Abstract
Carbapenem-resistant Klebsiella pneumoniae (CRKP) are of particular concern due to the spread of antibiotic resistance genes associated with mobile genetic elements. In this study, we collected 687 carbapenem-resistant strains recovered among clinical samples from 41 hospitals in nine Southern European countries (2016-2018). We identified 11 major clonal lineages, with most isolates belonging to the high-risk clones ST258/512, ST101, ST11, and ST307. blaKPC-like was the most prevalent carbapenemase-encoding gene (46%), with blaOXA-48 present in 39% of isolates. Through the combination and comparison of this EURECA collection with the previous EuSCAPE collection (2013-2014), we investigated the spread of high-risk clones circulating in Europe exhibiting regional differences. We particularly found blaKPC-like ST258/512 in Greece, Italy, and Spain, blaOXA-48 ST101 in Serbia and Romania, blaNDM ST11 in Greece, and blaOXA-48-like ST14 in Türkiye. Genomic surveillance across Europe thus provides crucial insights for local risk mapping and informs necessary adaptions for implementation of control strategies.
Similar content being viewed by others
Introduction
Gram-negative Klebsiella pneumoniae (Kp) are part of the microbiota in humans and are considered opportunistic pathogens able to cause hospital- and community-acquired infections such as pneumonia, bloodstream and urinary tract infections1. Clinically, these infections are treated with fluoroquinolones, aminoglycosides, cephalosporins, and, as a last resort, carbapenem antibiotics. However, Kp may become resistant to carbapenems mainly through the acquisition of resistance genes encoding carbapenemases, or the production of extended-spectrum beta-lactamases or cephalosporinases combined with porin alterations2. Carbapenemase genes are of particular concern as they can spread in association with mobile genetic elements (MGE) that are part of plasmids and transposons. These carbapenemase genes, mainly blaKPC, blaNDM, blaVIM, and blaOXA-48, are often associated with particular and successful nosocomial clones, sometimes with the close relationship between a lineage and the antibiotic resistance determinants3.
The increase of carbapenem-resistant Kp (CRKP) represents an unquestionable threat to the health of hospitalized patients globally, with a high mortality rate compared to patients infected with carbapenem-susceptible Kp4,5. Indeed, CRKP is a relevant public health problem with economic effects6, and it was recognized as a critical pathogen by the World Health Organization (WHO) for the need of new antimicrobials7. Different surveillance programs are collaboratively running at several hospitals in many countries. In Europe, national outbreaks of CRKP have been reported mainly in Southern European countries such as Greece, Spain, and Italy where also the highest prevalence is observed8,9. To limit the high incidence, increasing spread, and unraveling transmission pathways of CRKP, it is therefore relevant to study their characteristics, including distribution of lineages and resistance determinants at regional and local levels as well as internationally.
The COMBACTE consortium pursues the prevention and treatment of antibiotic-resistant-associated infections through four main projects. Among them, COMBACTE-CARE, seeks to support the development of new treatment options, together with the analysis of clinical and epidemiological datasets in all European member states and affiliated countries10,11. As part of this, the European prospective cohort study on Enterobacterales showing resistance to carbapenems (EURECA) aimed to understand how the patients across Europe are infected and currently treated for Enterobacterales-associated infections, but also which subgroups of patients responded well to different antibiotic treatments10. Local laboratories submitted carbapenem non-susceptible isolates to EURECA from May 2016 to November 2018 from cohorts of patients with bacterial infections in Southern European countries. Our task was to characterize and identify circulating clones of CRKP by analyzing 687 genomes. In addition, we contextualized the spread of CRKP for a broader population view by comparing our data to the previous EuSCAPE study8, which included the carbapenem-susceptible and non-susceptible isolates during 2013–2014 sample from a wider range of European countries.
Results
Presence of carbapenemase genes across different ST
In the EURECA collection, 683 isolates were classified by the phylogenetic analysis into Klebsiella pneumoniae sensu stricto, with two isolates each in Klebsiella variicola and Klebsiella quasipneumoniae. Of all the Kp sensu stricto, we identified 50 different sequence types (STs) and nine novel single locus variants (SLV) (Supplementary Data 1). Most isolates (n = 599, 87%) were grouped in one of 11 clonal lineages and their single-locus variants (Table 1), with ST258/512 being the dominant lineage (n=204; 30%), and ST512 the main single ST (n = 116, 17%) (Fig. 1). Other clonal lineages in order of prevalence were ST11 (n = 116, 17%), ST101 (n = 87, 15%), ST307 (n = 71, 10%), ST15 (n = 42, 6%), and ST147 (n = 40, 6%). Of the carbapenemase genes, blaKPC-like (n = 314, 46%) were the most abundant, mainly linked to ST258/512 (Fig. 1). blaOXA-48-like (n = 266, 39%) was the second most widespread gene found in several STs, for instance, ST15, ST101, ST11. blaNDM-1 was found in 96 (14%) isolates either on its own or in combination with blaOXA-48-like genes, and interestingly, it was present in half of the ST11 isolates. A combination of two carbapenemase genes was found in 38 isolates (5.5%), with the majority of isolates having a combination of blaOXA-48 + blaNDM-1 (n = 25, 66%) (Fig. 1, Table 1). Despite re-culturing under antibiotic selection, DNA extraction, and re-sequencing, we could not find a carbapenemase in 22 isolates. All of these were classed phenotypically as “non-KPC, non-metallo beta-lactamase”. As we were unable to recover a carbapenemase, we assume that the resistance mechanism is something other than an acquired carbapenemase or that any plasmid conferring resistance was lost before sending the strain for sequencing.
This phylogenetic tree contains 683 Klebsiella pneumoniae sensu stricto carbapenem resistant isolates collected as part of the EURECA study. The inner ring shows sequence type, and the different colors of the outer ring depict the carbapenemase genes. Main Sequence types with more than five isolates have been labeled.
