Abstract
Histone H3-mutant gliomas are deadly brain tumors characterized by a dysregulated epigenome and stalled differentiation. In contrast to the extensive datasets available on tumor cells, limited information exists on their tumor microenvironment (TME), particularly the immune infiltrate. Here, we characterize the immune TME of H3.3K27M and G34R/V-mutant gliomas, and multiple H3.3K27M mouse models, using transcriptomic, proteomic and spatial single-cell approaches. Resolution of immune lineages indicates high infiltration of H3-mutant gliomas with diverse myeloid populations, high-level expression of immune checkpoint markers, and scarce lymphoid cells, findings uniformly reproduced in all H3.3K27M mouse models tested. We show these myeloid populations communicate with H3-mutant cells, mediating immunosuppression and sustaining tumor formation and maintenance. Dual inhibition of myeloid cells and immune checkpoint pathways show significant therapeutic benefits in pre-clinical syngeneic mouse models. Our findings provide a valuable characterization of the TME of oncohistone-mutant gliomas, and insight into the means for modulating the myeloid infiltrate for the benefit of patients.
Similar content being viewed by others
Introduction
Despite multimodal therapy regimens, high-grade gliomas (HGG) have poor outcomes and remain one of the leading causes of cancer-related deaths in children1. A remarkable feature of pediatric HGG (pHGG) is the presence of two hotspot, gain-of-function, somatic mutations in coding histone 3 (H3) genes, which drive tumor formation and progression2,3. Indeed, in diffuse midline gliomas (DMG), lysine to methionine amino acid substitutions in H3 variants (H3K27M) are identified in ~80% of tumors, and inhibit the methyltransferase activity of the Polycomb Repressor Complex 2 (PRC2), inducing global loss and containment of the repressive H3K27me3 mark at the complex nucleation sites4,5. In hemispheric HGGs, glycine 34 to arginine or to valine substitutions (G34R/V) in H3F3A are found in ~20% of tumors and impair the modification and/or recognition of the adjacent H3.3K36 residue in cis6,7,8.
The TME, including the tumor-infiltrating immune cells, plays a major role in modulating tumorigenesis as it directly or indirectly regulates the growth and support of tumor cells and, consequently, impacts cancer treatment and outcomes9. For several aggressive cancers such as melanoma and non-small-cell lung cancers, modulating the immune system using immune checkpoint inhibitors (ICI) has shown profound benefits10,11. The same success has not been achieved for childhood cancer, presumably based on their lower tumor mutational burden compared to adult neoplasms12. However, recent studies on subsets of “immune cold” Atypical Teratoid Rhabdoid Tumors (ATRT), which are deadly high-grade brain tumors in children, have shown paradoxical lymphoid infiltrates and sustained responses to ICI13, further indicating that a better characterization of the TME, especially the immune infiltrate, is needed in pediatric cancers. This includes H3-mutant HGGs, where limited information on the TME exists to orient optimal adjuvant therapies, particularly those characterizing and modulating the immune system. The rare available studies that have used one or few immune markers, mainly in H3K27M-mutant DMGs, indicate these brain neoplasms have poor non-inflammatory immune infiltration, mainly enriched in tumor-associated macrophages with scarce T-cells14,15,16,17. No studies exist on the TME of H3.3G34R/V pHGGs.
To explore the nature of the TME, mainly the immune cell infiltration and their interactions with H3-mutant cells, we profile H3.3K27M and G34R tumors using single-cell transcriptomic and spatial proteomic by imaging mass cytometry (IMC) technologies. We validate our findings on several mouse models of H3.3-mutant HGGs ranging from patient-derived xenografts in humanized immune-deficient mouse models to several syngeneic mouse models, including our recently described highly penetrant mouse H3.3K27M and G34R glioma models18. In this work, we uncover the universal infiltration by heterogeneous populations of myeloid cells from resident microglia and newly recruited infiltrating macrophages, limited T cell infiltration, and high expression of immune checkpoint markers on immune and tumor cells in pHGG samples. This is specific to pHGG across age and tumor location in the brain as more mixed immune populations are identified in lower grade tumors and other brain tumor entities. These findings are recapitulated in all mouse models studied, demonstrating them as reliable pre-clinical tools to explore tumor immune modulation. We further show that the crosstalk between the infiltrating immune cells and tumor cells contributes to tumorigenesis and immune evasion in H3.3K27M and can be therapeutically targeted. Our data indicate that joint inhibition of myeloid cells and immune checkpoints (IC) is therapeutically beneficial, providing important findings for the development and design of future immunotherapies in deadly pHGG.
Results
Immune profiling reveals a common abundant myeloid population in the TME of pediatric gliomas
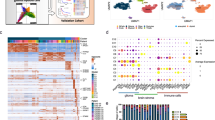
To characterize the immune landscape at single-cell resolution of H3-mutant and compare it to other pediatric gliomas, we profiled 66 pediatric glioma samples from all grades and from different subtypes using Chromium 10X technologies (27 samples from freshly dissociated single cells (sc) and 39 from single nuclei (sn), of which 17 are newly generated datasets) (Fig. 1A, Supplementary Data 1A, B). These included H3.3K27M (N = 19) and H3.3G34R/V (N = 16) mutant samples, low-grade gliomas (LGG, N = 9), and ependymomas (EP, N = 22, 20 posterior fossa group A and 2 hemispheric) (Fig. 1A). Cell filtering, normalization, integration and clustering were performed as previously described19. To annotate cells, we first identified clusters of non-immune cells using a consensus-based automated annotation pipeline based on 10 machine learning classifiers trained on a single-cell reference of the normal developing brain19. As expected, we identified various populations including OPC-like, astrocyte-like, and ependymal-like cells in each tumor subtype. Next, to annotate immune cells, we trained the classifiers on a reference of immune cells from the adult HGG microenvironment20 containing a large diversity of cell types, including a range of brain resident and infiltrating myeloid and lymphoid populations. After annotating individual cells, we validated these annotations by verifying high detection rate of PTPRC (CD45, pan-leukocyte marker) and the presence of appropriate canonical cell type-specific markers in each cell population (Supplementary Fig. 2A). Finally, we assessed batch effects by interrogating cells types expected to show low variation across samples: as expected, T cells and vascular cells clustered together independent of tumor entity (Supplementary Fig. 1E–G, see “Methods”). We identified 54431 immune cells in the single-cell dataset and only 8563 immune cells in the single-nuclei dataset split across the 4 tumor entities (Fig. 1B, Supplementary Fig. 1A–D). Indeed, as previously reported for human microglia21, data from tumor samples processed using snRNA-seq systematically underestimated the proportion and diversity of immune cells compared to samples processed using scRNA-seq in both human (Supplementary Fig. 1A–D) and mouse datasets, even when the two technologies were applied to the same tumor samples (Supplementary Fig. 1H–1). We thus focused our analysis on samples processed by scRNA-seq and mainly used snRNA-seq datasets as a validation set of trends when appropriate (Supplementary Fig. 1C, D).
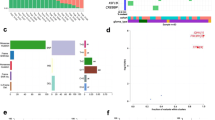
A Oncoprint summarizing single-cell tumor samples, including clinical information, immune cell proportions and genetic alterations. B UMAP of immune cells from 27 pediatric brain tumor samples (H3.3 K27M (N = 4) and G34R (N = 2) mutant gliomas, low-grade gliomas (LGG, N = 8) and ependymomas (EP, N = 13)) by scRNA-seq, with cells colored by projected tumor type. C UMAP of immune cells split by tumor entity. D Proportion of myeloid and lymphoid cells H3.3 K27M (N = 4) vs LGG (N = 8) and H3.3 K27M (N = 4) vs Ependymoma (N = 13) (* denotes adjusted p value < 0.05 and log fold change >1, permutation test; n = 10,000). E Proportions of myeloid cell types and activation states in H3.3 K27M (N = 4) vs LGG (N = 8) and H3.3 K27M (N = 4) vs Ependymoma (N = 13) (* denotes FDR adjusted p value < 0.05 and log fold change >1, permutation test; n = 10,000). F Proportions of lymphoid cell types and activation states in H3.3 K27M (N = 4) vs LGG (N = 8) and H3.3 K27M (N = 4) vs Ependymoma (N = 13) (* denotes adjusted p value < 0.05 and log fold change >1, permutation test; n = 10,000).
