Abstract
Type A γ-aminobutyric acid receptors (GABAARs) are inhibitory pentameric ligand-gated ion channels in the brain. Many anesthetics and neurosteroids act through binding to the GABAAR transmembrane domain (TMD), but the structural basis of their actions is not well understood and no resting-state GABAAR structure has been determined. Here, we report crystal structures of apo and the neurosteroid anesthetic alphaxalone-bound desensitized chimeric α1GABAAR (ELIC-α1GABAAR). The chimera retains the functional and pharmacological properties of GABAARs, including potentiation, activation and desensitization by alphaxalone. The apo-state structure reveals an unconventional activation gate at the intracellular end of the pore. The desensitized structure illustrates molecular determinants for alphaxalone binding to an inter-subunit TMD site. These structures suggest a plausible signaling pathway from alphaxalone binding at the bottom of the TMD to the channel gate in the pore-lining TM2 through the TM1–TM2 linker. The study provides a framework to discover new GABAAR modulators with therapeutic potential.
Similar content being viewed by others
Introduction
Type A γ-aminobutyric acid receptors (GABAARs) control neuronal excitability and are the primary inhibitory pentameric ligand-gated ion channels (pLGICs) in the central nervous system1. Naturally occurring GABAARs are mostly heteropentamers assembled by homologous subunits. (αβ)2δGABAARs are exclusively extrasynaptic and (αβ)2γGABAARs are both synaptic and extrasynaptic2 receptors that mediate prolonged tonic and short phasic inhibition, respectively1,3. Among them, the α1-containing receptors are the most abundant GABAARs in the brain2.
GABAAR-mediated inhibition of neuronal excitability results from membrane hyperpolarization due to Cl− flux upon GABAAR activation, which can be triggered by binding of the neurotransmitter GABA to the orthosteric site in the extracellular domain (ECD) or allosteric binding of endogenous neuroactive steroids1,3. In addition, inhibitory functions of GABAARs can also be regulated by a wide variety of synthetic drugs in different physiological and pathological contexts. GABAARs are targets for the treatment of neurological diseases and disorders, such as epilepsy, depression and insomnia3. GABAARs are also targets for general anesthetics. The underlying mechanisms of action of these drugs on GABAARs remain to be determined4.
Ample experimental evidence suggests that the transmembrane domain (TMD) of GABAARs harbors sites for the primary actions of general anesthetics and neurosteroids5,6,7,8,9,10,11,12. The TMD has an essential role in functional transitions among the resting, activated, and desensitized states of these Cl−-conducting channels. Three-dimensional GABAAR structures, particularly those revealing insights into how general anesthetics and neurosteroids exert their actions, are limited due to many technical challenges. For a long time, the crystal structure of a desensitized homomeric β3GABAAR13 was the only determined structure in the family of GABAARs. The successes in crystallographic structural determination of GLIC-α1GABAAR11 and β3-α5GABAAR12 chimeras open a new path to accelerate the process and demonstrate the feasibility of chimeric GABAARs in illustrating the structural basis underlying the actions of neurosteroids or general anesthetics. All of these structures represent the desensitized state. More recently, cryo-electron microscopy structures of the human α1β2γ2GABAAR in complex with GABA and flumazenil have been published14. However, no apo-state structure has yet been determined for GABAARs.
Here, we report crystal structures of an α1GABAAR chimera in the apo state and in an alphaxalone-bound desensitized state. Alphaxalone (5α-pregnan-3α-ol-11,20 dione) is a potent neurosteroid anesthetic. The anxiolytic, anticonvulsant, analgesic, and sedative-hypnotic effects of alphaxalone have been linked to its potentiation of GABA-evoked currents and direct activation of GABAARs15,16. However, the data about the alphaxalone binding site in GABAARs and the underlying structural basis of alphaxalone action are sparse17,18. Our crystal structures illustrate the molecular details of alphaxalone binding to the α1GABAAR TMD, reveal neurosteroid anesthetic action starting at the bottom of the TMD, and provide a structural basis for the rational discovery of new drugs.
Results
The α1GABAAR chimera resembles functions of GABAARs
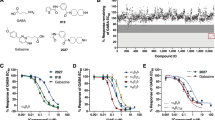
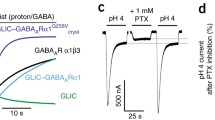
We constructed a GABAAR chimera by fusing the TMD of human α1GABAARs to the ECD of ELIC, a prokaryotic pLGIC from Erwinia chrysanthemi19 (Fig. 1a). The final residue R199 in the ECD of ELIC was fused with the first residue K222 in the TMD of human α1GABAAR. To facilitate crystallization, the large intracellular loop between TM3 and TM4 in α1GABAAR (G313 to N387) was replaced by the tripeptide linker from ELIC (GVE) and 12 residues at the C-terminus of α1GABAAR (R418 to Q429) were removed. Sequence details of the chimera are provided in the supplementary information (Supplementary Fig. 1). Probably because the human α1GABAAR and ELIC share 46.6% sequence similarity in their ECDs and because GABA also activates ELIC20, the α1GABAAR chimera is functional in Xenopus oocytes without sequence optimization at the ECD-TMD interface. This easy process of obtaining a functional α1GABAAR chimera is in contrast with difficulties in making the ELIC-α7 nicotinic acetylcholine receptor (nAChR) chimera, which required extensive sequence modification at the ECD-TMD interface in order to mimic the functions of α7nAChR21. ELIC agonists20, including propylamine (PPA), activate oocytes expressing the α1GABAAR chimera in a concentration dependent manner (Fig. 1b). The neurosteroid anesthetic alphaxalone potentiates the agonist-induced current (Fig. 1c), directly activates the α1GABAAR chimera in the absence of agonist and desensitizes the channel during its continued application (Fig. 1d). Picrotoxin, known to inhibit GABAARs, also inhibits the α1GABAAR chimera (Fig. 1e). These functional responses of the α1GABAAR chimera to alphaxalone and picrotoxin resemble those observed on the authentic α1-containing GABAARs22,23,24. Alphaxalone inhibits ELIC (Supplementary Fig. 2). The distinctly different functional and pharmacological properties of ELIC and GABAARs25,26 offer an opportunity to dissect the role of the GABAAR TMD in a chimeric form.
