Abstract
In animals, evolutionarily conserved Polycomb repressive complex 2 (PRC2) catalyzes histone H3 lysine 27 trimethylation (H3K27me3) and PRC1 functions in recruitment and transcriptional repression. However, the mechanisms underlying H3K27me3-mediated stable transcriptional silencing are largely unknown, as PRC1 subunits are poorly characterized in fungi. Here, we report that in the filamentous fungus Magnaporthe oryzae, the N-terminal chromodomain and C-terminal MRG domain of Eaf3 play key roles in facultative heterochromatin formation and transcriptional silencing. Eaf3 physically interacts with Ash1, Eed, and Sin3, encoding an H3K36 methyltransferase, the core subunit of PRC2, and a histone deacetylation co-suppressor, respectively. Eaf3 co-localizes with a set of repressive Ash1-H3K36me2 and H3K27me3 loci and mediates their transcriptional silencing. Furthermore, Eaf3 acts as a histone reader for the repressive H3K36me2 and H3K27me3 marks. Eaf3-occupied regions are associated with increased nucleosome occupancy, contributing to transcriptional silencing in M. oryzae. Together, these findings reveal that Eaf3 is a repressive H3K36me2 reader and plays a vital role in Polycomb gene silencing and the formation of facultative heterochromatin in fungi.
Similar content being viewed by others
Introduction
In eukaryotes, the repeating subunit of chromatin is the nucleosome, formed by 147 bp of DNA wrapped around a histone octamer containing two copies of each histone H2A, H2B, H3, and H41. The chromatin state of a target locus influences the accessibility of the transcriptional machinery through post-translational histone modifications and is closely associated with transcriptional activity2,3,4. Active and silent transcription at the chromatin level is accompanied by distinct post-translational modifications of histones or their combinations2,3. Active transcription is typically marked by high levels of histone acetylation or trimethylation of H3 lysine 4 and lysine 36 (H3K4me3 and H3K36me3), whereas repressed transcription is enriched with markers including H3K27me3 and H3K9me35. Notably, H3K27me3 serves as a hallmark of facultative heterochromatin, in which transcription can be reversibly turned on and off in response to environmental clues or during phases of cell differentiation and development2,6,7. Polycomb group proteins usually assemble with Polycomb repressive complex 1 (PRC1) and PRC2. PRC2 catalyzes H3K27me3, while PRC1 is involved in H3K27me3 recruitment and chromatin compaction in animals5,8,9. Although PRC2 is highly conserved in fungi, PRC1 subunits have been rarely identified so far7,8. Therefore, the mechanisms by which PRC2 recognizes H3K27me3 and mediates stable transcriptional repression remain largely unknown.
H3K36me is catalyzed by histone methyltransferase Set2 in yeast, an active histone marker of euchromatin essential for transcriptional regulation10, whereas two H3K36 methyltransferases, Ash1 and Set2, are responsible for H3K36me2/3 in many filamentous fungi11,12,13. The Trithorax group catalyzes both H3K36me and H3K4me, countering the repressive effects of Polycomb group complex-mediated H3K27me in yeast and higher eukaryotes5,6,14, whereas these results were different from that in filamentous fungi. In Neurospora crassa, Ash1-H3K36me2-marked chromatin can be further modified with H3K27me, and Ash1 modulates H3K27me2/3 accumulation11. Recently, we have found that Set2-catalyzed-H3K36me3 activates transcription, while Ash1-catalyzed-H3K36me2 co-localizes with H3K27me3 to repress transcription in Magnaporthe oryzae, which causes rice blast disease worldwide15. Ash1, rather than Set2, is required for the distribution of facultative heterochromatic modifications and coordinated maintenance of transcriptional silencing15. However, the detailed mechanisms by which Ash1 and Ash1-catalyzed-H3K36me2 contribute to transcriptional silencing, and the co-localization of Ash1-H3K36me2 with H3K27me3 require further elucidation.
The regulation of transcription by histone modifications, catalyzed by either histone methyltransferases or acetyltransferases, is mediated in part through the recruitment of specific histone readers5,16. H3K36me2/3, for instance, is recognized by chromodomain-containing EAF3/Mrg15, a subunit of NuA4 histone acetyltransferase complex and the Rpd3S histone deacetylase complex, which deacetylates histones and prevents spurious transcription initiation within gene bodies17. Mrg15 is also an integral subunit of the Ash1 complex in Drosophila, and its interaction enhances the catalytic activity of Ash118. In the plant pathogen Zymoseptoria tritici, an Eaf3 homolog has been reported to be important for Ash1 activity and required for transcriptional repression in facultative heterochromatin19. In addition, H3K36me is also recognized by a PWWP motif, which found in many nuclear chromatin-binding proteins and H3K36 methyltransferases10. However, the mechanisms by which repressive Ash1-H3K36me2 is recognized and specifically recruited to its targets remains unknown.
In this study, we find that the transcriptional regulator Eaf3 in M. oryzae, which contains an N-terminal chromodomain and a C-terminal MRG domain, plays a key role in the facultative heterochromatin formation and transcriptional silencing. Eaf3 co-localizes with a set of repressive H3K36me2 and H3K27me3 loci and mediates their transcriptional silencing by physically interacting with an H3K36 methyltransferase and a core subunit of PRC2. Eaf3 is also associated with a transcriptional corepressor, maintaining histone hypoacetylation at Ash1-H3K36me2 genes. Moreover, Eaf3 acts as a histone reader of repressive H3K36me2 and H3K27me3, and Eaf3-occupied regions associated with increased nucleosome occupancy, thereby coordinately contributing to transcriptional silencing in M. oryzae. Therefore, our findings reveal that Eaf3 plays important roles in facultative heterochromatin formation and Polycomb-mediated gene silencing in fungi.
Results
Eaf3 interacts with Ash1 in Magnaporthe oryzae
H3K36me is usually an active histone marker associated with transcriptional activation in eukaryotes4,10. However, studies have shown that Ash1-catalyzed H3K36me was associated with transcriptional silencing in filamentous fungi11,12,19. Recently, we have found that Ash1-catalyzed-H3K36me2 also co-localizes with H3K27me3 and is required for the normal distribution of H3K27me3 in M. oryzae15. However, several critical questions remain unanswered, including the mechanisms underlying the contribution of Ash1 and Ash1-catalyzed-H3K36me2 to transcriptional silencing, the specific recruitment of Ash1-H3K36me2 to its targets, and the co-localization of Ash1-H3K36me2 with H3K27me3.
To answer these questions, we performed affinity purification of an ASH1-Flag strain to identify the candidate interactors (Supplementary Fig. 1). As shown in Fig. 1a, two annotated and tandemly arranged proteins MGG_06716 and MGG_06717, containing a chromodomain (CD) and an MRG domain respectively, were captured (Fig. 1a, b). The two proteins are orthologs of Eaf3 in yeast and Mrg15 in Drosophila respectively, which recognize Set2-catalyzed H3K36me318,20. Therefore, we focused on these two proteins for further analyses. First, target deletion mutants of MGG_06716 and MGG_06717 were created in the wild-type (B157) strain of M. oryzae (Supplementary Fig. 2a, b). Although the Δmgg_06716 and Δmgg_06717 strains exhibited a similar phenotype with slightly smaller colonies (Fig. 1c), the growth defects of Δmgg_06716 and Δmgg_06717 could not be rescued by introducing their own complementary constructs respectively (Supplementary Fig. 2c). Moreover, transcriptome analysis revealed that Δmgg_06716 and Δmgg_06717 shared a significant overlap with up-regulated (Log2Fold Change > 1, P < 0.05) and down-regulated (Log2FC < -1, P < 0.05) genes (Fig. 1d, Supplementary Fig. 2d). Therefore, we speculated that MGG_06716 and MGG_06717 encode a single protein, hereby designated as EAF3 (a gene encompassing MGG_06716 and MGG_06717), supported by their tandem arrangement on the chromosome of M. oryzae.
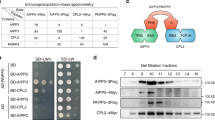
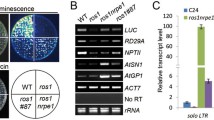
a Summary of Ash1-interactors identified by a mass spectrometry assay. Total proteins were extracted from an ASH1-Flag strain and immunoprecipitated with anti-Flag magnetic beads, followed by a mass spectrometry assay. b Gene structure of EAF3 showing tandemly arranged chromodomain (CD)-containing MGG_06716, MRG domain-containing MGG_06717, and EAF3. Integrative genome viewers (IGV) were used to show the expression pattern in the RNA-seq assays of WT, ∆eaf3, ∆mgg_06716 and ∆mgg _06717 strains. c Radial growth and statistical analysis of colony diameters in the indicated strains. Colonies of the indicated strains were grown on a complete medium (CM) for 7 d, then the top side of the colonies was imaged. Values represent means ± standard deviation (SD) from three biological replicates and statistical significance to the WT strain was assessed by a two-tailed Student’s test (**P < 0.01). n.s., no significance. d A Venn diagram showing significant overlap of gene sets among up-regulated genes in the indicated strains. A P-value indicates the significance of the overlaps between two gene sets using two-sided Fisher’s exact test. e Yeast-two-hybrid assays between Eaf3 and Ash1. Full-length and truncated fragments of EAF3 and ASH1 were cloned into pGBKT7 or pGADT7 respectively. The bait and prey plasmids were co-transformed into Y2Hgold yeast strain. Subsequently, co-transformants were grown on basal medium without tryptophan and leucine (SD-WL) and selective medium without tryptophan, leucine, histidine, and adenine (SD-WLHA). AD and BD, plasmid with empty pGADT7 or pGBKT7 used in the experiments. f Co-immunoprecipitation (Co-IP) assays were performed with the strains co-expressing EAF3-GFP and ASH1-Flag. Immunoprecipitation was performed with anti-Flag M2 magnetic beads and western blotting was conducted with anti-GFP and anti-Flag respectively. Two independent experiments were performed with similar results.
