Abstract
Sex chromosomes underlie the development of male or female sex organs across species. While systemic signals derived from sex organs prominently contribute to sex-linked differences, it is unclear whether the intrinsic presence of sex chromosomes in somatic tissues has a specific function. Here, we use genetic tools to show that cellular sex is crucial for sexual differentiation throughout the body in Drosophila melanogaster. We reveal that every somatic cell converts the intrinsic presence of sex chromosomes into the active production of a sex determinant, a female specific serine- and arginine-rich (SR) splicing factor. This discovery dismisses the mosaic model which posits that only a subset of cells has the potential to sexually differentiate. Using cell-specific sex reversals, we show that this prevalence of cellular sex drives sex differences in organ size and body weight and is essential for fecundity. These findings demonstrate that cellular sex drives differentiation programs at an organismal scale and highlight the importance of cellular sex pathways in sex trait evolution.
Similar content being viewed by others
Introduction
Living organisms utilise two major biological strategies to create sexual dimorphisms in somatic tissues1,2. The first involves the development of male and female-specific glands and organs that produce systemic signals. These sex-specific organs can shape and maintain somatic sex differences over time, regardless of the sex of the receiving cells3. The second strategy is based on sex chromosomes and relies on cell-intrinsic mechanisms. In this case, sex-chromosomal genes create distinctions in equivalent differentiated cells in both males and females. Cellular sex plays a critical role in this strategy.
Our understanding of the significance of cellular sex is primarily based on the study of invertebrate model systems4,5 such as Drosophila melanogaster6,7,8. In this particular species, sexual differentiation does not depend on gonadal hormones. Females possess two X chromosomes, while males have one X and one Y chromosome. The specific complement of sex chromosomes in a given cell triggers a splicing cascade that produces an RNA-binding protein called TransformerF (TraF)9,10,11,12,13,14,15. This protein is exclusively produced in female cells and regulates the splicing of exonic enhancers by indirect binding through its partner, Transformer 2 (Tra2)16,17,18,19,20,21,22,23,24,25,26. Unlike TraF, Tra2 is present in both sexes and binds directly to RNA through its RNA-binding domain. TraF and Tra2 influence splice site selection through their arginine- and serine-rich (RS) domains, with TraF’s RS domain serving as a bridge between spliceosome components22,27,28,29.
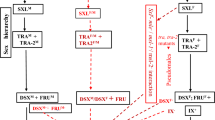
TraF is responsible for the sex-specific splicing of only two identified targets: doublesex (dsx)10,30,31 and fruitless (fru)32,33,34,35,36,37,38,39,40 (Fig. 1a). These two key factors shape the sexual dimorphisms observed in flies. Male-specific isoforms of fru (fruM) are essential for producing transcription factors expressed in specific neurons that underlie sexual orientation and aggression32,33,34,35,36,37,38,39. Both male (dsxM) and female (dsxF) forms of dsx RNA also encode transcription factors, but only a few direct Dsx targets are known7,41. DsxM plays a crucial role in regulating sexual behaviours by controlling the development of male-specific neurons42. DsxF is expressed in just 80 neurons in the female brain, while DsxM is expressed in around 300 neurons in males43. Interestingly, some of these male-specific neurons also co-express FruM.40,44. DsxF activates the expression of yolk protein (Yp) genes in fat body cells, necessary for egg growth45, and also promotes pheromone production by regulating a desaturase called desatF46. Sex-specific development of the gonads is the last and key aspect of sexual differentiation that is regulated by dsx/fru7,47 and conserved in diverse animal species. Both dsx and fru are only expressed in a subset of cells and are required at specific critical periods during development to build dimorphic structures44,48,49,50. Once established, their continuous expression in adulthood is sometimes needed to maintain sex differences. For example, thermos-sensitive alleles of tra2 underscored the necessity of the sex cascade in sustaining sex-specific Yp synthesis51,52 or desatF expression41. Adult gonadal cell types, intriguingly, also exhibit sexual plasticity. The continuous expression of DsxM in the adult testis’ cyst cells is vital for preserving their sexual identity. Indeed, adult-specific ectopic DsxF production feminises these cells into female follicle-like cells47,53. Moreover, dsx and fruM function during adulthood to inhibit male-male courtship and maintain sexual orientation, respectively54,55,56. These results led to the proposal of the two-gene model of sexual differentiation48. According to this model, both male and female flies are composed of a mosaic of dsx-/fru-positive and dsx-/fru-negative cells that, consequently, have and don’t have the potential to sexually differentiate. This mosaic expression of sexuality at the cellular level is believed to be a widespread feature of animals57,58.
a Diagrams displaying the genomic loci of tra, fru and dsx genes. A blue and red triangle indicates the male- and female-specific stop coding in the tra locus. b Expression pattern of dsx-Gal4 and fru-Gal4 (P1 promoter) reporters in female tissues at the third larval stage (DNA labelled with DAPI, blue; the membrane-bound form of GFP, green). c Expression pattern of tra promoter-Gal4 reporter in male and female tissues at the third larval stage (DNA labelled with DAPI, blue; nucleus-targeted GFP, green). d Expression pattern of traF-Gal4 reporter in male and female tissues at the third larval stage (DNA labelled with DAPI, blue; nucleus-targeted GFP, green). e Expression pattern of traF-Gal4 reporter, combined with the dsx-Gal80 and fru-Gal80 (P1 promoter) repressors, in female tissues at the third larval stage (DNA labelled with DAPI, blue; the nuclear form of GFP, green). The visualised cell populations and specific organs are green in the right and top diagrams. Diagrams of the new alleles generated are presented at the top of each panel. Immunohistochemical analyses were repeated at least three independent times.
However, recent studies59,60,61, including our work62,63, have revealed that sex chromosomes impact the function of some cells, even if they do not express dsx and fru, the final effectors of the sex determination cascade. We previously found62 that the sexual identity of adult intestinal stem cells, determined by the presence or absence of TraF, significantly impacts their proliferative capacity. These findings illustrate that somatic cells undergo sex-specific differentiation throughout adulthood, independently of dsx and fru. While we have established a proof-of-principle that TraF-dependent mechanisms impact intestinal stem cells, the full range of functions and phenotypic consequences of this sex pathway remains elusive.
Here, using newly generated Drosophila melanogaster genetic tools, we systematically assess the general significance of cellular sex at the organism-wide level. We found that every organ and somatic cell converts the intrinsic presence of sex chromosomes into the active production of a sex determinant. This ubiquity of cellular sex is crucial in determining sex differences in organ size and body weight. It is also essential for sex organ development and reproduction. Ultimately, our findings also show that these cellular sex pathways depend on the splicing activity of TraF and are active in dsx-expressing cells, where they control species-specific differentiation programmes vital for fertility. By testing the general role of cellular sex, we discover that it plays an essential function in the process of sexual differentiation.
Results
Every cell has a sexual identity
One outstanding question concerning the functioning of the sex determination cascade is whether all cells are sexually differentiated. Based on the expression of dsx and fru, the two terminal effectors of sexual differentiation (Fig. 1b)44,48,49,50, most somatic tissues are the same in males and females and cannot differentiate sexually. Those observations suggest that TraF, the upstream sex determinant, lacks function in most organs. However, this hypothesis has never been tested: although tra was discovered in 19459, the lack of a functional TraF reporter has prevented further examination of tra expression and activity in most organs.
To test this assumption, we replaced tra coding sequences with Gal4 sequences, generating a transgenic reporter line and enabling us to follow tra promoter activity (Fig. 1c). As anticipated, we found that this knock-in Gal4 created a mutant allele of tra that produced classical traKO morphological phenotypes, female to male sex reversals, when heterozygous with a null allele (Supplementary Fig. S1a). Using this genetic tool, we first characterised the transcription pattern of tra at the third larval stage in all major tissues. Unlike dsx and fru, which are only expressed in a restricted subset of tissues (Fig. 1b), this reporter revealed ubiquitous tra expression across organs in both sexes (Fig. 1c).
To further investigate the specification of genetic sex at the organ level, we next examined the translation profile of the TraF protein. Previous work indicates that sex-specific splicing of tra transcripts only leads to the production of functional TraF protein in XX cells7. Therefore, we introduced Gal4 in-frame just before the stop codon in the endogenous tra locus (Fig. 1d). The resulting allele, traF-Gal4, only produces wild-type TraF protein supporting female differentiation (Supplementary Fig. S1b) and Gal4 in females (Fig. 1d). We observed extensive TraF production in the tissues where dsx and fru are expressed, such as in the fat bodies (Fig. 1d), where DsxF activates yolk protein genes45. Interestingly, we also detected TraF in other female organs. For example, TraF is ubiquitously expressed in the female wing imaginal disc, the brain, and the tracheal system (Fig. 1d). We then characterised the temporal dynamics of traF-Gal4-driven expression. We found that TraF is ubiquitously produced at all developmental stages, from embryos to adults (Supplementary Fig. S1c).