Construction and function of the ELIC-α1GABAAR chimera. a Schematic representation of the ELIC-α1GABAAR chimera, constructed by fusing the ELIC extracellular domain (yellow) ending at residue R199 to the α1GABAAR transmembrane domain (orange) beginning at residue K222. See more details in the supporting information (Supplementary Fig. 1). b The ELIC agonist propylamine (PPA) activates the α1GABAAR chimera expressed in Xenopus oocytes in a concentration dependent manner (EC50 = 19.8 ± 1.5 μM, n = 4). c The neurosteroid alphaxalone potentiates the current of oocytes expressing the α1GABAAR chimera: (left) representative potentiation trace by 0.1 μM alphaxalone; (right) concentration-response potentiation curve with EC50 = 45.5 ± 10.7 nM (n = 6). d Representative traces showing that 3 or 1 μM alphaxalone activates and then quickly desensitizes the α1GABAAR chimera (current disappears during alphaxalone application as marked by arrows). e Picrotoxin, a known GABAAR blocker, inhibits the α1GABAAR chimera. Note the quick desensitization by 10 μM PPA. Error bars in b and c represent SEM. Scale bars in c, d, and e represent 30 s (horizontal) and 10 nA (vertical)
X-ray structures of apo and desensitized α1GABAAR chimeras
The α1GABAAR chimera was expressed in E. coli and purified in DDM micelles, in which the chimera forms homogenous pentamers suitable for crystallization (Supplementary Fig. 3). We determined x-ray structures of the α1GABAAR chimera in the apo and desensitized states with 3.2 and 3.45 Å resolutions, respectively (Table 1; Fig. 2). Alphaxalone activates and subsequently desensitizes the α1GABAAR chimera within a minute. Thus, on the crystallization time scale, the alphaxalone-bound α1GABAAR chimera is in a desensitized state (Fig. 1d).
Crystal structures of the ELIC-α1GABAAR chimera. a Side (left) and bottom (right) views of the α1GABAAR chimera in complex with alphaxalone. Alphaxalone binding to the inter-subunit sites in the TMD is indicated by the FO–FC omit electron density map contoured at 4 σ (green mesh). b Side view of the apo α1GABAAR chimera. One of the five subunits is covered with the 2FO–FC electron density map contoured at 1 σ (blue mesh). A zoom-in view of the interfacial region shows the representative residue contacts at the interface between the TM2–TM3 loop and the Cys-loop or β1–β2 linker in the ECD: F119-A285 (3.1 Å), F116-Y282 (2.4 Å), T28-K279 (4.5 Å)
The desensitized (Fig. 2a) and apo (Fig. 2b) α1GABAAR chimeras retain the common structural characteristics of pLGICs11,12,13,19,27,28,29,30,31,32,33,34, including well-folded β-sheets in the ECD and α-helices in the TMD. The pentameric assembly provides a central pore for ion permeation (Fig. 2a). In addition to the peptide bond linking the last ECD residue R199 of ELIC to the first TMD residue K222 of α1GABAAR (Fig. 1a), the ECD and TMD in the chimera are structurally coupled by extensive interface interactions (Fig. 2b) that have been suggested to be crucial for functional channels35,36. Notably, K279 in the TM2–TM3 loop, a conserved residue in GABAARs, engages in a polar interaction with T28 of the β1–β2 linker in the adjacent subunit. A cis-conformation of P120, a highly conserved residue in all pLGICs, leads to a Cys-loop orientation that enables the backbone carbonyl of neighboring residues (F119 and F116) to participate in polar contacts with the TM3 residue A285 and Y282 in the TM2–TM3 loop. These ECD–TMD interactions have also been reported for structures of β3GABAAR13 and GLIC-α1GABAAR11.
The apo α1GABAAR chimera shows little spontaneous leaking current in electrophysiology measurements and its x-ray structure offers a glance at the transmembrane pore of α1GABAAR in the apo state (Fig. 3). The pore-lining TM2 helix in each subunit is oriented with 10.2° radial and 2.3° lateral tilting angles so that the pore radius is largest at the extracelluar end and is reduced gradually toward the intracellular end. The most constricted radius (~2.0 Å) is at V257 (2′) (Fig. 3b, c). The pore radii at both V257 (2′) and P253 (−2′) are too small to allow passage of a hydrated Cl− ion. This closed channel shows no contraction at L264 (9′) because the L264 sidechains are tangential to the pore axis. The pore profile is considerably different from those reported previously for resting-state pLGICs (Fig. 3c, Supplementary Fig. 4), including ELIC19,37, GLIC38, GluCl29, and 5HT3AR33, for which the conserved leucine (9′) constitutes the narrowest pore or channel gate. However, a previous study of α1β1γ2GABAAR using the scanning-cysteine-accessibility method (SCAM) reported that, in the absence of GABA, charged sulfhydryl reagents applied from the extracellular end of the resting-state channel were able to penetrate to the level of α1V257 (2′)39, suggesting a similar pore profile to that shown in the structure of the apo α1GABAAR chimera (Fig. 3).
Crystal structures of the pore of the ELIC-α1GABAAR chimera. a Pore lining residues in the TM2 helices of the apo α1GABAAR chimera covered by the 2FO–FC electron density map contoured at 1 σ (blue mesh). b Overlaid structures of the pore-lining TM2 helices from apo (orange) and alphaxalone-bound (cyan) α1GABAAR chimeras. Blue and red dots define apo α1GABAAR chimera pore radii greater or less than the radius of a hydrated Cl− ion (3.2 Å), respectively. c Comparison of the pore radii of the α1GABAAR chimera in the apo (orange) and desensitized states (cyan) to the desensitized β3GABAAR (green), the desensitized GLIC-α1GABAAR chimera (blue), the desensitized α5GABAAR chimera (brown), and resting ELIC (yellow)
The pore profile of the α1GABAAR chimera desensitized by alphaxalone is similar to the pore profile of the apo chimera, except that the most constricted pore is located at P253 (−2′) in the desensitized structure (Fig. 3c). The putative gate at the −2′ position in the desensitized α1GABAAR chimera matches with observations from the structures of desensitized GLIC-α1GABAAR11, β3-α5GABAAR12, β3GABAAR13, α340, and α134 glycine receptors (GlyRs), and α4β2nAChR27 (Fig. 3c, Supplementary Fig. 4).
Global twisting and blooming movements of pLGICs have been proposed to accompany functional conformation changes29,38,41. Both the apo and alphaxalone-bound ELIC-α1GABAAR are closed channels (Fig. 3) and no significant blooming is observed. Relative to the aligned structure of the apo ELIC-α1GABAAR, the alphaxalone-bound structure shows 1.3° inward and 0.18° outward radial tilt from the pore axes in the ECD and TMD, respectively. The alphaxalone-bound structure shows a small counterclockwise twist (0.80°) of the ECD and a clockwise twist (1.37°) of the TMD around the pore axis (Supplementary Fig. 5). This direction of twist leads toward channel opening based on a structure survey of pLGICs under different functional states29,34,38. In the case of ELIC-α1GABAAR, however, the small magnitude of twist in the desensitized alphaxalone-bound structure with respect to the apo structure is consistent with the fact that both channels are closed (Fig. 3).
The binding mode of alphaxalone
The structure of the α1GABAAR chimera co-crystallized with alphaxalone shows that alphaxalone binds to five equivalent inter-subunit sites close to the intracellular end of the α1GABAAR TMD (Fig. 2a, Supplementary Fig. 6a). Residues within 4-Å of alphaxalone are from TM3 of the principal subunit (A305, T306, Y309, and F310) and from TM1 (Q242, V243, W246) and TM4 (P401) of the neighboring complementary subunit (Fig. 4, Supplementary Fig. 6b). The aromatic ring of the conserved residue W246 is parallel to the C ring of alphaxalone, establishing a ring stacking interaction that potentially stabilizes the binding. The polar sidechains of Q242 and T306 are within reach to form putative hydrogen bonds with alphaxalone (Fig. 4b).
