μm. Results are shown as mean ± SD. n = 3. * Indicates a significant difference from the control group, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Glycolysis pathway and PGC-1α are regulated by IRX5 overexpression
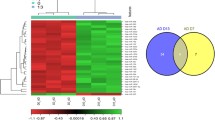
To gain a better understanding of the molecular mechanism of IRX5 in hMSCs, we determined global gene expression profiles by RNA sequencing (RNA-Seq). We analyzed genes which displayed significant differential expression identified by DEseq2 with adjusted p-value <0.05 and absolute log fold change >1. The volcano plot revealed that PGC-1α is one of the most significant genes changed in IRX5-overexpressing hMSCs (Fig. 4A). Heatmap analyses showed adipogenesis-associated genes were up-regulated in the Plvx/IRX5 group (Fig. 4B); whereas glycolysis-associated genes were down-regulated compared to control hMSCs (Fig. 4C). We tested and verified the results of RNA-seq through qRT-PCR and western blot. Consistently, overexpression of IRX5 promoted the expression of PGC-1α both in mRNA and protein levels significantly (Fig. 4D, F and Fig. S5); the major glycolysis-related genes were down-regulated by IRX5 overexpression as shown in qRT-PCR (Fig. 4E). Western blot showed that the protein expression of GLUT1, an important glucose transporter in the cell membrane, also decreased. These combined results suggest a reduced reliance on the glycolysis pathway (Fig. 4F and Fig. S5). In ST2 cells and HELA cells, we also found that PGC-1α were up-regulated and glycolysis were inhibited (Fig. S1C–F, S2C–E, S8B–C and S9B). Furthermore, we manually screened the potential promoter of PGC-1α for putative IRX5 binding sites. Human candidate PGC-1α promoter sequence from −1450 to +121 contains IRX5 binding sites (ACANNTGT). We then cloned the potential human PGC-1α promoter into the pGL3-basic luciferase reporter vector and performed a dual luciferase reporter assay. The results showed that IRX5 overexpression enhanced the transcriptional activity of PGC-1α (Fig. 4G), suggesting that PGC-1α is a possible pivotal transcriptional target of IRX5 in regulating adipogenic differentiation and energy metabolism.
A Volcano plot of differentially expressed genes (n = 3). B Heatmap plot of adipogenesis-associated genes (n = 3). C Heatmap of glycolysis-associated genes (n = 3). D qRT-PCR analysis of the mRNA level of PGC-1α in Plvx/CTRL and Plvx/IRX5 groups (n = 3). E qRT-PCR analysis of the mRNA level of GAPDH, GLUT1, HK2, LDHA, PFKP, PGK1 and PKM in Plvx/CTRL and Plvx/IRX5 groups (n = 3). F Western blot analysis of the protein levels of PGC-1α and GLUT1 (n = 3). G Dual luciferase reporter assay shows that overexpression of IRX5 promotes the transcription activity of PGC-1α in 293E cells (n = 5). Results are shown as mean ± SD. * indicates a significant difference from the control group, *p < 0.05, ** p < 0.01, *** p < 0.001, ****p < 0.0001.
Energy-targeted metabolomics analysis was performed to investigate the function of IRX5 in energy metabolism. Cell deposition of Plvx/CTRL and Plvx/IRX5 groups was assayed by LC-MS/MS. 2D-Principal component analysis (PCA) plots of the metabolites varied significantly between Plvx/CTRL and Plvx/IRX5 groups (Fig. 5A), indicating that both groups have very different metabolic profiles. Heatmap and violin plots showed that some glycolytic metabolites, especially lactate, were reduced while levels of tricarboxylic acid cycle (TCA) metabolites were increased in IRX5 overexpressing hMSCs (Fig. 5B, C). Glucose uptake and lactate production assays demonstrated that the rate of glucose uptake and lactate production was lower in Plvx/IRX5 group (Fig. 5D, E). Taken together, these data indicated that IRX5 inhibits the glycolytic pathway and promotes TCA cycle.
A Principal component analysis (PCA) (n = 3). Two-dimensional clustergram depicts the internal structure of the metabolomics data set with respect to variance (n = 3). B Heatmap of significantly changed metabolites in glycolysis and OxPhos pathway analyzed by LC-MS/MS (n = 3). C Violin plot compared the metabolites intensities in glycolysis and OxPhos pathway (n = 3). D Glucose uptake assay valued the rate of glucose consumption (n = 8). E Lactate production assay determined the rate of lactate production (n = 8). Results are shown as mean ± SD. *p < 0.05, ** p < 0.01, *** p < 0.001, ****p < 0.0001.