To test whether somatic sexual identity based on TraF expression is independent of dsx and fru, we took advantage of Gal80-based repression of Gal464. We generated a transgenic line where dsx-Gal80 and fru-Gal80 repress traF-Gal4-driven expression in dsx- and fru-positive cells. These two Gal80 alleles proved specific and efficient, suppressing dsx- and fru-Gal4-driven expression (Supplementary Fig. S1d, e). These tools were not able to suppress traF-Gal4-driven expression in most organs (Fig. 1e), validating the universality of cellular sex. While only a limited subset of tissues uses dsx and fru to differentiate sexually, our data indicate that cellular sex is ubiquitous, and most female organs have the potential to control their sexual differentiation through a mechanism downstream of TraF independent of classical dsx- and fru-based regulation.
We finally sought to characterise the dimorphic nature of traF-Gal4-driven expression at the cellular level. While our previous results demonstrated that every organ has its own sexual identity, it remains possible that some cells within a tissue (made of multiple cell types) do not have the capacity to sexually differentiate. To test this hypothesis, we introduced a different binary system to detect TraF production: traF-LexA. We combined this novel genetic tool with cell-type specific Gal4-UAS drivers and directly visualised the overlap between TraF expression and specific cell types. We selected two tissues for in-depth examination: the intestine and the brain. All the cell types studied (Fig. 2a and Supplementary Fig. S2a) expressed TraF in females only (Fig. 2b and Supplementary Fig. S2b). For example, all intestinal cell types (tracheal cells, visceral muscles, intestinal progenitors, and entero-endocrine cells) were negative for dsx and fru (Fig. 2c and Supplementary Fig. S2c) but positive for TraF during larval and adult stages (Fig. 2b and Supplementary Fig. S2b).
a Expression pattern of cell-type specific reporters in the female midgut at the third larval stage (DNA labelled with DAPI, blue; the nuclear form of GFP, green). b Expression pattern of cell-type specific reporters, combined with the traF-LexA marker, in the female midgut at the third larval stage (DNA labelled with DAPI, blue; the nuclear form of GFP, green, the nuclear form of tdTomato, red). Arrowheads indicate examples of cells expressing both GFP and tdTomato. c Expression pattern of cell-type specific reporters, combined with the dsx-Gal80 and fru-Gal80 repressors, in the female midgut at the third larval stage (DNA labelled with DAPI, blue; the nuclear form of GFP, green). The visualised cell populations and specific cell types are yellow or green in the right and top diagrams. The TraF-expressing cells alone and co-expressing cell-type specific markers are depicted in red and yellow, respectively. Diagrams of the alleles used are presented on the top of each panel. Immunohistochemical analyses were repeated at least three independent times.
Taken together, our findings establish that cellular sex is ubiquitous both spatially and across developmental stages. Every somatic cell converts the intrinsic presence of sex chromosomes into the active production of the sex determinant TraF. This molecular switch specifies sexual state in a static and binary manner, with no organ, cell type, temporal plasticity or heterogeneity. This refutes the idea that males and females are mosaics of sexually differentiated and sexually undifferentiated cells.
Cellular sex is essential and sufficient to drive sex differences
Our results reveal that all cells are competent to differentiate sexually. Next we tested whether TraF expression reflects physiological differences in these cells. Is the effective action of the sex determination hierarchy limited only to a subset of cells? In Drosophila, based on the two-gene model of sexual differentiation48, it is predicted that the conversion of both dsx- and fru-expressing cells should be enough to completely change an individual’s phenotypic sex. We decided to test this model and assess the global functional significance of ubiquitous TraF expression.
To do so, we developed two new tra alleles, allowing us to perform conditional cis-allele switching65 from a null allele of tra to the wild-type female isoform or the reverse. The first line, traKO FRT traF, allows the rescue of TraF activity following the expression of the Flp recombinase in any cell type of interest. Combining this allele with dsx-Gal4 and fru-Gal4-driven Flp expression generates tra knock-out females with restored traF expression, specifically in dsx- and fru-positive cells (Fig. 3a). The second line, traF FRT traKO, permits the production of TraF knock-in males. In this case, Flp expression in dsx- and fru-positive cells results in the expression of TraF in dsx- and fru-negative cells only (Fig. 3b). RT-qPCR and morphological analyses established that dsx and fru sex-specific splicing (Supplementary Fig. S3a, b), as well as sex-specific abdominal pigmentation and genitalia (Supplementary Fig. S3c, d), remained unaffected validating that our manipulations were indeed restricted to dsx- and fru-negative cells. Based on these findings, we concluded that the modifications introduced into the tra locus generated chimeric male/female animals with manipulation of sexual identity confined to dsx- and fru-negative cells.
a Diagrams representing the specific cell population with tra knock-out in females, the allele generated and the genotypes used. b Diagrams representing the specific cell population with traF knock-in in males, the allele generated and the genotypes tested. c–e Adult wing size (c), adult weight (d), female fertility (e) quantifications following tra knock-out specifically in dsx- and fru-negative cells. In all figures, the wing size and the weight of control females are set at 100%. All other sexes and genotypes are shown as a percentage of this female control. f Ovaries of control females and females with tra knock-out specifically in dsx- and fru-negative cells (DNA labelled with DAPI, blue; Actin stained with phalloidin, red). g–i Adult wing size (g), adult weight (h), and male fertility (i) quantifications following traF knock-in specifically in dsx- and fru-negative cells. j Testes of control males and males with traF knock-in specifically in dsx- and fru-negative cells (DNA labelled with DAPI, blue; Actin stained with phalloidin, red). k–m Adult wing size quantifications following tra knock-out and gain-of-function specifically in wing pouch cells (k), tra knock-down and overexpression in wing pouch cells (l), and tra knock-down specifically in wing cells outside the pouch (m). In this and all subsequent figures, boxplots display the minimum, the maximum, the sample median, and the first and third quartiles. Data was combined from at least three independent experiments. n = wing number measured per genotype in (c), (g), (k), (l), and (m), number of replicates (each repeat containing five flies) in (d) and (h) and number of flies in (e) and (i). Asterisks highlighting significant comparisons across sexes are displayed in grey boxes at the bottom of graphs; those highlighting significant comparisons within female and male datasets are displayed in red and blue boxes, respectively. When significant, the difference compared to the median of the controls is indicated. For all panels, p-values from one-sided Mann-Whitney-Wilcoxon tests are ***p < 0.0001, **p = 0.0043, *p = 0.035, (ns) p = 0.7322.
First, we investigated whether female differentiation requires cellular sex outside the cells expressing the only two known splicing targets of TraF. If so, females masculinised only in dsx- and fru-negative cells should display altered sexual differentiation. To test this prediction, we used our traKO FRT traF allele to specifically express TraF in dsx- and fru-expressing cells (Fig. 3a). We used three sexually distinct characteristics, organ size, body weight, and fertility, to assess the phenotypic sex of the flies59,66,67. Knock-out of tra in dsx- and fru-negative cells resulted in partial masculinisation of organ size (Fig. 3c) and body weight (Fig. 3d). Indeed, females were ~ 10% smaller and lighter. The same manipulation sterilised females (Fig. 3e) due to the presence of atrophic gonads (Fig. 3f). Importantly, the phenotypes observed for the three readouts fully recapitulated tra null mutant phenotypes (Fig. 3c–e). Together, these data establish that a non-canonical TraF-dependent mechanism is essential and controls important aspects of somatic sexual differentiation in dsx- and fru-negative cells.
If cellular sex is a switch mechanism, then males feminised only in dsx- and fru-negative cells should not only lose male identity, but they should also gain female characteristics (Fig. 3b). We used our traF FRT traKO allele to drive traF expression specifically in dsx- and fru-negative cells in males. This specific TraF expression resulted in increased wing size (Fig. 3g) and body weight (Fig. 3h), and induced male sterilisation (Fig. 3i) due to atrophic testes (Fig. 3j). Overexpression of TraF in dsx- and fru-negative cells recapitulated these observations and similarly induced female characteristics in otherwise normal males (Supplementary Fig. S3e–j). Interestingly, performing the reverse experiments, feminising males only in dsx- and fru-expressing cells (Supplementary Fig. S3k) did not affect organ size and body weight (Fig. 3c, d). These results confirm that the sexual identity of dsx- and fru-positive cells are not involved at all in the sex differences studied.
To investigate the mechanism of action of this new non-canonical sex pathway (i.e., independent of fru and dsx), we examined the relative contribution of cell-autonomous versus non-cell-autonomous effects of TraF on the sex differences in adult organ size. When we specifically eliminated or introduced traF function in the wing pouch cells, it significantly impacted wing size (Fig. 3k), mirroring whole-body traF manipulations (Fig. 3c, g). Interestingly, when we expressed traF only in the wing disc of a traF null mutant, it completely restored the standard size of female wings (Fig. 3k) despite all other organs being masculinised in these individuals. In fact, the adult wing was even slightly larger than that of wild-type females. We observed the same effects with both traF knockdown and over-expression, specifically in the wing disc (Fig. 3l). On the other hand, removing traF from all wing disc cells except the pouch didn’t affect wing size (Fig. 3m). These results show that TraF’s influence on the size of adult organs is primarily through cell-autonomous mechanisms. To strengthen our findings, we conducted genetic experiments targeting adipocytes, as a previous report suggested that the expression of TraF in the fat body could impact body size during larval stages in a non-cell-autonomous manner59. As anticipated from our earlier results, eliminating traF function in the fat body had no impact on wing size (Supplementary Fig. S3l). Furthermore, the expression of traF in adipocytes did not restore the size of female null mutants (Supplementary Fig. S3l). Similarly, both fat body-specific traF knockdown and over-expression did not affect wing size (Supplementary Fig. S3m), confirming that TraF controls the size of adult organs through cell-autonomous mechanisms.