To verify this hypothesis, a 4.7-kb genomic complementary fragment (EAF3-C), including a 2.0 kb upstream region and 2.7 kb coding region, was cloned and introduced into Δmgg_06716 and Δmgg_06717 respectively (Supplementary Fig. 2a, b). When grown on complete medium, EAF3-C fully restored the growth defects of Δmgg_06716 and Δmgg_06717, providing the evidence that MGG_06716 and MGG_06717 are part of one gene (Fig. 1c). Next, a deletion mutant of EAF3 was generated in the WT strain. Compared with WT, deletion of EAF3 exhibited a growth phenotype comparable to those observed in Δmgg_06716 and Δmgg_06717 (Fig. 1c and Supplementary Fig. 2c). Remarkably, the growth defect of Δeaf3 was fully rescued by the introduction of EAF3-C (Fig. 1c). By combining the transcriptional pattern from RNA-seq of the deletion mutants and complementary tests, we inferred that the genomic fragments containing CD and MRG domains encode only one gene.
Subsequently, we determined whether full-length Eaf3 is indeed physically associated with Ash1 in M. oryzae. In a yeast two hybrid (Y2H) assay, Eaf3 specifically interacted with Ash1, but not Set2, in yeast cells (Fig. 1e). To further determine whether the individual CD or MRG domain was sufficient for the interaction, we created truncated constructs of EAF3CD and EAF3MRG, including only CD and MRG domain respectively. Notably, a truncated protein only containing the MRG domain was necessary for the interaction between Eaf3 and Ash1 (Fig. 1e). Moreover, Co-IP assays conducted with the strains co-expressing EAF3-GFP (Supplementary Fig. 3a–c) and ASH1-Flag further confirmed this interaction in vivo (Fig. 1f). Together, these results indicated that Eaf3 interacts with Ash1 in M. oryzae.
Eaf3 is required for the normal distribution of Ash1-H3K36me2 occupancy and transcriptional silencing
To further explore the association between Ash1 and Eaf3, we compared transcriptomes of Δash1 and Δeaf3. In comparison with the WT transcriptome, the transcriptomes of Δash1 and Δeaf3 exhibited similar patterns of gene expression changes, sharing significant overlap in up-regulated and down-regulated genes (Fig. 2a and Supplementary Fig. 4a)15. The absolute majority (86.4%, 1911/2212) of differentially expressed genes (DEGs) in the Δeaf3 strain were up-regulated, indicating that EAF3 primarily functions to repress transcription, similar to Ash1 in M. oryzae15.
a A Venn diagram showing significant overlaps of gene sets among up-regulated genes in the ∆eaf3 and ∆ash1 strains, and Ash1-H3K36me2-marked genes. Significance of overlaps was determined using two-sided Fisher’s exact test and P-values were shown the degree of significance. b-c Box plots and violin plots showing average transcription levels of the genes with and not Ash1-H3K36me2-marked (b) and Ash1-H3K36me3-marked (c) respectively in the indicated strains. For box-plot, the horizontal lines from top to bottom represent the maximum, first quartile, median, third quartile, and minimum of the total data respectively. For b and c, statistical significance was assessed by a two-tailed Student’s test (** P < 0.01). n.s., no significance. FPKM (Fragments Per Kilobase of exon model per Million mapped fragments). d Relative abundance of H3K36me2 and H3K36me3 in the indicated strains. The ratios of H3K36me2 and H3K36me3 to H3 in western blots were calculated by ImageJ software respectively. Values are means ± SD from five biological replicates. Statistical significance to the WT strain was assessed by a two-tailed Student’s test (** P < 0.01). The exact P-values and western blots are shown in the Source Data. e Integrative genome viewers (IGV) showing H3K36me2 and H3K36me3 occupancy in the representative genomic regions of the WT, ∆eaf3, and ∆ash1 strains. f Metaplots and heatmaps showing H3K36me2 ChIP-seq signals in the Ash1-H3K36me2-marked regions of the indicated strains. The average H3K36me2 ChIP-seq signals within 3-kb genomic regions flanking peak summits are shown. CPM (Counts per million). g Metaplots and heatmaps showing H3K36me3 ChIP-seq signals in the Ash1-H3K36me3-marked regions of the indicated strains. For f and g, metaplots and heatmaps are shown with one biological replicate of ChIP-seq assays and similar results were gained in another replicate.
In M. oryzae, Ash1 and Set2 were responsible for full activity of H3K36me2/3, and regions marked by H3K36me2/3 were divided into Ash1-catalyzed and Set2-catalyzed15. Compared with WT, genomic regions with a significant reduction of H3K36me2 occupancy in the Δset2 and Δash1 strains were designated as Set2-H3K36me2 and Ash1-H3K36me2 peaks respectively15. Subsequently, we found that 26.8% (382/1423) of Ash1-H3K36me2-marked genes was de-repressed in Δeaf3 (Fig. 2a). The average transcription of Ash1-H3K36me2-marked genes in Δeaf3 were significantly higher than those in the WT strain, while no obvious differences were found in the average transcription of Ash1-H3K36me3 and Set2-H3K36me2 genes (Fig. 2b, c, Supplementary Fig. 4b)15. Collectively, these results suggest that Eaf3 is specifically required for the transcriptional repression of Ash1-H3K36me2-marked genes.
We further examined whether Eaf3 affected the enrichment of H3K36me2 and H3K36me3 occupancy. The relative abundances of H3K36me2 and H3K36me3 were firstly detected using western blotting in the WT, Δeaf3, and Δash1 strains. Compared with the level in the WT, disruption of EAF3 caused a significant reduction in H3K36me2 levels, without affecting H3K36me3, whereas a significant reduction in both H3K36me2 and H3K36me3 was observed in Δash1 (Fig. 2d). Moreover, the abundance of H3K36me2 in Δeaf3 was still slightly higher than that in Δash1 (Fig. 2d). Subsequently, we conducted genome-wide mapping of H3K36me2 and H3K36me3 occupancy using chromatin immunoprecipitation followed by high-throughput sequencing (ChIP-seq) assays in the Δeaf3 strain, as well as in the WT and Δash1 strains (Supplementary Fig. 5a–f). Collectively, the deletion of EAF3 not only resulted in a loss of H3K36me2 in 495 genes, but also led to significantly reduced occupancy in those Ash1-H3K36me2-marked regions15, whereas only a slight reduction in occupancy was observed in H3K36me3-marked regions in Δeaf3 (Fig. 2e–g). Consistent with the findings of western blotting, the average intensity of the Ash1-H3K36me2-marked regions in Δeaf3 exhibited a similar reduction with that in Δash1, when compared with those in the WT (Fig. 2e, f). However, the relative intensity of Ash1-H3K36me3-marked regions in Δeaf3 did not clearly differ from those in the WT strain (Fig. 2e, g)15. Thus, Eaf3 is specially required for the enrichment of Ash1-H3K36me2 occupancy in M. oryzae.
Eaf3 co-localizes and regulates a set of Ash1-H3K36me2-marked genes
Next, we determined the genome-wide occupancy of Eaf3 by ChIP-seq using a GFP antibody in the EAF3-GFP strain (Supplementary Fig. 3a–c and Supplementary Fig. 5k–o). In total, 1365 peaks (Log2FC > 1, P < 0.05), corresponding with 867 genes, were identified, of which 58% peaks were located at the intergenic regions (Fig. 3a–c). A GO analysis of the Eaf3-occupied genes revealed enrichment for various terms, such as chromatin remodeling and methylation (Supplementary Fig. 6). We further investigated whether the Eaf3-occupied regions were accompanied by the altered enrichment of H3K36me2 and H3K36me3 in Δeaf3. Compared with those in the WT, intensities of H3K36me2 in Eaf3-occupied regions were significantly reduced in Δeaf3, whereas the intensity of H3K36me3 was only slightly decreased (Fig. 3d, e). We also compared the gene sets between Eaf3-occupied and Ash1-H3K36me2-marked genes, revealing that 23.1% (200/867) of the Eaf3-occupied genes were co-localized with Ash1-H3K36me2 occupancy (Fig. 3f). Together, these results further indicated the role of Eaf3 in the specific recruitment and enrichment of Ash1-H3K36me2 occupancy.
Metaplots (a) and heatmaps (b) showing the Eaf3 ChIP-seq signals of Eaf3-occupied and random regions in the EAF3-GFP strain. The average signals within 3-kb genomic regions flanking peak summits and random regions are shown. c Distribution of Eaf3-occupied and random regions. The promoter of genes, TTS, exon, intron, and intergenic region are shown. d Metagene plots showing H3K36me2 and H3K36me3 ChIP-seq signals of the Eaf3-occupied regions in the WT and ∆eaf3 strains. e Heat maps showing H3K36me2 and H3K36me3 ChIP-seq signals of the Eaf3-occupied regions in the WT and ∆eaf3 strains. f A Venn diagram showing significant overlap of the gene sets between Ash1-H3K36me2-marked and Eaf3-occupied. The significance of overlap was determined using two-sided Fisher’s exact test. g Box plots and violin plots showing average transcriptions of the genes with and not Eaf3 occupancy in the indicated strains. For box-plot, the horizontal lines from top to bottom represent the maximum, first quartile, median, third quartile, and minimum of the total data respectively. Statistical significance was assessed by a two-tailed Student’s test. n.s., no significance. h Relative abundance of Eaf3-GFP in the indicated strains, which calculated by ImageJ software and that of WT was set as 1. Values are average ± SD from five replicates. i Metagene plots showing Eaf3 ChIP-seq signals of the Eaf3-occupied regions in the WT and ∆ash1 strains. j IGV showing Eaf3-GFP, H3K36me2, and H3K36me3 occupancy from ChIP-seq and RNA-seq assays in the indicated strains. k–m ChIP-qPCR assays were performed to validate the ChIP-seq results. Relative enrichments of Eaf3 (k), H3K36me2 (l), and H3K36me3 (m) are shown. I and II indicated the amplified fragments in the ChIP-qPCR assays. Values are means ± SD from three technical replicates. For h, k–m, statistical significance was assessed by a two-tailed Student’s test (**P < 0.01). For a–e, i, j, metaplots, heatmaps, and IGV snapshots are shown with one biological replicate, and similar results were gained in another replicate.
