Abstract
Here, we present the case of a 47-year-old woman diagnosed with luminal B breast cancer subtype and provide an in-depth analysis of her gene mutations, chromosomal alterations, mRNA and protein expression changes. We found a point mutation in the FGFR2 gene, which is potentially hyper-activating the receptor function, along with over-expression of its ligand FGF20 due to genomic amplification. The patient also harbors somatic and germline mutations in some mismatch repair (MMR) genes, with a strong MMR mutational signature. The patient displays high microsatellite instability (MSI) and tumor mutational burden (TMB) status and increased levels of CTLA-4 and PD-1 expression. Altogether, these data strongly implicate that aberrant FGFR signaling, and defective MMR system might be involved in the development of this breast tumor. In addition, high MSI and TMB in the context of CTLA-4 and PD-L1 positivity, suggest the potential benefit of immune checkpoint inhibitors. Accurate characterization of molecular subtypes, based on gene mutational and expression profiling analyses, will be certainly helpful for individualized treatment and targeted therapy of breast cancer patients, especially for those subtypes with adverse outcome.
Similar content being viewed by others
Introduction
Breast cancer is a global public health concern with a high incidence among women, leading to substantial morbidity and mortality [1, 2]. The treatment approach for breast cancer takes into account various factors such as cancer stage and biology, and patient preferences and tolerance. Surgery is the primary treatment option, complemented by radiation, chemotherapy, endocrine therapy, targeted therapy, and immunotherapy [3, 4]. It is essential to recognize that breast cancer exhibits significant heterogeneity in terms of its morphological and molecular characteristics [5, 6]. Gaining a comprehensive understanding of the distinct features associated with breast cancer is vital for accurate diagnosis, optimal treatment selection, and prognostic predictions for patients.
Breast cancer arises in the epithelium of the ducts (85%) or lobules (15%) in the glandular tissue of the breast. According to the PAM50 signature, breast cancer has been classified into five molecular subtypes: Luminal A [estrogen receptor (ER)+ and progesterone receptor (PR)+, HER2-, low levels of Ki-67], Luminal B [ER+, HER2-, and either Ki-67 high or PR low], Luminal B-like (ER+, HER2+, any Ki-67, and any PR, HER2-enriched [ER-, PR- and HER2+], triple negative (TNBC) or basal-like [ER−, PR-, and HER2-] [7, 8]. Heterogeneity of breast cancer makes its treatment complicated, involving a combination of different modalities including surgery, radiotherapy, chemotherapy, hormonal therapy, or targeted biological therapies [9]. Multiple genetic aberrations are present in breast cancer patients [10, 11], ranging from p53 [12,13,14,15], components of the apoptotic machinery [16] and of the ubiquitin proteasome system [17,18,19], transcriptional modifiers [20,21,22], to hypoxic or metabolic regulators [23,24,25,26]. These factors determine cancer progression [27,28,29] and affect therapeutic responses [30, 31]. Moreover, further analyzes of gene expression or mutational status might reveal additional tumor heterogeneity among subtypes and probably help to develop new personalized treatment strategies for individual patients.
Luminal B breast cancer typically affects women who are post-menopausal but can also occur in younger women; these tumors are more aggressive than subtype A ones. They usually display higher grades and have worse prognosis because of high expression of proliferation-related genes. They also benefit from hormone therapy along with chemotherapy [32]. Tumors with a high mutational load, for instance because of DNA mismatch repair deficiency, are generally highly responsive to immune checkpoint blockade [33]. Luminal-type breast cancers are normally less immunogenic than TNBCs because of a lower rate of tumor specific mutations and subsequent neoantigen load. They are therefore less frequently characterized by the presence of tumor-infiltrating lymphocytes that are often associated with favorable prognosis in TNBC and HER2+ breast cancers.
In this case report, we present a detailed analysis of a luminal B breast cancer case, with a high proliferation rate. Genome-wide mutational and global gene expression analyses were performed to characterize the molecular profile of this neoplasia. Microsatellite instability, cancer mutational signatures, and the tumor mutational burden were also evaluated. The aim of this case report is to provide insights into the distinct molecular characteristics of breast cancer and their potential clinical significance. By providing a comprehensive multi-omics analysis of this breast cancer patient, we have proposed potential prognostic indicators for the patient as well as avenues for therapeutic interventions. Ultimately, this knowledge will contribute to the continuous endeavors in combating breast cancer and enhancing the well-being of those affected by the disease.
