Abstract
How prostate cancer cells and their precursors mediate changes in the tumor microenvironment (TME) to drive prostate cancer progression is unclear, in part due to the inability to longitudinally study the disease evolution in human tissues. To overcome this limitation, we perform extensive single-cell RNA-sequencing (scRNA-seq) and molecular pathology of the comparative biology between human prostate cancer and key stages in the disease evolution of a genetically engineered mouse model (GEMM) of prostate cancer. Our studies of human tissues reveal that cancer cell-intrinsic activation of MYC signaling is a common denominator across the well-known molecular and pathological heterogeneity of human prostate cancer. Cell communication network and pathway analyses in GEMMs show that MYC oncogene-expressing neoplastic cells, directly and indirectly, reprogram the TME during carcinogenesis, leading to a convergence of cell state alterations in neighboring epithelial, immune, and fibroblast cell types that parallel key findings in human prostate cancer.
Similar content being viewed by others
Introduction
Like many human cancers, prostate cancer is a highly heterogeneous disease with respect to clinical, pathological, and molecular features. In particular, the ever-growing body of genomic and molecular studies of human prostate cancer has revealed numerous and heterogeneous molecular subtypes and their associations with clinicopathological features1,2,3.
Despite this heterogeneity, the early stages of prostate cancer development appear to have some convergent patterns, involving transitions from benign glands, to precursor lesions called prostatic intraepithelial neoplasia (PIN), to invasive carcinoma4,5. Luminal epithelial cells in PIN lesions exist within the confines of a normal glandular architecture but display molecular and cellular alterations characteristic of invasive carcinoma cells, albeit with somewhat lower rates. Such truncal alterations, which represent early convergent patterns across the known heterogeneity of prostate cancer, include nucleolar enlargement, MYC over-expression, telomere shortening, DNA hypermethylation-mediated silencing of GSTP1 and other tumor suppressor/caretaker genes, and formation of oncogenic Ets fusions6,7,8,9,10,11,12,13,14,15,16,17,18. Furthermore, widespread inflammatory lesions in the prostate, termed proliferative inflammatory atrophy (PIA), are thought to be a hotbed from which PIN and invasive cancer can emerge19,20. In the normal prostate, differentiated luminal cells can undergo a cell-state change resembling an intermediate-like phenotype with both luminal and basal cell characteristics as a consequence of androgen depletion (diffuse atrophy) and local inflammation (focal simple atrophy and post-atrophic atrophy)21. PIA is thought to represent a state of inflammatory injury and regeneration in which proliferative and atrophic epithelial cells take on an intermediate cell phenotype. PIA does not universally progress to cancer. However, consistent with a model implicating PIA as a hotbed from which prostate cancer can initiate, these lesions can be seen to merge directly with PIN and, less frequently, with invasive carcinoma22,23. A subset of PIA cells display some of the convergent molecular patterns seen in PIN and invasive carcinoma lesions, with a low frequency of PIA cells showing MYC over-expression, partial promoter hypermethylation, and silencing of GSTP114,19,20,24,25,26,27,28. Additionally, oncogenic chromosomal alterations have been reported in PIA, including Ets fusions, P53 alterations, and gain of chromosome 8 centromeric DNA26,27,29,30.
The convergent transitions in the luminal epithelial cells during the early development of prostate cancer are accompanied by multiple convergent patterns of alterations in the surrounding microenvironment. The atrophic (intermediate) cells observed in PIA lesions generally occur in the context of: i) widespread chronic inflammation composed of myeloid and lymphoid cells, and ii) a reactive stroma with morphologically altered fibroblast and smooth muscle cell compartments that are infiltrated with chronic inflammatory cells. Remarkably, as these PIA regions transition to PIN and invasive cancer lesions, there is a profoundly reduced inflammation and immune cell presence, consistent with the characterization of prostate cancer as “immunologically cold”31,32,33. Furthermore, basal cells in PIN lesions are relatively depleted compared to benign glands and are completely lost in invasive carcinoma lesions6,34. Additionally, the fibroblast, smooth muscle, vascular, and other stromal compartments are also thought to be significantly altered in the context of prostatic neoplasia35. However, the precise nature of the epithelial, stromal, and immune cell alterations during the early transitions in prostate carcinogenesis and how the alterations in the neoplastic cells may mediate these changes is poorly understood. This is in part because of a lack of studies focusing on the early stages of prostate cancer development at single-cell resolution and the inability to longitudinally follow these early stages of disease evolution in human tissues.
Here, we use extensive scRNA-seq and rigorous molecular pathology approaches to study the comparative biology between human prostate cancer tissues and landmark stages in the disease evolution of GEMM of murine prostate cancer to define alterations in the transcriptional programs and cellular states during early prostate cancer development. Consistent with the idea that MYC activation in cancer cells can drive changes to the TME36, we show that activation of MYC signaling is a common denominator across the well-known heterogeneity of prostate cancer and that its downstream sequelae mediate a chain of cell network interactions responsible for reprogramming the TME of prostate cancer. In general, MYC activation in various tumor types promotes an immunosuppressive TME, indirectly modulating cytokine expression37,38,39,40,41. Strikingly, we show that prostate luminal-specific activation of MYC signaling in the mouse initially creates a pro-inflammatory state, with many parallels to that seen in human PIA and early PIN lesions, with subsequent transition to an immunosuppressive state with downregulated tumor suppressor pathways during the progression to invasive carcinoma, reminiscent of the immunologically cold microenvironment of human invasive prostate cancer.
Results
A single-cell transcriptional atlas of localized primary prostate cancer tissues from ten subjects
To develop a detailed atlas of cell states and transcriptional programs in the TME of early prostate cancer, we used scRNA-seq to examine prostate tissues collected from 10 subjects with localized, treatment-naive prostate cancer that underwent radical prostatectomy. For each subject, tissues were collected from each zone of the prostate (peripheral, central, and transition), as well as from areas with visible tumors (Fig. 1A). Because prostate cancer is primarily observed in the peripheral zone42, cancer-enriched and benign-enriched tissue samples from the peripheral zone (Fig. 1B) were used for single-cell RNA-seq assessments, yielding >110,000 cells with high-quality genome-scale gene expression measurements (Fig. 1C, Supplementary Data 1). Dimensionality reduction and cluster analysis revealed the spectrum of cell types present in the prostate TME (Fig. 1C, D, Supplementary Data 2), including luminal and basal epithelial cells, endothelial cells, pericytes, smooth muscle cells, fibroblasts, macrophages, mast cells, T cells, and B cells. Interestingly, in a subset of luminal epithelial cells derived from cancer-enriched tissue samples, we observed a striking subject-specific clustering (Fig. 1C), consistent with the known inter-individual tumor heterogeneity in prostate cancer1,43. However, for the majority of other cell types, there was no subject-specific clustering or skewing; although some clusters, such as basal cells, fibroblasts, and macrophages, did appear to show some skewing in the cancer- vs benign-enriched tissues. These results raised the possibility that although there is significant inter-individual heterogeneity in cancer epithelial cells, there may be cancer-specific convergent alterations in other cells in the TME, evident as less inter-individual heterogeneity. We explored this hypothesis in significant detail in subsequent studies and analyses.
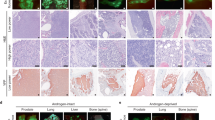
A For each prostatectomy (n = 10), tissue punches were collected from each zone of the prostate as well as the tumor site (4 punches per subject). For each tissue punch, part of the tissue was processed for scRNA-seq from freshly dissociated cells, and the remaining half was frozen and sectioned for histology, IHC, and in situ hybridization. Schematic created in Biorender.com. Panel A created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license. B) Representative examples of hematoxylin and eosin (H&E) staining (first two column panels) and immunohitostical PIN4 staining (third column panel) of fresh frozen peripheral zone tissue punches that were benign enriched (n = 10) or cancer enriched (n = 10). Brown marks tumor protein P63 (TP63) and high molecular weight cytokeratin expressed in basal epithelial cells. Red marks alpha-methylacyl-CoA racemase (AMACR), which is expressed in carcinoma and prostatic intraepithelial neoplasia (PIN). For 1x magnification, scale bar indicates 2.5 mm. For 10x magnification, scale bar indicates 200 μM. C Dimensionality reduction (uniform manifold approximation and projection, UMAP) and clustering analysis of scRNA-seq libraries from peripheral zone (PZ) tissue showed cells clustering by recognized cell types (n = 110,715 cells, 18 samples). Subject and cancer-enriched zones of luminal cells suggest inter-individual and intra-tumor heterogeneity in cancer cells. D Heatmap of cell type marker genes show cluster-specific expression.
Defining benign and neoplastic epithelial cell states and associated gene expression and copy number alterations
We next defined the epithelial cell states in the prostate tissues with greater granularity by subsetting the luminal and basal clusters and repeating dimensionality reduction and clustering analyses. We used a combination of differential gene expression analysis, well-known gene expression markers of known epithelial cell types, and inferred copy number alterations derived from the gene expression data to further refine the identification of neoplastic cells harboring copy number alterations known to be recurrent in human prostate cancer (see Supplementary Information for details)19,44,45,46,47,48. With these analyses, we identified benign basal and luminal cells, intermediate cells49,50,51,52, and eight distinct PIN/cancer clusters that we referred to as Cancer-1 through Cancer-8 (Fig. 2A, B, Supplementary Fig. 1, Supplementary Data 3). These intermediate cells enriched in the peripheral zone of the prostate share overlapping gene expression (PIGR, LCN2, OLFM4) to a previously characterized epithelial cell population referred to as club-like cells49,50,51,52. However, expression of SCGB1A1, a key marker gene of club cells originally described by Henry et al. 49, was rare in intermediate cells of the peripheral zone (Supplementary Information, Supplementary Fig. 1E). The cancer clusters identified here showed strong enrichment of specific subjects, again consistent with the known inter-individual molecular heterogeneity of prostate cancer. In contrast, the benign basal, luminal, and intermediate clusters had representation from all ten subjects, with little inter-individual heterogeneity (Supplementary Fig. 2A, B).
A Epithelial clusters were subsetted, and dimensionality reduction (UMAP) and clustering analysis were repeated (n = 78,621 cells, 18 samples). Benign and cancer cells clustered separately. B Stacked bar plots showing the proportion of cells for each cluster that are from (left panel) benign or cancer-enriched libraries and (right panel) which subject (1-10). C UMAP indicating cells from inferred copy number variation (InferCNV) analysis harboring cancer-associated mutations by subject. D Heatmap from inferCNV analysis of cancer cells from Subject 2 indicating regions with inferred CNV gain (red) and loss (blue). Cancer subgroup A shows 10q loss, while the remaining subgroups show 10q is intact, suggesting heterogeneous loss of PTEN (black box). E Example of intra-tumor heterogeneity from Subject 2. Immunohistochemical for frozen sections for PIN4, ERG, and PTEN staining shows cancer cells in top panels are negative for ERG and positive for PTEN. In contrast, lower panels show another group of cancer cells that are ERG-positive but PTEN-negative. Staining was performed across 39 tissue punches, with five specifically from Subject 2. For 2x magnification, scale bar indicates 2 mm. For 10x magnification, scale bar indicates 200 μM. F Heatmap of normalized enrichment scores (NES) from gene set enrichment analysis (GSEA) of Hallmark collection and “Dang MYC targets up gene set” comparing each cancer cluster with the luminal cluster. G Plot showing NES of top 20 pathways by adjusted p-value, comparing aggregated cancer clusters with the luminal cluster by GSEA. Statistics were derived using the fgsea implementation of a two-tailed GSEA. The adjusted p-value statistic is derived from multiple hypothesis testing using Benjamini-Hochberg procedure across all gene sets considered.