Based on our observations, it appears that TraF exerts its splicing activity in a wide range of cells, including some that do not express fru or dsx. The expression of TraF in these cells is necessary and sufficient to shape multiple phenotypic differences between the sexes, through autonomous mechanisms. Indeed, we found that the sexual differentiation of the dsx- and fru-negative cells controls sex differences in organ size, body weight and sex organ formation. This is the first time that the sex of general somatic cells has been implicated in the three phenotypes investigated. It is a common biological strategy to use the sexual identity of equivalent differentiated cells in both sexes to create sex differences in phenotypes. Contrary to the traditional two-gene model of sexual differentiation that only considers dsx- and fru-positive cells, our findings indicate that cellular sex has a crucial role in organs and suggest that an intrinsic sex pathway exists downstream of TraF.
The new TraF-dependent mechanism is active in dsx-expressing cells
Our previous genetic manipulations were restricted to dsx- and fru-negative cells, so we were curious to test whether TraF-dependent sex differentiation is active and operates in dsx- or fru-expressing cells.
We hypothesised that a TraF variant could potentially uncouple canonical and uncharacterised sex differentiation pathways, enabling us to test the latter’s significance in dsx- and fru-cells. Given the high degree of evolutionary divergence in sex-determining gene sequences between species68,69,70, we leveraged natural TraF variations among closely related fly species71 (Fig. 4a). We generated knock-in lines that replaced the coding sequences of D. melanogaster TraF with sequences from two closely related species, D. sechellia and D. virilis. These alleles were then evaluated to determine their ability to regulate various aspects of sexual differentiation under TraF control. D. sechellia protein was able to rescue all aspects of female differentiation, including genitalia formation (Fig. 4b), dsx, and fru splicing (Supplementary Fig. S4a), as well as organ size (Fig.4c) and female fertility (Fig. 4d). However, while the D. virilis protein triggered dsx and fru female-specific splicing (Fig. 4b and Supplementary Fig. S4a) and female size (Fig. 4c), it failed to support female fertility (Fig. 4d). To confirm these findings, we generated two additional tra knock-in lines. One line was a negative control and utilised a more distantly related species, the beetle Tribolium castaneum72. The other line used the TraF protein from D. pseudoobscura, which has a similar degree of divergence from D. melanogaster TraF as the D. virilis protein. The knock-in line from T. castaneum exhibited classical tra loss-of-function morphological phenotypes (Supplementary Fig. S4b), failing to induce female-specific splicing of dsx and fru (Supplementary Fig. S4a), as well as female organ size (Supplementary Fig. S4c) or fertility (Supplementary Fig. S4d). This allele served as a negative control for our cross-species transgenic approach. Interestingly, the D. pseudoobscura protein triggered female-specific splicing of dsx and fru (Supplementary Fig. S4a), as well as female size (Supplementary Fig. S4c), but it could not support female fertility (Supplementary Fig. S4d). This new allele independently replicated our results obtained with D. virilis. As anticipated, our approach identified TraF variants that can regulate dsx and fru splicing but can only partially activate the new TraF-based non-canonical sex differentiation pathway (i.e., independent of fru and dsx). This allows us to test its significance in dsx- and fru-expressing cells.
a TraF protein sequence divergence between Drosophila species and the red flour beetle Tribolium castaneum. b Sex transformations of abdominal pigmentation and genitalia induced by tra knock-out and knock-in of D. sechellia and D. virilis tra genes. c, d Adult wing size (c), and female fertility (d) quantifications following D. sechellia and D. virilis TraF protein expressions. e Diagrams representing the specific cell population with D. virilis tra expression, the allele generated, and the genotypes tested. f Female fertility quantifications following D. virilis TraF protein expression specifically in fru- or dsx-positive cells. Data was combined from at least three independent experiments. n = wing number measured per genotype in (c) and number of flies in (d) and (f). Asterisks highlighting significant comparisons across sexes are displayed in grey boxes at the bottom of graphs; those highlighting significant comparisons within female datasets are displayed in red. For all panels, p-values from one-sided Mann-Whitney-Wilcoxon tests are ***p < 0.0001, (ns) p > 0.066.
To establish how TraF divergence is linked to specific functions, we created a genetic tool to achieve tissue-specific interspecies gene exchange analyses. This transgenic fly line allowed us to replace the wild-type D. melanogaster tra gene with the divergent tra of virilis in any given genetically defined (Gal4-positive) cell population (Fig. 4e). While expression of virilis TraF specifically in fru-positive cells did not affect fertility (Fig. 4f), it sterilised females in dsx-positive cells (Fig. 4f). To rule out a potential effect of altered dsx splicing undetectable in RT-qPCR, we generated a dsxF constitutive allele (dsxF cons). As expected, this new reagent was able to carry out all the crucial functions mediated by DsxF during development and adulthood (Supplementary Fig. S4e-j), which are necessary for female differentiation. Indeed, dsxF cons effectively suppressed the expression of a male-specific reporter73 in the ventral nerve cord (Supplementary Fig. S4h) and simultaneously triggered the female-specific expression of Yp1 in adult adipocytes in both males and females (Supplementary Fig. S4g). It also successfully restored the fertility of dsx null mutant females (Supplementary Fig. S4j). However, this feminising allele did not rescue the sterility of females expressing D. virilis TraF protein (Supplementary Fig. S4j), indicating that a sex pathway downstream of TraF is active and crucial in dsx-cells. We observed that these female flies had normal ovaries with developing egg chambers at all stages, including late vitellogenic stages (Supplementary Fig. S4k). However, they were unable to lay eggs and had swollen abdomens (Fig. 4b). The Hox gene Abdominal-B (Abd-B) is responsible for forming the posterior segments, which include cells involved in egg-laying behaviour, such as the motoneurons that innervate the oviducts74 or the vaginal plate75,76. When D. virilis TraF was expressed specifically in Abd-B-expressing cells, it caused sterility in females (Supplementary Fig. S4l), suggesting that the egg retention phenotype might be due to problems in the sexual differentiation of these somatic cells.
These findings demonstrate that the cellular sex pathway we identified plays a crucial role not only during development in dsx- and fru-negative cells but also in dsx-positive cells. The cross-species transgenic tests provide evidence that the variation in TraF protein sequences between Drosophila species has functional importance and regulates lineage-specific cellular sex differentiation programmes.
Cellular sex requires the splicing activity of TraF and its cofactor Tra2
Previous experiments revealed that D. virilis TraF can splice dsx and fru, but only partially supports the activation of the cellular sex pathway. This led us to wonder if the new targets of TraF were also regulated at the splicing level. To gain further insights into the molecular mechanisms involved in cellular sex specification, we investigated whether TraF splicing activity is required. We therefore focused on the RS domain of TraF and its cofactor Tra2.
To eliminate TraF’s splicing activity, we designed a new tra knock-in allele with RS/SR repeats mutated to alanines. Our findings showed that the absence of female-specific dsx- and fru-splicing events (Supplementary Fig. S5a) caused sex-reversal of genitalia and secondary sexual characteristics (Fig. 5a). We also observed a reduction in organ size (Fig. 5b) and body weight (Fig. 5c) in these masculinised female flies lacking the TraF RS domain. In addition, these pseudo-males were found to be sterile (Fig. 5d). Notably, deletion of the C-terminal proline-rich domain, which is not implicated in the splicing activity of TraF27, did not impact the ability of TraF to drive female organ size, unlike RS domain mutation (Supplementary Fig. S5b). These results highlight that the RS domain is required for TraF splicing activity and its role in cellular sex-driven differentiation.
a Sex transformations of abdominal pigmentation and genitalia induced by tra knock-out and RS domain mutation. b–d Adult wing size (b), adult weight (c), and female fertility (d) quantifications following RS domain mutation. e Expression pattern of tra2-Gal4 reporter in the female tissues at the third larval stage (DNA labelled with DAPI, blue; the nuclear form of GFP, green). f Diagrams representing the specific cell population with tra2 knock-out, the alleles generated, and the genotype used. g–i Adult wing size (g), adult weight (h), and female fertility (i) quantifications following tra2 knock-out specifically in dsx- and fru-negative cells. Data was combined from at least three independent experiments. n = wing number measured per genotype in (b) and (g), number of replicates (each repeat containing five flies) in (c) and (h) and number of flies in (d) and (i). Asterisks highlighting significant comparisons across sexes are displayed in grey boxes at the bottom of graphs; those highlighting significant comparisons within female datasets are displayed in red. For all panels, p-values from one-sided Mann-Whitney-Wilcoxon tests are ***p < 0.0001, (ns) p > 0.99.