PGC-1α inhibitor and metformin activate glycolysis and rescue the excessive adipogenic differentiation of IRX5 overexpressing hMSCs
To confirm the involvement of PGC-1α in the excessive adipogenic differentiation of IRX5 overexpressing hMSCs, we treated cells with a selective chemical inhibitor of PGC-1α, SR-18292, which has been shown to repress PGC-1α activation by increasing its level of acetylation [17]. The toxicity of SR-18292 was tested with CCK-8 assays, which showed that concentrations of SR-18292 lower than 80uM have no toxic effect on proliferation (Fig. 6A). We validated that 10 μM and 20 μM of SR-18292 significantly inhibited the protein level of PGC-1α and increased the protein level of GLUT1 (Fig. 6B and Fig. S6A). 20 μM of SR-18292 was chosen for the following experiments. SR-18292 increased the rate of glucose uptake and lactate production in hMSCs (Fig. 6C, D). Meanwhile, reduced mRNA expression of glycolysis-related genes, PFKP and LDHA, and GLUT1 protein expression in Plvx/IRX5 group were reversed by SR-18292 (Fig. 6E–G and Fig. S6B–C). Accumulation of lipid droplets of the Plvx/IRX5 group after 16 days of adipogenic induction was reduced by SR-18292 (Fig. 6H). qRT-PCR also revealed that CEBP-α, PPAR-γ, FABP4, LPL and CD36 were down-regulated in the IRX5/Plvx upon SR-18292 treatment after 4 days (Figs. 6I) and 8 days (Fig. 6J) of adipogenic induction.
A CCK-8 assay evaluated the toxicity of SR-18292, a selective inhibitor of PGC-1α (n = 5). B Western blot analysis of the levels of PGC-1α and GLUT1 in hMSCs after treatment with 10 μM and 20 μM of SR-18292 (n = 3). C Glucose uptake assay with treatment of 20 μM SR-18292 (n = 5). D Lactate production assay with treatment of 20 μM SR-18292 (n = 5). E qRT-PCR analysis of the levels of PFKP and LDHA in Plvx/CTRL, Plvx/IRX5 and Plvx/IRX5 treated with 20 μM of SR-18292 (Plvx/IRX5 + SR-18292) groups during adipogenic differentiation (n = 3). F Western blot analysis of the levels of PGC-1α in Plvx/CTRL, Plvx/IRX5 and Plvx/IRX5 + SR-18292 groups (n = 3). G Western blot analysis of the levels of GLUT1 in Plvx/CTRL, Plvx/IRX5 and Plvx/IRX5 + SR-18292 groups (n = 3). H Oil red O staining of Plvx/CTRL, Plvx/IRX5 and Plvx/IRX5 + SR-18292 groups at D16 after adipogenic induction (n = 3). I mRNA levels of CEBP-α, PPAR-γ, FABP4, LPL and CD36 of Plvx/CTRL, Plvx/IRX5 and Plvx/IRX5 + SR-18292 groups at D4 after adipogenic induction (n = 3). J mRNA levels of CEBP-α, PPAR-γ, FABP4, LPL and CD36 of Plvx/CTRL, Plvx/IRX5 and Plvx/IRX5 + SR-18292 groups at D8 after adipogenic induction (n = 3). Scale bar, 500 μm. Results are shown as mean ± SD. *p < 0.05, ** p < 0.01, *** p < 0.001, ****p < 0.0001.
As a direct inhibitor of complex I in the electron transport chain (ETC) [18, 19], metformin has been used as an OxPhos inhibitor and glycolysis activator in many studies [20,21,22]. CCK-8 assays revealed that metformin did not affect the proliferation of hMSCs under concentrations of 1 mM (Fig. 7A). Levels of GLUT1 protein increased following treatment with 1 mM metformin for 24 hr (Fig. 7B and Fig. S7A). Activation of glycolysis was validated through glucose uptake and lactate production assay (Fig. 7C, D). mRNA levels of PFKP and LDHA were significantly higher in the Plvx/IRX5 group following metformin treatment (Fig. 7E). Using Western blot assays, we also found that GLUT1 protein expression was rescued by metformin (Fig. 7F and Fig. S7B). Protein levels of PGC-1α increased after metformin treatment in hMSCs (Fig. 7B and Fig. S7A), while mRNA expression of PGC-1α was only slighted affected by metformin in IRX5 overexpressing cells (Fig. 7G, H), indicating that glycolysis lies downstream of PGC-1α. Oil red O staining revealed that metformin reversed the increase of adipogenic differentiation in Plvx/IRX5 hMSCs (Fig. 7I). Moreover, mRNA levels of adipogenic-associated genes after 4 days (Fig. 7J) and 8 days (Fig. 7K) of adipogenic induction also revealed functional rescue of metformin. Overall, these findings demonstrate that IRX5 overexpression promotes adipogenic differentiation through PGC-1α regulation and glycolysis.