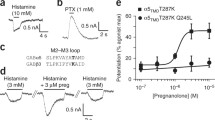
Alphaxalone binding mode in the α1GABAAR chimera. a 2D chemical structure of alphaxalone with rings and carbon atoms labeled according to the IUPAC standard for steroids. b Crystal structure of alphaxalone (cyan) bound to a pocket lined by residues in the transmembrane domain from the principal (yellow) and complementary (white) subunits of the α1GABAAR chimera. Alphaxalone is surrounded by the 2FO–FC electron density map contoured at 1 σ (blue mesh). Three residues in close contact with alphaxalone are highlighted. Functional validation of the alphaxalone-binding site was performed by c activation of Xenopus oocytes expressing WT (solid circles), T306A (open circles), Q242L (solid squares) and W246L (open squares) α1GABAAR chimeras by propylamine (PPA) with EC50 = 20 ± 1, 23 ± 2, 39 ± 3, and 3300 ± 300 μM, respectively; d alphaxalone (0.1 μM) potentiation at the EC10 concentration of PPA; e alphaxalone (3 μM) activation normalized to EC100 PPA activation for each construct. Error bars represent SEM (n ≥ 3 oocytes). Statistical significance was assessed by one-way ANOVA followed by Fisher’s LSD post-hoc test. Asterisks indicate statistical difference from WT at p < 0.001 (***) and p < 0.05 (*)
To assess the importance of these interactions to the functional modulation of α1GABAAR by alphaxalone, we performed electrophysiology measurements on the wild type (WT) and three mutants of the α1GABAAR chimera. Two of these mutants, Q242L and T306A, are similar to the WT α1GABAAR chimera in their response to the orthosteric agonist PPA, whereas the PPA EC50 of the W246L mutant is increased two orders of magnitude compared to WT (Fig. 4c), signaling the importance of this conserved residue W246 in the function of GABAARs. For the Q242L and T306A mutants, 0.1 μM alphaxalone still potentiates channel currents but with smaller magnitudes compared to the WT chimera (Fig. 4d). The W246L mutation has a stronger effect and almost completely abolishes potentiation by the same concentration of alphaxalone (Fig. 4d). As expected, these mutations also reduce channel activation by alphaxalone (Fig. 4e). These findings from the α1GABAAR chimera are consistent with the results from the full-length GABAARs. A number of previous studies on full-length GABAARs show that the α1-W246L and α1-Q242L mutations significantly suppressed allopregnanolone potentiation42,43. Similarly, alphaxalone potentiation of α1β3 GABAAR is also significantly reduced in the α1-T306A mutant compared to the WT receptor (Supplementary Fig. 7).
We performed molecular dynamics (MD) simulations (Fig. 5, Supplementary Fig. 8) to quantify the stability of alphaxalone interactions with W246, Q242, and T306. The stability of ring stacking is measured by the distances between two pairs of atoms in alphaxalone and W246 (Fig. 5a, b). The narrow distance distributions over the course of simulations suggest stable ring stacking. The same simulation data are used to estimate probabilities of forming hydrogen bonds between relevant hydroxyl and carbonyl groups in alphaxalone and Q242 or T306 (Fig. 5c), showing that alphaxalone has a ~ 55% probability to form a hydrogen bond with Q242, but only a ~23% probability with T306. The data agree with mutagenesis and functional results (Fig. 4) that show a stronger influence of Q242 than T306 on functional effects of alphaxalone.
Alphaxalone interactions with nearby residues in molecular dynamics (MD) simulations. a A representative snapshot from MD simulations shows alphaxalone contacts with T306, Q242, and W246. Distances between alphaxalone and the residues are measured as marked by the dash lines. b Histograms of distances between alphaxalone and W246 atoms (1 and 2) shown in a. c Histograms of distances between alphaxalone and Q242 or T306 atoms (3 and 4, respectively) shown in a. Data in b and c are from three replicate 50-ns simulations, where snapshots were collected every 100-ps for analysis. Distances were measured for each of the five alphaxalone molecules per replicate simulation (500 snapshots × 3 replicates × 5 alphaxalone = 7500 distances in total)
A path for alphaxalone activation and desensitization
In the absence of agonist binding to the orthosteric site in the ECD, alphaxalone allosterically triggers the activation and subsequent desensitization of ELIC-α1GABAAR (Fig. 1d). What is the structural basis for channel activation and desensitization by alphaxalone? In addition to the small twist movement of the ECD and TMD along the channel axis (Supplementary Fig. 5), we notice a set of conformational changes in the aligned x-ray structures of the apo and alphaxalone-bound ELIC-α1GABAAR (Fig. 6, Supplementary Fig. 9). Alphaxalone binding to the bottom of the TMD introduces structural changes, including the orientation of W246 in the TM1, presumably to optimize its interaction with alphaxalone (Fig. 6). The change passes to the TM1–TM2 linker and further to the TM2, particularly at P253 (−2′) and V257 (2′), resulting in a backbone RMSD of 0.95 Å for the residues covering from W246 in TM1 to V257 (2′) in TM2 (Fig. 6b). The structural changes further propagate up to the ECD–TMD interface and other ECD regions, as evidenced by the observed RMSDs: 0.66 Å for the TM2–TM3 loop (P278-T284), 0.61 Å for the pre-TM1 (R199-I223), 0.58 Å for the β1–β2 linker (V26-E30), 0.63 Å for the Cys-loop (N112-F126), 1.01 Å for loop A (N80-S84), and 0.84 Å for loop C (D172-N186) (Supplementary Fig. 9). Although these small changes do not necessarily reflect the actual magnitude of the structural changes involved in the functional transitions from the resting to activated and subsequent desensitized states, they suggest a plausible path starting at the bottom of the TMD for alphaxalone-induced channel activation and desensitization.