With the limitation of the small sample size for H3.3G34R tumors and the use of enzymatic dissociation which may dampen immune content22, all glioma samples were largely infiltrated by a different and abundant population of myeloid cells, comprised of macrophages, microglia and proliferating myeloid cells (Fig. 1E, Supplementary Fig. 1B, Supplementary Fig. 2B, Supplementary Fig. 2D). Notably, in contrast to LGG and EP samples which harbored a mixed lymphoid and myeloid infiltration (Fig. 1C–F, Supplementary Fig. 1B), H3.3K27M gliomas exhibited a lower proportion of lymphoid cells (Fig. 1D), with few T cells, regulatory T cells and NK cells (Fig. 1F, Supplementary Fig. 2D). As similar predominant myeloid infiltrates have been previously observed in adult IDH-mutant and H3-wild type HGGs23,24, the reduced lymphocyte infiltrates in these H3-mutant tumors may thus be a feature of the TME of most HGG regardless of grade, age, H3 mutations, brain location, or the glial cell-of-origin.
Since the immune infiltrate from H3.3K27M tumors showed striking differences compared to LGGs (Fig. 1D–F), we examined if gene signatures associated with different aspects of immune activation/evasion, inflammation, and cell recruitment23,24 were enriched in immune compartments in tumors (Supplementary Data 2A–C). Among the immune cell states identified, the majority of K27M-derived microglia showed an absence of activation signatures, highlighted by classical marker genes CSF1R and CX3CR1, and were depleted of disease-associated microglia signatures (Fig. 2A, B, Supplementary Fig. 2C). The H3.3K27M TME also contained rare populations of microglia and macrophages enriched for immunomodulatory genes. These included interferon-activated microglia (expressing IFIT1 and IFIT3) and phago lipid (expressing LPL) microglia and hypoxic macrophages (expressing BNIP3), which were observed to a lesser exctent in LGG or Ependymomas (Fig. 1E, Fig. 2A, B, Supplementary Fig. 2C, G). Pathway analysis using Hallmark gene sets revealed that in these tumor samples, microglia significantly downregulated TNF signaling, immune response, and complement pathways, indicating a more immunosuppressive state (Fig. 2C–E). Additionally, macrophages significantly upregulated metabolism pathways, including hypoxia and glycolysis (Supplementary Fig. 2E).
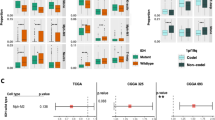
A Expression of differently expressed genes in H3.3K27M HGG microglia populations. B Enrichment (ssGSEA score) for immune gene sets in H3.3K27M HGG microglia populations (*** denotes adjusted p value < 0.0001, two-sided t test). C Differentialy expressed genes between H3.3K27M HGG and LGG microglia and macrophages. Genes with an adjusted p value < 0.05 and log2 fold change >0.5 are shown in red/blue (two-sided Wilcoxon ranked sum tets). D Expression of cytokines and immune checkpoints in H3.3K27M, Ependymoma and LGG. E Pathway analysis (MSigDB Hallmarks) for H3.3K27M microglia compared to LGG microglia. Significant pathways (adjusted p value < 0.05, GSEA Kolmogorov-Smirnov test) shown in red/blue. F Enrichment (ssGSEA score) for immune gene sets in H3.3K27M HGG and LGG microglia, macrophage and T-cell populations (*** denotes adjusted p value < 0.0001, two-sided t test).
Cells presenting high expression of ICs cannot exert proper anti-tumor effects as they are considered non-functional or exhausted subsets, and likely contribute to the immunosuppressive TME. Therefore, we investigated in our single-cell dataset the expression of genes encoding ICs and immunomodulatory chemokines and cytokines that can also modulate this TME. Microglia in H3.3K27M tumors significantly upregulated immunosuppressive genes such as CX3CR1, KLF2, TGFB1 compared to LGG and had higher levels of HAVCR2 (TIM3), LGALS9 (Galectin-9) and CD86 (Fig. 2C, D; Supplementary Fig. 2F, G). LGG tumors exhibited high expression of CLL3/4 and IL1B, which is consistent with a higher proportion of tumor-infiltrating lymphocytes and an increased tumor-fighting capacity compared to H3K27M-derived microglia (Fig. 2C, D; Supplementary Fig. 2F). Additionally, in LGG samples, both myeloid and lymphoid subtypes showed higher scores for immune signatures related to cytokines, myeloid recruitment, and antigen presentation (Fig. 2F). Interestingly, key immune-modulating factors were primarily expressed by the myeloid cells, especially in LGG and K27M HGG tumors (Supplementary Fig. 2F). These results highlight the distinct features of myeloid cells in pediatric glioma subtypes and how they contribute to promoting an immunosuppressive microenvironment, particularly in HGG.
Spatial analyses reveal significant communication between cancer and myeloid subsets
To characterize the immune architecture of H3-mutant gliomas, at the protein level and in a spatially resolved manner, we next applied Imaging Mass Cytometry (IMC) to profile the interaction between H3.3K27M and H3.3G34R tumor cells and their respective immune infiltrates, and validate our findings (Fig. 3A). We used a customized panel of 19 antibodies targeting tumor, immune, and stromal cells (Supplementary Data 3), on primary tissue samples from 12 H3-mutant patients (H3.3K27M, N = 7 and H3.3G34R, N = 5; Supplementary Data 1). IMC images were collected using a tissue microarray composed of these 12 tumors and a control human tonsil sample (Supplementary Fig. 3A). In total, 35,190 cells were annotated into 15 different cell subtypes, including lymphoid, myeloid and tumor cell populations (Fig. 3B). IMC in silico cell segmentation analysis was performed as previously described23 (Fig. 3C). Cancer cells were identified as H3.3K27M (marker for the mutation and additional loss of K27me3) or H3.3G34R positive cells (Fig. 3D, E; Supplementary Fig. 3B). Myeloid-derived monocytes were identified by CD14 and CD16 expression and classified into three categories: classic (CD14+CD16-), intermediary (CD14+CD16+) and non-classic (CD14-CD16+). Dendritic cells (DC) were annotated as CD11c+ and MHC Class II molecule (HLA-DR+) cells, endothelial cells as CD31+ and astrocytes as GFAP+ cells (Supplementary Fig. 3C). The lymphoid subsets were divided into four different subtypes, namely CD4 T cells (CD3+CD4+), CD8 T cells (CD3+CD8+), double-negative T cells (DN T cells; CD3+CD4-CD8-), and B cells (CD20+), identified by their canonical cell markers (Fig. 3B; Supplementary Fig. 3C).
A Schematic representation of the approaches used to profile the immune infiltration of human H3.3K27M and H3.3G34R mutant tumors by IMC. B Cell lineage assignment tree. Cancer cells were identified using K27M and G34R markers. Cell surface markers were used to identify and classify the immune cells. Glial fibrillary acidic protein (GFAP) positive cells were annotated as astrocytes and cells negative for the markers mentioned were annotated as undefined (not represented in the image). Cl Classic, Alt Alternative. Panels (A, B) created with BioRender.com, released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license. C Representative image of in silico cell annotation. All samples were annotated using the same approach. D–E Representative multichannel IMC images. Histone H3 mutation status was confirmed for all samples. K27M (green) and G34R (red) markers were used to identify cancer cells. K27me3 (red in D) is reduced in K27M positive cells. DNA in blue. F Cell composition and (G). Percentage of immune populations per tumor type identified by IMC, H3.3K27M (N = 7) and H3.3G34R (N = 5). H–I Representative multichannel IMC images of microglia (CD68+ P2Y12+ ), BMDM (CD68+ P2Y12-), monocytes (CD14+ CD16+ ) and T cells (CD3+ ) identified in H3.3K27M and H3.3G34R patient samples.