Geographic distribution of major clonal lineages and carbapenemase genes
We compared EURECA isolates to the EuSCAPE collection (n = 1717) recovered during 2013–2014 with a similar sampling framework (Fig. 2d)8. Carbapenem-susceptible isolates were only collected as part of the EuSCAPE collection, grouping into a variety of clonal lineages, with 27.2% (n = 467) not belonging to any of the major clonal lineages, apart from Serbia where half of the isolates were ST101 (Fig. 2a). In contrast, carbapenem resistant isolates were grouped in eleven common clonal lineages in both collections with minor changes in the order of prevalence (87% EuSCAPE, 88% EURECA of each collection; Fig. 2b and c), except for the ST35 present only in EuSCAPE. The most prominent clonal lineage was ST258/512, then ST11 and ST101, with noticeable regional differences in abundance (Table 1, Supplementary Table 1). Italy was dominated by ST258/512 and associated with blaKPC-like genes (Fig. 2b and e). Whilst ST258/512 was present in Greece as well, the country had a higher diversity of clones (17 different STs, vs. 13 STs in Italy in EURECA) with ST11-blaNDM as prominent as ST258/512 (41 and 58 isolates, respectively; Fig. 2c, Table 1). In Eastern Europe, in Serbia and Romania, the abundant clone was ST101, although Serbia and Romania had a second dominant clone (ST11, and ST15 in EURECA, respectively; Fig. 2b and c). In the carbapenemases present, Serbia had a higher percentage of isolates harboring blaNDM gene in the EuSCAPE collection (44%, n = 24), whereas the later EURECA collection showed a dominance of blaOXA-48 (75%, n = 82). In contrast, although numbers are small in the EURECA collection, Romania had a higher percentage of blaNDM isolates (35%, n = 11) compared to EuSCAPE (6%, n = 4) (Fig. 2e and f).
a Proportion of the EuSCAPE carbapenem susceptible (CSKP) clonal lineages. b and c Clonal lineages of CRKP isolates of EuSCAPE and EURECA, respectively. d European countries, which participated in EuSCAPE (green), EURECA (blue), and the overlap of both surveys (purple). Map created with mapchart.net and included under CC BY-SA 4.0 licence. e and f Distribution of carbapenem resistance genes in EuSCAPE and EURECA, respectively.
Other countries, such as Spain and Türkiye displayed a more mixed picture of different clonal lineages with several clones at near-equal proportions, yet changed over time. Though ST11, ST15, and ST147 still feature prominently in Spain (18%, 17%, 10%, respectively), ST258 is now dominant (28%), and 16% are ST307 (Fig. 2c). Türkiye showed a number of different lineages in the EuSCAPE collection, however in EURECA, ST14 is dominant (49%, n = 24; Fig. 2c). Common to Serbia, Spain and Türkiye is the presence of the blaOXA-48 gene (Fig. 2e and f). As this gene is often carried on a promiscuous plasmid12, this might explain the wide dispersal in various lineages.
ST258/512 – the worldwide spread of a successful lineage
The dissemination of the ST258/512 has been characterized largely in previous studies. We, therefore, analyzed the EURECA collection (Fig. 3, highlighted in the first column in light blue; https://microreact.org/project/gUrL4jULY1JEPNExrFK4zH-globalcollectionclonallineage258512) together with 652 publicly available genomes around the world13,14,15,16, including the EuSCAPE collection8.
A phylogenetic tree of 856 isolates of ST258, ST512 and single locus variants with 204 isolates from EURECA, 236 isolates submitted to EuSCAPE, and 415 isolates with publicly available sequence data. Reference genome: 30660/NJST258 (CP006923). Tree tip colors correspond to country. Columns indicate: Project, (1) country, (2) ST, (3) and KPC-variant. (4) For further metadata and interactive exploration, follow this Microreact link: https://microreact.org/project/gUrL4jULY1JEPNExrFK4zH-globalcollectionclonallineage258512.
KPC-producing K. pneumoniae were first reported as outbreaks in the USA, initially focussed on the East Coast before spreading more widely14,15,17. From the oldest genomes reported of ST258, we can see that these originate from the USA (light green, Fig. 3), and that they also show considerable diversity, which has been previously described as “paraphyletic”14. Emergence of ST258 has been dated to the mid-1990s, before showing in clinical presentations and outbreaks at the beginning of the 2000s14. These ST258 isolates are mainly capsule-type KL107, carry the blaKPC-3 variant, and have been predominantly isolated from the USA (Fig. 3)14,15.
Clade 1 is the first major export of ST258 outside of the USA. This clade simultaneously changed both the capsule as well as the KPC variant, now KL106 and blaKPC-2 (Supplementary Data 2)14,16. Previous reports8,14 noted that isolates in this clade have a close association with Greece, with isolates either being isolated there or being associated with recent travel or healthcare exposure in Greece. This clone was first reported sporadically in Greece between 2006-200818. This introduction and expansion in Greece is detected in the EURECA collection, with isolates from Serbia and Montenegro nested within the diversity of the Greek isolates.
Before the second major clade, previously labeled as clade 2, there are a handful of US isolates which are both KL107 and blaKPC-2. Given the characteristics of the paraphyletic isolates, it would appear that clade 2 emerged directly out of this background population. Clade 2 of ST258 was introduced into Israel, with early reports from 2005, where also the emergence of the single locus variant ST512 was documented8,13,19. Following this evolution and widespread dissemination within Israel, the clone was then introduced successfully into Italy (Fig. 3), likely in a single event8. Further spread of ST512 then occurred to Spain, which was not detected within the EUSCAPE dataset; this may be due to sampling in a different city than before (Fig. 3).
Microevolution of other single locus variants through mutations in rpoB is visible within the ST512 clade: ST868 (Fig. 3, red), has arisen on two independent occasions due to the same mutation in rpoB, making this ST not monophyletic. A new monophyletic ST1519 (orange) appears to be particularly prevalent in Bologna, as most isolates originate from that city.
Allelic variant mutations of blaKPC gene have been described that confer resistance to ceftazidime/avibactam. In this collection, however, we found only one ST1519 isolate carrying blaKPC-36 (Fig. 3).