The PIN/cancer clusters showed distinct gene expression programs and copy number alterations consistent with known prostate cancer subtypes that were enriched in specific subjects (Fig. 2A, B, Supplementary Fig. 1, Supplementary Information), including ERG fusion-positive cancers (Cancer-2, Cancer-4, and Cancer-5 clusters; enriched in subjects 5, 9, and 10 respectively) and SPINK1 positive cancers (Cancer-1, Cancer-3, Cancer-6, Cancer-7, and Cancer-8; enriched in subjects 2, 1, 7, 4, and 8 respectively) (Supplementary Fig. 1B)15,16,17,53,54. Consistent with known patterns in human prostate cancer, we observed ERG-positive cases with retained PTEN expression, ERG-positive cases with complete PTEN loss, and ERG-negative cases with PTEN-intact protein expression (Supplementary Fig. 2)16,17,55,56. Furthermore, inferred copy number alteration patterns showed significant heterogeneity across subjects, consistent with known regions of recurrent copy number alteration in primary prostate cancer, and strong parallels with the gene expression derived clustering (Fig. 2C, Supplementary Figs. 3–5, Supplementary Information)17,57.
In addition to the striking inter-individual heterogeneity in the cancer clusters, the scRNA-seq-derived copy number analysis also captured intra-individual tumor heterogeneity. For example, in subject 2, a subset of luminal cells had 10q loss, a common CNV in prostate cancer consistent with PTEN loss58,59,60,61, while other cancer cells from this subject did not harbor 10q/PTEN loss (Fig. 2D and Supplementary Fig. 3B). Tissue staining from subject 2 showed heterogeneous loss of PTEN, with one group of cancer cells with PTEN expression intact and another group with loss of PTEN (Fig. 2E), consistent with the intratumoral heterogeneity of PTEN deletion identified by the inferCNV analysis.
These in situ tissue analyses corroborated that our scRNA-seq analyses could capture the well-known inter- and intra-individual tumor heterogeneity in prostate cancer. Additionally, our scRNA-seq analyses showed very little inter-individual heterogeneity in benign basal and luminal epithelial cells, intermediate cells, and stromal cells.
MYC activation is a common denominator across the known molecular heterogeneity of prostate cancer
To identify pathways dysregulated in cancer clusters 1-8, we performed gene set enrichment analysis (GSEA) comparing benign luminal epithelial cells with each of the eight cancer clusters using the Hallmark gene set collection. As expected, there was variation in the upregulation and downregulation of several gene sets (Fig. 2F), consistent with the heterogeneity we observed in our multimodal investigation of prostate cancer tissues by scRNA-seq clustering analysis, inferred CNVs, and in situ tissue staining. Two gene sets were universally downregulated, including androgen response and cholesterol homeostasis. Three Hallmark gene sets were upregulated in all eight cancer clusters: MYC Targets V1, MYC Targets V2, and E2F Targets. Aggregating cancer clusters 1-8 and performing differential expression and pathway analysis of this aggregated cancer cluster against benign luminal cell clusters revealed that the Hallmark MYC Targets V1 gene set was the top upregulated pathway and androgen response the top downregulated pathway in the cancer clusters (Fig. 2G, and Supplementary Data 4). The universal upregulation of MYC targets and downregulation of the androgen response pathway across all cancer clusters are consistent with recent reports characterizing the antagonistic relationship between MYC and androgen response genes62,63,64. Including the Dang MYC Targets Up gene set in GSEA65 reaffirmed significant upregulation of MYC targets across all cancer clusters (Fig. 2F–G). To confirm that upregulated MYC activity is prevalent across prostate cancer subtypes, we examined the TCGA prostate cancer dataset, which was instrumental in defining molecular subtypes of prostate cancer16. We found upregulated Hallmark MYC Targets V1, Hallmark MYC Targets V2, and Dang MYC Targets in the vast majority of prostate cancer samples, regardless of Gleason score and molecular subtype (Supplementary Fig. 6).
Additionally, we observed elevated expression of MYC Targets V1 genes in the basal, intermediate, and cancer clusters compared to benign luminal epithelial clusters (Supplementary Fig. 7A–F). Validation studies with in situ tissue staining confirmed the over-expression of nuclear MYC in prostate cancer cells and basal cells from benign glands compared to normal luminal epithelial cells (Supplementary Fig. 7G). These findings are consistent with prior immunostaining studies showing that MYC expression is common in prostatic basal cells, negative in normal luminal cells, somewhat upregulated in atrophic luminal (intermediate) cells, and its overexpression is a highly frequent and early event in prostate cancer18,24.
Upregulated MYC activity in benign basal and intermediate cells suggests that at least some aspects of MYC activity and its oncogenic potential are likely context-specific. We further interrogated MYC activity in the prostate epithelial clusters by expanding our pathway analysis to include 20 MYC-associated gene sets. Several MYC upregulated gene sets and downregulated gene sets were significant in the cancer cluster (Supplementary Fig. 8A). Gene signature analysis of the leading edge genes of the Dang Bound by MYC geneset65 confirmed enrichment in basal, cancer, and intermediate clusters. However, gene signature analysis of the leading edge genes from the Dang Regulated by MYC Down geneset revealed cancer cluster-specific downregulation of MYC targets (Supplementary Fig. 8B). Furthermore, these downregulated MYC targets in cancer cells significantly overlapped with gatekeeper pathways P53 and Apoptosis, and the pro-inflammatory TNFA signaling via NFKB pathway (Supplementary Fig. 8C, D). Comparing the two MYC upregulated luminal clusters, cancer and intermediate, re-affirmed significant downregulation of TNFA signaling via NFKB, P53 pathway, and apoptosis (Supplementary Fig. 7C). Comparisons of cancer and intermediate also revealed significant upregulation of mTOR signaling and downregulation of the negative regulation of PI3K/AKT/mTOR signaling, including PTEN (Supplementary Fig. 8E). Interestingly, the Immediate Early Response 3 gene (IER3), a MYC downregulated gene in the cancer clusters (Supplementary Fig. 8D–F), was previously reported to be a negative regulator of the PI3K/AKT/mTOR signaling pathway66,67. Altogether, these findings suggest that MYC activity differs between neoplastic and intermediate cells. In the cancer clusters, MYC directly and indirectly downregulates P53 signaling, apoptosis, and TNFA signaling and upregulates mTOR signaling.
MYC activity reprograms neighboring benign epithelial cells in the TME
It is difficult to assess the causal sequelae of MYC activation in human tissues. Therefore, to investigate the downstream consequences of MYC activation in the prostate TME, we modeled MYC activation in mouse prostates using transgenic Hi-Myc mice62,68,69,70,71. Because the 6-month timepoint marks a critical stage in carcinogenesis, the switch from precursor to invasive carcinoma in the Hi-Myc model, we rigorously assessed the single-cell transcriptomes in each prostate lobe (anterior, dorsal, lateral, and ventral) of the MYC-driven mouse model of prostate cancer (Hi-Myc) in both FVB/NJ and C57BL/6 J mouse strains aged 6 months (Fig. 3A–C, Supplementary Fig. 9, Supplementary Information). As a reference, age-matched wild-type (WT) animals from our previously published single-cell atlas of the mouse prostate by lobe and strain were included in the analysis72.
A Tissue was collected from each prostate lobe (anterior, dorsal, lateral, and ventral) of the MYC-driven mouse model of cancer (Hi-Myc) in both FVB/NJ (n = 2) and C57BL/6 J (N = 2) mouse strains, as well as age-matched wild type animals (FVB/NJ N = 2, C57BL/6 J n = 3). Panel A created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license. B Representative H&E (upper panels) and chromogenic in situ hybridization (CISH, lower panels) staining of human MYC transgene in FFPE mouse prostate tissues from 6-month-old Hi-Myc and age-matched wild-type (WT) mice. Staining was performed across 32 samples. Scale bar used for WT samples indicates 200 μM. Scale bar used for Hi-Myc samples indicates 500 μM, and 200 μM zoomed in. C Representative CISH staining of human MYC transgene in each lobe of Hi-Myc mouse prostate from C57BL/6 J and FVB/NJ mice. Staining was performed across 32 samples. For C57BL/6 J, scale bars indicate 2 mm (Anterior), 1 mm (Dorsal), 500 μM (Lateral) and 1 mm (Ventral). For FVB/NJ, all scale bars indicate 1 mm. D UMAP of mouse scRNA-seq by cell type, strain, lobe, genotype, and MYC transgene expression (n = 58,435 cells, 36 samples). E Heatmap of each cluster and cell type marker gene expression. F Heatmap showing expression of genes positively correlated with MYC transgene expression in luminal cell populations stratified by genotype, strain, and lobe. Genes are grouped by associated biological pathways. The first column shows the scaled expression of the MYC transgene.
Dimensionality reduction and clustering analysis showed that cells grouped primarily by known cell types (Fig. 3D, E, Supplementary Fig. 10A). Consistent with previously reported single-cell atlases of the mouse prostate72,73, luminal epithelial cells from WT mice subclustered by lobe and strain (Supplementary Fig. 10A). However, Hi-Myc luminal cells clustered by genotype and lost strain and lobe-specific clustering, with the exception of the luminal cells from the ventral lobe (Fig. 3D and Supplementary Fig. 10). Genes positively correlated with MYC transgene expression in luminal cells were associated with several biological pathways linked to MYC activation (Fig. 3F), including cell cycle progression, ribosome biogenesis, RNA processing, protein degradation, and mitochondrial function65,74,75,76.
To investigate the MYC-driven changes in the epithelial population, we subsetted our analysis to the luminal and basal epithelial cells and identified four epithelial clusters disproportionately enriched in Hi-Myc mice (Fig. 4A–C, Supplementary Data 5, Supplementary Information). Two of the epithelial clusters that were strongly enriched in the Hi-Myc compared to WT animals showed strong expression of the human MYC transgene and upregulation of MYC targets (Fig. 4D). Given the in situ analysis of MYC transgene expression coinciding exclusively in PIN and cancer lesions in Hi-Myc mice (Fig. 3B, C), the cells of these two clusters represent the precursor and invasive carcinoma cells in the Hi-Myc animals. We therefore termed these clusters Luminal MYC 1 and Luminal MYC 2 and noted that they were enriched in the dorsal/lateral vs. ventral lobes, respectively (Fig. 4C).
A UMAP of mouse prostate scRNA-seq subsetted to epithelial clusters (n = 52,116 cells, 36 samples). B UMAPs of Psca and Ly6d show cluster-enriched gene expression. C Scatter plots showing the proportion of epithelial clusters for each sample. Each dot represents a sample (n = 36) colored by lobe (anterior, dorsal, lateral, ventral). Bar represents the mean cell proportion of samples for each genotype. Statistics were generated by comparing WT (n = 20) and Hi-Myc (n = 16) samples using linear regression analysis (limma) in scRNA-seq with multiple samples (RAISIN) cell proportions test. The adjusted p-value is derived from a two-sided t-test and adjusted for multiple hypothesis testing using Benjamini-Hochberg procedure. D Heatmap showing expression of endogenous Myc, transgene MYC, and Hallmark MYC targets V1 leading edge genes from GSEA for each epithelial cluster. E Representative example of Ly6d CISH staining basal compartment in PIN gland of Hi-Myc FFPE prostate tissue. Staining was performed across 10 samples. For 10x magnification, scale bar indicates 200 μM. For 40x magnification, scale bar indicates 50 μM. F Example of KRT6A CISH staining basal compartment of intraductal carcinoma in human prostate cancer from frozen tissue sections. Staining was performed in 10 tissue punches. Scale bar indicates 500 μM.