Alphaxalone-induced structural changes at the bottom of the TMD. a Bottom view of overlaid TM1-TM2 structures of the apo (orange) and alphaxalone-bound (cyan) ELIC-α1GABAAR. b Side view of overlaid structures of apo (principal subunit - gold; complementary subunit - orange) and alphaxalone-bound (principal subunit - blue; complementary subunit - cyan) ELIC-α1GABAAR. For clarity, only TM2 and TM3 are shown in the principal subunit and only TM1 and TM2 are shown in the complementary subunit. The arrow highlights structural perturbations originating from the alphaxalone binding site near W246 through the TM1–TM2 linker to the pore-lining residues P253 (−2′) and V257 (2′). c The 2FO-FC electron density maps (blue mesh, contoured at 1 σ) covering TM1–TM2 in the apo (left) and alphaxalone-bound (right) ELIC-α1GABAAR. The sidechains for residues W246 to V257 (2′) are highlighted
Structural cooperation and independence of the ECD and TMD
The structural independence of individual domains in pLGICs and cooperativity between these domains are essential for constructing functional chimeric channels11,12,44. In this study, ELIC-α1GABAAR provides a window into how the ECD and TMD both retain their individuality in the chimera and cooperate structurally to form a functional channel. Crystal structures of apo ELIC-α1GABAAR and apo ELIC (aligned along their common ECDs) show that even under the same ECD, the chimera TMD independently adopts the α1GABAAR structure that is significantly different from the ELIC TMD structure (Fig. 7a, b). The ECD influence on the TMD structure is rather limited. Indeed, even an isolated TMD of pLGICs can form functional channels that retain some characteristics of the parent channels45,46,47. The structural cooperativity between the individual domains is also observed (Fig. 7). Although most parts of the ECD in the two structures overlap well, there is an overall ECD backbone RMSD of 2.1 Å, resulting mainly from the displacement of several key regions in the chimera (Fig. 7c). The subtle and profound displacements of the respective pre-TM1 and the TM2–TM3 loop in the chimera (Fig. 7b) may have led to the conformational adjustments of the Cys-loop and β1–β2 linker at the ECD–TMD interface, as well as alterations further up at loop A and loop C (Fig. 7c). As an example of structural cooperation between the ECD and TMD, the conserved residue K279 in the TM2–TM3 of GABAARs and its equivalent residue R255 in ELIC are oriented in different directions so that they form respective inter-subunit and intra-subunit polar interactions with T28 in the β1–β2 linker (Supplementary Fig. 10). The inter-subunit K279–T28 interactions would not be possible without the β1–β2 linker relocation in the chimera. An equivalent inter-subunit polar interaction (K274-E52) is also observed in the crystal structure of β3GABAAR13. The importance of this lysine residue in channel gating of GABAARs has been well recognized35,36. Mutation of this lysine impairs function of GABAARs36 and is associated with epilepsy48,49.
Structural changes in the ECD due to different TMDs. a Top view of crystal structures of apo ELIC-α1GABAAR (orange) and apo ELIC (green), aligned along all ECD residues. b Displacements of Cα atoms of equivalent residues in apo ELIC-α1GABAAR and apo ELIC in the pre-TM1 and the TM2–TM3 loop are labeled. Both the pre-TM1 and TM2–TM3 loop regions may affect the ECD. c Structural changes in the ECD are measured by displacements of the highlighted residues in several key regions
Discussion
We have determined both the apo and desensitized structures for the ELIC-α1GABAAR chimera. Although our desensitized structure was obtained through alphaxalone binding that directly activates and subsequently desensitizes the chimera (Fig. 1), it demonstrates the same gate in the pore at the cytosolic portal as observed in the structures of β3GABAAR, α1GlyR, α3GlyR, and α4β2nAChR13,27,34,40 desensitized by orthosteric ligands (Supplementary Fig. 4) as well as the more recently determined GLIC-α1GABAAR and β3-α5GABAAR11,12 (Fig. 3c). The consistency of the pore constriction at the −2′ position among these desensitized pLGICs, independent of whether they are desensitized by orthosteric (in most cases) or allosteric (such as in this study) ligands, settles the debate about what causes non-conductivity in a desensitized pLGIC50,51,52,53,54.
No structure has been reported in the past for a resting GABAAR. The apo ELIC-α1GABAAR structure presents a look at the TMD of α1GABAAR in the resting state. The pore profile shows the most constriction at V257 (2′) (Fig. 3), not at the conserved L264 (9′) that is perceived as a gate of channel activation based on previously published structures of the resting pLGICs, including GluCl29, 5HT3AR31,33, GLIC38, and ELIC19,37 (Fig. 3, Supplementary Fig. 4). Because of the unconventional occlusion location in the pore of the apo ELIC-α1GABAAR, it is natural to question whether the apo structure reflects a true resting-state conformation of ELIC-α1GABAAR. We have analyzed the question from several different angles. Could the DDM detergent and crystallization conditions have introduced conformational biases to the resting ELIC-α1GABAAR? DDM has been used for purification and crystallization of many different pLGICs, including GluCl that exhibits different pore conformations for the closed/resting state and open/activated state channels29. Both ELIC and ELIC-α1GABAAR are crystallized with DDM, but show distinctly different pore conformations in the resting-state condition. Thus, it is not convincing to simply attribute the unconventional resting-state pore profile to detergent- and/or crystallization-related artifacts, even though one cannot completely rule out this possibility. Can we assign the apo ELIC-α1GABAAR to a desensitized state in the absence of agonist because of the similar structures of the apo and desensitized ELIC-α1GABAAR? The classic definition of desensitization requires the presence of agonists or channel modulators. Moreover, our electrophysiology functional data also do not support this assignment. Can a resting-state GABAAR adopt the non-conducting channel conformation similar to those observed for desensitized pLGICs? Our apo structure suggests such a possibility. In addition, this possibility has been suggested previously by a SCAM study39, in which the charged sulfhydryl reagents were able to penetrate from the ECD of α1β1γ2GABAAR to the pore at the level of α1V257 (2′) in the resting state. We also notice that GABAARs have a different pattern of hydrophobicity in pore-lining residues compared to those pLGICs with known resting structures (Supplementary Fig. 11, Supplementary Table 1), including the anion-conducting GluCl29 along with the cation-conducting 5HT3AR31,33, Torpedo nAChR55, GLIC38, and ELIC19,37, which have consecutive hydrophobic rings at the 9′, 13′ and 17′ (16′) positions. In contrast, GABAARs have mostly hydrophilic pore-lining residues except at the 9′, −2′, and 2′ positions (Supplementary Fig. 11). Combining all aforementioned information with the pore radius profiles for the apo and desensitized α1GABAAR chimera, we propose a channel gate for both activation and desensitization at the cytoplasmic end of the pore formed by the 2′ and −2′ residues. Whether the finding can be generalized to other GABAARs mandates additional structural investigations, particularly with apo GABAARs.
The binding site identified for the anesthetic alphaxalone overlaps well with the site elegantly illustrated previously for the endogenous potentiating neurosteroids tetrahydro-deoxycorticosterone (THDOC) and pregnanolone in the respective GLIC-α1GABAAR and β3-α5GABAAR chimeras11,12 (Supplementary Fig. 12), suggesting that neurosteroid potentiation and activation of GABAARs are mediated through this conserved inter-subunit site close to the base of the TMD. It is also noteworthy that photoreactive analogs of the intravenous general anesthetics propofol9 and etomidate56,57 have been photolabeled to the TM1 residues M236 and I239, showing the proximity of the binding sites for these intravenous anesthetics and alphaxalone (Supplementary Fig. 6). A more recent mutagenesis/electrophysiology study of α1β3γ2LGABAARs also suggest the proximity between the alphaxalone site and the sites for propofol and etomidate at the transmembrane β+-α− interface18. Among all the interactions between alphaxalone and α1GABAAR shown in our structure, the ring stacking of the conserved W246 in the TM1 with the C ring of alphaxalone is most critical. Eliminating this interaction in the W246L mutant completely abolishes potentiation and activation by alphaxalone (Fig. 4). Putative hydrogen bonding, especially through Q242 (one helical turn above W246) to the 3α-hydroxyl of alphaxalone, is also important as the potentiation by alphaxalone was reduced substantially in Q242L and only moderately in T306A (Fig. 4). The order of W246, Q242, and T306 in their impact to functional modulation of alphaxalone is associated with the stability of their interactions with alphaxalone as revealed in MD simulations (Fig. 5), in which the ring stacking with W246 is arguably the steadiest interaction. The probability for alphaxalone to form a hydrogen bond with Q242 is much higher than with T306 (Fig. 5). A more dominant role of the complementary α1 subunit TM1 containing W246 and Q242 in alphaxalone potentiation agrees with the finding in heteromeric GABAARs42, where allopregnanolone binding to the β+-α− interface is a major contributor to functional potentiation.