Case presentation
Case narration
A female, 47-year-old asymptomatic patient with a BMI of 25.7, non-smoker, non-vegetarian and normal defecation was diagnosed with breast cancer (Table 1). Her anamnesis included menarche at 10 years old and menopause at 40 years old because of bilateral oophorectomy. She had one pregnancy at the age of 30 and did not breastfeed. There is no history of hormone use before menopause. The histo-pathological diagnosis was breast invasive ductal carcinoma NST, tumor grading G3 (Fig. 1A, B). The TNM staging was pT2 pN0 (sn). The patient did not receive neoadjuvant therapy and underwent complete radicality. The patient was also diagnosed with autoimmune thyroiditis.
A Haematoxylin and eosin slide shows a ductal infiltrating breast carcinoma (G3) (scale bar represents 200 µm). B High magnification of panel A shows high cellular pleomorphism, absence of tubules and numerous mitoses (scale bar represents 20 µm). C Estrogen receptor expression in more than 95% of breast cancer cells (scale bar represents 200 µm). D Progesterone receptor expression in about 1% of breast cancer cells (scale bar represents 200 µm). E Image displays high proliferation index (Ki67 > 35%) (scale bar represents 200 µm). F In situ hybridization analysis of HER2 status demonstrates the amplification of HER2 gene (ErbB2/CEN17 ratio >2) (scale bar represents 10 µm). G Expression mRNA levels (TPM) of estrogen receptor 1 (ESR1), progesterone receptor (PR1), ERBB2 receptor tyrosine kinase 2 (HER2) and proliferation marker KI-67 (MKI67) for the patient (red triangle) and the clinical cohort (blue boxplot).
The analysis of the prognostic and predictive biomarkers by immunohistochemistry (IHC) and fluorescence in situ hybridization (FISH) enabled the classification of the breast cancer as luminal B (Fig. 1C–F). In fact, the tumor was found to be ER+ ( > 95%), PR+ (1%) and HER2 score 2 at IHC. FISH analysis demonstrated the amplification status of HER2 (ErbB2/CEN17 ratio >2 FISH). Proliferation indices evaluated in terms of percentage of Ki67 positive cancer cells were high (Ki-67 > 35%). Of note, ER1 and HER2 expression assessed by RNA-seq is consistent with IHC analysis. As shown in Fig. 1G, the patient indeed showed upregulation of ER1 and down-regulation of HER2 mRNAs as compared to the clinical cohort (354 ER+HER2- cases out of total 580 breast cancer patients). Intrinsic subtyping was carried out by using both the research-based 50-gene prediction analysis of a microarray (PAM50) subtype predictor and Absolute Intrinsic Molecular Subtyping. Overall, the diagnosis of luminal B subtype was consistent with both histopathological and molecular data.
Genetic mutation analysis
Breast cancers seem to be usually caused by the accumulation of multiple mutations and genetic aberrations that correlate with different treatment strategy and clinical outcome of patients [34,35,36]. In accordance with the high heterogeneity of breast cancer, the gene mutation pattern also varies between different subtypes, and can be unique for every single patient [5, 37,38,39,40,41]. Luminal/ER-positive subtypes are reported to be the most heterogeneous in terms of mutation spectrum, copy number changes and patient outcomes [8, 42].
A comprehensive genomic profiling revealed a high frequency of somatic mutations with single base substitutions in the patient (Table 2). Importantly, among them, there are mutations with available targeted treatment in different phases of clinical trials. It is worth mentioned that we found a new mutation (Asn550Asp) at the FGFR2 locus, a tyrosine kinase receptor that mediates FGF signaling. Although a different amino-acidic substitution (Asp550Lys) has been identified in breast cancer patients [43], this particular mutation has never been previously described in mammary tumors. The mutation is located within the tyrosine kinase domain (Fig. 2A), potentially leading to aberrant activation of the receptor. Of note, one ligand of FGFR2, FGF20 was also found to be amplified in the tumor tissue (Table 3). Accordingly, we observed an increase in FGF20 expression at mRNA level in this patient (Fig. 2B), suggesting that both hyper-activated FGFR2 and increased levels of its ligand could amplify the FGFR2 signaling. Noteworthy, FGF20 is found amplified and to lesser extent deleted in approximately 2% of breast cancer cases from METABRIC dataset (Fig. 2C).
