Abstract
The metabotropic glutamate receptors (mGluRs) are neuromodulatory family C G protein coupled receptors which assemble as dimers and allosterically couple extracellular ligand binding domains (LBDs) to transmembrane domains (TMDs) to drive intracellular signaling. Pharmacologically, mGluRs can be targeted at the LBDs by glutamate and synthetic orthosteric compounds or at the TMDs by allosteric modulators. Despite the potential of allosteric compounds as therapeutics, an understanding of the functional and structural basis of their effects is limited. Here we use multiple approaches to dissect the functional and structural effects of orthosteric versus allosteric ligands. We find, using electrophysiological and live cell imaging assays, that both agonists and positive allosteric modulators (PAMs) can drive activation and internalization of group II and III mGluRs. The effects of PAMs are pleiotropic, boosting the maximal response to orthosteric agonists and serving independently as internalization-biased agonists across mGluR subtypes. Motivated by this and intersubunit FRET analyses, we determine cryo-electron microscopy structures of mGluR3 in the presence of either an agonist or antagonist alone or in combination with a PAM. These structures reveal PAM-driven re-shaping of intra- and inter-subunit conformations and provide evidence for a rolling TMD dimer interface activation pathway that controls G protein and beta-arrestin coupling.
Similar content being viewed by others
Introduction
G protein coupled receptors (GPCRs) control a wide range of physiological and therapeutic processes by initiating intracellular signaling cascades via heterotrimeric G proteins1,2. Following activation, desensitization mechanisms tune GPCR signaling to maintain temporal precision and avoid excessive activation3. On the seconds to minutes time scale, acute desensitization can occur via G protein receptor kinases (GRKs), which bind to activated receptors to both sterically occlude G protein binding and sequester active G proteins4,5,6. GRKs also phosphorylate key intracellular residues of GPCRs to promote beta-arrestin (\(\beta\)-arr) coupling7,8 resulting in the recruitment of endocytic proteins to drive internalization and trafficking over the minutes to hours time scale9. A critical aspect of GPCR drug targeting is the ability of ligands to control both receptor activation and receptor desensitization. Different ligands for the same receptor are often able to stabilize distinct states within a receptor’s conformational landscape, which can lead to a relative bias toward modes of activation or desensitization10,11,12,13.
Determining the conformational basis of differential effects of ligands on GPCR activation and desensitization is a major challenge, particularly for the dimeric, multi-domain family C GPCRs. The metabotropic glutamate receptor (mGluRs or mGlu receptors) are prototypical family C GPCRs, which mediate synaptic neuromodulation and serve as prominent drug targets for a plethora of neurological and psychiatric disorders14,15,16. Structurally, mGluRs form constitutive dimers17,18,19,20 with each subunit containing a large extracellular domain consisting of a bi-lobed ligand binding domain (LBD) and a cysteine rich domain (CRD). The CRD connects to the canonical seven helix transmembrane domain (TMD) which is followed by a disordered C-terminal domain (CTD)21. The LBDs serve as the binding site for the native agonist glutamate as well as other synthetic orthosteric agonists and antagonists, while the TMDs both initiate transducer coupling and provide a binding site for synthetic positive (PAMs) and negative (NAMs) allosteric modulators22,23. PAMs are primarily defined by their ability to enhance the effect of agonists on G protein signaling23,24,25, but it has become increasingly clear that many mGluR PAMs can directly evoke activation in the full-length receptor in the absence of agonist26,27,28,29,30, thus serving as “ago-PAMs”. In addition to their ability to activate heterotrimeric G proteins, we have recently shown that a subset of mGluRs is capable of glutamate-evoked internalization by GRKs and \(\beta\)-arrs6,31. However, the capacity of different orthosteric and allosteric compounds to evoke mGluR internalization has not yet been addressed, raising the question of potential functional selectivity. This knowledge is critical in a therapeutic context as the relative ability of ligands to drive G protein versus GRK/\(\beta\)-arr coupling is likely a major determinant of efficacy and long-term compensatory effects, as is well-documented for a variety of family A GPCRs32,33,34.
Structural28,35,36,37,38,39,40 and spectroscopic18,19,41,42,43,44,45,46,47 studies of full-length receptors have revealed complex intra- and inter-subunit rearrangements that drive mGluR activation. Briefly, agonist-mediated activation has been shown to involve LBD closure, re-shaping of the LBD dimer interface, inter-CRD compaction, and eventual formation of a TM6 containing inter-TMD interface. However, a full understanding of the conformational pathways controlled by both orthosteric and allosteric drugs in the context of activation and desensitization remain poorly understood. Ultimately, understanding how agonists and PAMs control mGluR function and conformation is critical for both gaining a fundamental understanding of family C GPCRs and for harnessing the potential of PAMs as subtype-specific therapeutics48,49. Here we use a combination of functional assays to reveal unexpected complexity in the response of group II and III mGluRs to orthosteric versus allosteric ligands. We then resolve cryo-EM structures under a range of ligand conditions, enabling a model of mGluR3 activation and internalization involving a dynamic population of conformational intermediates differentially stabilized by orthosteric and allosteric ligands.
Results
Orthosteric agonists drive activation and internalization with a correlated maximum efficacy
We first asked how mGluRs respond to a range of orthosteric agonists (Supplementary Fig. 1a) in terms of both activation and internalization with an initial focus on mGluR3, the subtype with the most robust acute desensitization and endocytosis31. We assessed the agonists LY379268 and DCG-IV by first comparing their effects to glutamate in a G protein-dependent G protein-coupled inwardly rectifying potassium (GIRK) current assay (Fig. 1a). All three agonists produced dose-dependent, reversible GIRK currents (Fig. 1b; Supplementary Fig. 1b, c) but the maximal current evoked by saturating concentrations were larger than glutamate for LY379268 and smaller than glutamate for DCG-IV (Fig. 1c; Supplementary Fig. 1d). These results are consistent with prior studies of group II mGluRs which have characterized LY379268 as a “super agonist” and DCG-IV as a partial agonist41,46,50.
a Schematic showing mGluR-mediated GIRK activation through Gαi/o activation and Gβγ release. b Representative whole-cell patch clamp current trace showing agonist-evoked mGluR3-mediated GIRK potassium currents showing the relative maximum agonist efficacies. Gray dashed line represents peak glutamate current. c Bar plot quantifying the maximum mGluR3-mediated current normalized to the glutamate (100 µM; n = 11 cells) condition for DCG-IV (100 µM; n = 6 cells; p = 0.003) and LY379268 (1 µM; LY37; n = 7 cells; p = 0.007). d Schematic showing surface labeling assay to quantify ligand-evoked receptor internalization. Cells are incubated with agonist for 60 min followed by labelling of SNAP-tagged surface receptors with a membrane-impermeable fluorophore. e Quantification of the % internalization of mGluR3 following treatment with each agonist (same concentrations as panel c) (Glu: n = 6 experimental days, 10 averaged images per day; DCG-IV: n = 5, p = 0.025; LY37: n = 6; p = 0.509). f Representative widefield fluorescence images depicting internalization of SNAP-mGluR3 labeled with BG-Alexa-546 prior to agonist treatment for 30 min. Red arrows highlight endocytosed receptors. Scale Bar = 5 µm. g Scatter plot showing maximum GIRK activation versus internalization for a range of agonists normalized to saturating glutamate for three mGluR subtypes (mGluR3, mGluR7, mGluR8). Linear regression was applied to the entire data set and constrained to go through the glutamate reference point (data points are presented as mean values ± SEM). Bar-plots represent mean ± SEM For (c), (e): One-way ANOVA with Dunnet’s multiple comparisons test against WT. P-values are reported in the specific panel legend.
To assess receptor endocytosis, we used our previously established SNAP-tag-based surface labeling internalization assay (Fig. 1d) where glutamate drives a dose-dependent and β-arr dependent drop in surface fluorescence31 (Supplementary Fig. 1e, f). All three agonists drove mGluR3 internalization, but DCG-IV drove substantially less internalization than glutamate while LY379268 drove slightly more (Fig. 1e). This same trend was observed upon imaging of cells with fluorophore labeling prior to agonist treatment where a larger accumulation of intracellular receptors was seen following glutamate or LY379268 application compared with DCG-IV (Fig. 1f). As a control, all three agonists failed to elicit internalization of mGluR2 (Supplementary Fig. 1g,h).
To further test the correlation of agonist efficacy for activation and internalization, we extended our analysis to more agonists and receptor subtypes mGluR7 and mGluR8, which are also subject to \(\beta\)-arr-mediated internalization31,51,52 (Supplementary Fig. 2). Figure 1g shows a summary of the data, revealing a positive linear correlation between the efficacy of G protein activation and \(\beta\)-arr driven internalization. Notably, none of the agonists tested showed a clear efficacy bias toward one pathway relative to glutamate as the reference ligand. This suggests that shared agonist-driven conformational changes drive both activation and internalization and raises the question of how activation and internalization are related for allosteric ligands.
PAMs promote activation and internalization with a bias toward internalization
PAMs have been reported across mGluR subtypes as potential therapeutics for a range of disorders23,48,53, but few compounds have been shown to have PAM-like activity on mGluR354,55. One recently reported mGluR3 PAM compound, VU6023326, shows similar potency for mGluR2 and mGluR3 and contains general structural similarity to other mGluR PAMs (Supplementary Fig. 3a). In line with the agonism of mGluR2 we have seen with other PAMs27, VU6023326 produced dose-dependent GIRK currents upon application to mGluR3-expressing cells (Fig. 2a, b; Supplementary Fig. 3b). The maximal current evoked by VU6023326 was ~80% of that produced by saturating glutamate (Fig. 2b; Supplementary Fig. 3c), consistent with our prior study of mGluR2 where all PAMs tested showed a 60-80% efficacy27. We then asked if application of PAM alters the maximal response to agonists by applying VU6023326 in the presence of saturating glutamate or LY379268. In both cases, VU6023326 produced a clear inward current that boosted the maximum response to glutamate or LY379268 by 20–30% (Fig. 2c, d; Supplementary Fig. 3d–g). This result reveals that mGluR PAMs can not only serve as allosteric agonists and modulators of agonist affinity, but they can also boost agonist efficacy.
a GIRK current trace showing the response to a saturating dose of VU602 (10 µM; VU6023326) compared to glutamate. b Dose-response curve for VU602-evoked GIRK currents normalized to the response to saturating glutamate. LogEC50 = −8.10 ± 0.16 (error = SEM). Black dashed line shows the glutamate dose-response curve as a reference. c GIRK current trace showing that application of VU602 in the presence of saturating glutamate further potentiates mGluR3-driven currents. d Bar plot summarizing the maximum response for VU602 (10 µM; n = 24 cells), glutamate (100 µM; n = 27 cells; p < 0.001), or co-application of both (n = 6 cells; p < 0.001). Glutamate is used as the reference for normalization. e Representative images showing VU602 and VU602+Glu evoked internalization. Scale bar = 5 µm. f Bar plot showing mGluR3 internalization in response to VU602 (10 µM; n = 18 experimental days, 10 averaged images per day; p = 0.025), glutamate (100 µM; n = 21), or co-application of both (n = 8; p = 0.016). g Summary plots of normalized maximum GIRK activation versus maximum internalization for all PAMs used in this study on mGluR3, mGluR7, or mGluR8 with all values normalized to the effects of saturating glutamate (data points are presented as mean values ± SEM). h Schematic summarizing the effects or PAMs, glutamate or both on G protein activation versus GRK/\(\beta\)-arr internalization across different mGluR subtypes. i Representative images showing extent of mGluR2 internalization for saturating doses of VU602 (10 µM) alone or with 1 mM glutamate. Note: GRK2 was co-expressed for mGluR2 internalization experiments. This experiment was repeated across three different days. j Schematics summarizing the effects of PAMs, glutamate or both on G protein activation versus GRK/\(\beta\)-arr internalization for mGluR2. Bar-plots represent mean ± SEM. For (d, f): One-way ANOVA with Dunnet’s multiple comparisons test against glutamate condition.
