Abstract
Cholangiocarcinoma (CCA) is a malignant tumor that originates from the biliary epithelial cells. It is characterized by a difficult diagnosis and limited treatment options. Autophagy is a cellular survival mechanism that maintains nutrient and energy homeostasis and eliminates intracellular pathogens. It is involved in various physiological and pathological processes, including the development of cancer. However, the role, mechanism, and potential therapeutic targets of autophagy in CCA have not been thoroughly studied. In this review, we introduce the classification, characteristics, process, and related regulatory genes of autophagy. We summarize the regulation of autophagy on the progression of CCA and collect the latest research progress on some autophagy modulators with clinical potential in CCA. In conclusion, combining autophagy modulators with immunotherapy, chemotherapy, and targeted therapy has great potential in the treatment of CCA. This combination may be a potential therapeutic target for CCA in the future.
Similar content being viewed by others
Facts
-
Role of autophagy in the progression of CCA.
-
Autophagy modulators with clinical potential in CCA.
-
Combining autophagy modulators with immunotherapy, chemotherapy, and targeted therapy in the treatment of CCA.
Open questions
-
What’s future use of autophagy in the treatment of CCA?
-
How will autophagy work for CCA?
-
What will be the impact of combined therapy.
Introduction
Cholangiocarcinoma(CCA), arising from the biliary epithelium, is the most common bile duct malignancy and the second most common liver cancer after hepatocellular carcinoma (HCC) accounting for 10–20% of all primary liver cancers [1,2,3]. Fluke infection, inflammatory bowel disease, intrahepatic bile duct stones, choledochal cysts, and primary sclerosing cholangitis are well-known risk factors for CCA. Radical surgical resection is the only effective treatment for early-stage CCA [4, 5]. With the progression of the disease, chemotherapy is one of the effective treatments for advanced CCA. However, chemotherapy has poor inhibition effect on advanced malignant bile duct cells, and the 5-year survival rate is even less than 5% [6, 7].
Autophagy was first proposed by Ashford and Porter in 1962. It refers to the formation of autophagosome by the double-layer membrane covering part of the cytoplasm and the organelles, proteins and other components that are decomposed inside the cell by falling off from the free area of the rough endoplasmic reticulum and fusion with lysosome to form autophagolysosome to degrade the contents of the cell for its own metabolic needs and the renewal of specific organelles. It is usually a survival mechanism that maintains nutrient and energy homeostasis and eliminates intracellular pathogens. However, since the process has multi-steps conditions and control points, once several sites are out of control, it can lead to various human diseases, including cancer [8]. Recent studies have shown that autophagy plays a variety of pathophysiological roles in diseases such as cancer, autoimmune diseases and infection-related diseases. For example, it can help cells clear damaged proteins and pathogens. As a double-edged sword, autophagy can be used as both tumor promoter and tumor suppressor in the occurrence and development of cancer. BNIP3, an autophagy signaling protein (a pro-apoptotic member of Bcl-2), is expressed at high levels in colorectal and gastric epithelial cancers, suggesting that increased expression of BNIP3 may be necessary for the development of these cancers [9]. Beclin1 allele deletion has been found in human breast cancer [10] and prostate cancer [11]. In mouse models, lack of Beclin1 gene is more likely to develop lymphoma, liver cancer and other tumors [12]. In summary, autophagy regulation has emerged as a promising strategy for cancer treatment [13,14,15,16,17].
Autophagy
Classification of autophagy
Autophagy can be divided into three types: micro-autophagy (MI), chaperone-mediated autophagy (CMA), and macro-autophagy(M). MI refers to the process in which lysosomes or vacuole intima directly invaginate and wrap intracellular substances, and degrade in the lumen of organelles after membrane rupture [18]. CMA is highly selective and often degrades target proteins with uniquely recognized pentapeptide motifs (KFERQ) by means of cytoplasmic chaperone Hsc70. The receptor LAMP2A on the lysosome membrane recognizes the KFERQ group exposed to the binding protein and transfers the selected protein to the lysosome for degradation [19]. By contrast, MA is an evolutionarily conserved metabolic process. Proteins and other cytoplasmic components are sequestered in an isolation membrane, which expands and closes to form a double-layer vesicle structure, namely autophagosome. Then the autophagosome and lysosome fuse and eventually degrade. The typical process of MA is triggered by the inactivation of the mechanistic target of the mammalian target protein of the rapamycin complex 1 (mTORC1) protein complex. Under physiological conditions, MA can serve as a protective mechanism to maintain the stability of the genome under stress conditions. Then once malignant changes occur, MA will in turn protect tumor cells, maintain their proliferation, metastasis and drug resistance [13, 20,21,22].
Process of autophagy
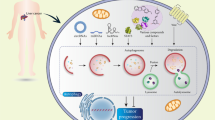
The whole process is highly regulated by a limited number of autophagy-related genes (ATGs) [23,24,25], which can be activated under a variety of stress conditions, such as starvation [26, 27], hypoxia [28, 29], infection, oxidative stress [30, 31], etc. Initiation, nucleation, fusion with lysosomes, and cargo degradation are four stages of process of autophagy [32]. In mammalian cells, the initiation of autophagy is regulated by the Unc-51-like kinase 1 (ULK1) complex, a complex of ULK1, ULK2, ATG13, FIP200 and ATG101. MTOR kinase is a major regulator of this process. Under nutrient-rich conditions, mTOR can phosphorylate ULK1/2 and block the activity of ULK1 complex. When nutrition, energy deficiency or other stress conditions, mTOR activity decreases after inhibition, and ULK1 complex is dephosphorylated and activated. In the autophagy nucleation step, after the activated complex is transferred to phagocytes, the PI3K-Beclin1 complex is activated and vesicle nucleation is induced [33]. During this period, two independent ubiquitin-like binding systems play a key role in the elongation and maturation of autophagy membranes. One is the ATG12-conjugation system. ATG12 is first activated by E1-like enzyme ATG7 [34], which is coupled with ATG5 [35] by E2-like enzyme ATG10 [36] to generate ATG5-ATG12 complex. The complex then further interacts with ATG16L to form ATG16L complex. The other is the LC3-conjugation system. ATG4 cleaves LC3 to produce LC3-I (soluble form). It then binds to phosphatidylethanolamine (PE) via ATG3 and ATG7 [35] to form LC3-II (present on the autophagosome membrane). LC3-II is crucial to the expansion and completion of the autophagy membrane. It is often used as a marker of autophagy progress in research. Then, mature autophagosomes fuse with lysosomes to form autophagic lysosomes. At last, the sequenced cytoplasmic components are degraded by lysosomal hydrolases. The degradation products are recycled to the cytoplasm for cell reuse. The adaptor protein sequestosome 1 (p62) can target specific products of autophagosomes and degrade with other cargo proteins, which is commonly used to measure autophagy flow [35] (Fig. 1).
A Initiation: ULK1 complex regulates autophagy initiation. B Nucleation: The activated ULK1 complex was transferred to phagocytes and activated PI3K-Beclin1 complex to induce vesicle nucleation; Elongation: The cytoplasm and organelles are enveloped and engulfed during elongation; Maturation: Completion and transport of autophagosomes. C Fusion: Autophagosomes fuse with lysosomes to form autophagolysosomes. D Degradation: Cytoplasmic components are degraded by lysosomal hydrolases. E ULK1 complex including ULK1, ULK2, ATG13, FIP200, and ATG101. F PI3K-Beclin1 complex including Beclin1, VPS34, VPS15, and ATG14L. G The ATG12-conjugation including ATG12, ATG7, ATG10, ATG5, and ATG16L. H The LC3-conjugation including LC3, ATG4, LC3-I, ATG7, ATG3, PE, and LC3-II.
The role of autophagy in CCA
CCA is a highly heterogeneous malignant tumor. Many risk factors can lead to chronic inflammation and/or cholestasis, which can alter the heredity and epigenetic inheritance of bile duct cells and ultimately lead to malignant transformation of bile duct cells [37]. Further research on CCA heredity and epigenetic inheritance, CCA occurrence, drug resistance, metastasis and recurrence mechanism will provide more specific therapies for CCA in the future.
Based on anatomic location, CCA are classified as intrahepatic (iCCA), perihilar (pCCA), or distal (dCCA) [38]. Various high-throughput spatial sequencing techniques showed that CCA at different anatomical sites had different molecular profiles and demonstrated heterogeneity of different subtypes. Studies have divided the genome of iCCA into two categories: one is inflammation and the other is proliferation. The former is mainly the activation of inflammatory signals, while the latter is the activation of oncogenic signaling pathways and oncogenes [39]. Most cancers have at least one driver gene mutation, and KRAS is the most common mutation, especially in dCCA [40].