Cells negative for normal brain, endothelial, and cancer cell markers were annotated as immune cells. H3.3G34R tumors presented about 34% of immune cells compared to 14% in H3.3K27M tumors, accounting for an average of 24% of all the annotated cells. All major cell subtypes were observed in most patients, with variations in the cell number and absence of lymphoid subsets in some H3-mutant samples (Fig. 3F, G; Supplementary Fig. 3D), consistent with data obtained in our single-cell analysis. Strikingly, both H3.3 K27M and G34R human tumors were enriched in different subsets of myeloid cells (95% of the immune cells). These included resident classical microglia (CD68+CD163-P2Y12+), bone marrow-derived macrophages (BMDM, CD68+CD163-P2Y12-) and two different populations of monocytes (CD3-CD14+CD16- and CD3-CD14-CD16+) (Fig. 3G, I; Supplementary Fig. 3D). B cell (CD20+) and T cell populations were very rare in tumor samples, representing barely ~4% of tumor-infiltrating cells (Supplementary Fig. 3E).
To characterize proximity and communication between tumor and immune cells, we performed permutation analysis on our IMC dataset to investigate the spatial co-occurrence patterns of different cells and identify interactions or avoidance behaviors between two populations in both of the H3-mutant tumor types. Different myeloid subsets were found to interact with H3-mutant cells. Classical BMDMs showed strong interactions with both H3.3K27M- and G34R-mutant cells while monocyte subsets tended to interact mainly with H3.3G34R cells (Fig. 4A; Supplementary Fig. 4A). Except for astrocytes, H3.3K27M cancer cells were more likely to avoid most non-cancer lineages within the TME, whereas H3.3G34R cells had strong likelihood of interacting with other cancer cells, monocytes subsets and astrocytes (Fig. 4A). With the limitation of their low number in samples, no differences in the interaction of lymphoid cells with cancer cells were observed. Interestingly, in H3.3K27M-mutant tumors, endothelial cells interact significantly with cancer and myeloid cells (microglia, BMDMs and classical monocytes), and astrocytes (Fig. 4A; Supplementary Fig. 3B), indicating the important role of vascular structures in the tumor microenvironment.
A Heat map indicates IMC spatial analysis among cell phenotypes and their patterns of cell-cell interactions (red) or avoidance (blue) for H3.3K27M (top) and H3.3G34R (bottom) tumors, determined by pairwise scores. Only interaction/avoidance >50% (random chance) are shown. Associations should be read row-to-column. Rows represent the relationship of a cell type of interest. Columns represent the relationship of other surrounding cell types. Panel illustrating cell interactions created with BioRender.com, released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license. B Representative multichannel IMC images of myeloid cells (CD68+ ) interacting with K27M and G34R cells. C UMAP plots colored by expression of ligand or receptor pairs in H3.3K27M tumors (N = 4). Both immune and non-immune cells were used to predict ligand-receptor interactions. D Expression of ligand-receptor pairs in H3.3K27M immune cell populations.
Both scRNA-seq and IMC results consistently revealed that H3-mutant gliomas are enriched and interacting with a large heterogeneous myeloid population (Figs. 1D–F, 3G, 4A; Supplementary Fig. 1A, B, 4A). Given their heterogeneity and complex unknown roles, we further analyzed ligand-receptor expression in immune-infiltrating cells in the H3.3K27M scRNA-seq data using CellPhoneDB25. We identified several ligand-receptor pairs predicted to mediate communications between immune and neural-like cell types in these samples. This included the OSM/LIFR pair, which was previously shown in adult HGG to modulate behavior of myeloid and tumor cells16, while the CD47 receptor and its ligand galectin-9 (LGALS9) were also strongly expressed by the immune/myeloid cells, further indicating active immune suppression in H3.3K27M DMGs and H3.3G34R tumors (Fig. 4C, D; Supplementary Fig. 4C, D).
In summary, we help resolve the immune landscape of H3-mutant gliomas using single cell transcriptomics and spatial IMC. Our data indicate that these tumors have a diverse myeloid population, including resident macrophages and BMDMs, which predominates in the TME. We further identify their complex interactions with tumor cells, and show these myeloid cells closely interact with glioma cancer cells, and their cell-to-cell interactions promote an immune evasive environment.
H3.3K27M mouse models have a rich immunosuppressive myeloid-infiltrating tumor microenvironment
The main challenge of therapeutically modulating the immune microenvironment of these tumors is the lack of validated pre-clinical models. To this effect, we investigated the immune composition of the TME in several H3.3K27M mouse models. We explored the TME in a range of existing mouse models using cytometry by time-of-flight (CyTOF) and/or immunohistochemistry (IHC). These included two previously published H3.3K27M mouse models, where concurrent expression of H3.3K27M and loss of Tp53 led to HGG formation26,27, and a patient-derived orthotopic xenograft model (PDOX) in a CD34+ humanized NOD scid gamma (NSG) mouse, which contains a reconstituted human immune system (including T, B and NK cells) from CD34+ stem cells. In both syngeneic models, we observed high infiltration of myeloid cells and scarce infiltration of T/B cells (Fig. 5A). Likewise, high myeloid infiltration was observed in the humanized PDOX mouse model engrafted with SU-DIPG-XIII, a representative H3.3K27M DMG line4,28. Similar to what we observed in human samples, limited T cell infiltration was found in the humanized NSG mice, further confirming that the majority of H3.3K27M tumor immune-infiltrated cells are of myeloid origin (Fig. 5B).
A IHC immune markers from previous established H3K27M models (Pathania et al. and Golbourn et al). KD knockdown, oe overexpression. B IHC immune markers of tumors arose in humanized CD34+ mice (N = 2) engrafted with human SU-DIPGXIII cells (700,000 cells injected in the pons). C Schematic representation of the generation of H3.3K27M syngeneic mouse model. In utero electroporation (IUE) was used to deliver piggyBac transposable elements and gRNA/Cas9 into lower rhombic lip NPCs, using E12.5 embryos of C57Bl6/J (BL6) mice. IUE de novo tumors were used to establish cell line models. GFP+ cells were isolated and sorted. Cells were expanded in vitro and able to be engrafted and grown in immunocompetent C57Bl6/J mice. 4-hit established cell line was used for serial transplantations experiments in C57Bl6/J. GFP+ tumor regions were dissociated and single cells re-engrafted into secondary hosts. Panel created with BioRender.com, released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license. D IHC immune markers using de novo 4-hit engrafted model. E Overall survival curve of engrafted C57Bl6/J mice. 4-hit cells (150 or 350k, N = 5 each) were injected in the pons or thalamus (thal) of mice. F Cell frequencies by flow CyTOF analysis (percentage of CD45+ live cells) from 4-hit tumors injected in the pons or thalamus of mice (N = 3). G Representative IHC images of lymphocytic (CD3), and myeloid (F4/80) markers using H3.3K27M 4-hit model, engrafted in the pons and thalamus of C57Bl6/J mice.
Even though they reliably mimic human DMGs, the models tested above have long tumor latency and variable penetrance, making them less amenable for pre-clinical testing. We therefore used our recently described syngeneic mouse models18, which have several advantages: shorter tumor latency, full tumor penetrance, and expression of GFP and the luciferin analog AkaLumine-HCl sequences, enabling in vivo and in vitro tumor tracking. In this syngeneic model, in utero electroporation (IUE) of piggyBac transposon-based and CRISPR vectors targeting neural stem cells and their progeny in the lower rhombic lip of the developing hindbrain (at E12.5) led systematically to DMG formation. We used here two models: a 2-hit model, with H3.3 K27M expression and Tp53 loss, and a 4-hit model, with H3.3K27M and Pdgfra expression with Atrx/Trp53 loss (Fig. 5C, Supplementary Fig. 5A, B)18. We characterized the tumor immune infiltrate by flow CyTOF and IHC and confirmed that de novo H3.3K27M mouse tumors also contained abundant myeloid infiltration (Fig. 5D, Supplementary Fig. 5D–F), comprised mainly of P2 and P3/P4 macrophages, DCs, and myeloid-deveried suppressor cells (MDSC).