Neurosteroids and general anesthetics, including alphaxalone, not only potentiate but also directly activate GABAARs at higher concentrations10,15,16,58,59. The structures of the α1GABAAR chimera determined in this study suggest a mechanism suitable for direct activation by alphaxalone, in which the TM1–TM2 linker at the bottom of the TMD plays a crucial role in transducing conformational changes that originate from alphaxalone binding at the TM1 (W246 and Q242) to the pore-lining TM2 helices (Fig. 6). Because the same neurosteroid binding site in the TMD is also responsible for potentiation11, the signaling pathway starting at the bottom of the TMD revealed in our crystal structures likely is relevant for both potentiation and activation of GABAARs. Similar conformational changes are also found in neurosteroid potentiation of the β3-α5GABAAR chimera, though the changes induced by neurosteroids in that case may be diluted by the channel’s high propensity for spontaneous opening12. Our structures also show that conformational changes introduced by binding of the neurosteroid anesthetic alphaxalone and mediated by the TM1–TM2 linker can propagate beyond the TM2 (Supplementary Figs. 5, 9). Together, these results underscore three conclusions. First, alphaxalone binding to the inter-subunit site at the bottom of the TMD can introduce global conformational changes involved in channel potentiation, activation, and desensitization. Second, the TM1–TM2 linker, noted previously for its involvement in channel desensitization60, plays a key role in mediating activation and potentiation by neurosteroids. Finally, this mechanism of activation or potentiation starting at the bottom of the TMD can be exploited in the rational search for new GABAAR modulators with better potency and efficacy.
Methods
Protein expression and purification
The ELIC-α1GABAAR chimera was constructed using overlapping PCR by fusing the ELIC ECD ending at R199 with the human α1GABAAR TMD starting at K222 (Supplementary Fig. 1). Primer sequences are provided in Supplementary Table 2. To facilitate crystallization, we replaced the lengthy intracellular loop (G314 to N387) of α1GABAAR with the short linker (G290-V291-E292) connecting TM3 and TM4 in ELIC, and also deleted 12 residues at the C terminus of α1GABAAR (REPQLKAPTPHQ). ELIC-α1GABAAR, similar to the ELIC construct designed for E. coli expression37, was cloned in pET26b vector under an IPTG-inducible promoter. N-terminally His-tagged maltose binding protein (MBP) was fused to the N terminus of ELIC-α1GABAAR and a TEV enzyme cleavage site was inserted between MBP and ELIC-α1GABAAR.
For protein expression, the plasmid was transformed into Rosetta(DE3)pLysS (Novagen) cells under double selection with kanamycin (50 μg/mL) and chloramphenicol (35 μg/mL). Three to five isolated colonies were used to inoculate 100 mL LB media with the antibiotics and grown overnight at 37 °C in an environmental shaker at 250 rpm. The overnight culture was then diluted 1:100 into 6 × 1 L of LB media with the antibiotics and grown to an optical density of ~0.7–0.8. All six liters were harvested (5000 rpm, 20 min, 4 °C) and suspended into 2 L of LB media supplemented with 0.5 M sorbitol. The concentrated cells were equilibrated in an environmental shaker at 15 °C and 250 rpm for 1 h before inducing expression with 0.2 mM isopropyl β-d-1-thiogalactopyranoside. The cells were harvested after ~20 h expression, re-suspended in 50 mM Tris pH 8, 300 mM NaCl and flash frozen in liquid N2. For protein purification, the frozen cells were thawed and homogenized with the addition of 2 mg/mL lysozyme and 1 μL benzonase before lysis using an M-110Y microfluidizer processor (Microfluidics). Membranes were pelleted by ultracentrifugation at 45 krpm at 4 °C in a Type 45Ti rotor. The fusion protein was extracted with 2% (w/v) n-dodecyl-β-d-maltoside (DDM, Anatrace), purified using a 5-mL HisTrap HP column (GE Healthcare), eluted by 250 mM imidazole, and then desalted in a buffer of 50 mM sodium phosphate at pH 8, 250 mM NaCl, and 0.05% (w/v) DDM. MBP was cleaved by TEV protease overnight and separated from ELIC-α1GABAAR using a 1-mL HisTrap HP column. The pentameric fraction of ELIC-α1GABAAR was collected in a buffer containing 10 mM sodium phosphate at pH 8, 150 mM NaCl, 0.05% (w/v) DDM by size exclusion chromatography using a Superdex 200 10/300GL column (GE Healthcare). The purified pentameric ELIC-α1GABAAR was concentrated to ~4 mg/mL for crystallization.
Crystallography and data analysis
Crystals were obtained using the sitting-drop vapor diffusion method at 4 °C. All chemicals used for crystallization were purchased from Sigma-Aldrich (St. Louis, MO) unless stated otherwise. The reservoir solution contained 19–21.5% PEG 400, 400 mM NaSCN, 100 mM MES buffer at pH 6.1 and was mixed with the protein in a 1:1 ratio for crystallization of apo ELIC-α1GABAAR. For co-crystallization, alphaxalone in the concentration of 10–1000 μM was mixed with the protein for ~30 min before setting up trays. The crystals were obtained typically after 3–4 weeks and cryo-protected by using up to 35% PEG400 before being flash-frozen in liquid nitrogen for storage.
The x-ray diffraction data of apo crystals were collected on the Southeast Regional Collaborative Access Team (SERCAT) beamline 22-ID at the Advanced Photon Source, Argonne National Laboratory. The data of alphaxalone-bound co-crystals were collected on the beamline 12–2 at the Stanford Synchrotron Radiation Lightsource (SSRL). The collected datasets were indexed, integrated, and scaled using the XDS program61. The scaled datasets were merged with Aimless62. Details of crystal parameters and data collection statistics for both crystals are provided in Table 1.
The molecular replacement method was used for the initial structure determination. The ECD of ELIC (PDB code: 4Z90)63 and the TMD of β3GABAAR (PDB code: 4COF)13 were used as the isolated search ensembles for the molecular replacement solution by PHASER64. The TMD of the initial solution was then mutated to α1GABAAR and used as the new search model for molecular replacement. The results clearly indicated two pentamers in the crystallographic asymmetric unit. This initial model was then refined in both the BUSTER65 and Phenix66 programs. Non-crystallographic symmetry (NCS) restraints were used throughout refinement. After the initial refinements, 2FO-FC composite and simulated annealing omit maps were calculated and examined to improve the model. The entire structure was iteratively analyzed, rebuilt with the program Coot67 and refined in Phenix66.
