Abstract
To date, colorectal cancer (CRC) still has limited therapeutic efficacy and poor prognosis and there is an urgent need for novel targets to improve the outcome of CRC patients. The highly conserved ubiquitination modification mediated by E3 ubiquitin ligases is an important mechanism to regulate the expression and function of tumor promoters or suppressors in CRC. In this review, we provide an overview of E3 ligases in modulating various biological processes in CRC, including proliferation, migration, stemness, metabolism, cell death, differentiation and immune response of CRC cells, emphasizing the pluripotency of E3 ubiquitin ligases. We further focus on the role of E3 ligases in regulating vital cellular signal pathways in CRC, such as Wnt/β-catenin pathway and NF-κB pathway. Additionally, considering the potential of E3 ligases as novel targets in the treatment of CRC, we discuss what aspects of E3 ligases can be utilized and exploited for efficient therapeutic strategies.
Similar content being viewed by others
Facts
-
Colorectal cancer is the third most common cancer diagnosed worldwide, but with limited therapeutic efficacy and poor prognosis.
-
E3 ligases mediated-ubiquitination regulates the expression and function of tumor promoters or suppressors in the development of CRC.
-
E3 ligases play multifaceted roles in various biological processes and several vital signal pathways of CRC in a context-depending manner.
Open questions
-
Can E3 ligases be efficiently targeted to improve the treatment and prognosis of CRC?
-
How can we identify the bi-functional E3 ligases in CRC and amplify their tumor suppressive effects?
-
What is the mechanism that determines E3 ligases to accurately recognize and ubiquitinate different substrates in CRC cells?
Introduction
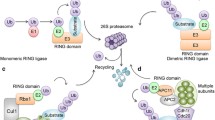
Protein ubiquitination is one of the predominantly post-translational modifications, and an overview of the ubiquitin-proteasome system (UPS) is provided in Fig. 1. E3 ubiquitin ligases are the key enzymes in the UPS to recognize specific substrates and covalently connect ubiquitin (Ub) to a protein substrate. E3 ligases-mediated ubiquitination modification exhibits different functions depending on the kind of ubiquitin conjugation. For instance, the most common K48-linked polyubiquitination promotes protein degradation by the 26S proteasome [1], while K63-linked polyubiquitination mainly regulates signal transduction and endocytosis [2]. The dysfunction of E3 ubiquitin ligases leads to the illegitimate degradation or abnormal accumulation of target protein, and the loss of tumor suppressive protein or the overactivation of oncogenic protein is always pathological and carcinogenic. The multifaceted impact of E3 ubiquitin ligases has been confirmed in many cancers, including breast cancer, ovarian cancer cells [3] and glioblastoma [4].
The ubiquitination cascade begins with the ATP-dependent activation of Ub by E1 ubiquitin-activating enzymes. After E2 ubiquitin-conjugating enzymes obtain activated-Ub, E3 ubiquitin ligases combine with Ub-E2 complex and substrate, and transfer Ub from E2 to the substrate directly or indirectly. In contrast, the deubiquitinating enzymes (DUBs) remove Ub from substrates and keep the balance of ubiquitin level of target protein together with E3 ligases to maintain cellular homeostasis. E3 ubiquitin ligases specifically recognize 7 key Lysine sites (K6, K11, K27, K33, K48 and K63) or N-terminal Methionine site (Met1) of ubiquitin and forms different chain linkages, such as mono-ubiquitination, multi-ubiquitination, linear or branched polyubiquitination chain, to exert multiple biological functions.
Colorectal cancer (CRC) is the third most common cancer diagnosed worldwide and remains the third leading cause of cancer mortality [5]. Despite existing treatment for CRC patients have achieved great advances, such as surgery, chemotherapy, radiotherapy, targeted therapy or immunotherapy, high recurrence rate and drug resistance constrain therapeutic efficacy and lead to poor prognosis, indicating a true desire for a more effective therapeutic modality. With growing interest of protein ubiquitination in CRC, mounting evidences have revealed that E3 ubiquitin ligases plays a pivotal and sophisticated role in the progression of CRC and suggested the promising therapeutic value of E3 ligases.
In this article, we [1] review the accumulative evidence about E3 ligases involved in modulating the proliferation, migration, stemness, metabolism, cell death and immune response of CRC cells; and [2] underline the regulatory effects of E3 ligases on classical signal pathway transduction in CRC; [3] highlight the potential of E3 ligases as molecular biomarkers and potent therapeutic targets in CRC.
E3 ubiquitin ligases
To date, over 700 E3 ligases can be categorized into 3 main type enzymes on the basis of the structure and the process of conjugating ubiquitin to substrate: homologous to the E6-AP C terminus (HECT) domain-containing E3s, really interesting new gene (RING) finger domain-containing E3s and RING-between-RING (RBR) family E3s [6]. RING-type E3s, the largest E3 ligases family, directly transfer the ubiquitin from E2 to the substrate, while HECT-type E3s accept the E2-bound ubiquitin and then transfer it to the substrate. The smallest E3 ligases family RBR E3s contains 12 members, which have RING domain but with a transfer mechanism of ubiquitin similar to HECT-type E3s, Cullin-RING ubiquitin ligases (CRL) are multi-subunit E3 ligase complexes and consist the scaffold protein (Cullin family), the RING box protein (RBX1/2) and different substrate recognition components. For example, CRL4DCAF1 contains Cullin4A, RBX1, damage-specific DNA binding protein 1 (DDB1) and DDB1- and CUL4-associated factor 1 (DCAF1).
The role of E3 ubiquitin ligases in CRC
The tumorigenesis of colorectal cancer is complicated and multi-faceted. Here we focus on the essential regulatory roles of E3 ligases in multiple biological processes closely related to CRC development, including proliferation, motility, stemness, metabolism, cell death and immune response.
E3 ubiquitin ligases in the proliferation of CRC
Persistent proliferation of tumor cells is a hallmark of CRC, and many E3 ligases exhibits pro- or anti-proliferation effects through ubiquitinating specific proteins involved in aberrant cell cycle progression, DNA damage repair functions and other cellular responses.
As a member of tripartite motif-containing proteins (TRIM) E3 family, the oncogenic TRIM6 promotes cell proliferation through increasing the ubiquitination and degradation of TIS21, an anti-proliferative protein that arrest CRC cells in G0/G1 phase [7]. In contrast, itchy E3 ubiquitin-protein ligase (ITCH) and HECT domain and RCC1-like domain-containing protein 3 (HERC3) suppress CRC cell proliferation and lead to G0/G1 phase arrest by targeting at CDK4 and RPL23A for K48-linked ubiquitination degradation respectively [8, 9]. The oncogenic ubiquitin-protein ligase E3 component n-recognin 5 (UBR5) promotes the degradation of ECRG4 via the ubiquitin-proteasome pathway, contributing to increased S-phase cells and augmented CRC growth [10]. E3 ligase neuronal precursor cell-expressed developmentally downregulated 4 (NEDD4) is responsible for the proteasomal degradation of p21, a key negative regulator of tumor proliferation that associates with G1 arrest [11].
What’s more, E3 ligase PDZ domain containing ring finger 3 (PDZRN3) is reported to mediate K63-linked ubiquitination and degradation of TBX20, a tumor-suppressive protein that impedes Non-homologous End Joining DNA damage repair and suppresses CRC cell proliferation [12]. E3 ligase complex CRL4ADCAF1-mediated degradation of the ribonuclease Dicer1 promotes CRC growth [13], while β-TrCP1-induced ubiquitination and degradation of HuR, an oncoprotein that binds to proliferation-related RNA, suppresses CRC cell growth [14]. Knockdown of E3 ligase Mule exhibits anti-proliferation effect in CRC cells partially attributed to Mule-induced degradation of MIZ1, a MYC-associated protein that inhibits MYC function [15]. E3 ligase ring finger protein (RNF) 111 shows tumor-suppressive properties in tumorigenesis of CRC by mediating the proteasomal degradation of SnoN/Ski, co-repressors that block the transcription of Smad2/3 [16].
E3 ubiquitin ligases related to the migration and metastasis of CRC
Cancer metastasis is a complicated and precisely modulated biological multi-steps process, in which epithelial–mesenchymal transition (EMT) is significant for CRC cells to acquire the invasion and migration capability.
E3 ligases F-box protein (FBXO) 11 and TRIM16 inhibit the EMT in CRC by mediating the ubiquitination and degradation of Snail, a transcription factor that represses E-cadherin expression [17, 18]. Constitutive photomorphogenic 1 (COP1) is the E3 ligase of ETV4, another transcription factor promoting CRC migration [19]. E3 ligase HERC3 induces the K27 and K48-linked ubiquitination degradation of EIF5A2, which negatively regulates EMT and suppresses CRC metastasis [20]. CRL4ADCAF16 complex catalyzes the mono-ubiquitination of PHGDH at K146, thereby enhances PHGDH activity to upregulate S-adenosylmethionine and promotes tumor cell migration [21]. It’s ascertained that carboxy terminus of Hsc70 interacting protein (CHIP) is the E3 ligase of autophagy-related protein 9B and myosin-9, which both contributes to CRC metastasis [22]. CHIP also mediates the ubiquitination and degradation of Mortalin-2, an oncoprotein that promotes CRC cell migration ability [23]. E3 ligase Smad ubiquitin regulatory factor (Smurf2) is responsible for the ubiquitination and degradation of RhoA, a key GTPase that contributes to metastasis in CRC [24]. TRIM65 targets at Rho GTPase activating protein 35 for proteasomal degradation, leading to increased migration-related structures in CRC cells [25].