Next, we used the de novo 2-hit and 4-hit model tumors to establish cell lines that could be expanded and used for further in vivo experiments. For this, we collected tumor GFP+ regions of our syngeneic models, dissociated the cells, sorted GFP+ cells, and expanded them in vitro (Fig. 5C). Using C57BL/6 immunocompetent syngeneic mice (BL6), we validated that dissociated cells were able to generate highly penetrant tumors when injected either in the pons or the thalamus, the primary brain regions where DMGs normally occur (Fig. 5E; Supplementary Fig. 5C). Tumors were observed with similar kinetics in all mice injected, regardless of the cell dose and tumor location (Fig. 5E), indicating that the cells generated from the first tumor obtained in mice retained tumor-forming capacity. Interestingly, when we additionally engrafted these tumor cells in immunocompromised NSG mice, tumors arose at a faster rate (22 days) compared to BL6 (37 days) (Supplementary Fig. 5G), indicating that the immune pressure in the immunocompetent BL6 hosts delays initial tumor establishment, even if it is not able to fully prevent tumor formation.
In all engraftments using the 2-hit and 4-hit models, and in both tumor locations, we observed a preponderance of myeloid cells (F4/80+ cells) and limited B- and T-cell infiltration in the TME (Fig. 5F, G; Supplementary Fig. 5H, I). Immune profiling by flow CyTOF showed that regardless of the brain location in the pons or the thalamus, CCR2+ P3/P4 macrophages were the most abundant cell subtype (Fig. 5F). Given the high consistency across models and locations, we further assessed whether this immune infiltrate was also seen in the H3.3G34R mouse models, using our previously described G34R-PAD mouse model (H3.3G34R and PdgfraD842V expression with Atrx/Trp53 loss18), obtaining similar results (Supplementary Fig. 5J).
These results show the syngeneic models are reliable pre-clinical tools based on their ability to expand in vitro and to form highly penetrant tumors in immunocompetent mice, reproducing DMG and HGG pathology, including the TME. They reinforce the importance of myeloid infiltration in these HGGs and demonstrate that this is an early event in tumor development, as it is identified in the de novo tumors and in the 2-hit as well as the 4-hit models.
Myeloid-infiltrating cells are central to H3.3K27M tumorigenesis and can be therapeutically modulated
Newly injected tumor cells need to adapt and recruit specific cell types that would allow them to establish and support their growth in new environments. To understand this phenomenon, we performed in vivo serial engraftments to assess the evolution of the TME. Using the cell line established from the initial tumor formed from the 4-hit de novo model, we orthotopically engrafted BL6 mice. Once mice showed tumor burden symptoms, we dissociated tumors and directly engrafted dissociated cells into new host BL6 mice (Engraftment 2) and repeated this process in new BL6 mice (Engraftment 3) (Fig. 5C). Tumor engraftment was faster, and survival was lower with each consecutive engraftment (Fig. 6A). Once tumors formed, we collected GFP+ areas from each engraftment cohort and profiled their tumor immune populations by CyTOF, as antibodies for IMC are less available and poorly validated for mouse immune cells. We used a panel of 37 markers to identify immune cell lineages and their activation/inhibitory markers, 29 of which were used for phenotyping (Supplementary Data 4; Supplementary Fig. 6A). After automated clustering with FlowSOM, using bona fide markers for immunophenotyping 19 different immune clusters, we visualized the immune populations in two dimensions using t-distributed stochastic neighbor embedding (Fig. 6B; Supplementary Fig. 6A, B). The total number of CD45+ cells was not different between tumor engraftments (Fig. 6C). Tumors from Engraftment 1 were rich in myeloid cells, with a large number of granulocytic MDSC cells, and microglia, while their lymphocytic population was mainly composed of Tregs (Fig. 6B; Supplementary Fig. 6B). With each engraftment, we observed an increase in the proportion of myeloid cells, leading to much less diverse immune infiltration at Engraftment 3 (Fig. 6B–D), which best mirrored the TME of human H3.3K27M HGGs. In all serial transplants, myeloid cells were comprised of resident macrophages but also of newly recruited BMDMs (macrophages CCR2+ populations) (Fig. 6B), showing active recruitment of myeloid cells to the tumors. These results suggest that the more aggressive and faster growing tumors in serial engraftments are associated with an expansion of myeloid cells, including newly recruited BMDM, at the expense of other immune populations.
A Overall survival of C57Bl6/J mice from serial transplantation experiments using the 4-hit established cell line model. Log-rank test ***p = 0.001. Engraftment 1 (N = 12) cohort was generated using in vitro cultured cells. Engraftment 2 (N = 10) cohort was generated from Engraftment 1 dissociated tumors. Lastly, Engraftment 2 tumors were dissociated and injected into new hosts, generating the Engraftment 3 (N = 5) cohort. B t-SNEs from CyTOF immune infiltration profiles. t-SNEs map representing entire cell populations identified (N = 3). C Percentage of CD45+ live cells in 4-hit serial engraftment by flow CyTOF analysis (Engraftment 1 N = 3, Engraftment 2 N = 5, Engraftment 3 N = 3). Data are presented as mean values ± SEM. D Cell frequencies from Engraftment 1, 2 and 3 tumors. E Schema of drug treatment and overall survival of C57Bl6/J mice treated with anti-CSF1R and/or anti-PD1. Mice were treated with isotypes (Vehicle, N = 16), anti-CSF1R antibody (N = 13) (three doses of 300 µg, during one week), anti-PD1 antibody (4H2, N = 6) (two doses of 200 µg per week, during four weeks), and the combination of anti-CSF1R+anti-PD1 (N = 14). F Mice were also treated for four weeks with anti-CSF1R antibody alone (N = 8) (three doses of 300 µg), anti-CSF1R+anti-PD1 (N = 7) or the Vehicles (N = 7). Two mice treated with the combination developed tumors and were used to assess the tumor immune infiltrate. Panels (E, F) created with BioRender.com, released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license. G IHC representative images for CD3+ lymphocytic and F4/80+ myeloid markers and (H). Immune clusters identified by flow CyTOF analysis from tumors found at endpoint in mice treated with anti-CSF1R alone or combined with anti-PD1, during four weeks (cohort from panel F). Vehicle: N = 4, anti-CSF1R: N = 3, anti-CSF1R + PD1: N = 2. Data are presented as mean values ± SEM.