Studies have found that chronic inflammation, epithelial mesenchymal transformation and epigenetic abnormalities are the driving factors of bile duct cell deterioration and play an important role in the progression of CCA. Autophagy is particularly important among these drivers.CCA, with its highly fibrous stroma, is the archetype of inflammatory cancer. Malignant transformation of bile duct cells is associated with chronic cholangitis [41]. Recent studies have found that pro-inflammatory cytokines such as IL-6, endotoxin and TNF are overexpressed in chronic bile duct inflammation. Inflammatory bile duct cells and overexpression of IL-6 contribute to malignant transformation of bile duct cells. During cancer formation, chronic inflammation is involved in a number of signaling pathways that influence the process of autophagy. Qi et al. reported that exposure of human lung epithelial cells to arsenic resulted in cell malignancy, resulting in persistent overexpression of IL6 and inhibition of autophagy [42]. IL6 affects the STAT3 signaling pathway by inhibiting the Beclin1-Bcl-2 complex, leading to carcinogenesis. On the contrary, overexpression of Beclin-1 can enhance autophagy and prevent IL-6-mediated cell malignancy. This correlation between IL6-mediated cellular malignancy and autophagy activity during tumorigenesis may provide a novel therapeutic approach for inflammatory iCCA.
Epithelial-to-mesenchymal transition (EMT) has been suggested as a driver of epithelial tumor spread and evidence showed that autophagy and EMT are correlated. EMT is a complex process. First, a wide range of factors including cytokines and growth factors (such as growth factors with affinity for receptor tyrosine kinases and transforming growth factor (TGF)-β1) [43, 44], morphogenetic signals (namely Hedgehog, Notch, and Wnt signaling) [45], and post-transcriptional gene regulator microRNAs [46]activate EMT transcription factors such as Snail(Snail1), Twist1/2, ZEB1/2 and Slug(Snail2). Subsequently, EMT transcription factors drive EMT. EMT allows cells to show mobility and invasiveness, which is related to tumor progression, metastasis and drug resistance [47, 48]. In human hepatocellular carcinoma (HCC), HepG2 cells can induce autophagy by EMT and TGF-β1/SMad3 [49]. TGF-β1 was found to increase the invasion of bile duct cancer cells through the EMT mechanism [50]. Nitta and colleagues indicated that autophagy occurred in nutrient deficiency-induced bile duct cancer cells, and immunohistochemical staining of CCA tissues showed that the expression of autophagy-associated protein Ambra1 was positively correlated with the expression of Snail, one of the major transcription factors of EMT. They found that inhibition of autophagy by chloroquine (CQ) attenuated the invasion of CCA cells under nutrient deficiency conditions and also reduced the invasion of CCA cells induced by TGF- β1 [51]. Similarly, in bladder cancer cells exhibiting nutrient deficiency, EMT induces autophagy through the TGF-β1/Smad3 signaling pathway and promotes the invasion of bladder cancer cells [52].
The role of epigenetic modifications such as DNA hypermethylation, microRNA and histone modification in the pathophysiology of CCA has attracted more and more attention [53,54,55]. It has been suggested that epigenetic abnormalities play an important role in the development of CCA [53] and also can regulate autophagy [56]. DNA methylation is a reversible chemical modification of promoter CpG island cytosine catalyzed by DNA methyltransferase family, which leads to gene inactivation through transcriptional inhibition [57, 58]. DNA methylation mediated silencing of tumor suppressor genes can often be seen in CCA. Wang et al. found that IDH1/2 DNA hypermethylation (called mutant IDH1/2) was found in 10% of iCCA. This mutant IDH1/2 had higher expression level of p53 and longer recurrence time and survival time than wild-type IDH1/2, suggesting that hypermethylated CCA may represent a different molecular subclass with a better prognosis [59]. Some studies have found the relationship between autophagy inhibition and histone methylation in IDH mutant gliomas [60], which provides a new treatment approach for CCA with autophagy inhibitors. Many microRNAs, such as miR-21, miR-200b and miR-29b, are up-regulated or down-regulated in CCA. They are not only biomarkers of CCA, but also therapeutic targets [61]. MicroRNAs are also involved in autophagy regulation in tumors and have been shown to alter the levels of several key proteins in the autophagy pathway, including Beclin1, LC3, ULK2, ATG4 and ATG9 [62, 63]. Studies found that the expression of miR-124 was down-regulated in CCA. miR-124 induces autophagy-related cell death in CCA cells by down-regulating the anti-apoptotic factor Bcl-2 and activating the autophagy-promoting protein Beclin1 by inhibiting STAT3 [64]. Overexpression of histone deacetylase 1 (HDAC) in iCCA has been reported to be associated with lymph node metastasis, vascular invasion and low survival [65]. Other HDACs has also been found in CCA. HDAC6 participates in the autophagy degradation pathway. HDAC6 inhibitors can inhibit autophagy in multiple myeloma and neuroblastoma. In a mouse model of colon cancer, the HDAC6 inhibitor was used in combination with bortezomib to inhibit tumor growth in vivo. Gradilone and his colleagues found that the overexpression of HDAC6 in CCA can promote the shortening of primary cilia and the malignant transformation of normal bile duct cells. Inhibiting HDAC6 can restore cilia expression and inhibit tumor growth [66].
The role of autophagy key proteins in CCA
Although the pathological role and regulatory mechanism of autophagy in the development of CCA are not fully understood, some recent reports have revealed important autophagy key proteins in CCA, emphasizes the significance of these autophagy-related proteins in CCA, and provided therapeutic pathway for CCA.
Beclin1 is the first identified autophagy effector of mammalian cells and a key factor in the initiation of autophagy. Studies have found that Beclin1 inactivation and autophagy defects can lead to malignant transformation of cells. The prognosis of Beclin1 is inconsistent for different tumors. When the expression of Beclin1 is different, the tumor metastasis, prognosis and survival rate are also different [12, 67, 68]. In nasopharyngeal carcinoma, highly expressed Beclin1 predicts poor prognosis [69]. In breast cancer, low expression of Beclin1 may contribute to tumor occurrence and development [68]. In CCA, studies have demonstrated the importance and prognostic value of Beclin1 [70, 71]. Dong and colleagues evaluated Beclin1 expression levels in iCCA samples. Compared with normal bile duct epithelial cells, the expression of Beclin1 was increased in most iCCA samples. In Beclin1 positive samples, low expression of Beclin1 was significantly associated with lymph node metastasis, lower overall survival and disease-free survival [70].
LC3 is another key autophagy regulator and a potential prognostic biomarker for cancer [72, 73]. It contains two isoforms: one is the soluble form LC3B(LC3B-I), the other is the lipidized form LC3B(LC3B-II). When LC3-I is converted to LC3-II, it indicates autophagy induction. During autophagy, the soluble form is transformed into the lipidized form and becomes a part of the autophagy membrane [74]. In a study, Chen et al. demonstrated for the first time that LC3B is an independent predictive biomarker for overall survival and disease-free survival of iCCA, and that high expression of LC3B indicates poor tumor differentiation, early recurrence and short long-term survival [75].
In CCA, FOXO1 is associated with autophagy flux. FOXO1 is an important transcription factor with a highly conserved DNA binding domain and is widely expressed in the spleen, liver and lung [76,77,78]. He et al. demonstrated for the first time that FOXO1 exists in autophagy regulation and can cause oxidative stress in human CCA cells by impairing autophagy flux, suggesting that FOXO1 may be a potential therapeutic target for CCA [79].
Death associated protein kinase 1 (DAPK1), one of Ser/Thr kinase family, is an important tumor suppressor. It mainly inhibits tumor cell growth by inducing autophagy and apoptosis [80]. We know that Beclin1 is a tumor suppressor, which affects the stability of p53 by affecting deubiquitination enzymes. DAPK1 inhibits tumor proliferation by different mechanisms, including upregulation of p53 function, etc [81]. As an autophagy inducer, however, how to inhibit tumor remains unclear.
Autophagy modulator in CCA
Autophagy activators
The procedure of autophagy is activated and regulated by numerous activator and mechanisim. Following are some activators for iniation of autophagy (Table 1).
Piperlongumine (PL)
PL, a natural product derived from piper longum plant, induces high levels of ROS production [82]. It passes through NF-κB, P38/JNK and other signaling pathways lead to apoptosis or autophagy in colon cancer, breast cancer, pancreatic cancer, kidney cancer, ovarian cancer, head and neck cancer and prostate cancer [83, 84]. Chen et al. found that PL also induced autophagy in CCA cells. In an in vitro experiment, they found that PL induced autophagy and apoptosis of HuCCT-1 cells via ROS-activated Erk signaling, and autophagy was inhibited when the Erk signaling pathway was inhibited [85]. This is the first report to demonstrate that PL can treat CCA by inducing cell autophagy. However, the specific efficacy has not been thoroughly studied, and a large number of studies are still needed if PL is applied to treat CCA in the future.
Pterostilbene
Pterostilbene is a natural methoxylated analogue of resveratrol, which has higher natural utilization, cell absorption, better lipophilicity, and a longer half-life than resveratrol [86]. Rimando et al. demonstrated that it has anti-inflammatory, anticancer and antioxidant biological effects [87]. Many studies have reported that pterostilbene has autophagy induction properties for various tumors [88,89,90,91]. In one study, after treating CCA cells with pterostilbene, it was found that pterostilbene did not induce cell apoptosis, but inhibited the cell proliferation by activating autophagy. It increased the expression of ATG5, Beclin1, LC3-II and inhibited the expression of P62 in vitro, which suggested the enhancement of autophagy activity. When CCA cells were treated with autophagy inhibitor 3-MA, autophagy induction and anti-tumor activity were inhibited [86].