The refined pentameric structure of apo ELIC-α1GABAAR was used as the molecular replacement model for the structure of alphaxalone-bound crystals. The solution also contained two pentamers in the crystallographic asymmetric unit. After initial rounds of refinement, the electron density difference map (FO-FC) showed strong density for alphaxalone in the TMD. The initial structure of alphaxalone was obtained from a previous publication68 and the alphaxalone molecule was fit to the FO-FC difference density in Coot. The Translation-Libration-Screw-rotation model (TLS) and torsional non-crystallographic symmetry (NCS) restraints were applied to all subunits of two pentamers in the asymmetric unit and the final model was refined in Phenix66.
The geometry and stereochemistry of the final structures were validated by the program MolProbity69. The refinement statistics are given in Table 1. All molecular graphics were prepared using PyMol70 or VMD71.
Electrophysiology
Functional properties of the α1GABAAR chimera, its mutants, and ELIC were measured using two-electrode voltage clamp (TEVC) electrophysiology of Xenopus laevis oocytes expressing the channels of interest. All procedures involving Xenopus laevis oocytes were approved by the University of Pittsburgh Institutional Animal Care and Use Committee. A T7 promoter followed by DNA encoding a selected channel was inserted into the pCMV-mGFP Cterm S11 Neo Kan vector (Theranostech, NM). Capped complementary RNA was synthesized with the mMessage mMachine T7 kit (Ambion), purified with the RNeasy kit (Qiagen), and injected (4–25 ng) into Xenopus laevis oocytes (stages 5–6). Site-directed mutagenesis was performed using the QuickChange Lightning Kit (Agilent) and confirmed by sequencing at the University of Pittsburgh Health Sciences Genomics Research Core. Oocytes were maintained at 10 or 18 °C in modified Barth’s solution containing 88 mM NaCl, 1 mM KCl, 2.4 mM NaHCO3, 15 mM HEPES, 0.3 mM Ca(NO3)2, 0.41 mM CaCl2, 0.82 mM MgSO4, 10 μg mL−1 sodium penicillin, 10 μg mL−1 streptomycin sulfate, and 100 μg mL−1 gentamycin sulfate at pH 6.7. Two-electrode voltage clamp experiments were performed at room temperature 1–4 days after injection with an OC-725C Amplifier (Warner Instruments) and Digidata 1440 A digitizer (Axon Instruments) in a 20-μL oocyte recording chamber (Automate Scientific). Oocytes were clamped to a holding potential of −60 mV. Oocytes expressing the α1GABAAR chimera and its mutants were recorded using ND96 solution (96 mM NaCl, 2 mM KCl, 1.8 mM CaCl2, 1 mM MgCl2 and 5 mM HEPES at pH 7.4). The recording solution for oocytes expressing ELIC contained 130 mM NaCl, 0.1 mM CaCl2, and 10 mM HEPES at pH 7.4 to minimize inhibition by divalent cations72. Alphaxalone stock solution was prepared in DMSO and the final DMSO concentration used for experiments was no more than 0.01%. Data were collected and processed using Clampex 10 (Molecular Devices). Non-linear regressions were performed using Prism 7.0 (Graphpad).
Molecular dynamics simulations
The crystal structure of the alphaxalone-bound ELIC-α1GABAAR (PDB code: 6CDU) was used as initial coordinates in MD simulations. The alphaxalone-bound ELIC-α1GABAAR was embedded into a pre-equilibrated lipid bilayer composed of POPC/cholesterol in a 5:1 molar ratio73,74 using the GROMACS g_membed tool75. The system was solvated in TIP3P water, ionized with 100 mM NaCl, and contained ∼172,000 atoms.
Three replicate 50-ns MD simulations were run using GROMACS 201676 and the CHARMM36 force field77. Alphaxalone geometry and parameters were assigned by analogy using the CHARMM General Force Field (CGenFF) for drug-like molecules77 and refined using the Force Field Toolkit (ffTK) protocol78. Optimized alphaxalone structure and parameters are provided in the Supporting Materials (Supplementary Fig. 13, Supplementary Tables 3–5). The system was energy minimized for 10,000 steps with harmonic position restraints of 10 K kJ/mol/nm2 on the protein backbone atoms, followed by 3 ns of equilibration, during which position restraints on the protein backbone were gradually reduced from 10,000 to 0 kJ/mol/nm2. Production simulations were performed at a constant pressure and temperature (NPT) of 1 atm and 310 K with a 2-fs time step. Bonds were constrained using the LINCS algorithm79. The particle mesh Ewald method was used for long-range electrostatic interactions80. A 12-Å cutoff was used for nonbonded interactions. Full electrostatic and non-bonded interactions were evaluated every 2 fs and the neighbor list was updated every 10 fs. Systems were simulated with periodic boundary conditions in three dimensions.
Data availability
Data supporting the findings of this manuscript are available from the corresponding author upon reasonable request. Crystal structures of apo and alphaxalone-bound α1GABAAR chimeras are deposited in the Protein Data Bank with accession codes 6D1S and 6CDU, respectively.
References
Farrant, M. & Nusser, Z. Variations on an inhibitory theme: phasic and tonic activation of GABA(A) receptors. Nat. Rev. Neurosci. 6, 215–229 (2005).
Olsen, R. W. & Sieghart, W. GABA A receptors: subtypes provide diversity of function and pharmacology. Neuropharmacology 56, 141–148 (2009).
Brickley, S. G. & Mody, I. Extrasynaptic GABA(A) receptors: their function in the CNS and implications for disease. Neuron 73, 23–34 (2012).
Franks, N. P. General anaesthesia: from molecular targets to neuronal pathways of sleep and arousal. Nat. Rev. Neurosci. 9, 370–386 (2008).
Jurd, R. et al. General anesthetic actions in vivo strongly attenuated by a point mutation in the GABA(A) receptor beta3 subunit. FASEB J. 17, 250–252 (2003).
Mihic, S. J. et al. Sites of alcohol and volatile anaesthetic action on GABA(A) and glycine receptors. Nature 389, 385–389 (1997).
Forman, S. A. & Miller, K. W. Anesthetic sites and allosteric mechanisms of action on Cys-loop ligand-gated ion channels. Can. J. Anaesth. 58, 191–205 (2011).
Eaton, M. M. et al. Multiple non-equivalent interfaces mediate direct activation of GABAA receptors by propofol. Curr. Neuropharmacol. 14, 772–780 (2016).
Jayakar, S. S. et al. Multiple propofol-binding sites in a gamma-aminobutyric acid type A receptor (GABAAR) identified using a photoreactive propofol analog. J. Biol. Chem. 289, 27456–27468 (2014).
Hosie, A. M., Wilkins, M. E., da Silva, H. M. & Smart, T. G. Endogenous neurosteroids regulate GABAA receptors through two discrete transmembrane sites. Nature 444, 486–489 (2006).
Laverty, D. et al. Crystal structures of a GABAA-receptor chimera reveal new endogenous neurosteroid-binding sites. Nat. Struct. Mol. Biol. 24, 977–985 (2017).
Miller, P. S. et al. Structural basis for GABAA receptor potentiation by neurosteroids. Nat. Struct. Mol. Biol. 24, 986–992 (2017).
Miller, P. S. & Aricescu, A. R. Crystal structure of a human GABAA receptor. Nature 512, 270–275 (2014).