Myeloid cells play multiple roles in the immune response in brain tumors, which include impairment of T-cell recruitment/activation, tumor recognition and lysis. Furthermore, IC-expressing cells are not able to exert a proper anti-tumor effect29,30,31. PD1 inhibition has been successful in multiple tumor types, however, similar results have not been reproduced in pediatric gliomas outside of patients with constitutive mismatch repair defects who have hypermutation syndromes. Notably, immune populations including the rare lymphoid populations from our 4-hit model expressed different immune regulator markers, such as PD1 (Supplementary Fig. 6C, D), further highlighting the immune suppressive TME in H3.3K27M tumors. As a proof of concept, we evaluated if tumor growth would be impacted if the myeloid infiltration was targeted on its own, as previously described in proneural adult HGG models31,32,33, and whether joint targeting of its immunosuppressive network would prove synergistic. For this, we targeted CSF1R, a receptor kinase critical for survival and proliferation of CNS microglia, peripheral tissue macrophages and infiltrating blood myeloid cells (Fig. 6D), and PD1 which is expressed by the rare lymphocytes within the TME. Tumor-bearing mice were treated with anti-CSF1R (3x during one week) or anti-PD1 antibodies alone (4H2, 2x/week for four weeks), or in combination (Fig. 6E). No effect on tumor formation was observed when we used each inhibitor on its own. Strikingly, combined targeting of myeloid cells through CSF1R inhibition and the immune checkpoint PD1 significantly extended mouse survival and increased CD3+ infiltration (Fig. 6E, Supplementary Fig. 6E). As we still observed newly recruited BMDMs in tumors after anti-CSF1R treatment (alone or combined), we further investigated whether a more prolonged treatment (3x/week for four weeks) would prove more beneficial to improve survival rates (Fig. 6F). Here as well, CSF1R inhibitor alone failed to impact tumor formation, while anti-CSF1R combined with PD1 extended mouse survival, with 4/7 mice surviving for more than four months without visible clinical signs of disease. We performed IHC and flow CyTOF on tumors from mice treated with anti-CSF1R alone or combined with PD1 harvested at endpoint to assess the tumor immune infiltrate, compared to tumors treated with vehicle alone (Fig. 6G, H, Supplementary Fig. 6F, G). We observed increased CD3+ T cell infiltration in tumors from mice treated with the combination, and, despite CSF1R inhibition, residual F4/80+ myeloid cells, including microglial cells and macrophages, albeit at a lower level (Fig. 6G, H, Supplementary Fig. 6G). Myeloid modulation may thus have resulted in improved T cell-recruitment and mediated cytotoxic killing unhampered by the PD1 checkpoint, accounting for the increased survival we observe in mice.
These results confirm the diverse and complex roles of myeloid cells in the H3.3K27M TME, which may actively prevent recruitment and function of T cells in tumors. Notably, they suggest that the myeloid infiltrate may facilitate tumor establishment and progression and may represent a potential factor in immunotherapy resistance and/or failure that can be manipulated jointly with immune checkpoint inhibition for the benefit of patients.
Discussion
We profiled the immune populations in H3-mutant pediatric gliomas using a combination of single-cell transcriptomics and spatial single-cell proteomic on primary tumor samples. Our data show universal infiltration of tumor samples by a complex myeloid infiltrate and limited lymphoid cells in all pHGG. We further show that H3.3K27M mouse models recapitulate this immune infiltrate, which seems to sustain tumor formation and maintenance in H3.3K27M syngeneic mouse models. Last, we show that targeting these immune cells can be of therapeutic benefit.
The TME is a key regulator of cancer establishment and progression. We show that a striking feature of gliomas is the preponderance of heterogeneous myeloid cells from resident microglia to BMDM, regardless of grade, age, brain location, and H3 mutations. Indeed, our data indicate that the immune infiltrate is dominated by a diverse population of microglia/bone-marrow macrophages and a low number of infiltrating lymphocytes in H3.3K27M and G34R-mutant pediatric gliomas. Similar results were obtained in a recent study which profiled five H3.3K27M DMG using single-cell transcriptomics34 and in adult HGG including IDH-mutant gliomas, which are wild type for H3 genes23,24, indicating that this large preponderance of myeloid cells may be a feature of pHGG. Indeed, even if LGGs also have a diverse myeloid immune infiltrate, they also harbor a large number of lymphoid cells as we show here, while the TME of ATRTs, an aggressive embryonal brain tumor, can include a high immune T cell infiltrate13. This further suggests that the limited lymphoid infiltration is not due solely to the brain location or a glial or neuronal subtype, but rather is a characteristic of all pHGG. Several studies have shown that HGG across the lifespan are sustained by cancer neural stem cells, which mimic to some level the behavior of normal neural stem cells. Neural stem cells and myeloid cells emerge in the brain at the same developmental time points and physically co-exist within the brain parenchyma. In contrast, lymphoid cells emerge much later in development and rarely migrate to the brain parenchyma, remaining at the brain periphery, possibly because they could be damaging to non-renewable brain progenitors and neurons (reviewed in35). This avoidance of lymphoid cells orchestrated by neural stem cells and their myeloid entourage may be co-opted by the cancer stem cells in HGG, accounting for the low lymphoid infiltrates in pHGG. This warrants further investigations especially as more studies on the spatial proteome of the immune infiltrate in the developing brain become available.
IMC analyses allowed us to accurately categorize cellular identity and functional modalities at the single-cell resolution, with a clear separation between resident and infiltrating immune cells. It showed that both H3.3K27M and G34R pHGG are composed of resident microglia, bone-marrow derived macrophages, and monocytes, and revealed a strong interaction between tumor cancer cells and the myeloid population, while single-cell sequencing analysis indicated a heterogeneous and immunosuppressive profile for myeloid cells. Ligand-receptor analyses highlighted several candidate signaling pairs that are associated with antitumor immunity, tumor cell recognition, suppression of immune response and expansion of the immunosuppressive myeloid population16,36,37,38. Thus, our data suggest that tumor-myeloid cell interactions, including expression of several immune checkpoint markers, and resulting cytokine profiles promote an immunosuppressive milieu that acts to limit recruitment and survival of T and B cells. The spatial organization of tumors has shown to be imperative for understanding tumorigenic and immunological mechanisms ruling cancer16,23 and our study reveals predominant features of tumor evolution that involve direct cell–cell contact, including the induction of multiple immunosuppression programs, accounting for the limited recruitment of lymphoid cells within tumors.
We further show that myeloid infiltration is recapitulated by multiple H3.3K27M mouse models, including our recently described immunocompetent preclinical mouse models18. Our data indicate that all H3.3K27M models recapitulate the immune infiltrate identified in human H3.3K27M DMGs. Notably, we demonstrate that serially engrafted tumors are more aggressive and show progressive reduction in the immune infiltrate diversity which becomes largely skewed towards myeloid infiltration, recapitulating what we observe in human tumors. Similar findings have been observed in adult HGG models where cancer stem cells acquired the capacity to escape immune surveillance in serial transplants through epigenetic editing, which leads to the active recruitment of immunosuppressive myeloid cells39. Further studies are needed to identify the precise mechanisms by which histone mutant cells may be able to regulate myeloid recruitment and rely on them to grow.
Targeting myeloid cells in HGG using an anti-CSF1R regimen seems promising to improve overall survival in several adult HGG mouse models, especially in combination with other agents33,39,40. In contrast to other studies on adult HGG, myeloid depletion using anti-CSF1R antibodies produced no therapeutic benefit on its own in our H3.3K27M models, while its joint inhibition with PD1 was beneficial. Indeed, we observed increased survival and infiltration of tumors with CD3 + T cells when jointly inhibiting CSF1R and PD1. Several factors may account for the observed differences when only inhibiting CSF1R, including variations in the TME composition based on the oncogenic driver, its interactions with tumor cells, brain location, optimal scheduling, and the brain penetrance of inhibitors. Since we used the same inhibitor, and similar dosage and scheduling as described in ref. 39, brain penetrance alone does not explain the lack of survival benefit from exclusive CSF1R inhibition. An epigenetic driver mutation like H3.3K27M and the midline brain location may elicit different immune infiltrates and tumor-TME interactions, potentially explaining the observed lack of therapeutic effect. Further studies are needed to improve TME targeting and understand the lack of antitumor effects from CSF1R blockade alone in the context of H3.3K27M. It is also important to investigate the reprogramming and reshaping of myeloid cells, rather than their depletion, to better address their role in brain tumorigenesis and understand how these populations evolve in tumors during therapy.