Pristimerin
Pristimerin is a triterpenoid and can be isolated from Celastraceae and Hippocrateaceae. Sun et al. found that pristimerin significantly reduced the expression of apoptoses-related proteins procaspase-3, Bcl-2 and Bcl-XL, increased the expression of Bax, and promoted Beclin1 activation and autophagy initiation in vivo and in vitro [92]. In addition, they found that when the eCCA cell line QBC939 cells and the iCCA cell line RBE cells were treated with different concentrations of pristimerin, the inhibitory effect of this compound on QBC939 cells was stronger than that on RBE cells. The reason for the selectivity needed to be further studied [92].
Dihydroartemisinin (DHA)
DHA is an anti-malaria drug. Consistent with previous reports, DHA can increase the expression of autophagy-related genes such as ATG12 and BNIP3 while decreasing mTOR expression [93, 94]. Previous studies found that DHA killed CCA cells mainly through two pathways: caspase-dependent apoptosis and autophagy-dependent death pathways. Ser/Thr kinase DAPK is the main common mediator of these two pathways, which can regulate both apoptosis and autophagy [95,96,97]. Thongchot and colleagues reported that DHA induced ROS-mediated ER stress by activating DAPK, promoted the destruction of Beclin1-Bcl-2 complex and induced the death of CCA cells, thereby activating autophagy. Interestingly, DHA is only mildly toxic to normal bile duct cells but cytotoxic to cancer cells. In CCA cells lacking Beclin1, autophagy activation is blocked, which shows that DHA also needs Beclin1 activation [98].
Rapamycin and its analogues
Rapamycin and its analogues have been used as cancer therapeutic agents. One of their potential therapeutic mechanisms is the induction of autophagy, which may become a new molecular target for CCA therapy [99, 100]. The PI3K/Akt/mTOR pathway is a crucial signaling cascade regulating cell growth, proliferation and apoptosis [101]. Mammalian rapamycin target protein (mTOR) is a downstream effector of PI3K/Akt signaling pathway and is considered to be a therapeutic target for many diseases. The first definite mTOR inhibitor is rapamycin (sirolimus), also widely known as an autophagy inducer. Rapamycin and its analogue everolimus (RAD001) showed anti-tumor activity in previous studies [102,103,104]. In a study, rapamycin increased the sensitivity of PTEN mutant testicular cancer cells to radiotherapy by inducing autophagy [102, 105]. In a phase IB clinical study, RAD001 inhibited the growth of endometrial cancer cells and had synergistic effects with other anticancer drugs [106].
ABTL0812
ABTL0812 is a kind of small molecule drugs with anti-tumor activity. Currently, some phase 2 clinical trials are evaluating the efficacy of this drug in endometrial cancer and lung cancer [107, 108]. Previous studies have shown that ABTL0812 induces AKT-MRORC1 inhibition and autophagy-mediated tumor cell death by promoting the expression of TRIB3, a pseudokinase that binds and prevents AKT activation by upstream kinases PDPK1 and MTORC2 [107, 109]. In CCA, ABTL0812 has inhibitory effect on CCA cells. It can induce cytotoxic autophagy of CCA cells by inducing ER stress and TRIB3 mediated inhibition of Akt/mTOR axis [109, 110]. Similar to DHA, normal bile duct cells can survive at the concentration of ABTL0812 and are fatal to CCA cells, suggesting that ABTL0812 may provide a new and safe method for CCA treatment.
Phenformin
Phenformin is a biguanide compound, similar to metformin, used to treat type 2 diabetes [111]. In a study, Hu and his colleagues found that when CCA cells were treated with phenformin, autophagy-related genes ATG5, ATG7 and Beclin1 were up-regulated and cell death was increased, suggesting that phenformin can inhibit the growth of bile duct cancer cells by inducing autophagy in CCA cells [112]. Therefore, phenformin is expected to be a treatment for CCA.
ABC294640
Sphingosine kinase (Sphk) is a lipid kinase with a carcinogenic effect [113,114,115,116,117]. Studies have shown that Sphk subtype Sphk2 can produce mitogenic lipid sphingosine-1-phosphoric acid (S1P) to promote the growth of CCA [118]. ABC294640 is a novel specific inhibitor of Sphk2. Ding and colleagues found that ABC294640 can induce autophagy in CCA cells and can enhance the cytotoxicity and apoptosis induced by ABC294640 through the use of autophagy inhibitors bafilomycin A1 and chloroquine, thus inducing protective autophagy [119].
Urolithin A
The polyphenols ellagitannins and ellagic acid, which are found in nature, produce urolithin A (UA), a substance with anticancer action against a variety of cancers [120].
Studies have shown that UA can have several effects on gastric cancer (GC) cells in vitro. It can restrict the movement and invasion of these cancer cells, promote cell death (apoptosis), and induce autophagy by influencing the phosphatidylinositol-3-kinase/protein kinase B/mammalian target of rapamycin (PI3K/AKT/mTOR) pathway [120,121,122]. UA is expected to be used for GC treatment.
Autophagy inhibitors
There are also numerous inhibitors of autophagy procedure (Table 1).
Capsaicin
Capsaicin, the main pungent component in chili pepper, can inhibit human liver cancer, breast cancer and other malignant tumors [123, 124]. Among the natural compounds that can inhibit CCA autophagy, capsaicin is the only one that inhibits autophagy by activating mTOR. Existing studies have shown that capsaicin negatively regulates cell survival, adhesion, inflammation, differentiation and growth by interfering with nuclear factor-κB (NF-κB) and activator protein-1 (AP-1) signal transduction [125]. In CCA, capsaicin can reverse Beclin-1 and ATG5 upregulation and activate the PI3K/AKT/mTOR pathway to inhibit 5-Fu-induced autophagy [126]. Capsaicin in combination with 5-Fu also enhanced the sensitivity of QBC939 cells in vitro and in vivo [126], suggesting that capsaicin is promising as a new treatment option for CCA patients in combination with chemotherapy agents. However, the mechanism of the combination of capsaicin and chemotherapy drugs still needs more in-depth research.
Chloroquine (CQ)
CQ is an antimalarial drug that blocks the combination of lysosomes and autophagosomes by changing the acidic environment of lysosome, so as to accumulate a large number of degraded proteins and induce ER stress [127,128,129]. In CCA model, CQ could reduce CCA cell invasion and TGF-β 1-induced CCA cell invasion under starvation [129]. In addition, CQ can also improve the sensitivity of drug-resistant bile duct cancer cells to chemotherapy drugs [127]. Qu et al. compared the activity of QBC939 cells and HepG2 cells treated with CQ and found that the activity of QBC939 cells was more significantly inhibited[127]. The cell activity of CQ in combination with cisplatin was much lower than that of CQ alone. They further confirmed that the levels of autophagic proteins P62 and LC3-II/I in QBC939 cells were significantly increased after treatment with CQ, suggesting that QBC939 cells had high autophagy ability. This provides a new therapeutic idea for CQ as an autophagy inhibitor to inhibit autophagy and thus inhibit CCA cell growth in the future.
Hydroxychloroquine (HCQ)
HCQ acts similarly to chloroquine, both of which were initially used only as an adjuvant in combination with other antineoplastic agents [130, 131]. It was later found that HCQ or CQ alone can also produce good anti-tumor effects. HCQ induced apoptosis of CCA cells by upregulating ROS. With the addition of ROS scavger GSH, HCQ mainly inhibits the final stage of autophagy, namely the membrane fusion between autophagosome and lysosome [132, 133]. According to the results of Chen et al., when CCA was treated with IC50 concentration of HCQ, the number of apoptotic cells and the expression of apoptotic-related proteins in CCA cells increased, and G1 phase was blocked. At the same time, autophagy continued smoothly upstream, and the expression of autophagy-related proteins increased [134]. These results suggest that HCQ can inhibit the proliferation of bile duct cancer cells and induce apoptosis by inhibiting the accumulation of ROS induced by autophagy. However, this is only shown to be effective in animals, and the detailed mechanism in vivo needs to be further studied.
Resveratrol
Resveratrol is a natural polyphenolic antitoxin found in grapes. It has a variety of benefits for humans, especially in cardiovascular aspects. Previous studies have shown that resveratrol can induce autophagy-mediated cell death in gastric cancer and leukemia cells [135, 136]. In CCA, resveratrol inhibits autophagy by inhibiting FOXO1 acetylation, impairing the binding of FOXO1 and ATG7, leading to apoptosis [79]. He et al. found in vitro that one of the mechanisms of resveratrol induced apoptosis of QBC939 cells is that the inhibition of autophagy leads to oxidative stress and mitochondrial dysfunction (MTD) [79].
Oblongifolin C (OC)
OC is a natural small molecule compound extracted from Garcinia yunnanensis. Previous studies have demonstrated that OC is a novel inhibitor of autophagy flux, which can inhibit autophagosome-lysosomal fusion and autophagic degradation [137]. However, the effect of OC on CCA remains to be clarified. Zhang and colleagues treated CCA QBC939 cells with OC, and found that OC could inhibit autophagy and promote mitochondrial dysfunction (MtD) by blocking the fusion of autophagosome and lysosome, thus inducing apoptosis of CCA cells [138].