E3 ligases and their role in CSC stemness maintenance
The multipotent cancer stem cells (CSCs) are vital to tumorigenesis, chemotherapy resistance as well as cancer recurrence. Emerging evidences have suggested that the regulation of ubiquitination levels in CRC cells may endow cells self-renewal abilities and the E3 ligases involved in the maintenance of stemness are listed below.
E3 ligase F-box and WD repeat domain-containing (FBXW) 11 targets at tumor suppressor HIC1 for ubiquitination degradation, which contributes to the maintenance of stem-cell-like properties in CRC cells [26]. Another study found that loss of FBXW7-mediated proteasomal degradation of transcription factor ZEB2 increases CSC properties [27]. E3 ligase mouse double minute 2 (MDM2)-induced ubiquitination of p53 is enhanced by WD40 repeat domain-containing protein 35 (WDR35), leading to reduced stability of p53 and oxaliplatin resistance [28]. Overexpression of E3 ligase TRIM25 also promotes oxaliplatin resistance and the stem cell properties in CRC by blocking the K63-linked ubiquitination and degradation of EZH2 mediated by E3 ligase TNF receptor-associated factor 6 (TRAF6) [29]. TRAF6 also suppresses CSC properties by conjugating K27- and K33-linked ubiquitination to the cancer stemness marker aldehyde dehydrogenase 1 family member B1 (ALDH1B1) and decreasing its enzyme activity [30]. The autoubiquitination of E3 ligase TRIM21 at K214 and K217 is promoted by the regulator CSN6, thereby decreasing TRIM21-mediated ubiquitination and degradation of Oct1, a transcription activator of another cancer stemness marker ALDH1A1 [31]. Additionally, elevated expression of E3 ligase RANBP2-type and C3HC4-type zinc finger-containing 1 (RBCK1) is found to enhance CRC cell stemness and modulate chemosensitivity [32].
E3 ligases in CRC cell metabolism
E3 ligases in glucose metabolism
Upregulated PKM2-mediated aerobic glycolysis (Warburg effect) is a hallmark of colorectal cancer cells to meet the huge energy demand for substance anabolism. E3 ligase CHIP destabilizes PKM2 via mediating its ubiquitination and degradation [33], whereas TRIM29 triggers the K48-linked ubiquitination and proteasomal degradation of PKM1 to decrease PKM1/PKM2 ratio, motivating PKM2-mediated aerobic glycolysis to become the dominate energy source of CRC cells [34]. E3 ligase FBXW7 suppresses lactate production and CRC progression by directly promoting the degradation of enolase 1 through ubiquitin-proteasome pathway [35]. FBXW7 also ubiquitinates and downregulates c-Myc via proteasomal pathway, which thereby suppresses the expression of downstream target hexokinase-2, a rate-limiting enzyme of glycolysis [36]. Moreover, Smurf2 reduce aerobic glycolysis in CRC cells through mediating ubiquitination and degradation of ChREBP, a transcription factor that can reprogram glucose metabolism [37].
E3 ligases in amino acid metabolism
Serine–Glycine–One-Carbon (SGOC) pathway is consisted of folate cycle and methionine cycle and supports essential metabolic processes related to the survival of CRC cells. The K95 acetylation of SHMT2, a one-carbon metabolic enzyme in folate cycle, inhibits its activity and promotes the K63-linked ubiquitination and lysosomal degradation mediated by E3 ligase TRIM21 [38]. Similarly, CRL3 E3 ligase complex-induced ubiquitination and proteasomal degradation of MAT IIα, an essential enzyme in methionine cycle, is promoted by K81 acetylation [39]. E3 ligase speckle-type POZ protein (SPOP) mediates polyubiquitination and degradation of ILF3, an oncoprotein that stabilizes the mRNA of genes involved in SGOC pathway [40]. It’s reported that E3 ligase COP1 mediates the K48-linked ubiquitination and degradation of FOXO4, which directly binds and suppresses the promoters of SGOC-related genes to attenuate SGOC metabolism [41]. E3 ligase TRAF2 decreases arginine synthesis by mediating the ubiquitination and degradation of the rate-limiting enzyme ASS1 [42].
E3 ligases determining CRC cell fate
E3 ligases related to apoptosis and autophagy
Apoptosis deficiency contributes to CRC malignant phenotype. In this section, we focus on the E3 ligases involved in the complicated mechanisms of apoptosis escape and autophagy enhancement strategies in CRC cells.
E3 ligase X-linked inhibitor of apoptosis protein (XIAP) participates in the resistance against TRAIL-induced apoptosis in CRC cells through inhibiting caspase-3 activity and mediating the ubiquitination and degradation of active p17 fragment of caspase-3 [43]. In contrast, E3 ligase Mule facilitates TRAIL-induced apoptosis through inducing proteasomal degradation of Mcl-1, a member of anti-apoptotic Bcl-2 family [44]. Irradiation-induced apoptosis also involves the destabilization of Mcl-1, as irradiation decreases AKT activity and then promotes FBXW7-mediated ubiquitination of Mcl-1. Simultaneously knocking down S-phase kinase-associated protein 2 (Skp2), which mediates K63-linked ubiquitination and activation of Akt, further augments the interaction between FBXW7 and Mcl-1 as well as strengthens irradiation-induced CRC cell apoptosis [45]. FBXW7 also targets at Cyclin E for proteasomal degradation, while kinase Plk2 leads to the destabilization of FBXW7 as well as accumulation of Cyclin E and decreases apoptotic activity of CRC cells [46].
RNF152 is the E3 ligase of Bcl-xL, a highly expressed anti-apoptotic protein in CRC [47]. E3 ligase complex CRL5 is responsible for K11-linked polyubiquitination and degradation of pro-apoptotic protein Noxa in oxidative stress-induced apoptosis, which can be enhanced by peroxiredoxin 1 oligomers and contributes to inhibition of ROS-induced apoptosis [48]. Seven in absentia homolog 1 (SIAH1)-mediated proteasomal degradation of HIPK2 is attenuated by genotoxic stress, which stabilizes HIPK2 and activates apoptosis in CRC cells [49]. HECT Domain E3 Ubiquitin Protein Ligase 2 (HECTD2) mediated proteasomal degradation of EHMT2 is enhanced by propionate, leading to facilitated cell apoptosis [50]. The expression of E3 ligase HERC5 is found downregulated in CRC cells, which attenuates the ubiquitination and degradation of CtBP1, leading to suppressed apoptosis [51]. EZH2 deficiency-induced upregulation of E3 ligase ITCH enhances proteasomal degradation of the pro-survival protein c-FLIP, promoting TNFα-mediated apoptosis [52].
Autophagy is also closely related to apoptosis. CRC cell autophagy flux can be promoted by E3 ligase TRIM39, which attenuates the ubiquitination of Rab7 at Lys191, instead of augmenting its ubiquitination and degradation, and contributes to increased autophagosome-lysosome fusion and CRC progression [53]. E3 ligase TRAF6 catalyzes K63-linked polyubiquitination of the recognized marker LC3B at Lys51, contributing to autophagy activation and metastasis inhibition of CRC [54]. XIAP is the E3 ligase of p62 and inhibits p62-mediated autophagy of CRC cells [55]. E3 ligase RNF216 dampens the autophagy of CRC cells by promoting the ubiquitination and degradation of Beclin1, an indispensable regulator of autophagy [56].
E3 ligases related to CRC cell differentiation
Poorly differentiated CRCs are more aggressive and correlates with poor patient survival and prognosis. E3 ligase FBXW7 promotes the ubiquitination and degradation of CDX2, a transcription factor critical to intestinal differentiation in CRC progression [57]. Smurf2 mediates the poly-ubiquitination and degradation of onco-protein ID1, which inhibits cell differentiation by hindering the transcription of differentiation-related genes [58]. In addition, Smurf2 also catalyzes multiple mono-ubiquitination of Smad3 and inhibits its transcription activity, thereby downregulating ID1 and facilitating CRC cell differentiation [59]. High rate of G1-S transition is a main characteristic of poorly differentiated cells. E3 ligase Skp2 promotes CRC differentiation through mediating the ubiquitination and destabilization of p21 and p27, two CDK inhibitors that diminish G1-S transition [60]. The transcription factor RORγt is known to augment IL-17 transcription and increase Th17 cell proportion. E3 ligase ITCH targets at RORγt for ubiquitination degradation and perturbs differentiation of naive T cells, while impeding the combination of ITCH with RORγt promotes RORγt-dependent Th17 cell differentiation [61].
E3 ligases regulating cell immune response
Overexpression of the inhibitory receptor PD-L1 on CRC cells, which contributes to silenced T cell activity and immune escape, is one of the main causes of poor response to immunotherapy of CRC patients. E3 ligase SPOP binds with the intracellular segment of PD-L1 and mediates the proteasome-dependent degradation of PD-L1, while the inactivity of SPOP contributes to immune escape in CRC patients [62]. PD-1 expressed on macrophages in CRC is ubiquitinated and degraded by casitas B lineage lymphoma (c-Cbl) to modulate the tumor microenvironment [63]. What’s more, PD-1 on tumor-infiltrating T cells is reported to be ubiquitinated and degraded by CRL3KLHL22 complex before its transportation to the cell surface, preventing excessive T cell suppression [64].