In summary, we depict the spatial architecture and describe the contribution of the infiltrated-immune populations to H3.3K27M and G34R tumorigenesis and immune evasion, charting the immune populations present in H3-mutant pHGG and their interactions with tumor cells. Our data represent a rich resource for future studies and a guide for immunotherapeutic development for oncohistone-mutant brain tumors and possibly other pediatric brain malignancies. Since tumor infiltration can be modulated in a relevant mouse model, further investigations are warranted on the relationship of myeloid populations, immune checkpoint blockade therapies and tumorigenesis, to orient future development and design of immunotherapies for these deadly tumors.
Methods
Ethics approval and informed consent
This study was conducted in accordance with the Institutional Review Board of the respective institutions from which the samples were collected. All patients signed a written consent prior to any procedures performed, approved by the Research Ethics and Review Board of McGill University and Affiliated Hospitals Research Institutes.
All procedures on mice were performed in accordance with protocols approved by the Animal Care Committee of the RI-MUHC, McGill University Health Center Animal Care Committee and in compliance with the Canadian Council on Animal Care (CCAC) guidelines. Housing, breeding and procedures performed were in keeping with CCAC guidelines. Briefly, mice were housed under a cycle of 14-h light and 10-h dark (14:10). Room temperature was continuously monitored by a building automation system equipped with an alarm or notification system, and maintained in the range of 20–26 °C. Room relative humidity was kept at between 40 and 60%.
In utero electroporation
In utero electroporation was performed as previously described26, with minor modifications. Timed-mated, pregnant C57BL/6 J (RRID:IMSR_JAX:000664) mice were acquired from Charles River Laboratories (Kent, United Kingdom) and maintained under pathogen-free conditions, in individually ventilated cages, with food and water provided ad libitum. All procedures were approved by the UK Home Office (PPL PP2303899) and carried out in accordance with the Animals (Scientific Procedures) Act 1986. Pregnant females at E12.5 (hindbrain) or E13.5 (forebrain) were anesthetized using 2.5% isoflurane and 1.5 L O2/min, with analgesic support provided pre-operatively via subcutaneous delivery of Buprevet at 0.1 mg/kg. Uterine horns were exposed through a 1 cm incision and individual embryos were digitally manipulated into the correct orientation for intraventricular injection. Pulled borosilicate capillaries were loaded with endotoxin-free DNA and Fast Green dye (0.05%, Sigma) for visualization, and a microinjector (Eppendorf) was used to inject either the lateral or fourth ventricles with the DNA-dye mixture. 3–5 plasmids were injected simultaneously, each up to a final concentration of 2 µg/µl and 1–2 µl of total solution was injected per embryo. DNA was electroporated into ganglionic eminence progenitors using 5 mm tweezertrodes (BTX), or into lower rhombic lip progenitors using 3 mm tweezertrodes, applying 5 square pulses at 35 V and 25 V respectively, 50 ms each with 950 ms intervals. The embryos were returned into the abdominal cavity, the muscle and skin were sutured, and the animal was monitored until fully recovered from the procedure.
Cell line derivation and culture
Tumor-bearing C57BL/6 J mice were euthanized by CO2 exposure. Brains were rapidly dissected in ice-cold dissociation medium (20 mM glucose, 81.8 mM Na2SO4, 30 mM K2SO4, 5.8 mM MgCl2, 250 µM CaCl2, 1 mM HEPES, 160 µM NaOH, 0.8 mM kynurenic acid, 50 µM D-APV, 100 U/mL penicillin, 100 µg/mL streptomycin, 5 µg/ml plasmocin and 100 µg/ml primocin). Coronal sections were cut using a brain matrix and GFP+ (tumor) and GFP- regions (stroma) were microdissected under an epifluorescence stereomicroscope (Leica M205, Leica Biosystems). Microdissected tissue was then enzymatically digested into a single-cell suspension using the Papain Dissociation System according to the manufacturer’s instructions (Worthington Biochemicals, LK003150). The dissociated cell solution was separated on an OptiPrep density gradient to remove debris, followed by GFP+ cell sorting using FACSAria II (BD Biosciences). Sorted cells were plated into NeuroCult NSC proliferation media (STEMCELL Technologies, 05701) containing 20 ng/ml EGF (Miltenyi Biotec), 20 ng/ml bFGF (Miltenyi Biotech,130-093-243), 10 ng/ml PDGF-AA (Shenandoah Biotechnology), 10 ng/ml PDGF-BB (Shenandoah Biotechnology) and 2 μg/ml Heparin (STEMCELL Technologies, 07980). Cells were grown as spheroids (gliomaspheres) using ultra-low attachment plates (Corning). Once cell lines were established, they were cultured as monolayer prior experiments using Neurobasal media (Gibco-Life Technologies, 21103-049) containing 1% of GlutaPlus (200 mM,Wisent, 609-066-EL), 1% N2 supplement (100X, Wisent, 305-116-1 L) and 2% of NeuroCult SM1 (50X, StemCell Technologies, 05711), 10 ng/mL of bFGF (Miltenyi Biotech,130-093-243) and EGF (Miltenyi Biotech, 130-097-751).
Mouse cell line H3.3K27M/ATRX-TP53KD/PDGFRAoe was provided by Dr Paolo Salomoni, and the cell line H3.3K27M/TP53DN/PDGFRAoe by Dr Sameer Agnihotri, and maintained as previously described26,27. SU-DIPGXIII cell line was provided by Dr Michelle Monje, and were cultured as previously described4,28. All cell lines were authenticated using STR profiling and negative for mycoplasma.
Orthotopic injection and treatment
Engrafted mouse tumors were obtained using intracranial stereotaxic injections. For this, 150-700,000 single cells resuspended in cold HBSS were injected into the pons of 5- to 6-week-old male or female BL6 (C57BL/6 J), NSG (NOD.Cg-Prkdcscid Il2rgtm1Wjl/SzJ), or Hu-CD34 + NSG (NOD.Cg-Prkdcscid Il2rgtm1Wjl) mice, Jackson Laboratory. For tumor serial transplantation experiments, GFP+ tumors were dissected from brains once mice showed tumor burden symptoms, mechanically disaggregated and followed by enzymatic digestion using DNase and Liberase T (Roche, 05401020001). Briefly, tissue was incubated at 37 °C for 10–20 min and reaction was inhibited by adding PBS containing 75U DNase and 1% FBS. Cells were spinned and resuspended in neurobasal media. Intracranial injections were performed as described above.
For in vivo treatment, anti-CSF1R (CD115, BioXCell, BP0213) and anti-PD1 4H2 (kindly provided by Bristol-Meyer Squibb) antibodies were administered by intraperitoneal (IP) injection. Animals were randomly assigned. Mice received 300 µg of anti-CSF1R three times during one or four week(s), and/or 200 µg of anti-PD1 twice a week during four weeks, 3 days post cell injection.
Immunohistochemistry
Brains were fixed in 10% PFA overnight then transferred into 70% ethanol media. Following fixation, the samples were dehydrated and embedded in paraffin for processing. For IHC, samples were sectioned in 3–5 µm slices. IHC was performed using Ventana Discovery Ultra Instrument (Ventana Medical Systems) or Leica automated Bond-III platform in conjunction with the Polymer Refine Detection System (DS9800). Sections were incubated with primary antibodies: mouse anti-CD3 (Roche/Ventana, 790-4341), human anti-CD4 (Abcam, ab133616, 1:100), human anti-CD8 (ThermoFisher, MA5-14548, 1:100), anti-CD68 (Cell Signaling, 76437, 1:100), anti-H3K27M (RevMab Bioscience, 31-1175-00, 1:100), anti-F4/80 (Cell Signaling, 70076 T, 1:100). Slides were counterstained with hematoxylin and Bluing Reagent, and finalized with mounting medium. Sections were scanned using the Aperio AT Turbo Scanner (Leica Biosystems). Images were analyzed using ImageJ software, using multiple high-power fields. The positively stained cells were counted using automated method after adjusting particle size, circularity and intensity threshold.