We then tested whether Tra2 protein, the binding partner of TraF, is involved in the regulation of cellular sex. To study its expression pattern, we generated a tra2-Gal4 knock-in reporter line (Fig. 5e). Our findings showed that when tra2 knockdown was induced through tra2-Gal4, female-to-male sex reversal transformations occurred in all sexually dimorphic structures controlled by dsx and fru (Supplementary Fig. S5c). This effect could be rescued when tra2-Gal4 was combined with dsx- and fru-Gal80 (Supplementary Fig. S5c). All of our findings were consistent with tra2-Gal4 accurately reporting tra2 expression. We also found that this newly generated reporter was expressed ubiquitously in both sexes (Fig. 5e and Supplementary Fig. S5d), suggesting that the sex differentiation pathway could also rely on Tra2.
In order to evaluate the role of Tra2, we introduced tra2 into cells expressing dsx and fru in tra2 null mutants. This approach allowed us to generate flies that specifically lack Tra2 in cells that do not express the two established splicing targets of TraF. We accomplished this using a tra2 CRISPR null mutant and two lines that reinstated tra2 expression in dsx- and fru-expressing cells (Fig. 5f). These lines were created by exchanging MiMIC lines77 with a trojan exon encoding tra2. This genetic manipulation fully rescued sex-specific splicing of dsx and fru (Supplementary Fig. S5e) as well as genitalia formation (Supplementary Fig. S5f). In addition, we drove tra2 knockdown using the newly generated traF-Gal4 combined with dsx- and fru-Gal80 as an alternative strategy (Supplementary Fig. S5g–k). Both genetic manipulations partially abrogated the sex bias in organ size (Fig. 5g and Supplementary Fig. S5j) and body weight (Fig. 5h and Supplementary Fig. S5k). The tra2 mutation downregulates size and weight in masculinised female flies. Indeed, females were 8% smaller and 11% lighter, whereas males were unaffected. Moreover, we were able to abolish the feminisation of organ size and body weight in TraF overexpressing males by downregulating tra2 expression (Supplementary Fig. S5l). These results suggest that the presence of Tra2 in somatic cells is crucial for the control of organ size and body weight by cellular sex. However, we observed that tra2 mutation (Fig. 5i) and downregulation (Supplementary Fig. S5m) did not affect female fertility when confined to dsx- and fru-negative cells. This indicates that a distinct tra2-independent mechanism is involved in fertility control by cellular sex.
Thus, our data point to the existence of two cellular sex pathways that contribute to sex differences and depend on the RS domain of TraF. One mechanism is tra2-dependent and controls sex-specific differences in organ size and body weight, while the other mechanism is tra2-independent and is essential for female fertility.
Discussion
Our findings invalidate the two-gene model of sexual differentiation in Drosophila and demonstrate that cellular sex is ubiquitous and essential for shaping differences between males and females during development and evolution.
We found that every somatic cell expresses the primary sex determinant, TraF, indicating that all male and female cells are sexually differentiated. It is worth noting that among the thousand Gal4 reporter lines generated in Drosophila over decades, our traF-Gal4 is the first and only sex-specific and ubiquitous line. Obtaining this tool was quite challenging since TraF production is controlled by female-specific alternative splicing of a very short exon, an event easily disrupted by the introduction of additional sequences. The ubiquitous expression of the fly sex determinant has significant implications beyond sex differentiation. TraF is a splicing factor, thus it appears crucial to sex biological samples when studying alternative splicing processes in Drosophila. In the brain, for example, all neuronal and glial cells expressed TraF, and this tissue has the maximum mRNA isoform diversity, but most splicing studies78,79 still use unisex or mixed samples. Besides fundamental research, our observation could be beneficial for genetic engineering. For example, individual sex-sorting remains a crucial issue in male sterile techniques developed to control pests and disease vector insect populations80. Our traF-Gal4 line design could serve as a model for developing reporters that combine fluorescent proteins downstream of the ubiquitous female-specific traF exon. This approach could be adapted to various insect species since TraF is conserved.
While recent studies suggest that non-cell-autonomous mechanisms also contribute to male-female size differences in flies59,81,82,83, our data reveal that intrinsic sex pathways downstream of TraF account for 40% of sex difference in organ size and 30% of the sex gap in body weight. Previous studies have primarily concentrated on analysing the targets and expression sites of dsx and fru7, assuming that these two effectors account for all TraF functions. However, our discoveries emphasise that it is crucial to look beyond these two factors and characterise the complete range of sex-specific isoforms since the overlooked sex pathways depend on the splicing activity of TraF.
We discovered that TraF targets beyond dsx are also crucial for specifying sex organs. Our experiments, involving TraF from various fly species, further confirmed this conclusion. We found that one TraF protein from D. virilis could splice dsx and fru but could not support female reproduction. These findings prove that there are intrinsic cellular sex pathways that differ between closely related species, which operate in dsx-positive cells. Altogether, our experiments demonstrate the global requirement of cellular sex for sexual differentiation and the presence of essential sex pathways. The genetic tools we have developed will help researchers further explore the functional significance of cellular sex in various sex differences related to immunity84, metabolism85, or development85 that have yet to be fully understood.
Our results make Drosophila the first animal where the presence of cellular sex and its functional importance have been tested extensively at the organism level. Could the ubiquitous nature of sex-determinant expression and function be conserved in other animals, including mammals? While the general importance of intrinsic sex pathways remains to be determined, clear examples already highlight the critical role of cellular sex in these animals. In humans, the functional equivalent of TraF is SRY which instructs embryonic gonads to develop into testes86. Interestingly, SRY is also present in various adult tissues outside the sex organs, including the lungs, heart, and liver, where its putative functions and targets remain to be fully explored87. Beyond SRY, while it’s commonly believed that Y-chromosome genes are only expressed in the testis, recent studies have shown elevated Y-chromosome gene expression in non-reproductive tissues88,89,90,91,92. For example, EIF1AY, which encodes an essential translation initiation factor, is more abundant in male heart tissue than its X-linked homologue EIF1AX in female heart tissue at the protein level92. Combined with the recent discoveries linking Y-linked genes to cancer growth and immunotherapy93,94,95, it is becoming increasingly evident that cellular sex also plays an essential role in mammalian development and physiology. Despite these few examples, the global importance of cellular sex in vertebrates at the organism level and its integration with better-characterised hormonal effects remain to be elucidated.
Methods
Fly strains and media
Gal4 drivers
dsx-Gal4 (gift from C. Rezaval, generated by ref. 44, FlyBase ID: FBti0168641), fruP1-Gal4 (BDSC: 66696, FlyBase ID: FBti0168666), mex1-Gal4 (BDSC: 91368, FlyBase ID: FBti0213076), prosV1-Gal4 (BDSC: 84276, FlyBase ID: FBti0010694), Dl-Gal4 (BDSC: 45136, FlyBase ID: FBti0134190), vm-Gal4 (BDSC: 48547, FlyBase ID: FBti0133125), btl-Gal4 (BDSC: 66790, FlyBase ID: FBti0185389), repo-Gal4 (BDSC: 7415, FlyBase ID: FBti0018692), nSyb-Gal4 (BDSC: 51635, FlyBase ID: FBti0150361), tra promoter-Gal4 (this study, see below for details), traF-Gal4 (this study, see below for details), dsx-Gal80 (this study, see below for details), fruP1-Gal80 (this study, see below for details), traF-LexA (this study, see below for details), tra2-Gal4 (this study, see below for details), nubbin-Gal4 (BDSC: 84330, FlyBase ID: FBti0016825), cg-Gal4 (BDSC: 7011, FlyBase ID: FBti0027802), Yolk1-Gal4 (BDSC: 58814, FlyBase ID: FBti0164887), R17G11-Gal4 (BDSC: 49275, FlyBase ID: FBti0133549), Abd-B-Gal4 (BDSC: 55848, FlyBase ID: FBti0074266), pdm2-Gal4 (BDSC: 49828, FlyBase ID: FBtp0057439), tsh-Gal4 (BDSC: 3040, FlyBase ID: FBti0002787).
UAS transgenes
UAS-Stinger (BDSC: 84277, FlyBase ID: FBti0074589), UAS-flp (BDSC: 4539, FlyBase ID: FBti0012284), UAS-traF (BDSC: 4590, FlyBase ID: FBti0010566), UAS-traF (this study, see below for details), UAS-traFΔRS (this study, see below for details), UAS-traFΔP (this study, see below for details), LexAop2-IVS-tdTomato.nls (BDSC:66680, FlyBase ID: FBti0185294), UAS-IVS-mCD8::GFP (BDSC: 32186, FlyBase ID: FBti0131963), UAS-IVS-mCD8::GFP (BDSC: 32185, FlyBase ID: FBti0131931), UAS-dicer2 (VDRC: 60007).
RNAi transgenes
UAS-tra2RNAi (BDSC: 28018, FlyBase ID: FBti0128004), UAS-traRNAi (BDSC: 28512, FlyBase ID: FBti0127269).