Furthermore, we evaluated whether Eaf3 occupancy correlated with transcriptional repression by integrating RNA-seq and ChIP-seq. In the WT strain, the average transcription level of Eaf3-occupied genes was significantly lower than that of not-occupied genes, confirming the role of EAF3 as a transcriptional repressor in M. oryzae (Fig. 3g). Moreover, average transcription levels of Eaf3-occupied genes in the Δeaf3 and Δash1 strains, was both higher than that in the WT strain, indicating that both Eaf3 and Ash1 were required for transcriptional repression of Eaf3-occupied genes (Fig. 3g). Subsequently, we deleted ASH1 in the EAF3-GFP strain and checked whether Ash1 affected Eaf3 activity or occupancy. Compared with WT, the deletion of ASH1 led to substantially lower Eaf3 accumulation and decreased intensity of Eaf3 occupancy at its targeted loci (Fig. 3h-i). Integrative genome viewers (IGV) illustrated that representative genes, such as MGG_09150 (a FAD-binding protein) and MGG_04404 (a cytochrome P450), were Eaf3-occupied, and accompanied by reduced H3K36me2 occupancy and increased transcription in Δeaf3 (Fig. 3j), which were further validated by individual ChIP-qPCR assays (Fig. 3k–m). Together, these results further demonstrate the important role of EAF3 in the transcriptional silencing and the recruitment of Ash1-H3K36me2 in M. oryzae.
Eaf3 recruits EED, a core subunit of PRC2, and contributes to enrichment and transcriptional silencing of Ash1-H3K36me2-marked genes
Although Ash1-H3K36me2 co-localizes with H3K27me3 and Ash1 is required for the normal distribution of H3K27me311, the underlying molecular mechanism remains unknown. Given that Eaf3 co-localizes and regulates a set of Ash1-H3K36me2-marked genes, we asked whether this co-localization was mediated by Eaf3. Subsequently, affinity purification assays with GFP beads were conducted using the EAF3-GFP strain to identify Eaf3 interactors. As shown in Fig. 4a, Ash1 was highly enriched among the top hit of Eaf3-interactors, further confirming the association between Ash1 and Eaf3. Besides, two other proteins, Eed and Sin3, drew our further attention, as the disruption of EED and SIN3 resulted in a significant overlap of DEGs with those in the Δeaf3 strain in M. oryzae (Supplementary Fig. 7a, b).
a Summary of Eaf3-interactors identified by a mass spectrometry assay in the EAF3-GFP strain. b Eed specifically interacts with CD domain of Eaf3 in yeast cells. Indicated fragments were cloned into pGBKT7 or pGADT7 respectively. AD and BD, plasmid with empty pGADT7 or pGBKT7 used in the experiments. c Co-IP assays were performed with the strains co-expressing EED-GFP and EAF3-Flag. Three independent repeats were performed with similar results. d Relative abundance of H3K36me2 and H3K36me3 in the indicated strains, which were calculated by ImageJ software respectively. Values are means ± SD from five biological replicates. e Metagene plots and heatmaps showing H3K36me2 ChIP-seq signals of the Ash1-H3K36me2-marked regions within 3.0-kb flanking the peak summits in the indicated strains. f Box plots and violin plots showing average transcriptions of the genes with and not Ash1-H3K36me2-marked in the indicated strains. For box-plot, the horizontal lines from top to bottom represent the maximum, first quartile, median, third quartile, and minimum of the total data respectively. g Metagene plots and heatmaps showing H3K36me2 ChIP-seq signals of the Eaf3-occupied regions in the indicated strains. h Box plots and violin plots showing average transcription levels of the genes with and not Eaf3-occupied in the indicated strains. i Relative abundance of Eaf3-GFP in the indicated strains, which calculated by ImageJ software and that of WT was set as 1. Values are means ± SD from five biological replicates. j IGV showing Eaf3-GFP, H3K36me2 and H3K27me3 occupancy in the WT and ∆eaf3 strains. ChIP-qPCR assays were performed to validate the ChIP-seq results. Relative enrichments of Eaf3 (k), H3K36me2 (l), and H3K27me3 (m) are shown. I and II indicated the amplified fragments in the ChIP-qPCR assays. Values are means ± SD from three technical replicates. For d, f, h, i, k–m, statistical significance was assessed by a two-tailed Student’s test (** P < 0.01). For e, g, and j, metaplots, heatmaps, and IGV snapshots are shown with one biological replicate and similar results were gained in another replicate.
Eed, a core subunit of PRC2, is known to recognize H3K27me3 and promote the spread of this histone marker6,8. Next, the interaction between Eaf3 and Eed was further confirmed both in vitro and in vivo. Using Y2H assays, Eaf3 was found to specifically interact with Eed, but not with Kmt6 and Suz12, other core subunits of PRC2, in yeast cells (Fig. 4b). Unlike the requirement of MRG domain for interacting with Ash1, CD domain of Eaf3 is specially required for interacting with Eed (Fig. 4b). Consistently, Co-IP assays using the strains co-expressing EAF3-Flag (Supplementary Fig. 3d–f) and EED-GFP further confirmed the interaction in vivo (Fig. 4c). Together, Eaf3 was physically associated with Eed in M. oryzae.
To explore the genetic effects of the association between Eed and Eaf3, we investigated whether Eed is required for the enrichment and transcriptional silencing of Ash1-H3K36me2-marked genes. First, the relative abundances of H3K36me2 and H3K36me3 were detected in Δeed. Compared with that of WT, the abundance of H3K36me2, but not H3K36me3, was significantly lower in Δeed (Fig. 4d). Subsequently, the genome-wide occupancy of H3K36me2 was further mapped with a ChIP-seq assays. The intensity of H3K36me2 in the Ash1-catalyzed regions was significantly lower in Δeed than that of the WT strain (Fig. 4e). Moreover, 50.5% (719/1423) of Ash1-H3K36me2-marked genes were de-repressed with significantly higher transcription in Δeed than that in the WT strain (Fig. 4f and Supplementary Fig. 7c). Therefore, Eed was required for the H3K36me2 enrichment and transcriptional silencing of those Ash1-H3K36me2-marked genes in M. oryzae.
Next, we explored whether Eed is required for the enrichment and transcriptional silencing of Eaf3-occupied genes. In the Eaf3-occupied regions, the intensity of H3K36me2 was also lower in Δeed than that in the WT strain (Fig. 4g). Moreover, 28.3% (245/867) of Eaf3-occupied genes were de-repressed in Δeed (Supplementary Fig. 7c). The average transcription of Eaf3-occupied genes in Δeed was significantly higher than in the WT strain (Fig. 4h), implying that Eed was also required for Eaf3-mediated transcriptional silencing. We further deleted EED in the EAF3-GFP strain and checked whether Eed was required for Eaf3 activity or occupancy. Compared with that in the WT strain, deletion of EED also resulted in a substantially lower Eaf3 accumulation (Fig. 4i). IGV further illustrated that representative genes, such as MGG_15534 and MGG_08944, were occupied by Eaf3, and accompanied by reduced H3K36me2 and H3K27me3 in Δeaf3 (Fig. 4j and Supplementary Fig. 5g, h), which were further validated by individual ChIP-qPCR assays (Fig. 4k–m). Together, these results further demonstrate that recruiting Eed contributes to the enrichment of Eaf3 and Ash1-H3K36me2 occupancy, affecting their transcriptional expression in M. oryzae.
Eaf3 is associated with transcriptional corepressors, maintaining histone hypoacetylation at Ash1-H3K36me2 genes
In affinity purification assays, Sin3, a component of the histone deacetylation complex (HDAC) in M. oryzae, was also identified as an Eaf3-partner21,22,23,24. Next, we validated the interaction between Eaf3 and Sin3. The CD domain, but not MRG domain, of Eaf3 was found to interact with Sin3, and the interaction was further confirmed by Co-IP assays (Fig. 5a-b).
a Y2H assays between Eaf3 and Sin3. b Co-IP assays were performed with the strains co-expressing SIN3-GFP and EAF3-Flag. Two independent experiments had similar results. c Relative abundance of H3K4ac, H3K27ac, and H4K5ac in the indicated strains. Ratios of H3K4ac, H3K27ac, and H4K5ac to H3 in western blots were calculated by ImageJ software respectively and that of WT was set as 1. Values are means ± SD from five biological replicates. d, e Metagene plots and heatmaps showing H3K27ac ChIP-seq signals of the Eaf3-occupied regions in the WT and ∆eaf3 strains. f A Venn diagram showing significant overlap of gene sets between H3K27 hyperacetylated and Eaf3-occupied. P-values were calculated by two-sided Fisher’s exact test. g Box plots and violin plots showing average transcription of the genes with and not Eaf3 occupancy in the indicated strains. h Box plots and violin plots showing average transcription levels of the genes with and not Ash1-H3K36me2 in the indicated strains. For box-plot, the horizontal lines from top to bottom represent the maximum, first quartile, median, third quartile, and minimum of the total data, respectively. i IGV showing representative genes with Eaf3-GFP, H3K36me2, and H3K27ac occupancy from ChIP-seq assay and RNA-seq in the indicated strains. ChIP-qPCR assays were performed to validate the ChIP-seq results. Relative enrichments of Eaf3 (l), H3K36me2 (m), and H3K27ac (n) are shown. I and II indicated the amplified fragments in the ChIP-qPCR assays. Values are means ± SD from three technical replicates. For c, g, h, j–l, statistical significance was assessed using a two-tailed Student’s test (**P < 0.01). The exact P-values and western blots are shown in the Source Data. ns, no significance. For d, e, i, metaplots, heatmaps and IGV snapshots are shown with one biological replicate, and similar results were gained in other replicates.