A CCK-8 assay evaluated the toxicity of metformin in hMSCs (n = 5). B Western blot analysis of the levels of PGC-1α and GLUT1 in hMSCs after treatment with 1 mM metformin (n = 3). C Glucose uptake assay was performed for the medium harvested from hMSCs after treatment with 1 mM metformin (n = 5). D Lactate production assay (n = 5). E qRT-PCR analysis of the levels of PFKP and LDHA in IRX5-control (Plvx/CTRL), IRX5-overexpressing (Plvx/IRX5) and IRX5-overexpressing treating with 1 mM metformin (Plvx/IRX5 + Metformin) groups during adipogenic differentiation (n = 3). F Western blot analysis of the levels of GLUT1 in Plvx/CTRL, Plvx/IRX5 and Plvx/IRX5 + Metformin groups (n = 3). G mRNA levels of PGC-1α in Plvx/CTRL, Plvx/IRX5 and Plvx/IRX5 + Metformin groups at D4 after adipogenic induction (n = 3). H mRNA levels of PGC-1α in Plvx/CTRL, Plvx/IRX5 and Plvx/IRX5 + Metformin groups at D8 after adipogenic induction (n = 3). I Oil red O staining of Plvx/CTRL, Plvx/IRX5 and Plvx/IRX5 + Metformin groups at D16 after adipogenic induction (n = 3). J mRNA levels of CEBP-α, PPAR-γ, FABP4, LPL and CD36 in Plvx/CTRL, Plvx/IRX5 and Plvx/IRX5 + Metformin groups at D4 after adipogenic induction (n = 3). K mRNA levels of CEBP-α, PPAR-γ, FABP4, LPL and CD36 in Plvx/CTRL, Plvx/IRX5 and Plvx/IRX5 + Metformin groups at D8 after adipogenic induction (n = 3). Scale bar, 500 μm. Results are shown as mean ± SD. *p < 0.05, ** p < 0.01, *** p < 0.001, ****p < 0.0001.
Discussion
In this study, we demonstrated that IRX5 promotes adipogenesis of hMSCs by increasing the expression of PGC-1α and inhibiting glycolysis. These changes can be reversed with PGC-1α inhibitor and metformin.
Marrow adipose tissue (MAT) accumulates naturally with age and in metabolic diseases like obesity [23]. Understanding the lineage commitment of BMSCs is critical for understanding bone homeostasis and for improvements in regenerative medicine. Adipogenesis of hMSCs involves two steps: commitment to adipogenic lineage cells followed by maturation into a functional adipocyte. Adipogenesis of hMSCs was induced with an induction cocktail containing l-methyl-3-isobutylxanthine (IBMX), dexamethasone, insulin, and Rosiglitazone [24]; maintained in an insulin-only medium. The accumulation of lipid-rich vacuoles within cells was readily visible once stained red with Oil Red O solution. We first verified osteogenic and adipogenic differentiation of hMSCs, which displayed classical differentiation patterns, indicating that hMSCs can be used as an appropriate cell model for investigating cell differentiation.
In the current study, we found that expression of IRX5 was upregulated during adipogenic induction. Overexpression of IRX5 in hMSCs promotes the expression of adipogenic-related genes and significantly improves the accumulation of lipid droplets. The pro-adipogenic effect of IRX5 is consistent with the study on the peripheral adipose tissues which showed that adult IRX5 knock-out mice had reduced fat mass and were protected from high fat diet-induced fat accumulation [25]. Large-scale clinical research studies demonstrated that a variant of FTO enhances the expression of IRX5 and IRX3 leading to excessive adipocyte differentiation and obesity [15, 26, 27]. However, this is contrary to one mouse study which showed that loss of Irx3 and Irx5 increases the hypertrophic chondrocytes transition toward adipocytes, and that the amount of marrow fat tissue in the neonatal distal tibia increases in these animals [28]. It has been revealed that there are two types of adipocytes in BM. Constitutive MAT (cMAT) develops early on in the distal skeletal region, contains large adipocytes resembling WAT, and is relatively devoid of active hematopoiesis; Regulated MAT (rMAT) containing single adipocytes interspersed throughout is colocalized with active hematopoiesis in proximal skeletal regions [29]. Based on the description in this study [28], more cMAT are accumulated to the distal tibia (cMAT) in Irx3-null or Irx5-null animals. While it is still unknown if rMAT and cMAT share the same progenitors, the lipid composition and gene expression are definitely different. This may help to explain previous inconsistency in the relationship between IRX5 and different adipose tissues. Two mutations in human IRX5 cause a recessive congenital disorder, Hamamy syndrome, resulting in skeletal abnormalities, osteopenia and severe cardiac defects [30]. The phenotype present in humans is not observed in Irx5−/− mice, suggesting that the function of IRX5 in humans is distinct from its function in mice [30,31,32,33,34,35]. Furthermore, the osteopenia in Hamamy Syndrome couldn’t be explained by the promoting adipogenic effect revealed in our study. However, accumulating evidence shows that the inverse correlation between MAT and bone mass is not always causally linked. For example, a subset of patients with congenital generalized lipodystrophy 1 (CGL1) or CGL2 (who lack MAT) develop osteolytic cyst-like lesions in the long bones [36]. Therefore, it is important to dissect the role of IRX5 on differentiation using hMSCs cell model.