Strikingly, VU6023326 produced a larger extent of mGluR3 internalization than glutamate and the response was slightly boosted when ligands were co-applied (Fig. 2e, f; Supplementary Fig. 3h). Importantly, VU6023326-driven mGluR3 internalization was \(\beta\)-arr dependent (Supplementary Fig. 3i). These data indicate that VU6023326 is capable of both initiating G protein-dependent activation and GRK and \(\beta\)-arr-dependent internalization, but that the relative efficacy is higher for internalization.
Motivated by the apparent internalization bias of VU6023326 on mGluR3, we tested a panel of PAMs on group III mGluRs in both GIRK activation and internalization assays (Supplementary Fig. 4). Figure 2g shows a summary of the relative efficacy of G protein activation (i.e. maximal GIRK current amplitude) versus \(\beta\)-arr coupling (i.e. maximum internalization) for all mGluR3, mGluR7, and mGluR8 PAMs tested, highlighting that all PAMs serve as partial agonists toward G protein and show a clear \(\beta\)-arr bias (Fig. 2h). Interestingly, only a subset of PAMs was able to substantially boost the maximal efficacy of glutamate in terms of either G protein activation or internalization. We also investigated the effects of PAMs on mGluR2, a subtype which is resistant to agonist-driven internalization31, but responsive to PAMs as a partial agonist in the GIRK current assay27 (Supplementary Fig. 5a–h). In the internalization assays, neither PAMs nor glutamate alone evoked clear effects (Fig. 2i; Supplementary Fig. 5i). However, co-application of PAM and glutamate produced a clear internalization of mGluR2 with the largest effects for VU6023326 and TASP 0433864 (Fig. 2i; Supplementary Fig. 5i). Importantly, the Glu/PAM-evoked internalization of mGluR2 was GRK and \(\beta\)-arr-dependent (Supplementary Fig. 5j), revealing that PAMs can drive \(\beta\)-arr coupling even in subtypes that do not typically recruit \(\beta\)-arr in response to agonists (Fig. 2j).
PAMs drive LBD-dependent and independent conformational and functional effects
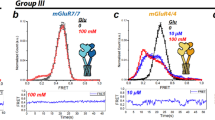
To probe the conformational effects of allosteric modulation, we used inter-subunit mGluR FRET sensors that detect activation-associated rearrangement between LBDs46 or TMDs44. Application of VU6023326 produced a modest, but clear decrease in inter-LBD FRET when applied to SNAP-mGluR3 expressing cells (Fig. 3a, b; Supplementary Fig. 6a–e). Consistent with prior work27, VU6023326 did not produce a response in SNAP-mGluR2 expressing cells (Supplementary Fig. 6e). We reasoned that this subtype discrepancy may be due to the elevated basal inter-LBD conformational dynamics of mGluR341,56. Consistent with this, introduction of the S152D mutant, which blunts basal inter-LBD dynamics41, or application of the orthosteric antagonist LY341495 reduced and abolished the VU6023326 response, respectively (Fig. 3c; Supplementary Fig. 6e). The NAM MNI137 also fully blocked the VU6023326 response (Fig. 3c), likely through a competitive binding mechanism. When co-applied with glutamate, VU6023326 did not further boost the maximal response (Fig. 3b) as it did in the GIRK current assay (Fig. 2c).
a Schematic showing inter-LBD FRET sensor where LBD closure and reorientation from relaxed to active dimer states upon agonist exposure decrease FRET efficiency. b Bar graph showing normalized mGluR3 inter-LBD FRET decrease following application of saturating VU6023326 (10 µM; VU602; n = 6; p < 0.0001), glutamate (n = 19) or both (n = 5; p = 0.5802). c Bar graph showing the inhibition of VU602 inter-LBD FRET responses by competitive antagonist LY341495 (5 µM; LY34; n = 4) or NAM MNI-137 (60 µM; n = 5). d Schematic showing inter-TMD FRET sensor where agonist application receptor increases the FRET efficiency as TMD domains undergo activation-associated rearrangements. e Bar graph showing normalized mGluR3 inter-TMD FRET increase following application of saturating PAM VU6023326 (10 µM; VU602; n = 6; p < 0.0001), glutamate (n = 10) or both (n = 4; p < 0.0001). f Bar graph showing the inhibition of VU602 inter-TMD FRET responses by saturating competitive antagonist LY341495 (10 µM; LY34; n = 5) or NAM MN1-137 (60 µM; n = 5; p < 0.0001). g Representative GIRK current trace for mGluR3 showing the lack of response to PAM (VU602) when applied in the presence of competitive antagonist. h Same as in (g) but for mGluR2, which maintains a clear PAM response in the presence of competitive antagonist. i Bar graph summarizing inhibition of PAM-evoked GIRK currents when applied in the presence of competitive antagonist LY341495 (5 µM) or NAM MNI137 (10 µM) for mGluR3 (LY341495, n = 10 cells; MNI137, n = 5 cells; p = 0.391) and mGluR2 (LY341495, n = 7 cells; MNI137, n = 5 cells; p = 0.0001). Values are normalized to the response to saturating glutamate. Dotted line shows the response to PAM alone for each receptor. j Bar graph showing inhibition of PAM-induced mGluR3 internalization by competitive antagonist LY341495 (20 µM; n = 14 experimental days) or NAM MNI137 (10 µM; n = 7 experimental days; p < 0.0001). k Representative images showing mGluR3 internalization in the presence of antagonist+PAM but not NAM + PAM conditions. Red arrows highlight clusters of internalized receptors. Scale bar = 10 µm. l Schematic summarizing the effects of competitive antagonist on PAM-induced functional effects for mGluR3 and mGluR2. Bar-plots represent mean ± SEM for n = separate preparations for FRET experiments (b, c, e, f), n = cells for electrophysiology experiments (i), and n = biological replicates (separate days) for imaging experiments (j). For (b, e, i): One-way ANOVA with Dunnet’s multiple comparisons against.
We next turned to inter-TMD FRET where we found that VU6023326 can elicit a clear, partial response when applied alone (Fig. 3d, e; Supplementary Fig. 6f–h). The inter-TMD FRET response was only partially reduced by saturating LY341495 application but was nearly abolished by MNI137 (Fig. 3f). Unlike what was seen with inter-LBD FRET, VU6023326 was able to boost the response to saturating glutamate (Fig. 3e; Supplementary Fig. 6i). VU6023326 was also able to boost the maximal response to saturating LY379268 (Supplementary Fig. 6j, k). Together these data show that PAMs can drive inter-LBD and inter-TMD conformational changes but reveal a larger effect on TMD than LBD conformation.
Given the ability of PAMs to drive conformational changes that are independent of LBD closure and rearrangement, we asked what the functional properties are of antagonist/PAM-bound states. Strikingly, co-application of LY341495 abolished the GIRK current response to VU6023326 in mGluR3 (Fig. 3g) but not in mGluR2 expressing cells (Fig. 3h). The latter is consistent with our prior study of mGluR2 PAM-driven currents27. Both mGluR2- or mGluR3-mediated VU6023326 responses were fully blocked by the NAM MNI137 (Fig. 3i; Supplementary Fig. 7a, b). These results suggest that distinct conformational coupling properties between TMDs and LBDs exist in different group II mGluR subtypes.
Unlike what was seen with GIRK currents, LY341495 was only able to weakly inhibit VU6023326-driven internalization of mGluR3 (Fig. 3j, k; Supplementary Fig. 7c). In key controls, LY341495 fully blocked glutamate-driven internalization (Supplementary Fig. 7c) and MNI137 fully blocked both VU6023326 and glutamate-driven internalization (Fig. 3j, k; Supplementary Fig. 7d). The ability of a PAM to drive mGluR3 internalization, but not G protein-activation, in the LY341495-bound state indicates that a GRK/\(\beta\)-arr biased conformation is occupied. Overall, these studies show both ligand and subtype-specific effects, suggesting underappreciated conformational complexity in mGluR activation and desensitization (Fig. 3l).
Agonist-bound Cryo-EM structures reveal PAM-evoked structural reshaping
We turned to single particle cryo-EM to gain more insight into the activation and internalization-associated structural states of mGluR3. We first determined the structure of detergent-solubilized, full length mGluR3 in the presence of LY379268 alone or both LY379268 and VU6023326 (Fig. 4a; Supplementary Fig. 8). LY379268 only and LY379268/VU6023326 structures share a similar global domain arrangement to each other, and to other agonist/PAM bound mGluR structures28,36,37,38,39, with increasing RMSD from LBDs to CRDs to TMDs (Fig. 4a−c; Supplementary Fig. 9a). Due to the high resolution at the LBDs, we were able to clearly place the LY379268 agonist in both structures and to assign density to ions (Supplementary Fig. 9b, c). These ion densities appear in overlapping locations to Cl- ions seen in an mGluR3 LBD crystal structure57, although we cannot unambiguously define their identity.
a Full length cryo-EM density map and protein model for LY379268 (agonist; LY37) and LY37/VU6023326 (agonist/PAM; LY37/VU602) bound mGluR3. b Full-length mGluR3 LY37/VU602 model with LY37 in purple and ions in black highlighting the inter LBD bottom lobe (LB2) distance measured from the Cα of E224 and inter-CRD distance measured from the Cα of C546. c Plot comparing inter-LB2 versus inter-CRD distance across reported agonist-bound homodimeric mGluR subtypes measured via conserved positions highlighted in (b). d mGluR3 LY37 (green) and mGluR3 LY37/VU602 (blue) structures aligned via the top lobe (LB1) of chain A. The angle between helix B and helix O and the angle between helix F of chain A and chain B is reported. Structural offsets are reported for residues T410 (2.7 Å), R206(1.8 Å), I530(2.2 Å), M547(3.9 Å), and E567 (5.2 Å). e Top and bottom views of single TMD alignments between LY37 and LY37/VU602 structures with offsets highlighted by red arrows. Alignments are performed for the entire TMD from residue 571 to the end of the model, including loops. f, g Side views comparing aligned TMDs for mGluR3 LY37 and mGluR3 LY37/VU602 structures. TM4 is hidden from the panel (g) for clarity. h Alignment of TMDs for mGluR3 LY37, mGluR3 LY37/VU602, mGluR2 agonist/PAM (7MTR), and mGluR2 agonist/PAM/G protein (7MTS). For clarity, TM4 is hidden from view. i Side views of LY37 and LY37/VU602 TMD dimer with TM6 shown in dark blue and the Cα of key residues highlighted. j Inter-TM6 interface profile measured by intersubunit Cα distance of outward facing residues for LY37 and LY37/VU602 structures. k Plot of inter TM6 distance across full-length homodimeric mGluR structures using two conserved positions.