Salinomycin (Sal)
Sal is a polyether antibiotic, which has anti-tumor effects in liver cancer, leukemia, colon cancer and other tumors [139,140,141]. It has been proved that Sal can inhibit autophagy and play an anticancer role in CCA. Klose et al. treated CCA cells with different concentrations of Sal and found that the autophagy flux of CCA cells was inhibited in a dose-dependent manner. They also found that Sal also led to the accumulation of dysfunctional mitochondria and the increase of ROS after inhibiting autophagy in CCA cells, suggesting that it can influence autophagy and lysosomal fusion just as CQ does [142].
Discussion
Autophagy is a closely coordinative process that can maintain average cell growth and metabolism, prevent energy depletion after stress, and sustain protein and nutrient balance [143, 144]. Due to the multiple regulatory points in autophagy, studying the precise molecular mechanisms between autophagy and tumor is challenging. Due to the multiple regulatory sites of autophagy, it is challenging to study the precise molecular mechanisms between autophagy and tumors. In CCA, the current study seems to have little understanding of the relationship between autophagy and CCA. Although preclinical studies on targeted autophagy in the treatment of CCA have some results, its clinical studies are still uncertain. We can now take a little inspiration from studies of lung cancer, breast cancer and other cancers. The combination of autophagy and immunotherapy has been found in non-small cell lung cancer [145]. Autophagy plays an important role in adaptive immune responses. When ATG5 is lost, it not only damages autophagy, but also disrupts B cell development and T cell homeostasis [146, 147]. Combining autophagy regulation with immunotherapy will be a key area of future research. In clinical studies, targeted therapy with Traditional Chinese medicine can also regulate autophagy. Furanodiene is a natural compound isolated from the Rhizoma Curcumae. Xu et al. pointed out that it can stimulate ER stress and induce autophagy and apoptosis of lung cancer cells, indicating that autophagy can regulate the treatment of Traditional Chinese medicine [148]. In clinical treatment, Chinese medicine treatment can also regulate autophagy. In lung cancer, Chinese medicine affects the pro-death activity and protection of autophagy of lung cancer cells. Isodeoxycholin (ESI) shows potential anticancer effect [149]. After ESI treatment, autophagy markers such as ATG3, LC3-II and Beclin1 in lung cancer cells increased significantly, and therapeutic effects were produced through various channels. Marsdenia tenacissima (MTE) is a TCM that has been used to treat asthma, rheumatism, and tracheitis for thousands of years. MTE may induce apoptosis and autophagy inhibition coinstantaneously in lung cancer cells. Extracellular signal-regulated kinases (ERK) activation is partially linked with apoptotic and autophagic cell death after MTE treatment. Therefore, the mechanism of MTE induces apoptosis and suppresses autophagy via ERK activation [150]. Bu-Zhong-Yi-Qi Decoction (BZYQD) as a potential anti-tumor TCM. ROS accumulation may activate apoptosis and autophagy via oxidative stress [151].
The combination of autophagy and immunotherapy has been found in non-small cell lung cancer [145]. In contrast, autophagy has been well studied in the treatment of breast cancer. The effect of autophagy on breast tumor cells mainly depends on the stage of breast cancer development and previous treatment in the treatment process. Commonly used chemotherapeutic agents induce sustained autophagy and mediate autophagic cell death pathways. At this point, the use of the novel mTORC1/2 kinase increases autophagic flux and thus increase cytotoxic effects [152]. On the contrary, forced autophagy after chemotherapy may induce drug resistance, and tumors are prone to relapse. Therefore, drugs that disrupt autophagy mechanisms can be selected. In conclusion, the treatment strategies and directions for breast cancer at different stages are quite different.
Combined with the study of CCA, autophagy plays a multifaceted role in it. Even in different subtypes of CCA, autophagy has an environmentally dependent or conflicting role, suggesting a need to integrate the stages of disease development with autophagy characteristics. Given the heterogeneity of CCA and the potential need for long-term use of autophagy modulators during treatment, it is necessary to develop new biomarkers to further monitor autophagy status in patients with CCA and adjust medication according to this autophagy status. At present, various autophagy modulators for the treatment of CCA are still in the stage of preclinical research. Using preclinical models can further reveal the specific mechanism of autophagy in CCA. Comparing the autophagy levels of different mutated CCA cells can help us study how the mutation further regulates autophagy, and which subtype of patients with CCA will have a higher therapeutic effect if different autophagy modulators are applied. Therapies that combine autophagy regulation with immunotherapy, chemotherapy, and targeted therapies appear to be promising strategies for the treatment of CCA in the future.
References
Razumilava N, Gores GJ. Cholangiocarcinoma. Lancet. 2014;383:2168–79. https://doi.org/10.1016/s0140-6736(13)61903-0
Banales JM, Cardinale V, Carpino G, Marzioni M, Andersen JB, Invernizzi P, et al. Expert consensus document: Cholangiocarcinoma: current knowledge and future perspectives consensus statement from the European Network for the Study of Cholangiocarcinoma (ENS-CCA). Nat Rev Gastroenterol Hepatol. 2016;13:261–80. https://doi.org/10.1038/nrgastro.2016.51
Nakanuma Y, Sato Y, Harada K, Sasaki M, Xu J, Ikeda H. Pathological classification of intrahepatic cholangiocarcinoma based on a new concept. World J Hepatol. 2010;2:419–27. https://doi.org/10.4254/wjh.v2.i12.419
Doherty B, Nambudiri VE, Palmer WC. Update on the diagnosis and treatment of cholangiocarcinoma. Curr Gastroenterol Rep. 2017;19:2 https://doi.org/10.1007/s11894-017-0542-4
Blechacz B. Cholangiocarcinoma: current knowledge and new developments. Gut Liver. 2017;11:13–26. https://doi.org/10.5009/gnl15568
Bridgewater J, Galle PR, Khan SA, Llovet JM, Park JW, Patel T, et al. Guidelines for the diagnosis and management of intrahepatic cholangiocarcinoma. J Hepatol. 2014;60:1268–89. https://doi.org/10.1016/j.jhep.2014.01.021
Rizvi S, Gores GJ. Current diagnostic and management options in perihilar cholangiocarcinoma. Digestion. 2014;89:216–24. https://doi.org/10.1159/000360791
Galluzzi L, Bravo-San Pedro JM, Levine B, Green DR, Kroemer G. Pharmacological modulation of autophagy: therapeutic potential and persisting obstacles. Nat Rev Drug Discov. 2017;16:487–511. https://doi.org/10.1038/nrd.2017.22
Lee SH, Jeong EG, Yoo NJ, Lee SH. Mutational and expressional analysis of BNIP3, a pro-apoptotic Bcl-2 member, in gastric carcinomas. Apmis. 2007;115:1274–80. https://doi.org/10.1111/j.1600-0643.2007.00795.x
Futreal PA, Söderkvist P, Marks JR, Iglehart JD, Cochran C, Barrett JC, et al. Detection of frequent allelic loss on proximal chromosome 17q in sporadic breast carcinoma using microsatellite length polymorphisms. Cancer Res. 1992;52:2624–7.
Gao X, Zacharek A, Salkowski A, Grignon DJ, Sakr W, Porter AT, et al. Loss of heterozygosity of the BRCA1 and other loci on chromosome 17q in human prostate cancer. Cancer Res. 1995;55:1002–5.
Qu X, Yu J, Bhagat G, Furuya N, Hibshoosh H, Troxel A, et al. Promotion of tumorigenesis by heterozygous disruption of the beclin 1 autophagy gene. J Clin Investig. 2003;112:1809–20. https://doi.org/10.1172/jci20039
Yun CW, Lee SH. The roles of autophagy in cancer. Int J Mol Sci. 2018;19. https://doi.org/10.3390/ijms19113466
Chude CI, Amaravadi RK. Targeting autophagy in cancer: update on clinical trials and novel inhibitors. Int J Mol Sci. 2017;18. https://doi.org/10.3390/ijms18061279
Ávalos Y, Canales J, Bravo-Sagua R, Criollo A, Lavandero S, Quest AF. Tumor suppression and promotion by autophagy. Biomed Res Int. 2014;2014:603980. https://doi.org/10.1155/2014/603980
Marinković M, Šprung M, Buljubašić M, Novak I. Autophagy modulation in cancer: current knowledge on action and therapy. Oxid Med Cell Longev. 2018;2018:8023821. https://doi.org/10.1155/2018/8023821
Pietrocola F, Pol J, Vacchelli E, Baracco EE, Levesque S, Castoldi F, et al. Autophagy induction for the treatment of cancer. Autophagy. 2016;12:1962–4. https://doi.org/10.1080/15548627.2016.1214778
Yoshii SR, Mizushima N. Monitoring and measuring autophagy. Int J Mol Sci. 2017;18. https://doi.org/10.3390/ijms18091865
Kaushik S, Bandyopadhyay U, Sridhar S, Kiffin R, Martinez-Vicente M, Kon M, et al. Chaperone-mediated autophagy at a glance. J Cell Sci. 2011;124:495–9. https://doi.org/10.1242/jcs.073874.