A Schematic structural features of the FGFR2 protein. Patient’s mutations are indicated by an arrow. Data were obtained from cBioPortal. B Expression of FGF20 in the patient (red triangle) compared to the clinical cohort (RNA-Seq). C FGFR2 and FGF20 genomic alterations in the METABRIC dataset. Samples were clustered based on their molecular subtype. Data were obtained from cBioPortal.
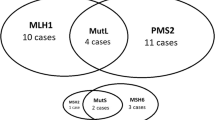
We also identified a somatic mutation in the MSH6 gene (Arg1024Trp) with a high variant allele frequency (VAF) (50%) and two germline mutations in the MSH2 and PMS2 genes (Table 2). These three genes, together with MLH1 constitutes the mismatch repair (MMR) system that is in charge of recognizing and repairing deletions, insertion, and misincorporation of bases during DNA replication and recombination. MSH2 mutations have been associated with an increased risk of developing several tumors including colon, breast, ovarian, and endometrial cancers [44], however these mutations in breast cancer are quite rare, while amplifications are more frequent with 1% of cases showing overlapping amplification of MSH6 and MSH2 and up to 1.5% PMS2 amplification in other cases from METABRIC database (Fig. 3A). The mutation in the MSH6 gene has been already reported in colorectal cancer patients in which was associated with medium/high TMB [45]. In this patient, the mutation c.20754G>A (Gly692Glu) of the MSH2 gene lies within 5th domain (V) (Fig. 3B), is novel and has not been reported previously. We therefore put forward the hypothesis that a mutation within this domain may inhibit MMR [46]. We also found a deletion of the MSH2 gene (Table 3). Although already described in other tumors as a somatic variant [47], to the best of our knowledge, this PMS2 germline mutation has never been previously reported in breast cancer patients. The PMS2 alteration c.1243G>A (Val415Met), is a transition mutation located between the DNA mismatch repair and the dimerization domains (Fig. 3C). Consistent with the observed alterations, mutational signature analysis revealed a strong MMR signature (Fig. 4A, B).
A MSH2, MSH6 and PMS2 genomic alterations in the METABRIC dataset. Samples were clustered based on their molecular subtype. Data were obtained from cBioPortal. B Schematic structural features of the MSH2 protein. Patient’s mutation is indicated by an arrow. Data were obtained from cBioPortal. C Schematic structural representation of the PMS2 protein. Patient’s mutation is indicated by an arrow. Data were obtained from cBioPortal.
The patient (red triangle) is compared to the clinical cohort (blue boxplot). A, B Mutational contribution of MMR related signatures. C MSI score (MSI High: score >0.901). The patient is observed as having MSI High status. D The patient has a higher TMB as compared to the cohort median (>95. percentile).
Since the MMR system assists in the maintenance of the genomic fidelity and reduces general gene mutations, its impairment facilitates high frequency somatic mutations and can lead to the insertion or deletion at microsatellites. Coherently, we observed high TMB (34.5) and MSI (9.5) values in this patient as compared to the clinical cohort (Fig. 4C, D). MMR is closely correlated to treatment selection. Chemotherapy and radiation treatment-induced mutagenesis may be accelerated in patients having deficiency in the MMR system. Some novel mutated genes may be driver genes of cancer, which means that MMR inactivation can lead to disease progression and resistance to chemotherapy and radiotherapy. From this point of view, chemotherapy and radiotherapy is probably risky for patients with MMR. On the other hand, previous studies demonstrated that hypermutation and MMR are associated with high tumor-specific neoantigen burden inducing high T cell infiltration that is therefore predictive of a high response to immune checkpoint inhibitors (ICI) [48,49,50,51].
To find further potential treatment strategies of immune therapy for this case, we assessed the expression of immune checkpoint genes, including cytotoxic T-lymphocyte-associated protein 4 (CTLA4), programmed cell death protein 1 (PD-1), programmed death-ligand 1 (PD-L1), programmed death-ligand 2 (PD-L2) by RNA-sequencing. As shown in Fig. 5, CTLA-4 and PD-1 expression was moderately higher in this patient compared with the median expression in the clinical cohort, while for the other checkpoint genes, the differences were less pronounced.
Furthermore, we also assessed potential risk factors using several risk models. Comprehensive analysis of all these models indicated that this patient is at high risk of recurrence and decreased relapse-free survival. As for therapeutic strategy, the prediction showed increased sensitivity to neoadjuvant paclitaxel plus fluorouracil, adriamycin, and cyclophosphamide chemotherapy, and low risk of tamoxifen resistance.