Zhu, S. et al. Structure of a human synaptic GABAA receptor. Nature 559, 67–72 (2018).
Cottrell, G. A., Lambert, J. J. & Peters, J. A. Modulation of GABAA receptor activity by alphaxalone. Br. J. Pharmacol. 90, 491–500 (1987).
Siegwart, R., Krahenbuhl, K., Lambert, S. & Rudolph, U. Mutational analysis of molecular requirements for the actions of general anaesthetics at the gamma-aminobutyric acidA receptor subtype, alpha1beta2gamma2. BMC Pharmacol. 3, 13 (2003).
Nourmahnad, A. et al. Tryptophan and cysteine mutations in M1 helices of alpha1beta3gamma2L gamma-aminobutyric acid type A receptors indicate distinct intersubunit sites for four intravenous anesthetics and one orphan site. Anesthesiology 125, 1144–1158 (2016).
Ziemba, A. M. et al. Alphaxalone binds in inner transmembrane beta+-alpha- interfaces of alpha1beta3gamma2 gamma-aminobutyric acid type A receptors. Anesthesiology 128, 338–351 (2018).
Hilf, R. J. & Dutzler, R. X-ray structure of a prokaryotic pentameric ligand-gated ion channel. Nature 452, 375–379 (2008).
Zimmermann, I. & Dutzler, R. Ligand activation of the prokaryotic pentameric ligand-gated ion channel ELIC. PLoS Biol. 9, e1001101 (2011).
Tillman, T. S., Seyoum, E., Mowrey, D. D., Xu, Y. & Tang, P. ELIC-alpha7 Nicotinic acetylcholine receptor (alpha7nAChR) chimeras reveal a prominent role of the extracellular-transmembrane domain interface in allosteric modulation. J. Biol. Chem. 289, 13851–13857 (2014).
Weir, C. J. et al. The interaction of anaesthetic steroids with recombinant glycine and GABAA receptors. Br. J. Anaesth. 92, 704–711 (2004).
Maitra, R. & Reynolds, J. N. Modulation of GABA(A) receptor function by neuroactive steroids: evidence for heterogeneity of steroid sensitivity of recombinant GABA(A) receptor isoforms. Can. J. Physiol. Pharmacol. 76, 909–920 (1998).
Krishek, B. J., Moss, S. J. & Smart, T. G. A functional comparison of the antagonists bicuculline and picrotoxin at recombinant GABAA receptors. Neuropharmacology 35, 1289–1298 (1996).
Kinde, M. N. et al. Common anesthetic-binding site for inhibition of pentameric ligand-gated ion channels. Anesthesiology 124, 664–673 (2016).
Chen, Q. et al. Structural basis of alcohol inhibition of the pentameric ligand-gated ion channel ELIC. Structure 25, 180–187 (2017).
Morales-Perez, C. L., Noviello, C. M. & Hibbs, R. E. X-ray structure of the human alpha4beta2 nicotinic receptor. Nature 538, 411–415 (2016).
Hibbs, R. E. & Gouaux, E. Principles of activation and permeation in an anion-selective Cys-loop receptor. Nature 474, 54–60 (2011).
Althoff, T., Hibbs, R. E., Banerjee, S. & Gouaux, E. X-ray structures of GluCl in apo states reveal a gating mechanism of Cys-loop receptors. Nature 512, 333–337 (2014).
Huang, X., Chen, H., Michelsen, K., Schneider, S. & Shaffer, P. L. Crystal structure of human glycine receptor-alpha3 bound to antagonist strychnine. Nature 526, 277–280 (2015).
Hassaine, G. et al. X-ray structure of the mouse serotonin 5-HT3 receptor. Nature 512, 276–281 (2014).
Hilf, R. J. & Dutzler, R. Structure of a potentially open state of a proton-activated pentameric ligand-gated ion channel. Nature 457, 115–118 (2009).
Basak, S. et al. Cryo-EM structure of 5-HT3A receptor in its resting conformation. Nat. Commun. 9, 514 (2018).
Du, J., Lu, W., Wu, S., Cheng, Y. & Gouaux, E. Glycine receptor mechanism elucidated by electron cryo-microscopy. Nature 526, 224–229 (2015).
Kash, T. L., Jenkins, A., Kelley, J. C., Trudell, J. R. & Harrison, N. L. Coupling of agonist binding to channel gating in the GABA(A) receptor. Nature 421, 272–275 (2003).
Hales, T. G. et al. An asymmetric contribution to gamma-aminobutyric type A receptor function of a conserved lysine within TM2-3 of alpha1, beta2, and gamma2 subunits. J. Biol. Chem. 281, 17034–17043 (2006).
Pan, J. et al. Structure of the pentameric ligand-gated ion channel ELIC cocrystallized with its competitive antagonist acetylcholine. Nat. Commun. 3, 714 (2012).
Sauguet, L. et al. Crystal structures of a pentameric ligand-gated ion channel provide a mechanism for activation. Proc. Natl Acad. Sci. USA 111, 966–971 (2014).
Xu, M. & Akabas, M. H. Identification of channel-lining residues in the M2 membrane-spanning segment of the GABA(A) receptor alpha1 subunit. J. Gen. Physiol. 107, 195–205 (1996).
Huang, X., Chen, H. & Shaffer, P. L. Crystal structures of human GlyRalpha3 bound to ivermectin. Structure 25, 945–950 e942 (2017).
Cecchini, M. & Changeux, J. P. The nicotinic acetylcholine receptor and its prokaryotic homologues: structure, conformational transitions & allosteric modulation. Neuropharmacology 96, 137–149 (2015).
Bracamontes, J. R., Li, P., Akk, G. & Steinbach, J. H. A neurosteroid potentiation site can be moved among GABAA receptor subunits. J. Physiol. 590, 5739–5747 (2012).
Hosie, A. M., Clarke, L., da Silva, H. & Smart, T. G. Conserved site for neurosteroid modulation of GABA A receptors. Neuropharmacology 56, 149–154 (2009).
Moraga-Cid, G. et al. Allosteric and hyperekplexic mutant phenotypes investigated on an alpha1 glycine receptor transmembrane structure. Proc. Natl Acad. Sci. USA 112, 2865–2870 (2015).
Bondarenko, V. et al. NMR structures of the human alpha7 nAChR transmembrane domain and associated anesthetic binding sites. Biochim. Biophys. Acta 1838, 1389–1395 (2014).
Mowrey, D. D. et al. Open-channel structures of the human glycine receptor alpha1 full-length transmembrane domain. Structure 21, 1897–1904 (2013).
Bondarenko, V. et al. NMR structures of the transmembrane domains of the alpha4beta2 nAChR. Biochim. Biophys. Acta 1818, 1261–1268 (2012).
Epi, K. C. et al. De novo mutations in epileptic encephalopathies. Nature 501, 217–221 (2013).
Kodera, H. et al. De novo GABRA1 mutations in Ohtahara and West syndromes. Epilepsia 57, 566–573 (2016).
Keramidas, A. & Lynch, J. W. An outline of desensitization in pentameric ligand-gated ion channel receptors. Cell. Mol. life Sci. 70, 1241–1253 (2013).