Poillet-Perez L, White E. Role of tumor and host autophagy in cancer metabolism. Genes Dev. 2019;33:610–9. https://doi.org/10.1101/gad.325514.119
White E, Mehnert JM, Chan CS. Autophagy, metabolism, and cancer. Clin Cancer Res. 2015;21:5037–46. https://doi.org/10.1158/1078-0432.Ccr-15-0490
Li X, Zhou Y, Li Y, Yang L, Ma Y, Peng X, et al. Autophagy: a novel mechanism of chemoresistance in cancers. Biomed Pharmacother. 2019;119:109415 https://doi.org/10.1016/j.biopha.2019.109415
Wirawan E, Vanden Berghe T, Lippens S, Agostinis P, Vandenabeele P. Autophagy: for better or for worse. Cell Res. 2012;22:43–61. https://doi.org/10.1038/cr.2011.152
Chen Y, Klionsky DJ. The regulation of autophagy—unanswered questions. J Cell Sci. 2011;124:161–70. https://doi.org/10.1242/jcs.064576. Pt 2
Parzych KR, Klionsky DJ. An overview of autophagy: morphology, mechanism, and regulation. Antioxid Redox Signal. 2014;20:460–73. https://doi.org/10.1089/ars.2013.5371
Su Z, Wang T, Zhu H, Zhang P, Han R, Liu Y, et al. HMGB1 modulates Lewis cell autophagy and promotes cell survival via RAGE-HMGB1-Erk1/2 positive feedback during nutrient depletion. Immunobiology. 2015;220:539–44. https://doi.org/10.1016/j.imbio.2014.12.009
Joy S, Thirunavukkarasu L, Agrawal P, Singh A, Sagar BKC, Manjithaya R, et al. Basal and starvation-induced autophagy mediates parasite survival during intraerythrocytic stages of Plasmodium falciparum. Cell Death Discov. 2018;4:43. https://doi.org/10.1038/s41420-018-0107-9
Yang Z, Goronzy JJ, Weyand CM. Autophagy in autoimmune disease. J Mol Med. 2015;93:707–17. https://doi.org/10.1007/s00109-015-1297-8
Yang Z, Zhong L, Zhong S, Xian R, Yuan B. Hypoxia induces microglia autophagy and neural inflammation injury in focal cerebral ischemia model. Exp Mol Pathol. 2015;98:219–24. https://doi.org/10.1016/j.yexmp.2015.02.003
Tang Y, Jacobi A, Vater C, Zou L, Zou X, Stiehler M. Icariin promotes angiogenic differentiation and prevents oxidative stress-induced autophagy in endothelial progenitor cells. Stem Cells. 2015;33:1863–77. https://doi.org/10.1002/stem.2005
Li SJ, Sun SJ, Gao J, Sun FB. Wogonin induces Beclin-1/PI3K and reactive oxygen species-mediated autophagy in human pancreatic cancer cells. Oncol Lett. 2016;12:5059–67. https://doi.org/10.3892/ol.2016.5367
Kaur J, Debnath J. Autophagy at the crossroads of catabolism and anabolism. Nat Rev Mol Cell Biol. 2015;16:461–72. https://doi.org/10.1038/nrm4024
Itakura E, Kishi C, Inoue K, Mizushima N. Beclin 1 forms two distinct phosphatidylinositol 3-kinase complexes with mammalian Atg14 and UVRAG. Mol Biol Cell. 2008;19:5360–72. https://doi.org/10.1091/mbc.e08-01-0080
Tanida I, Mizushima N, Kiyooka M, Ohsumi M, Ueno T, Ohsumi Y, et al. Apg7p/Cvt2p: a novel protein-activating enzyme essential for autophagy. Mol Biol Cell. 1999;10:1367–79. https://doi.org/10.1091/mbc.10.5.1367
Mizushima N, Noda T, Yoshimori T, Tanaka Y, Ishii T, George MD, et al. A protein conjugation system essential for autophagy. Nature. 1998;395:395–8. https://doi.org/10.1038/26506
Shintani T, Mizushima N, Ogawa Y, Matsuura A, Noda T, Ohsumi Y. Apg10p, a novel protein-conjugating enzyme essential for autophagy in yeast. EMBO J. 1999;18:5234–41. https://doi.org/10.1093/emboj/18.19.5234
Labib PL, Goodchild G, Pereira SP. Molecular pathogenesis of cholangiocarcinoma. BMC Cancer. 2019;19:185. https://doi.org/10.1186/s12885-019-5391-0
Rizvi S, Khan SA, Hallemeier CL, Kelley RK, Gores GJ. Cholangiocarcinoma—evolving concepts and therapeutic strategies. Nat Rev Clin Oncol. 2018;15:95–111. https://doi.org/10.1038/nrclinonc.2017.157
Sia D, Hoshida Y, Villanueva A, Roayaie S, Ferrer J, Tabak B, et al. Integrative molecular analysis of intrahepatic cholangiocarcinoma reveals 2 classes that have different outcomes. Gastroenterology. 2013;144:829–40. https://doi.org/10.1053/j.gastro.2013.01.001
Simbolo M, Fassan M, Ruzzenente A, Mafficini A, Wood LD, Corbo V, et al. Multigene mutational profiling of cholangiocarcinomas identifies actionable molecular subgroups. Oncotarget. 2014;5:2839–52. https://doi.org/10.18632/oncotarget.1943
Cardinale V, Semeraro R, Torrice A, Gatto M, Napoli C, Bragazzi MC, et al. Intra-hepatic and extra-hepatic cholangiocarcinoma: New insight into epidemiology and risk factors. World J Gastrointest Oncol. 2010;2:407–16. https://doi.org/10.4251/wjgo.v2.i11.407
Qi Y, Zhang M, Li H, Frank JA, Dai L, Liu H, et al. Autophagy inhibition by sustained overproduction of IL6 contributes to arsenic carcinogenesis. Cancer Res. 2014;74:3740–52. https://doi.org/10.1158/0008-5472.Can-13-3182
Katsuno Y, Lamouille S, Derynck R. TGF-β signaling and epithelial-mesenchymal transition in cancer progression. Curr Opin Oncol. 2013;25:76–84. https://doi.org/10.1097/CCO.0b013e32835b6371
Lamouille S, Xu J, Derynck R. Molecular mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell Biol. 2014;15:178–96. https://doi.org/10.1038/nrm3758
Gonzalez DM, Medici D. Signaling mechanisms of the epithelial-mesenchymal transition. Sci Signal. 2014;7:re8. https://doi.org/10.1126/scisignal.2005189
Guo F, Parker Kerrigan BC, Yang D, Hu L, Shmulevich I, Sood AK, et al. Post-transcriptional regulatory network of epithelial-to-mesenchymal and mesenchymal-to-epithelial transitions. J Hematol Oncol. 2014;7:19. https://doi.org/10.1186/1756-8722-7-19
Saitoh M. Involvement of partial EMT in cancer progression. J Biochem. 2018;164:257–64. https://doi.org/10.1093/jb/mvy047
Krebs AM, Mitschke J, Lasierra Losada M, Schmalhofer O, Boerries M, Busch H, et al. The EMT-activator Zeb1 is a key factor for cell plasticity and promotes metastasis in pancreatic cancer. Nat Cell Biol. 2017;19:518–29. https://doi.org/10.1038/ncb3513
Li J, Yang B, Zhou Q, Wu Y, Shang D, Guo Y, et al. Autophagy promotes hepatocellular carcinoma cell invasion through activation of epithelial-mesenchymal transition. Carcinogenesis. 2013;34:1343–51. https://doi.org/10.1093/carcin/bgt063
Sato Y, Harada K, Itatsu K, Ikeda H, Kakuda Y, Shimomura S, et al. Epithelial-mesenchymal transition induced by transforming growth factor-{beta}1/Snail activation aggravates invasive growth of cholangiocarcinoma. Am J Pathol. 2010;177:141–52. https://doi.org/10.2353/ajpath.2010.090747
Nitta T, Sato Y, Ren XS, Harada K, Sasaki M, Hirano S, et al. Autophagy may promote carcinoma cell invasion and correlate with poor prognosis in cholangiocarcinoma. Int J Clin Exp Pathol. 2014;7:4913–21.
Tong H, Yin H, Hossain MA, Wang Y, Wu F, Dong X, et al. Starvation-induced autophagy promotes the invasion and migration of human bladder cancer cells via TGF-β1/Smad3-mediated epithelial-mesenchymal transition activation. J Cell Biochem. 2019;120:5118–27. https://doi.org/10.1002/jcb.27788
Sandhu DS, Shire AM, Roberts LR. Epigenetic DNA hypermethylation in cholangiocarcinoma: potential roles in pathogenesis, diagnosis and identification of treatment targets. Liver Int. 2008;28:12–27. https://doi.org/10.1111/j.1478-3231.2007.01624.x
Andersen JB, Thorgeirsson SS. Genetic profiling of intrahepatic cholangiocarcinoma. Curr Opin Gastroenterol. 2012;28:266–72. https://doi.org/10.1097/MOG.0b013e3283523c7e
Isomoto H. Epigenetic alterations associated with cholangiocarcinoma (review). Oncol Rep. 2009;22:227–32.