Discussion
Breast cancer is a highly heterogeneous disease that can be characterized not only by pathological features, but also according to differences in genetic and molecular characteristics. The heterogeneity of breast cancer has significant implications for the diagnosis, treatment, and prognosis of this disease and personalized treatment for individual patients is recommended. Accurate classification of breast cancer subtypes and further molecular diagnosis are helpful to identify potential targets for personalized therapies and determine the most appropriate treatment options.
This patient was diagnosed with Luminal B subtype breast cancer, predicted with high risk of recurrence and decreased relapse-free survival. By performing global genomic and transcriptomic analyses, we proposed new neoadjuvant treatments for this patient based on identified molecular signatures. The genomic alteration analysis can also help in re-subtyping at molecular level Luminal B breast cancer. At first, the risk model showed low risk of tamoxifen resistance, suggesting that tamoxifen could be probably the basic treatment. We found that FGFR2 and its ligand FGF20 are potentially aberrantly activated and over-expressed, due to mutational and genomic alterations respectively. FGFR2 amplification has also been found in other tumors, such as gastric, esophageal, squamous cell lung, liver, and bladder cancers [52]. Drugs targeting FGFR2 have been already used in clinical trials for treating FGFR2 amplified tumors, including Bemarituzumab and AZD 4547(ClinicalTrials.gov). In this case, both FGFR2 and FGF20 are proposed as potential targets.
In conclusion, a high MSI and TMB pattern and loss of wild-type MSH2, MSH6 and PMS2 genes, strongly suggest that, in this patient, the mismatch repair defect might have contributed to the development of breast cancer. High frequencies of somatic mutations may result in increased expression of neoantigens, and tumor-specific T-cell reactivity. Hence, considering the high TMB and MSI-H values, and the MMR signature, the patient would be probably sensitive to ICIs. Immune checkpoint detection indicated that ICI targeting CTLA-4 and PD-1 would be another option of neoadjuvant treatment. Fremd and colleagues reported a case of a woman with hormone receptor-positive, HER2 negative metastatic breast cancer, who achieved a durable complete remission after treatment with pembrolizumab [53]. This finding strongly supports the importance of testing the MMR in breast cancer patients and searching for biomarkers to predict the success of the immune therapy. However, whether targeting FGFR2/FGF20 or immune checkpoint genes should be considered only after much more convinced verification of genes expression at protein level. It is worth mentioning that chemotherapy and radiotherapy should be considered carefully because of the possibility of novel mutations of cancer driver genes, which can lead to selection of treatment-resistant cells.
Altogether, we present a comprehensive multi-omics analysis of a case with luminal B breast cancer and propose potential therapeutic strategies based on identified molecular targets.
Materials and methods
Collection of samples
Tumor tissues were globally collected using a standardized protocol, minimizing the ischemia time until freezing in liquid nitrogen. To ensure the quality of the samples, all tissues were hematoxilin and eosin stained and subjected to a pathological QC. Samples need to be invasive, have a tumor content of ≥30 % and Necrosis ≤30 %. Normal tissues were processed in parallel and need to be free of tumor and representative regarding the tumor tissue to be included.
Approximately 10 mg tissue were taken for nucleic acid extraction and protein lysate preparation each. To account for tumor heterogeneity, pathological QCs were performed on two sections, before and after taking the analysis material. The tissues stay frozen during the entire process.
Immunohistochemical analysis
Approximately 1 × 1 × 0.5 cm of tissue was formalin-fixed and paraffin-embedded. Serial sections were used to evaluate prognostic and predictive biomarkers including ER, PR, Ki67, and HER2 through immunohistochemistry. Briefly, sections were stained using the automated Leica Bond IHC platform (Leica Biosystems, Deer Park, IL). After antigen retrieval, 4-μm thick sections were incubated with the following primary monoclonal antibodies: mouse monoclonal anti-ER (clone 6F11; Leica Biosystems), mouse monoclonal anti-PR (clone 16; Leica Biosystems), mouse monoclonal anti-Ki67 (clone MM1; Leica Biosystems) and mouse monoclonal anti-HER2 (clone CB11, Leica Biosystems). Reactions were revealed using BOND-PRIME Polymer DAB Detection System (Leica Biosystems, Deer Park, IL). Immunohistochemistry was evaluated by two blind pathologists.
Nucleic acid extraction and quality assessment
Frozen tissue slices were mixed with β-mercaptoethanol containing sample buffer and homogenized using the BeadBug system. DNA and RNA were extracted in parallel from the same sample using the Qiagen AllPrep Universal Kit according to the manufacturer’s instructions.