ST307–the emergence of a new threat
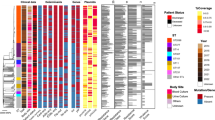
Similarly to ST258/512, ST307 (n = 68,10%) is an important globally distributed clone20,21, therefore we analyzed the phylogenetic relationship of 113 ST307 isolates and three single locus variants (SLV) from the EuSCAPE and EURECA collections. This clone was frequently found in Greece, Italy, and Spain (n = 81, 70%; Table 1, Fig. 4). We investigated the previously proposed cutoff of 21 SNPs8 across the different major clonal lineages, and found it to be comparable across all (Supplementary Fig. 1). We thus use this as a guiding value to identify putative within- or between-hospital clusters. In ST307, we found considerable diversity within this clone (Fig. 4), but found that isolates with less than 21 SNPs were indeed isolated from the same hospitals. This then proposed two small outbreaks in Spain within single hospitals (Fig. 4), one in which most isolates carried the blaOXA-48 gene (n = 10), and a second in which blaKPC-3 (n = 7; n = 1 combination with blaOXA-162) was found (Fig. 4 purple boxes). Further putative transmission events using this cut-off were detected in Greece and Italy (orange branches). In Italy, we also found a closely related clade (pink box) composed of both collections as well as different hospitals, indicating that this had been circulating for several years and is more widely disseminated. Isolates carried either blaKPC-2, blaKPC-3, or no carbapenemase, indicating fluidity with respect to high-level resistance. Overall, in this European collection, the most prevalent carbapenemase genes were blaKPC-like, particularly in Italy and Greece, which also corresponded to the most prevalent carbapenemase type in these countries (Figs. 2b and 4).
Combined analysis of 117 isolates from EuSCAPE and EURECA. Reference genome CPKp1825 (WMHT01). Columns present the information about: 1 – Project, 2 – single locus variants, 3 – Country, 4 – Different hospitals (shades for each country), 5 – Carbapenemase genes. Putative outbreaks are highlighted with orange branches, and clades of interest discussed in the text are shaded with boxes (pink for Italy and purple for Spain).
In a global context, including EuSCAPE and EURECA collections as well as globally available isolates, the phylogenetic analysis showed a high diversity, with multiple small clades (Supplementary Fig. 2, https://microreact.org/project/rFyHR2a3iN1FDh7kJKj8pf-clonallineagest307globalcollectionproject). Two larger clades from the USA and Zambia stand out, particularly as they do not carry carbapenemases. Apart from these, the smaller clades are often country-specific, for example for Italy, Spain, France, and Germany. In ST307, various carbapenemases were found, alone and in combinations. ST258/512 shows a similar global diversity in average SNP distances between isolates as ST307 (116 SNPs and 110 SNPs, respectively). In contrast to ST258/512, ST307 shows no single carbapenemase or sublineage that is predominantly globally distributed.
ST11 and ST101–the importance of knowing local conditions
A total of 330 isolates of ST11 and the single locus variants ST340 and ST437 from the EuSCAPE and EURECA collections were analyzed. The phylogenetic tree formed three clades (Fig. 5a): clade 1 containing the isolates belonging exclusively to ST437, clade 2 with ST11 isolates plus a nested ST340 clade, and clade 3 with ST11 isolates only. The distribution of carbapenemase genes differed between the countries, in clade 3 in Greece ST11 was associated with the blaNDM-1 gene, in contrast, in Spain blaOXA-48 as well as blaOXA48-like genes were present, whereas Serbian isolates in clades 1 and 2 carried either blaNDM-1 or blaOXA-48 (Fig. 5a). Overall, the isolates clustered largely by country and no transmission events were observed between countries based on the 21 SNP cutoff. For an in-depth study, we reconstructed a phylogenetic tree only with the Serbian isolates, due to most of them belonging to two different STs to the founder ST11 (Fig. 5b). Interestingly, the ST340 clade contained predominantly carbapenem-non-susceptible EuSCAPE isolates without carbapenemase genes but with extended-spectrum beta-lactamase blaCTX-M-15. In this clade, both highly clonal isolates within the same hospital (0-21 SNPs), and signs of a nationally spreading clone (22–100 SNPs for isolates collected from different hospitals) are visible. Isolates within the ST437 clade frequently carried either blaNDM-1 or blaOXA-48. Two of the subclades belonged exclusively to the EURECA collection and might represent a better approximation of the current circulating clones of carbapenem-non-susceptible ST437 in Serbia (Fig. 5b).
a Combined analysis of 329 isolates from EuSCAPE and EURECA. Reference genome F64 (VILG01). Inner ring: project, (1) middle ring: geographic origin, (2) outer ring: carbapememase genes. (3) b Phylogenetic relationship between isolates ST11 from Serbia only. Columns display the information about project, (1) different colors representing different hospitals in Serbia, (4) SNP distance, (5) and carbapenemase genes (3).
In a global contextualization of ST11 and variants using 1648 genomes publicly available in Pathogenwatch from different countries, the phylogenetic tree was clustered into several clades, with carbapenemase-producing EuSCAPE and EURECA ST11 isolates grouped in one particular clade (Supplementary Fig. 3b). Within this clade, there appears to be a particular subclade with blaOXA-48 spreading in Spain, whereas the other European isolates carried blaNDM-1. For isolates of ST437, the Serbian EURECA isolates clustered closely with isolates from Croatia and Slovenia, potentially signifying an Eastern European subclade (Supplementary Fig. 3c).
In ST101 (n = 87) (Supplementary Fig. 4a), EURECA isolates mainly came from Serbia (n = 56, 64%) and Romania (n = 17, 19%) and formed separate clades. We found that some of the Romanian isolates appeared to be more diverse and were interspersed with isolates from Italy, Spain, and Türkiye, carrying different carbapenemase genes, however there was also a particular Romanian-only clade carrying blaOXA-48 (Supplementary Fig. 4a, blue clade). In contrast, isolates from Serbia formed distinct clades, so we investigated them in more detail. To define the ancestral relationships of the ST101 in Serbia, we analyzed 135 genomes: 57 from EURECA collection, 42 from the EuSCAPE survey8, and 36 genomes from Palmieri et al.22. The latter study recovered colistin resistance CRKP of ST101 between 2013 and 2017 and confirmed mgrB mutations as the major cause of colistin resistance in these isolates. They also reported blaOXA-48 as the carbapenemase gene endemic in Serbia, which agrees with our results. The phylogenetic tree showed several clades, with the EuSCAPE isolates being the oldest and forming the most basal clade (Supplementary Fig. 4b). These isolates were also largely carbapenem-susceptible and did not harbor a carbapenemase (with one exception) (Supplementary Fig. 1b). There are further EuSCAPE isolates with either blaOXA-48, blaNDM-1 or a combination of both, however, the EURECA and Palmieri collections are intermingling in a separate, highly related clade, which was characterized by the presence of blaOXA-48 (Supplementary Fig. 4b, shaded box). We found less than 21 SNPs between isolates of these two collections, and isolates had been obtained from a number of different hospitals, which might be indicative of a locally circulating clone. However, we cannot confirm this since we have no further information about the isolates of either collection. Importantly, we can rule out that these isolates contributed twice to the different collections because the sampling time frames did not overlap.