Baek SH, Kim KI. Epigenetic control of autophagy: nuclear events gain more attention. Mol Cell. 2017;65:781–5. https://doi.org/10.1016/j.molcel.2016.12.027
Baylin SB, Herman JG. DNA hypermethylation in tumorigenesis: epigenetics joins genetics. Trends Genet. 2000;16:168–74. https://doi.org/10.1016/s0168-9525(99)01971-x
Okano M, Bell DW, Haber DA, Li E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell. 1999;99:247–57. https://doi.org/10.1016/s0092-8674(00)81656-6
Wang P, Dong Q, Zhang C, Kuan PF, Liu Y, Jeck WR, et al. Mutations in isocitrate dehydrogenase 1 and 2 occur frequently in intrahepatic cholangiocarcinomas and share hypermethylation targets with glioblastomas. Oncogene. 2013;32:3091–3100. https://doi.org/10.1038/onc.2012.315
Trejo-Solís C, Serrano-Garcia N, Escamilla-Ramírez Á, Castillo-Rodríguez RA, Jimenez-Farfan D, Palencia G, et al. Autophagic and apoptotic pathways as targets for chemotherapy in glioblastoma. Int J Mol Sci. 2018;19. https://doi.org/10.3390/ijms19123773
Howell JA, Khan SA. The role of miRNAs in cholangiocarcinoma. Hepat Oncol. 2016;3:167–80. https://doi.org/10.2217/hep-2015-0003
Frankel LB, Lund AH. MicroRNA regulation of autophagy. Carcinogenesis. 2012;33:2018–25. https://doi.org/10.1093/carcin/bgs266
Gozuacik D, Akkoc Y, Ozturk DG, Kocak M. Autophagy-regulating microRNAs and cancer. Front Oncol. 2017;7:65. https://doi.org/10.3389/fonc.2017.00065
Roch AM, Maatman T, Carr RA, Easler JJ, Schmidt CM, House MG, et al. Evolving treatment of necrotizing pancreatitis. Am J Surg. 2018;215:526–9. https://doi.org/10.1016/j.amjsurg.2017.11.020
Morine Y, Shimada M, Iwahashi S, Utsunomiya T, Imura S, Ikemoto T, et al. Role of histone deacetylase expression in intrahepatic cholangiocarcinoma. Surgery. 2012;151:412–9. https://doi.org/10.1016/j.surg.2011.07.038
Gradilone SA, Radtke BN, Bogert PS, Huang BQ, Gajdos GB, LaRusso NF. HDAC6 inhibition restores ciliary expression and decreases tumor growth. Cancer Res. 2013;73:2259–70. https://doi.org/10.1158/0008-5472.Can-12-2938
Kong XX, Zhang HY, Chen ZQ, Fan XF, Gong YS. [Inhibition of Beclin 1 enhances apoptosis by H2O2 in glioma U251 cells]. Sheng Li Xue Bao. 2011;63:238–44.
Liang XH, Jackson S, Seaman M, Brown K, Kempkes B, Hibshoosh H, et al. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature. 1999;402:672–6. https://doi.org/10.1038/45257
Wan XB, Fan XJ, Chen MY, Xiang J, Huang PY, Guo L, et al. Elevated Beclin 1 expression is correlated with HIF-1alpha in predicting poor prognosis of nasopharyngeal carcinoma. Autophagy. 2010;6:395–404. https://doi.org/10.4161/auto.6.3.11303
Dong LW, Hou YJ, Tan YX, Tang L, Pan YF, Wang M, et al. Prognostic significance of Beclin 1 in intrahepatic cholangiocellular carcinoma. Autophagy. 2011;7:1222–9. https://doi.org/10.4161/auto.7.10.16610
Wang TT, Cao QH, Chen MY, Xia Q, Fan XJ, Ma XK, et al. Beclin 1 deficiency correlated with lymph node metastasis, predicts a distinct outcome in intrahepatic and extrahepatic cholangiocarcinoma. PLoS ONE. 2013;8:e80317. https://doi.org/10.1371/journal.pone.0080317
Adams O, Dislich B, Berezowska S, Schläfli AM, Seiler CA, Kröll D, et al. Prognostic relevance of autophagy markers LC3B and p62 in esophageal adenocarcinomas. Oncotarget. 2016;7:39241–55. https://doi.org/10.18632/oncotarget.9649
Ladoire S, Chaba K, Martins I, Sukkurwala AQ, Adjemian S, Michaud M, et al. Immunohistochemical detection of cytoplasmic LC3 puncta in human cancer specimens. Autophagy. 2012;8:1175–84. https://doi.org/10.4161/auto.20353
Mizushima N. Methods for monitoring autophagy. Int J Biochem Cell Biol. 2004;36:2491–502. https://doi.org/10.1016/j.biocel.2004.02.005
Chen L, Fu H, Lu T, Cai J, Liu W, Yao J, et al. An integrated nomogram combining clinical factors and microtubule-associated protein 1 light chain 3B expression to predict postoperative prognosis in patients with intrahepatic cholangiocarcinoma. Cancer Res Treat. 2020;52:469–80. https://doi.org/10.4143/crt.2019.423
Calnan DR, Brunet A. The FoxO code. Oncogene. 2008;27:2276–88. https://doi.org/10.1038/onc.2008.21
Li Y, Ma Z, Jiang S, Hu W, Li T, Di S, et al. A global perspective on FOXO1 in lipid metabolism and lipid-related diseases. Prog Lipid Res. 2017;66:42–49. https://doi.org/10.1016/j.plipres.2017.04.002
Xing YQ, Li A, Yang Y, Li XX, Zhang LN, Guo HC. The regulation of FOXO1 and its role in disease progression. Life Sci. 2018;193:124–31. https://doi.org/10.1016/j.lfs.2017.11.030
He W, Zhang A, Qi L, Na C, Jiang R, Fan Z, et al. FOXO1, a potential therapeutic target, regulates autophagic flux, oxidative stress, mitochondrial dysfunction, and apoptosis in human cholangiocarcinoma QBC939 cells. Cell Physiol Biochem. 2018;45:1506–14. https://doi.org/10.1159/000487576
Bialik S, Kimchi A. The death-associated protein kinases: structure, function, and beyond. Annu Rev Biochem. 2006;75:189–210. https://doi.org/10.1146/annurev.biochem.75.103004.142615
Raveh T, Droguett G, Horwitz MS, DePinho RA, Kimchi A. DAP kinase activates a p19ARF/p53-mediated apoptotic checkpoint to suppress oncogenic transformation. Nat Cell Biol. 2001;3:1–7. https://doi.org/10.1038/35050500
Möhler H, Pfirrmann RW, Frei K. Redox-directed cancer therapeutics: taurolidine and piperlongumine as broadly effective antineoplastic agents (review). Int J Oncol. 2014;45:1329–36. https://doi.org/10.3892/ijo.2014.2566
Zheng J, Son DJ, Gu SM, Woo JR, Ham YW, Lee HP, et al. Piperlongumine inhibits lung tumor growth via inhibition of nuclear factor kappa B signaling pathway. Sci Rep. 2016;6:26357. https://doi.org/10.1038/srep26357
Makhov P, Golovine K, Teper E, Kutikov A, Mehrazin R, Corcoran A, et al. Piperlongumine promotes autophagy via inhibition of Akt/mTOR signalling and mediates cancer cell death. Br J Cancer. 2014;110:899–907. https://doi.org/10.1038/bjc.2013.810
Chen SY, Huang HY, Lin HP, Fang CY. Piperlongumine induces autophagy in biliary cancer cells via reactive oxygen species-activated Erk signaling pathway. Int J Mol Med. 2019;44:1687–96. https://doi.org/10.3892/ijmm.2019.4324
Wang D, Guo H, Yang H, Wang D, Gao P, Wei W. Pterostilbene, an active constituent of blueberries, suppresses proliferation potential of human cholangiocarcinoma via enhancing the autophagic flux. Front Pharmacol. 2019;10:1238 https://doi.org/10.3389/fphar.2019.01238
Rimando AM, Cuendet M, Desmarchelier C, Mehta RG, Pezzuto JM, Duke SO. Cancer chemopreventive and antioxidant activities of pterostilbene, a naturally occurring analogue of resveratrol. J Agric Food Chem. 2002;50:3453–7. https://doi.org/10.1021/jf0116855
Kang KY, Shin JK, Lee SM. Pterostilbene protects against acetaminophen-induced liver injury by restoring impaired autophagic flux. Food Chem Toxicol. 2019;123:536–45. https://doi.org/10.1016/j.fct.2018.12.012
Ko CP, Lin CW, Chen MK, Yang SF, Chiou HL, Hsieh MJ. Pterostilbene induce autophagy on human oral cancer cells through modulation of Akt and mitogen-activated protein kinase pathway. Oral Oncol. 2015;51:593–601. https://doi.org/10.1016/j.oraloncology.2015.03.007
Wang Y, Ding L, Wang X, Zhang J, Han W, Feng L, et al. Pterostilbene simultaneously induces apoptosis, cell cycle arrest and cyto-protective autophagy in breast cancer cells. Am J Transl Res. 2012;4:44–51.