Surprisingly, the other two clusters, Ly6d-expressing luminal and basal cells, did not show robust expression of the human MYC transgene despite being significantly enriched in the Hi-Myc mice. This suggested that MYC activation in neoplastic luminal cells led to the expansion of a pre-existing population or an induction of these Ly6d-expressing epithelial cells in the surrounding microenvironment, referred to as Reactive Luminal and Reactive Basal cells. We, therefore, further explored these cell clusters (Supplementary Information) and sought to identify parallels in the human prostate cancer tissues.
Gene expression analysis showed that Krt6a was one of the genes uniquely upregulated in Ly6d-expressing luminal and basal cells and strongly co-expressed with Ly6d (Supplementary Figs. 11B and 12A, D). Expression analysis of epithelial cells from prostatectomy samples revealed that a subset of basal cells expressed KRT6A and LY6D, suggesting the mouse Ly6d-expressing Reactive Basal cells enriched in Hi-Myc mouse prostates were also observed in the human prostate (Supplementary Fig. 12). Chromogenic in situ hybridization (CISH) for Ly6d in Hi-Myc mouse prostate tissues revealed a subset of PIN lesions expressing Ly6d in the basal compartment (Fig. 4E). Likewise, CISH staining in human prostatectomy tissues revealed that KRT6A was expressed primarily in a subset of basal cells of prostate glands, including PIA, PIN, intraductal carcinoma, and to a much lesser extent, a subset of atrophic (intermediate) cells in PIA (Fig. 4F, Supplementary Fig. 13).
Although the Reactive Basal cluster was highly enriched in the Hi-Myc prostate, the lack of MYC transgene expression and in situ tissue staining pattern of Ly6d in mouse prostates and KRT6A in human prostates suggest that this cell type may be induced by MYC-transformed neoplastic cells through paracrine signaling (Fig. 4C, D). Inferred ligand-receptor paired interactions revealed that the Luminal MYC 1 cluster had the strongest inferred interactions with the Reactive Basal cluster, expressing several of the top ligands targeting the Reactive Basal cluster, including the type I interferon ligand, interferon kappa (Ifnk) (Fig. 5A, B)77,78. Interestingly, expression of the type I interferon receptor subunits Ifnar1 and Ifnar2 in the Reactive Basal cells was positively correlated with the activity of the interferon regulatory factor IRF5 as measured by expression of its downstream targets (Fig. 5C, D). In line with upregulated IRF5 activity, the Reactive Basal cluster also had increased Irf5 expression (Supplementary Fig. 14B). Consistent with activated interferon signaling in the TME, the Luminal MYC 1 and Reactive Basal clusters were both found to be upregulated in interferon alpha and gamma response pathways (Fig. 5E). This cell communication network and pathway analyses suggested that MYC-expressing neoplastic cells induce enrichment of neighboring Ly6d-expressing epithelial cells in part via type I interferon signaling and other ligand-receptor interactions (Fig. 5A–E, Supplementary Information).
A Cell communication plot of epithelial clusters from inferred ligand-receptor-transcription factor network analysis (Domino). Nodes represent individual clusters and are scaled based on ligand expression. Edges indicate signaling between two clusters and are weighted based on the strength of signaling. The edge color matches the cluster expressing the ligand. The ligands from the Luminal MYC 1 cluster is marked in yellow, while all other clusters are colored grey to highlight specific interactions from the Luminal MYC 1 cluster to all other epithelial clusters. B Heatmap showing which epithelial clusters express the top ligands targeting receptors expressed on the Reactive Basal cluster from inferred ligand-receptor pair interactions. C Heatmap showing the correlation of transcription factor activity scores (from SCENIC analysis) and receptor expression. Transcription factors were selected based on top activated scores in the Reactive Basal cluster. The receptors represented in this heatmap are known to bind to the top ligands targeting the Reactive Basal cluster. D Expression of downstream targets of the transcription factor interferon response factor 5 (IRF5) in epithelial clusters. E Plot showing NES of top 15 pathways by adjusted p-value. Statistics derived using fgsea implementation of a two-tailed GSEA, and the adjusted p-value from multiple hypothesis testing using Benjamini-Hochberg procedure. Comparisons include Luminal and Luminal MYC 1 clusters (top panel) and Basal and Reactive Basal clusters (bottom panel). Representative immunohistochemical staining in WT and Hi-Myc mouse prostate tissues of F CD3 to mark T cells (40 tissues) and G F4/80 to mark macrophages (36 tissues). In F, the 1x magnification scale bar indicates 1 mm and the 10x magnification scale bar indicates 100 μM. In G, the 1x magnification scale bar indicates 1 mm for WT and 2 mm for Hi-Myc. The 10x magnification scale bar indicates 200 μM. H Heatmaps of top 20 genes upregulated in the Luminal MYC 1 cluster at 6 months and downregulated at 10 months. Genes are grouped by associated gene sets from the GSEA Hallmark collection.
Proinflammatory to immunosuppressive switch in MYC-expressing luminal cluster accompanying progression to invasive cancer
In addition to upregulated interferon response pathways, GSEA comparing Luminal and Luminal MYC 1 cells also revealed upregulation of another pro-inflammatory pathway, tumor necrosis factor alpha (TNFA) signaling via nuclear factor kappa-light-chain-enhancer of activated B cells (NF-κB) (Fig. 5E). In both FVB/NJ and C57BL/6 J strains, we observed a significant enrichment of various immune cells, including mast cells, T cells, and Trem2-expressing macrophages in Hi-Myc prostates compared to age-matched WT prostates (Supplementary Fig. 15, Supplementary Information). Prior studies have suggested TREM2 Macrophages have an immunosuppressive role in the TME79,80. Immunostaining for T cells (CD3) and macrophages (F480) in WT and Hi-Myc mouse prostates showed enrichment of T cells and macrophages in the prostate TME of Hi-Myc mice (Fig. 5F–G), consistent with the upregulation of inflammatory pathways observed by GSEA in luminal epithelial cells (Fig. 5E).
To better understand the dynamics of the epithelial and immune cell phenotypes in response to MYC activation and cancer progression over time, we expanded our analysis to include a later time point (10 months +/− 6 weeks) of dorsal and lateral lobes from Hi-Myc and aged-matched WT prostates from FVB/NJ mice. The FVB/NJ dorsal and lateral lobes in Hi-Myc mice are known to progress to more penetrant invasive cancer phenotypes by the 10-month time point, representing a critical stage of invasive carcinoma and allows us to understand the dynamics of the TME during cancer progression (Supplementary Fig. 9). We merged scRNA-seq datasets from the dorsal and lateral lobes of Hi-Myc and aged-matched WT prostates from FVB/NJ mice aged 6 months and 10 months and integrated by age (Supplementary Fig. 16A, B, Supplementary Data 6). GSEA of MYC-expressing Luminal MYC 1 cluster comparing 10-month and 6-month mice revealed a shift in multiple pathways (Supplementary Fig. 16C). In particular, the Luminal MYC 1 cluster in 10-month-old mice downregulated several genes associated with critical pathways that serve as gatekeepers of malignant transformation, including P53 and DNA repair (Fig. 5H, Supplementary Fig. 16C). Strikingly, the Luminal MYC 1 cluster was also downregulated for several inflammation-related genes, including inflammatory response and IL6 JAK STAT3 signaling (Fig. 5H, Supplementary Fig. 16C). Collectively, these pathway analyses suggest that a cancer-cell intrinsic pro-inflammatory to immunosuppressive switch coincides with pathway alterations associated with cancer progression.
We next examined whether the cancer cell-intrinsic switch from pro-inflammatory to immunosuppressive programs was accompanied by alterations in the composition of the immune microenvironment between the 6-month, precursor-enriched, and 10-month, invasive cancer-enriched time points. We first defined the major immune cell clusters (Fig. 6A, B) and noted several distinct differences between the two time points, representing the precursor and invasive carcinoma stages. First, we observed an enrichment in the 10-month vs. 6-month timepoint of multiple cell populations known to have immunosuppressive function, including TREM2 positive macrophages, myeloid-derived suppressor cells (MDSCs), and regulatory T cells (T regs), which could also be confirmed by in situ hybridization analysis of the mouse prostate tissues (Fig. 6C, Supplementary Fig. 17). In the human prostatectomy dataset, we observed a subset of T cells that expressed regulatory T-cell-associated markers in a gene signature analysis (UCell) (Fig. 6D–F). We also observed that a subset of macrophages expressed TREM2 (Fig. 6G), suggesting that immune composition changes driven by MYC oncogene activity in mice are similar to those observed in human prostate cancer. Notably, a recent scRNA-seq prostate cancer study focused on invasive cribriform carcinoma and intraductal carcinoma also reported increased TREM2 macrophages in the TME compared to benign tissues81.
A Immune cell clusters were subsetted from dorsal and lateral lobes of FVB/NJ mice from WT and Hi-Myc at 6 months and 10 months, and dimensionality reduction (UMAP) and clustering analysis were repeated (n = 4,474 cells, 12 samples). B Heatmap of immune cell type marker genes shows cluster-specific expression in merged scRNA-seq data. C Scatter plots showing the proportion of select immune cell populations for each sample (n = 12). Each dot represents a sample colored by genotype and age. Bars represent the mean cell proportion of samples for each group. Statistics were generated by comparing Hi-Myc samples at 6 months (n = 4) and 10 months (n = 2) using RAISIN’s cell proportions test based on linear regression modeling (limma). The adjusted p-value is derived from a two-sided t-test and adjusted for multiple hypothesis testing using Benjamini-Hochberg procedure. D UMAP of human immune cell clusters from prostatectomy scRNA-seq dataset (n = 12,108 cells, 18 samples). E Heatmap of immune cell type marker gene expression of immune cells from prostatectomy samples. Expression of F) regulatory T cell gene signature (IL2RA, FOXP3, CD4, IKZF2, CCR4, CTLA4) and G) TREM2 in human immune cell clusters.
Stromal cells are significantly altered in the TME of MYC-driven prostate cancer
In the merged 6-month and 10-month FVB/NJ dorsal and lateral datasets, the stromal populations included smooth muscle, pericytes, endothelial, glial, and fibroblast cells (Fig. 7A, Supplementary Fig. 18A–D, Supplementary Data 7). Hi-Myc animals showed increased fractions of endothelial cells, consistent with induction of neovasculature in MYC-driven neoplasia (Fig. 7B).
A Immune clusters were subsetted from dorsal and lateral lobes of FVB/NJ mice from WT and Hi-Myc at 6 months and 10 months (n = 6,162 cells, 12 samples). A fibroblast subcluster enriched in Hi-Myc tissue expresses Timp1. B Scatter plots showing the proportion of select stromal cell populations significantly altered in Hi-Myc compared to WT. Each dot represents a sample (n = 12) colored by genotype and age. Bars represent the mean cell proportion of samples for each group. RAISIN’s cell proportions test used to statistically compare WT (n = 6) and Hi-Myc (n = 6) samples. The adjusted p-value is derived from a two-sided t-test and adjusted for multiple hypothesis testing using Benjamini-Hochberg. C Violin plot showing expression of Pdgfra, Timp1, and fibrosis-associated collagens (Col1a1, Col1a2, Col3a1, Col5a2) by genotype (WT and Hi-Myc) and age (6 months and 10 months) in fibroblast cells. D UMAP of human stromal cell clusters (n = 18,837 cells, 18 samples). Right panel shows the expression of the Hi-Myc Fibroblast Timp1 gene signature (TIMP1, MFAP5, SERPINA3, IGF1, SFRP1, MMP2, SERPINF1, COL1A1, COL5A2, COL3A1). E Scatter plot showing proportion of Macrophages Trem2 and Fibroblast Timp1 clusters for each sample with corresponding two-tailed Pearson correlation coefficient and p-value. F Inferred ligand-receptor-transcription factor network analysis (Domino) of epithelial, macrophage, and fibroblast clusters from FVB/NJ aggregated scRNA-seq data. Luminal MYC 1 cluster is marked in red, Macrophages Trem2 cluster is marked in yellow, Fibroblast Timp1 node is marked in black, and all other clusters are colored grey. G Heatmap showing which clusters express the top ligands targeting the Fibroblast Timp1 cluster. H Heatmap showing the correlation of transcription factor activity scores (derived from SCENIC) and receptor expression. Transcription factors were selected based on top activated scores in the Fibroblast Timp1 cluster. Receptors for TNF, PTPRC, CD72, OSM, and TGFB1 are shown.