Other clonal lineages
Regarding other clonal lineages, one to highlight is the ST147 (n = 40), which was mainly recovered from Spain (n = 20, 50%) and Greece (n = 15, 37%) in the EURECA collection. An outbreak in Tuscany, Italy, in 201823, and therefore we contextualized our isolates plus the EuSCAPE isolates (Supplementary Fig. 5). EuSCAPE and EURECA isolates were mostly diverse, with only two small clusters in Greece and Spain (EURECA collection) indicative of local clones (shaded boxes), whereas the Tuscany outbreak formed a separate, highly clonal expansion. Interestingly, we found six different carbapenemase genes within this ST, and their combination of two of these carbapenemase genes, even in the local lineages in Spain (isolates with blaOXA-48 with and without blaNDM) and Greece (isolates with blaKPC-2 with and without blaVIM), only the Tuscany outbreak was very homogenous in carrying the blaNDM-1 gene.
The ST15 was also diverse, recovered from a wide range of countries, and with different carbapenemase combinations (Supplementary Fig. 6). In Spain, we identified three clades (purple boxes); in two clades isolates carried blaOXA-48 and the other, highly clonal clade with 14 isolates from the same hospital exhibited blaOXA245 exclusively (EURECA collection).
Lastly, the ST14 from the EURECA collection highlighted a local expansion with ST14 and the single variant ST2096 in Türkiye. Most of the isolates were from the same hospital except two, carrying the blaOXA-232 and blaOXA-48 gene on its own or in combination with blaNDM-1 gene (Supplementary Fig. 7). Contextual analysis within a global collection showed that these isolates, especially ST2096, are part of a particular clade containing isolates from Türkiye, Saudi Arabia and a number of other countries (Supplementary Fig. 7; https://microreact.org/project/3mPo67JrFnL6Xt3cVSVrR1-st142096lineageglobalcollectionproject).
Discussion
Due to their rapid and efficient spread, CRKP is considered a public health problem worldwide. In order to monitor the clonal evolution and geographical expansion over time, new tools such as WGS have been implemented. The characterization of carbapenem-resistant isolates is essential for infection control purposes because of their impact on therapeutic decisions in clinics, hence laboratory detection providing high accuracy and fast diagnosis is key for accurate antibiotic delivery24. The entire collection of 683 isolates in this present study (COMBACTE-CARE EURECA) exhibited phenotypic resistance to at least one carbapenem commonly used to treat infections caused by multiple drug-resistant pathogens. The collection was split into 11 clonal lineages, the most prominent being ST258/512, ST11, ST101, and ST307, as the major high-risk clones distributed worldwide (Fig. 2a, b). The CRKp isolates in these 11 clonal lineages corresponded to 87% and 88% of the EuSCAPE and EURECA collections, respectively. In the EURECA collection, 27 singleton STs were found, with 150 found in EuSCAPE, as this collection encompasses the more diverse carbapenem susceptible population. As the EuSCAPE collection temporally preceded EURECA by two years, we contextualized and compared both collections to better illustrate the clonal distribution of CRKP in Europe. This showed that CSKP, only collected during EuSCAPE, is very diverse and contains many different STs, however, CRKP are grouped into the well-known high-risk clones circulating around the world20 (Supplementary Table 1, Fig. 2). Some changes were observed in the prevalence in different countries: ST307 carrying 4 different carbapenemase genes increased in Italy, Romania and Spain, and a particular ST14 lineage was prominent in Türkiye. Differences were also observed with respect to carbapenemase genes: In Romania and Türkiye blaNDM-1 was more prominent, in Spain more blaKPC-like genes were detected, and in Serbia blaOXA-48 were found in a considerable proportion (Supplementary Table 1, Fig. 2).
Since the first report in 2001 of the clonal lineage ST258/512 in the USA associated with blaKPC, its international spread has been rapid and extensive8,14,15. Using isolates belonging to ST258/512 of our current collection along with previously published ones (Fig. 3), we confirmed the continental spill-over of ST258/512 into Europe from the USA, which began with several introductions into Greece and Italy, the expansion into Serbia and Montenegro, Romania and finally into Spain. EURECA helped to highlight more clearly the second introduction of this clone into Spain and Serbia (Fig. 3). We confirmed that this lineage linked to blaKPC-like is still prevalent in Italy, Greece, and Spain (Fig. 2)25,26. One isolate of the single locus variant ST1519 harbored blaKPC-36 and was recovered from the blood of a hospitalized patient in Bologna, Italy, with a previous study reporting this novel blaKPC-3 variant in the same city27. This new mutation confers resistance to ceftazidime/avibactam use for the treatment of CRKP28.
ST258 was not the most abundant clone in other European countries such as Romania, Serbia, and Türkiye in the collections we compared (Fig. 2); this could be due to the rise of other high-risk clones like ST307, ST101, and ST11.
The emerging high-risk clone ST307 was first described in the mid-1990s, and it has since been reported worldwide with different carbapenemase resistance determinants29,30,31. This clone has also been responsible for nosocomial outbreaks in European countries such as Portugal, Spain, France, the UK, Germany, Netherlands, Slovenia, and Romania32, and has been considered as a replacement for other high-risk clones, such as ST258 in countries like Colombia33, South Africa34, and Italy35. Furthermore, it was also responsible for nosocomial outbreaks harboring blaKPC variants conferring ceftazidime-avibactam resistance24 and, more recently, cefiderocol resistance36. The presence of several chromosomal and plasmid-encoded factors associated with hypervirulence make ST307 a superior clone that can easily share large plasmids with other Kp and Enterobacterales species, such as the Enterobacter cloacae complex31. Indeed, the presence of a distinct capsule structure (rare in the Kp population) may contribute to the successful propagation of this lineage37. In the EuSCAPE and EURECA collections, we observed a regional circulation of this clone in Greece, Italy, and Spain (Fig. 4). In our two observed collections, the carbapenemase genes found in this clone often appeared to reflect the dominant carbapenemase in that country, namely blaKPC-2, and blaKPC-3 in Greece and Italy, blaOXA-48 for the Spanish isolates. This may be a hint that the adaptability of ST307 with respect to carbapenemase uptake could be influenced by geographic prevalence, although interestingly, the Romanian ST307 isolates carried blaNDM-1 (Fig. 4).