In both structures, the LBDs are stabilized in a closed/closed-active conformation with a similar inter-LB2 (lower lobe) distance to prior agonist-bound cryo-EM structures of mGluRs (Fig. 4b, c). Furthermore, both structures show compact CRDs with an inter-subunit distance of ~20-25 Å (Fig. 4b, c). Analysis of inter-CRD distance across agonist-bound mGluR structures revealed more variability compared to inter-LBD distances, with our mGluR3 structures more closely resembling mGluR2 than mGluR5 structures (Fig. 4c). Close analysis of the LBDs revealed rearrangements in the LY379268/VU6023326-bound structure compared to LY379268 only, which may explain the effects of PAM on agonist affinity and/or efficacy (Fig. 4d; Supplementary Fig. 6i–k). In the presence of VU6023326, a compaction occurs at the LBD dimer interface, characterized in the upper lobe (LB1) as a change in angle between helix B of one protomer and helix O of the other protomer, from 136.9o to 139.1o in the LY379268 and LY379268/VU6023326 structures, respectively (Fig. 4d). At LB2, a change in angle from 63.6o (LY379268) to 62.5o (LY379268/VU6023326) is seen between helix F on each protomer, ultimately creating a tighter inter-LBD interface (Fig. 4c, d). This inter-LBD rearrangement subtly reshapes the LY379268 binding site, bringing top lobe residues S151 and R68 ~ 0.5 Å closer to the ligand and decreasing inter-lobe distance in the LY379268/VU6023326 structure (Supplementary Fig. 9b). Differences are also observed in CRD compaction between the LY379268 and LY379268/VU6023326 structures, with a general CRD rearrangement that decreases the inter-subunit distance by 2–5 Å in the LY379268/VU6023326 structure (Fig. 4d). Finally, extracellular loop 2 (EL2), also exhibits a reshaping and positional shift in the presence of VU6023326 (Supplementary Fig. 9d, e), which may contribute to inter-domain allostery as has been proposed in previous mGluR structures36,37,38.
The VU6023326-containing structure shows intra- and inter-subunit rearrangement of the TMD, including a widening of the extracellular face of the TMD bundle due to an outward shift of TM1, TM5, and TM6, and a repositioning of TM3 (Fig. 4e; Supplementary Fig. 10a). This repositioning opens an apparent cavity where density is observed (Supplementary Fig. 10b) that aligns with the location of a previously resolved mGluR2 PAM37 (Supplementary Fig. 10a). Using a cavity detection tool on complete receptor models (see Methods section for details), it is apparent that VU6023326 can fit between TM5 and TM6 in the LY379268/VU6023326 structure, but no cavity is detected in the same region in the LY379268 only structure (Supplementary Fig. 10c). To further understand this likely PAM binding site, we performed molecular docking (see Methods). The top three scoring poses showed VU6023326 within the expected pocket and allow us to predict key stabilizing residues on TM3 (F652, Y656), TM5 (D744), and TM6 (W789, F785, F789) in the mGluR3 TMD (Supplementary Fig. 10d). We mutated each of these positions to alanine and observed impaired PAM-driven GIRK current activation in terms of both potency and/or maximum efficacy for F652A, Y7656A, W782A, and the Y656A/F785A double mutant (Supplementary Fig. 10e–g), supporting this putative binding site.
In the VU6023326-bound structure, re-organization is also seen at the intracellular face of the TMD, with offsets for TM2, TM3, TM4, and TM6, as well as lengthened helical character and an outward bend for TM7 (Fig. 4e–h; Supplementary Fig. 10a, b). Notably, in the LY379268/VU6023326 model TM2 and TM3 reposition to resemble the orientation seen in a G protein-bound mGluR2 structure (Fig. 4h). Together these PAM-induced conformational changes likely reposition intracellular loops, a key step in transducer coupling35,37,40.
A TM6 containing dimer interface is observed in both structures (Fig. 4i), consistent with agonist/PAM-bound structures from other mGluR subtypes28,36,37,38,39 and the high degree of TM6 conservation (Supplementary Fig. 11a, b). However, between structures we observe a difference in both the shape of TM6 and the inter-TMD interface profile. In the absence of VU6023326, TM6 curves such that its closest inter-TMD distance occurs at the middle of TM6 and then widens towards the extracellular face. In contrast, the VU6023326-containing structure shows a straighter TM6 with the tightest inter-TMD distance occurring at the most extracellular region of TM6, as seen in other agonist/PAM family C GPCR structures (Fig. 4i–k; Supplementary Fig. 11c). The change in the inter-TMD interface initiated by PAM likely stabilizes TM6 in a way that contributes to intra-TMD reorganization of helices (Fig. 4e–h) and, ultimately, transducer coupling.
Reshaping of the TMD by PAMs is also supported by three dimensional variability analysis (3DVA)58,59,60. Overall, more distinct motions are observed for the LY379268 structure compared to the LY379268/VU6023326 structure, indicating that the LY379268/VU6023326-bound state is less dynamic, especially in the TMDs and CRDs (Supplementary Fig. 12; Supplementary Movies 1, 2, 3). One motion component only observed from the LY379268 data set shows a straightening and compaction of the top of TM6 and the CRDs (Supplementary Fig. 12b; Supplementary Movie 3), suggesting that agonist bound mGluR3 samples different TM6-containing interfaces and the addition of PAM stabilizes one of the more compact states. This is consistent with a conformational selection model of allosteric modulation and our observation that PAMs increase the maximal inter-TMD FRET response to agonist (Fig. 3). Additionally, the LY379268 data reveals highly dynamic TMD density in which breathing at the TMD dimer interface is observed such that TM5 moves towards and away from the interface (Supplementary Fig. 12c; Supplementary Movie 4). This motion is consistent with a rolling interface from TM5 to TM6 along the activation pathway. Overall, this analysis suggests that the LY379268 data set samples a range of conformations and the addition of VU6023326 stabilizes a subset of these states.
Antagonist-bound cryo-EM structures reveal multiple CRD and TMD positions
To gain structural insight into the effects of PAMs in the absence of agonist, we prepared samples with the antagonist LY341495 with or without VU6023326 (Supplementary Figs. 13, 14). We reasoned that a VU6023326 only condition would be difficult to interpret and likely highly dynamic (i.e. poor resolution), due to basal dynamics of the mGluR3 LBD41,56. In addition, we anticipated that the LY341495/VU6023326 ligand combination may provide an opportunity to stabilize intermediates along the activation/desensitization pathway.
The LY341495 only condition resulted in a heterogeneous data set with three distinct full length cryo-EM density maps (Fig. 5a). All three classes show open LBDs in a relaxed inter-subunit conformation, as expected given the presence of an orthosteric antagonist. The class 1 map represents a relatively high-resolution asymmetric state with a TM3 and TM4-containing interface. The class 2 map contains high resolution LBDs but poor resolution at the CRDs and TMDs. The best resolved state, class 3, is asymmetric and shows a TM5-containing TMD interface. The LY341495/VU6023326 data set also represents a heterogeneous, but more structurally similar, ensemble of states. We resolved two distinct full length cryo-EM density maps with equivalent particle numbers. “Class 1” is a lower resolution asymmetric map with limited TMD density, while the “class 2” map is higher resolution and C2 symmetric (Fig. 5a). Both class 1 and class 2 maps appear to show TM5-containing interfaces, although the improved resolution makes this unambiguous only for class 2.
a Full-length cryo-EM density maps and protein model, when applicable, for LY341495 (antagonist; LY34) and LY34/VU6023326 (antagonist/PAM; LY34/VU602) bound mGluR3. LY34 class 2 and LY34/VU602 class 1 density maps are depicted with a low pass filter of 6 Å. b Plot comparing inter-LB2 versus inter-CRD distance across reported antagonist and or NAM-bound homodimeric mGluR subtypes measured via conserved positions as in Fig. 5b. c LY34 class 1 (gold), LY34 class 3 (raspberry) and LY34/VU602 class 2 (lavender) structures aligned via the top lobe (LB1) of chain A. Structural offsets are reported for residues, R206 (4.4 Å and 1.1 Å), I530 (8.3 Å and 2.7 Å), and P566 (21.5 Å and 5.8 Å). d, e TMD alignments between LY34 class 3 and LY34/VU602 class 2 structures with offsets highlighted by red arrows. Alignments are performed for the entire TMD from residue 571 to the end of the model, including loops. f TMD alignments between LY34/VU602 class 2, LY37/VU602, and agonist/PAM/G protein-bound mGluR2 (PDB: 7TMS) structures showing subtle differences in TM1, TM6, and TM7 positioning. g Top view of TMDs with inter-TMD distance measured from the alpha carbon of Cα V639 (TM3) and I708 (TM4) for LY34 class 1 and between V746 (TM5) for LY34 class 3 and LY34/VU602 class 2. h Inter-TM5 interface profile measured by intersubunit C\(\alpha\) distance of outward facing residues for LY34 class 3 and LY34/VU602 class 2. i Plot of inter-TM5 distance across full length homodimeric mGluR structures using a conserved TM5 position (y-axis) and the center of mass distance of the TMD bundles (x-axis).
The high-resolution class 1 and class 3 LY341495 structures show major differences in domain orientations with much of the difference stemming from TMD positioning (Supplementary Fig. 15a). In contrast, the LY341495 class 3 structure produces a much better global alignment with the LY341495/VU6023326 class 2 structure (Supplementary Fig. 15a). LY341495-bound mGluR3 structures show generally similar LBD conformations to each other and to previously reported antagonist and/or NAM-bound mGluR structures, as measured by LB2 distance (Fig. 5b). However, alignment of LY341495-bound mGluR3 structures revealed differences in inter-LBD and inter-CRD organization. Most notably, LY341495 class 3 and LY341495/VU6023326 class 2 show a large repositioning of LB2 and CRD compared to LY341495 class 1 (Fig. 5c; Supplementary Fig. 15b). A smaller helix F/F angle (52.8o) is seen for LY341495 class 1 compared to LY341495 class 3 (58.3o) and LY341495/VU6023326 (57.0o), indicative of the altered orientation between LB2 of adjacent subunits (Fig. 5c; Supplementary Fig. 15b). No changes in LBD closure or LY341495 binding site coordination are seen in the presence or absence of PAM (Supplementary Fig. 15c). Aligning all LY341495-containing maps reveals that the CRD position for LY341495 class 2 falls between that of class 1 and class 3, supporting the idea that class 2 represents a conformational intermediate (Supplementary Fig. 15d). When aligning the maps obtained for LY341495/VU6023326 class 1 and class 2, only a minor CRD offset is seen, suggesting that these two conformations are closely related, and that PAM constrains the range of CRD motion (Supplementary Fig. 15e). 3DVA on pooled data either for LY341495 class 1 and class 2 or LY341495/VU6023326 class 1 and class 2, reveals LBD movements in both data sets leading to changes in LB2 angles (Supplementary Fig. 15f, g; Supplementary Movie 5, 6). These angle changes are consistent with interconversion between the different helix F angles observed for our LY341495 classes (Fig. 5c), which may accommodate the variety of CRD placements we observe across structures.