The fibroblasts sub-clustered into three groups, including two fibroblasts previously characterized in WT mouse prostates, Sult1e1-expressing interstitial fibroblasts and Rorb-expressing subglandular fibroblasts72,82, and a novel Timp1-expressing fibroblast cluster (Fibroblast Timp1) that was highly enriched in Hi-Myc mice (Fig. 7B, Supplementary Fig. 18C). This cluster showed significantly increased expression of fibrosis-associated collagens Col1a1, Col1a2, Col3a1, and Col5a283,84,85, consistent with a reactive stroma and the desmoplastic pan cancer-associated fibroblast gene expression signature (pan-dCAF)35,86 (Fig. 7C, Supplementary Fig. 19, Supplementary Information). More broadly, the enrichment of fibrosis-associated extracellular matrix (ECM) gene expression in the TME of prostate cancer is consistent with a desmoplastic response, a feature associated with invasive carcinoma in tissues87,88,89. Interestingly, individual cells in the Timp1 Fibroblast cluster showed mutually exclusive expression of Rorb or Sult1e1, suggesting that the Fibroblast Timp1 cells are potentially induced from either subglandular or interstitial fibroblasts (Supplementary Fig. 18D). Genes differentially expressed in the Fibroblast Timp1 cluster in mice were evaluated in the human stroma. Gene signature analysis revealed that a subset of fibroblasts in human prostatectomy samples was enriched for the Fibroblast Timp1 gene signature (Fig. 7D, Supplementary Fig. 18, Supplementary Data 9).
In the mouse prostate TME, this increase in the Fibroblast Timp1 CAFs was accompanied by decreases in the normal interstitial and subglandular fibroblasts and smooth muscle clusters in the Hi-Myc mice (Fig. 7B). Similar stromal changes were also observed in C57BL6/J Hi-Myc mice, with enrichment of endothelial cells and Timp1-expressing fibroblasts and a reduced proportion of interstitial fibroblasts (Supplementary Fig. 21, Supplementary Data 8). Intriguingly, these Fibroblast Timp1 CAFs increased in the 10-month invasive cancer enriched compared to 6-month precursor-enriched tissues (Fig. 7B).
Moreover, the fraction of cells in the Fibroblast Timp1 cluster was significantly correlated with the fraction of cells in the Macrophage Trem2 cluster (Fig. 7E), suggesting that these two clusters may be connected through paracrine signaling. Inferred ligand-receptor-transcription factor network analysis in the mouse dataset showed strong inferred interactions of the Macrophage Trem2 cluster with the Fibroblast Timp1 cluster (Fig. 7F). In our scRNA-seq data, many ligands targeting the Fibroblast Timp1 cluster were expressed in the Macrophage Trem2 cluster, including transforming growth factor beta 1 (Tgfb1) (Fig. 7G). Notably, transcription factors such as early growth response 4 (EGR4), EGR2, and Wilms Tumor 1 (WT1) had activation scores positively correlated with TGFB1 receptors Tgfbr1, Tgfbr2, Tgfbr3, and activin A receptor type 1 (Acvr1) in the Fibroblast Timp1 cluster (Fig. 7H). While multiple transcription factors may contribute to the upregulation of the fibrosis-associated genes in the Fibroblast Timp1 cluster, EGR4, in particular, is known to regulate Timp1 and the fibrosis-associated collagens Col1a1, Col1a2, Col3a1, and Col5a290. Importantly, TGFB signaling is a key instigator of tissue fibrosis and is thought to be mediated by immune cells91,92. Indeed, scRNA-seq analysis coupled with in situ tissue validation is consistent with a sequence of events involving infiltration and expansion of Tgfb1 expressing macrophages in tissues with MYC oncogene expressing luminal cells, coinciding with dramatic stromal remodeling enriched for a unique fibroblast population with robust collagen expression.
Upregulation of mTOR signaling following PTEN loss accelerates MYC-driven alterations to the prostate TME
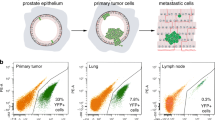
The pro-inflammatory to immunosuppressive switch observed in human prostate cancer was also found in the Hi-Myc neoplastic cells. Compared to 6-month-old Hi-Myc mice, the Luminal MYC 1 cells in 10-month-old Hi-Myc had downregulation of multiple pro-inflammatory pathways, including TNFA signaling via NFKB, inflammatory response, interferon gamma response, and interferon alpha response, with several genes being MYC downregulated targets (Supplementary Fig. 21A, B). Analysis of human prostate cancer also implicated the upregulation of mTOR signaling and downregulation of negative regulators of mTOR signaling, such as IER3 and PTEN, coinciding with the immunosuppressive switch (Supplementary Figs. 7C, 8E, F). Similarly, in Hi-Myc mice, the Luminal MYC 1 cells from 10-month-old mice were significantly upregulated for mTOR signaling and downregulated for negative regulators of mTOR signaling (Supplementary Fig. 21C, D). Collectively, these findings in human prostate cancer and Hi-Myc mice suggested that negative regulators of mTOR signaling delay MYC-mediated malignant transformation. Notably, PTEN, a well-established tumor suppressor and negative regulator of PI3K/AKT/mTOR signaling, is lost biallelically in ~20% of primary prostate cancer and half of aggressive castration-resistant prostate cancer56. We hypothesized that genetic disruption of PTEN would accelerate the MYC-induced pro-inflammatory to immunosuppressive switch in prostate carcinogenesis and associated TME alterations. To test this hypothesis, we employed a transgenic mouse model of MYC activation and PTEN loss driven by the prostate luminal-specific HoxB13 regulatory locus (BMPC). In contrast to the Hi-Myc mice, BMPC mice have accelerated prostate cancer progression with extensive PIN-like lesions at 3 months and develop metastatic disease at 5-6 months93. Tissues from BMPC mice were dissected from 3-month-old prostate lobes and two lymph node metastatic lesions from a 6-month-old BMPC mouse. We performed scRNA-seq analysis and integrated it with the FVB WT and Hi-Myc scRNA-seq datasets (Supplementary Fig. 21E–H). The TME of the 3-month BMPC mouse prostate resembled the TME of the Hi-Myc 6-month prostate, containing Ly6d-expressing Reactive Basal cells, Trem2 macrophages, and Timp1 fibroblasts (Supplementary Fig. 21I–L). The TME of metastatic lesions collected from the lymph nodes of 6-month BMPC mice were also enriched for Luminal MYC cells, Trem2 macrophages, and Timp1 fibroblasts. However, Reactive Basal cells were absent from the TME of metastatic lesions as expected since the site of metastasis would not have basal cells that can become reactive. Gene signature analysis across the various stages of prostate cancer progression revealed an enhanced upregulation of MYC programs in metastatic prostate cancer cells (Supplementary Fig. 22A). Additionally, suppression of the P53 pathway and the cell-intrinsic pro-inflammatory to immunosuppressive switch observed in Hi-Myc (6 months vs 10 months) occurred in a much-accelerated timeline in the BMPC mice (3 months vs 6 months) (Supplementary Fig. 22B–D), suggesting that programmed loss of the PTEN tumor suppressor pathway and consequent upregulated mTOR signaling accelerated MYC-driven alterations to the prostate TME.
Discussion
Primary prostate cancer is a heterogeneous disease with multiple genomic alterations driving carcinogenesis. For example, common alterations include fusion and overexpression of E26 transformation-specific (ETS) transcription factors, as well as mutations in speckle type BTB/POZ protein (SPOP), forkhead box A1 (FOXA1), isocitrate dehydrogenase (NADP( + )) 1 (IDH1), and phosphatase and tensin homolog (PTEN)1,15,16,94,95. Yet, despite the underlying genomic heterogeneity typical of this disease, there are several key characteristics of the TME shared across most primary prostate cancers. The glandular architecture characteristic of the normal prostate, which consists of cuboidal basal cells aligning the perimeter of differentiated columnar epithelial cells, devolves in carcinoma, with cancer cells invading the surrounding fibromuscular stroma96,97,98. In digital rectal exams, areas of the prostate suspected of harboring cancer cells can stiffen and feel indurated99, likely due to the desmoplastic response driven by an inflammatory reactive stroma characterized by infiltrating immune cells and increased deposition of extracellular matrix proteins35.
Our single-cell and complementary in situ tissue analysis of prostatectomy tissues collected from 10 patients diagnosed with primary prostate cancer are consistent with the notion that prostate cancer is a heterogeneous disease. Yet, despite this tumor heterogeneity, we observed MYC activation as a common denominator across these cancer clusters. This observation was supported when performing differential gene expression and pathway analysis of the publicly available TCGA dataset of primary prostate cancer16.
Evaluating the direct effects of MYC activation in the prostate TME is challenging in human tissues. Therefore, we modeled MYC activation using the transgenic Hi-Myc mouse model of prostate cancer68,69 and used computational approaches to evaluate changes in the TME. The Hi-Myc model has 34 additional amino acids from the cloning vector on the C-terminal end of the expressed MYC protein68. To account for potential strain-specific and model-specific effects, we evaluated the Hi-Myc mouse model of prostate cancer in both FVB/NJ and C57BL/6 J mouse strains and examined the TME of BMPC mice, another MYC-driven model of prostate cancer. Key insights from mouse model studies were evaluated in prostatectomy tissue samples to confirm that observations from mouse studies were not a mouse model-specific phenomenon but also relevant to human prostate cancer.
Based on these analyses, we found that MYC activation in neoplastic cells sets off a cascade of alterations that fundamentally alters the composition and transcriptional programs of surrounding cell types in the TME (Fig. 8). Initially, MYC activation is associated with an induction of a pro-immunogenic transcriptional program in a precursor-enriched state. This leads to propagation of interferon and inflammatory mediators that induce a new basal cell state, likely through engagement of type I interferon receptors. These newly programmed Reactive Basal Ly6d/Krt6a-expressing cells in turn propagate a pro-immunogenic program with resulting influx of pro-inflammatory cells including macrophages, mast cells, and T cells. Over time, the MYC-activated luminal neoplastic cells in the precursor state undergo a switch to significantly downregulate the pro-immunogenic transcriptional programs alongside downregulation of other key tumor suppressor pathways during progression to invasive cancer. This change is accompanied by an enrichment of immunosuppressive cells in the microenvironment, including MDSCs, Tregs, and TREM2 macrophages. This immunosuppressive microenvironment is consistent with the well-established observation that prostate cancer TME is immunologically cold31,32,33,100. This immunosuppressive stage in prostate cancer tumor progression, following the pro-inflammatory precursor stage, is consistent with previous reports demonstrating that cell-intrinsic MYC activation promotes immune evasion by suppressing cGAS-STING and MHCI-mediated antigen presentation, and upregulating CD47, PD-L1 and a glycoimmune checkpoint ligand101,102,103,104.