Glucose, a major source of ATP for mammalian cells, can be metabolized by oxidative phosphorylation (OxPhos) and/or glycolysis [37]. BMSCs predominantly reside in regions of the BM cavity, which provides a hypoxic niche with the oxygen concentration of 1%-6% [38]. Under this condition, BMSCs tend to produce energy through glycolysis, reflecting their adaptation to low oxygen levels and their relatively low energy demand [39, 40]. The rewiring of glucose metabolism has also been implicated in cell lineage commitment [37]. For example, the Wnt/LRP5 pathway could promote osteogenic differentiation of mouse BMSCs through glycolysis [41], and notch signaling is known to partially suppress osteoblast differentiation by restricting glycolysis [42]. Additionally, the rate of glycolysis and increased formation of OxPhos supercomplexes seem to be characteristic of hMSC adipogenic differentiation [43,44,45,46,47,48]. Thus, adipogenesis of hMSCs is closely associated with changes in glucose metabolism. The data in this study indicate that IRX5 promotes PGC-1α expression and inhibits glycolysis. PGC-1α is a transcriptional co-activator of PPAR-γ and acts as a key regulator of mitochondrial biogenesis and oxidative metabolism in many tissues [49,50,51]. Some studies have revealed that PGC-1α modulates adipogenic differentiation [46, 52,53,54]. Acetylation of PGC-1α, which can be induced by SR-18292 [17], is used as a means of controlling its transcriptional activity and metabolic function. Metformin at suprapharmacological concentration could inhibit mitochondrial respiration, thereby compensatorily boosting glycolysis to produce ATP [55]. Here, we show that IRX5 overexpression in hMSCs led to increased PGC-1α expression and reduced glycolysis. The increased adipogenesis in hMSCs overexpressing IRX5 could be partially reversed by SR-18292 and metformin. Together, these observations implicate PGC-1α as an upstream driver of glycolysis and adipogenesis. Importantly, glycolysis is also a driver for adipogenesis, rather than a consequence of increased PGC-1α.
Energy metabolism during cell commitment is a multifaceted process. Therefore, understanding cell adaptation during differentiation and its molecular regulation could shed light on the application and therapy of BMSCs. Overall, the data presented in this manuscript suggest that hMSCs with IRX5 overexpression may shift their energy production preference from glycolysis to OxPhos, which is beneficial for adipogenesis.
In conclusion, IRX5 is accompanied by the adipogeneis of hMSCs; this adipogenesis is increased by IRX5 overexpression. PGC-1α inhibition and glycolysis activation rescue IRX5 modulated adipogenesis (Fig. 8). Moreover, further in-depth study of the relationship between IRX5 and bone homeostasis under physical and pathological conditions, as well as the underlying regulatory mechanism, are needed.
Materials and methods
Cell cultures and reagents
Human bone marrow-derived mesenchymal stem cells (hMSCs) were purchased from ScienCell (CA, USA) and cultured in mesenchymal stem cell medium (MSCM; ScienCell). hMSCs were incubated at 37 °C in a humidified atmosphere of 5% CO2 and were used at passage 3–7. SR-18292 (PGC-1α inhibitor) was purchased from Cayman (Michigan, USA) and metformin was purchased from Selleck (Texas, USA). Adipogenic differentiation induction medium and adipogenic differentiation maintenance medium were used for adipogenic induction (Cyagen, Guangdong, China).
Adipogenic differentiation
hMSCs were seeded in 6 well plate or 24 well plate at 5 × 104 cells/cm2. When the cells reached 100% confluence, the growth medium was replaced with an induction medium for 3 days and then a maintenance medium for an additional day. The cycle of induction and maintenance was repeated at least three times for optimal induction. After the cells differentiated, they were fixed with 4% paraformaldehyde for 10 min and stained with oil red O solution (Cyagen) for over 2 h. The lipid droplets were observed and photographed by an inverted microscope (Leica, Hessen, Germany).
Osteogenic differentiation
When the cells reached 70%-80% confluence, osteogenic differentiation medium, containing 10 mM β-glycerophosphate (Sigma-Aldrich, MO, USA), 50 μg/mL ascorbic acid (Sigma-Aldrich), and 1% Pen/Strep (Hyclone, UT, USA), was added and changed every day. ALP activity assay and ALP staining were conducted after osteogenic differentiation for 7 days as previously described [56].
After osteogenic induction for 21 days, mineral nodules were detected by staining with alizarin red S (pH=4.2; Servicebio, Hubei, China) for 1 hr.