Alignment of intra-TMD conformation between LY341495 bound class 1 and class 3 models shows an RMSD of 4.4 Å with modest repositioning of all helices (Supplementary Fig. 16a). Alignment of LY341495 class 3 with the LY341495/VU6023326 structure reveals an improved alignment (RMSD = 4.0 Å), but with subtle outward shifts at the top of TM6 (Supplementary Fig. 16a), which are consistent with the effects of PAM observed in agonist-bound structures. This comparison also reveals differences at the cytoplasmic face of the TMD, which may contribute to transducer coupling. These differences are seen in the form of TM6 curving towards the helical bundle and TM7 lengthening and outward extension in the LY341495/VU6023326 structure, as well as additional offsets are observed at the bottom of TM1 and TM3 (Fig. 5d, e). In line with common conformational effects of PAM in the presence of antagonist or agonist, the LY341495/VU6023326 TMD aligns more closely with the LY379268/VU6023326 TMD (RMSD = 4.0 Å) than the LY379268 only TMD (RMSD = 6.9 Å) (Supplementary Fig. 16a) suggesting that VU6023326 stabilizes similar intra-TMD rearrangements independent of the orthosteric ligand. However, conformational differences are observed for LY341495/VU6023326 compared to LY379268/VU6023326, including a repositioning of the intracellular face at TM1, TM6, and TM7 (Fig. 5f; Supplementary Fig. 16b). Additionally, when aligning these two structures with a G protein bound mGluR2 structure37, the LY379268/VU6023326 structure shares more features with the G protein bound TMD, especially regarding the placement of TM1, TM5, and TM6 (Fig. 5f). These subtle rearrangements may reshape intracellular loops and underlie the apparent lack of PAM-induced G protein coupling seen in the presence of LY341495 (Fig. 3).
The most dramatic differences between LY341495-bound states are seen at the dimeric interface between the TMD bundles. The LY341495 class 1 interface is asymmetric with the extracellular end of TM3 of one protomer and TM4 of the other protomer making contact (Fig. 5g; Supplementary Fig. 16b). The interface is not extensive due to bowing of TM4 and thus is tighter at the extracellular and intracellular ends. By aligning TMD bundles into the LY341495 class 2 density (Supplementary Fig. 16c), we deduce that the inter-TMD interface of this state generally resembles that observed for class 1 but is slightly tighter and more symmetric such that the extracellular side of both TM3 helices contribute. In contrast, the LY341495 class 3 interface only contains TM5 and extends throughout the length of the helix (Fig. 5g; Supplementary Fig. 16b). A TM5 interface is also observed for LY341495/VU6023326 class 2 (Fig. 5g; Supplementary Fig. 16b) and is likely also present for LY341495/VU6023326 class 1 (Supplementary Fig. 16d). While the LY341495/VU6023326 class 2 interface is like that observed with LY341495 class 3, there is a reshaping throughout the length of the interface (Fig. 5h). 3DVA of the LY341495 class 3, but not the LY341495/VU603 class 2, data shows TMD motion in which density assigned to TM4 moves towards the dimeric interface, suggesting the possibility of interconversions between a TM4 and TM5 containing interface (Supplementary Fig. 16e; Supplementary Movie 7).
Notably, our structures represent a tight TM5 containing interface observed for mGluR homodimers (Fig. 5i; Supplementary Fig. 16f), although some prior mGluR homodimer structures show TM5 pointing towards each other at a distance that is too far to form interfacial contacts28,36,38. Furthermore, a recent cryo-EM study revealed a TM5 containing interface for an mGluR2/3 heterodimer40. Additionally, inactive GABAB receptor structures also have been reported to contain variable TM5 interfaces61,62,63,64 (Supplementary Fig. 16f). Overall, our structures and dynamics analyses reveal that in the presence of LY341495, the LBD samples conformations that are compatible with multiple CRD orientations and TMD interfaces, while the addition of PAM reduces the possible orientations sampled and restrains the receptor to a TM5 inter-TMD interface.
A working conformational model of mGluR activation, internalization, and allosteric modulation
Together our structural analyses allow a visualization of the complex conformational changes that occur across activation- and internalization-associated states of mGluR3. Full-length, single-subunit alignments of the five structural models reported here reveal multiple classes of conformational shifts across domains (Fig. 6a). In brief, agonist binding stabilizes a closed LBD which leads to a large CRD repositioning such that LY341495- and LY379268-containing structures show very different TMD positions. Within each cluster of agonist and antagonist-bound structures, the presence of PAM can fine-tune the position of the CRD and TMD. Repositioning of the CRD and TMD under different ligand-associated states leads to a range of inter-TMD dimer interfaces that exists along a corridor on the TMD surface starting with TM3 and TM4 (antagonist-bound), converting to a TM5 interface (antagonist-bound) and converging on a TM6 interface (agonist-bound) (Fig. 6b). PAMs stabilize and reshape the TM5 or TM6 interface in the presence of antagonist or agonist, respectively.
a Single chain alignment for five mGluR3 structures aligned to LB1 of chain A with key rearrangements indicated via red arrows. b Top view of TMD interfaces when aligned to the TMD of chain A with LY34/VU602 TMD shown. Interfacial residues within 11 Å are shown as spheres. c TMDs of LY34 class1 (left), LY34/VU602 class 2 (middle), and LY37/VU602 (right) with the interface and key residues highlighted. d Scatter plot of the effects of alanine mutation to intersubunit interface interacting residues in terms of the response to glutamate or VU602 in G protein activation (x-axis) or internalization (y-axis) assays. Error bars represent the standard error of the mean of the difference between the mutant and the WT for ∆Log EC50 or the maximum internalization difference. e Working model of a ligand-stabilized rolling TMD interface mechanism of activation and internalization.
To test the role of the identified TMD interfaces, we performed alanine scanning mutagenesis of interfacial residues observed in our structures (Fig. 6c). All alanine mutants expressed to normal levels (Supplementary Fig. 16a) and were tested in response to either glutamate or VU6023326 in both the G protein-dependent GIRK current assay and in the surface labeling internalization assay (Supplementary Fig. 17b–v). Mutation to the asymmetric TM3-TM4 interface or symmetric TM6 interface typically enhanced or impaired both activation and internalization, respectively (Fig. 6d). This is consistent with the TM3/TM4 interface representing a fully inactive state and prior work showing that the TM6 interface is involved in activation38,44,65. Notably, the most extracellular residue mutated on TM6, V791A, showed a larger impairment of internalization in response to glutamate than PAM (Fig. 6d; Supplementary Fig. 17s-v). Interestingly, in the internalization assay the effects of mutations were primarily observed in the maximum efficacy compared to WT, with minimal effects on potency.
Mutations to interfacial TM5 residues had more complex results including some divergent responses to glutamate versus PAM (Fig. 6d). Glutamate-driven GIRK activation was left-shifted for all mutants (Supplementary Fig. 17i) while PAM-driven GIRK currents were unaffected, except for L750A which showed a modestly reduced maximal response (Supplementary Fig. 17j, k). In response to glutamate, all variants showed clear internalization with enhancement for T742A and L750A. In contrast, V746A and V753A showed impaired internalization (Supplementary Fig. 17l, n). Similar trends were observed in PAM-induced internalization but with more subtle effects (Supplementary Fig. 17m, o). It’s notable that in response to glutamate V746A and V753A showed enhanced G protein activation and impaired GRK/\(\beta\)-arr coupling (Fig. 6d), suggesting the presence of biased conformations with preferential G protein-coupling. Together, these data show diversity in the role of TM5 residues in shaping the ligand response. While our functional data support the importance of TM5 and TM6 interfaces, it is important to note that TM5 and TM6 are both known to contain key residues that form PAM binding sites26,35,37,66 and critical molecular switches involved in intra-subunit TMD activation28,67,68. Thus, the functional effects of our mutants are likely due to a combination of effects on dimer interface strength/shape, intra-TMD conformational dynamics, and PAM binding. Despite this ambiguity, these data clearly highlight the inter-TMD interface as a hotspot for controlling the response to both PAMs and agonists in terms of both G protein-dependent activation and GRK/\(\beta\)-arr-dependent internalization.
Discussion
Together these functional and structural data are in line with a working model of mGluR3 activation and internalization based on a rolling, stepwise inter-TMD reorientation (Fig. 6e). In the presence of antagonist-stabilized open-open/relaxed LBDs, an ensemble of TM3-, TM4-, and TM5-containing interfaces is populated. With the addition of PAM, the TM5- containing interfaces become the dominant class, which is consistent with the ability of PAM to drive inter-TMD FRET changes even in the presence of LY341495 (Fig. 3; Supplementary Fig. 6). In the presence of agonist-stabilized closed-closed/active LBDs, a variety of TM6-containing interfaces are occupied, including an asymmetric TM5/TM6 interface observed with 3DVA, with PAM stabilizing the re-shaped “6/6*” interface with tighter interactions at the extracellular end of TM6. Our model is supported by a seminal biochemical crosslinking study of mGluR2 which showed ligand-sensitive crosslinking of outward facing residues in TM4, TM5, and TM665.
The ability of PAMs to both serve as modulators of orthosteric agonists as well as direct allosteric agonists, even in the presence of orthosteric antagonists, is consistent with a variety of inactive and active conformations that are differentially populated depending on ligand occupancy. Based on our functional data where the antagonist/PAM ligand combination drives strong internalization, but not G protein-coupling, we reason that the TM5 interface states we observe are intrinsically GRK/\(\beta\)-arr biased (Fig. 6e). In the presence of agonists, PAMs can enhance both G protein activation and internalization, consistent with the TM6 interface comprising a state or ensemble of states with strong coupling to both G protein and GRK/\(\beta\)-arrs (Fig. 6e). In conditions where PAM is applied alone in the absence of agonist or antagonist, it is likely that a more complex combination of conformations, including intermediate states not observed in cryo-EM, are occupied. Presumably, the ability of PAM to stabilize both TM5 (“internalization-biased”) and TM6 (“balanced”) interfaces enables a net bias toward GRK and \(\beta\)-arr mediated desensitization processes relative to glutamate (Fig. 2). In contrast, the variable ability of orthosteric agonists to stabilize the same “balanced” TM6 interface-containing state likely explains the correlation between activation and internalization efficacies (Fig. 1).
The distinct TMD interface stabilized by a given ligand or ligand combination can likely control transducer coupling in two ways. First, different TMD interfaces provide steric constraints that determine the ability of transducers (G proteins, GRKs, \(\beta\)-arrs) to access their binding sites on the intracellular face of the receptor. Indeed, in analysis of a TM4-containing interface of mGluR2, Seven et al. proposed that this conformation was incompatible with G protein coupling to either subunit due to steric clashes with either the other subunit or the plasma membrane37. GRKs and \(\beta\)-arrs may be less constrained by the TMD interface given that their key interactions are with flexible C-terminal domains11,69,70 and \(\beta\)-arrs have been shown to bind GPCR TMD cores with variable angles71,72. Second, different TMD interfaces likely stabilize distinct intra-TMD conformations with intrinsic preferences for G proteins, GRKs, or \(\beta\)-arrs, as has been proposed for family A GPCRs10,11,12,13. Consistent with this, intra-TMD conformational differences are seen in LY341495/VU6023326 (TM5 interface) and LY379268/VU6023326 (TM6 interface) structures which may underlie distinct transducer coupling properties (Supplementary Fig. 16a). Ultimately, transducer-bound structures of mGluRs in the presence of different ligand combinations are needed to test this working model.