A MYC activation in luminal epithelial cells leads to the induction of benign Ly6d/Krt6a-expressing epithelial population (Reactive Epithelial cells) in the basal compartment of prostate glands. B At the precursor stage, the TME is pro-inflammatory, with upregulated interferon signaling in both MYC-expressing luminal cells and Reactive Basal cells, and (C) enrichment of various immune cells in the TME, including mast cells, T cells, and Trem2-expressing macrophages. D As MYC-expressing luminal cells progress to invasive carcinoma, there is a pro-inflammatory to immunosuppressive switch with downregulation of inflammatory response pathways and IL6 JAK STAT3 signaling, and enrichment of immunosuppressive cell types, including regulatory T cells (Tregs), myeloid-derived suppressive cells (MDSCs), and Trem2-expressing macrophages. E Secretion of TGFB by Trem2-expressing macrophages activates TGFB signaling and EGR4 transcription factor activity in fibroblasts, resulting in a desmoplastic CAF population expressing Timp1 and reactive-stroma-associated ECM proteins such as collagen. Graphical abstract created in Biorender.com. Figure 8 created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license.
The TREM2 macrophages express high levels of Tgfb1, which likely signals through TGFbeta receptors to induce transcription factor (e.g., Egr4, Egr2, Wt1) mediated expression of reactive-stroma-associated collagen genes and Timp1 in a population of cancer-associated fibroblasts that dominate the stromal microenvironment in the invasive cancer-enriched tissues. Consistent with these findings, a recent single-cell study of fibrosis in human liver tissues reported that TREM2 macrophages promoted fibrosis-associated changes in the stroma105.
This cascade of TME alterations to cell states in MYC-driven murine prostate cancer had several key parallels to that in human prostate cancer. The initial pro-immunogenic programs induced in the Luminal MYC 1 cells in the mouse have strong parallels to the intermediate cell clusters in the human prostate, which also show activation of MYC signaling alongside strong activation of immunogenic programs. The Reactive Basal Ly6d/Krt6a cells seen in the precursor lesions have strong parallels to a LY6D/KRT6A cell population seen in the basal compartment of PIA, PIN, and intraductal cancer lesions in the human tissues. The Macrophage Trem2 and Fibroblast Timp1 populations also had strong parallels in the human cancer-enriched prostate tissues. These findings establish the human relevance of the TME alterations accompanying MYC-driven prostate cancer in the mouse model.
While these findings provide some fundamental insights into convergent aspects of prostate cancer cells and their microenvironment, multiple important questions are raised that will require significant additional studies. First, what are the specific mechanisms by which MYC activation leads to the initial induction of pro-immunogenic programs? Numerous prior studies have implicated multiple pathways connecting MYC activation to immunosuppressive signaling, including suppression of interferon signaling gene expression and subsequent inflammatory response, downregulation of MHC class I antigen expression, and secretion of pro-tumorigenic cytokines (CCL2, CCL9, IL13, IL23, etc.) that recruit and polarize macrophages36,106,107,108,109. However, the precise mechanisms at play prior to immune suppression in the pro-inflammatory early stages of prostate carcinogenesis are not well established. Our data suggest that MYC downregulated targets, specifically in cancer cells, suppress the expression of inflammatory-associated genes. Second, what specific mechanisms are responsible for the switch from the pro-immunogenic signals to an immunosuppressive state despite ongoing MYC activation and signaling? While prior studies in other tumor types have reported that MYC activation promotes an immunosuppressive TME37,38,39,40,41, our findings show that MYC activation in neoplastic precursors initially promotes a pro-inflammatory TME that progresses to an immunosuppressive TME following disease evolution from neoplastic precursors and early invasive adenocarcinoma to more advanced adenocarcinoma during malignant progression. With the identification of the specific types of changes to the immune and stromal microenvironment associated with this switch in this study, it will be possible to uncover the mechanisms responsible for these changes in future studies.
These alterations induced by MYC-driven prostate cancer may also represent convergent sequelae of other cancer drivers, perhaps via cross-talk through MYC activation. For example, a recent scRNA-seq study in mouse models of prostate cancer driven by PTEN, RB1, and/or TP53 deletion reported upregulation of JAK/STAT inflammatory signaling was a key instigator of lineage plasticity in prostate cancer cells110. Notably, adenocarcinoma cells in this model were also upregulated for inflammatory response, interferon signaling, and TNFA signaling via NF-κB.
In conclusion, MYC activation is a common denominator across all molecular subtypes of prostate cancer. MYC activation in luminal cells, directly and indirectly, reprograms the local TME, with widespread effects on neighboring epithelial, stromal, and immune cell types. These changes in the TME of prostate cancer may be informative for future therapeutic strategies. For example, there is an ongoing clinical trial with antibodies targeting TREM2 macrophages in cancer111. Our study shows that one of the key changes in the TME of MYC-driven prostate cancer includes the enrichment of TREM2 macrophages with implications for driving an immunosuppressive and reactive stromal microenvironment. Additionally, several ongoing pre-clinical and clinical trials targeting MYC may ultimately lead to a new treatment option for prostate cancer patients112.
Methods
Human prostatectomy tissue punches
Prostate tissue specimens were collected from men diagnosed with primary prostate cancer undergoing radical prostatectomy at Johns Hopkins University (Supplementary Data 1) and consented under IRB approved protocols NA_00048544 and NA_00087094. Prostatectomies were sectioned fresh from apex to base, and fresh tissue samples were collected using an 8 mm punch biopsy tool from each zone of the prostate, as well as apparent sites of tumor (approximately 4 punches per subject). As described below, for each tissue punch, a portion of the punched tissue core was used to cut frozen sections for H&E staining, immunostaining (PIN4, ERG, PTEN, and MYC), and in situ hybridization for human MYC mRNA. The adjacent portion of the tissue punch was processed for scRNA-seq.
Mouse prostate dissection
All mice were handled in accordance under the guidelines of the Johns Hopkins University Animal Care and Use Committee (JHU ACAUC) and the approved protocols MO23M187 and MO22M223. Mice were housed under a 12 hour light and dark cycle (on at 6 AM and off at 6 PM), temperatures between 20 to 25 °C, and humidity level between 30% to 70%. The prostate cancer GEMMs used in these studies were euthanised before reaching the maximum tumor burden guidelines approved by the JHU ACAUC. Specifically, GEMMs were euthanized no later than ten months of age, when any dimension of the solid tumor reached 2.0 cm, or when mice displayed signs of clinical complications, ulceration, or ascites. Wildtype C57BL/6 J and FVB/NJ mice were purchased from Jackson Laboratory. The FVB Hi-Myc mice, FVB-Tg(ARR2/Pbsn-MYC)7Key, were purchased from NCI Mouse Repository. The laboratory of Brian Simons provided the C57BL/6 J Hi-Myc mice (B6 Hi-Myc). The C57BL/6 J Hi-Myc mice were generated by backcrossing the Hi-Hyc transgenic mice originally developed in FVB/NJ mice68 with C57BL/6 J for more than ten generations69. Hi-Myc Mice were maintained until they reached six months and ten months (+/− 6 weeks) of age. Individual prostate lobes were dissected, and scRNA-seq libraries for each lobe were prepared separately. The Bieberich laboratory provided the BMPC mice93. Tissues from BMPC mice were dissected from 3-month-old prostate lobes and two lymph node metastatic lesions from a 6-month-old BMPC mouse. All mice were euthanized using carbon dioxide asphyxiation, and the urogenital tract was removed and placed into a petri dish containing 50 ml of Hanks’ Balanced Salt Solution (HBSS, Gibco 14175-079). Under a dissection microscope, adipose tissues were removed from the urogenital tract to isolate the prostate. The four pairs of lobes (anterior, dorsal, lateral, and ventral lobes) were separated and dissected from the urethra by forceps. One pair was dissected in half for single-cell RNA-Seq, and the other half was formalin-fixed & paraffin-embedded (FFPE). The remaining lobe pair was frozen.
Dissociation of mouse and human prostate tissues
Dissected tissues were processed as previously described72. Briefly, tissues were minced with razor blades and digested in 0.25% Trypsin-EDTA (Gibco 25200-072) for 10 min at 37 °C, followed by incubation for 2.5 hours at 37 °C with gentle agitation in DMEM containing 10% FBS, 1 mg/mL Collagenase Type I (Gibco 17100-017), and 0.1 mg/mL of DNase I (Roche 10104159001). Digested tissues were centrifuged at 400 x g for 5 minutes, washed with HBSS, and further incubated in 0.25% Trypsin-EDTA for 10 min at 37 °C. Cells were suspended in DMEM containing 10% FBS and 0.4 mg/mL of DNase I and filtered through a 40 μm cell strainer.
Single-cell RNA-sequencing and data pre-processing
Libraries for scRNA-seq were prepared using the 10x Genomics Chromium Single Cell 3’ Library and Gel bead Kit V2 (CG00052_RevF) according to the manufacturer’s protocol for each dissected mouse prostate lobe and each human prostatectomy tissue punch. The cDNA libraries were sequenced (150 bp paired-end) on the Illumina HiSeqX platform. Each sequenced library was demultiplexed to FASTQ files using Cell Ranger (10x Genomics). For human tissues, Cell Ranger (version 3.2.0) count pipeline was used to align reads to the GRCh38 transcriptome and create a gene-by-cell count matrix. The human sequencing files can be accessed on NCBI dbGaP (accession phs003480.v1.p1). For mouse tissues, Cell Ranger (version 2.2.0) count pipeline was used to align reads to the mm10 mouse transcriptome and create a gene-by-cell count matrix. To align transgene expression, Kallisto quant (v0.46.0) was used to align the reads against human MYC and mouse Myc reference sequences. Samtools (v0.1.19) was used to extract alignments from BAM files for each sample. An in-house Perl script was used to map the alignments to the original FASTQ reads and create a feature-barcode count matrix. The Cell Ranger matrices were merged with the custom count matrices containing human MYC and mouse Myc. The mouse sequencing files can be accessed on NCBI GEO (GSE228945). For both mouse and human datasets, Seurat (version 4.3.0)113 was used to pre-process the data further. Cells with mitochondrial gene percentages >25% were removed.
Dimensionality reduction and cell type identification
Normalization and variance stabilization of scRNA-seq data were performed using Seurat (version 4.3.0), specifically SCTransform (version 2) with gamma-Poisson generalized linear model fitting113,114,115,116,117. For dimensionality reduction and clustering analysis, a principal component analysis was performed, and a range of 30 to 50 principal components was used to compute the Uniform Manifold Approximation & Projection (UMAP) dimensions and perform Louvain clustering at varying resolutions, ranging from 0.1 to 1.0. Differential gene expression analysis of previously characterized cell type-specific genes was used to identify the cell type for each cluster72,118. Mouse prostate data for aggregated 6-month WT and Hi-Myc scRNA-seq data were split by strain and normalized by SCTransform. To integrate datasets split by strain, 2000 anchor features were chosen, canonical correlation analysis was selected for dimensionality reduction with 20 dimensions used to specify the neighbor search space, and 200 neighbors (k-filter) were specified for anchor filtering. Mouse prostate scRNA-seq data for FVB/NJ dorsal and lateral lobes of 6-month and 10-month mice were split by individual samples, and each sample library was normalized by SCTransform. A WT sample and a Hi-Myc sample from 10-month-old mice were selected as references for integration. Similarly, BMPC samples were integrated with all scRNA-seq libraries generated from FVB/NJ WT and Hi-Myc samples, with the 10-month-old Hi-Myc and WT samples as references.