Lentivirus production and transduction
The lentivirus vector (pGLV8/ mCherry/ Puro) overexpressing human or mouse IRX5 gene and vector (pGLV10/ mCherry/ Puro) containing shRNAs targeting IRX5 were packaged and purified by GenePharma (Shanghai, China). hMSCs, 293E cells, HELA cells and ST2 cells were infected with lentivirus in the presence of 5 μg/mL polybrene (Sigma-Aldrich) for 12–24 h. 48 h later, the infection efficiency was observed by fluorescent microscopy. Puromycin (2ug/ml) was used to select infected cells. For brevity, Cells with IRX5-overexpressing/empty lentivirus and IRX5-shRNA/Ctrl lentivirus infection were referred as Plvx/IRX5, Plvx/CTRL, IRX5-shRNA, and Ctrl-shRNA, respectively.
CCK-8 assay
hMSCs were seeded into 96-well plates with 10,000 cells for each well and the cell viability was tested as previously described [57].
qRT-PCR
Total RNA was extracted from cultured cells using a Trizol reagent kit (CWBIO, Beijing, China). The concentration of total RNA was measured by NanoDrop 2000 (Thermo Fisher., MA, USA). cDNA was transcribed from 1 μg of RNA using PrimeScriptTM Reverse Transcription Reagent Kit (Takara Bio, Kyoto, Japan). qRT-PCR was performed with TB Green Premix Ex Taq II Kit (Takara Bio) using QuantStudio 6 (Applied Biosystems, MA, USA). The primer sequences are listed in Table S1.
Western blot analysis
Proteins were collected on ice using RIPA (Beyotime, Shanghai, China) with phosphatase inhibitor (Roche Applied Science, Baden-Württemberg, Germany) and protease inhibitor (PMSF; Roche Applied Science). The concentration of protein was measured with a BCA kit (Biosharp, Anhui, China). 30 μg of protein were electrophoresed on 8%-10% SDS-polyacrylamide gel and transferred onto methanol-treated PVDF membrane (Millipore, MA, USA). The membranes were blocked with 5% non-fat milk for over 1 h and incubated with anti-IRX5 (Sigma-Aldrich, SAB1404807, MO, USA), anti-PGC-1α (Santa Cruz, SC-517380, Texas, USA), anti-GLUT1 (Abcam, ab115730, Cambridge, UK), anti-PPAR-γ (Abcam, ab178806, Cambridge, UK), anti-CEBP-α (Cell Signaling Technology, D56F10, Cambridge, UK), anti-β-actin (BioPM, PMK081W, Hubei, China), anti-β-Tubulin (Abmart, M20005F, Shanghai, China) antibodies at 4 °C overnight. An ECL system (Millipore) was used to visualize the target bands. Image J software (National Institutes of Health, Maryland, USA) was used for quantification by densitometry.
Transcriptome analysis
An RNeasy mini kit (Qiagen, Düsseldorf, Germany) was used to isolate total RNA from hMSCs. Library construction and sequencing were performed at Shanghai Sinomics Corporation. An Average of 49 million reads per sample were obtained. Trim Galore was applied for automatic quality and adapter trimming (https://www.bioinformatics.babraham.ac.uk/projects/trim_galore). The processed reads were aligned to the human reference genome HG38 by HISAT2 [58]. As a result, an average alignment rate of 85% was obtained. Sample read counts were summarized by StringTie2 [59] with the GENCODE V32 annotation file. Differential expression analysis was determined based on read counts by using DESeq2 [60]. P-values of DESeq2 were corrected using the Benjamini-Hochberg [61] procedure for multiple testing adjustment. The genes were ranked by the Wald statistics reported by DESeq2 followed by the pre-ranked GO term GSEA analysis [62].
Dual luciferase assay
Potential PGC-1α promoter (-1450bp to +121 bp) was amplified and then cloned into pGL3-basic vectors as reporter plasmids (See Table S2 for PGC-1α promoter sequences). 293E cells transfected stably with blank lentivirus or IRX5-overexpressing lentivirus were established and seeded on 24-well plates at a density of 1 × 105 cells per well. Then, the plasmids were transfected as previously described [57] and PGC-1α transcriptional activity (the light units of firefly luciferase/Renilla luciferase, fRLU/rRLU) was calculated.
LC–MS/MS assay
Cell collection
3 × 106 cells were seeded in 75 cm2 culture flasks and cultured until the cells reached over 90% confluence. Cells were washed with cold PBS twice, harvested by cell scrapers, and centrifuged at 1000 g for 10 min at 4 °C. The supernatant was discarded and the deposition was quick-frozen in liquid nitrogen and kept at −80 °C.
Sample preparation
Samples were thawed on ice. 500 μl pre-cooled 80% methanol aqueous solution was added and then whirled for 2 min. The mixture was frozen for 5 min in liquid nitrogen and thawed again. The thaw-frozen steps were repeated 3 times. Once the samples were centrifuged at 15,000 rpm at 4 °C for 20 min, the supernatant was transferred into the sample bottles for LC–MS/MS analysis.