Velisetty, P., Chalamalasetti, S. V. & Chakrapani, S. Conformational transitions underlying pore opening and desensitization in membrane-embedded Gloeobacter violaceus ligand-gated ion channel (GLIC). J. Biol. Chem. 287, 36864–36872 (2012).
Velisetty, P. & Chakrapani, S. Desensitization mechanism in prokaryotic ligand-gated ion channel. J. Biol. Chem. 287, 18467–18477 (2012).
Dellisanti, C. D. et al. Site-directed spin labeling reveals pentameric ligand-gated ion channel gating motions. PLoS Biol. 11, e1001714 (2013).
Kinde, M. N. et al. Conformational changes underlying desensitization of the pentameric ligand-gated ion channel ELIC. Structure 23, 995–1004 (2015).
Unwin, N. Refined structure of the nicotinic acetylcholine receptor at 4 A resolution. J. Mol. Biol. 346, 967–989 (2005).
Li, G. D. et al. Identification of a GABAA receptor anesthetic binding site at subunit interfaces by photolabeling with an etomidate analog. J. Neurosci. 26, 11599–11605 (2006).
Chiara, D. C. et al. Mapping general anesthetic binding site(s) in human alpha1beta3 gamma-aminobutyric acid type A receptors with [(3)H]TDBzl-etomidate, a photoreactive etomidate analogue. Biochemistry 51, 836–847 (2012).
Eaton, M. M. et al. Mutational analysis of the putative high-affinity propofol binding site in human beta3 homomeric GABAA receptors. Mol. Pharmacol. 88, 736–745 (2015).
Stewart, D., Desai, R., Cheng, Q., Liu, A. & Forman, S. A. Tryptophan mutations at azi-etomidate photo-incorporation sites on alpha1 or beta2 subunits enhance GABAA receptor gating and reduce etomidate modulation. Mol. Pharmacol. 74, 1687–1695 (2008).
Gielen, M., Thomas, P. & Smart, T. G. The desensitization gate of inhibitory Cys-loop receptors. Nat. Commun. 6, 6829 (2015).
Kabsch, W. Xds. Acta Crystallogr. D 66, 125–132 (2010).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D 69, 1204–1214 (2013).
Chen, Q. et al. Direct pore binding as a mechanism for isoflurane inhibition of the pentameric ligand-gated ion channel ELIC. Sci. Rep. 5, 13833 (2015).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
BUSTER v. 2.10.0 (UK Global Phasing Ltd., Cambridge, 2011).
Adams, P. D. et al. PHENIX: a comprehensive python-based system for macromolecular structure solution. Acta Crystallogr. Sect. D 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010).
Bandyopadhyaya, A. K. et al. Neurosteroid analogues. 15. A comparative study of the anesthetic and GABAergic actions of alphaxalone, Delta16-alphaxalone and their corresponding 17-carbonitrile analogues. Bioorg. Med. Chem. Lett. 20, 6680–6684 (2010).
Davis, I. W., Murray, L. W., Richardson, J. S. & Richardson, D. C. MOLPROBITY: structure validation and all-atom contact analysis for nucleic acids and their complexes. Nucleic Acids Res. 32, W615–W619 (2004).
The PyMOL Molecular Graphics System. Schrödinger, LLC (Delano Scientific LLC, Palo Alto, CA, 2002).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996). 27-38.
Zimmermann, I., Marabelli, A., Bertozzi, C., Sivilotti, L. G. & Dutzler, R. Inhibition of the prokaryotic pentameric ligand-gated ion channel ELIC by divalent cations. PLoS Biol. 10, e1001429 (2012).
Wells, M. M. et al. Ensemble-based virtual screening for cannabinoid-like potentiators of the human glycine receptor alpha1 for the treatment of pain. J. Med. Chem. 58, 2958–2966 (2015).
Wennberg, C. L., van der Spoel, D. & Hub, J. S. Large influence of cholesterol on solute partitioning into lipid membranes. J. Am. Chem. Soc. 134, 5351–5361 (2012).
Wolf, M. G., Hoefling, M., Aponte-Santamaria, C., Grubmuller, H. & Groenhof, G. g_membed: efficient insertion of a membrane protein into an equilibrated lipid bilayer with minimal perturbation. J. Comput. Chem. 31, 2169–2174 (2010).
Abraham, M. J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2, 19–25 (2015).
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010).
Mayne, C. G., Saam, J., Schulten, K., Tajkhorshid, E. & Gumbart, J. C. Rapid parameterization of small molecules using the Force Field Toolkit. J. Comput. Chem. 34, 2757–2770 (2013).
Hess, B., Bekker, H. & Berendsen, H. J. C. Fraaije JGEM. LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald—an N.Log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Acknowledgements
The authors thank Prof. William Furey for his help and discussion in data collection and analysis, Mr. Devin Adell for his participation in the early electrophysiology data collection, and Ms. Sandy Hirsh for her suggestions on the manuscript. Use of the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, is supported by the U.S. Department of Energy, Office of Science, and Office of Basic Energy Sciences under Contract No. DE-AC02-76SF00515. The SSRL Structural Molecular Biology Program is supported by the DOE Office of Biological and Environmental Research, and by the National Institutes of Health, National Institute of General Medical Sciences (including P41GM103393). Use of the Advanced Photon Source was supported by the U. S. Department of Energy, Office of Science, Office of Basic Energy Sciences, under Contract No. W-31-109-Eng-38. The MD simulations were supported in part by the National Science Foundation through XSEDE resources (TG-MCB050030N). The research was supported by NIH funding (R01GM056257). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
P.T. and X.Y. designed the project. Q.C. conducted protein preparation and crystallization. Q.C., P.A., A.E.C., and P.T. contributed to x-ray data collection. Q.C. and P.A. processed diffraction data and refined the structures. Q.C., M.M.W., P.A., and P.T. analyzed structures. Q.C., T.S.T., and M.M.W. performed TEVC functional measurements and data analysis. M.M.W. and Q.C. performed molecular dynamics simulations and they, along with P.T., analyzed data. P.T., Q.C., and M.M.W. wrote the manuscript with input from other authors. All authors contributed and reviewed the results and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, Q., Wells, M.M., Arjunan, P. et al. Structural basis of neurosteroid anesthetic action on GABAA receptors. Nat Commun 9, 3972 (2018). https://doi.org/10.1038/s41467-018-06361-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-018-06361-4
- Springer Nature Limited
This article is cited by
-
Neurosteroids and their potential as a safer class of general anesthetics
Journal of Anesthesia (2024)
-
Cryo-EM structures reveal native GABAA receptor assemblies and pharmacology
Nature (2023)
-
The effects of a competitive antagonist on GABA-evoked currents in the presence of sedative-hypnotic agents
Pharmacological Reports (2020)
-
GABAA receptor signalling mechanisms revealed by structural pharmacology
Nature (2019)
-
Synthesis, sciatic nerve block activity evaluation and molecular docking of fluoro-substituted lidocaine analogs as local anesthetic agents
Medicinal Chemistry Research (2019)