Mutants
traKO (BDSC: 67412, FlyBase ID: FBti0186559), traF FRT KO (generated by ref. 63, FlyBase ID: FBti0211844), traKO FRT traF (this study, see below for details), traΔRS (this study, see below for details), tra2KO (this study, see below for details), dsx>tra2 (this study, see below for details), fruP1>tra2 (this study, see below for details), Dsech tra (this study, see below for details), Dvir tra (this study, see below for details), traDmel FRT Dvir (this study, see below for details), dsx Fcons (this study, see below for details), dsx1 (BDSC: 1679, FlyBase ID: FBal0003154), dsxΔ (gift from B. Prudhomme, FlyBase ID: FBal0325111), tra21 (BDSC: 66712, FlyBase ID: FBal0017020), Df(3 L)st-j7 (BDSC: 5416, FlyBase ID: FBab0002416), Dpseudo tra (this study), Tcas tra (this study).
Animals were reared on fly food containing (per liter): 10 g of agar, 82.5 g polenta, 34 g dry yeast and 3,75 g Moldex (per liter, diluted in ethanol). All experimental flies were kept in incubators at 25 °C, and on a 12 hr light/dark cycle. Flies were transferred to fresh vials every 3 days, and fly density was kept to a maximum of 15 flies per vial.
Fertility tests
For fertility experiments, males or females were collected and aged for 3–5 days. For female fertility experiments, females were mated over five days to CantonS males (1 female with 1 male per vial). For male fertility experiments, males were mated over five days to CantonS females (1 male with 3 females per vial). Flies were then removed, and progeny was counted.
Generation of the tra promoter-Gal4 knock-in allele
To generate the tra promoter Gal4 knock-in allele, the 1917 nucleotides (nt) upstream of the tra start codon were cloned into the RIV FRTnMCS1FRT white vector (DGRC: Plasmid#1333) using the NotI and NheI restriction sites by gene synthesis (Genscript). The Gal4 sequences and the 3’UTR of tra, along with the 967 downstream nt, were added by PCR between the NheI and XhoI sites. The construct was sequence-verified, and a transgenic line was established through ΦC-31 integrase mediated transformation (Bestgene), using a recently generated amorphic allele of tra63 in which tra locus has been replaced by an attP site (BDSC: 67412, FlyBase ID: FBti0186559). The generated allele behaves as a tra null mutant.
Generation of the traF-Gal4 and traF-LexAp65 knock-in alleles
To generate the traF-Gal4 and traF-LexAp65 knock-in alleles, the T2A-Gal4 sequences (Addgene: Plasmid #62893) or the T2A-nls-LexA::p65 sequences (Addgene: Plasmid #26230) were cloned upstream of the tra STOP codon into an RIV FRTnMCS1FRT white vector (DGRC: Plasmid#1333, containing the tra locus (3869 nt containing: tra coding region, 1910 nt upstream and the 967 nt downstream63). The constructs were sequence-verified and transgenic lines were established through ΦC-31 integrase mediated transformation (Bestgene), using a recently generated amorphic allele of tra63 in which tra locus has been replaced by an attP site (BDSC: 67412, FlyBase ID: FBti0186559).
Generation of the traF KO -Gal4 knock-in allele
To generate the traF KO-Gal4 knock-in allele, the T2A-Gal4 sequences (Addgene: Plasmid #62893) were cloned 26 nt downstream of the tra female-specific exon into a RIV FRTnMCS1FRT white vector (DGRC: Plasmid#1333, containing the tra locus (674 nt containing: tra coding region deleted of 582 nt of the female-specific exons, the 353 nt upstream and the 310 nt downstream63). The constructs were sequence-verified and transgenic lines were established through ΦC-31 integrase mediated transformation (Bestgene), using a recently generated amorphic allele of tra63 in which tra locus has been replaced by an attP site (BDSC: 67412, FlyBase ID: FBti0186559).
Generation of the dsx-Gal80 knock-in allele
To generate a knock-in Gal80 under the control of dsx regulatory sequences, recombination-mediated cassette exchange of the following insertion was performed: Mi{y[+mDint2]=MIC}dsx[MI03050] (BDSC: 36182, FlyBase ID: FBti0143242). The swapping strategy was previously described in ref. 77. Midiprep plasmid DNA of pBS-KS-attB2-SA(1)-T2A-3X Gal80-Hsp70 (addgene: Plasmid#62952) was injected together with ΦC31 plasmid DNA into the embryos of flies bearing MI03050. The orientation of the Gal80 exon inserts was then determined by PCR amplification.
Generation of the fru-Gal80 knock-in allele
To generate a knock-in Gal80 under the control of fru regulatory sequences (P1 promoter), recombination-mediated cassette exchange of the following insertion was performed: Mi{y[+mDint2]=MIC}fru[MI05459] (BDSC: 42086, FlyBase ID: FBti0149046). Midiprep plasmid DNA of pBS-KS-attB2-SA(1)-T2A-3X Gal80-Hsp70 (addgene: Plasmid#62952) was injected together with ΦC31 plasmid DNA into the embryos of flies bearing MI05459. The orientation of the Gal80 exon inserts was then determined by PCR amplification.
Generation of the UAS-traF lines
To generate a wild type UAS-traF line, tra coding regions (687 nt, FlyBase ID: FBtr0075364) were cloned by PCR into the pUASt-attB vector (DGRC: Plasmid#1419) between the EcoRI and XhoI restrictions sites. In the UAS-traFΔRS line, the 17 RS/SR repeats are mutated in alanine (RS/SR > AA). In the UAS-traFΔP line, the 35 last amino-acids of tra are deleted. The constructs were sequence-verified and transgenic lines were established through ΦC-31 integrase mediated transformation (Bestgene), using the attP40 (BDSC: 36304, FlyBase ID: FBti0114379) attP site line.
Generation of the tra KO FRT traF knock-in allele
To generate the traKO FRT traF knock-in allele, the following sequences were cloned into the RIV FRTnMCS1FRT white vector (DGRC: Plasmid#1333) using the EcoRI and BglII restriction sites by gene synthesis (Genscript): tra promoter-truncated tra cDNA –FRT site – traF cDNA. The construct was sequence-verified, and transgenic lines were established through ΦC-31 integrase mediated transformation (Bestgene), using a recently generated amorphic allele of tra63 in which tra locus has been replaced by an attP site (BDSC: 67412, FlyBase ID: FBti0186559).
Generation of the D. virilis tra knock-in allele
To generate the D. virilis tra knock-in allele, the tra locus (tra coding region, the 353 nt upstream and the 310 nt downstream) was cloned by gene synthesis (Genscript) into the RIV FRTnMCS1FRT white vector (DGRC: Plasmid#1333) using the EcoRI and XhoI restriction sites. The tra-coding regions were replaced with the tra-coding regions of D. virilis. The introns, 5’UTR, and 3’UTR were not swapped. The construct was sequence-verified, and a transgenic line was established through ΦC-31 integrase-mediated transformation (Bestgene) using a recently generated amorphic allele of tra63 in which the tra locus has been replaced by an attP site (BDSC: 67412).
Generation of the D. sechellia tra knock-in allele
To generate the D. sechellia tra knock-in allele, the tra locus (tra coding region, the 353 nt upstream and the 310 nt downstream) was cloned by gene synthesis (Genscript) into the RIV FRTnMCS1FRT white vector (DGRC: Plasmid#1333) using the EcoRI and XhoI restriction sites. The tra-coding regions were replaced with the tra-coding regions of D. sechellia. The introns, 5’UTR, and 3’UTR were not swapped. The construct was sequence-verified, and a transgenic line was established through ΦC-31 integrase mediated transformation (Bestgene) using a recently generated amorphic allele of tra63 in which tra locus has been replaced by an attP site (BDSC: 67412).
Generation of the D. pseudoobscura tra knock-in allele
To generate the D. pseudoobscura tra knock-in allele, the tra locus (tra coding region, the 353 nt upstream and the 310 nt downstream) was cloned by gene synthesis (Genscript) into the RIV FRTnMCS1FRT white vector (DGRC: Plasmid#1333) using the EcoRI and NheI restriction sites. The tra-coding regions were replaced with the tra-coding regions of D. pseudoobscura. The introns, 5’UTR, and 3’UTR were not swapped. The construct was sequence-verified, and a transgenic line was established through ΦC-31 integrase mediated transformation (Bestgene) using a recently generated amorphic allele of tra63 in which tra locus has been replaced by an attP site (BDSC: 67412).
Generation of the T. castaneum tra knock-in allele
To generate the T. castaneum tra knock-in allele, the tra locus (tra coding region, the 353 nt upstream and the 310 nt downstream) was cloned by gene synthesis (Genscript) into the RIV FRTnMCS1FRT white vector (DGRC: Plasmid#1333) using the EcoRI and XhoI restriction sites. The tra-coding regions were replaced with the tra-coding regions of T. castaneum. The introns, 5’UTR, and 3’UTR were not swapped. The construct was sequence-verified, and a transgenic line was established through ΦC-31 integrase mediated transformation (Bestgene) using a recently generated amorphic allele of tra63 in which tra locus has been replaced by an attP site (BDSC: 67412).
Generation of the tra Dmel FRT Dvir knock-in allele
To generate the traDmel FRT Dvir knock-in allele, the following sequences were cloned into the RIV FRTnMCS1FRT white vector (DGRC: Plasmid#1333) using the EcoRI and XhoI restriction sites by gene synthesis (Genscript): D. melanogaster tra locus–FRT site – D. virilis tra locus. The construct was sequence-verified, and transgenic lines were established through ΦC-31 integrase-mediated transformation (Bestgene) using a recently generated amorphic allele of tra63 in which tra locus has been replaced by an attP site (BDSC: 67412, FlyBase ID: FBti0186559).