DNA and RNA concentration were quantified using Qubit fluorometer with the Qubit dsDNA BR assay or Qubit RNA BR assay respectively.
DNA and RNA quality were assessed using the Agilent Tapestation with the Agilent Genomic DNA kit or Agilent High-Sensitivity RNA ScreenTape kit respectively. RNAs need to have a RIN ≥ 4 or a DV200 ≥ 60 to be selected for library preparation.
Library preparation and NGS sequencing
Libraries for whole genome sequencing (WGS) were prepared using the PCR-free KAPA Hyper Prep Kit (Roche). For whole transcriptome sequencing, RNA samples were depleted of the ribosomal RNA using the Ribo Zero Kit (Illumina) and library preparation was performed using the TruSeq Stranded Total RNA Kit (Qiagen). For small RNA sequencing the QIAseq miRNA Kit (Qiagen) was used All library preparation kits were used according to manufacturer’s instructions. Sequencing was performed on a NovaSeq6000 system (Illumina).
For WGS, average coverage for tumor samples was ≥60× and ≥30× for normal samples with a total genomic coverage of ≥95%.
Whole transcriptome sequencing datasets have ≥100 million total reads with less than 20% of ribosomal origin and ≥20 million reads mapping to mRNAs according to Ensembl reference. Ribosomal depletion was performed to remove nuclear rRNA and mt-rRNA.
NGS data processing
NGS data were aligned against Grch38 genome assembly. Identification and annotation of short genomic variations in normal sample was done using Haplotype Caller (genome analysis toolkit; GATK) [54]. WGS somatic variation were called using a consensus of Mutect2 [55], Strelka [56], Varscan [57], and Somatic Sniper [58]. Structural variations were called using R packages TitanCNA [59] and DellyCNV [60].
RNA-Seq differential expression was based on normalized readcount data (TPM: transcripts per million).
Bioinformatical analyses
Mutational signatures were calculated using the R package MutationalPatterns [61]. MSI classification was done using R package MSIseq [62]. PAM50 subtyping as well as risk scores were investigated using R package genefu [63]. TMB was calculated as the number of non-synonymous mutations of protein coding genes divided by exome size in Megabases.
References
Nagarajan D, McArdle SEB. Immune landscape of breast cancers. Biomedicines. 2018;6:20.
Makhoul I, Atiq M, Alwbari A, Kieber-Emmons T. Breast cancer immunotherapy: an update. Breast Cancer Basic Clin Res. 2018;12:1178223418774802.
McDonald ES, Clark AS, Tchou J, Zhang P, Freedman GM. Clinical diagnosis and management of breast cancer. J Nucl Med. 2016;57:9S–16S.
Kerr AJ, Dodwell D, McGale P, Holt F, Duane F, Mannu G, et al. Adjuvant and neoadjuvant breast cancer treatments: a systematic review of their effects on mortality. Cancer Treat Rev. 2022;105:102375.
Cancer Genome Atlas N. Comprehensive molecular portraits of human breast tumours. Nature. 2012;490:61–70.
Heng YJ, Lester SC, Tse GM, Factor RE, Allison KH, Collins LC, et al. The molecular basis of breast cancer pathological phenotypes. J Pathol. 2017;241:375–91.
Gao JJ, Swain SM. Luminal a breast cancer and molecular assays: a review. Oncologist. 2018;23:556–65.
Parker JS, Mullins M, Cheang MC, Leung S, Voduc D, Vickery T, et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol. 2009;27:1160–7.
Lukasiewicz S, Czeczelewski M, Forma A, Baj J, Sitarz R, Stanislawek A. Breast cancer-epidemiology, risk factors, classification, prognostic markers, and current treatment strategies-an updated review. Cancers. 2021;13:4287.
Marchetti P, Antonov A, Anemona L, Vangapandou C, Montanaro M, Botticelli A, et al. New immunological potential markers for triple negative breast cancer: IL18R1, CD53, TRIM, Jaw1, LTB, PTPRCAP. Discov Oncol. 2021;12:6.
Romera-Giner S, Andreu Martinez Z, Garcia-Garcia F, Hidalgo MR. Common pathways and functional profiles reveal underlying patterns in Breast, Kidney and Lung cancers. Biol direct. 2021;16:9.
Panatta E, Zampieri C, Melino G, Amelio I. Understanding p53 tumour suppressor network. Biol Direct. 2021;16:14.