Our findings have multiple implications of relevance to the physiological and therapeutic contexts in which mGluR PAMs have been applied. Preclinical studies have pointed to positive allosteric modulation of mGluR3 as a promising strategy for the treatment of cognitive deficits associated with neuropsychiatric diseases, including schizophrenia73,74,75, for neuroprotective effects in the context of neurodegenerative diseases76,77,78, and for the treatment of addiction79. However, the myriad effects of PAMs on both mGluR activation and internalization reported here complicates interpretation of pharmacological studies and therapeutic application of such ligands. Depending on the context, the strong GRK/\(\beta\)-arr coupling driven by PAMs may limit or enhance their therapeutic effects relative to orthosteric agonists. For example, PAMs may boost signaling in the short term but drive a long-term down regulation of mGluRs which limits efficacy and leads to tolerance. Alternatively, the enhanced \(\beta\)-arr coupling produced by PAMs may initiate signaling pathways that drive the beneficial effects of mGluR activation, as has been proposed for some family A GPCR agonists33,34. Our findings that PAMs are also internalization-biased for mGluR7 and mGluR8 and can even drive the internalization of mGluR2, an otherwise internalization-resistant subtype, suggests that this is a critical aspect of allosteric pharmacology to be considered for all mGluR-targeting compounds. Indeed, prior work has also shown that PAMs can strongly drive internalization of mGluR580. Furthermore, the complexity of TMD conformational changes revealed by our work suggests that it may be possible to develop \(\beta\)-arr or G protein-biased allosteric compounds for mGluRs. Overall, extensive work is needed to better understand the effects of G protein-dependent and independent signaling and trafficking of mGluRs in neurological and behavioral contexts and to harness this for improved therapeutic approaches.
Methods
Molecular biology
Rat SNAP-mGluR2, SNAP-mGluR3, SNAP-mGluR7 and human SNAP-mGluR8 constructs used in this study were previously generated20,31. The SNAP-mGluR3-Halo-ICL2 FRET sensors was previously reported44. All SNAP-mGluR constructs contain an N-terminal signal sequence from rat mGluR5 followed by an HA epitope and the SNAP-tag self-labelling enzyme upstream of the mGluR sequence and are inserted in a pRK5 vector backbone. Single point mutagenesis was performed using PCR-mediated site-directed mutagenesis. Previously described GIRK1-F137S homotetramerization mutant81 and tdTomato (Addgene #30530) were used for patch-clamp experiments. GRK2-GFP plasmid5 was used in internalization experiments with mGluR2. Dominant negative β-arr1 (S412D) construct31,82 was used to confirm clathrin-dependent, β-arr-driven internalization of mGluR3 in response to agonist or PAM. Glutamate transporter (EAAT1) plasmid was kindly gifted by Prof. Miriam Stoeber (University of Geneva, Switzerland). For cryo-EM studies, full length rat mGluR3 with a C-terminal thrombin cleavage site followed by a C-terminal mVenus and Twin-Strep-tag83 was generated in the pEZT-BM BacMam expression vector84.
Cell culture and transfection
HEK293 (ATCC: CRL-1573) and HEK293T (ATCC: CRL-3216) cells were grown and maintained in Dulbecco’s Modified Eagle’s Medium (DMEM; #10-013-CV Corning, USA) supplemented with 10% fetal bovine serum (FBS; Thermo Fisher #A5256) and cultured at 37 °C/5% CO2. HEK293 GRK2/3 knock-out cells were kindly gifted by Asuka Inoue (Tohoku University, Japan). For electrophysiology and imaging experiments, cells were seeded on poly-L-lysine (Sigma #P2636) coated rounded 18 mm glass coverslips and transfected using Lipofectamine 2000 (Thermo Fisher Scientific). 6 h after transfection media was replaced and supplemented with antagonist until the time of the experiment (5 µM LY341495 for mGluR2 and mGluR3, 20 µM CPPG for mGluR7 and mGluR8) to maintain cell health. Experiments were performed 24-48 hr post-transfection.
Compound Synthesis and Medicinal Chemistry
Detailed in vitro evaluation of VU6023326 can be found in Yamada et al.54.
All NMR spectra were recorded on a 400 MHz AMX Bruker NMR spectrometer. 1H and 13 C chemical shifts are reported in δ values in ppm downfield with the deuterated solvent as the internal standard. Data are reported as follows: chemical shift, multiplicity (s = singlet, d = doublet, t = triplet, q = quartet, b = broad, m = multiplet), integration, coupling constant (Hz). Low resolution mass spectra were obtained on an Agilent 6120 or 6150 with ESI source. Method A: MS parameters were as follows: fragmentor: 70, capillary voltage: 3000 V, nebulizer pressure: 30 psig, drying gas flow: 13 L/min, drying gas temperature: 350 °C. Samples were introduced via an Agilent 1290 UHPLC comprised of a G4220A binary pump, G4226A ALS, G1316C TCC, and G4212A DAD with ULD flow cell. UV absorption was generally observed at 215 nm and 254 nm with a 4 nm bandwidth. Column: Waters Acquity BEH C18, 1.0 × 50 mm, 1.7 um. Gradient conditions: 5% to 95% CH3CN in H2O (0.1% TFA) over 1.4 min, hold at 95% CH3CN for 0.1 min, 0.5 mL/min, 55 °C. Method B: MS parameters were as follows: fragmentor: 100, capillary voltage: 3000 V, nebulizer pressure: 40 psig, drying gas flow: 11 L/min, drying gas temperature: 350 °C. Samples were introduced via an Agilent 1200 HPLC comprised of a degasser, G1312A binary pump, G1367B HP-ALS, G1316A TCC, G1315D DAD, and a Varian 380 ELSD (if applicable). UV absorption was generally observed at 215 nm and 254 nm with a 4 nm bandwidth. Column: Thermo Accucore C18, 2.1 × 30 mm, 2.6 um. Gradient conditions: 7% to 95% CH3CN in H2O (0.1% TFA) over 1.6 min, hold at 95% CH3CN for 0.35 min, 1.5 mL/min, 45 °C. High resolution mass spectra were obtained on an Agilent 6540 UHD Q-TOF with ESI source. MS parameters were as follows: fragmentor: 150, capillary voltage: 3500 V, nebulizer pressure: 60 psig, drying gas flow: 13 L/min, drying gas temperature: 275 °C. Samples were introduced via an Agilent 1200 UHPLC comprised of a G4220A binary pump, G4226A 3 ALS, G1316C TCC, and G4212A DAD with ULD flow cell. UV absorption was observed at 215 nm and 254 nm with a 4 nm bandwidth. Column: Agilent Zorbax Extend C18, 1.8 µm, 2.1 × 50 mm. Gradient conditions: 5% to 95% CH3CN in H2O (0.1% formic acid) over 1 min, hold at 95% CH3CN for 0.1 min, 0.5 mL/min, 40 °C. Optical specific rotations were obtained using JASCO P-2000 Digital Polarimeter equipped with Tungsten-Halogen lamp (WI), 589 nm wavelength, photomultiplier tube (1P28-01) detector and CG2-100 Cylindrical glass cell, 2.5ø x 100 mm. For compounds that were purified on a Gilson preparative reversed-phase HPLC, the system comprised of a 333 aqueous pump with solvent-selection valve, 334 organic pump, GX271 or GX-281 liquid hander, two column switching valves, and a 155 UV detector. UV wavelength for fraction collection was user-defined, with absorbance at 254 nm always monitored. Method 1: Phenomenex Axia-packed Luna C18, 30 × 50 mm, 5 µm column. Mobile phase: CH3CN in H2O (0.1% TFA). Gradient conditions: 0.75 min equilibration, followed by user defined gradient (starting organic percentage, ending organic percentage, duration), hold at 95% CH3CN in H2O (0.1% TFA) for 1 min, 50 mL/min, 23 °C. Method 2: Phenomenex Axia packed Gemini C18, 50 × 250 mm, 10 um column. Mobile phase: CH3CN in H2O (0.1% TFA). Gradient conditions: 7 min equilibration, followed by user defined gradient (starting organic percentage, ending organic percentage, duration), hold at 95% CH3CN in H2O (0.1% TFA) for 7 min, 120 mL/min, 23 °C. Chiral separation was performed on a Thar (Waters) Investigator SFC Column: Chiral Technologies CHIRALPAK IF, 4.6 × 250 mm, 5 µm column. Gradient conditions: 20% to 50% IPA in CO2 over 7 min, hold at 50% CO2 for 1 min. Flow rate: 3.5 mL/min. Column temperature: 40 °C. System backpressure: 100 bar. Solvents for extraction, washing and chromatography were HPLC grade. All reagents were purchased from Aldrich Chemical Co. and were used without purification.
Preparation of (R)−3-(3-(4-(3,3-dimethylbutanoyl)−3-hydroxy-2-methylphenoxy)−2-methyl-propoxy)−4-methoxybenzoic acid (VU6023326)
In a microwave vial equipped with a stir bar, (S)−1-(2-hydroxy-4-(3-hydroxy-2-methylpropoxy)−3-methylphenyl)−3,3-dimethylbutan-1-one (41 mg, 0.14 mmol), methyl vanillate (13 mg, 0.07 mmol), di-tert-butyl azodicarboxylate (35 mg, 0.15 mmol) and triphenylphosphine (40 mg, 0.15 mmol) in THF (0.5 mL). The reaction was heated to 120 °C for 30 minutes in a Biotage microwave reactor. After completion, the reaction was diluted with water and the desired product extracted with dichloromethane (3X) and the organic layer passed through a phase separator. The product was concentrated and purified using reverse phase HPLC: 30x100mm column, 35–100% acetonitrile gradient (w/ 0.1% TFA) over 10 minutes. Fractions containing the desired product were concentrated to give compound the title compound: VU6023326 (29% yield), HR-MS observed m/z = 445.2228; calculated m/z = 445.2221. LC-MS Retention time: 1.638 min, [M + H]+: 445.2. 1H NMR (400 MHz, CDCl3) δ 7.68 (dd, J = 8.4, 2.0 Hz, 1H), 7.57 – 7.51 (m, 2H), 6.88 (d, J = 8.5 Hz, 1H), 6.39 (d, J = 9.0 Hz, 1H), 4.17 – 4.00 (m, 4H), 3.84 (s, 3H), 2.71 (s, 2H), 2.54 (s, 1H), 2.04 (s, 3H), 1.18 (d, J = 6.9 Hz, 3H), 1.00 (s, 9H).
Patch-clamp electrophysiology
Whole-cell patch-clamp recordings of cells co-expressing mGluRs with GIRK1-F137S and tdTomato were performed 24–36 h post-transfection as previously described in refs. 27,44. GIRK currents were recorded at −60 mV in voltage clamp mode using an Axopatch 200B amplifier and a Digidata 1550B interface commanded via pClampex software (Molecular Devices). Bath solution contained (in mM): 120 KCl, 25 NaCl, 10 HEPES, 2 CaCl2, 1 MgCl2. In this bath solution, different agonists, antagonists, or allosteric modulators were added via gravity-driven perfusion at the concentrations specified for each experiment. For all experiments, drug responses were normalized to glutamate as all recordings concluded with application of saturating glutamate (100 μM for mGluR3; 1 mM for mGluR2, mGluR8; 10 mM for mGluR7). All data was obtained from at least three separate experimental days for each condition. Each condition was measured from at least 3 different cells. For dose-response curves, data points were fitted with a four-parameter non-linear regression model. LogEC50 shifts of the different mutants in Supplementary Fig. 17, were compared by an extra sum-of-squares F-test, where the null hypothesis was that the Log EC50 was the same between WT and each of the mutants.