It’s known that activated tumor-infiltrating T cells mainly rely on glycolysis to ensure energy requirement, and the disorder of glycolytic metabolism will impair antitumor function as well as lead to immune evasion [65]. E3 ligase zinc finger protein (ZFP91) mediates K63-linked ubiquitination of the catalytic subunit of serine/threonine protein phosphatase 2A (PP2Ac) which restricts mTORC1-mediated T cell glycolytic metabolism, and facilitates the assembly of PP2A, ultimately disrupting the glycolysis and antitumor activity of T cells [66]. Different from activated T cells, activated T reg cells are commonly with lower glycolytic activity in tumor. It’s noteworthy that ZFP91 contributes to T reg cell homeostasis via inducing Beclin1 K63-linked ubiquitination, leading to inhibited mTORC1-dependent glycolysis as well as the development of immune tolerance [67].
The pluripotency of E3 ligases
E3 ligases that are associated with a plethora of biological processes in CRC, including cell proliferation, metastasis, stemness maintenance, cell metabolism, cell fate determination and cell immune response, are listed in Table 1. It is obviously that E3 ligases shows versatility and complexity in CRC, as aberrant expressed E3 ligases simultaneously modulate various events related to tumorigenesis through recognizing different molecules and changing the activity of target proteins via mono- or poly-ubiquitination. For instance, Smurf2 negatively influences CRC progression and is identified to be the E3 ligase of RhoA, ChREBP, SIRT1, SATB1 and Smad3 [24, 37, 68,69,70]. E3 ligase RNF40 suppresses apoptosis by mediating H2B mono-ubiquitination of anti-apoptotic proteins genes, including Mcl-1 and BIRC5, and maintaining their expression [71]. Similarly, RNF2 inhibits CRC cell apoptosis by suppressing EGR1 expression via mediating the mono-ubiquitination of H2A at K119 of EGR1 promoter [72].
What’s more, instead of synergistic effects, a specific E3 ligase involved in CRC progression could be oncogenic or tumor-suppressive or have opposing functions in a context-dependent manner. For example, FBXW11 plays a dual role in tumorigenesis of CRC, which suppresses on CRC proliferation and migration through targeting at ZNF281 [73], and meanwhile contributes to stemness by downregulating HIC1 [26]. RNF41 acts as an oncogenic E3 ligase by promoting the ubiquitination and degradation of Disheveled 2 (Dvl2) in the presence of KITENIN, rather than as a suppressor of CRC through targeting at ErbB3/ErbB4 for degradation [74]. What’s more, RNF41 suppresses CRC stemness and metastasis through mediating the ubiquitination and degradation of ASB6 [75].
E3 ligases regulating important cellular signaling pathway
EGFR signaling pathway
The aberrant overexpression of the canonical RTK receptor EGFR is very common in CRC patients and contributes to poor prognosis. Activated EGFR initiates several phosphorylation cascades, including Ras-MEK-MAPK signaling and PI3K-AKT-mTOR signaling. Tyr1045 phosphorylation of EGFR triggers its internalization, followed by c-Cbl-mediated ubiquitination and lysosomal degradation as shown in Fig. 2, which is a common mechanism for downregulating EGFR excessive activation [76]. E3 ligase β-TrCP and anaphase-promoting complex subunit 2 (ANACP2) both suppress EGFR signaling and contribute to the destabilization of Ras in the ubiquitin–proteasome pathway [77, 78].
The ubiquitination modification marked in red destabilizes the substrate, while the ubiquitination marked in blue promotes the stability. Pink icons represent upregulated or oncogenic E3 ligases observed in CRC models. Blue icons represent downregulated or tumor-suppressive E3 ligases observed in CRC models.
AKT activity is downregulated by E3 ligase SIAH1 through mediating K48-linked ubiquitination and proteasomal degradation of AKT in CRC cells [79]. In contrast, SIAH2 and RNF146 positively regulates PI3K-AKT signaling pathway by promoting the ubiquitination and degradation of PTEN, a suppressor of AKT that blocks its phosphorylation [80, 81]. AKT signaling is also negatively modulated by phosphatase PHLPP, which is ubiquitinated and degraded by TRIM29 [82]. Activated AKT directly phosphorylates and activates downstream mTORC1, whereas RNF167 mediates the K63-linked ubiquitination of Sestrin2, which inhibits mTORC1 activation by sequestering GATOR2 [83]. It’s worth noting that the mutant E3 ligase also play a role in regulating PI3K-AKT pathway, as RNF43_p.G659fs exhibits oncogenic ability through mediating the ubiquitination and degradation of PI3KRS1 and ultimately activates PI3K-AKT pathway, rather than the canonical Wnt pathway [84].
Wnt/β-catenin pathway
Abnormal activation of Wnt signaling is widely present in CRC. The classical Wnt/β-catenin signal is initiated by the binding of Wnt ligands to a receptor complex composed of the Frizzled (FZD) receptors and the low-density lipoprotein receptors (LRPs), which is negatively regulated by E3 ligases RNF43 and its homolog zinc and ring finger protein 3 (ZNRF3) through ubiquitinating FZD receptor and inducing FZD endocytosis [85]. R-spondins (RSPO) are secreted Wnt agonists and generally neutralizes RNF43/ZNRF3 by interacting with leucine-rich repeat containing G protein-coupled receptors (LGRs), enhancing Wnt signaling under low-dose Wnt, which is attenuated by tumor-suppressive E3 ligases NEDD4 and its homologue NEDD4L through targeting LGR5 for both lysosomal and proteasomal degradation [86]. Interestingly, RSPO2 functions as a tumor suppressor by increasing the accumulation of ZNRF3 and suppressing Wnt signaling [87]. Additionally, RSPO2-augmented ZNRF3-mediated ubiquitination and degradation of FZD7 counteracts Wnt5a stimulation and inhibits noncanonical Wnt signaling [88].
As the main effector of Wnt/β-catenin signal, β-catenin is ubiquitinated and degraded by β-TrCP after phosphorylation mediated by a destruction complex consisted of Axin, APC, CK1 and GSK3β in the absence of Wnt stimulation [89]. RNF146 mediates the ubiquitination and degradation of Axin1 and positively regulates β-catenin signaling [90]. The constitutive destruction of cytosolic β-catenin is usually impeded by Dvl2 after Wnt stimulation, which can be targeted by NEDD4 and NEDD4L for ubiquitination and degradation [86]. What’s more, RNF128, c-Cbl, SIAH1 and Mule are novel E3 ligases that promote the ubiquitin-mediated degradation of β-catenin respectively in CRC [91,92,93,94]. In contrast, RNF8 conjugates K63-linked ubiquitination to β-catenin and facilitates its nuclear translocation [95]. E3 ligase FBXW7 ubiquitinates and degrades KLF5, an oncoprotein that facilitates the nuclear localization of β-catenin [96]. RNF43 also mediates the ubiquitination and degradation of protease-activated receptor 2 (PAR2) that promotes β-catenin nuclear localization [97]. Surprisingly, independent of its E3 ligase activity, RNF220 promotes the deubiquitination and stabilization of β-catenin by bridging the interaction of deubiquitinase USP7 and β-catenin [98]. Similarly, RNF14 acts as a binding partner of T-cell factor/lymphoid enhancer-binding factor (TCF/LEF) and facilitates the nuclear recruitment of β-catenin without altering H2B ubiquitination [99]. What’s more, RNF6 activates the Wnt/β-catenin pathway via accelerating the ubiquitination and proteasomal degradation of TLE3, an inhibitor of β-catenin/TCF4 complex [100].
NF-κB pathway
In TNF-activated canonical NF-κB pathway, HECT domain and Ankyrin repeat containing E3 ubiquitin-protein ligase 1 (HACE1)-induced K63-linked ubiquitination of TRAF2 within the TNFR complex, along with cIAP1/2-dependent K63-linked ubiquitination of RIP1, both promotes the formation of TAK1/TAB2/TAB3 complex, which activates downstream IκB kinase (IKK) complex [101, 102]. E3 ligase TRAF6 is recruited after toll-like receptor 4 (TLR4) activation and mediates K63-linked ubiquitination of itself, further promoting NF-κB signal transduction [103]. NF-κB1 (p50) and RelA (p65) are ultimately liberated and act as the main effector of NF-κB pathway via transcription regulation. Under the signal activation, the inhibitor of p50/p65 dimer, IκBα, is targeted by E3 ligase RB binding protein 6 (RBBP6) and β-TrCP for ubiquitination and degradation, enabling the nuclear translocation of p50/p65 dimer [104, 105]. In contrast, E3 ligase CHIP and PDZ and LIM domain 2 (PDLIM2) promotes the ubiquitination and destabilization of p65 and inhibits NF-κB signaling [106, 107]. Downregulated RNF20/RNF40 decreases histone H2B monoubiquitylation in CRC, which recruits p65 and augments the transcription of p50/p65 dimer, consequently increasing NF-κB signaling [108].