Differential gene expression analysis
To identify marker genes, differential gene expression analysis of all clusters was performed using Seurat (version 4.3.0) in merged datasets without integration. Genes considered in the analysis needed to be expressed in at least 25% of cells in a cluster with a minimum natural log fold change of 0.25. All outputs of this analysis are included as supplementary data. The gene filtering parameters used to generate each heatmap or dot plot from differential gene expression analysis are described in the legends.
Inferred copy number variation (inferCNV)
We applied computational methodologies to infer copy number variations (CNVs) in our prostatectomy scRNA-seq data. In this approach, transcripts from neighboring genes on chromosomes with consistent up- or down-regulated expression are inferred to have a genomic gain or loss, respectively119. For each subject, luminal cells from the transition zone were used as a reference to infer CNVs in the luminal cells of peripheral zone tissue punches. To determine which genes were used for inferCNV analysis, genes with a mean count of 0.1 were selected. The residual expression intensities were denoised using default dynamic thresholding. A three-state (deletion, neutral, amplification) hidden Markov model-based method was used to predict CNVs. Cells were grouped based on similar CNV patterns using the “subclusters” analysis mode with the default Leiden method to partition hierarchical clustering trees. Subclusters with inferred CNVs previously reported in prostate cancer were identified and incorporated in the Seurat object as metadata to indicate predicated cancer cells on the UMAP19,44,45,46,47,48.
Pathway and gene set enrichment analysis (GSEA)
Gene sets from the Molecular Signatures Database (msigdb), including the Hallmark Collection, KEGG, Reactome, and C2 curated gene sets, were imported using the R package msigdbr (version 7.5.1)65,120,121,122,123,124. The Wilcoxon rank test was performed on genes differentially expressed between clusters using Presto (version 1.0)125. The specific comparisons are indicated for each plot and figure legend. Genes were ranked by the statistical output, and GSEA was performed using the R package fgsea to determine normalized enrichment scores (NES) and associated adjusted p-values (version 1.22.0)126. In the mouse scRNA-seq dataset, the Pearson correlation of human MYC transgene was determined for all genes. We used the Investigate Gene Sets tool, available on msigdb, to compute significant overlaps between genes that positively correlated with human MYC transgene expression and biological pathways annotated by KEGG and Reactome.
Gene signature analysis
Gene signature scores were computed using UCell (version 2.0.1) to evaluate cluster-specific marker gene expression127. Gene lists for gene signature analysis included the leading edge genes from GSEA of MYC V1 targets comparing luminal and cancer clusters, regulatory T cell markers (IL2RA, FOXP3, CD4, IKZF2, CCR4, CTLA4), and Luminal MYC 1 cluster marker genes (Supplementary Data 9). The resulting scores ranging from 0 to 1 for each cell are indicated in UMAP plots.
Immunohistochemistry (IHC)
Frozen and formalin-fixed and paraffin-embedded (FFPE) tissues were immunostained using the Ventana Discovery Ultra Autostainer IHC (Roche Diagnostics, Basel, Switzerland). Tissue slides were steamed for 32 minutes in Cell Conditioning 1 (CC1) solution (Roche Diagnostics, Cat. No. 950-500) for antigen retrieval. For multiplex PIN4 cocktail staining, tissues were incubated with antibodies diluted 1:50, including CK903 (Enzo, Cat. No. ENZ-C34903) for 40 minutes, P63 (Biocare, Cat. No. CM163A) for 40 minutes, and AMACR (Zeta, Cat No. Z2001L) for 32 minutes. Chromogenic staining of target antigens was performed using the DISCOVERY anti-HQ HRP kit (903/p63 cocktail) and the Discovery anti-HQ-NP kit (AMACR). Similarly, PTEN, ERG, CD3, F4/80, FOXp3 (Cell Signaling, Cat. No 12653) staining was performed on the Ventana Discovery Ultra Autostainer IHC. FFPE slides were steamed for 48 minutes in Cell Conditioning 1 (CC1) solution (Roche Diagnostics, Cat. No. 950-500) for antigen retrieval. Tissues were incubated with primary antibody using the following conditions, PTEN (Cell Signaling, Cat. No. 9188) diluted 1:100 with 1-hour incubation, ERG (Roche, Cat. No 6478450001) undiluted with 12 minutes of incubation, CD3 (FisherSci, Cat. No RM9107S) diluted 1:200 with 40 minutes of incubation, F4/80 (Cell Signaling, Ca. No 70076) diluted 1:200 with 36 minutes of incubation, and FOXp3 Cell Signaling (Cat. No 12653) diluted 1:250 with 1-hour incubation. Chromogenic staining of target antigens was performed using the DISCOVERY anti-HQ HRP kit.
Chromogenic in situ hybridization (CISH)
To prepare frozen and FFPE tissue for CISH staining, slides were incubated for 30 minutes at 60 °C and deparaffinized by incubating slides at room temperature (RT) for 10 minutes in xylene twice and then subsequently incubated in 100% ethanol twice and finally left to air dry. Hydrogen peroxide solution was added to the slides for 10 minutes at RT. Slides were steamed in 1 x RNAscope Target retrieval reagent at 100 °C for 18 minutes, followed by protease plus digestion for 30 minutes at 40 °C to allow target accessibility. The following probes were used for CISH staining, including Hs-MYC (RNAscope, Cat No. 311761) and Mm-Ly6d (RNAscope, Cat No. 532071) in mouse tissues. Probes were added to slides and incubated in the HybEZ TM Oven for 2 hours at 40 °C. Frozen human prostatectomy tissue was cut into 5 μm sections and fixed in 10% neutral buffered formalin for 15 minutes at 4 °C and dehydrated in alcohol. Hydrogen peroxide solution was added to the slides for 10 minutes at RT. Slides were steamed in 1 x RNAscope Target retrieval reagent at 100 °C for 5 minutes, followed by protease IV digestion for 10 minutes at RT to allow target accessibility. Hs-KRT6A (RNAscope, Cat No. 520721) probe was added to slides and incubated in the HybEZ TM Oven for 2 hours at 40 °C. Signal amplification and detection were performed using the RNAscope 2.5 HD Reagent kit-Brown (Cat.No 322300, ACD) according to the manufacturer’s protocol. The probe signal was detected with DAB. Slides were counter-stained with 50% Gill’s Hematoxylin for 2 min, rinsed in 0.02% ammonia water for 15 seconds, treated in 100% ethanol and xylene, and coverslipped with Cytoseal mounting medium.
Cell communication network analysis
A computational strategy was used to construct inferred ligand-receptor-transcription factor cell communication networks among cell clusters in our mouse scRNA-seq datasets. Transcription factor activation scores were determined using the Single-Cell rEgulatory Network Inference and Clustering (SCENIC) pipeline90. A Docker image provided by developers was used to run the Python implementation of SCENIC. The R-based package Domino was used to compile ligand-receptor interactions using the CellphoneDB repository128, calculate the transcription factor activity enrichment for each cluster based on SCENIC-defined activation scores and correlate receptor expression and transcription factor scores129. The parameters for constructing cell communication networks included a maximum of 10 transcription factors for each cluster with a minimum p-value of 1 × 10−4 and a correlation ≥ 0.25 for receptor expression and transcription factor activation.
Statistics and reproducibility
For both human and mouse tissues, approximately 5000 cells were sequenced per sample to capture all cell types that constitute the tissue microenvironment, including scarce cell types. For the prostatectomy scRNA-seq and immunostaining studies, ten subjects were randomly selected to sufficiently capture the well-known tumor heterogeneity of prostate cancer (Fig. 1A). For the GEMM experiments, approximately 10,000 or more cells were collectively represented for each comparison by genotype, strain, and time point. At least two mice were included in each group, with each prostate lobe dissected (4 samples per mouse) for most mice (Fig. 3A), except for the 10-month-old Hi-Myc mice, where the dorsal and lateral lobes were combined. At ten months, a large tumor forms in both the dorsal and lateral lobes of the Hi-Myc GEMM. No statistical method was used to predetermine sample sizes. GSEA was performed to identify significantly upregulated and down-regulated pathways. The statistics were derived using the fgsea implementation of a two-tailed GSEA. Nominal p-values are based on an adaptive, multilevel splitting Monte Carlo scheme. The adjusted p-value statistic was derived from multiple hypothesis testing using the Benjamini-Hochberg procedure across all gene sets considered. To identify marker genes, differential gene expression analysis of all clusters was performed using Seurat (version 4.3.0) in merged datasets without integration. The statistics were derived using a two-tailed Wilcoxon Rank-Sum test and adjusted for multiple hypothesis testing using the Bonferroni correction. Differences in cell type proportions across groups were determined using RAISIN (version 1.0). Statistics were derived using linear regression analysis (limma), and the adjusted p-value was generated from a two-sided t-test and adjusted for multiple hypothesis testing using the Benjamini-Hochberg procedure. All statistical analyses were performed in R (version 4.3.2). No data were excluded from all analyses performed. However, poor-quality cells with > 25% mitochondrial ratio, contaminated from cell-free RNA, or mixed from the capture of multiple cells were filtered from scRNA-seq datasets.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The publicly available human prostate adenocarcinoma RNA-seq data used in this study are available in the cbioportal database TCGA Prostate Adenocarcinoma (Cell 2015) [https://www.cbioportal.org/study/summary?id=prad_tcga_pub]16. The publicly available mouse prostate and urethral scRNA-seq used to examine stromal and epithelial cell types are available on the CZ CellxGene platform [https://cellxgene.cziscience.com/collections/fbc5881f-1ee3-4ffe-8095-35e15e1a08fc]130. The human prostatectomy data generated in this study have been deposited in the NCBI dbGaP database under accession code phs003480.v1.p1 and access to data can be requested using the following link: [https://www.ncbi.nlm.nih.gov/projects/gap/cgi-bin/study.cgi?study_id=phs003480.v1.p1]. To comply with the NIH Genomic Data Sharing (GDS) policies and protect the interests of research participants, access to the human sequencing data is controlled via dbGaP. Investigators who are permanent employees of their institution, tenure track professor or senior scientist, are eligible to request data authorization, which is approved through the Data Access Committee (DAC). Response to data requests will be completed within one month. Approved investigators will have access to data for up to one year and have the option to renew access. The mouse prostate data are deposited in the NCBI GEO database under accession code GSE228945. All processed data frames are also available as R objects and can be accessed as a SharePoint link described in the GitHub repository [https://github.com/YegnaLab/MYC_Prostate_TME]. The remaining data are available within the Article, Supplementary Information, or Supplementary Data.
Code availability
The code for the analysis performed in this study can be accessed as Rmd files or knitted HTML files on a GitHub repository [https://github.com/YegnaLab/MYC_Prostate_TME], which includes a SharePoint link to access R objects and data frames to run the code.
References
Haffner, M. C. et al. Genomic and phenotypic heterogeneity in prostate cancer. Nat. Rev. Urol. 18, 79–92 (2021).
Mitchell, T. & Neal, D. E. The genomic evolution of human prostate cancer. Br. J. Cancer 113, 193–198 (2015).
Spratt, D. E., Zumsteg, Z. S., Feng, F. Y. & Tomlins, S. A. Translational and clinical implications of the genetic landscape of prostate cancer. Nat. Rev. Clin. Oncol. 13, 597–610 (2016).
Shen, M. M. & Abate-Shen, C. Molecular genetics of prostate cancer: new prospects for old challenges. Genes Dev. 24, 1967–2000 (2010).
Sandhu, S. et al. Prostate cancer. Lancet 398, 1075–1090 (2021).
Zhou, M. High-grade prostatic intraepithelial neoplasia, PIN-like carcinoma, ductal carcinoma, and intraductal carcinoma of the prostate. Mod. Pathol. 31, S71–S79 (2018).