Widefield fluorescence imaging and surface labeling internalization assay
To visualize internalization, HEK293T cells were transfected with 0.7 µg of SNAP-tagged receptors per well of a 12-well plate. To reduce ambient glutamate levels and prevent basal internalization in the mGluR3 conditions, cells were co-transfected with 0.7 µg of the glutamate transporter (EAAT1). 24 h post-transfection, cells were labeled for 30 min at 37 °C with 1 µM Surface BG-Alexa546 (impermeable dye; New England Biolabs) for SNAP in extracellular buffer (EX) containing (in mM): 10 HEPES, 135 NaCl, 5.4 KCl, 2 CaCl2, 1 MgCl2 and pH 7.4. After labelling, cells were thoroughly washed with fresh EX buffer and incubated in agonist/antagonist and/or PAM/NAM for 30 min. Live cells were imaged on an Olympus IX83 inverted microscope through an CMOS ORCA Science Flash4v3.0 camera (Hamamatsu) and using a 100X immersion oil objective (NA 1.45). The fluorophore was excited using a coupled 561 nm laser. A minimum of three days of independent transfections and a minimum of 7 images were acquired per condition.
For the surface labeling assay, we followed a previously standardized protocol6,31. Briefly, after 24 h expression, SNAP-tagged receptors were incubated in prewarmed culture media containing the desired combination of agonist, antagonist, PAM, and/or NAM for 60 min at 37 °C. Afterwards, cells were labeled with 1 µM Surface BG-Alexa 546 in EX for 20 min at RT and imaged on an Olympus IX83 microscope using a 60X immersion oil objective (NA 1.49). 10–12 snapshots per condition were acquired per day and a minimum of three independent replicates on different days were used per condition. Quantification of the total membrane fluorescence of cells was done using a Li algorithm threshold method in Image J (Fiji). The mean intensity of cells was calculated by subtracting the background for each condition and normalizing against the control antagonist condition. mGluR2 internalization experiments (Supplementary Fig. 5) were performed with co-expression of wild-type GRK2.
Ensemble FRET measurements
Live cell ensemble, intersubunit FRET measurements were performed as previously described in ref. 44. Briefly, 36–48 h post-transfection, cells were washed with EX buffer, labeled with fluorophores for 45 min at 37 °C, and washed again in EX. For inter-LBD FRET experiments, cells expressing N-terminal SNAP-tagged mGluRs were labeled with 1 µM BG-LD555 (Lumidyne Technologies) and 3 µM BG-LD655 (Lumidyne Technologies). For inter-TMD FRET experiments, cells expressing mGluR3 with Halo tag in ICL2 were labeled with 1 µM CA-JF549 (Promega) and 2–3 µM CA-JF646 (Promega). Cells were then imaged at room temperature with a 60x objective (N.A. = 1.45) on an inverted Olympus IX83 microscope with donor excitation via a 561 nm laser and imaging of both donor and acceptor channels at 0.5–1 Hz with an exposure time of 100 ms using two separate sCMOS cameras (Hamamatsu ORCA-Flash4v3.0). A gravity-driven perfusion system was used to deliver drugs diluted in EX solution. Fluorescence intensity traces were extracted from each movie by drawing a region of interest (ROI) around a cell cluster (3–15 cells) from donor and acceptor channels using CellSens software (Olympus). FRET was calculated as FRET = IAcceptor / (IDonor + IAcceptor), in which I = fluorescence intensity and the FRET value was normalized based on the initial baseline before any drug application. For all experiments, data was obtained from multiple cell clusters and averaged from at least three replicates from at least two separate transfections.
Expression and purification of mGluR3
cDNA for rat mGluR3 was cloned into the pEZT-BM BacMam expression vector84 and the C-terminus was fused to a thrombin cleavage site preceding a C-terminal mVenus and Twin-Strep-tag83. The expression construct was transformed into DH10Bac cells to produce bacmid which was then transfected into Spodoptera frugiperda (Sf9) cells grown in ESF 921 media (Expression Systems). P1 and P2 virus production was monitored using GFP fluorescence from the pEZT-BM vector until virus harvesting. HEK293S GnTI- cells (ATCC CRL-3022) were grown at 37 °C and 8% CO2 to a density of 3.5 × 106 cells/mL in FreeStyle suspension media (Gibco) supplemented with 2% fetal bovine serum (Gibco) and Anti-Anti (Gibco). P2 virus was added to cells, the suspension was incubated at 37 °C for 24 h, then sodium butyrate (Sigma) was added to a final concentration of 10 mM and flasks were shifted to 30 °C and 8% CO2. Cells were collected 72 h after transduction by low-speed centrifugation, flash-frozen in liquid nitrogen, and stored at −80 °C.
Cell pellets were resuspended in cold lysis buffer containing 10 mM HEPES (pH 7.5),10 mM MgCl2, 20 mM KCl, 1 M mM NaCl, EDTA-free protease inhibitor cocktail tablet (Sigma), and 25 ug/ml DNase (50 mL buffer per ~1 g of cell pellet). Following manual pipetting, the cells were disrupted using a QSonica Q700 sonicator (5 × 15 s, power level 60). The lysate was then clarified by low-speed centrifugation at 7200 × g for 20 min and membranes were isolated by ultracentrifugation at 125,000 × g for 2 h. All steps were performed on ice or at 4 oC. Membranes were then collected and covered with 1 mL membrane freezing buffer containing 10 mM HEPES (pH 7.5),10 mM MgCl2, 20 mM KCl, and 30% glycerol, flash-frozen in liquid nitrogen, and stored at −80 °C. For all LY341495 bound structures, 20 \(\mu\)M LY341495 was present in all steps.
Membranes were resuspended and Dounce homogenized in buffer containing 100 mM HEPES (pH 7.5), 300 mM NaCl, 2 mM CaCl2, and the appropriate ligand (10 \(\mu\)M LY379268, 20 \(\mu\)M LY341495, 10 \(\mu\)M VU6023326). For the LY341495 plus VU6023326 preparation only, the sample was incubated with an additional 100 \(\mu\)M VU6023326 for 30 minutes on ice prior to proceeding to the next step. Protein was extracted via the addition of n-Dodecyl-β-D-maltopyranoside (DDM) with cholesteryl hemisuccinate (CHS) (Anatrace, D310-CH210) to a final concentration of 50 mM DDM / 4 mM CHS and rotated for 1 h at 4 °C. Following ultracentrifugation at 4 oC at 125,000 × g for 50 min the supernatant was filtered and bound to a 5 mL Strep-Tactin column (GE) equilibrated with running buffer containing 100 mM HEPES (pH 7.5), 300 mM NaCl, 2 mM CaCl2, 0.1% glyco-diosgenin (GDN), and the appropriate ligand. To facilitate detergent exchange, the column was washed with running buffer for 5x column volumes. To remove bound heat shock proteins, the column was washed with 5x column volume of running buffer supplemented with 10 mM MgCl2 and 5 mM ATP and then 5x more washes of running buffer. Protein was subsequently eluted via running buffer supplemented with 5 mM D-Desthiobiotin (IBA). Elution fractions were collected, analyzed by SDS-PAGE and concentrated using 100KD cut-off. The sample was loaded onto a Superose 6 Increase 10/300 GL (Cytiva) column equilibrated in buffer containing 100 mM HEPES (pH 7.5), 300 mM NaCl, 2 mM CaCl2, 0.01% GDN, and the appropriate ligand. Elution fractions were collected, analyzed by SDS-PAGE, and key fractions were concentrated using a 100 KDa cut off and used for cryo-EM studies.
Cryo-EM sample preparation, data acquisition, and processing
mGluR3 samples ranging from 2.0 to 4.7 mg/mL were applied at a volume of 3 \({{\mu }}\)L to plasma treated UltrAuFoil 1.2/1.3 300 mesh grids (Quantifoil). Vitrified samples were prepared using a Vitrobot Mk IV (Thermo Fisher), blot time of 2 s, blot force of −2, at 100% humidity and 23 oC. Grids were imaged with a Titan Krios electron microscope (Thermo Fisher) operated at 300 kV and a nominal magnification of 105,000× and equipped with a K3 camera (Gatan) set in super-resolution mode (0.426 Å pixel size). Images were collected using Leginon.
Beam induced motion was corrected using MotionCor2 in Relion 3.185 using two-fold binning and dose-weighting. These images were then used for contrast transfer function estimation via CTFFIND4.186. In Relion, particles were picked using a 3D initial model template of the receptor and extracted using a box size of 416. Further processing was then performed in CryoSPARC version 3.3.287. Ab initio followed by multiple rounds of heterogeneous refinement resulted in one good class that was further refined via subsequent ab initio and heterogeneous refinement jobs (Supplementary Figs. 8, 13, and 14). For the LY341495 data set only, an additional heterogenous refinement step with C2 symmetry enforced was performed to separate class 1 and class 2 particles, and then symmetry was no longer enforced for subsequent steps in this data set. The best heterogeneous refinement class/classes were then used for homogeneous refinement followed by non-uniform refinement. For the LY379268, LY379268/VU6023326, and LY341495/VU6023326 class 2 data sets, we enforced C2 symmetry at the last non-uniform processing of the full-length map, as it improved resolution while maintaining all features seen without symmetry constraints. These particles were then imported back into Relion for Bayesian polishing88 followed by non-uniform refinement to obtain the best full length map. As we anticipated that flexibility between the extracellular domain (LBD and CRD) and the TMDs contributes to heterogeneity within our full-length map, we performed local refinement. Using a mask of the extracellular domain (LBDs and CRDs), local refinement was performed to obtain a map of the extracellular domain at improved resolution. Additionally using a mask of the TMD or TMD and CRD, particle subtraction was performed to generate a particle stack only containing TMD information followed by 2D classification to exclude a small number of poorly resolved particles, and then an ab initio job to generate a 3D volume and non-uniform refinement. This results in a map of the TMD with improved density for helices that are not well resolved in the full-length map. When applicable, 3D variability analysis was performed on full-length maps with a mask that excluded solvent and micelle, with 5 variability components and a low-pass filter resolution of 5 Å.
Model building
To ensure proper alignment of maps, the ECD and TMD maps were aligned to their respective full-length map. Initial models were built using mGluR3 homology models for 7MTR (used for LY379268 and LY379268/VU6023326), 7EPA (used for LY341495 class 1), 7WI6 (used for LY341495/VU6023326) or our own previous models. Each model was then refined via iterative rounds of real-space refinement using the full length map in Phenix89 and manual refinement in Coot90 using information from both the full length and domain maps. Regions with poor resolution were either deleted, as in the case of some loops, or stubbed, as is the case of the TMDs. Visual representations were prepared using ChimeraX91 and Pymol92.
To prepare the models for cavity analysis, the missing atomic coordinates in chain A of the LY379268 and LY379268/VU6023326 cryo-EM structures without N- and C-termini (residues 31-822) were generated using the MODELLER software version 9.1993 and the rat mGluR3 sequence (UniProt ID P31422). Specifically, missing segments from the extracellular domain (ECD; residues 120-139) and intracellular loop 2 (IC2; residues 664-685) were modeled based on the mGluR2 structures corresponding to PDB IDs 7EPB and 7E9G, respectively, while missing TMD side chains and segments from intracellular loop 1 (IC1; residues 600-610), intracellular loop 3 (IC3; residues 759-766), and extracellular loop 3 (EC3; residues 794-800) were based on the mGluR2 structure corresponding to PDB ID 7MTR. All remaining segments with fewer than four residues were built ab initio. Reconstructed models of chain A for each structure were selected based on the lowest Discrete Optimized Protein Energy (DOPE) score94, and then copied and overlapped onto chain B using the PyMOL Molecular Graphics System (version 2.5.2). The final models were prepared for analysis using the Protein Preparation Wizard within the Schrödinger Maestro software package95 and the default protocol. The latter involved adding missing side chains and hydrogen atoms, capping the two protomers at residues R31 and K822, assigning correct bond orders and side chain protonation states at pH 7.4, and performing a short, restrained minimization in vacuo using the OPLS3e force field96.