Hedgehog pathway
Abnormal activation of Hedgehog signaling is commonly considered essential to CRC development. E3 ligases such as P300/CBP-associated factor (PCAF) and β-TrCP participate in the regulation of Hedgehog pathway through mediating the ubiquitination and degradation of GLI proteins, the main downstream effectors of Hedgehog pathway [109, 110]. SuFu is a classical suppressor of Hedgehog pathway that prevents the nuclear accumulation of GLI. Ligand of Num-protein X 1 (LNX1) is responsible for the ubiquitination and degradation of SuFu, which is promoted by SuFu negating protein 1 (SNEP1). SNEP1 is also a target gene of GLI2, thus forming a positive feedback loop and strengthening Hedgehog signaling [111].
Hippo pathway
The famous tumor-suppressive Hippo pathway comprises a series of phosphorylation cascade. Activated MST1/2 and SAV1 phosphorylates and activates LATS1/2 complex, which phosphorylates two oncogenic transcriptional co-activators, YAP and its orthologue TAZ, preventing their nuclear translocation and promoting their ubiquitination and degradation [112]. E3 ligase RNF128 mediates the ubiquitination and degradation of MST protein and suppresses Hippo signal [113]. Another study identified that the E3 ligase praja2 facilitates the proteasomal degradation of MOB1A, a strengthener of Hippo signal by activating LATS1/2 complex, and inhibits the Hippo signaling cascade in CRC [114]. SIAH1 directly interacts with YAP and promotes its K48-linked ubiquitination and proteasomal degradation in CRC cells [79]. HACE1 also targets YAP for degradation via proteasome pathway [115].
The therapeutic value and application prospects of E3 ligases in CRC
In the past few years, extensive efforts have been conducted to clarify the pivotal role of E3 ligases in many biological processes and signal pathways, driving a deeper understanding of the nonnegligible roles of E3 ligases in CRC development and progression. Many E3 ligases are considered potential diagnostic markers and survival predictors of CRC patients, for instance, patients with stage II/III CRC and high-expression of an E3 ligase of c-Myc, membrane-associated guanylate kinase, WW and PDZ domain containing 3 (MAGI3), had a satisfying recurrence-free survival (~80%, 5-year) and with no necessity for further adjuvant chemotherapy [116].
E3 ligases as novel effectors of treatment methods
Marvelous studies attempt to explore the full power of E3 ligases as novel targets of cancer therapeutics and emphasize its potential clinical significance. Researchers found the subunit of CRL complex, KLHL22, involves in the synergistic effect of 5-FU and anti-PD-1 therapy, in that 5-FU inhibits the transcription of KLHL22 and reduces the subsequent degradation of PD-1 [64]. Another study clarified that the combination of artesunate with WNT974 (a WNT inhibitor) induces the proteasomal degradation of overexpressed oncoprotein KRAS in CRC via significantly upregulating E3 ligase ANACP2 and β-TrCP [78]. Moreover, E3 ligases are the effectors of several anti-tumor active ingredients, including Scutellarein and Kurarinone, which respectively enhances cell division cycle 4 (CDC4)-mediated degradation of RAGE [117] and WDR76-mediated degradation of KRAS [118] in ubiquitin-proteasome pathway.
E3 ligases as potential targets of cancer therapeutics
Modulating the activity of E3 ligases improves the tumor suppressive effect of current inmmuno- or chemo-therapies in CRC cells, which has great potential to be exploited therapeutically. For instance, knockout of C-cbl prevents the exhaustion of CAR-T cells and heightens the efficacy of immunotherapy [119]. MDM4/MDM2 double knockdown in CRC cells revives p53 activity and has synergistic effect with anti-tumor drugs, including cytotoxic 5-FU [120] and MEK inhibitor trametinib [121], but another study found that impaired activity of MDM2 confers irradiation resistance to CRC cells [122], suggesting that regulating E3 enzyme activity is a double-edged sword and deserves further investigations before the translation into clinical trials. Apart from silencing the expression of E3 ligase to repress the catalytic activity, a recent study defined Hakin-1, the first small molecule inhibitor targeting at the E3 ligase Hakai, blocks Hakai-dependent ubiquitination of E-cadherin in CRC cells, presenting promising therapeutic potential against CRC progression [123]. The oncogenic CRL4DCAF4 complex, composed of Cullin4, RBX1, DDB1 and DCAF4, mediates the ubiquitination and degradation of tumor suppressor ST7 [124]. A novel compound NSC1892 is identified to strongly disrupt the Cullin4-DDB1 interaction and cause the degradation of DDB1, thereby leading to stabilization of ST7 and significant inhibition of CRC growth [125]. Additionally, inhibiting components of CARM1-p300-c-Myc-Max (CPCM) transcriptional complex that binds to the promoter of Cullin4, also decreases the ubiquitination and degradation of ST7 [126], suggesting that it’s practical to regulate E3 ligase complex activity by targeting at its subunits.
E3 ligases-based strategies: proteolysis targeting chimera (PROTAC) and molecular glue
Considering the fact that the 20 S proteasome inhibitors (bortezomib, carfifilzomib and ixazomib) that target ubiquitin-proteasome pathway are only satisfying in haematological malignancies and have disappointing results from clinical trials on CRC patients [127], scientists have been committed to exploring new therapeutic strategies based on ubiquitin pathway and have defined two novel techniques that targets specific oncogenic proteins based on several E3 ligases, the proteolysis targeting chimera (PROTAC) and molecular glue. PROTACs are synthetic bifunctional compounds containing a chemical linker and two ligands, with one targeting an E3 ligase and the other binding to the protein of interest (POI), artificially inducing the ubiquitination and degradation of the target. The end that recruits POI can be peptides, small molecules and antibodies depending on the aberrantly expressed proteins in CRC. Many PROTACs have entered clinical testing in breast and prostate cancer [128], and the major anti-tumor PROTACs developed in CRC are listed in Table 2. Cereblon (CRBN), the substrate receptor of CRL4, is one of the most widely used E3 ligases and can be recruited by Thalidomide and its derivatives. Von Hippel-Lindau (VHL) is another frequently targeted E3 ligase and acts as an adaptor protein of CRL2. Similarly, the molecular glue, a smaller molecule with higher bioavailability and without the requirement of a unique binding pockets in target proteins, also promotes selective protein degradation via acting as a scaffold of protein-E3 ligase interactions [129]. Overall, although it’s a highly challenging task to translate laboratory achievements into ultimately clinical efficacy, drug-induced proteolysis of target proteins based on E3 ligases-mediated ubiquitination and degradation is a promising therapeutic strategy in colorectal cancer and worth more attention.
Conclusions and future perspectives
The dynamic balance of protein is important to cellular homeostasis in colorectal cancer and E3 ubiquitin ligases are key factors mediating protein ubiquitination and degradation. The aberrant expression or regulation of E3 ligases leads to the excessive degradation or accumulation of target protein, which contributes to cell malignant transformation. In this review, we summarized the E3 ubiquitin ligases found in CRC and reveal their role in the development of CRC, including cell proliferation, migration, stemness maintenance, metabolism, cell fate determination, immune response as well as several typical signal pathways in CRC. We pointed out the pluripotency of E3 ubiquitin ligases that E3 ligases and substrates are not just biunique in CRC backgrounds. We also highlighted that it’s of great potential that E3 ubiquitin ligases become potent cancer therapeutic targets or biomarkers and it’s promising to include E3 ligase-based therapeutics in future clinical practice for the benefit of CRC patients.
Data availability
Data sharing is not applicable to this review as no data sets are generated and analyzed in this study.
References
Chau V, Tobias JW, Bachmair A, Marriott D, Ecker DJ, Gonda DK, et al. A multiubiquitin chain is confined to specific lysine in a targeted short-lived protein. Science. 1989;243:1576–83.
Chen ZJ. Ubiquitination in signaling to and activation of IKK. Immunol Rev. 2012;246:95–106.
Li X, Yang K-B, Chen W, Mai J, Wu X-Q, Sun T, et al. CUL3 (cullin 3)-mediated ubiquitination and degradation of BECN1 (beclin 1) inhibit autophagy and promote tumor progression. Autophagy. 2021;17:4323–40.
Hou J, Deng Q, Zhou J, Zou J, Zhang Y, Tan P, et al. CSN6 controls the proliferation and metastasis of glioblastoma by CHIP-mediated degradation of EGFR. Oncogene. 2017;36:1134–44.
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2022. CA Cancer J Clin. 2022;72:7–33.
Buetow L, Huang DT. Structural insights into the catalysis and regulation of E3 ubiquitin ligases. Nat Rev Mol Cell Biol. 2016;17:626–42.
Zheng S, Zhou C, Wang Y, Li H, Sun Y, Shen Z. TRIM6 promotes colorectal cancer cells proliferation and response to thiostrepton by TIS21/FoxM1. J Exp Clin Cancer Res. 2020;39:23.
Wang C, Li H, Wu L, Jiao X, Jin Z, Zhu Y, et al. Coiled-coil domain-containing 68 downregulation promotes colorectal cancer cell growth by inhibiting ITCH-mediated CDK4 degradation. Front Oncol. 2021;11:668743.
Zhang Z, Wu Q, Fang M, Liu Y, Jiang J, Feng Q, et al. HERC3 directly targets RPL23A for ubiquitination degradation and further regulates Colorectal Cancer proliferation and the cell cycle. Int J Biol Sci. 2022;18:3282–97.
Wang J, Zhao X, Jin L, Wu G, Yang Y. UBR5 contributes to colorectal cancer progression by destabilizing the tumor suppressor ECRG4. Dig Dis Sci. 2017;62:2781–9.