Large surveillance and sequencing efforts such as the current study not only improve our understanding of global clones but may also highlight regional circumstances that need to be taken into account in local infection control efforts. In Serbia, two different single locus variants of ST11 circulate: ST340, largely non-susceptible to carbapenems but without carbapenemases from the EuSCAPE collection, and ST437, mainly from the later EURECA collection, whose isolates often harbor either blaOXA-48 or blaNDM-1 (Fig. 5b). The founder ST11 is of minor importance as we found few isolates belonging to this ST, and putatively only signifies a single small introduction event from Greece (Fig. 5a). Comparing all non-susceptible isolates in EuSCAPE and EURECA from Serbia, the percentage of ST101 isolates has stayed the same, however the ST437 lineage has doubled in proportion. Since EURECA sampled fewer but completely overlapping hospitals with EuSCAPE, this may be an emerging trend amongst Serbian isolates. Also, in Serbia, a particular ST101 clone appears to have emerged, that is characterized by the presence of blaOXA-48 (Supplementary Fig. 1a, b). From the current collections, we are not able to say whether the susceptible clones are still circulating in Serbia, or whether ST101 isolates would now exclusively be part of the non-susceptible clade. Similarly, in Spain, several different lineages of ST15 were co-circulating; two lineages were isolates carried blaOXA-48 and one clade with blaOXA-245 (Supplementary Fig. 3).
Regional differences exist in terms of the local dominant CRKP clones. In Greece, Italy, and Spain, ST258/512 was the dominant clone, while, in Serbia and Romania, ST101 was predominant, and ST14 was potentially expanding in Türkiye. The respective carbapenemase genes within these STs are likewise diverse. Overall, blaKPC-like was the most prevalent carbapenemase gene (46%) associated with the most abundant ST258/512. The second most frequent carbapenemase gene was blaOXA-48 (39%), widely spread between different STs. Moreover, a relatively high proportion of isolates (5.5%) harbored two carbapenemases.
The combination of both EURECA and EuSCAPE collections helped to elucidate the high-risk clones circulating in Europe and the evolution of Kp. As both surveys had similar collection strategies, we were able to compare temporal changes from 2012–14 to 2016–18 through a comparison of dominant lineages, their associated carbapenem resistance genes, and the potential acquisitions of locally circulating genes. Collections such as these demonstrate that continuous surveys are necessary, and the sampling should always include susceptible background populations as well as highly resistant isolates in order to discern introduction from de-novo emergence.
These surveys also provide a blueprint for important European initiatives towards better planning for public health and infection control interventions.
Methods
Isolate collection and antimicrobial susceptibility
Isolates were recovered from patients diagnosed with bloodstream, intra-abdominal, pneumonia, and complicated urinary tract infection in 41 hospitals in nine countries (Albania, Croatia, Greece, Italy, Montenegro, Romania, Serbia, Spain, and Türkiye). The collection period was from May 2016 to November 2018. Overall, 49% of the samples were from blood (n = 334), 31% from urine (n = 213) and 20% from other sources (n = 140) (Supplementary Data 1). The countries with the major sample contributions to this study were Spain (n = 202), Greece (n=174), Italy (n=111), and Serbia (n=110). Enterobacterales isolates with MIC ≥1 mg/L (dilution methods) or ≤22 mm (disc diffusion, 10 μg disks) for meropenem or imipenem isolated from patients with the above infections were considered as putative CRE and studied; those producing carbapenemases and/or showing resistance to imipenem or meropenem according to EUCAST breakpoints were included. Collected isolates were phenotypically confirmed by disc diffusion at the EURECA central laboratory at the University of Antwerp. Further, the MIC values were determined by broth microdilution at SERMAS laboratory in the Hospital Ramon y Cajal in Madrid. The isolates were identified to the species level with MALDI-ToF, and then confirmed by whole genome sequencing.
Whole genome sequencing and quality control
DNA extraction was performed using the Roche High Pure Template Preparation Kit. DNA concentration was measured with the Qubit fluorometer, followed by the sequencing library preparation using Nextera™ DNA Flex Library Prep (flow cell for 2x150bp paired-end sequencing).
Raw sequence reads were mapped to the reference genome MGH78578 (CP000647) using Smalt38 and Samtools39, with subsequent filtering with Genome Analysis Toolkit40. The minimum accepted average coverage was 30X per sample. The reads were de novo assembled using SPAdes v3.13.141 with Kmer sizes 21, 33, 55, 77, 99, 109, and 123. The expected size for the assemblies was between 5–7 Mb, 50–100 contigs, and N50 >100,000 bp. Kraken with minikraken database was used to check the species and potential contamination of samples42. Sequence types (ST) were determined using the multilocus sequences typing (MLST) software (https://github.com/tseemann/mlst). The average coverage was 50x with a genome size of 5.6 Mb and N50 of 259,285 bp. The median number of contigs was 86 (Supplementary Data 1). Sequencing data has been submitted to the ENA project PRJEB63349, with individual accession identifiers for reads and assemblies given in Supplementary Data 1.
Acquired antimicrobial resistance genes were identified using Abricate v0.9.8 (https://github.com/tseemann/abricate), with a local database based on ResFinder43. Capsular (K) serotypes were predicted by Kaptive, as implemented in the Kleborate genotyping pipeline (https://github.com/katholt/Kleborate)44. For variations of the blaKPC gene, we extracted the nucleotide sequence of all KPC-positive isolates. We then compared variations within the gene with known mutations associated with ceftazidime/avibactam resistance (GenBank accession number MH593787).
Phylogenetic analyzes
Phylogenetic trees of the whole collection and subsets of particular sequence types were estimated using RAxML v8.2.1245 based on SNP alignments after mapping to the reference genomes and removal of recombinant regions using Gubbins v2.4.146. Reference genomes were chosen based on ST of interest with long-read sequencing and genome coverage >80X. Reference genomes for species overview: MGH78578 (GenBank accession number CP000647), ST11: F64 (VILG01), ST14: KPN528 (CP020856), ST15: P35 (CP053041), ST101: Kp_Goe_33208 (CP018447), ST147: HKP0064 (JACTAR01), ST258: 30660 (CP006923), ST307: CPKp1825 (WMHT01). Phylogenetic tree visualization was done with iToL47. Public datasets of interest (EuSCAPE8, Italian collection of ST14723, Serbian collection of ST10122, and worldwide collection of ST25813,14,15,16 were downloaded from the European Nucleotide Archive (ENA) and included in the analysis.