Immunofluorescence
For immunofluorescence, free-floating sections were incubated in a blocking solution (10% goat or donkey serum, 3% BSA, 0.3% Triton-X in PBS) for 1 h at RT and then incubated with primary antibodies at 4 °C overnight. Sections were washed in PBS-Tween (0.05%) before addition of Hoechst 33342 and Alexa Fluor-conjugated secondary antibodies in blocking solution for 1 h at RT. Following washing, sections were mounted in ProLong Diamond Antifade mountant (Thermo Fisher Scientific) and imaged on a confocal microscope (Leica SP8, Leica Biosystems).
CyTOF
Tissue digestion
Tumor samples were processed and dissociated utilizing the Brain Tumor Dissociation Kit (Miltenyi Biotec, 130-095-942) into single cell suspensions per the manufacturer’s instructions.
Staining
CyTOF staining was performed per previously published protocol41. Briefly, cells were washed in cell-staining buffer (CSB:DPBS with 0.5% bovine serum albumin (Sigma, USA) and 0.02% sodium azide) and then incubated with Human TruStain FcX (BioLegend, USA). The cells were then stained for viability with Rh103 (Fluidigm, USA). After an additional wash with CSB, cells were stained with a surface staining antibody cocktail (Supplementary Data 4). For intracellular staining, cells were washed again with CSB and incubated with the FOXP3 fixation and permeabilization buffer solution (Invitrogen, USA). After fixation and permeabilization, cells were washed with 1X FOXP3 wash buffer (Invitrogen, USA) and stained with intracellular antibody cocktail. Cells were then washed with CSB, re-fixed with 1.6% paraformaldehyde (Sigma, USA), and kept in CSB overnight at 4 °C. On the following day, samples were labeled with 191Ir/193Ir DNA Intercalator (Fluidigm, USA) and then washed with CSB. Cells were pelleted and shipped overnight to the Longwood Medical Area CyTOF Core of the Dana-Farber Cancer Institute with refrigeration in CSB. On the day of the analysis, cells were washed with MilliQ water. Beads were added for normalization in a 1:10 dilution (Fluidigm, USA). Analysis was done on a Fluidigm mass cytometer at a rate of 250 events/s. Data were normalized using the beads and exported as FCS files.
Analysis
FCS files were uploaded to the Premium Cytobank platform (Beckman Coulter, USA) and manually gated. Beads were gated out and cells were gated as positive for DNA, single events, and negative for Rh103 for viable cells. Immune cells and microglia were further gated on CD45. The resulting events were exported as new FCS files and further analyzed using FlowJo (BD Life Sciences, USA) and the FlowSOM plug-in feature. For the initial tumor analysis, FCS files were grouped and concatenated based on sample type (i.e., de novo tumors and subsequent retransplanted tumors). The resulting concatenated files had equal numbers of events from each sample (n = 644 for de novo tumors and n = 821 for retransplanted tumors) to ensure equal representation of each sample across the experiment and used for FlowSOM clustering. Subsequently, a new concatenated file was created using all events from all files and subjected to FlowSOM clustering generated based on the balanced original concatenated file. For the treatment analysis, the CD45 positive events for those samples were gated on CD3, and the CD3 positive and CD3 negative events were concatenated and analyzed separately (n = 4785 for the CD3 negative cells and n = 4593 for CD3 positive cells). Dimensionality reduction for populations was done with t-distributed stochastic neighbor embedding (tSNE). FlowSOM clustering was performed using preset k = 20 for the tumor analysis and presets k = 15 and k = 5 for the treatment CD3 negative and CD3 positive events respectively. Clusters were manually annotated using the expression heatmap generated through FlowSOM. The concatenated dataset was then re-gated into the original samples and the percent abundance of each cluster per sample was calculated. Similar clusters were grouped together and the non-immune cell clusters were removed for the analysis. tSNE plots and heatmaps were extracted from FlowJo. Mean metal intensities (MMI) for specific markers and cluster abundance data were used to generate plots using Graphpad Prism 9. Differences in cluster abundance and MMIs between samples were calculated using unpaired t-tests with Welch’s correction in Graphpad Prism 9.
Single-cell and single nuclei transcriptomic analysis
Data processing and cell annotation of human tumor samples
Alignment and data preprocessing
Cell Ranger (10x Genomics) (v2.2.0 and 3.0.1)42 was used with default parameters for read alignment, demultiplexing, and quantification of gene counts per cell. Alignment was performed using the hg19 reference genome build, coupled with the Ensembl transcriptome (v75)43. Downstream data processing was performed as previously described19 using Seurat (v4.0.0)44. Briefly, cells were filtered using the following quality control (QC) metrics: mitochondrial content (indicative of cell damage), number of genes, and number of unique molecular identifiers (UMIs). Libraries were scaled to 10,000 UMIs per cell and log-normalized. UMI counts and mitochondrial content were regressed from normalized gene counts and the residuals Z-scored gene-wise. Dimensionality reduction was performed using principal-component analysis applied to the top 2000 most variant genes. The first 30 principal components were then used as input for projection to two dimensions using UMAP45 and for clustering using a shared nearest neighbor (SNN) modularity optimization algorithm based on the Louvain algorithm on a k-nearest neighbors graph with k = 20. Mitochondrial and ribosomal percentages per sample were calculated by dividing counts coming from ribosomal (defined as having gene symbols matching ‘RPS’, ‘RPL’, ‘MRPS’, ‘MRPL’) or mitochondrial genes (defined as having gene symbol beginning with ‘MT-’) by the total counts of the sample. Cell cycle scores for G2/M and S phases were obtained as implemented in Seurat by calculating the average expression of G2/M phase- and S phase-associated gene lists in each single cell and subtracting the average expression of control gene lists. Control gene lists were derived by binning genes in each input list into 24 bins according to expression levels and randomly selecting 100 control genes from within each expression bin.
Cell type annotation
Cells were first annotated at the individual cell level using scCoannotate, a consensus annotation pipeline using five machine learning methods (Spearman correlation, support vector machines, ACTINN46, SingleCellNet47, SciBet48) trained on a reference of the developing mouse forebrain and pons19,49. Predicted labels from each method were aggregated into broader categories based on a pre-defined cell type ontology (Supplementary Data 2), and a consensus label was determined using a majority vote based on the five methods. Cells with ties between methods, or no majority, were labeled as “Unresolved”. Clusters with a majority (>50%) of cells bearing an immune label (microglia, macrophages) were labeled as immune. The immune cells were then re-annotated at the single-cell level using scCoAnnotate, with 10 machine learning methods, Spearman correlation, support vector machines, ACTINN46, SingleCellNet47, SciBet48 scPred50, SingleR51, scHPL52, scLearn53, scClassify54, trained on a reference of GBM-associated brain immune cells from a cohort of adult glioblastoma samples20. A consensus label for each cell was computed using a modified version of the CAWPE algorithm55 as follows: first, the F1 accuracy measure was computed for each method by performing a 5-fold cross validation in the reference data set. Second, each machine learning method was trained on the reference data set and used to produce a confidence score for each reference cell type in each immune cell. Finally, the class label for each immune cell was selected as follows:
Where \({W}_{j}\) is the cross-validation accuracy (F1) measure for method j, \(\alpha\) is a hyper parameter (set to \(\alpha=4\)), and \({C}_{i}\) is the prediction confidence score for the reference cell type i in the immune cell. Predicted labels from each method were aggregated into broader categories based on a pre-defined cell type ontology (Supplementary Data 2).
Sample integration
Cells from all single-cell RNA-seq samples (N = 27) and single-nuclei RNA-seq samples (N = 39) were integrated separately for each technology using Harmony with default parameters, 30 principal components as input, and regressing out difference between samples56. The integrated space was processed as previously described for individual samples. After integration and clustering of the single-cell RNA-seq immune cells, post-clustering QC was performed, and three clusters were removed: one cluster defined by a high mitochondrial content, a cluster containing a single cell, and a cluster of potentially misclassified cells that lacked immune cell markers and expressed the non-immune genes PDGFRA, OLIG2, SOX2, NOVA1.