Subsequently, we investigated the genetic effects of Sin3 recruitment on Eaf3 and Ash1-H3K36me2 occupancy and transcriptional regulation. First, the relative abundance of histone acetylation was detected in the WT and Δeaf3 strains. Compared with the level in the WT, the loss of EAF3 led to nearly two-fold increases in the level of examined H3K4ac, H3K27ac, and H4K5ac (Fig. 5c). Furthermore, genome-wide mapping of H3K27ac occupancy demonstrated an increase in Δeaf3 compared with WT strain, particularly in regions marked by Ash1-H3K36me2 and Eaf3 (Fig. 5d, e and Supplementary Fig. 5i, j). Although loss of EAF3 caused such amounts of direct and indirect changes in examined H3K27ac occupancy, 70.9% (615/867) of Eaf3-occupied genes were associated with hyperacetylated H3K27 in Δeaf3 (Fig. 5f). Compared with those in the WT, average transcription of Eaf3-occupied and Ash1-H3K36me2-marked genes were significantly higher in Δsin3 (Fig. 5g, h). Furthermore, IGV illustrated that representative genes, such as MGG_02592 and MGG_02191, were occupied with Eaf3, accompanied by reduced H3K36me2 and increased H3K27ac occupancy in Δeaf3 (Fig. 5i), which were further validated by individual ChIP-qPCR assays (Fig. 5j–l). Thus, recruiting Sin3 by Eaf3 maintains a histone hypoacetylated chromatin state and contributes to the transcriptional silencing of Eaf3-occupied and Ash1-H3K36me2-marked genes.
Eaf3-occupancy is required for nucleosome organization
Given that loss of EAF3 results in a reduction in histone methylation and an increase in histone acetylation and term of chromatin remodeling is highly enriched in the GO analysis of Eaf3-occupied genes, we speculated that the disruption of EAF3 is associated with local nucleosome changes. In WT, nucleosome distribution is centered on the most preferred positions, whereas in mutants, alterations such as shifts (forward or backward) or occupancy changes (increased or decreased) are expected to be detected25,26. Micrococcal nuclease (MNase) shows a much higher preference for naked DNA than for nucleosomal DNA and digestion of chromatin by MNase, followed by high-throughput sequencing (MNase-seq), is a powerful tool for investigating nucleosome organization across the genome27. Thus, MNase-seq was conducted in the WT and Δeaf3 strains with two biological replicates (Supplementary Fig. 8).
Compared with WT, 31297 changes in nucleosome shifts (including 15260 forward and 16037 backward) and 4437 changes in nucleosome occupancy (including 150 increased and 4287 decreased) were detected in Δeaf3 according to the DANPOS screening criteria of nucleosome changes (Fig. 6a). Eaf3-occupied regions from ChIP-seq were corresponding with 5008 units of nucleosomes. We integrated ChIP-seq and MNase-seq to investigate whether Eaf3 occupancy is associated with local nucleosome changes. The nucleosome set of Eaf3-occupied was compared with the nucleosomes sets carrying forward or backward shifts, as well as those with increased or decreased occupancy in Δeaf3 (Fig. 6a). Remarkably, over 8-fold (822/98, Observed/Expected) occurrence of nucleosomes was found with both decreased occupancy in Δeaf3 and occupied by Eaf3, (Fig. 6a), indicating that the loss of EAF3 associated with decreased occupancy in Eaf3-occupied nucleosomes.
a Venn diagrams showing overlaps of the indicated nucleosome sets. The sets of nucleosome changes, including changed shifts or occupancy, were calculated with that of the Δeaf3 strain versus that of WT. b Integrated analysis of RNA-seq, ChIP-seq, and MNase-seq showing the overlap of the indicated gene sets. For example, “14.7%, 773/703, P < 1.687e-04” indicates that transcription of 14.7% (773/5258) genes with forward shifted nucleosomes was up-regulated in the ∆eaf3 strain, which showed statistically significant overlaps (773/703, Observed/Expected) between two gene sets. For a and b, P-values were calculated by two-sided Fisher’s exact test. c Box plots and violin plots showing average transcriptions of the genes with down or up-regulated nucleosome occupancy in the WT and ∆eaf3 strains. d Box plots and violin plots comparing average transcriptions of genes with down-regulated nucleosome occupancy with or not Eaf3 occupancy in the WT and ∆eaf3 strains. For box-plot, the horizontal lines from top to bottom represent the maximum, first quartile, median, third quartile, and minimum of the total data, respectively. e IGV showing MNase-seq and RNA-seq of representative genes with one biological replicate in the indicated strains. Similar results were gained in other replicates. f MNase-qPCR assay was performed to verify the results of MNase-seq. Values are means ± SD from three technical replicates. For c and d, statistical significance was assessed using a two-tailed Student’s test (* P < 0.05; ** P < 0.01). The exact P-values and western blots are shown in the Source Data.
Nucleosome occupancy is a barrier to transcription and decreased occupancy within the genes is usually associated with transcriptional activation25,26. Next, we integrated RNA-seq and MNase-seq to investigate whether changes in the nucleosome positions within genes in Δeaf3 associated with changes in transcription. As a single gene usually has several nucleosomes with altered position, we re-mapped these changes on a gene-by-gene basis. Compared with that in the WT, 5258 genes with a forward nucleosome shift and 5481 genes with a backward shift were identified in Δeaf3. Additionally, 59 genes with increased nucleosome occupancy and 695 genes with decreased occupancy were found in Δeaf3 (Fig. 6b). In the Δeaf3 strain, up-regulated genes, but not down-regulated genes, were significantly enriched with changed nucleosome shifts and decreased occupancy (Fig. 6b). Consistently, 53.5% (372/695) of genes with decreased nucleosome occupancy were also occupied by Eaf3, implying more relaxed chromatin at those Eaf3-occupied loci in Δeaf3 than that in WT (Fig. 6b). Indeed, the average transcription of those genes with decreased nucleosome occupancy in Δeaf3 was significantly lower than those with increased occupancy, especially for Eaf3-occupied (Fig. 6c, d). IGV revealed that representative genes, such as MGG_17586 and MGG_16989, were up-regulated in Δeaf3, accompanied by decreased nucleosome occupancy, which was further verified by MNase-qPCR (Fig. 6e, f). These results further confirm the role of EAF3 as a transcriptional repressor that was required for nucleosome organization in M. oryzae.
Eaf3 recognizes H3K36 and H3K27 methylation, via its MRG and chromodomain domain, respectively
Previous studies have elucidated that MRG15/EAF3 recognizes Set2-catalyzed H3K36me35,10. As Eaf3 interacts with Ash1 and is required for the normal distribution of Ash1-H3K36me2 occupancy, we further investigated whether Eaf3 recognizes H3K36me2 in M. oryzae. Moreover, a chromodomain is usually found in the Pc subunit of PRC1 that acts as an H3K27me3 reader7,28, prompting us to investigate whether Eaf3 also recognizes H3K27me3. Subsequently, recombinant full-length and truncated Eaf3 fused with MBP were purified and incubated with histone peptides H3K36me2/3, H3K27me2/3, H3K4me2/3, and H3K9me2/3. Pulldown assays showed that MBP-Eaf3 has higher binding activity with H3K36me2/3 peptides, but relatively weaker binding with H3K27me2/3 and no binding with H3K4me2/3 and H3K9me2/3 (Fig. 7a). Moreover, truncated Eaf3 with chromodomain exhibited specific binding activity to H3K27me3, whereas the MRG domain was specifically responsible for binding to H3K36me2/3 (Fig. 7b). The histone-binding activity of full-length Eaf3 was further confirmed by an MST analysis (Fig. 7c). Together, these results indicate that Eaf3 acts as a reader of H3K36 and H3K27 methylation in M. oryzae.
a Pulldown assays using MBP-Eaf3 and biotinylated histone peptides carrying the indicated methylation. Two independent experiments had similar results. b Pull-down assays using truncated Eaf3 and biotinylated histone peptides carrying the indicated methylation. c MST measuring the affinity of binding between MBP-Eaf3 and peptides with H3K36, H3K27, H3K9 and H3K4 methylation. Kd, dissociation constant. NBD, no binding detected. Data points are represented as means ± SD from three independent replicates. d A Venn diagram showing significant overlap among gene sets of Ash1-H3K36me2, H3K27me3, and Eaf3. The significance of overlap was determined using two-sided Fisher’s exact test. e Relative abundance of H3K27me3 in the indicated strains. Relative abundance of H3K27me3 to H3 was calculated by ImageJ software and that of WT was set as 1. Values are means ± SD from five biological replicates. Statistical significance was assessed by a two-tailed Student’s test (**P < 0.01). The exact P-values and western blots are shown in the Source Data. f A Venn diagram showing the significant overlap between gene sets of H3K27me3 (WT/∆kmt6) and H3K27me3 (∆eaf3/∆kmt6)). P-values were calculated by two-sided Fisher’s exact test. g, h Heatmaps and metagene plots showing H3K27me3 ChIP-seq signals of the H3K27me3-marked regions in the WT and ∆eaf3 strains. i Box plots and violin plots showing average transcriptions levels of the genes with and not H3K27me3 in the indicated strains. For box-plot, the horizontal lines from top to bottom represent the maximum, first quartile, median, third quartile, and minimum of the total data respectively. Statistical significance was assessed by a two-tailed Student’s test (**P < 0.01). The exact P-values are shown in the Source Data. For g and h, metaplots and heatmaps are shown with one biological replicate of ChIP-seq or RNA-seq assays, and similar results were gained in other replicates.