Koh, C. M. et al. Alterations in nucleolar structure and gene expression programs in prostatic neoplasia are driven by the MYC oncogene. Am. J. Pathol. 178, 1824–1834 (2011).
McNeal, J. E. & Bostwick, D. G. Intraductal dysplasia: a premalignant lesion of the prostate. Hum. Pathol. 17, 64–71 (1986).
Jerónimo, C. et al. Quantitation of GSTP1 methylation in non-neoplastic prostatic tissue and organ-confined prostate adenocarcinoma. J. Natl Cancer Inst. 93, 1747–1752 (2001).
Emmert-Buck, M. R. et al. Allelic Loss on Chromosome 8p12–21 in Microdissected Prostatic Intraepithelial Neoplasia. Cancer Res. 55, 2959–2962 (1995).
Häggman, M. J., Wojno, K. J., Pearsall, C. P. & Macoska, J. A. Allelic loss of 8p sequences in prostatic intraepithelial neoplasia and carcinoma. Urology 50, 643–647 (1997).
Bostwick, D. G. et al. Independent origin of multiple foci of prostatic intraepithelial neoplasia: comparison with matched foci of prostate carcinoma. Cancer 83, 1995–2002 (1998).
Graham, M. K. & Meeker, A. Telomeres and telomerase in prostate cancer development and therapy. Nat. Rev. Urol. 14, 607–619 (2017).
Gupta, H. et al. Progressive Spreading of DNA Methylation in the GSTP1 Promoter CpG Island across Transitions from Precursors to Invasive Prostate Cancer. Cancer Prev. Res. 16, 449–460 (2023).
Tomlins, S. A. et al. Recurrent fusion of TMPRSS2 and ETS transcription factor genes in prostate cancer. Science 310, 644–648 (2005).
Cancer Genome Atlas Research Network. The Molecular Taxonomy of Primary Prostate Cancer. Cell 163, 1011–1025 (2015).
Taylor, B. S. et al. Integrative genomic profiling of human prostate cancer. Cancer Cell 18, 11–22 (2010).
Trabzonlu, L. et al. Molecular Pathology of High-Grade Prostatic Intraepithelial Neoplasia: Challenges and Opportunities. Cold Spring Harb. Perspect. Med. 9, a030403 (2019).
De Marzo, A. M., Marchi, V. L., Epstein, J. I. & Nelson, W. G. Proliferative inflammatory atrophy of the prostate: implications for prostatic carcinogenesis. Am. J. Pathol. 155, 1985–1992 (1999).
Sfanos, K. S., Yegnasubramanian, S., Nelson, W. G. & De Marzo, A. M. The inflammatory microenvironment and microbiome in prostate cancer development. Nat. Rev. Urol. 15, 11–24 (2018).
De Marzo, A. M. et al. A working group classification of focal prostate atrophy lesions. Am. J. Surg. Pathol. 30, 1281–1291 (2006).
Putzi, M. J. & De Marzo, A. M. Morphologic transitions between proliferative inflammatory atrophy and high-grade prostatic intraepithelial neoplasia. Urology 56, 828–832 (2000).
Wang, W., Bergh, A. & Damber, J.-E. Morphological transition of proliferative inflammatory atrophy to high-grade intraepithelial neoplasia and cancer in human prostate. Prostate 69, 1378–1386 (2009).
Gurel, B. et al. Nuclear MYC protein overexpression is an early alteration in human prostate carcinogenesis. Mod. Pathol. 21, 1156–1167 (2008).
Koh, C. M. et al. MYC and Prostate Cancer. Genes Cancer 1, 617–628 (2010).
Mani, R. S. et al. Inflammation-Induced Oxidative Stress Mediates Gene Fusion Formation in Prostate Cancer. Cell Rep. 17, 2620–2631 (2016).
Shrestha, E. et al. Oncogenic gene fusions in nonneoplastic precursors as evidence that bacterial infection can initiate prostate cancer. Proc. Natl Acad. Sci. USA 118, e2018976118 (2021).
De Marzo, A. M., Haffner, M. C., Lotan, T. L., Yegnasubramanian, S. & Nelson, W. G. Premalignancy in Prostate Cancer: Rethinking What we Know. Cancer Prev. Res. 9, 648–656 (2016).
Tsujimoto, Y., Takayama, H., Nonomura, N., Okuyama, A. & Aozasa, K. Postatrophic hyperplasia of the prostate in Japan: histologic and immunohistochemical features and p53 gene mutation analysis. Prostate 52, 279–287 (2002).
Shah, R., Mucci, N. R., Amin, A., Macoska, J. A. & Rubin, M. A. Postatrophic hyperplasia of the prostate gland: neoplastic precursor or innocent bystander? Am. J. Pathol. 158, 1767–1773 (2001).
Stultz, J. & Fong, L. How to turn up the heat on the cold immune microenvironment of metastatic prostate cancer. Prostate Cancer Prostatic Dis. 24, 697–717 (2021).
Venturini, N. J. & Drake, C. G. Immunotherapy for Prostate Cancer. Cold Spring Harb. Perspect. Med. 9, a030627 (2019).
Strasner, A. & Karin, M. Immune Infiltration and Prostate Cancer. Front. Oncol. 5, 128 (2015).
Wojno, K. J. & Epstein, J. I. The utility of basal cell-specific anti-cytokeratin antibody (34 beta E12) in the diagnosis of prostate cancer. A review of 228 cases. Am. J. Surg. Pathol. 19, 251–260 (1995).
Barron, D. A. & Rowley, D. R. The reactive stroma microenvironment and prostate cancer progression. Endocr.-Relat. Cancer 19, R187–R204 (2012).
Dhanasekaran, R. et al. The MYC oncogene - the grand orchestrator of cancer growth and immune evasion. Nat. Rev. Clin. Oncol. 19, 23–36 (2022).
Li, J., Dong, T., Wu, Z., Zhu, D. & Gu, H. The effects of MYC on tumor immunity and immunotherapy. Cell Death Discov. 9, 103 (2023).
Bisso, A. et al. Cooperation Between MYC and β-Catenin in Liver Tumorigenesis Requires Yap/Taz. Hepatology 72, 1430–1443 (2020).
Sun, S. et al. Nuclear Aurora kinase A triggers programmed death-ligand 1-mediated immune suppression by activating MYC transcription in triple-negative breast cancer. Cancer Commun. 41, 851–866 (2021).
Ruiz de Galarreta, M. et al. β-Catenin Activation Promotes Immune Escape and Resistance to Anti-PD-1 Therapy in Hepatocellular Carcinoma. Cancer Discov. 9, 1124–1141 (2019).
Li, Q. et al. The role of exosomal miR-375-3p: A potential suppressor in bladder cancer via the Wnt/β-catenin pathway. FASEB J. 34, 12177–12196 (2020).
Ali, A. et al. Prostate zones and cancer: lost in transition? Nat. Rev. Urol. 19, 101–115 (2022).
Tolkach, Y. & Kristiansen, G. The Heterogeneity of Prostate Cancer: A Practical Approach. Pathobiology 85, 108–116 (2018).
Lee, W.-H. et al. Cytidine methylation of regulatory sequences near the pi-class glutathione S-transferase gene accompanies human prostatic carcinogenesis. Proc. Natl Acad. Sci. 91, 11733–11737 (1994).
Brooks, J. D. et al. CG island methylation changes near the GSTP1 gene in prostatic intraepithelial neoplasia. Cancer Epidemiol. Biomark. Prev. 7, 531–536 (1998).
Porter, C. M. et al. Lactoferrin CpG Island Hypermethylation and Decoupling of mRNA and Protein Expression in the Early Stages of Prostate Carcinogenesis. Am. J. Pathol. 189, 2311–2322 (2019).
Bussemakers, M. J. et al. DD3: a new prostate-specific gene, highly overexpressed in prostate cancer. Cancer Res. 59, 5975–5979 (1999).
Luo, J. et al. Alpha-methylacyl-CoA racemase: a new molecular marker for prostate cancer. Cancer Res. 62, 2220–2226 (2002).
Henry, G. H. et al. A Cellular Anatomy of the Normal Adult Human Prostate and Prostatic Urethra. Cell Rep. 25, 3530–3542.e5 (2018).
Song, H. et al. Single-cell analysis of human primary prostate cancer reveals the heterogeneity of tumor-associated epithelial cell states. Nat. Commun. 13, 141 (2022).
Tuong, Z. K. et al. Resolving the immune landscape of human prostate at a single-cell level in health and cancer. Cell Rep. 37, 110132 (2021).
Huang, F. W. et al. Club-like cells in proliferative inflammatory atrophy of the prostate. J. Pathol. https://doi.org/10.1002/path.6149 (2023).
Perner, S. et al. TMPRSS2:ERG fusion-associated deletions provide insight into the heterogeneity of prostate cancer. Cancer Res. 66, 8337–8341 (2006).
Tomlins, S. A. et al. The role of SPINK1 in ETS rearrangement-negative prostate cancers. Cancer Cell 13, 519–528 (2008).
Gumuskaya, B. et al. Assessing the order of critical alterations in prostate cancer development and progression by IHC: further evidence that PTEN loss occurs subsequent to ERG gene fusion. Prostate Cancer Prostatic Dis. 16, 209–215 (2013).
Jamaspishvili, T. et al. Clinical implications of PTEN loss in prostate cancer. Nat. Rev. Urol. 15, 222–234 (2018).
Williams, J. L., Greer, P. A. & Squire, J. A. Recurrent copy number alterations in prostate cancer: an in silico meta-analysis of publicly available genomic data. Cancer Genet. 207, 474–488 (2014).
Cairns, P. et al. Frequent inactivation of PTEN/MMAC1 in primary prostate cancer. Cancer Res. 57, 4997–5000 (1997).
Li, J. et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 275, 1943–1947 (1997).
Teng, D. H. et al. MMAC1/PTEN mutations in primary tumor specimens and tumor cell lines. Cancer Res. 57, 5221–5225 (1997).
Vlietstra, R. J., van Alewijk, D. C., Hermans, K. G., van Steenbrugge, G. J. & Trapman, J. Frequent inactivation of PTEN in prostate cancer cell lines and xenografts. Cancer Res. 58, 2720–2723 (1998).
Qiu, X. et al. MYC drives aggressive prostate cancer by disrupting transcriptional pause release at androgen receptor targets. Nat. Commun. 13, 2559 (2022).
Guo, H. et al. Androgen receptor and MYC equilibration centralizes on developmental super-enhancer. Nat. Commun. 12, 7308 (2021).
Sena, L. A. et al. Androgen receptor activity in prostate cancer dictates efficacy of bipolar androgen therapy through MYC. J. Clin. Invest. 132, e162396 (2022).
Zeller, K. I., Jegga, A. G., Aronow, B. J., O’Donnell, K. A. & Dang, C. V. An integrated database of genes responsive to the Myc oncogenic transcription factor: identification of direct genomic targets. Genome Biol. 4, R69 (2003).
Arlt, A. & Schäfer, H. Role of the immediate early response 3 (IER3) gene in cellular stress response, inflammation and tumorigenesis. Eur. J. Cell Biol. 90, 545–552 (2011).
Chen, Y. et al. Immediate early response 3 gene promotes aggressive progression and autophagy of AML by negatively regulating AKT/mTOR. Transl. Oncol. 35, 101711 (2023).
Ellwood-Yen, K. et al. Myc-driven murine prostate cancer shares molecular features with human prostate tumors. Cancer Cell 4, 223–238 (2003).
Simons, B. W. et al. A mouse model of prostate cancer bone metastasis in a syngeneic immunocompetent host. Oncotarget 10, 6845–6854 (2019).