Generation of the dsxF cons knock-in allele
To generate a dsxFcons knock-in, recombination-mediated cassette exchange of the following insertion was performed: Mi{y[+mDint2]=MIC}dsx[MI03050] (BDSC: 36182, FlyBase ID: FBti0143242). The pBS-KS-attB2-SA(1)-T2A-3X Gal80-Hsp70 (addgene: Plasmid#62952) vector was modified. The last common exon of dsx followed by the female-specific exon and the SV40 poly(A) replaced the Gal80 sequences, and a 3xP3 promoter driving DsRed was added to facilitate the knock-in event screening. Midiprep plasmid DNA of this construct was injected together with ΦC31 plasmid DNA into the embryos of flies bearing MI03050. The orientation of the tra2 exon inserts was then determined by PCR amplification.
Generation of the tra ΔRS knock-in allele
To generate the traΔRS knock-in allele, the tra locus (1648 nt containing: tra coding region, the 353 nt upstream and the 310 nt downstream) was cloned by gene synthesis (Genscript) into the RIV FRTnMCS1FRT white vector (DGRC: Plasmid#1333) using the EcoRI and XhoI restriction sites. In the tra coding region, the 17 RS/SR repeats are mutated in alanine (RS > AA). The construct was sequence-verified, and a transgenic line was established through ΦC-31 integrase mediated transformation (Bestgene) using a recently generated amorphic allele of tra63 in which tra locus has been replaced by an attP site (BDSC: 67412).
Generation of the tra2 KO CRISPR null mutant
To generate a tra2 null mutant, two gRNAs targeting the tra2 coding sequence (gRNA 1: tcgatgcttcatgtcaaaagagg, and gRNA 2: gatgaagttcgacctaatcgtgg) were cloned into the pCFD5 vector (Addgene: Plasmid #73914). A 1 kb homology arm flanking the cleavage site 1 was PCR-amplified from genomic DNA using the Q5 high-fidelity polymerase from New England Biolabs (M0491S). The PCR product was digested with NheI and SacII prior to cloning into the pDsRedattP vector (Addgene: Plasmid #51019). A 1 kb homology arm flanking the cleavage site 2 was PCR-amplified from genomic DNA. The PCR product was digested with AvrII and XhoI prior to cloning into the pDsRedattP vector, containing the first homology arm. The constructs were sequence-verified and a mutant line was established through injection (Bestgene) of the 2 generated vectors (pCFD5 gRNAs and pDsRedattP homology arms) in yw;nos-Cas9 (FlyBase ID: FBti0156858) embryos. The generated deletion removed 1513 nt of the tra2 coding sequence and replaced it with an attP landing site and a loxP-flanked 3xP3-DsRed marker.
Generation of the tra2-Gal4 knock-in allele
To generate the tra2-Gal4 knock-in allele, the 589 nt of the tra2 locus (containing half of the second intron and the third exon) were cloned into the RIV FRTnMCS1FRT white vector (DGRC: Plasmid#1333) using the EcoRI and AsiSI restriction sites by gene synthesis (Genscript). The T2A-Gal4 sequences (addgene: Plasmid #62893), along the SV40 poly(A), were added by PCR between the AsiSI and XhoI sites. The construct was sequence-verified, and a transgenic line was established through ΦC-31 integrase mediated transformation (Bestgene), using a newly generated amorphic allele of tra2KO in which tra2 locus has been replaced by an attP site. The generated allele behaves as tra2 null mutant.
Generation of the fru-tra2 knock-in allele
To generate a knock-in of tra2 under the control of fru regulatory sequences (P1 promoter), recombination-mediated cassette exchange of the following insertion was performed: Mi{y[+mDint2]=MIC}fru[MI05459] (BDSC: 42086, FlyBase ID: FBti0149046). The pBS-KS-attB2-SA(1)-T2A-3X Gal80-Hsp70 (addgene: Plasmid#62952) vector was modified. The tra2 cDNA replaced the Gal80 sequences, and a 3xP3 promoter driving DsRed was added to facilitate the knock-in event screening. Midiprep plasmid DNA of this construct was injected together with ΦC31 plasmid DNA into the embryos of flies bearing MI05459. The orientation of the tra2 exon inserts was then determined by PCR amplification.
Generation of the dsx-tra2 knock-in allele
To generate a knock-in of tra2 under the control of dsx regulatory sequences, recombination-mediated cassette exchange of the following insertion was performed: Mi{y[+mDint2]=MIC}dsx[MI03050] (BDSC: 36182, FlyBase ID: FBti0143242). The pBS-KS-attB2-SA(1)-T2A-3X Gal80-Hsp70 (addgene: Plasmid#62952) vector was modified. The tra2 cDNA replaced the Gal80 sequences, and a 3xP3 promoter driving DsRed was added to facilitate the knock-in event screening. Midiprep plasmid DNA of this construct was injected together with ΦC31 plasmid DNA into the embryos of flies bearing MI03050. The orientation of the tra2 exon inserts was then determined by PCR amplification.
Immunohistochemistry
Larval and adult tissues were dissected in 1xPBS, placed on poly-L-lysine (Sigma-Aldrich, P1524-1G) coated slides, fixed in 3.7% formaldehyde (Polyscience) in 1xPBS for 20 min at room temperature (RT) and then washed several times in PBS containing 0,3% Triton X-100 (PBT). Dissected tissues were blocked in PBT + 4% Horse serum (HS) at RT for at least one h. The primary antibodies incubation was performed in PBT + HS for 48 h at 4 °C. After several washes, secondary antibodies were incubated for two hours at RT. Then, dissected tissues were mounted into Vectashield with DAPI (Vector Labs) to stain DNA. Fluorescence images were acquired using a Leica SP5 DS confocal microscope. The following primary antibodies (see also Table S1) were used: chicken anti-GFP (1/10000) (ab13970 Abcam), goat anti-tdTomato (1/500) (LS-C340696, LSBio).
Wing size
Adult flies were collected in ethanol at least 12 hours following emergence to ensure the wings were fully expanded. Right wings were dissected in ethanol and mounted in a drop of Euparal mounting medium (Carl Roth #7356.1). Slides were dried overnight on a heating plate at 60 °C. Then, wings were imaged using a Leica M205 FA associated with Leica DFC7000T camera. Wing areas were quantified using ImageJ by manually selecting the Cartesian coordinates of six landmarks representing junctions of veins with the wing contour, then measuring the number of pixels included in the resulting outline (method adapted from96). For a given experiment, all values were normalised to one control condition.
Body weight
2–5 days old male and female flies were frozen at −20 °C. During the two following days, flies were weighed by groups of 5 adults on an XPR analytical balance (Mettler Toledo). For a given experiment, all values were normalised to one control condition.
Reverse transcription and quantitative-PCR
RNAs were extracted from 10 whole flies (or 15 heads) using TRIzol (Invitrogen). RNAs were cleaned using an RNAeasy mini Kit (Qiagen, 74-104). cDNAs were synthesised using the iScript cDNA synthesis kit (Bio-Rad, 170-8889) from 500 ng of total RNAs. Quantitative PCRs were performed by mixing cDNA samples (5 ng) with iTaq Universal SYBR® Green Supermix (Bio-Rad, 172-5124) and the relevant primers in 384-well plates. Expression abundance was calculated using a standard curve for each gene and normalised to the expression of the rp49 control gene. For data display purposes, the median of the expression abundance was arbitrarily set at 100% for control males or females, and the percentage of that expression is displayed for all the tested genotypes. qPCR primer pairs used are listed in table S1.
Statistics and data presentation
All statistical analyses were carried out in GraphPad Prism 7.04. Comparisons between two genotypes or conditions were analysed with the Mann-Whitney-Wilcoxon rank sum test. This non-parametric test does not require the assumption of normal distributions, so no methods were used to determine whether the data met such assumptions. All graphs were generated using GraphPad Prism 7.04. In all figures, when significant, the differences between the medians of the control and the tested conditions are indicated.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data is available in the main text or the supplementary materials. Materials generated for the study are available from the corresponding author on request. Source data are provided with this paper.
References
Arnold, A. P. Rethinking sex determination of non-gonadal tissues. Curr. Top. Dev. Biol. 134, 289–315 (2019).
Arnold, A. P. Sexual differentiation of brain and other tissues: five questions for the next 50 years. Horm. Behav. 120, 104691 (2020).
Mauvais-Jarvis, F., Arnold, A. P. & Reue, K. A guide for the design of pre-clinical studies on sex differences in metabolism. Cell Metab. 25, 1216–1230 (2017).
García, L. R. & Portman, D. S. Neural circuits for sexually dimorphic and sexually divergent behaviors in Caenorhabditis elegans. Curr. Opin. Neurobiol. 38, 46–52 (2016).
Meyer, B. J. Mechanisms of sex determination and X-chromosome dosage compensation. Genetics 220, iyab197 (2022).