Zhang S, Yu C, Yang X, Hong H, Lu J, Hu W, et al. N-myc downstream-regulated gene 1 inhibits the proliferation of colorectal cancer through emulative antagonizing NEDD4-mediated ubiquitylation of p21. J Exp Clin Cancer Res. 2019;38:490.
Luo J, Chen JW, Zhou J, Han K, Li S, Duan JL, et al. TBX20 inhibits colorectal cancer tumorigenesis by impairing NHEJ-mediated DNA repair. Cancer Sci. 2022;113:2008–21.
Ren W, Shen S, Sun Z, Shu P, Shen X, Bu C, et al. Jak-STAT3 pathway triggers DICER1 for proteasomal degradation by ubiquitin ligase complex of CUL4A(DCAF1) to promote colon cancer development. Cancer Lett. 2016;375:209–20.
Lan Y, Xiao X, He Z, Luo Y, Wu C, Li L, et al. Long noncoding RNA OCC-1 suppresses cell growth through destabilizing HuR protein in colorectal cancer. Nucleic Acids Res. 2018;46:5809–21.
Peter S, Bultinck J, Myant K, Jaenicke LA, Walz S, Muller J, et al. Tumor cell-specific inhibition of MYC function using small molecule inhibitors of the HUWE1 ubiquitin ligase. EMBO Mol Med. 2014;6:1525–41.
Sharma V, Antonacopoulou AG, Tanaka S, Panoutsopoulos AA, Bravou V, Kalofonos HP, et al. Enhancement of TGF-beta signaling responses by the E3 ubiquitin ligase Arkadia provides tumor suppression in colorectal cancer. Cancer Res. 2011;71:6438–49.
Ruan L, Liu W, Yang Y, Chu Z, Yang C, Yang T, et al. TRIM16 overexpression inhibits the metastasis of colorectal cancer through mediating Snail degradation. Exp Cell Res. 2021;406:112735.
Wei M, Ma Y, Shen L, Xu Y, Liu L, Bu X, et al. NDRG2 regulates adherens junction integrity to restrict colitis and tumourigenesis. EBioMedicine. 2020;61:103068.
Xiao J, Yang S, Shen P, Wang Y, Sun H, Ji F, et al. Phosphorylation of ETV4 at Ser73 by ERK kinase could block ETV4 ubiquitination degradation in colorectal cancer. Biochem Biophys Res Commun. 2017;486:1062–8.
Zhang Z, He G, Lv Y, Liu Y, Niu Z, Feng Q, et al. HERC3 regulates epithelial-mesenchymal transition by directly ubiquitination degradation EIF5A2 and inhibits metastasis of colorectal cancer. Cell Death Dis. 2022;13:74.
Zhang Y, Yu H, Zhang J, Gao H, Wang S, Li S, et al. Cul4A-DDB1-mediated monoubiquitination of phosphoglycerate dehydrogenase promotes colorectal cancer metastasis via increased S-adenosylmethionine. J Clin Invest. 2021;131:e146187.
Zhong Y, Long T, Gu CS, Tang JY, Gao LF, Zhu JX, et al. MYH9-dependent polarization of ATG9B promotes colorectal cancer metastasis by accelerating focal adhesion assembly. Cell Death Differ. 2021;28:3251–69.
Sane S, Hafner A, Srinivasan R, Masood D, Slunecka JL, Noldner CJ, et al. UBXN2A enhances CHIP-mediated proteasomal degradation of oncoprotein mortalin-2 in cancer cells. Mol Oncol. 2018;12:1753–77.
Wu Y, Liu B, Lin W, Zhao R, Han W, Xie J. AAMP promotes colorectal cancermetastasis by suppressing SMURF2-mediatedubiquitination and degradation of RhoA. Mol Ther Oncolytics. 2021;23:515–30.
Chen D, Li Y, Zhang X, Wu H, Wang Q, Cai J, et al. Ubiquitin ligase TRIM65 promotes colorectal cancer metastasis by targeting ARHGAP35 for protein degradation. Oncogene. 2019;38:6429–44.
Yao J, Yang J, Yang Z, Wang XP, Yang T, Ji B, et al. FBXW11 contributes to stem-cell-like features and liver metastasis through regulating HIC1-mediated SIRT1 transcription in colorectal cancer. Cell Death Dis. 2021;12:930.
Li N, Babaei-Jadidi R, Lorenzi F, Spencer-Dene B, Clarke P, Domingo E, et al. An FBXW7-ZEB2 axis links EMT and tumour microenvironment to promote colorectal cancer stem cells and chemoresistance. Oncogenesis. 2019;8:13.
Di Y, Jing X, Hu K, Wen X, Ye L, Zhang X, et al. The c-MYC-WDR43 signalling axis promotes chemoresistance and tumour growth in colorectal cancer by inhibiting p53 activity. Drug Resist Updat : Rev Comment Antimicrob Anticancer Chemother. 2023;66:100909.
Zhou S, Peng J, Xiao L, Zhou C, Fang Y, Ou Q, et al. TRIM25 regulates oxaliplatin resistance in colorectal cancer by promoting EZH2 stability. Cell Death Dis. 2021;12:463.
Baek SH, Jang YK. AMBRA1 negatively regulates the function of ALDH1B1, a cancer stem cell marker, by controlling its ubiquitination. Int J Mol Sci. 2021;22:12079.
Qin B, Zou S, Li K, Wang H, Wei W, Zhang B, et al. CSN6-TRIM21 axis instigates cancer stemness during tumorigenesis. Br J Cancer. 2020;122:1673–85.
Liu ML, Zang F, Zhang SJ. RBCK1 contributes to chemoresistance and stemness in colorectal cancer (CRC). Biomed Pharmacother. 2019;118:109250.
Zhao G, Yuan H, Li Q, Zhang J, Guo Y, Feng T, et al. DDX39B drives colorectal cancer progression by promoting the stability and nuclear translocation of PKM2. Signal Transduct Target Ther. 2022;7:275.
Han J, Zhao Z, Zhang N, Yang Y, Ma L, Feng L, et al. Transcriptional dysregulation of TRIM29 promotes colorectal cancer carcinogenesis via pyruvate kinase-mediated glucose metabolism. Aging (Albany NY). 2021;13:5034–54.
Zhan P, Wang Y, Zhao S, Liu C, Wang Y, Wen M, et al. FBXW7 negatively regulates ENO1 expression and function in colorectal cancer. Lab Invest. 2015;95:995–1004.
Wu Z, Han X, Tan G, Zhu Q, Chen H, Xia Y, et al. Dioscin inhibited glycolysis and induced cell apoptosis in colorectal cancer via promoting c-myc ubiquitination and subsequent Hexokinase-2 suppression. Oncol Targets Ther. 2020;13:31–44.
Li Y, Yang D, Tian N, Zhang P, Zhu Y, Meng J, et al. The ubiquitination ligase SMURF2 reduces aerobic glycolysis and colorectal cancer cell proliferation by promoting ChREBP ubiquitination and degradation. J Biol Chem. 2019;294:14745–56.
Wei Z, Song J, Wang G, Cui X, Zheng J, Tang Y, et al. Deacetylation of serine hydroxymethyl-transferase 2 by SIRT3 promotes colorectal carcinogenesis. Nat Commun. 2018;9:4468.
Wang J, Zhu ZH, Yang HB, Zhang Y, Zhao XN, Zhang M, et al. Cullin 3 targets methionine adenosyltransferase IIalpha for ubiquitylation-mediated degradation and regulates colorectal cancer cell proliferation. FEBS J. 2016;283:2390–402.
Li K, Wu JL, Qin B, Fan Z, Tang Q, Lu W, et al. ILF3 is a substrate of SPOP for regulating serine biosynthesis in colorectal cancer. Cell Res. 2020;30:163–78.
Choi HH, Zou S, Wu JL, Wang H, Phan L, Li K, et al. EGF relays signals to COP1 and facilitates FOXO4 degradation to promote tumorigenesis. Adv Sci. 2020;7:2000681.
Jia H, Yang Y, Li M, Chu Y, Song H, Zhang J, et al. Snail enhances arginine synthesis by inhibiting ubiquitination-mediated degradation of ASS1. EMBO Rep. 2021;22:e51780.
Ehrenschwender M, Bittner S, Seibold K, Wajant H. XIAP-targeting drugs re-sensitize PIK3CA-mutated colorectal cancer cells for death receptor-induced apoptosis. Cell Death Dis. 2014;5:e1570.
Park SH, Lee D-H, Kim JL, Kim BR, Na YJ, Jo MJ, et al. Metformin enhances TRAIL-induced apoptosis by Mcl-1 degradation via Mule in colorectal cancer cells. Oncotarget. 2016;7:59503–18.
Yu X, Zhou L, Liu W, Liu L, Gao F, Li W, et al. Skp2 stabilizes Mcl-1 and confers radioresistance in colorectal cancer. Cell Death Dis. 2022;13:249.
Ou B, Zhao J, Guan S, Wangpu X, Zhu C, Zong Y, et al. Plk2 promotes tumor growth and inhibits apoptosis by targeting Fbxw7/Cyclin E in colorectal cancer. Cancer Lett. 2016;380:457–66.