Panatta E, Butera A, Celardo I, Leist M, Melino G, Amelio I. p53 regulates expression of nuclear envelope components in cancer cells. Biol Direct. 2022;17:38.
Butera A, Roy M, Zampieri C, Mammarella E, Panatta E, Melino G, et al. p53-driven lipidome influences non-cell-autonomous lysophospholipids in pancreatic cancer. Biol Direct. 2022;17:6.
Rozenberg JM, Zvereva S, Dalina A, Blatov I, Zubarev I, Luppov D, et al. The p53 family member p73 in the regulation of cell stress response. Biol Direct. 2021;16:23.
Vitale I, Pietrocola F, Guilbaud E, Aaronson SA, Abrams JM, Adam D, et al. Apoptotic cell death in disease-current understanding of the NCCD 2023. Cell Death Differ. 2023;30:1097–154.
Zhao Y, Huang X, Zhu D, Wei M, Luo J, Yu S, et al. Deubiquitinase OTUD6A promotes breast cancer progression by increasing TopBP1 stability and rendering tumor cells resistant to DNA-damaging therapy. Cell Death Differ. 2022;29:2531–44.
Yin X, Teng X, Ma T, Yang T, Zhang J, Huo M, et al. RUNX2 recruits the NuRD(MTA1)/CRL4B complex to promote breast cancer progression and bone metastasis. Cell Death Differ. 2022;29:2203–17.
Ren C, Han X, Lu C, Yang T, Qiao P, Sun Y, et al. Ubiquitination of NF-kappaB p65 by FBXW2 suppresses breast cancer stemness, tumorigenesis, and paclitaxel resistance. Cell Death Differ. 2022;29:381–92.
Jiang D, Qiu T, Peng J, Li S, Tala, Ren W, et al. YB-1 is a positive regulator of KLF5 transcription factor in basal-like breast cancer. Cell Death Differ. 2022;29:1283–95.
Kudo M, Zalles N, Distefano R, Nigita G, Veneziano D, Gasparini P, et al. Synergistic apoptotic effect of miR-183-5p and Polo-Like kinase 1 inhibitor NMS-P937 in breast cancer cells. Cell Death Differ. 2022;29:407–19.
Qian J, Lei X, Sun Y, Zheng L, Li J, Zhang S, et al. Long non-coding RNA SNHG8 enhances triple-negative breast cancer cell proliferation and migration by regulating the miR-335-5p/PYGO2 axis. Biol Direct. 2021;16:13.
Yan Y, He M, Zhao L, Wu H, Zhao Y, Han L, et al. A novel HIF-2alpha targeted inhibitor suppresses hypoxia-induced breast cancer stemness via SOD2-mtROS-PDI/GPR78-UPR(ER) axis. Cell Death Differ. 2022;29:1769–89.
Han X, Ren C, Lu C, Qiao P, Yang T, Yu Z. Deubiquitination of MYC by OTUB1 contributes to HK2 mediated glycolysis and breast tumorigenesis. Cell Death Differ. 2022;29:1864–73.
Liu YY, Liu HY, Yu TJ, Lu Q, Zhang FL, Liu GY, et al. O-GlcNAcylation of MORC2 at threonine 556 by OGT couples TGF-beta signaling to breast cancer progression. Cell Death Differ. 2022;29:861–73.
Xie W, Jiang Q, Wu X, Wang L, Gao B, Sun Z, et al. IKBKE phosphorylates and stabilizes Snail to promote breast cancer invasion and metastasis. Cell Death Differ. 2022;29:1528–40.
Cassandri M, Butera A, Amelio I, Lena AM, Montanaro M, Mauriello A, et al. ZNF750 represses breast cancer invasion via epigenetic control of prometastatic genes. Oncogene. 2020;39:4331–43.
Memmi EM, Sanarico AG, Giacobbe A, Peschiaroli A, Frezza V, Cicalese A, et al. p63 Sustains self-renewal of mammary cancer stem cells through regulation of Sonic Hedgehog signaling. Proc Natl Acad Sci USA. 2015;112:3499–504.
Melino G, Memmi EM, Pelicci PG, Bernassola F. Maintaining epithelial stemness with p63. Sci Signal. 2015;8:re9.
Zhang J, Zhang G, Zhang W, Bai L, Wang L, Li T, et al. Loss of RBMS1 promotes anti-tumor immunity through enabling PD-L1 checkpoint blockade in triple-negative breast cancer. Cell Death Differ. 2022;29:2247–61.