Hopkins, B. R. & Kopp, A. Evolution of sexual development and sexual dimorphism in insects. Curr. Opin. Genet Dev. 69, 129–139 (2021).
Camara, N., Whitworth, C. & Van Doren, M. The creation of sexual dimorphism in the Drosophila soma. Curr. Top. Dev. Biol. 83, 65–107 (2008).
Christiansen, A. E., Keisman, E. L., Ahmad, S. M. & Baker, B. S. Sex comes in from the cold: the integration of sex and pattern. Trends Genet 18, 510–516 (2002).
Sturtevant, A. H. A gene in drosophila melanogaster that transforms females into males. Genetics 30, 297–299 (1945).
Baker, B. S. & Ridge, K. A. Sex and the single cell. I. On the action of major loci affecting sex determination in Drosophila melanogaster. Genetics 94, 383–423 (1980).
Belote, J. M. et al. Control of sexual differentiation in Drosophila melanogaster. Cold Spring Harb. Symp. Quant. Biol. 50, 605–614 (1985).
Butler, B., Pirrotta, V., Irminger-Finger, I. & Nöthiger, R. The sex-determining gene tra of Drosophila: molecular cloning and transformation studies. EMBO J. 5, 3607–3613 (1986).
McKeown, M., Belote, J. M. & Baker, B. S. A molecular analysis of transformer, a gene in Drosophila melanogaster that controls female sexual differentiation. Cell 48, 489–499 (1987).
Boggs, R. T., Gregor, P., Idriss, S., Belote, J. M. & McKeown, M. Regulation of sexual differentiation in D. melanogaster via alternative splicing of RNA from the transformer gene. Cell 50, 739–747 (1987).
McKeown, M., Belote, J. M. & Boggs, R. T. Ectopic expression of the female transformer gene product leads to female differentiation of chromosomally male Drosophila. Cell 53, 887–895 (1988).
Ryner, L. C. & Baker, B. S. Regulation of doublesex pre-mRNA processing occurs by 3’-splice site activation. Genes Dev. 5, 2071–2085 (1991).
Hoshijima, K., Inoue, K., Higuchi, I., Sakamoto, H. & Shimura, Y. Control of doublesex alternative splicing by transformer and transformer-2 in Drosophila. Science 252, 833–836 (1991).
Inoue, K., Hoshijima, K., Higuchi, I., Sakamoto, H. & Shimura, Y. Binding of the Drosophila transformer and transformer-2 proteins to the regulatory elements of doublesex primary transcript for sex-specific RNA processing. Proc. Natl Acad. Sci. USA 89, 8092–8096 (1992).
Tian, M. & Maniatis, T. Positive control of pre-mRNA splicing in vitro. Science 256, 237–240 (1992).
Tian, M. & Maniatis, T. A splicing enhancer complex controls alternative splicing of doublesex pre-mRNA. Cell 74, 105–114 (1993).
Amrein, H., Hedley, M. L. & Maniatis, T. The role of specific protein-RNA and protein-protein interactions in positive and negative control of pre-mRNA splicing by Transformer 2. Cell 76, 735–746 (1994).
Wu, J. Y. & Maniatis, T. Specific interactions between proteins implicated in splice site selection and regulated alternative splicing. Cell 75, 1061–1070 (1993).
Tian, M. & Maniatis, T. A splicing enhancer exhibits both constitutive and regulated activities. Genes Dev. 8, 1703–1712 (1994).
Lynch, K. W. & Maniatis, T. Synergistic interactions between two distinct elements of a regulated splicing enhancer. Genes Dev. 9, 284–293 (1995).
Lynch, K. W. & Maniatis, T. Assembly of specific SR protein complexes on distinct regulatory elements of the Drosophila doublesex splicing enhancer. Genes Dev. 10, 2089–2101 (1996).
Heinrichs, V., Ryner, L. C. & Baker, B. S. Regulation of sex-specific selection of fruitless 5’ splice sites by transformer and transformer-2. Mol. Cell Biol. 18, 450–458 (1998).
Sciabica, K. S. & Hertel, K. J. The splicing regulators Tra and Tra2 are unusually potent activators of pre-mRNA splicing. Nucleic Acids Res 34, 6612–6620 (2006).
Zuo, P. & Maniatis, T. The splicing factor U2AF35 mediates critical protein-protein interactions in constitutive and enhancer-dependent splicing. Genes Dev. 10, 1356–1368 (1996).
Dauwalder, B. & Mattox, W. Analysis of the functional specificity of RS domains in vivo. EMBO J. 17, 6049–6060 (1998).
Baker, B. S. & Wolfner, M. F. A molecular analysis of doublesex, a bifunctional gene that controls both male and female sexual differentiation in Drosophila melanogaster. Genes Dev. 2, 477–489 (1988).
Burtis, K. C. & Baker, B. S. Drosophila doublesex gene controls somatic sexual differentiation by producing alternatively spliced mRNAs encoding related sex-specific polypeptides. Cell 56, 997–1010 (1989).
Ito, H. et al. Sexual orientation in Drosophila is altered by the satori mutation in the sex-determination gene fruitless that encodes a zinc finger protein with a BTB domain. Proc. Natl Acad. Sci. USA 93, 9687–9692 (1996).
Ryner, L. C. et al. Control of male sexual behavior and sexual orientation in Drosophila by the fruitless gene. Cell 87, 1079–1089 (1996).
Lee, G. et al. Spatial, temporal, and sexually dimorphic expression patterns of the fruitless gene in the Drosophila central nervous system. J. Neurobiol. 43, 404–426 (2000).
Goodwin, S. F. et al. Aberrant splicing and altered spatial expression patterns in fruitless mutants of Drosophila melanogaster. Genetics 154, 725–745 (2000).
Demir, E. & Dickson, B. J. fruitless splicing specifies male courtship behavior in Drosophila. Cell 121, 785–794 (2005).
Manoli, D. S. et al. Male-specific fruitless specifies the neural substrates of Drosophila courtship behaviour. Nature 436, 395–400 (2005).
Kimura, K.-I., Ote, M., Tazawa, T. & Yamamoto, D. Fruitless specifies sexually dimorphic neural circuitry in the Drosophila brain. Nature 438, 229–233 (2005).
Shirangi, T. R., Taylor, B. J. & McKeown, M. A double-switch system regulates male courtship behavior in male and female Drosophila melanogaster. Nat. Genet 38, 1435–1439 (2006).
Rideout, E. J., Billeter, J.-C. & Goodwin, S. F. The sex-determination genes fruitless and doublesex specify a neural substrate required for courtship song. Curr. Biol. 17, 1473–1478 (2007).
Clough, E. et al. Sex- and tissue-specific functions of Drosophila doublesex transcription factor target genes. Dev. Cell 31, 761–773 (2014).
Sato, K. & Yamamoto, D. Molecular and cellular origins of behavioral sex differences: a tiny little fly tells a lot. Front Mol. Neurosci. 16, 1284367 (2023).
Nojima, T. et al. A sex-specific switch between visual and olfactory inputs underlies adaptive sex differences in behavior. Curr. Biol. 31, 1175–1191.e6 (2021).
Rideout, E. J., Dornan, A. J., Neville, M. C., Eadie, S. & Goodwin, S. F. Control of sexual differentiation and behavior by the doublesex gene in Drosophila melanogaster. Nat. Neurosci. 13, 458–466 (2010).
Burtis, K. C., Coschigano, K. T., Baker, B. S. & Wensink, P. C. The doublesex proteins of Drosophila melanogaster bind directly to a sex-specific yolk protein gene enhancer. EMBO J. 10, 2577–2582 (1991).
Shirangi, T. R., Dufour, H. D., Williams, T. M. & Carroll, S. B. Rapid evolution of sex pheromone-producing enzyme expression in Drosophila. PLoS Biol. 7, e1000168 (2009).
Camara, N., Whitworth, C., Dove, A. & Van Doren, M. Doublesex controls specification and maintenance of the gonad stem cell niches in Drosophila. Development 146, dev170001 (2019).
Robinett, C. C., Vaughan, A. G., Knapp, J.-M. & Baker, B. S. Sex and the single cell. II. There is a time and place for sex. PLoS Biol. 8, e1000365 (2010).
Pan, Y., Robinett, C. C. & Baker, B. S. Turning males on: activation of male courtship behavior in Drosophila melanogaster. PLoS One 6, e21144 (2011).
Stockinger, P., Kvitsiani, D., Rotkopf, S., Tirián, L. & Dickson, B. J. Neural circuitry that governs Drosophila male courtship behavior. Cell 121, 795–807 (2005).
Belote, J. M., Handler, A. M., Wolfner, M. F., Livak, K. J. & Baker, B. S. Sex-specific regulation of yolk protein gene expression in Drosophila. Cell 40, 339–348 (1985).
Bownes, M., Scott, A. & Blair, M. The use of an inhibitor of protein synthesis to investigate the roles of ecdysteroids and sex-determination genes on the expression of the genes encoding the Drosophila yolk proteins. Development 101, 931–941 (1987).
Grmai, L., Hudry, B., Miguel-Aliaga, I. & Bach, E. A. Chinmo prevents transformer alternative splicing to maintain male sex identity. PLoS Genet 14, e1007203 (2018).