Yu CL, Yang SF, Hung TW, Lin CL, Hsieh YH, Chiou HL. Inhibition of eIF2α dephosphorylation accelerates pterostilbene-induced cell death in human hepatocellular carcinoma cells in an ER stress and autophagy-dependent manner. Cell Death Dis. 2019;10:418. https://doi.org/10.1038/s41419-019-1639-5
Sun JM, Xu HT, Zhao L, Zhang YB, Kang PC, Song ZF, et al. Induction of cell-cycle arrest and apoptosis in human cholangiocarcinoma cells by pristimerin. J Cell Biochem. 2019. https://doi.org/10.1002/jcb.28485
Qin G, Zhao C, Zhang L, Liu H, Quan Y, Chai L, et al. Dihydroartemisinin induces apoptosis preferentially via a Bim-mediated intrinsic pathway in hepatocarcinoma cells. Apoptosis. 2015;20:1072–86. https://doi.org/10.1007/s10495-015-1132-2
Du XX, Li YJ, Wu CL, Zhou JH, Han Y, Sui H, et al. Initiation of apoptosis, cell cycle arrest and autophagy of esophageal cancer cells by dihydroartemisinin. Biomed Pharmacother. 2013;67:417–24. https://doi.org/10.1016/j.biopha.2013.01.013
Celik S, Akcora D, Ozkan T, Varol N, Aydos S, Sunguroglu A. Methylation analysis of the DAPK1 gene in imatinib-resistant chronic myeloid leukemia patients. Oncol Lett. 2015;9:399–404. https://doi.org/10.3892/ol.2014.2677
Eisenberg-Lerner A, Kimchi A. PKD is a kinase of Vps34 that mediates ROS-induced autophagy downstream of DAPk. Cell Death Differ. 2012;19:788–97. https://doi.org/10.1038/cdd.2011.149
Inbal B, Bialik S, Sabanay I, Shani G, Kimchi A. DAP kinase and DRP-1 mediate membrane blebbing and the formation of autophagic vesicles during programmed cell death. J Cell Biol. 2002;157:455–68. https://doi.org/10.1083/jcb.200109094
Thongchot S, Vidoni C, Ferraresi A, Loilome W, Yongvanit P, Namwat N, et al. Dihydroartemisinin induces apoptosis and autophagy-dependent cell death in cholangiocarcinoma through a DAPK1-BECLIN1 pathway. Mol Carcinog. 2018;57:1735–50. https://doi.org/10.1002/mc.22893
Cao C, Subhawong T, Albert JM, Kim KW, Geng L, Sekhar KR, et al. Inhibition of mammalian target of rapamycin or apoptotic pathway induces autophagy and radiosensitizes PTEN null prostate cancer cells. Cancer Res. 2006;66:10040–7. https://doi.org/10.1158/0008-5472.Can-06-0802
Alonso MM, Jiang H, Yokoyama T, Xu J, Bekele NB, Lang FF, et al. Delta-24-RGD in combination with RAD001 induces enhanced anti-glioma effect via autophagic cell death. Mol Ther. 2008;16:487–93. https://doi.org/10.1038/sj.mt.6300400
Sabatini DM. mTOR and cancer: insights into a complex relationship. Nat Rev Cancer. 2006;6:729–34. https://doi.org/10.1038/nrc1974
Di Paolo S, Teutonico A, Ranieri E, Gesualdo L, Schena PF. Monitoring antitumor efficacy of rapamycin in Kaposi sarcoma. Am J Kidney Dis. 2007;49:462–70. https://doi.org/10.1053/j.ajkd.2006.11.037
Moreno A, Akcakanat A, Munsell MF, Soni A, Yao JC, Meric-Bernstam F. Antitumor activity of rapamycin and octreotide as single agents or in combination in neuroendocrine tumors. Endocr Relat Cancer. 2008;15:257–66. https://doi.org/10.1677/erc-07-0202
Motzer RJ, Escudier B, Oudard S, Hutson TE, Porta C, Bracarda S, et al. Efficacy of everolimus in advanced renal cell carcinoma: a double-blind, randomised, placebo-controlled phase III trial. Lancet. 2008;372:449–56. https://doi.org/10.1016/s0140-6736(08)61039-9
Shinohara ET, Cao C, Niermann K, Mu Y, Zeng F, Hallahan DE, et al. Enhanced radiation damage of tumor vasculature by mTOR inhibitors. Oncogene. 2005;24:5414–22. https://doi.org/10.1038/sj.onc.1208715
Wang H, Li D, Li X, Ou X, Liu S, Zhang Y, et al. Mammalian target of rapamycin inhibitor RAD001 sensitizes endometrial cancer cells to paclitaxel-induced apoptosis via the induction of autophagy. Oncol Lett. 2016;12:5029–35. https://doi.org/10.3892/ol.2016.5338
Felip I, Moiola CP, Megino-Luque C, Lopez-Gil C, Cabrera S, Solé-Sánchez S, et al. Therapeutic potential of the new TRIB3-mediated cell autophagy anticancer drug ABTL0812 in endometrial cancer. Gynecol Oncol. 2019;153:425–35. https://doi.org/10.1016/j.ygyno.2019.03.002
López-Plana A, Fernández-Nogueira P, Muñoz-Guardiola P, Solé-Sánchez S, Megías-Roda E, Pérez-Montoyo H, et al. The novel proautophagy anticancer drug ABTL0812 potentiates chemotherapy in adenocarcinoma and squamous nonsmall cell lung cancer. Int J Cancer. 2020;147:1163–79. https://doi.org/10.1002/ijc.32865
Erazo T, Lorente M, López-Plana A, Muñoz-Guardiola P, Fernández-Nogueira P, García-Martínez JA, et al. The new antitumor drug ABTL0812 inhibits the Akt/mTORC1 axis by upregulating tribbles-3 pseudokinase. Clin Cancer Res. 2016;22:2508–19. https://doi.org/10.1158/1078-0432.Ccr-15-1808
Muñoz-Guardiola P, Casas J, Megías-Roda E, Solé S, Perez-Montoyo H, Yeste-Velasco M, et al. The anti-cancer drug ABTL0812 induces ER stress-mediated cytotoxic autophagy by increasing dihydroceramide levels in cancer cells. Autophagy. 2021;17:1349–66. https://doi.org/10.1080/15548627.2020.1761651
Davidson MB, Peters AL. An overview of metformin in the treatment of type 2 diabetes mellitus. Am J Med. 1997;102:99–110. https://doi.org/10.1016/s0002-9343(96)00353-1
Hu S, Ouyang Q, Cheng Q, Wang J, Feng F, Qiao L, et al. Phenformin inhibits cell proliferation and induces cell apoptosis and autophagy in cholangiocarcinoma. Mol Med Rep. 2018;17:6028–32. https://doi.org/10.3892/mmr.2018.8573
Chen MH, Yen CC, Cheng CT, Wu RC, Huang SC, Yu CS, et al. Identification of SPHK1 as a therapeutic target and marker of poor prognosis in cholangiocarcinoma. Oncotarget. 2015;6:23594–608. https://doi.org/10.18632/oncotarget.4335
Datta A, Loo SY, Huang B, Wong L, Tan SS, Tan TZ, et al. SPHK1 regulates proliferation and survival responses in triple-negative breast cancer. Oncotarget. 2014;5:5920–33. https://doi.org/10.18632/oncotarget.1874
Le Scolan E, Pchejetski D, Banno Y, Denis N, Mayeux P, Vainchenker W, et al. Overexpression of sphingosine kinase 1 is an oncogenic event in erythroleukemic progression. Blood. 2005;106:1808–16. https://doi.org/10.1182/blood-2004-12-4832
Li W, Yu CP, Xia JT, Zhang L, Weng GX, Zheng HQ, et al. Sphingosine kinase 1 is associated with gastric cancer progression and poor survival of patients. Clin Cancer Res. 2009;15:1393–9. https://doi.org/10.1158/1078-0432.Ccr-08-1158
Wallington-Beddoe CT, Powell JA, Tong D, Pitson SM, Bradstock KF, Bendall LJ. Sphingosine kinase 2 promotes acute lymphoblastic leukemia by enhancing MYC expression. Cancer Res. 2014;74:2803–15. https://doi.org/10.1158/0008-5472.Can-13-2732
Liu R, Zhao R, Zhou X, Liang X, Campbell DJ, Zhang X, et al. Conjugated bile acids promote cholangiocarcinoma cell invasive growth through activation of sphingosine 1-phosphate receptor 2. Hepatology. 2014;60:908–18. https://doi.org/10.1002/hep.27085
Ding X, Chaiteerakij R, Moser CD, Shaleh H, Boakye J, Chen G, et al. Antitumor effect of the novel sphingosine kinase 2 inhibitor ABC294640 is enhanced by inhibition of autophagy and by sorafenib in human cholangiocarcinoma cells. Oncotarget. 2016;7:20080–92. https://doi.org/10.18632/oncotarget.7914
Zhang Y, Jiang L, Su P, Yu T, Ma Z, Liu Y, et al. Urolithin A suppresses tumor progression and induces autophagy in gastric cancer via the PI3K/Akt/mTOR pathway. Drug Dev Res. 2023;84:172–84. https://doi.org/10.1002/ddr.22021
Gu Y, Fei Z, Zhu R. miR-21 modulates cisplatin resistance of gastric cancer cells by inhibiting autophagy via the PI3K/Akt/mTOR pathway. Anticancer Drugs. 2020;31:385–93. https://doi.org/10.1097/cad.0000000000000886
Zhang Y, Zhang Y, Halemahebai G, Tian L, Dong H, Aisker G. Urolithin A, a pomegranate metabolite, protects pancreatic β cells from apoptosis by activating autophagy. J Ethnopharmacol. 2021;272:113628. https://doi.org/10.1016/j.jep.2020.113628
Wu TT, Peters AA, Tan PT, Roberts-Thomson SJ, Monteith GR. Consequences of activating the calcium-permeable ion channel TRPV1 in breast cancer cells with regulated TRPV1 expression. Cell Calcium. 2014;56:59–67. https://doi.org/10.1016/j.ceca.2014.04.006
Huang SP, Chen JC, Wu CC, Chen CT, Tang NY, Ho YT, et al. Capsaicin-induced apoptosis in human hepatoma HepG2 cells. Anticancer Res. 2009;29:165–74.