Pairwise minimum SNP differences analysis
We used a PHP script based on the converted matrix from a DNA sequence alignment with snp-dist (https://github.com/tseemann/snp-dists) to determine the minimum SNP difference between pairs of isolates of the same ST. Based on their geographical origin, isolate pairs were analyzed in three different settings: different countries, the same country but different hospitals, and the same hospital. Plots were generated in R (v4.3.0)48, using the ggplot2 (v3.4.4)49 package.
Global contextualization of lineages
We used the Pathogenwatch (https://pathogen.watch/) public genomes for a global contextualization of lineages. From the 48000 genome database associated with K. pneumoniae, we filtered by ST and created collections of up to 2000 genomes, corresponding to the maximum allowed.
Regardless of the geographical location, we excluded the samples that were highly clonal to include more diverse isolates, and we formatted the metadata in the same way we did for the EURECA collection. Microreact (https://microreact.org/) was used for interactive visualization.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
References
Hansen, D. S., Gottschau, A. & Kolmos, H. J. Epidemiology of Klebsiella bacteraemia: a case control study using Escherichia coli bacteraemia as control. J. Hosp. Infect. 38, 119–132 (1998).
Bush, K. Past and present perspectives on β-Lactamases. Antimicrob Agents Chemother. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6153792/ (2018).
Munoz-Price, L. S. et al. Clinical epidemiology of the global expansion of Klebsiella pneumoniae carbapenemases. Lancet Infect. Dis. 13, 785–796 (2013).
Borer, A. et al. Attributable mortality rate for carbapenem-resistant Klebsiella pneumoniae bacteremia. Infect. Control Hosp. Epidemiol. 30, 972–976 (2009).
Wu, C., Zheng, L. & Yao, J. Analysis of risk factors and mortality of patients with carbapenem-resistant Klebsiella pneumoniae infection. Infect. Drug Resist. 15, 2383–2391 (2022).
Cassini, A. et al. Attributable deaths and disability-adjusted life-years caused by infections with antibiotic-resistant bacteria in the EU and the European Economic Area in 2015: a population-level modelling analysis. Lancet Infect. Dis. 19, 56–66 (2019).
Tacconelli, E. Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development. https://policycommons.net/artifacts/1818147/global-priority-list-of-antibiotic-resistant-bacteria-to-guide-research-discovery-and-development/ (South Africa: Infection Control Africa Network; 2017).
David, S. et al. Epidemic of carbapenem-resistant Klebsiella pneumoniae in Europe is driven by nosocomial spread. Nat. Microbiol. 4, 1919–1929 (2019).
Lo Fo Wong, D., Diaz-Högberg, L. & Dominique L. M. European Centre for Disease Prevention and Control. Antimicrobial resistance surveillance in Europe 2022–2020 data. https://www.ecdc.europa.eu/en/publications-data/antimicrobial-resistance-surveillance-europe-2022-2020-data
Gutiérrez-Gutiérrez, B. et al. EUropean prospective cohort study on Enterobacteriaceae showing REsistance to CArbapenems (EURECA): a protocol of a European multicentre observational study. BMJ Open. 7, e015365 (2017).
Kostyanev, T. et al. The Innovative Medicines Initiative’s New Drugs for Bad Bugs programme: European public–private partnerships for the development of new strategies to tackle antibiotic resistance. J. Antimicrob. Chemother. 71, 290–295 (2016).
David, S. et al. Integrated chromosomal and plasmid sequence analyses reveal diverse modes of carbapenemase gene spread among Klebsiella pneumoniae. Proc. Natl Acad. Sci. Usa. 117, 25043–25054 (2020).
Adler, A. et al. Evolution and dissemination of the Klebsiella pneumoniae clonal group 258 throughout Israeli post-acute care hospitals, 2008–13. J. Antimicrob. Chemother. 72, 2219–2224 (2017).
Bowers, J. R. et al. Genomic Analysis of the Emergence and Rapid Global Dissemination of the Clonal Group 258 Klebsiella pneumoniae Pandemic. Bengoechea J. A., editor. PLOS ONE 10, e0133727 (2015).
Cerqueira, G. C. et al. Multi-institute analysis of carbapenem resistance reveals remarkable diversity, unexplained mechanisms, and limited clonal outbreaks. Proc. Natl Acad. Sci. 114, 1135–1140 (2017).
DeLeo, F. R. et al. Molecular dissection of the evolution of carbapenem-resistant multilocus sequence type 258 Klebsiella pneumoniae. Proc. Natl Acad. Sci. 111, 4988–4993 (2014).
Cuzon, G. et al. Worldwide Diversity of Klebsiella pneumoniae That Produce β-Lactamase blaKPC-2 Gene. Emerg. Infect. Dis. 16, 1349–1356 (2010).
Afolayan, A. O. et al. Three Klebsiella pneumoniae lineages causing bloodstream infections variably dominated within a Greek hospital over a 15 year period. Micro. Genomics. 9, mgen001082 (2023).
Warburg, G. et al. A carbapenem-resistant Klebsiella pneumoniae epidemic clone in Jerusalem: sequence type 512 carrying a plasmid encoding aac(6′)-Ib. J. Antimicrob. Chemother. 67, 898–901 (2012).
Wyres, K. L. et al. Distinct evolutionary dynamics of horizontal gene transfer in drug resistant and virulent clones of Klebsiella pneumoniae. PLOS Genet. 15, e1008114 (2019).
Villa, L. et al. Diversity, virulence, and antimicrobial resistance of the KPC-producing Klebsiella pneumoniae ST307 clone. Microb Genomics. https://www.microbiologyresearch.org/content/journal/mgen/10.1099/mgen.0.000110 (2021).
Palmieri, M. et al. Genomic Epidemiology of Carbapenem- and Colistin-Resistant Klebsiella pneumoniae Isolates From Serbia: Predominance of ST101 Strains Carrying a Novel OXA-48 Plasmid. Front Microbiol. 11, 294 (2020).