Peng, Q., Chen, J. & Pan, Y. From fruitless to sex: On the generation and diversification of an innate behavior. Genes Brain Behav. 20, e12772 (2021).
Chen, J. et al. fruitless tunes functional flexibility of courtship circuitry during development. Elife 10, e59224 (2021).
Peng, Q. et al. The sex determination gene doublesex is required during adulthood to maintain sexual orientation. J. Genet Genomics 49, 165–168 (2022).
Bayer, E. A., Sun, H., Rafi, I. & Hobert, O. Temporal, spatial, sexual and environmental regulation of the master regulator of sexual differentiation in C. elegans. Curr. Biol. 30, 3604–3616.e3 (2020).
Lawson, H. N., Wexler, L. R., Wnuk, H. K. & Portman, D. S. Dynamic, non-binary specification of sexual state in the c. elegans nervous system. Curr. Biol. 30, 3617–3623.e3 (2020).
Rideout, E. J., Narsaiya, M. S. & Grewal, S. S. The sex determination gene transformer regulates male-female differences in drosophila body size. PLoS Genet 11, e1005683 (2015).
Wat, L. W., Chowdhury, Z. S., Millington, J. W., Biswas, P. & Rideout, E. J. Sex determination gene transformer regulates the male-female difference in Drosophila fat storage via the adipokinetic hormone pathway. Elife 10, e72350 (2021).
Regan, J. C. et al. Sex difference in pathology of the ageing gut mediates the greater response of female lifespan to dietary restriction. Elife 5, e10956 (2016).
Hudry, B., Khadayate, S. & Miguel-Aliaga, I. The sexual identity of adult intestinal stem cells controls organ size and plasticity. Nature 530, 344–348 (2016).
Hudry, B. et al. Sex differences in intestinal carbohydrate metabolism promote food intake and sperm maturation.Cell 178, 901–918.e16 (2019).
Ma, J. & Ptashne, M. The carboxy-terminal 30 amino acids of GAL4 are recognized by GAL80. Cell 50, 137–142 (1987).
Baena-Lopez, L. A., Alexandre, C., Mitchell, A., Pasakarnis, L. & Vincent, J.-P. Accelerated homologous recombination and subsequent genome modification in Drosophila. Development 140, 4818–4825 (2013).
Cline, T. W. Autoregulatory functioning of a Drosophila gene product that establish es and maintains the sexually determined state. Genetics 107, 231–277 (1984).
Testa, N. D., Ghosh, S. M. & Shingleton, A. W. Sex-specific weight loss mediates sexual size dimorphism in Drosophila melanogaster. PLoS One 8, e58936 (2013).
Geuverink, E. & Beukeboom, L. W. Phylogenetic distribution and evolutionary dynamics of the sex determination genes doublesex and transformer in insects. Sex. Dev. 8, 38–49 (2014).
Whitfield, L. S., Lovell-Badge, R. & Goodfellow, P. N. Rapid sequence evolution of the mammalian sex-determining gene SRY. Nature 364, 713–715 (1993).
Pamilo, P. & O’Neill, R. J. Evolution of the Sry genes. Mol. Biol. Evol. 14, 49–55 (1997).
O’Neil, M. T. & Belote, J. M. Interspecific comparison of the transformer gene of Drosophila reveals an unusually high degree of evolutionary divergence. Genetics 131, 113–128 (1992).
Shukla, J. N. & Palli, S. R. Sex determination in beetles: production of all male progeny by parental RNAi knockdown of transformer. Sci. Rep. 2, 602 (2012).
Meissner, G. W., Luo, S. D., Dias, B. G., Texada, M. J. & Baker, B. S. Sex-specific regulation of Lgr3 in Drosophila neurons. Proc. Natl Acad. Sci. USA 113, E1256–E1265 (2016).
Bussell, J. J., Yapici, N., Zhang, S. X., Dickson, B. J. & Vosshall, L. B. Abdominal-B neurons control Drosophila virgin female receptivity. Curr. Biol. 24, 1584–1595 (2014).
Estrada, B. & Sánchez-Herrero, E. The Hox gene Abdominal-B antagonizes appendage development in the genital disc of Drosophila. Development 128, 331–339 (2001).
Foronda, D., Estrada, B., de Navas, L. & Sánchez-Herrero, E. Requirement of Abdominal-A and Abdominal-B in the developing genitalia of Drosophila breaks the posterior downregulation rule. Development 133, 117–127 (2006).
Diao, F. et al. Plug-and-play genetic access to drosophila cell types using exchangeable exon cassettes. Cell Rep. 10, 1410–1421 (2015).
Zhang, Z., Bae, B., Cuddleston, W. H. & Miura, P. Coordination of alternative splicing and alternative polyadenylation revealed by targeted long read sequencing. Nat. Commun. 14, 5506 (2023).
Alfonso-Gonzalez, C. et al. Sites of transcription initiation drive mRNA isoform selection. Cell 186, 2438–2455.e22 (2023).
Zheng, X. et al. Incompatible and sterile insect techniques combined eliminate mosquitoes. Nature 572, 56–61 (2019).
Millington, J. W. et al. Female-biased upregulation of insulin pathway activity mediates the sex difference in Drosophila body size plasticity. Elife 10, e58341 (2021).
Sawala, A. & Gould, A. P. The sex of specific neurons controls female body growth in Drosophila. PLoS Biol. 15, e2002252 (2017).
Sawala, A. & Gould, A. P. Sex-lethal in neurons controls female body growth in Drosophila. Fly. (Austin) 12, 133–141 (2018).
Belmonte, R. L., Corbally, M.-K., Duneau, D. F. & Regan, J. C. Sexual Dimorphisms in Innate Immunity and Responses to Infection in Drosophila melanogaster. Front Immunol. 10, 3075 (2019).
Mank, J. E. & Rideout, E. J. Developmental mechanisms of sex differences: from cells to organisms. Development 148, dev199750 (2021).
Goodfellow, P. N. & Lovell-Badge, R. SRY and sex determination in mammals. Annu Rev. Genet 27, 71–92 (1993).
Lee, J. & Harley, V. R. The male fight-flight response: a result of SRY regulation of catecholamines? Bioessays 34, 454–457 (2012).
Bellott, D. W. et al. Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 508, 494–499 (2014).
Trabzuni, D. et al. Widespread sex differences in gene expression and splicing in the adult human brain. Nat. Commun. 4, 2771 (2013).
Xu, J., Burgoyne, P. S. & Arnold, A. P. Sex differences in sex chromosome gene expression in mouse brain. Hum. Mol. Genet 11, 1409–1419 (2002).
Johansson, M. M. et al. Spatial sexual dimorphism of X and Y homolog gene expression in the human central nervous system during early male development. Biol. Sex. Differ. 7, 5 (2016).
Godfrey, A. K. et al. Quantitative analysis of Y-Chromosome gene expression across 36 human tissues. Genome Res 30, 860–873 (2020).
Qi, M., Pang, J., Mitsiades, I., Lane, A. A. & Rheinbay, E. Loss of chromosome Y in primary tumors. Cell S0092-8674(23)00646–3 https://doi.org/10.1016/j.cell.2023.06.006 (2023).
Li, J. et al. Histone demethylase KDM5D upregulation drives sex differences in colon cancer. Nature 619, 632–639 (2023).
Abdel-Hafiz, H. A. et al. Y chromosome loss in cancer drives growth by evasion of adaptive immunity. Nature 619, 624–631 (2023).
Trotta, V., Calboli, F. C. F., Ziosi, M. & Cavicchi, S. Fitness variation in response to artificial selection for reduced cell area, cell number and wing area in natural populations of Drosophila melanogaster. BMC Evol. Biol. 7, S10 (2007).
Acknowledgements
We thank Carolina Rezaval, and Benjamin Prud’homme for fly lines; the Bloomington Drosophila Stock Centre for providing Drosophila fly lines; the iBV platforms: Baptiste Monterroso and Sameh Ben Aicha from the imaging facility, Florence Besse and Romain Levayer for comments and Anita Mencser for fly food; and all the members of the BH laboratory for fruitful discussions and comments. We would like to express our gratitude to Irene Miguel-Aliaga for her support at the start of the project. This work was supported by the Université Côte d’Azur, CNRS (ATIP-Avenir programme), INSERM, European Research Council (ERC starting grant CellSex, Grant number: ERC-2019-STG#850934), and the French Government (National Research Agency, ANR) through the “Investments for the Future” programmes LABEX SIGNALIFE ANR-11-LABX-0028-01 and IDEX UCAJedi ANR-15-IDEX-01.
Author information
Authors and Affiliations
Contributions
Conceptualisation: C.H. and B.H. Methodology: C.H. and B.H. Investigation: C.H. and T.P. Visualisation: C.H. Funding acquisition: B.H. Project administration: B.H. Supervision: B.H. Writing – original draft: B.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hérault, C., Pihl, T. & Hudry, B. Cellular sex throughout the organism underlies somatic sexual differentiation. Nat Commun 15, 6925 (2024). https://doi.org/10.1038/s41467-024-51228-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-51228-6
- Springer Nature Limited