Cavity analysis
MOL2 files for chain A of the LY379268 and LY379268/VU6023326 structures were generated using ChimeraX97. The IChem toolkit (version 5.2.9)98,99 was used to perform a cavity analysis on each structure. Specifically, the cavity detection and druggability/ligandability prediction tool called Volsite was used in unrestricted mode. The identified cavities were then assessed based on a measured druggability score (drug score) with positive values indicating druggable cavities and negative values indicating non-druggable cavities. Molecular graphics were created using PyMOL.
Molecular docking
VU6023326 was prepared for docking using the LigPrep utility included in the Schrödinger 2022-4 suite. The ionization state was chosen with the carboxylate group in anionic form as is expected at physiological pH. Chain A of the agonist-PAM mGluR2 structure (PDB 7MTR) was superimposed onto the TMD of the LY379268/VU6023326 mGluR3 structure using TM3, TM5, and TM6 for alignment. The residues in the LY379268/VU6023326 structure corresponding to the binding pocket of the mGluR2 PAM, ADX55164, were optimized around ADX55164 prior to docking. Attempts were made to explore multiple rotamer orientations of Trp6,46, based on the agonist-bound structures of mGluR2 and mGluR3, both with and without PAM, as available in the Protein Data Bank (PDB IDs 7MTR, 7MTS and 8JD5 with PAM; 7E9G, 8JD2 and 8JD3 without PAM). Subsequently, a cubic docking grid with an edge of 30 Å was generated at the center of mass of ADX55164 using Schrödinger’s Receptor Grid Generation utility. The mGluR3 PAM VU6023326 was docked using Schrödinger’s Glide tool with the Extra Precision (XP) scoring function. ADX55164 was docked as a control using the same procedure.
Quantification and statistical analysis
Statistical analysis for all experiments is indicated in figure legends, with error bars representing standard error of the mean (SEM) and each data point being repeated in triplicate or more. Analysis was performed using GraphPad Prism 9.0.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The atomic coordinates and cryo-EM density maps for mGluR3 bound to LY379268, LY379268/VU6023326, LY341495 class 1, LY341495 class 3, and LY341495/VU6023326 class 2 have been deposited in the PDB under accession codes 8TR2, 8TQB, 8TRD, 8TR0, and 8TRC, respectively, and in the Electron Microscopy Data Bank under accession codes EMD-41568, EMD-41501, EMD-41578, EMD-41567, and EMD-41577 respectively. Cryo-EM density maps for mGluR3 bound to LY341495 (class 2) and LY341495/VU6023326 class 1 have been deposited in the Electron Microscopy Data Bank under accession codes EMD-44861 and EMD-45242, respectively. All maps and models are available for download: https://wcm.box.com/s/1ou6btegy7i45cbtn5cgli40g5taui7c. All other datasets generated during and/or analyzed during the current study are available in the Source Data file. Source data are provided with this paper.
References
Pierce, K. L., Premont, R. T. & Lefkowitz, R. J. Seven-transmembrane receptors. Nat. Rev. Mol. Cell Biol. 3, 639–650 (2002).
Palczewski, K. & Orban, T. From Atomic Structures to Neuronal Functions of G Protein–Coupled Receptors. Annu. Rev. Neurosci. 36, 139–164 (2013).
Rajagopal, S. & Shenoy, S. K. GPCR desensitization: Acute and prolonged phases. Cell. Signal. 41, 9–16 (2018).
Tesmer, V. M., Kawano, T., Shankaranarayanan, A., Kozasa, T. & Tesmer, J. J. G. Structural biology: Snapshot of activated G proteins at the membrane: The Gαq-GRK2-Gβγ complex. Science 310, 1686–1690 (2005).
Raveh, A., Cooper, A., Guy-David, L. & Reuveny, E. Nonenzymatic Rapid Control of GIRK Channel Function by a G Protein-Coupled Receptor Kinase. Cell 143, 750–760 (2010).
Xiang, G. et al. Control of Gαq signaling dynamics and GPCR cross-talk by GRKs. Sci. Adv. 8, eabq3363 (2023).
Zhou, X. E. et al. Identification of Phosphorylation Codes for Arrestin Recruitment by G Protein-Coupled Receptors. Cell 170, 457–469.e13 (2017).
Chen, Q. & Tesmer, J. J. G. G protein–coupled receptor interactions with arrestins and GPCR kinases: The unresolved issue of signal bias. J. Biol. Chem. 298, 102279 (2022).
von Zastrow, M. & Sorkin, A. Mechanisms for Regulating and Organizing Receptor Signaling by Endocytosis. Annu. Rev. Biochem. 90, 709–737 (2021).
Smith, J. S., Lefkowitz, R. J. & Rajagopal, S. Biased signalling: From simple switches to allosteric microprocessors. Nat. Rev. Drug Discov. 17, 243–260 (2018).
Hilger, D., Masureel, M. & Kobilka, B. K. Structure and dynamics of GPCR signaling complexes. Nat. Struct. Mol. Biol. 25, 4–12 (2018).
Wingler, L. M. & Lefkowitz, R. J. Conformational Basis of G Protein-Coupled Receptor Signaling Versatility. Trends Cell Biol 30, 736–747 (2020).
Wootten, D., Christopoulos, A., Marti-Solano, M., Babu, M. M. & Sexton, P. M. Mechanisms of signalling and biased agonism in G protein-coupled receptors. Nat. Rev. Mol. Cell Biol. 19, 638–653 (2018).
Reiner, A. & Levitz, J. Glutamatergic Signaling in the Central Nervous System: Ionotropic and Metabotropic Receptors in Concert. Neuron 98, 1080–1098 (2018).
Niswender, C. M. & Conn, P. J. Metabotropic Glutamate Receptors: Physiology, Pharmacology, and Disease. Annu. Rev. Pharmacol. Toxicol. 50, 295–322 (2010).
Nicoletti, F. et al. Metabotropic glutamate receptors: From the workbench to the bedside. Neuropharmacology 60, 1017–1041 (2011).
Romano, C., Yang, W. L. & O’Malley, K. L. Metabotropic glutamate receptor 5 is a disulfide-linked dimer. J. Biol. Chem. 271, 28612–28616 (1996).
Levitz, J. et al. Mechanism of Assembly and Cooperativity of Homomeric and Heteromeric Metabotropic Glutamate Receptors. Neuron 92, 143–159 (2016).
El Moustaine, D. et al. Distinct roles of metabotropic glutamate receptor dimerization in agonist activation and G-protein coupling. Proc. Natl. Acad. Sci. USA. 109, 16342–16347 (2012).
Doumazane, E. et al. A new approach to analyze cell surface protein complexes reveals specific heterodimeric metabotropic glutamate receptors. FASEB J. 25, 66–77 (2011).
Ellaithy, A., Gonzalez-Maeso, J., Logothetis, D. A. & Levitz, J. Structural and Biophysical Mechanisms of Class C G Protein-Coupled Receptor Function. Trends Biochem. Sci. 45, 1049–1064 (2020).
Gregory, K. J. & Goudet, C. International Union of Basic and Clinical Pharmacology. CXI. Pharmacology, Signaling, and Physiology of Metabotropic Glutamate Receptors. Pharmacol. Rev. 73, 521 LP–521569 (2021).
Lindsley, C. W. et al. Practical Strategies and Concepts in GPCR Allosteric Modulator Discovery: Recent Advances with Metabotropic Glutamate Receptors. Chem. Rev. 116, 6707–6741 (2016).
Wacker, D., Stevens, R. C. & Roth, B. L. How Ligands Illuminate GPCR Molecular Pharmacology. Cell 170, 414–427 (2017).
May, L. T., Leach, K., Sexton, P. M. & Christopoulos, A. Allosteric Modulation of G Protein–Coupled Receptors. Annu. Rev. Pharmacol. Toxicol. 47, 1–51 (2007).
Rovira, X. et al. Overlapping binding sites drive allosteric agonism and positive cooperativity in type 4 metabotropic glutamate receptors. FASEB J 29, 116–130 (2015).
Gutzeit, V. A. et al. Conformational dynamics between transmembrane domains and allosteric modulation of a metabotropic glutamate receptor. Elife 8, 1–29 (2019).
Nasrallah, C. et al. Agonists and allosteric modulators promote signaling from different metabotropic glutamate receptor 5 conformations. Cell Rep. 36, 109648 (2021).
Mitsukawa, K. et al. A selective metabotropic glutamate receptor 7 agonist: Activation of receptor signaling via an allosteric site modulates stress parameters in vivo. Proc. Natl. Acad. Sci. 102, 18712–18717 (2005).
O’Brien, D. E. et al. Differential Pharmacology and Binding of mGlu2 Receptor Allosteric Modulators. Mol. Pharmacol. 93, 526 LP–526540 (2018).
Abreu, N., Acosta-Ruiz, A., Xiang, G. & Levitz, J. Mechanisms of differential desensitization of metabotropic glutamate receptors. Cell Rep. 35, 109050 (2021).
Che, T., Dwivedi-Agnihotri, H., Shukla, A. K. & Roth, B. L. Biased ligands at opioid receptors: Current status and future directions. Sci. Signal. 14, eaav0320 (2021).
Slosky, L. M. et al. β-Arrestin-Biased Allosteric Modulator of NTSR1 Selectively Attenuates Addictive Behaviors. Cell 181, 1364–1379.e14 (2020).
Ko, M. J. et al. β-Arrestin–dependent ERK signaling reduces anxiety-like and conditioned fear–related behaviors in mice. Sci. Signal. 14, eaba0245 (2021).
Lin, S. et al. Structures of Gi-bound metabotropic glutamate receptors mGlu2 and mGlu4. Nature 594, 583–588 (2021).
Du, J. et al. Structures of human mGlu2 and mGlu7 homo- and heterodimers. Nature 594, 589–593 (2021).
Seven, A. B. et al. G-protein activation by a metabotropic glutamate receptor. Nature https://doi.org/10.1038/s41586-021-03680-3 (2021).
Koehl, A. et al. Structural insights into the activation of metabotropic glutamate receptors. Nature 566, 79–84 (2019).
Fang, W. et al. Structural basis of the activation of metabotropic glutamate receptor 3. Cell Res. 32, 695–698 (2022).
Wang, X. et al. Structural insights into dimerization and activation of the mGlu2–mGlu3 and mGlu2–mGlu4 heterodimers. Cell Res. https://doi.org/10.1038/s41422-023-00830-2 (2023).
Vafabakhsh, R., Levitz, J. & Isacoff, E. Y. Conformational dynamics of a class C G-protein-coupled receptor. Nature 524, 497–501 (2015).
Habrian, C. H. et al. Conformational pathway provides unique sensitivity to a synaptic mGluR. Nat. Commun. 10, 1–13 (2019).
Liauw, B. W. H., Afsari, H. S. & Vafabakhsh, R. Conformational rearrangement during activation of a metabotropic glutamate receptor. Nat. Chem. Biol. 17, 291–297 (2021).
Thibado, J. K. et al. Differences in interactions between transmembrane domains tune the activation of metabotropic glutamate receptors. Elife 10, e67027 (2021).
Lecat-Guillet, N. et al. Concerted conformational changes control metabotropic glutamate receptor activity. Sci. Adv. 9, eadf1378 (2023).
Doumazane, E. et al. Illuminating the activation mechanisms and allosteric properties of metabotropic glutamate receptors. Proc. Natl. Acad. Sci 110, E1416–E1425 (2013).