Eaf3 is required for the normal distribution of H3K27me3 and its transcriptional silencing
As Eaf3 interacts with Eed and recognizes H3K27me3, we further investigated whether Eaf3 is required for the normal distribution of H3K27me3 occupancy. As shown in Figs. 7d, 21.0% (205/976) of H3K27me3-marked regions were also occupied by Eaf3, implying that Eaf3 might play a role in H3K27me3 occupancy. Subsequently, the relative abundance and intensity of H3K27me3 were detected in Δeaf3 using western blotting and ChIP-seq assays. Deletion of EAF3 accumulated less H3K27me3 level and led to a loss of H3K27me3 in 529 genes (Fig. 7e, f). The resulting H3K27me3-marked regions occupied 7.36% of the genome in the Δeaf3 strain, compared with 10.08% in the WT strain. Moreover, the intensity of H3K27me3 in the H3K27me3-marked regions in Δeaf3 were lower than those in WT (Fig. 7g, h). The average transcription of H3K27me3-marked genes in Δeaf3 was significantly higher than those in WT, but it was still lower than those in Δeed (Fig. 7i). Thus, Eaf3 is also required for the normal distribution of H3K27me3 occupancy and for maintaining transcriptional silencing in M. oryzae.
CD and MRG domains are indispensable for the full function of Eaf3
Owing to the differential requirements of the CD and MRG domains of Eaf3 for interactions with Ash1, Eed, and Sin3, we investigated whether these two domains are necessary for the biological functions of EAF3. Truncated complementary constructs with full-length EAF3 fused with GFP but lacking the genomic CD (EAF3MRG-GFP) or MRG fragment (EAF3CD-GFP), were cloned and introduced into the Δeaf3 strain respectively (Fig. 8a, b). Subsequently, mycelial growth, conidiation formation, and pathogenicity were investigated in deletion mutants and their complementary transformants. Compared with that of WT, the disruption of EAF3 significantly impaired fungal growth, conidiation formation, and pathogenicity in rice hosts (Fig. 8b–f). These defects can be fully rescued by the reintroduction of full-length EAF3-GFP, while transformants carrying either truncated EAF3MRG-GFP or EAF3CD-GFP failed to rescue those defects in Δeaf3 (Fig. 8b–f).
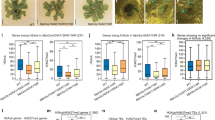
a A structure diagram showing the indicated complementary constructs. Truncated complementary constructs were cloned and introduced into Δeaf3 respectively. b, c Radial growth and statistical analysis of colony diameters of the indicated strains on the CM plate. Values are means ± SD from three biological replicates. d Statistical analysis of conidia number of the indicated strains. Values are means ± SD from three biological repeats. e Observation and statistical analysis of invasive hyphae growth in rice sheath cells at 40 hpi (hours post-inoculation). Four types of invasive hypha were quantified: no penetration (I), penetration with primary hyphae (II), penetration with secondary invasive hyphae in the first invaded cell (III), and invasive hyphae spreading into neighbouring cells (IV). Scale bar = 10 μm. Values are means ± SD from three biological replicates. f Rice seedlings infection assay. Susceptible rice seedlings (Oryza sativa L., cultivar CO39) were used for infection assays. Three independent experiments had similar results. g Growth and statistical analysis of colony diameters of the indicated strains under tested stresses. The growth inhibition rate was calculated by (1-diameter in stressed medium/diameter in CM) ×100%. Values are means ± SD from three biological repeats. h Metagene plots and heatmaps showing the Eaf3 ChIP-seq signals at the Ash1-H3K36me2 and H3K27me3-marked regions in the indicated strains. Similar results were gained in other replicates. i Relative abundance of H3K36me2, H3K36me3, H3K27me3, and H3K27ac in the indicated strains, which calculated by ImageJ software, and that of WT was set as 1. Values are means ± SD from four biological replicates. For c, d, g, and i, statistical significance to WT was assessed by a two-tailed Student’s test (**P < 0.01). j A schematic illustrating the mechanism of Eaf3-mediated transcriptional silencing. Eaf3 physically interacts with H3K36 methyltransferase Ash1, a core subunit of PRC2 Eed and histone deacetylation co-suppressor Sin3. Moreover, Eaf3 acts as a reader of the repressive H3K36me2, H3K27me3, together with H4K20me3, and Eaf3-occupied regions associated with increased nucleosome occupancy, implicating a new mechanism of facultative heterochromatin formation and Polycomb silencing.
H3K36me plays an essential role in chromatin-based regulation, such as transcriptional elongation, DNA damage repair, and DNA replication10. We further examined the role of EAF3 in DNA damage repair under genotoxic stresses including hydroxyurea (HU, an inhibitor of DNA replication) and methyl methane sulfonate (MMS, a DNA-alkylating agent). Compared with that in the WT, deletion of EAF3 led to increased sensitivity to DNA damage stresses (Fig. 8g). Moreover, the sensitivity in Δeaf3 could be fully recovered with full-length EAF3, whereas the absence of either CD or MRG domain could not rescue the defects (Fig. 8g). Collectively, these results indicated that the CD and MRG domains of Eaf3 are indispensable for the fungal development, pathogenicity, and DNA damage repair.
To further examine whether CD and MRG domains of Eaf3 are required for the recruitment of Eaf3 targets, the genome-wide occupancy of Eaf3 was determined in the EAF3MRG-GFP and EAF3CD-GFP strains respectively by ChIP-seq assays. In the Ash1-H3K36me2 and H3K27me3-marked regions, both the lack of CD and MRG domains significantly attenuated the intensity of Eaf3 enrichment (Fig. 8h). Moreover, the absence of either domain correlated with lower levels of H3K36me2 and H3K27me3, along with increased H3K27ac, as observed in Δeaf3 by western blotting (Fig. 8i). Together, the CD and MRG domains of Eaf3 are required for its occupancy and establishment of local histone modification.
To further expand our investigation, we retrieved the sequences of EAF3 orthologs from several fungal species, as well as other higher eukaryotes. Eaf3 from M. oryzae showed 33-65% sequence identity of amino acids with orthologs from fungi and relatively lower identity with higher eukaryotes, such as Drosophila and Arabidopsis (Supplementary Fig. 9). Interestingly, all Eaf3 orthologs in fungi possess a chromodomain and MRG domain, except for ortholog from Verticilium alfalfae that only had a chromodomain (Supplementary Fig. 9). Together, these results highlight the divergence of EAF3 orthologs across different fungal species.
Discussion
In this study, we identified a key transcriptional regulator EAF3, which acts as a reader of the repressive H3K36me2 and H3K27me3 by recruiting Ash1 and Eed. Eaf3 co-localizes with a set of repressive H3K36me2 and H3K27me3 loci and mediates their transcriptional silencing. Notably, Eaf3 acts as a histone reader for the repressive H3K36me2, H3K27me3, together with the reported H4K20me319, and Eaf3-occupied regions associated with increased nucleosome occupancy, which collectively contributes to transcriptional silencing in M. oryzae. Therefore, our findings reveal that Eaf3 plays a PRC1-like role in Polycomb gene silencing and facultative heterochromatin formation in fungi (Fig. 8j).
The specific recognition of histone marks by “reader” proteins constitutes an important mechanism for establishing and maintaining proper chromatin landscapes4,5. In animals, PRC2 catalyzes H3K27me3 and PRC1 participates in H3K27me3 recruitment and contributes to chromatin compaction6,8. A prevalent model for the functional transduction of H3K27me3 in mammals is that H3K27me3 is recognized and bound by the chromodomain-harboring component of PRC1, which induces gene silencing, at least partly, through mono-ubiquitination of histone H2A lysine 119 (H2AK119ub1) and chromatin compaction16,28. However, this pathway is generally absent in fungi and plants. In model plant Arabidopsis, two BAH-PHD proteins, EBS and SHL, were firstly identified as a H3K27me readers29,30. EBS and SHL, form plant-specific complexes with EMBRYONIC FLOWER 1 (EMF1), read H3K27me3, and mediate genome-wide transcriptional repression29,30. This finding suggests that BAH-EMF1c in Arabidopsis fulfills PRC1-like roles, representing the convergent evolution of PRC1 activity across plants and animals29,30.