Li W, Ma Y, He L, Li H, Chu Y, Jiang Z, et al. Protease-activated receptor 2 stabilizes Bcl-xL and regulates EGFR-targeted therapy response in colorectal cancer. Cancer Lett. 2021;517:14–23.
Xu S, Ma Y, Tong Q, Yang J, Liu J, Wang Y, et al. Cullin-5 neddylation-mediated NOXA degradation is enhanced by PRDX1 oligomers in colorectal cancer. Cell Death Dis. 2021;12:265.
Kurokawa K, Akaike Y, Masuda K, Kuwano Y, Nishida K, Yamagishi N, et al. Downregulation of serine/arginine-rich splicing factor 3 induces G1 cell cycle arrest and apoptosis in colon cancer cells. Oncogene. 2014;33:1407–17.
Ryu TY, Kim K, Han TS, Lee MO, Lee J, Choi J, et al. Human gut-microbiome-derived propionate coordinates proteasomal degradation via HECTD2 upregulation to target EHMT2 in colorectal cancer. ISME J. 2022;16:1205–21.
Zhu L, Wu J, Liu H. Downregulation of HERC5 E3 ligase attenuates the ubiquitination of CtBP1 to inhibit apoptosis in colorectal cancer cells. Carcinogenesis. 2021;42:1119–30.
Liu Y, Peng J, Sun T, Li N, Zhang L, Ren J, et al. Epithelial EZH2 serves as an epigenetic determinant in experimental colitis by inhibiting TNFalpha-mediated inflammation and apoptosis. Proc Natl Acad Sci USA. 2017;114:E3796–E805.
Hu J, Ding X, Tian S, Chu Y, Liu Z, Li Y, et al. TRIM39 deficiency inhibits tumor progression and autophagic flux in colorectal cancer via suppressing the activity of Rab7. Cell Death Dis. 2021;12:391.
Wu H, Lu XX, Wang JR, Yang TY, Li XM, He XS, et al. TRAF6 inhibits colorectal cancer metastasis through regulating selective autophagic CTNNB1/beta-catenin degradation and is targeted for GSK3B/GSK3beta-mediated phosphorylation and degradation. Autophagy. 2019;15:1506–22.
Huang X, Wang XN, Yuan XD, Wu WY, Lobie PE, Wu Z. XIAP facilitates breast and colon carcinoma growth via promotion of p62 depletion through ubiquitination-dependent proteasomal degradation. Oncogene. 2019;38:1448–60.
Wang H, Wang Y, Qian L, Wang X, Gu H, Dong X, et al. RNF216 contributes to proliferation and migration of colorectal cancer via suppressing BECN1-dependent autophagy. Oncotarget. 2016;7:51174–83.
Kumar Y, Shukla N, Thacker G, Kapoor I, Lochab S, Bhatt ML, et al. Ubiquitin Ligase, Fbw7, Targets CDX2 for degradation via two phosphodegron motifs in a GSK3beta-dependent manner. Mol Cancer Res. 2016;14:1097–109.
Kong Y, Cui H, Zhang H. Smurf2-mediated ubiquitination and degradation of Id1 regulates p16 expression during senescence. Aging Cell. 2011;10:1038–46.
Tang LY, Yamashita M, Coussens NP, Tang Y, Wang X, Li C, et al. Ablation of Smurf2 reveals an inhibition in TGF-beta signalling through multiple mono-ubiquitination of Smad3. EMBO J. 2011;30:4777–89.
Shen L, Qu X, Li H, Xu C, Wei M, Wang Q, et al. NDRG2 facilitates colorectal cancer differentiation through the regulation of Skp2-p21/p27 axis. Oncogene. 2018;37:1759–74.
Sun J, Jia H, Bao X, Wu Y, Zhu T, Li R, et al. Tumor exosome promotes Th17 cell differentiation by transmitting the lncRNA CRNDE-h in colorectal cancer. Cell Death Dis. 2021;12:123.
Zhang H, Xia Y, Wang F, Luo M, Yang K, Liang S, et al. Aldehyde Dehydrogenase 2 mediates alcohol-induced colorectal cancer immune escape through stabilizing PD-L1 Expression. Adv Sci. 2021;8:2003404.
Lyle C, Richards S, Yasuda K, Napoleon MA, Walker J, Arinze N, et al. c-Cbl targets PD-1 in immune cells for proteasomal degradation and modulates colorectal tumor growth. Sci Rep. 2019;9:20257.
Zhou XA, Zhou J, Zhao L, Yu G, Zhan J, Shi C, et al. KLHL22 maintains PD-1 homeostasis and prevents excessive T-cell suppression. Proc Natl Acad Sci USA. 2020;117:28239–50.
Rivadeneira DB, Delgoffe GM. Antitumor T-cell reconditioning: improving metabolic fitness for optimal cancer immunotherapy. Clin Cancer Res. 2018;24:2473–81.
Wang F, Zhang Y, Yu X, Teng XL, Ding R, Hu Z, et al. ZFP91 disturbs metabolic fitness and antitumor activity of tumor-infiltrating T cells. J Clin Invest. 2021;131:e144318.
Wang A, Ding L, Wu Z, Ding R, Teng XL, Wang F, et al. ZFP91 is required for the maintenance of regulatory T cell homeostasis and function. J Exp Med. 2021;218:e20201217.
Yu L, Dong L, Li H, Liu Z, Luo Z, Duan G, et al. Ubiquitination-mediated degradation of SIRT1 by SMURF2 suppresses CRC cell proliferation and tumorigenesis. Oncogene. 2020;39:4450–64.
Yu L, Dong L, Wang Y, Liu L, Long H, Li H, et al. Reversible regulation of SATB1 ubiquitination by USP47 and SMURF2 mediates colon cancer cell proliferation and tumor progression. Cancer Lett. 2019;448:40–51.
Niu B, Liu J, Lv B, Lin J, Li X, Wu C, et al. Interplay between transforming growth factor-beta and Nur77 in dual regulations of inhibitor of differentiation 1 for colonic tumorigenesis. Nat Commun. 2021;12:2809.
Schneider D, Chua RL, Molitor N, Hamdan FH, Rettenmeier EM, Prokakis E, et al. The E3 ubiquitin ligase RNF40 suppresses apoptosis in colorectal cancer cells. Clin Epigenet. 2019;11:98.
Wei F, Jing H, Wei M, Liu L, Wu J, Wang M, et al. Ring finger protein 2 promotes colorectal cancer progression by suppressing early growth response 1. Aging. 2020;12:26199–220.
Zhu Y, Zhou Q, Zhu G, Xing Y, Li S, Ren N, et al. GSK-3β phosphorylation-dependent degradation of ZNF281 by β-TrCP2 suppresses colorectal cancer progression. Oncotarget. 2017;8:88599–612.
Sun EG, Lee KH, Ko YS, Choi HJ, Yang JI, Lee JH, et al. KITENIN functions as a fine regulator of ErbB4 expression level in colorectal cancer via protection of ErbB4 from E3-ligase Nrdp1-mediated degradation. Mol Carcinog. 2017;56:1068–81.
Zeng W, Zhu JF, Guo J, Huang GJ, Ai LS, Zeng Y, et al. m(6)A-modified circFNDC3B inhibits colorectal cancer stemness and metastasis via RNF41-dependent ASB6 degradation. Cell Death Dis. 2022;13:1008.
Yao S, Zheng P, Wu H, Song LM, Ying XF, Xing C, et al. Erbin interacts with c-Cbl and promotes tumourigenesis and tumour growth in colorectal cancer by preventing c-Cbl-mediated ubiquitination and down-regulation of EGFR. J Pathol 2015;236:65–77.
Jeong W-J, Yoon J, Park J-C, Lee S-H, Lee S-H, Kaduwal S, et al. Ras stabilization through aberrant activation of Wnt/β-catenin signaling promotes intestinal tumorigenesis. Sci Signal. 2012;5:ra30.
Gong RH, Chen M, Huang C, Wong HLX, Kwan HY, Bian Z. Combination of artesunate and WNT974 induces KRAS protein degradation by upregulating E3 ligase ANACP2 and beta-TrCP in the ubiquitin-proteasome pathway. Cell Commun Signal. 2022;20:34.
Xiao Z, Wei Z, Deng D, Zheng Z, Zhao Y, Jiang S, et al. Downregulation of Siah1 promotes colorectal cancer cell proliferation and migration by regulating AKT and YAP ubiquitylation and proteasome degradation. Cancer Cell Int. 2020;20:50.
Hu Y, He Y, Liu W, Yu S, Wei Y, Bai S, et al. SIAH2 regulates colorectal cancer tumorigenesis via PI3K/ATK signaling pathway. Tissue Cell. 2022;78:101878.
Li N, Zhang Y, Han X, Liang K, Wang J, Feng L, et al. Poly-ADP ribosylation of PTEN by tankyrases promotes PTEN degradation and tumor growth. Genes Dev. 2015;29:157–70.
Jiang T, Wang H, Liu L, Song H, Zhang Y, Wang J, et al. CircIL4R activates the PI3K/AKT signaling pathway via the miR-761/TRIM29/PHLPP1 axis and promotes proliferation and metastasis in colorectal cancer. Mol Cancer. 2021;20:167.
Wang D, Xu C, Yang W, Chen J, Ou Y, Guan Y, et al. E3 ligase RNF167 and deubiquitinase STAMBPL1 modulate mTOR and cancer progression. Mol Cell. 2022;82:770–84.e9.