Martin, M. J. et al. Anatomy of an extensively drug-resistant Klebsiella pneumoniae outbreak in Tuscany, Italy. Proc. Natl Acad. Sci. Usa. 118, e2110227118 (2021).
Hernández-García, M. et al. Impact of ceftazidime-avibactam treatment in the emergence of novel KPC variants in the ST307-Klebsiella pneumoniae high-risk clone and consequences for their routine detection. J. Clin. Microbiol. 60, e0224521 (2022).
Cano-Martín, E. et al. A study in a regional hospital of a mid-sized Spanish city indicates a major increase in infection/colonization by carbapenem-resistant bacteria, coinciding with the COVID-19 pandemic. Antibiotics 10, 1127 (2021).
Chatzidimitriou, M. et al. Carbapenem-resistant Klebsiella pneumoniae in the Balkans: clonal distribution and associated resistance determinants. Acta Microbiol Immunol. Hung. 71, 10–24 (2024).
Gaibani, P., Ambretti, S., Campoli, C., Viale, P. & Re, M. C. Genomic characterization of a Klebsiella pneumoniae ST1519 resistant to ceftazidime/avibactam carrying a novel KPC variant (KPC-36). Int J. Antimicrob. Agents 55, 105816 (2020).
Haidar, G. et al. Mutations in blaKPC-3 That confer ceftazidime-avibactam resistance encode novel KPC-3 variants that function as extended-spectrum β-Lactamases. antimicrob agents chemother. 61: https://doi.org/10.1128/aac.02534-16 (2017).
Castanheira, M. et al. Rapid expansion of KPC-2-producing Klebsiella pneumoniae Isolates in two texas hospitals due to clonal spread of ST258 and ST307 lineages. Micro. Drug Resist. 19, 295–297 (2013).
Habeeb, M. A., Haque, A., Nematzadeh, S., Iversen, A. & Giske C. G. High prevalence of 16 S rRNA methylase RmtB among CTX-M extended-spectrum β-lactamase-producing Klebsiella pneumoniae from Islamabad, Pakistan. Int J Antimicrob Agent https://pubmed.ncbi.nlm.nih.gov/23622882/ (2022).
Heiden, S. E. et al. A Klebsiella pneumoniae ST307 outbreak clone from Germany demonstrates features of extensive drug resistance, hypermucoviscosity, and enhanced iron acquisition. Genome Med. 12, 113 (2020).
Peirano, G., Chen, L., Kreiswirth, B. N. & Pitout J. D. D. Emerging Antimicrobial-Resistant High-Risk Klebsiella pneumoniae Clones ST307 and ST147. Antimicrob Agents Chemother. https://doi.org/10.1128/AAC.01148-20 (2021).
Ocampo, A. M. et al. A two-year surveillance in five Colombian tertiary care hospitals reveals high frequency of non-CG258 clones of carbapenem-resistant klebsiella pneumoniae with distinct clinical characteristics. Antimicrob. Agents Chemother. 60, 332–342 (2015).
Lowe, M. et al. Klebsiella pneumoniae ST307 with blaOXA-181, South Africa, 2014–2016. Emerg. Infect. Dis. 25, 739 (2019).
Bonura, C. et al. An update of the evolving epidemic of bla KPC carrying Klebsiella pneumoniae in Sicily, Italy, 2014: emergence of multiple Non-ST258 clones. PLoS One. 15, e0132936 (2015).
Castillo-Polo, J. A. et al. Outbreak by KPC-62-producing ST307 Klebsiella pneumoniae isolates resistant to ceftazidime/avibactam and cefiderocol in a university hospital in Madrid, Spain. J. Antimicrob. Chemother. 78, 1259–1264 (2023).
Wyres, K. L., Lam, M. M. C. & Holt, K. E. Population genomics of Klebsiella pneumoniae. Nat. Rev. Microbiol. 18, 344–359 (2020).
Ponstigl, H. SMALT aligns DNA sequencing reads with a reference genome. https://www.sanger.ac.uk/tool/smalt/ (2015).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008 (2021).
McKenna, A. et al. The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Bankevich, A. et al. SPAdes: a new genome assembly algorithm and its applications to single-cellsSequencing. J. Comput Biol. 19, 455–477 (2012).
Wood, D. E. & Salzberg, S. L. Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 15, R46 (2014).
Zankari, E. et al. Identification of acquired antimicrobial resistance genes. J. Antimicrob. Chemother. 67, 2640–2644 (2012).
Lam, M. M. C. et al. A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex. Nat. Commun. 12, 4188 (2021).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Croucher, N. J. et al. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 43, e15–e15 (2015).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v5: an online tool for phylogenetic tree display and annotation. Nucleic Acids Res. 49, W293–W296 (2021).
RStudio Team. RStudio: integrated development for R. RStudio, PBC, Boston, MA. https://www.rstudio.com/ (2020).
Wickham H. Elegant graphics for data analysis:ork. ISBN 978-3-319-24277-4, https://ggplot2.tidyverse.org (2016).