Various orthologs, such as BAHCC1 and BAHD1 from mammals, BP1 and EPR1 from fungi, were identified as having conserved roles in Polycomb silencing31,32,33,34. BAHD1-induced heterochromatin formation is likely context-dependent and involves a multilevel mechanism, in which the direct binding of H3K27me3 by BAHD1BAH represents a critical and primary event33. BP1 interacts with the core PRC2 subunit Suz12 and directly binds to methylated H3K27 in the cereal fungal pathogen Fusarium graminearum32. Moreover, BP1 exhibits DNA binding activity through the PHD domain, contributing to the maintenance of transcriptional repression in H3K27me3-marked targets32. In our study, Eaf3 acted as a multifunctional ‘scaffold’, interacting with the H3K36 methyltransferase Ash1, core subunit of PRC2 Eed, and histone deacetylation co-suppressor Sin3. There may be other recognized proteins or recruiting mechanisms yet to be identified, and Eaf3 only co-localizes a subset of H3K36me2 and H3K27me3 regions. Loss of EAF3 resulted in a reduction in H3K36me2 and H3K27me3, and an increase in histone acetylation. This establishes a link between H3K27me3 with H3K36me2 and histone deacetylation, even H4K20me3, integral steps in stable gene repression. However, whether the increased histone acetylation in Δeaf3 is due to imbalance of histone modification or disrupted interaction with Sin3 is currently unknown. Notably, Eaf3 is required for nucleosome organization, and its disruption decreases nucleosome occupancy in the targeted genes. Together, these findings reveal a new mechanism by which Eaf3 contributes to H3K27me3 readout and facultative heterochromatin formation. Further studies are needed to determine how these chromodomain-MRG and BAH-PHD-containing proteins work together and whether they function cooperatively or competitively.
In addition to histone modifications, nucleosome positioning and occupancy may be critical for establishing facultative heterochromatin. In Drosophila, the ATP-utilizing chromatin assembly and remodeling factor (ACF) complex has been indirectly linked to the repression of Polycomb targets, and the loss of ACF1 disrupts nucleosome spacing in facultative heterochromatin35,36. In N. crassa, homologs of ISWI and ACF1, which form an ACF chromatin-remodeling complex, are required for the transcriptional repression of a subset of H3K27-methylated genes, accompanied by a downstream shift in the +1 nucleosome37,38. However, the direct connection between these proteins and PRC2 subunits has not been established. In our study, term as chromatin remodeling was highly presented in the genes with Eaf3-occupied. We further found that the loss of EAF3 led to numerous local nucleosomes shifts and occupancy changes, resulting in decreased nucleosome occupancy in Eaf3-occupied genes. Thus, Eaf3 is associated with higher nucleosome occupancy, which further maintain the stable transcription silencing of H3K27-methylated genes. Through physical interactions with Ash1, Eed, and Sin3, Eaf3 builds a direct linkage facilitating facultative heterochromatin in M. oryzae, through mechanisms involving H3K36me2 deposition, histone deacetylation, and nucleosome organization.
In M. oryzae, the roles of histone modifications such as histone (de) methylation and (de)acetylation have been extensively explored in various biological processes, including fungal growth, conidiation, and pathogenicity2,39,40. As a transcriptional regulator, the loss of EAF3 leads to abnormal fungal growth, reduced conidiation, impaired pathogenicity to rice, and increased sensitivity to DNA damage. Therefore, these findings explain the requirement of EAF3 in fungal growth, conidiation, and pathogenicity. During infection of rice, dynamic chromatin changes including histone methylation and acetylation usually affect transcription of targeted genes40. Most importantly, Eaf3 links H3K27me3 with H3K36me2 and histone deacetylation, facilitating the transcriptional silencing of genes on target chromatin. Therefore, it is important to explore the role of EAF3 during its interaction with rice.
Methods
Fungal strains and culture conditions
M. oryzae strain B157 was used as the wild-type (WT), which was a kind gift from the Indian Institute of Rice Research (Hyderabad, India), for obtaining deletion mutants41. Strains were grown on the complete medium (CM) at 25 °C for 7 d for growth measurement. For DNA damage stress experiments, strains were grown on the CM supplemented with 10 mmol/L hydroxyurea (HU) and 0.02% methyl methanesulfonate (MMS) at 25 °C for 7 d in the darkness. All experiments were conducted with three biological repeats and at least three technical replicates in each repeat.
Plasmid construction
To create the deletion mutants, about 1.0-kb of 5’ untranslated region (UTR) and 3’UTR were sequentially cloned adjacent to the resistance gene cassette within plasmids pFGL820 and pFGL821 (Addgene, 58223) according to the standard one-step gene replacement strategy41. To construct a complementary plasmid of EAF3-C, a 4.7-kb genomic fragment containing 2.0-kb of upstream sequence and 2.7-kb coding sequence was amplified and cloned into pFGL823. To construct EAF3-GFP, EAF3MRG-GFP, EAF3CD-GFP, full-length, and truncated regions of EAF3 driven by the H3 promoter and fused with the eGFP and TrpC terminator, were cloned to pKD5. After confirmation by sequencing, the resulting plasmids were introduced into the WT or their deletion mutants with a method of Agrobacterium tumefaciens-mediated transformation (ATMT). Strains and primers used in the experiments are listed in Table S1 and S2 respectively.
Conidial morphology observation assay
To assess conidiation, strains were grown on CM at 25 °C for 7 d under a 16 h light/8 h dark cycle. The conidia were collected from petri dishes and spore concentration was adjusted to about 104 mL-1. Then, about 10 µL of the spore solution was cultured on hydrophobic slides for 4 h. Subsequently, the microscopic morphology of conidia was observed and photographed.
Rice seedling and sheath infection assay
Infection assays on rice seedlings were conducted with a 1 × 105 mL-1 conidia suspension. After 7 d of infection, disease symptoms were assessed with three repeats. For the development assay of in planta invasive hyphal growth, 21-d-old rice seedlings (CO39) were used for sheath preparation. A conidial suspension was inoculated into the rice sheath and incubated in the growth chamber with a photoperiod of 16 h light/8 h dark at 25 °C. The inoculated sheath was manually trimmed and observed with an Olympus wide-field microscope at 40 hpi.
Yeast two-hybrid
Yeast two-hybrid assays were performed using the Y2Hgold yeast strain with the Matchmaker two-hybrid system (Clontech) according to the manufacturer’s instructions. The bait and prey constructs were cloned into pGBKT7 and pGADT7 vectors respectively. The co-transformants were grown on a basic medium without tryptophan and leucine (SD-WL) at 30 °C for 2 d. Subsequently, the direct protein-protein interactions were tested by growing the co-transformants on a selective medium without tryptophan, leucine, histidine, and adenine (SD-WLHA) for 4-6 d.
Western blotting
For detecting histone, 0.5-g mycelia cultured in the liquid CM for 2 d were harvested. The nuclei were isolated with extraction buffer (20 mM Tris-HCl pH 7.5, 20 mM KCl, 2 mM MgCl2, 25% glycerol, 250 mM sucrose, 0.1 mM phenylmethylsulfonyl fluoride, 5 mM beta-mercaptoethanol, and 1 × cocktail proteinase inhibitor freshly (MedChemExpress, HY-K0010) and filtered through two layers of Miracloth (Millipore, 475855-1 R). Subsequently, total histones were extracted with lysis buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 1 mM EDTA, 1% Triton X-100, and 1 × cocktail proteinase inhibitor freshly) and separated by 15% SDS-PAGE gel. For detecting Flag or GFP-tagged proteins, total proteins were directly extracted with lysis buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 1 mM EDTA, 1% Triton-100 and 1 × cocktail protein inhibitor freshly) and separated by 10% SDS-PAGE gel.
Immunoblotting was detected with anti-H3 (HuaBio, M1309-1), anti-H3K36me2 (Abcam, ab9049), anti-H3K36me3 (Abcam, ab9050), anti-H3K27me3 (Abcam, ab6002), anti-H3K27ac (Abcam, ab177178), anti-FLAG (Sigma, A8592), anti-GFP (Miltenyi, 130-091-833) and anti-GAPDH (HuaBio, R1208-3) respectively. The relative intensity of immunoblots was quantified with ImageJ software.
Co-immunoprecipitation (Co-IP)
To perform Co-IP assays, 0.5 g mycelia cultured in the liquid CM for 2 d were harvested. Total proteins were extracted with lysis buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 1 mM EDTA, 1% Triton-100, and 1 × cocktail proteinase inhibitor freshly (MedChemExpress, HY-K0010). Subsequently, the extracted proteins were precipitated with anti-FLAG M2 magnetic beads (Sigma, M8823) or GFP-Trap magnetic beads (Chromotek, gtma-20) according to the manufacturer’s instructions. The precipitated and input proteins were detected by immunoblotting with anti-FLAG (Sigma, A8592) or anti-GFP (Miltenyi, 130-091-833) respectively.
Affinity capture-mass spectrometry and data analysis
Fresh mycelia (1.0 g) from ASH1-FLAG and EAF3-GFP strains cultured in liquid CM for 2 d were harvested. Total proteins were extracted with lysis buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 1 mM EDTA, 1% Triton-100, and 1 × cocktail proteinase inhibitor freshly (MedChemExpress, HY-K0010). The supernatant was precipitated using anti-FLAG M2 magnetic beads (Sigma, M8823) or GFP-Trap magnetic beads (Chromotek, gtma-20) according to the manufacturer’s instructions. Precipitated proteins were digested with trypsin at 37 °C for 14 h and the resulting peptides were acidified with 0.1% formic acid (FA) before mass spectrometric analysis. Peptides were analyzed using the EASY-nLC 1200 HPLC system and Orbitrap Fusion mass spectrometry (Thermo Fisher Scientific) and separated on a 15 cm in-house C18 capillary column (Polymicro; 75 μm inner diameter) packed with 2.2 μm C18 resin (Michrom Bioresources) at a flow rate of 300 μL/min. Subsequently, peptides were eluted using mobile phases consisting of solvent A (0.1% FA) and solvent B (80% ACN/0.1% FA) through a linear gradient from 5% to 35%, and then 50% of solvent B at a duration of 65 min. The instrument was operated in the data-dependent mode with dynamic exclusion. The mass spectrometry survey scan was performed by an Orbitrap at a resolution of 60,000 over the 350–1500 m/z range, with the top-speed mode was selected for MS/MS measurements using HCD scans. The target values of the automatic gain controls were set to 200,000 for Orbitrap MS and 100,000 for Orbitrap MS/MS. Dynamic exclusion was performed for 45 s.