Glucose uptake assay and lactate production assay
Cells were seeded on a 6 well plate at a density of 5 × 105 cells per well. After 24 h, the medium was replaced with no phenol red medium. After incubation for 24 h, the medium was collected, and the cell numbers were counted.
The glucose uptake assay and lactate production assay were performed as previously described [63].
Statistical analysis
All experiments were repeated in triplicate. The data were analyzed through Graphpad Prism Version 8.0. Results were expressed as mean ± SD and comparisons were analyzed by Student’s t test for two groups and ANOVA for more than two groups. P values < 0.05 were considered statistically significant.
Data availability
The RNA-seq raw data have been deposited in the GEO database. Geo number is GSE195679.
References
Horowitz MC, Berry R, Holtrup B, Sebo Z, Nelson T, Fretz JA, et al. Bone marrow adipocytes. Adipocyte 2017;6:193–204.
Cornish J, Wang T, Lin JM. Role of marrow adipocytes in regulation of energy metabolism and bone homeostasis. Curr Osteoporos Rep. 2018;16:116–22.
Bianco P, Riminucci M, Gronthos S, Robey PG. Bone marrow stromal stem cells: nature, biology, and potential applications. Stem Cells. 2001;19:180–92.
Horwitz EM, Le Blanc K, Dominici M, Mueller I, Slaper-Cortenbach I, Marini FC, et al. Clarification of the nomenclature for MSC: The International Society for Cellular Therapy position statement. Cytotherapy 2005;7:393–5.
Dominici M, Le Blanc K, Mueller I, Slaper-Cortenbach I, Marini F, Krause D, et al. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy 2006;8:315–7.
Muraglia A, Cancedda R, Quarto R. Clonal mesenchymal progenitors from human bone marrow differentiate in vitro according to a hierarchical model. J Cell Sci. 2000;113:1161–6.
Tencerova M, Kassem M. The bone marrow-derived stromal cells: commitment and regulation of adipogenesis. Front Endocrinol (Lausanne). 2016;7:127.
Beresford JN, Bennett JH, Devlin C, Leboy PS, Owen ME. Evidence for an inverse relationship between the differentiation of adipocytic and osteogenic cells in rat marrow stromal cell cultures. J Cell Sci. 1992;102:341.
Pei L, Tontonoz P. Fat’s loss is bone’s gain. J Clin Invest. 2004;113:805–6.
Kerwin E. Bone loss and inhaled glucocorticoids. N Engl J Med. 2002;346:533–5.
van Staa TP, Leufkens HG, Cooper C. The epidemiology of corticosteroid-induced osteoporosis: a meta-analysis. Osteoporos Int. 2002;13:777–87.
Bredella MA, Fazeli PK, Miller KK, Misra M, Torriani M, Thomas BJ, et al. Increased bone marrow fat in anorexia nervosa. J Clin Endocrinol Metab. 2009;94:2129–36.
Justesen J, Stenderup K, Ebbesen EN, Mosekilde L, Steiniche T, Kassem M. Adipocyte tissue volume in bone marrow is increased with aging and in patients with osteoporosis. Biogerontology 2001;2:165–71.
Gomez-Skarmeta JL, Modolell J. Iroquois genes: genomic organization and function in vertebrate neural development. Curr Opin Genet Dev. 2002;12:403–8.
Claussnitzer M, Dankel SN, Kim K, Quon G, Meuleman W, Haugen C, et al. FTO obesity variant circuitry and adipocyte browning in humans. N Engl J Med. 2015;373:895–907.
Sobreira DR, Joslin AC, Zhang Q, Williamson I, Hansen GT, Farris KM, et al. Extensive pleiotropism and allelic heterogeneity mediate metabolic effects of IRX3 and IRX5. Science. 2021;372:1085–91.
Sharabi K, Lin H, Tavares C, Dominy JE, Camporez JP, Perry RJ, et al. Selective chemical inhibition of PGC-1alpha gluconeogenic activity ameliorates type 2 diabetes. Cell 2017;169:148–60.
Foretz M, Guigas B, Bertrand L, Pollak M, Viollet B. Metformin: from mechanisms of action to therapies. Cell Metab. 2014;20:953–66.
Luengo A, Sullivan LB, Heiden MG. Understanding the complex-I-ty of metformin action: limiting mitochondrial respiration to improve cancer therapy. Bmc Biol. 2014;12:82.
Nian Y, Iske J, Maenosono R, Minami K, Heinbokel T, Quante M, et al. Targeting age-specific changes in CD4(+) T cell metabolism ameliorates alloimmune responses and prolongs graft survival. Aging Cell. 2021;20:e13299.
Pasquale V, Ducci G, Campioni G, Ventrici A, Assalini C, Busti S, et al. Profiling and targeting of energy and redox metabolism in grade 2 bladder cancer cells with different invasiveness properties. Cells. 2020;9:2669.