Díaz-Laviada I, Rodríguez-Henche N. The potential antitumor effects of capsaicin. Prog Drug Res. 2014;68:181–208. https://doi.org/10.1007/978-3-0348-0828-6_8
Hong ZF, Zhao WX, Yin ZY, Xie CR, Xu YP, Chi XQ, et al. Capsaicin enhances the drug sensitivity of cholangiocarcinoma through the inhibition of chemotherapeutic-induced autophagy. PLoS ONE. 2015;10:e0121538. https://doi.org/10.1371/journal.pone.0121538
Qu X, Sheng J, Shen L, Su J, Xu Y, Xie Q, et al. Autophagy inhibitor chloroquine increases sensitivity to cisplatin in QBC939 cholangiocarcinoma cells by mitochondrial ROS. PLoS ONE. 2017;12:e0173712. https://doi.org/10.1371/journal.pone.0173712
Jia B, Xue Y, Yan X, Li J, Wu Y, Guo R, et al. Autophagy inhibitor chloroquine induces apoptosis of cholangiocarcinoma cells via endoplasmic reticulum stress. Oncol Lett. 2018;16:3509–16. https://doi.org/10.3892/ol.2018.9131
Nitta T, Mitsuhashi T, Hatanaka Y, Miyamoto M, Oba K, Tsuchikawa T, et al. Prognostic significance of epithelial-mesenchymal transition-related markers in extrahepatic cholangiocarcinoma: comprehensive immunohistochemical study using a tissue microarray. Br J Cancer. 2014;111:1363–72. https://doi.org/10.1038/bjc.2014.415
Dragowska WH, Weppler SA, Wang JC, Wong LY, Kapanen AI, Rawji JS, et al. Induction of autophagy is an early response to gefitinib and a potential therapeutic target in breast cancer. PLoS ONE. 2013;8:e76503. https://doi.org/10.1371/journal.pone.0076503
Maycotte P, Aryal S, Cummings CT, Thorburn J, Morgan MJ, Thorburn A. Chloroquine sensitizes breast cancer cells to chemotherapy independent of autophagy. Autophagy. 2012;8:200–12. https://doi.org/10.4161/auto.8.2.18554
Klionsky DJ, Abdelmohsen K, Abe A, Abedin MJ, Abeliovich H, Acevedo Arozena A, et al. Guidelines for the use and interpretation of assays for monitoring autophagy. Autophagy. 2016;12:1–222.
Mauthe M, Orhon I, Rocchi C, Zhou X, Luhr M, Hijlkema KJ, et al. Chloroquine inhibits autophagic flux by decreasing autophagosome-lysosome fusion. Autophagy. 2018;14:1435–55. https://doi.org/10.1080/15548627.2018.1474314
Chen J, Pan Q, Bai Y, Chen X, Zhou Y. Hydroxychloroquine induces apoptosis in cholangiocarcinoma via reactive oxygen species accumulation induced by autophagy inhibition. Front Mol Biosci. 2021;8:720370. https://doi.org/10.3389/fmolb.2021.720370
Puissant A, Robert G, Fenouille N, Luciano F, Cassuto JP, Raynaud S, et al. Resveratrol promotes autophagic cell death in chronic myelogenous leukemia cells via JNK-mediated p62/SQSTM1 expression and AMPK activation. Cancer Res. 2010;70:1042–52. https://doi.org/10.1158/0008-5472.Can-09-3537
Signorelli P, Munoz-Olaya JM, Gagliostro V, Casas J, Ghidoni R, Fabriàs G. Dihydroceramide intracellular increase in response to resveratrol treatment mediates autophagy in gastric cancer cells. Cancer Lett. 2009;282:238–43. https://doi.org/10.1016/j.canlet.2009.03.020
Lao Y, Wan G, Liu Z, Wang X, Ruan P, Xu W, et al. The natural compound oblongifolin C inhibits autophagic flux and enhances antitumor efficacy of nutrient deprivation. Autophagy. 2014;10:736–49. https://doi.org/10.4161/auto.28034
Zhang A, He W, Shi H, Huang X, Ji G. Natural compound oblongifolin C inhibits autophagic flux, and induces apoptosis and mitochondrial dysfunction in human cholangiocarcinoma QBC939 cells. Mol Med Rep. 2016;14:3179–83. https://doi.org/10.3892/mmr.2016.5591
Klose J, Eissele J, Volz C, Schmitt S, Ritter A, Ying S, et al. Salinomycin inhibits metastatic colorectal cancer growth and interferes with Wnt/β-catenin signaling in CD133(+) human colorectal cancer cells. BMC Cancer. 2016;16:896 https://doi.org/10.1186/s12885-016-2879-8
Klose J, Stankov MV, Kleine M, Ramackers W, Panayotova-Dimitrova D, Jäger MD, et al. Inhibition of autophagic flux by salinomycin results in anti-cancer effect in hepatocellular carcinoma cells. PLoS ONE. 2014;9:e95970 https://doi.org/10.1371/journal.pone.0095970
Lu D, Choi MY, Yu J, Castro JE, Kipps TJ, Carson DA. Salinomycin inhibits Wnt signaling and selectively induces apoptosis in chronic lymphocytic leukemia cells. Proc Natl Acad Sci USA. 2011;108:13253–7. https://doi.org/10.1073/pnas.1110431108
Klose J, Guerlevik E, Trostel T, Kühnel F, Schmidt T, Schneider M, et al. Salinomycin inhibits cholangiocarcinoma growth by inhibition of autophagic flux. Oncotarget. 2018;9:3619–30. https://doi.org/10.18632/oncotarget.23339
Glick D, Barth S, Macleod KF. Autophagy: cellular and molecular mechanisms. J Pathol. 2010;221:3–12. https://doi.org/10.1002/path.2697
Yu L, Chen Y, Tooze SA. Autophagy pathway: cellular and molecular mechanisms. Autophagy. 2018;14:207–15. https://doi.org/10.1080/15548627.2017.1378838
Liu G, Pei F, Yang F, Li L, Amin AD, Liu S, et al. Role of autophagy and apoptosis in non-small-cell lung cancer. Int J Mol Sci. 2017;18. https://doi.org/10.3390/ijms18020367
Jia W, Pua HH, Li QJ, He YW. Autophagy regulates endoplasmic reticulum homeostasis and calcium mobilization in T lymphocytes. J Immunol. 2011;186:1564–74. https://doi.org/10.4049/jimmunol.1001822
Miller BC, Zhao Z, Stephenson LM, Cadwell K, Pua HH, Lee HK, et al. The autophagy gene ATG5 plays an essential role in B lymphocyte development. Autophagy. 2008;4:309–14. https://doi.org/10.4161/auto.5474
Xu WS, Dang YY, Guo JJ, Wu GS, Lu JJ, Chen XP, et al. Furanodiene induces endoplasmic reticulum stress and presents antiproliferative activities in lung cancer cells. Evid Based Complement Alternat Med. 2012;2012:426521. https://doi.org/10.1155/2012/426521
Wang Y, Zhang J, Huang ZH, Huang XH, Zheng WB, Yin XF, et al. Isodeoxyelephantopin induces protective autophagy in lung cancer cells via Nrf2-p62-keap1 feedback loop. Cell Death Dis. 2017;8:e2876. https://doi.org/10.1038/cddis.2017.265
Jiao YN, Wu LN, Xue D, Liu XJ, Tian ZH, Jiang ST, et al. Marsdenia tenacissima extract induces apoptosis and suppresses autophagy through ERK activation in lung cancer cells. Cancer Cell Int. 2018;18:149 https://doi.org/10.1186/s12935-018-0646-4
Yu N, Xiong Y, Wang C. Bu-Zhong-Yi-Qi decoction, the water extract of Chinese traditional herbal medicine, enhances cisplatin cytotoxicity in A549/DDP cells through induction of apoptosis and autophagy. Biomed Res Int. 2017;2017:3692797. https://doi.org/10.1155/2017/3692797
Zhou Y, Rucker EB 3rd, Zhou BP. Autophagy regulation in the development and treatment of breast cancer. Acta Biochim Biophys Sin. 2016;48:60–74. https://doi.org/10.1093/abbs/gmv119
Funding
This study was supported by grants from the Zhejiang Health Committee (2021ZH003), Hangzhou Science and Technology Commission (202004A14), and the Construction Fund of Medical Key Disciplines of Hangzhou (OO20190001).
Author information
Authors and Affiliations
Contributions
HY and HK: wrote the original paper, data collection and correction. YW: figure settings. JY: provided article ideas and article review.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Approval and consent to publish
All authors have approved this submission to this journal.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Khizar, H., Hu, Y., Wu, Y. et al. The role and implication of autophagy in cholangiocarcinoma. Cell Death Discov. 9, 332 (2023). https://doi.org/10.1038/s41420-023-01631-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-023-01631-7
- Springer Nature Limited