MS/MS data were searched against the M. oryzae protein database (UniProt) using Proteome Discoverer 2.2 (Thermo Fisher Scientific). The analysis was configured for tryptic peptides, allowing up to two missed cleavages per peptide. Cysteine carbamido methylation was designated as a fixed modification. Methionine oxidation was considered as a variable modification. The precursor mass tolerance was set to 10 ppm, and the MS/MS fragment tolerance was set to 0.1 Da. The false discovery rate (FDR) of peptides was set to less than 1%.
Protein expression and pull-down assay
The coding sequences of full-length, CD, or MRG domain of EAF3 were fused with MBP and cloned into pMAL-c5X respectively, and then introduced into E. coli strain BL21. Recombinant proteins of MBP-Eaf3, MBP- Eaf3CD and MBP- Eaf3MRG were expressed under 37 °C for 2 h with 1 mM IPTG and purified with Amylose Resin (NEB, E8021V) according to the manufacturer’s instructions. Peptide pull-down assays were performed as previously described29. Biotinylated histone peptides were incubated with magnetic streptavidin beads (MedChemExpress, HY-K0208) in a binding buffer (20 mM Tris-HCl pH 7.9, 0.2 mM EDTA, 1 mM DTT, 0.2 mM PMSF, 20% Glycerol, 200 mM KCl and 1 × cocktail proteinase inhibitor freshly) for 4 h at 4 °C. Subsequently, the beads were washed twice to remove unbound peptides and further incubated with the purified recombinant proteins for 3 h. After extensive washing, protein-beads were separated by SDS-PAGE, and immunoblotting was conducted with anti-MBP antibody (GeneScript, A00190).
mRNA expression and RNA sequencing (RNA-seq) analysis
Fresh mycelia (0.2 g) cultured in liquid CM for 2 d were collected. Total RNAs were extracted with NGZOL reagent (HLingene, NG303M) and reverse-transcribed to cDNAs with commercial kits (Toyobo, FSQ-301). Real-time PCR (RT-qPCR) was performed with SYBR Green qPCR Master Mix (Toyobo, QST-100) in a LightCycler480 system (Roche). The constitutively expressed Tubulin (MGG_00604) was used as endogenous control to normalize the number of cDNA templates. Primers used in the experiments were listed and described in Table S2.
RNA-seq experiments were performed on Illumina NovaSeq 6000 system and the Hiseq-PE150 strategy by the Novogene Corp (Beijing, China). RNA-seq analysis was conducted as previously described21. Briefly, genes with at least a one-fold change in expression levels (padj <0.05) were considered as differentially expressed genes (DEGs). Detailed analysis was conducted and mapped with R Studio software (R version 4.3.1). Gene Ontology (GO) analysis, aimed at identifying enriched biological processes among the DEGs, was performed using the DAVID Bioinformatics Resources 6.8 (https://david.ncifcrf.gov) with default settings. The quantification of gene expression was calculated by DESeq2 software and expression levels were normalized using FPKM (Fragments Per Kilobase of exon model per Million mapped fragments).
Chromatin immunoprecipitation (ChIP) and ChIP sequencing (ChIP-seq) analysis
ChIP assays were conducted with mycelia as previously described21. Briefly, 1.0-g mycelia were crosslinked with 1% formaldehyde and stopped with 137.5 mM glycine. After grinding and nuclei isolation, chromatin was sonicated into 200-500 bp fragments with Diagenode Bioruptor (high setting, 16 cycles with 30 s on/30 s off) and incubated with anti-GFP (Abcam, ab290), H3K36me2 (Abcam, ab9049), H3K36me3 (Abcam, ab9050), H3K27me3 (Abcam, ab6002) and H3K27ac (Abcam, ab177178) antibodies for 8–12 h. Subsequently, the recovered DNAs were purified with phenol–chloroform extraction and used as templates for following ChIP-seq and ChIP-qPCR. Two biological repeats were conducted for each experiment.
For ChIP-seq assays, the recovered DNAs were used for library construction with the Scale ssDNA-seq Lib Prep Kit (ABclonal, RK20228). High-throughput sequencing was conducted using Illumina Hiseq PE150 system by Novogene. After sequencing, clean reads were mapped to the reference genome using Bowtie2 (version 2.2.5) software with default parameters, and reads with low mapping quality or mapping to multiple positions in the genome were removed using SAMtools (version 1.13). Enriched peaks were called by HOMER (v.4.9.1) and MACS2 (version 2.2.6). Generally, broad marks such as H3K27me3 and H3K36me2/3 use HOMER to call peak, but sharp marks such as Eaf3 or H3K27ac use MACS2. HOMER using ‘getDifferentialPeaksReplicates.pl -style histone, -f 2, -q 0.05, -DESeq2’. MACS2 using ‘macs2 callpeak,-f, -g, -B, -q 0.05 ’42. The BEDTOOLS (version 2.26.0) and HOMER (v.4.9.1) were used for annotating peaks. Peaks include in gene bodies (from TSS to TTS) are considered as occupied genes. DEEPTOOLS (version 3.5.2) was used for the following43: (1) BAMCOVERAGE program was used with ‘-bs 10’ to convert the data into bigwig files, which were imported into Integrative Genomics Viewer (IGV) for visualization. (2) BAMCOMPARE was used to normalize and obtain log2ratios from two BAM files with ‘-bs 10, --operation log2, --pseudocount 0.001, -of bigwig’. (3) COMPUTEMATRIX and PLOTHEATMAP programs were used to create meta-plots and heatmaps to compare the average signal intensities at defined genomic loci. To assign peaks to proximal genes, 3.0-kb flanking the peak summit was extracted. See Supplementary Data 1 for quality control data and antibodies used.
Micrococcal nuclease sequencing (MNase-seq) and analysis
For MNase-seq, 1.0 g mycelia cultured in the liquid CM for 2 d were collected. Nuclei were extracted in extraction buffer (20 mM Tris-HCl, pH 7.5, 20 mM KCl, 2 mmol/L MgCl2, 25% glycerol, 250 mM sucrose, 0.1 mM phenylmethylsulfonyl fluoride, 5 mM beta-mercaptoethanol, and 1 × cocktail proteinase inhibitor freshly (MedChemExpress, HY-K0010) and filtered with two-layer Miracloth (Millipore, 475855-1R). Subsequently, nuclei were washed and resuspended with MNase buffer (50 mM Tris-HCl pH 7.5, 25 mM MgCl2, 1 mM CaCl2) and digested with 2 units of micrococcal nuclease (Sigma, N3755) for 10 min at 37 °C. After incubation with stop buffer (100 mM EDTA, 100 mM EGTA) and 10% SDS and proteinase K at 60 °C for 1 h, nucleosomal DNA was recovered after removing RNAs and proteins, and used as templates for following MNase-seq and MNase-qPCR.
MNase-seq was performed on the Illumina NovaSeq 6000 system and the Hiseq-PE150 strategy by Novogene Corp (Beijing, China). Data analysis was performed as previously described44. Briefly, raw reads were trimmed by FASTP (version 0.23.2). SAMtools (version 1.13) was used to extract the unique and non-redundant mapped reads for further analysis45. The filtered reads were mapped to the reference Magnaporthe oryzae genome using the Bowtie2 (version 2.2.5) program42,45. The normalized signals (Counts Per Million, CPM) were calculated with DEEPTOOLS (version 3.5.2)43. All aggregated plots and heatmaps were generated with DANPOS (version 2.2.2)44, which was used to position nucleosome occupancy and perform statistic tests for significant changes. Nucleosomes with shift distance of 50-90 bp are denoted as position shift. Nucleosome regions with an “smt_diff_FDR” <0.05 and absolute value of “smt_log2FC” > 2 are denoted as occupancy changes.
Microscale thermophoresis (MST) assay
MST assay was performed using Monolith NT.115 (NanoTemper Technologies). Purified proteins were labeled with RED-NHS and loaded into NT.115 capillaries (NanoTemper, MO-L011, MO-K022). Affinity measurements were performed with a constant concentration of 50 nM of labeled protein and increasing concentrations of histone peptide protein in a PBS-T MST buffer at 25 °C, with 15% Nano-RED at medium power. Nano Temper Analysis Software (MO.Control v2.2.1, MO.Affinity Analysis v3.0.5) was used for data analysis (NanoTemper, Germany).
Phylogenetic analysis
Protein sequences of Eaf3 orthologs from different species were retrieved from NCBI using the Basic Local Alignment Search Tool (BLAST) as a query. Sequences were first aligned with ClustalW algorithm and phylogenetic analysis was conducted with MEGA software (v.11.0.10). Subsequently, schematic protein diagrams were drawn by SMART.
Statistical analysis
For statistical analysis, a two-tailed Student’s test was calculated with Excel. P-values were calculated with the two-sided Fisher’s exact test for overlapping using online tools to determine the significance of the overlap of two gene sets (the total number of genes used in the M. oryzae genome was 14317 and total number of nucleosomes used in the M. oryzae was 220167) (http://nemates.org/MA/progs/overlap_stats.html).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The mass spectrometry proteomics data was deposited in the ProteomeX change Consortium (http://proteomecentral.proteomexchange.org) via the iProX partner repository under accession no. PXD047046 [https://proteomecentral.proteomexchange.org/cgi/GetDataset?ID = PXD047046]. The ChIP-seq, MNase-seq and RNA-seq datasets generated in this study were deposited in the Gene Expression Omnibus (GEO) under the accession number GSE246679. The ChIP-seq and RNA-seq datasets of Ash1-H3K36me2/3 and Set2-H3K36me2/3 were generated from the published data and deposited in the Gene Expression Omnibus (GEO) under the accession number GSE235415, GSE235260 and GSE235261[https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE235260]15. Source data are provided with this paper.