Fang L, Ford-Roshon D, Russo M, O’Brien C, Xiong X, Gurjao C, et al. RNF43 G659fs is an oncogenic colorectal cancer mutation and sensitizes tumor cells to PI3K/mTOR inhibition. Nat Commun. 2022;13:3181.
Giebel N, de Jaime-Soguero A, Garcia Del Arco A, Landry JJM, Tietje M, Villacorta L, et al. USP42 protects ZNRF3/RNF43 from R-spondin-dependent clearance and inhibits Wnt signalling. EMBO Rep. 2021;22:e51415.
Novellasdemunt L, Kucharska A, Jamieson C, Prange-Barczynska M, Baulies A, Antas P, et al. NEDD4 and NEDD4L regulate Wnt signalling and intestinal stem cell priming by degrading LGR5 receptor. EMBO J. 2020;39:e102771.
Wu C, Qiu S, Lu L, Zou J, Li WF, Wang O, et al. RSPO2-LGR5 signaling has tumour-suppressive activity in colorectal cancer. Nat Commun. 2014;5:3149.
Dong X, Liao W, Zhang L, Tu X, Hu J, Chen T, et al. RSPO2 suppresses colorectal cancer metastasis by counteracting the Wnt5a/Fzd7-driven noncanonical Wnt pathway. Cancer Lett. 2017;402:153–65.
Ranes M, Zaleska M, Sakalas S, Knight R, Guettler S. Reconstitution of the destruction complex defines roles of AXIN polymers and APC in beta-catenin capture, phosphorylation, and ubiquitylation. Mol Cell. 2021;81:3246–61.e11.
Shen J, Yu Z, Li N. The E3 ubiquitin ligase RNF146 promotes colorectal cancer by activating the Wnt/beta-catenin pathway via ubiquitination of Axin1. Biochem Biophys Res Commun. 2018;503:991–7.
Zhu Y, Gan Y, Zou R, Sha H, Lu Y, Zhang Y, et al. RNF128 suppresses the malignancy of colorectal cancer cells via inhibition of Wnt/β-catenin signaling. Am J Transl Res. 2021;13:13567–78.
Richards S, Walker J, Nakanishi M, Belghasem M, Lyle C, Arinze N, et al. Haploinsufficiency of Casitas B-Lineage Lymphoma augments the progression of colon cancer in the background of adenomatous polyposis coli inactivation. Am J Pathol. 2020;190:602–13.
Jumpertz S, Hennes T, Asare Y, Vervoorts J, Bernhagen J, Schutz AK. The beta-catenin E3 ubiquitin ligase SIAH-1 is regulated by CSN5/JAB1 in CRC cells. Cell Signal. 2014;26:2051–9.
Dominguez-Brauer C, Khatun R, Elia AJ, Thu KL, Ramachandran P, Baniasadi SP, et al. E3 ubiquitin ligase Mule targets beta-catenin under conditions of hyperactive Wnt signaling. Proc Natl Acad Sci USA. 2017;114:E1148–E57.
Ren L, Zhou T, Wang Y, Wu Y, Xu H, Liu J, et al. RNF8 induces beta-catenin-mediated c-Myc expression and promotes colon cancer proliferation. Int J Biol Sci. 2020;16:2051–62.
Bialkowska AB, Liu Y, Nandan MO, Yang VW. A colon cancer-derived mutant of Kruppel-like factor 5 (KLF5) is resistant to degradation by glycogen synthase kinase 3beta (GSK3beta) and the E3 ubiquitin ligase F-box and WD repeat domain-containing 7alpha (FBW7alpha). J Biol Chem. 2014;289:5997–6005.
Nag JK, Appasamy P, Sedley S, Malka H, Rudina T, Bar-Shavit R. RNF43 induces the turnover of protease-activated receptor 2 in colon cancer. FASEB J. 2023;37:e22675.
Ma P, Yang X, Kong Q, Li C, Yang S, Li Y, et al. The ubiquitin ligase RNF220 enhances canonical Wnt signaling through USP7-mediated deubiquitination of beta-catenin. Mol Cell Biol. 2014;34:4355–66.
Wu B, Piloto S, Zeng W, Hoverter NP, Schilling TF, Waterman ML. Ring Finger Protein 14 is a new regulator of TCF/beta-catenin-mediated transcription and colon cancer cell survival. EMBO Rep. 2013;14:347–55.
Liu L, Zhang Y, Wong CC, Zhang J, Dong Y, Li X, et al. RNF6 promotes colorectal cancer by activating the Wnt/beta-Catenin pathway via ubiquitination of TLE3. Cancer Res. 2018;78:1958–71.
Tortola L, Nitsch R, Bertrand MJM, Kogler M, Redouane Y, Kozieradzki I, et al. The tumor suppressor Hace1 is a critical regulator of TNFR1-mediated cell fate. Cell Rep. 2016;15:1481–92.
Wu P, Shi KJ, An JJ, Ci YL, Li F, Hui KY, et al. The LEF1/CYLD axis and cIAPs regulate RIP1 deubiquitination and trigger apoptosis in selenite-treated colorectal cancer cells. Cell Death Dis. 2014;5:e1085.
Guangwei Z, Zhibin C, Qin W, Chunlin L, Penghang L, Ruofan H, et al. TRAF6 regulates the signaling pathway influencing colorectal cancer function through ubiquitination mechanisms. Cancer Sci. 2022;113:1393–405.
Xiao C, Wu G, Zhou Z, Zhang X, Wang Y, Song G, et al. RBBP6, a RING finger-domain E3 ubiquitin ligase, induces epithelial-mesenchymal transition and promotes metastasis of colorectal cancer. Cell Death Dis. 2019;10:833.
Ougolkov A, Zhang B, Yamashita K, Bilim V, Mai M, Fuchs SY, et al. Associations among beta-TrCP, an E3 ubiquitin ligase receptor, beta-catenin, and NF-kappaB in colorectal cancer. J Natl Cancer Inst 2004;96:1161–70.
Wang Y, Ren F, Wang Y, Feng Y, Wang D, Jia B, et al. CHIP/Stub1 functions as a tumor suppressor and represses NF-kappaB-mediated signaling in colorectal cancer. Carcinogenesis. 2014;35:983–91.
Liu S, Sun X, Wang M, Hou Y, Zhan Y, Jiang Y, et al. A microRNA 221- and 222-mediated feedback loop maintains constitutive activation of NFkappaB and STAT3 in colorectal cancer cells. Gastroenterology. 2014;147:847–859.e11.
Tarcic O, Pateras IS, Cooks T, Shema E, Kanterman J, Ashkenazi H, et al. RNF20 links Histone H2B ubiquitylation with inflammation and inflammation-associated cancer. Cell Rep. 2016;14:1462–76.
Kim BR, Jeong YA, Na YJ, Park SH, Jo MJ, Kim JL, et al. Genipin suppresses colorectal cancer cells by inhibiting the Sonic Hedgehog pathway. Oncotarget. 2017;8:101952–64.
Kim BR, Na YJ, Kim JL, Jeong YA, Park SH, Jo MJ, et al. RUNX3 suppresses metastasis and stemness by inhibiting Hedgehog signaling in colorectal cancer. Cell Death Differ. 2020;27:676–94.
Yan Z, Cheng M, Hu G, Wang Y, Zeng S, Huang A, et al. Positive feedback of SuFu negating protein 1 on Hedgehog signaling promotes colorectal tumor growth. Cell Death Dis. 2021;12:199.
Zhou D, Zhang Y, Wu H, Barry E, Yin Y, Lawrence E, et al. Mst1 and Mst2 protein kinases restrain intestinal stem cell proliferation and colonic tumorigenesis by inhibition of Yes-associated protein (Yap) overabundance. Proc Natl Acad Sci USA. 2011;108:E1312–20.
Ning S, Chen Y, Wang G, Liu Y, Yang Y, Zhang Z. Ring finger protein 128 promotes, rather than inhibits, colorectal cancer progression by regulating the Hippo signaling pathway. Front Oncol. 2022;12:1031160.
Yang P, Zhang D, Wang T, Ji J, Jin C, Peng C, et al. CAF-derived exosomal WEE2-AS1 facilitates colorectal cancer progression via promoting degradation of MOB1A to inhibit the Hippo pathway. Cell Death Dis. 2022;13:796.
Zhou Z, Zhang H-S, Zhang Z-G, Sun H-L, Liu H-Y, Gou X-M, et al. Loss of HACE1 promotes colorectal cancer cell migration via upregulation of YAP1. J Cell Physiol. 2019;234:9663–72.
Wang H, Yang W, Qin Q, Yang X, Yang Y, Liu H, et al. E3 ubiquitin ligase MAGI3 degrades c-Myc and acts as a predictor for chemotherapy response in colorectal cancer. Mol Cancer. 2022;21:151.
Li Y, Wang J, Zhong S, Li J, Du W. Scutellarein inhibits the development of colon cancer via CDC4‑mediated RAGE ubiquitination. Int J Mol Med. 2020;45:1059–72.
Kwon M, Oh T, Jang M, Kim GH, Kim JH, Ryu HW, et al. Kurarinone induced p53-independent G0/G1 cell cycle arrest by degradation of K-RAS via WDR76 in human colorectal cancer cells. Eur J Pharm. 2022;923:174938.