Acknowledgements
The authors wish to thank the microbiologists from all participating laboratories in the EURECA study, and the members of the EURECA/WP1B group for their dedicated work and substantial contribution during the isolate collection phase of this study. The EURECA/WP1B group members are: Silva Tafaj (University Hospital of Lung Diseases ‘Shefqet Ndroqi’, Tirana, Albania), Arjana Tambic Andrasevic (University Hospital for Infectious Diseases, Zagreb, Croatia), Sophia Vourli (National and Kapodistrian University of Athens, Athens, Greece), Efthymia Protonotariou (General University Hospital of Thessaloniki ‘Ahepa’, Thessaloniki, Greece), Eleni Vagdatli (Hippokration General Hospital, Thessaloniki, Greece), Ioanna Voulgaridi (General Hospital of Larissa, Larissa, Greece), Iris Spiliopoulou (University of Patras, Patras, Greece), Ioannis Deliolanis (Laiko General Hospital, Athens, Greece), Maria Panopoulou (University Hospital of Alexandroupolis, Alexandroupoulis, Greece), Ergina Malli and Efthymia Petinaki (University Hospital of Larissa, Larissa, Greece), Efstathia Perivolioti (Evangelismos General Hospital of Athens, Athens, Greece), Theodora Biniari (“Agioi Anargyroi” General Oncology Hospital, Athens, Greece), Nicholas Legakis (Metropolitan General Hospital, Athens, Greece), Levantia Zachariadou (Aghia Sophia Children’s Hospital, Athens, Greece), Anastassios Doudoulakakis (Panagiotis & Aglaia Kyriakou Children’s Hospital, Athens, Greece), Vassiliki Pitiriga (Henry Dunant Hospital, Athens, Greece), Antonino Di Caro (National Institute for Infectious Diseases Lazzaro Spallanzani, Rome, Italy), Sara Giordana Rimoldi (Hospital Luigi Sacco, Milan, Italy), GianMaria Rossolini (Florence University Hospital, Florence, Italy), Maria Paola Landini (Policlinico Sant’Orsola Malpighi, Bologna, Italy), Teresa Spanu (Policlinico Universitario A. Gemelli, Rome, Italy), Milena Arghittù (IRCCS Fondazione Ca Granda Ospedale Maggiore Policlinico, Milan, Italy), Rossana Cavallo (Molinette Teaching Hospital, Torino, Italy), Anna Marchese (University of Catania, Azienda Ospedaliero Universitaria “Policlinico-Vittorio Emanuele”, Catania, Italy), Susanna Cuccurullo (University of Naples S.U.N./Monaldi Hospital, Naples, Italy), Paola Bernaschi (Ospedale Pediatrico Bambino Gesù, Rome), Roberto Bandettini (IRCCS Istituto Giannina Gaslini, Genova, Italy), Arsim Kurti (National Institute of Public Health of Kosova, Pristhina, Kosovo), Milena Lopicic (Clinical Center of Montenegro, Podgorica, Montenegro), Olivia Dorneanu (Clinical Hospital of Infectious Diseases of Iasi, Iasi, Romania), Mirela Flonta (Cluj Napoca Infectious disease Clinical Hospital, Cluj Napoca, Romania), Alma Kosa (“Dr. Victor Babes” Clinical Hospital of Infectious and Tropical Diseases, Bucharest, Romania), Daniela Talapan (The National Institute of Infectious Diseases Matei Bals, Bucharest, Romania), Mariana Buzea (Elias University emergency hospital, Bucharest, Romania), Camelia Ghita (Fundeni Clinical hospital Romania, Bucharest, Romania), Edit Székely (Spitalul Clinic Judetean de Urgenta Tg. Mures, Targu Mures, Romania), Snezana Jovanovic (Clinical Center Serbia, Belgrade, Serbia), Teodora Vitorovic (Clinical center of „Dragisa Misovic“, Belgrade, Serbia), Deana Medic (Institute for Public Health Vojvodina, Novi Sad, Serbia), Branislava Kocic (Institute for Public Health, Nis, Serbia), Ana Perucica (Clinical Center “Zvezdara”, Belgrade, Serbia), Begoña Palop-Borrás (Hospital Universitario Carlos Haya Málaga, Spain), Irene Gracia Ahufinger (Hospital Universitario Reina Sofia, Cordoba, Spain), Fe Tubau (Microbiology Department, Hospital Universitari de Bellvitge, Institut Investigacions Biomèdiques de Bellvitge (IDBELL), Barcelona, Spain and CIBER de Enfermedades Respiratorias (CIBERes), Institute de Salud Carlos III, Madrid, Spain), Fernando Chaves (Hospital Universitario 12 de Octubre, Madrid, Spain), Julio Garcia Rodriguez (Hospital Universitario La Paz, Madrid, Spain), Rafael Cantón (Servicio de Microbiología, Hospital Universitario Ramón y Cajal and Instituto Ramón y Caja de Investigación Sanitaria (IRYCIS), Madrid, Spain and CIBER de Enfermedades Infecciosas (CIBERINFEC), Institute de Salud Carlos III, Madrid, Spain), Francisca Portero (Hospital Universitario Puerta de Hierro Majadahonda, Madrid, Spain), Patricia Muñoz (Hospital Universitario Gregorio Marañón (SERMAS-HGM), Madrid, Spain), Cüneyt Özakin (Uludag University Turkey, Uludag, Türkiye), Banu Sancak (Hacettepe University School of Medicine, Ankara, Türkiye), Can Bicmen (Dr. Suat Seren Chest Diseases and Surgery Training Hospital, Izmir, Türkiye), Ufuk Hasdemir and Guner Soyletir (Marmara University School of Medicine, Istanbul, Turkey), and Zeynep Ceren Karahan (Ankara University Faculty of Medicine Department of Medical Microbiology). We thank Valentina Valenzuela Dallos, Sarah Klassen and Leonardo Duarte dos Santos for excellent technical support at the Medical Center – University of Freiburg, Germany. This research was funded by: Innovative Medicines Initiative Joint Undertaking (https://www.imi.europa.eu/) under Grant Agreement Nos. 115523 [COMBACTE-NET (Combatting Bacterial Resistance in Europe)] and 115620 (COMBACTE-CARE) to TK, JBF-A, MG-C, RC, HGo, JR-B, and HGr; Plan Nacional de I+D+I, Instituto de Salud Carlos III, Subdirección General de Redes y Centros de Investigación Cooperativa, Ministerio de Ciencia, Innovación y Universidades, Spanish Network for Research in Infectious Diseases (REIPI RD16/0016/0001 and RD16/0016/0011) and CIBERINFEC (21/13/00012, 21/13/00084) to JR-B and RC; JPI-AMR call (K-STaR 01KI1910) to HGr and SR; Federal Ministry of Education and Research (BMBF; TAPIR 01KI2018) to SR.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
M.B.S., S.A.M. and S.R. performed the bioinformatics analysis. S.R. supervised the bioinformatics work. T.K. and H.Go. were responsible for sample handling and phenotypic confirmation. J.B.F.-A., M.G.-C., R.C., J.R.-B. performed the MIC testing. R.C., H.Go., J.R.-B. and H.Gr. conceived the project and acquired funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Geetha Nagaraj and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Budia-Silva, M., Kostyanev, T., Ayala-Montaño, S. et al. International and regional spread of carbapenem-resistant Klebsiella pneumoniae in Europe. Nat Commun 15, 5092 (2024). https://doi.org/10.1038/s41467-024-49349-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-49349-z
- Springer Nature Limited