Jiang A, Du P, Liu Y, Pu J, Shi J, Zhang H. Metformin regulates the Th17/Treg balance by glycolysis with TIGAR in hepatic ischemia-reperfusion injury. J Pharm Sci. 2021;146:40–48.
Boroumand P, Klip A. Bone marrow adipose cells - cellular interactions and changes with obesity. J Cell Sci. 2020;133:jcs238394.
Zebisch K, Voigt V, Wabitsch M, Brandsch M. Protocol for effective differentiation of 3T3-L1 cells to adipocytes. Anal Biochem. 2012;425:88–90.
Bjune JI, Haugen C, Gudbrandsen O, Nordbo OP, Nielsen HJ, Vage V, et al. IRX5 regulates adipocyte amyloid precursor protein and mitochondrial respiration in obesity. Int J Obes. 2019;43:2151–62.
Sobalska-Kwapis M, Suchanecka A, Slomka M, Siewierska-Gorska A, Kepka E, Strapagiel D. Genetic association of FTO/IRX region with obesity and overweight in the Polish population. Plos ONE. 2017;12:e180295.
Hunt LE, Noyvert B, Bhaw-Rosun L, Sesay AK, Paternoster L, Nohr EA, et al. Complete re-sequencing of a 2Mb topological domain encompassing the FTO/IRXB genes identifies a novel obesity-associated region upstream of IRX5. Genome Med. 2015;7:126.
Tan Z, Kong M, Wen S, Tsang KY, Niu B, Hartmann C, et al. IRX3 and IRX5 inhibit adipogenic differentiation of hypertrophic chondrocytes and promote osteogenesis. J Bone Min Res. 2020;35:2444–57.
Scheller EL, Doucette CR, Learman BS, Cawthorn WP, Khandaker S, Schell B, et al. Region-specific variation in the properties of skeletal adipocytes reveals regulated and constitutive marrow adipose tissues. Nat Commun. 2015;6:7808.
Bonnard C, Strobl AC, Shboul M, Lee H, Merriman B, Nelson SF, et al. Mutations in IRX5 impair craniofacial development and germ cell migration via SDF1. Nat Genet. 2012;44:709–13.
Hamamy HA, Teebi AS, Oudjhane K, Shegem NN, Ajlouni KM. Severe hypertelorism, midface prominence, prominent/simple ears, severe myopia, borderline intelligence, and bone fragility in two brothers: new syndrome? Am J Med Genet. 2007;143A:229–34.
Hamamy HA, Masri AT, Al-Hadidy AM, Ajlouni KM. Consanguinity and genetic disorders. Profile Jordan Saudi Med J 2007;28:1015–7.
Bai M, Han Y, Wu Y, Liao J, Li L, Wang L, et al. Targeted genetic screening in mice through haploid embryonic stem cells identifies critical genes in bone development. Plos Biol. 2019;17:e3000350.
Cain CJ, Gaborit N, Lwin W, Barruet E, Ho S, Bonnard C, et al. Loss of Iroquois homeobox transcription factors 3 and 5 in osteoblasts disrupts cranial mineralization. Bone Rep. 2016;5:86–95.
Costantini DL, Arruda EP, Agarwal P, Kim KH, Zhu Y, Zhu W, et al. The homeodomain transcription factor Irx5 establishes the mouse cardiac ventricular repolarization gradient. Cell 2005;123:347–58.
Gregory JM, Arkader A, Bokhari A, Dormans JP. Case report: unicameral bone cysts in a young patient with acquired generalized lipodystrophy. Clin Orthop Relat Res. 2010;468:1440–6.
Liu Y, Ma T. Metabolic regulation of mesenchymal stem cell in expansion and therapeutic application. Biotechnol Progr. 2015;31:468–81.
Chow DC, Wenning LA, Miller WM, Papoutsakis ET. Modeling pO(2) distributions in the bone marrow hematopoietic compartment. I. Krogh’s model. Biophys J. 2001;81:675–84.
Nuschke A, Rodrigues M, Wells AW, Sylakowski K, Wells A. Mesenchymal stem cells/multipotent stromal cells (MSCs) are glycolytic and thus glucose is a limiting factor of in vitro models of MSC starvation. Stem Cell Res Ther. 2016;7:179.
Mylotte LA, Duffy AM, Murphy M, O’Brien T, Samali A, Barry F, et al. Metabolic flexibility permits mesenchymal stem cell survival in an ischemic environment. Stem Cells. 2008;26:1325–36.
Esen E, Chen J, Karner CM, Okunade AL, Patterson BW, Long F. WNT-LRP5 signaling induces Warburg effect through mTORC2 activation during osteoblast differentiation. Cell Metab. 2013;17:745–55.
Lee SY, Long F. Notch signaling suppresses glucose metabolism in mesenchymal progenitors to restrict osteoblast differentiation. J Clin Invest. 2018;128:5573–86.