Grushevskyi, E. O. et al. Stepwise activation of a class C GPCR begins with millisecond dimer rearrangement. Proc. Natl. Acad. Sci. USA. 116, 10150–10155 (2019).
Foster, D. J. & Conn, P. J. Allosteric Modulation of GPCRs: New Insights and Potential Utility for Treatment of Schizophrenia and Other CNS Disorders. Neuron 94, 431–446 (2017).
Luessen, D. J. & Conn, P. J. Allosteric Modulators of Metabotropic Glutamate Receptors as Novel Therapeutics for Neuropsychiatric Disease. Pharmacol. Rev. 74, 630 LP–630661 (2022).
Olofsson, L. et al. Fine tuning of sub-millisecond conformational dynamics controls metabotropic glutamate receptors agonist efficacy. Nat. Commun 5, 5206 (2014).
Lee, J. et al. Distinct beta-arrestin coupling and intracellular trafficking of metabotropic glutamate receptor homo- and heterodimers. Sci. Adv. 9, eadi8076 (2023).
Lee, S. et al. Nedd4 E3 ligase and beta-arrestins regulate ubiquitination, trafficking, and stability of the mGlu7 receptor. Elife 8, 1–27 (2019).
Stansley, B. J. & Conn, P. J. Neuropharmacological Insight from Allosteric Modulation of mGlu Receptors. Trends Pharmacol. Sci. 40, 240–252 (2019).
Yamada, Y. et al. Positive allosteric modulators (PAMs) of the group II metabotropic glutamate receptors: Design, synthesis, and evaluation as ex-vivo tool compounds. BMCL 50, 128342 (2021).
Schann, S. et al. Chemical Switch of a Metabotropic Glutamate Receptor 2 Silent Allosteric Modulator into Dual Metabotropic Glutamate Receptor 2/3 Negative/Positive Allosteric Modulators. J. Med. Chem. 53, 8775–8779 (2010).
Tora, A. S. et al. Chloride ions stabilize the glutamate-induced active state of the metabotropic glutamate receptor 3. Neuropharmacology 140, 275–286 (2018).
Monn, J. A. et al. Synthesis and Pharmacological Characterization of C4-(Thiotriazolyl)-substituted-2-aminobicyclo[3.1.0]hexane-2,6-dicarboxylates. Identification of (1R,2S,4R,5R,6R)−2-Amino-4-(1H-1,2,4-triazol-3-ylsulfanyl)bicyclo[3.1.0]hexane-2,6-dicarboxylic Acid (LY2812. J. Med. Chem 58, 7526–7548 (2015).
Marino, J. & Schertler, G. F. X. A set of common movements within GPCR-G-protein complexes from variability analysis of cryo-EM datasets. J. Struct. Biol. 213, 107699 (2021).
Gao, Y. et al. Asymmetric activation of the calcium-sensing receptor homodimer. Nature 595, 455–459 (2021).
Punjani, A. & Fleet, D. J. 3D variability analysis: Resolving continuous flexibility and discrete heterogeneity from single particle cryo-EM. J. Struct. Biol. 213, 1–25 (2021).
Park, J. et al. Structure of human GABAB receptor in an inactive state. Nature https://doi.org/10.1038/s41586-020-2452-0 (2020).
Shaye, H. et al. Structural basis of the activation of a metabotropic GABA receptor. Nature. 2020 1–6 https://doi.org/10.1038/s41586-020-2408-4 (2020).
Papasergi-Scott, M. M. et al. Structures of metabotropic GABAB receptor. Nature https://doi.org/10.1038/s41586-020-2469-4 (2020).
Kim, Y., Jeong, E., Jeong, J.-H., Kim, Y. & Cho, Y. Structural Basis for Activation of the Heterodimeric GABAB Receptor. J. Mol. Biol. 432, 5966–5984 (2020).
Xue, L. et al. Major ligand-induced rearrangement of the heptahelical domain interface in a GPCR dimer. Nat. Chem. Biol. 11, 134–140 (2015).
Pérez-Benito, L. et al. Molecular Switches of Allosteric Modulation of the Metabotropic Glutamate 2 Receptor. Structure 25, 1153–1162.e4 (2017).
Zhou, Q. et al. Common activation mechanism of class a GPCRs. eLife 8, e50279 (2019).
Doré, A. S. et al. Structure of class C GPCR metabotropic glutamate receptor 5 transmembrane domain. Nature 511, 557–562 (2014).
Cahill, T. J. et al. Distinct conformations of GPCR-β-arrestin complexes mediate desensitization, signaling, and endocytosis. Proc. Natl. Acad. Sci. USA. 114, 2562–2567 (2017).
Shukla, A. K. et al. Visualization of arrestin recruitment by a G-protein-coupled receptor. Nature 512, 218–222 (2014).
Huang, W. et al. Structure of the neurotensin receptor 1 in complex with β-arrestin 1. Nature 579, 303–308 (2020).
Maharana, J., Banerjee, R., Yadav, M. K., Sarma, P. & Shukla, A. K. Emerging structural insights into GPCR-β-arrestin interaction and functional outcomes. Curr. Opin. Struct. Biol. 75, 102406 (2022).
Dogra, S. & Conn, P. J. Targeting metabotropic glutamate receptors for the treatment of depression and other stress-related disorders. Neuropharmacology 196, 108687 (2021).
Lainiola, M., Procaccini, C. & Linden, A.-M. mGluR3 knockout mice show a working memory defect and an enhanced response to MK-801 in the T- and Y-maze cognitive tests. Behav. Brain Res. 266, 94–103 (2014).
De Filippis, B. et al. The role of group II metabotropic glutamate receptors in cognition and anxiety: Comparative studies in GRM2−/−, GRM3−/− and GRM2/3−/− knockout mice. Neuropharmacology 89, 19–32 (2015).
Dogra, S. & Conn, P. J. Metabotropic glutamate receptors as emerging targets for the treatment of schizophrenia. Mol. Pharmacol. MOLPHARM-MR-2021-000460 https://doi.org/10.1124/molpharm.121.000460 (2022).
Durand, D. et al. Amyloid-beta neurotoxicity and clearance are both regulated by glial group II metabotropic glutamate receptors. Neuropharmacology 123, 274–286 (2017).
Corti, C. et al. The Use of Knock-Out Mice Unravels Distinct Roles for mGlu2 and mGlu3 Metabotropic Glutamate Receptors in Mechanisms of Neurodegeneration/Neuroprotection. J. Neurosci. 27, 8297 LP–8298308 (2007).
Busceti, C. L. et al. Behavioural and biochemical responses to methamphetamine are differentially regulated by mGlu2 and mGlu3 metabotropic glutamate receptors in male mice. Neuropharmacology 196, 108692 (2021).
Arsova, A. et al. Positive Allosteric Modulators of Metabotropic Glutamate Receptor 5 as Tool Compounds to Study Signaling Bias. Mol. Pharmacol. 99, 328–341 (2021).
Vivaudou, M. et al. Probing the G-protein Regulation of GIRK1 and GIRK4, the Two Subunits of the KACh Channel, Using Functional Homomeric Mutants*. J. Biol. Chem. 272, 31553–31560 (1997).
Lin, F.-T. et al. Clathrin-mediated Endocytosis of the β-Adrenergic Receptor Is Regulated by Phosphorylation/Dephosphorylation of β-Arrestin1. J. Biol. Chem. 272, 31051–31057 (1997).
Schmidt, T. G. M. et al. Development of the Twin-Strep-tag® and its application for purification of recombinant proteins from cell culture supernatants. Protein Expr. Purif. 92, 54–61 (2013).
Morales-Perez, C. L., Noviello, C. M. & Hibbs, R. E. Manipulation of Subunit Stoichiometry in Heteromeric Membrane Proteins. Structure 24, 797–805 (2016).
Zivanov, J. et al. RELION-3: New tools for automated high-resolution cryo-EM structure determination. bioRxiv 7, 1–22 (2018).
Rohou, A. & Grigorieff, N. CTFFIND4: Fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. CryoSPARC: Algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zivanov, J., Nakane, T. & Scheres, S. H. W. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCr J. 6, 5–17 (2019).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: Recent developments in Phenix. Acta Crystallogr. Sect. D Struct. Biol. 75, 861–877 (2019).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of {\it{Coot}}. Acta Crystallogr. Sect. D 66, 486–501 (2010).
Goddard, T. D. et al. UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Schrödinger, LLC. The {PyMOL} Molecular Graphics System, Version~1.8. (2015).
Šali, A. & Blundell, T. L. Comparative Protein Modelling by Satisfaction of Spatial Restraints. J. Mol. Biol. 234, 779–815 (1993).
Shen, M. & Sali, A. Statistical potential for assessment and prediction of protein structures. Protein Sci. 15, 2507–2524 (2006).
Schrödinger. Maestro, Schrödinger, LLC. (2022).
Harder, E. et al. OPLS3: A Force Field Providing Broad Coverage of Drug-like Small Molecules and Proteins. J. Chem. Theory Comput. 12, 281–296 (2016).
Pettersen, E. F. et al. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Da Silva, F., Desaphy, J. & Rognan, D. IChem: A Versatile Toolkit for Detecting, Comparing, and Predicting Protein–Ligand Interactions. Chem. Med. Chem. 13, 507–510 (2018).
Tran-Nguyen, V.-K., Da Silva, F., Bret, G. & Rognan, D. All in One: Cavity Detection, Druggability Estimate, Cavity-Based Pharmacophore Perception, and Virtual Screening. J. Chem. Inf. Model. 59, 573–585 (2019).
Acknowledgements
We thank Drs. David Eliezer, Jeremy Dittman, Changhao He, and Nandish Khanra for helpful discussions. We thank all Levitz Lab members for experimental assistance. We also thank Dr. Carl Fluck and Dr. Devrim Acehan at the Weill Cornell Medicine Cryo-EM Core facility for assistance with sample preparation as well as the Weill Cornell Scientific Computing Unit for their resources. Additionally, we are grateful to Drs. William Rice, Bing Wang, Huihui Kuang, and Alice Paquette at the NYU Langone Health’s Cryo–Electron Microscopy Laboratory for their assistance with data collection. We thank Dr. Craig Lindsley, Director of the Warren Center for Neuroscience Drug Discovery for his support. This work is supported by NIH grants F31NS129320 (A.S.), F32GM148001 (D.M.), and R01NS129904 (J. Levitz), the Margarita Salas Fellowship from the Ministry of Universities of Spain (A.G.-H), an NSF Graduate Research Fellowship (N.A), the Rohr Family Research Scholar Award (J. Levitz), and the Monique Weill-Caulier Award (J. Levitz).
Author information
Authors and Affiliations
Contributions
Protein purification, cryo-EM, and structural analysis were performed by A.S. with contributions from P.S., D.M., J.M., L.S., M.F. and J. Levitz. Live cell functional and conformational experiments were designed, performed, and analyzed by A.G.-H., J. Lee, N.A, M.K., A.S., and J. Levitz. K.G. and B.J.M. synthesized and characterized VU602. A.A. and K.H. assisted with molecular biology and L.S. and M.F. designed and performed computational analyses. A.S., A.G.-H., and J. Levitz wrote the paper with feedback from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Karen Gregory, Ryoji Suno and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Strauss, A., Gonzalez-Hernandez, A.J., Lee, J. et al. Structural basis of positive allosteric modulation of metabotropic glutamate receptor activation and internalization. Nat Commun 15, 6498 (2024). https://doi.org/10.1038/s41467-024-50548-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-50548-x
- Springer Nature Limited