Butera A, Cassandri M, Rugolo F, Agostini M, Melino G. The ZNF750-RAC1 axis as potential prognostic factor for breast cancer. Cell Death Discov. 2020;6:135.
Lafci O, Celepli P, Seher Oztekin P, Kosar PN. DCE-MRI radiomics analysis in differentiating luminal A and luminal B breast cancer molecular subtypes. Acad Radiol. 2023;30:22–9.
Chan TA, Wolchok JD, Snyder A. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N Engl J Med. 2015;373:1984.
Yadav S, Couch FJ. Germline genetic testing for breast cancer risk: the past, present, and future. Am Soc Clin Oncol Educ book Am Soc Clin Oncol Annu Meet. 2019;39:61–74.
Gallo M, Paludi D, Cicero DO, Chiovitti K, Millo E, Salis A, et al. Identification of a conserved N-capping box important for the structural autonomy of the prion alpha 3-helix: the disease associated D202N mutation destabilizes the helical conformation. Int J Immunopathol Pharmacol. 2005;18:95–112.
Melino S, Nepravishta R, Bellomaria A, Di Marco S, Paci M. Nucleic acid binding of the RTN1-C C-terminal region: toward the functional role of a reticulon protein. Biochemistry. 2009;48:242–53.
Yang L, Ye F, Bao L, Zhou X, Wang Z, Hu P, et al. Somatic alterations of TP53, ERBB2, PIK3CA and CCND1 are associated with chemosensitivity for breast cancers. Cancer Sci. 2019;110:1389–400.
Garrido-Castro AC, Lin NU, Polyak K. Insights into molecular classifications of triple-negative breast cancer: improving patient selection for treatment. Cancer Discov. 2019;9:176–98.
Slavin TP, Maxwell KN, Lilyquist J, Vijai J, Neuhausen SL, Hart SN, et al. The contribution of pathogenic variants in breast cancer susceptibility genes to familial breast cancer risk. NPJ Breast Cancer. 2017;3:22.
Favaloro B, Tamburro A, Angelucci S, Luca AD, Melino S, di Ilio C, et al. Molecular cloning, expression and site-directed mutagenesis of glutathione S-transferase from Ochrobactrum anthropi. Biochem J. 1998;335:573–9.
Angelucci S, Sacchetta P, Moio P, Melino S, Petruzzelli R, Gervasi P, et al. Purification and characterization of glutathione transferases from the sea bass (Dicentrarchus labrax) liver. Arch Biochem Biophys. 2000;373:435–41.
Paik S, Shak S, Tang G, Kim C, Baker J, Cronin M, et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med. 2004;351:2817–26.
Lei H, Deng CX. Fibroblast growth factor receptor 2 signaling in breast cancer. Int J Biol Sci. 2017;13:1163–71.
Westenend PJ, Schutte R, Hoogmans MM, Wagner A, Dinjens WN. Breast cancer in an MSH2 gene mutation carrier. Hum Pathol. 2005;36:1322–6.
Riviere P, Goodman AM, Okamura R, Barkauskas DA, Whitchurch TJ, Lee S, et al. High tumor mutational burden correlates with longer survival in immunotherapy-naive patients with diverse cancers. Mol Cancer Ther. 2020;19:2139–45.
Edelbrock MA, Kaliyaperumal S, Williams KJ. Structural, molecular and cellular functions of MSH2 and MSH6 during DNA mismatch repair, damage signaling and other noncanonical activities. Mutat Res. 2013;743-744:53–66.
Vargas-Parra GM, Gonzalez-Acosta M, Thompson BA, Gomez C, Fernandez A, Damaso E, et al. Elucidating the molecular basis of MSH2-deficient tumors by combined germline and somatic analysis. Int J Cancer. 2017;141:1365–80.
Matsushita H, Vesely MD, Koboldt DC, Rickert CG, Uppaluri R, Magrini VJ, et al. Cancer exome analysis reveals a T-cell-dependent mechanism of cancer immunoediting. Nature. 2012;482:400–4.
DuPage M, Mazumdar C, Schmidt LM, Cheung AF, Jacks T. Expression of tumour-specific antigens underlies cancer immunoediting. Nature. 2012;482:405–9.
Rooney MS, Shukla SA, Wu CJ, Getz G, Hacohen N. Molecular and genetic properties of tumors associated with local immune cytolytic activity. Cell. 2015;160:48–61.