Kumar J, Kumar R, Kumar Singh A, Tsakem EL, Kathania M, Riese MJ, et al. Deletion of Cbl-b inhibits CD8(+) T-cell exhaustion and promotes CAR T-cell function. J Immunother Cancer. 2021;9:e001688.
Imanishi M, Yamamoto Y, Wang X, Sugaya A, Hirose M, Endo S, et al. Augmented antitumor activity of 5-fluorouracil by double knockdown of MDM4 and MDM2 in colon and gastric cancer cells. Cancer Sci. 2019;110:639–49.
Wang X, Yamamoto Y, Imanishi M, Zhang X, Sato M, Sugaya A, et al. Enhanced G1 arrest and apoptosis via MDM4/MDM2 double knockdown and MEK inhibition in wild-type TP53 colon and gastric cancer cells with aberrant KRAS signaling. Oncol Lett. 2021;22:558.
Niu X, Cui H, Gu X, Wu T, Sun M, Zhou C, et al. Nuclear receptor PXR confers irradiation resistance by promoting DNA damage response through stabilization of ATF3. Front Oncol. 2022;12:837980.
Martinez-Iglesias O, Casas-Pais A, Castosa R, Diaz-Diaz A, Roca-Lema D, Concha A, et al. Hakin-1, a new specific small-molecule inhibitor for the E3 ubiquitin-Ligase Hakai, inhibits carcinoma growth and progression. Cancers. 2020;12:1340.
Liu H, Lu W, He H, Wu J, Zhang C, Gong H, et al. Inflammation-dependent overexpression of c-Myc enhances CRL4(DCAF4) E3 ligase activity and promotes ubiquitination of ST7 in colitis-associated cancer. J Pathol. 2019;248:464–75.
Yang C, Wu J, He H, Liu H. Small molecule NSC1892 targets the CUL4A/4B-DDB1 interactions and causes impairment of CRL4(DCAF4) E3 ligases to inhibit colorectal cancer cell growth. Int J Biol Sci. 2020;16:1059–70.
Lu W, Yang C, He H, Liu H. The CARM1-p300-c-Myc-Max (CPCM) transcriptional complex regulates the expression of CUL4A/4B and affects the stability of CRL4 E3 ligases in colorectal cancer. Int J Biol Sci. 2020;16:1071–85.
Mackay H, Hedley D, Major P, Townsley C, Mackenzie M, Vincent M, et al. A phase II trial with pharmacodynamic endpoints of the proteasome inhibitor bortezomib in patients with metastatic colorectal cancer. Clin Cancer Res : Off J Am Assoc Cancer Res. 2005;11:5526–33.
Lier S, Sellmer A, Orben F, Heinzlmeir S, Krauss L, Schneeweis C, et al. A novel Cereblon E3 ligase modulator with antitumor activity in gastrointestinal cancer. Bioorg Chem. 2022;119:105505.
Dieter SM, Siegl C, Codo PL, Huerta M, Ostermann-Parucha AL, Schulz E, et al. Degradation of CCNK/CDK12 is a druggable vulnerability of colorectal cancer. Cell Rep. 2021;36:109394.
Zhang B, Liu Q, Wen W, Gao H, Wei W, Tang A, et al. The chromatin remodeler CHD6 promotes colorectal cancer development by regulating TMEM65-mediated mitochondrial dynamics via EGF and Wnt signaling. Cell Discov. 2022;8:130.
Liang Q, Ma D, Zhu X, Wang Z, Sun TT, Shen C, et al. RING-Finger Protein 6 amplification activates JAK/STAT3 pathway by modifying SHP-1 ubiquitylation and associates with poor outcome in colorectal cancer. Clin Cancer Res. 2018;24:1473–85.
Wen P, Wang H, Li Y, Sui X, Hou Z, Guo X, et al. MICALL2 as a substrate of ubiquitinase TRIM21 regulates tumorigenesis of colorectal cancer. Cell Commun Signal. 2022;20:170.
Liang Q, Tang C, Tang M, Zhang Q, Gao Y, Ge Z. TRIM47 is up-regulated in colorectal cancer, promoting ubiquitination and degradation of SMAD4. J Exp Clin Cancer Res. 2019;38:159.
Naudin C, Sirvent A, Leroy C, Larive R, Simon V, Pannequin J, et al. SLAP displays tumour suppressor functions in colorectal cancer via destabilization of the SRC substrate EPHA2. Nat Commun. 2014;5:3159.
Seo HG, Kim HB, Yoon JY, Kweon TH, Park YS, Kang J, et al. Mutual regulation between OGT and XIAP to control colon cancer cell growth and invasion. Cell Death Dis. 2020;11:815.
Cheng J, Li Y, Wang X, Dong G, Sheng C. Discovery of novel PDEdelta degraders for the treatment of KRAS mutant colorectal cancer. J Med Chem. 2020;63:7892–905.
Liao H, Li X, Zhao L, Wang Y, Wang X, Wu Y, et al. A PROTAC peptide induces durable beta-catenin degradation and suppresses Wnt-dependent intestinal cancer. Cell Discov. 2020;6:35.
Jin J, Wu Y, Zhao Z, Wu Y, Zhou YD, Liu S, et al. Small-molecule PROTAC mediates targeted protein degradation to treat STAT3-dependent epithelial cancer. JCI Insight. 2022;7:e160606.
Zhou Q, Wu W, Jia K, Qi G, Sun XS, Li P. Design and characterization of PROTAC degraders specific to protein N-terminal methyltransferase 1. Eur J Med Chem. 2022;244:114830.
He Y, Ju Y, Hu Y, Wang B, Che S, Jian Y, et al. Brd4 proteolysis-targeting chimera nanoparticles sensitized colorectal cancer chemotherapy. J Control Rel. 2023;354:155–66.
Bian Y, Alem D, Beato F, Hogenson TL, Yang X, Jiang K, et al. Development of SOS1 inhibitor-based degraders to target KRAS-mutant colorectal cancer. J Med Chem. 2022;65:16432–50.
Qin AC, Jin H, Song Y, Gao Y, Chen YF, Zhou LN, et al. The therapeutic effect of the BRD4-degrading PROTAC A1874 in human colon cancer cells. Cell Death Dis. 2020;11:805.
Tong J, Tan X, Risnik D, Gao M, Song X, Ermine K, et al. BET protein degradation triggers DR5-mediated immunogenic cell death to suppress colorectal cancer and potentiate immune checkpoint blockade. Oncogene. 2021;40:6566–78.
Lu J, Huang Y, Huang J, He R, Huang M, Lu X, et al. Discovery of the first examples of Threonine Tyrosine Kinase PROTAC degraders. J Med Chem. 2022;65:2313–28.
Chen S, Li X, Li Y, Yuan X, Geng C, Gao S, et al. Design of stapled peptide-based PROTACs for MDM2/MDMX atypical degradation and tumor suppression. Theranostics. 2022;12:6665–81.
Madak JT, Cuthbertson CR, Chen W, Showalter HD, Neamati N. Design, synthesis, and characterization of Brequinar conjugates as probes to study DHODH Inhibition. Chemistry. 2017;23:13875–8.
Marei H, Tsai WK, Kee YS, Ruiz K, He J, Cox C, et al. Antibody targeting of E3 ubiquitin ligases for receptor degradation. Nature. 2022;610:182–9.
Uehara T, Minoshima Y, Sagane K, Sugi NH, Mitsuhashi KO, Yamamoto N, et al. Selective degradation of splicing factor CAPERalpha by anticancer sulfonamides. Nat Chem Biol. 2017;13:675–80.
Dunbar K, Macartney TJ, Sapkota GP. IMiDs induce FAM83F degradation via an interaction with CK1alpha to attenuate Wnt signalling. Life Sci Alliance. 2021;4:e202000804.
Ling X, Wu W, Aljahdali IAM, Liao J, Santha S, Fountzilas C, et al. FL118, acting as a ‘molecular glue degrader’, binds to dephosphorylates and degrades the oncoprotein DDX5 (p68) to control c-Myc, survivin and mutant Kras against colorectal and pancreatic cancer with high efficacy. Clin Transl Med. 2022;12:e881.
Acknowledgements
We thank all members of Chu lab for stimulating discussions and helpful comments. We also apologize to those whose work is not referenced due to space limitations.
Funding
This work was supported by the National Natural Science Foundation of China 82072725 and 81872042 (to Xiaoyuan Chu), 81702442 (to Zengjie Lei), 81972332 (to Yitian Chen), 82002583 (to Mengxi Huang), 82002591 (to Gongbo Fu), the Key Program of Jiangsu Provincial Health Commission ZD2021039 (to Xiaoyuan Chu), the Natural Science Foundation of Jiangsu province BK20170623 (to Zengjie Lei), BK20200273 (to Gongbo Fu), the China Postdoctoral Science Foundation 2020M670090ZX (to Zengjie Lei), 2021MD703958 (to Gongbo Fu), the Postdoctoral Science Found of Jiangsu province 2018K090B (to Zengjie Lei).
Author information
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, J., Feng, H., Wang, Y. et al. The involvement of E3 ubiquitin ligases in the development and progression of colorectal cancer. Cell Death Discov. 9, 458 (2023). https://doi.org/10.1038/s41420-023-01760-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-023-01760-z
- Springer Nature Limited