Code availability
Bioinformatics pipelines in this study used for RNA-seq, ChIPseq and MNase-seq analyses can be found in Supplementary Note 1.
References
Luger, K. et al. Crystal structure of the nucleosome core particle at 2.8 A resolution. Nature 389, 251–260 (1997).
Freitag, M. Histone methylation by SET domain proteins in fungi. Annu. Rev. Microbiol. 71, 413–439 (2017).
Cavalli, G. & Heard, E. Advances in epigenetics link genetics to the environment and disease. Nature 571, 489–499 (2019).
Michalak, E. M., Burr, M. L., Bannister, A. J. & Dawson, M. A. The roles of DNA, RNA and histone methylation in ageing and cancer. Nat. Rev. Mol. Cell Biol. 20, 573–589 (2019).
Zhang, T., Cooper, S. & Brockdorff, N. The interplay of histone modifications - writers that read. EMBO Rep. 16, 1467–1481 (2015).
Bieluszewski, T., Xiao, J., Yang, Y. & Wagner, D. PRC2 activity, recruitment, and silencing: a comparative perspective. Trends Plant Sci. 26, 1186–1198 (2021).
Ridenour, J. B., Möller, M. & Freitag, M. Polycomb repression without bristles: facultative heterochromatin and genome stability in fungi. Genes (Basel). 11, 638 (2020).
Wiles, E. T. & Selker, E. U. H3K27 methylation: a promiscuous repressive chromatin mark. Curr. Opin. Genet. Dev. 43, 31–37 (2017).
King, H. W., Fursova, N. A., Blackledge, N. P. & Klose, R. J. Polycomb repressive complex 1 shapes the nucleosome landscape but not accessibility at target genes. Genome Res. 28, 1494–1507 (2018).
Lam, U. T. F., Tan, B. K. Y., Poh, J. J. X. & Chen, E. S. Structural and functional specificity of H3K36 methylation. Epigenet. Chromatin 15, 17 (2022).
Bicocca, V. T., Ormsby, T., Adhvaryu, K. K., Honda, S. & Selker, E. U. ASH1-catalyzed H3K36 methylation drives gene repression and marks H3K27me2/3-competent chromatin. Elife 7, e41497 (2018).
Janevska, S. et al. Elucidation of the two H3K36me3 Histone Methyltransferases Set2 and Ash1 in Fusarium fujikuroi unravels their different chromosomal targets and a major impact of Ash1 on genome stability. Genetics 208, 153–171 (2018).
Carrozza, M. J. et al. Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123, 581–592 (2005).
Shirane, K., Miura, F., Ito, T. & Lorincz, M. C. NSD1-deposited H3K36me2 directs de novo methylation in the mouse male germline and counteracts Polycomb-associated silencing. Nat. Genet 52, 1088–1098 (2020).
Xu, M. et al. Two H3K36 methyltransferases differentially associate with transcriptional activity and enrichment of facultative heterochromatin in rice blast fungus. aBIOTECH 5, 1–16 (2024).
Wang, C., Zhu, B. & Xiong, J. Recruitment and reinforcement: maintaining epigenetic silencing. Sci. China Life Sci. 61, 515–522 (2018).
Joshi, A. A. & Struhl, K. Eaf3 chromodomain interaction with methylated H3-K36 links histone deacetylation to Pol II elongation. Mol. Cell 20, 971–978 (2005).
Huang, C. et al. Mrg15 stimulates Ash1 H3K36 methyltransferase activity and facilitates Ash1 Trithorax group protein function in Drosophila. Nat. Commun. 8, 1649 (2017).
Möller, M., Ridenour, J. B., Wright, D. F., Martin, F. A. & Freitag, M. H4K20me3 is important for Ash1-mediated H3K36me3 and transcriptional silencing in facultative heterochromatin in a fungal pathogen. PLOS Genet. 19, e1010945 (2023).
Schmähling, S. et al. Regulation and function of H3K36 di-methylation by the trithorax-group protein complex AMC. Development 145, dev163808 (2018).
Lin, C. et al. The additional PRC2 subunit and Sin3 histone deacetylase complex are required for the normal distribution of H3K27me3 occupancy and transcriptional silencing in Magnaporthe oryzae. N. Phytol. 236, 576–589 (2022).
Grzenda, A., Lomberk, G., Zhang, J. & Urrutia, R. Sin3: Master scaffold and transcriptional corepressor. Biochim. et. Biophys. Acta (BBA) - Gene Regul. Mech. 1789, 443–450 (2009).
Guan, H. et al. Diverse modes of H3K36me3-guided nucleosomal deacetylation by Rpd3S. Nature 620, 669–675 (2023).
Markert, J. W., Vos, S. M. & Farnung, L. Structure of the complete Saccharomyces cerevisiae Rpd3S-nucleosome complex. Nat. Commun. 14, 8128 (2023).
Reyes, A. A., Marcum, R. D. & He, Y. Structure and function of chromatin remodelers. J. Mol. Biol. 433, 166929 (2021).
Bieluszewski, T., Prakash, S., Roulé, T. & Wagner, D. The role and activity of SWI/SNF chromatin remodelers. Annu. Rev. Plant Biol. 74, 139–163 (2023).
Mansisidor, A. R. & Risca, V. I. Chromatin accessibility: methods, mechanisms, and biological insights. Nucleus 13, 238–278 (2022).
Connelly, K. E. et al. Engagement of DNA and H3K27me3 by the CBX8 chromodomain drives chromatin association. Nucleic Acids Res. 47, 2289–2305 (2019).
Li, Z., Fu, X., Wang, Y., Liu, R. & He, Y. Polycomb-mediated gene silencing by the BAH–EMF1 complex in plants. Nat. Genet. 50, 1254–1261 (2018).
Qian, S. et al. Dual recognition of H3K4me3 and H3K27me3 by a plant histone reader SHL. Nat. Commun. 9, 2425 (2018).
Wiles, E. T. et al. Evolutionarily ancient BAH–PHD protein mediates Polycomb silencing. Proc. Natl Acad. Sci. 117, 11614–11623 (2020).
Tang, G. et al. Fusarium BP1 is a reader of H3K27 methylation. Nucleic Acids Res. 49, 10448–10464 (2021).
Fan, H. et al. A conserved BAH module within mammalian BAHD1 connects H3K27me3 to Polycomb gene silencing. Nucleic Acids Res. 49, 4441–4455 (2021).
Fan, H. et al. BAHCC1 binds H3K27me3 via a conserved BAH module to mediate gene silencing and oncogenesis. Nat. Genet 52, 1384–1396 (2020).
Fyodorov, D. V., Blower, M. D., Karpen, G. H. & Kadonaga, J. T. Acf1 confers unique activities to ACF/CHRAC and promotes the formation rather than disruption of chromatin in vivo. Genes Dev. 18, 170–183 (2004).
Scacchetti, A. et al. CHRAC/ACF contribute to the repressive ground state of chromatin. Life Sci. Alliance 1, e201800024 (2018).
Wiles, E. T., Mumford, C. C., McNaught, K. J., Tanizawa, H. & Selker, E. U. The ACF chromatin-remodeling complex is essential for Polycomb repression. eLife 11, e77595 (2022).
Kamei, M. et al. IMITATION SWITCH is required for normal chromatin structure and gene repression in PRC2 target domains. Proc. Natl Acad. Sci. 118, e2010003118 (2021).
Shen, Y., Wei, W. & Zhou, D. Histone acetylation enzymes coordinate metabolism and gene expression. Trends Plant Sci. 20, 614–621 (2015).
Mittelsten Scheid, O., Zhang, W., Huang, J. & Cook, D. E. Histone modification dynamics at H3K27 are associated with altered transcription of in planta induced genes in Magnaporthe oryzae. PLOS Genet. 17, e1009376 (2021).
Wu, Z. et al. Polycomb repressive complex 2-Mediated H3K27 trimethylation is required for pathogenicity in Magnaporthe oryzae. Rice Sci. 29, 363–374 (2022).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Chen, K. et al. DANPOS: Dynamic analysis of nucleosome position and occupancy by sequencing. Genome Res. 23, 341–351 (2013).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Acknowledgements
This research was supported by the National Natural Science Foundation of China (32370200 and 32170192), Zhejiang Xianghu Laboratory, and National Youth Talent Support Program. We thank Ms Shujun Shao (Department of horticulture, Zhejiang University) for her assistance with MST assays and Dr Guochen Qin (Peking University Institute of Advanced Agricultural Sciences) for his assistance with mass spectrometry assays.
Author information
Authors and Affiliations
Contributions
Z.T. and M.X. conceived and designed the experiments; M.X., Q.Z., H.S., Z.W., and W.Z. performed the experiments and bioinformatics analysis; F.L. and Y.K. provided technical assistance and contributed materials; M.X., Q.Z. and Z.T. analyzed the data and wrote the paper, Z.T., and Y.K. revised the paper. All authors have discussed the results of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, M., Zhang, Q., Shi, H. et al. A repressive H3K36me2 reader mediates Polycomb silencing. Nat Commun 15, 7287 (2024). https://doi.org/10.1038/s41467-024-51789-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-51789-6
- Springer Nature Limited