Giannakis M, Mu XJ, Shukla SA, Qian ZR, Cohen O, Nishihara R, et al. Genomic correlates of immune-cell infiltrates in colorectal carcinoma. Cell Rep. 2016;15:857–65.
Consortium APG. AACR Project GENIE: powering precision medicine through an international consortium. Cancer Discov. 2017;7:818–31.
Fremd C, Hlevnjak M, Zapatka M, Zoernig I, Halama N, Fejzibegovic N, et al. Mismatch repair deficiency drives durable complete remission by targeting programmed death receptor 1 in a metastatic luminal breast cancer patient. Breast Care. 2019;14:53–9.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010;20:1297–303.
Geraldine van der Auwera BDOC. Genomics in the cloud: using docker, GATK, and WDL in terra.: O’Reilly Media, Incorporated; USA, 2020.
Kim S, Scheffler K, Halpern AL, Bekritsky MA, Noh E, Kallberg M, et al. Strelka2: fast and accurate calling of germline and somatic variants. Nat Methods. 2018;15:591–4.
Koboldt DC, Chen K, Wylie T, Larson DE, McLellan MD, Mardis ER, et al. VarScan: variant detection in massively parallel sequencing of individual and pooled samples. Bioinformatics. 2009;25:2283–5.
Larson DE, Harris CC, Chen K, Koboldt DC, Abbott TE, Dooling DJ, et al. SomaticSniper: identification of somatic point mutations in whole genome sequencing data. Bioinformatics. 2012;28:311–7.
Ha G, Roth A, Khattra J, Ho J, Yap D, Prentice LM, et al. TITAN: inference of copy number architectures in clonal cell populations from tumor whole-genome sequence data. Genome Res. 2014;24:1881–93.
Rausch T, Zichner T, Schlattl A, Stutz AM, Benes V, Korbel JO. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics. 2012;28:i333–i9.
Manders F, Brandsma AM, de Kanter J, Verheul M, Oka R, van Roosmalen MJ, et al. MutationalPatterns: the one stop shop for the analysis of mutational processes. BMC Genom. 2022;23:134.
Bonneville R, Krook MA, Kautto EA, Miya J, Wing MR, Chen HZ, et al. Landscape of microsatellite instability across 39cancer types. JCO Precis Oncol. 2017;2017:PO.17.00073.
Huang MN, McPherson JR, Cutcutache I, Teh BT, Tan P, Rozen SG. MSIseq: software for assessing microsatellite instability from catalogs of somatic mutations. Sci Rep. 2015;5:13321.
Funding
This work has been supported by the MUR-PNRR M4C2I1.3 PE6 project PE00000019 Heal Italia (CUP: E83C22004670001) to GM, EC, FB. The research leading to these results has received funding from AIRC under IG 2022 - ID. 27366 project – P.I. Melino Gennaro; under IG 2019 - ID. 22206 – P.I. Candi Eleonora; under IG 2019 – 23232 – P.I. Bernassola Francesca. Fondazione Luigi Maria Monti IDI-IRCCS (RC to EC). Work has been also supported by Regione Lazio through LazioInnova Progetto Gruppo di Ricerca n 85-2017-14986; n 33 & 55-2021-T0002E0001.
Author information
Authors and Affiliations
Consortia
Contributions
GM and FB conceived the project, XY, GM, YS, EC, AS, JB, and FB wrote the manuscript; XY, AS, and JB prepared figures. All the authors have approved this submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All the procedures carried out in the research with participation of humans were in compliance with the ethical standards of the institutional and/or national ethics committee and with the Helsinki Declaration of 1964 and its subsequent changes or with comparable ethics standards. Informed voluntary consent was obtained from every participant of the study: approval on 09-2019, number 96-19.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, X., Smirnov, A., Buonomo, O.C. et al. A primary luminal/HER2 negative breast cancer patient with mismatch repair deficiency. Cell Death Discov. 9, 365 (2023). https://doi.org/10.1038/s41420-023-01650-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-023-01650-4
- Springer Nature Limited
This article is cited by
-
Molecular profiling of a bladder cancer with very high tumour mutational burden
Cell Death Discovery (2024)
-
Genomic and transcriptomic profiling of hepatocellular carcinoma reveals a rare molecular subtype
Discover Oncology (2024)
-
Genetically driven predisposition leads to an unusually genomic unstable renal cell carcinoma
Discover Oncology (2024)