Ellis, L. et al. Generation of a C57BL/6 MYC-driven mouse model and cell line of prostate cancer. Prostate 76, 1192–1202 (2016).
Iwata, T. et al. MYC Overexpression Induces Prostatic Intraepithelial Neoplasia and Loss of Nkx3.1 in Mouse Luminal Epithelial Cells. PLoS ONE 5, e9427 (2010).
Graham, M. K. et al. Single-cell atlas of epithelial and stromal cell heterogeneity by lobe and strain in the mouse prostate. Prostate https://doi.org/10.1002/pros.24460 (2022).
Crowley, L. et al. A single-cell atlas of the mouse and human prostate reveals heterogeneity and conservation of epithelial progenitors. eLife 9, e59465 (2020).
Dang, C. V. et al. The c-Myc target gene network. Semin. Cancer Biol. 16, 253–264 (2006).
Chen, J. et al. MYC-driven increases in mitochondrial DNA copy number occur early and persist throughout prostatic cancer progression. bioRxiv 8, e169868 (2023).
Li, F. et al. Myc stimulates nuclearly encoded mitochondrial genes and mitochondrial biogenesis. Mol. Cell. Biol. 25, 6225–6234 (2005).
Harris, B. D., Schreiter, J., Chevrier, M., Jordan, J. L. & Walter, M. R. Human interferon-ϵ and interferon-κ exhibit low potency and low affinity for cell-surface IFNAR and the poxvirus antagonist B18R. J. Biol. Chem. 293, 16057–16068 (2018).
LaFleur, D. W. et al. Interferon-kappa, a novel type I interferon expressed in human keratinocytes. J. Biol. Chem. 276, 39765–39771 (2001).
Pittet, M. J., Michielin, O. & Migliorini, D. Clinical relevance of tumour-associated macrophages. Nat. Rev. Clin. Oncol. 19, 402–421 (2022).
Molgora, M. et al. TREM2 Modulation Remodels the Tumor Myeloid Landscape Enhancing Anti-PD-1 Immunotherapy. Cell 182, 886–900.e17 (2020).
Wong, H. Y. et al. Single cell analysis of cribriform prostate cancer reveals cell intrinsic and tumor microenvironmental pathways of aggressive disease. Nat. Commun. 13, 6036 (2022).
Joseph, D. B. et al. Single-cell analysis of mouse and human prostate reveals novel fibroblasts with specialized distribution and microenvironment interactions. J. Pathol. 255, 141–154 (2021).
Wu, F. & Chakravarti, S. Differential expression of inflammatory and fibrogenic genes and their regulation by NF-kappaB inhibition in a mouse model of chronic colitis. J. Immunol. 179, 6988–7000 (2007).
Mak, K. M., Png, C. Y. M. & Lee, D. J. Type V Collagen in Health, Disease, and Fibrosis. Anat. Rec. 299, 613–629 (2016).
Parra, E. R. et al. Interstitial and vascular type V collagen morphologic disorganization in usual interstitial pneumonia. J. Histochem. Cytochem. 54, 1315–1325 (2006).
Galbo, P. M. Jr, Zang, X. & Zheng, D. Molecular Features of Cancer-associated Fibroblast Subtypes and their Implication on Cancer Pathogenesis, Prognosis, and Immunotherapy Resistance. Clin. Cancer Res. 27, 2636–2647 (2021).
Abbas, O. & Mahalingam, M. Desmoplasia: not always a bad thing. Histopathology 58, 643–659 (2011).
DeClerck, Y. A. Desmoplasia: a response or a niche? Cancer Discov. 2, 772–774 (2012).
Chandler, C., Liu, T., Buckanovich, R. & Coffman, L. G. The double edge sword of fibrosis in cancer. Transl. Res. 209, 55–67 (2019).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Meng, X.-M., Nikolic-Paterson, D. J. & Lan, H. Y. TGF-β: the master regulator of fibrosis. Nat. Rev. Nephrol. 12, 325–338 (2016).
Zhao, M. et al. Targeting fibrosis: mechanisms and clinical trials. Signal Transduct. Target. Ther. 7, 1–21 (2022).
Hubbard, G. K. et al. Combined MYC Activation and Pten Loss Are Sufficient to Create Genomic Instability and Lethal Metastatic Prostate Cancer. Cancer Res. 76, 283–292 (2016).
Tomlins, S. A. et al. Distinct classes of chromosomal rearrangements create oncogenic ETS gene fusions in prostate cancer. Nature 448, 595–599 (2007).
Han, B. et al. Fluorescence in situ hybridization study shows association of PTEN deletion with ERG rearrangement during prostate cancer progression. Mod. Pathol. 22, 1083–1093 (2009).
Epstein, J. I. Prostate cancer grading: a decade after the 2005 modified system. Mod. Pathol. 31, S47–S63 (2018).
Bailar, J. C. 3rd, Mellinger, G. T. & Gleason, D. F. Survival rates of patients with prostatic cancer, tumor stage, and differentiation-preliminary report. Cancer Chemother. Rep. 50, 129–136 (1966).
Mellinger, G. T. Prognosis of prostatic carcinoma. Recent Results Cancer Res. 60, 61–72 (1977).
Chodak, G. W., Keller, P. & Schoenberg, H. W. Assessment of Screening for Prostate Cancer Using the Digital Rectal Examination. J. Urol. 141, 1136–1138 (1989).
Hirz, T. et al. Dissecting the immune suppressive human prostate tumor microenvironment via integrated single-cell and spatial transcriptomic analyses. Nat. Commun. 14, 663 (2023).
Dhanasekaran, R. et al. MYC Overexpression Drives Immune Evasion in Hepatocellular Carcinoma That Is Reversible through Restoration of Proinflammatory Macrophages. Cancer Res. 83, 626–640 (2023).
Wu, S.-Y. et al. MYC suppresses STING-dependent innate immunity by transcriptionally upregulating DNMT1 in triple-negative breast cancer. J. Immunother. Cancer 9, e002528 (2021).
Casey, S. C. et al. MYC regulates the antitumor immune response through CD47 and PD-L1. Science 352, 227–231 (2016).
Smith, B. A. H. et al. MYC-driven synthesis of Siglec ligands is a glycoimmune checkpoint. Proc. Natl Acad. Sci. USA 120, e2215376120 (2023).
Ramachandran, P. et al. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature 575, 512–518 (2019).
Zimmerli, D. et al. MYC promotes immune-suppression in triple-negative breast cancer via inhibition of interferon signaling. Nat. Commun. 13, 6579 (2022).
Kortlever, R. M. et al. Myc Cooperates with Ras by Programming Inflammation and Immune Suppression. Cell 171, 1301–1315.e14 (2017).
Braun, J., Felsher, D. W. & Goodglick, L. A. c-myc, MHCI, and NK resistance in immunodeficiency lymphomas. Ann. N. Y. Acad. Sci. 651, 467–469 (1992).
Versteeg, R. et al. c-myc down-regulates class I HLA expression in human melanomas. EMBO J. 7, 1023–1029 (1988).
Chan, J. M. et al. Lineage plasticity in prostate cancer depends on JAK/STAT inflammatory signaling. Science 377, 1180–1191 (2022).
Patnaik, A. et al. A phase 1a dose-escalation study of PY314, a TREM2 (Triggering Receptor Expressed on Macrophages 2) targeting monoclonal antibody. J. Clin. Orthod. 40, 2648–2648 (2022).
Llombart, V. & Mansour, M. R. Therapeutic targeting of ‘undruggable’ MYC. EBioMedicine 75, 103756 (2022).
Stuart, T. et al. Comprehensive Integration of Single-Cell Data. Cell 177, 1888–1902.e21 (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Ahlmann-Eltze, C. & Huber, W. glmGamPoi: fitting Gamma-Poisson generalized linear models on single cell count data. Bioinformatics 36, 5701–5702 (2021).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Franzén, O., Gan, L.-M. & Björkegren, J. L. M. PanglaoDB: a web server for exploration of mouse and human single-cell RNA sequencing data. Database 2019, baz046 (2019).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Dolgalev. msigdbr: MSigDB gene sets for multiple organisms in a tidy data format. R package version.
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Gillespie, M. et al. The reactome pathway knowledgebase 2022. Nucleic Acids Res. 50, D687–D692 (2022).
Korsunsky, I., Nathan, A., Millard, N. & Raychaudhuri, S. Presto scales Wilcoxon and auROC analyses to millions of observations. bioRxiv 653253 https://doi.org/10.1101/653253 (2019).
Sergushichev. An algorithm for fast preranked gene set enrichment analysis using cumulative statistic calculation. bioRxivorg.
Andreatta, M. & Carmona, S. J. UCell: Robust and scalable single-cell gene signature scoring. Comput. Struct. Biotechnol. J. 19, 3796–3798 (2021).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: inferring cell-cell communication from combined expression of multi-subunit ligand-receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Cherry, C. et al. Computational reconstruction of the signalling networks surrounding implanted biomaterials from single-cell transcriptomics. Nat. Biomed. Eng. 5, 1228–1238 (2021).
Joseph, D. B. et al. Urethral luminal epithelia are castration-insensitive cells of the proximal prostate. Prostate 80, 872–884 (2020).
Acknowledgements
We thank the members of the Sidney Kimmel Comprehensive Cancer Center’s Experimental and Computational Genomics Core, supported by Cancer Center Support Grant P30CA006973 (W.G.N.), for support with the single-cell sequencing studies and data pre-processing and analysis. This work was supported in part by NIH/NCI grants P50CA058236 (W.G.N.), U01CA196390 (A.M.D.), P01CA247886, U54CA274370 (A.M.D., S.Y.), P50CA180995 (C.E.P. awarded to M.K.G.), DOD PCRP grant W81XWH-19-1-0450 (S.E.L.), and by the Prostate Cancer Foundation, The Allegheny Health Network Johns Hopkins Pilot Project Grant (S.Y., T.L.D.), The Patrick C. Walsh Fund (S.Y.), The Irving Hansen Foundation (S.Y.), The Commonwealth Foundation (S.Y.), and the Maryland Cigarette Restitution Fund (S.Y.).
Author information
Authors and Affiliations
Contributions
S.Y., A.M.D., T.L.D., and W.G.N. conceived the project ideas and aims. S.Y., A.M.D., B.W.S., C.H.B., and S.E.L. developed the methodology and created some of the GEMMs used in this study. M.K.G., R.W., R.C., B.W., and A.G. used various computational tools to write code for scRNA-seq data. M.K.G., R.W., A.V., B.A., A.G., Q.Z., J.H., P.S.S., X.P., N.C., J.L., J.M., and M.R. validated results and verified overall replication and reproducibility of the results and experimental outputs. M.K.G., R.W., R.C., and B.A. performed the formal analysis and investigation. B.W.S., C.J.B., W.G.N., S.E.L., T.L.D., A.M.D., and S.Y. provided resources, study materials, reagents, and samples. M.K.G., A.G., A.S., Y.Z., and K.S. performed data curation and managed code and data. M.K.G. wrote and prepared the original draft of the manuscript. R.W., W.G.N., S.E.L., T.L.D., A.M.D., and S.Y. reviewed and edited the manuscript. M.K.G. and R.W. performed data visualisation. S.Y. supervised the overall research effort. M.K.G., K.S., and A.M.D. managed and coordinated research activities.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
: Nature Communications thanks Dean Felsher, Luc Furic and Douglas Strand for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Graham, M.K., Wang, R., Chikarmane, R. et al. Convergent alterations in the tumor microenvironment of MYC-driven human and murine prostate cancer. Nat Commun 15, 7414 (2024). https://doi.org/10.1038/s41467-024-51450-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-51450-2
- Springer Nature Limited