Hofmann AD, Beyer M, Krause-Buchholz U, Wobus M, Bornhauser M, Rodel G. OXPHOS supercomplexes as a hallmark of the mitochondrial phenotype of adipogenic differentiated human MSCs. Plos ONE. 2012;7:e35160.
Guntur AR, Gerencser AA, Le PT, DeMambro VE, Bornstein SA, Mookerjee SA, et al. Osteoblast-like MC3T3-E1 cells prefer glycolysis for ATP production but adipocyte-like 3T3-L1 cells prefer oxidative phosphorylation. J Bone Min Res. 2018;33:1052–65.
Ryu MJ, Kim SJ, Choi MJ, Kim YK, Lee MH, Lee SE, et al. Mitochondrial oxidative phosphorylation reserve is required for hormone- and PPARgamma agonist-induced adipogenesis. Mol Cells. 2013;35:134–41.
Zhang Y, Marsboom G, Toth PT, Rehman J. Mitochondrial respiration regulates adipogenic differentiation of human mesenchymal stem cells. Plos ONE. 2013;8:e77077.
Cen S, Li J, Cai Z, Pan Y, Sun Z, Li Z, et al. TRAF4 acts as a fate checkpoint to regulate the adipogenic differentiation of MSCs by activating PKM2. Ebiomedicine 2020;54:102722.
Jiang Y, Guo L, Xie LQ, Zhang YY, Liu XH, Zhang Y, et al. Proteome profiling of mitotic clonal expansion during 3T3-L1 adipocyte differentiation using iTRAQ-2DLC-MS/MS. J Proteome Res. 2014;13:1307–14.
Puigserver P, Wu Z, Park CW, Graves R, Wright M, Spiegelman BM. A cold-inducible coactivator of nuclear receptors linked to adaptive thermogenesis. Cell 1998;92:829–39.
Lin J, Handschin C, Spiegelman BM. Metabolic control through the PGC-1 family of transcription coactivators. Cell Metab. 2005;1:361–70.
Liu C, Lin JD. PGC-1 coactivators in the control of energy metabolism. Acta Biochim Biophys Sin. 2011;43:248–57.
Forni MF, Peloggia J, Trudeau K, Shirihai O, Kowaltowski AJ. Murine mesenchymal stem cell commitment to differentiation is regulated by mitochondrial dynamics. Stem Cells. 2016;34:743–55.
Huang PI, Chen YC, Chen LH, Juan CC, Ku HH, Wang ST, et al. PGC-1alpha mediates differentiation of mesenchymal stem cells to brown adipose cells. J Atheroscler Thromb. 2011;18:966–80.
Huang PI, Chou YC, Chang YL, Chien Y, Chen KH, Song WS, et al. Enhanced differentiation of three-gene-reprogrammed induced pluripotent stem cells into adipocytes via adenoviral-mediated PGC-1alpha overexpression. Int J Mol Sci. 2011;12:7554–68.
Owen MR, Doran E, Halestrap AP. Evidence that metformin exerts its anti-diabetic effects through inhibition of complex 1 of the mitochondrial respiratory chain. Biochem J. 2000;348:607–14.
Song F, Sun H, Huang L, Fu D, Huang C. The role of Pannexin3-modified human dental pulp-derived mesenchymal stromal cells in repairing rat cranial critical-sized bone defects. Cell Physiol Biochem. 2017;44:2174–88.
Huang L, Song F, Sun H, Zhang L, Huang C. IRX5 promotes NF-κB signalling to increase proliferation, migration and invasion via OPN in tongue squamous cell carcinoma. J Cell Mol Med. 2018;22:3899–910.
Kim D, Paggi JM, Park C, Bennett C, Salzberg SL. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat Biotechnol. 2019;37:907–15.
Kovaka S, Zimin AV, Pertea GM, Razaghi R, Salzberg SL, Pertea M. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol. 2019;20:278.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Hochberg Y, Benjamini Y. More powerful procedures for multiple significance testing. Stat Med. 1990;9:811–8.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Lee WC, Ji X, Nissim I, Long F. Malic enzyme couples mitochondria with aerobic glycolysis in osteoblasts. Cell Rep. 2020;32:108108.
Acknowledgements
The authors thank Yaoting Ji (Wuhan University) for his kind help in this research. This work was supported by the National Natural Science Foundation of China (81800945 and 81970918).
Author information
Authors and Affiliations
Contributions
BJ performed most of the experiments and figures, analyzed the data, and wrote the manuscript. LH provided technical support. TT contributed to the analysis of data. HW and HY collaborated on conducting the experiments. TM edited the manuscript. FS contributed to the design of the study. CH supervised the work. All authors contributed to the work and have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jiang, B., Huang, L., Tian, T. et al. IRX5 promotes adipogenesis of hMSCs by repressing glycolysis. Cell Death Discov. 8, 204 (2022). https://doi.org/10.1038/s41420-022-00986-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-022-00986-7
- Springer Nature Limited