Testing proportions of cell types
Proportions of cell types between the tumor entities were tested using the R-packgage scProportionsTest (v1.0.0), with default parameters and the number of permutations set to 10000.
Differential gene expression analysis
Differentially expressed genes (DEGs) between tumor entities were computed per cell type using the Wilcoxon rank sum test implemented in the FindMarkers function from the Seurat package (logFC.threshold=1, min.pct=0.05). Gene set enrichment analysis for DEGs ranked by log2 fold change was performed using enrichGO function from the clusterProfiler package (v4.6.2) and the Hallmarks57 gene sets (MSigDBr v 7.5.1) with adjusted p-values < 0.05 as the significance threshold.
Immune activation signature enrichment analysis
Single-sample gene set enrichment analysis (ssGSEA)58 was performed at the individual cell level using the escape R package59. In order to characterize immune cell subsets and evaluate immune activation, we performed ssGSEA for a number of published microglial, macrophage, and T-cell activation signatures from previous single-cell RNA-seq studies60,61,62. In addition, we evaluated 10 general signatures of immune activation and myeloid recruitment from a bulk RNA-seq study on the adult glioblastoma immune component24.
Differential gene expression analysis
Cell-cell communication inference. Cell-cell communications between immune and neural cells were inferred in individual tumor samples using CellphoneDB v225. This computational tool predicts communication between groups of cells using gene expression of ligands and receptors. Cell groups used for immune cells were as described in Broad celltype annotation. Labels for non-immune cells (RGC, glial progenitors, OPC, oligodendrocytes, astrocytes, ependymal, neuronal progenitors, neurons, non-neuroectoderm, other) were retrieved from the first round of annotation with a developing mouse reference. Cells with label “No consensus” were removed. CellphoneDB was run individually in each single-cell H3.3K27M sample (n = 4) with gene names, in statistical method mode set to default parameters. In each sample, predicted ligand-receptor interactions (i.e., ligand-receptor pair between a given pair of cell groups) with p value > 0.01 and predictions between two immune cell groups or two non-immune cell groups were removed. The resulting list of ligand-receptor pairs predicted between immune and non-immune cell groups from each sample was intersected to retain only the pairs appearing in all four samples.
Data processing and cell annotation of mouse tumor samples
Alignment and data preprocessing
For mouse data, a custom genome reference was built, consisting of the mm10 reference plus the different vectors used in the mouse models. Five custom chromosomes were added to the mm10 genome: the first one containing the piggybac transferase; the second one containing Akaluc and the GFP sequences; the third one containing the human H3F3A gene with HA tag; the fourth one containing the p53 guide RNA and the last one containing the shATRX sequence. Reads were aligned to this reference using Cellranger (v2.2.0 and 3.0.1), and reads mapping to the additional chromosomes were quantified using a custom GTF file supplied to Cellranger.
Cell type annotation
Mouse cell types were annotated as described for the human tumor data. Annotation of immune subsets was performed using a reference of brain immune cells from wild-type mouse GBM models20.
Expression of immune signatures in Single Cell and Single Nuclei data
Immune signatures were generated from GBM-associated brain immune cells (45) by computing differently expressed genes from each immune cell type with the Seurat (v4.0.0) function FindAllMarkers. For each cell type a signature was generated by first filtering for genes with adjusted p value < 0.05 and then taking the top 100 genes ranked by log fold change. The average expression of each gene in signature was plotted for single cell vs single nuclei data.
Imaging mass cytometry
Samples
A total of 12 formalin-fixed, paraffin-embedded (FFPE) patient tumor samples (H3.3K27M, N = 7 and H3.3G34R/V, N = 5) was processed. A human tonsil sample was also added and used as a control for the immune markers. Cores (1.5 mm in diameter) were removed from FFPE tissue blocks and assembled into tissue microarrays. Patient samples’ information can be found in Supplementary Data 1.
Antibody panel and staining
Antibody optimization, panel and staining were prepared as previously described23. An antibody panel of 24 targets was designed to distinguish cell types and states, including immune, mesenchymal, proliferative and immune checkpoint proteins (Supplementary Data 3). Five antibodies failed and were not included in the analysis. Nine antibodies were already purchased metal-labeled (Fluidigm) and the remaining unlabeled antibodies were conjugated with metals using Maxpar X8 Multimetal Labeling Kit (Fluidigm, 201300) following the manufacturer’s protocol. All antibody titration and specificity were tested by IMC using human tumor samples, spleen and/or tonsil. Details about metals, antibodies, and concentration are described in Supplementary Data 3.
Tissue slides were stained following the IMC staining protocol23. Briefly, FFPE TMA slides underwent deparaffinization and heat-mediated antigen retrieval using the Ventana Discovery Ultra auto-stainer platform (Roche Diagnostics), according to the manufacturer’s instructions. Slides were rinsed and blocked (Dako Serum-free Protein Block solution, Agilent) and stained with a cocktail containing metal-conjugated primary antibodies at 4 °C overnight. A secondary antibody cocktail containing metal-conjugated anti-biotin was added. Slides were counterstained with Cell-ID Intercalator-Ir (Fluidigm), rinsed, and air-dried prior to IMC acquisition. IMC acquisition was performed using the Hyperion Imaging System (Fluidigm).
Image processing and analysis
Analyses were based on not transformed and raw measurements. Single-cell marker expressions are summarized by mean pixel values for each channel. For heat map visualization, expression data were normalized to the 95th percentile and z-scored cluster means were plotted. All lineage and functional markers underwent a staining quality check prior to cell segmentation. A subset of functional markers did not stain consistently with IMC and were subsequently removed from analysis (PD-L1-Biotin, FoxP3, PD-1, CD45RA, CD45RO; see Supplementary Data 2). Cell segmentation was done using machine learning-based computer vision algorithms as previously described23.
IMC cell–cell pairwise interaction analysis
To identify significant pairwise interaction and avoidance behaviors between cell types, we performed permutation tests of single-cell interactions as previously described23. Cells within a 6-pixel radius (6 μm) were considered interacting. Significant interaction or avoidance between G34R and K27M samples were determined by comparing the mean interaction/avoidance score for each pairwise comparison using a two-sample t test (MATLAB). P-values less than 0.01 were considered significant where at least one of the groups had a tendency for interaction/avoidance >50% (random chance).
Statistics and reproducibility
The description of statistical details for each experiment is indicated in the figure legends. Experiments were validated in at least three biological replicates unless stated otherwise. Significance was assigned by tests deriving a p value less than 0.05. All the data shown are represented either as Mean ± standard deviation (SD) or Mean ± standard error mean (SE), as detailed in the figure legends or Methods section.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
scRNA-seq and snRNA-seq data for pediatric human tumors have been deposited in the European Genome-phenome Archive (EGA) under accession number EGAS00001007510. Previously published data used in this study is deposited in the European Genome-phenome Archive (EGA) under accession number EGAS00001005773 (https://ega-archive.org/dacs/EGAC00001002411), EGAS00001003170 and EGAS00001004301 (https://ega-archive.org/dacs/EGAC00001001571) and Gene Expression Omnibus (GEO) under accession number GSE125969. Accession numbers for previously published samples used in this study are also provided in Supplementary Data 1. scRNA-seq and snRNA-seq data for H3.3K27M mouse models have been deposited in Gene Expression Omnibus (GEO) under accession number GSE241985. All other data are available in the article and its Supplementary files or from the corresponding author upon request. Source data are provided with this paper.
Code availability
Code for cell type annotation is available at https://github.com/fungenomics/scCoAnnotate.
References
Ostrom, Q. T., Cioffi, G., Waite, K., Kruchko, C. & Barnholtz-Sloan, J. S. CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2014–2018. Neuro Oncol. 23, iii1–iii105 (2021).
Schwartzentruber, J. et al. Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482, 226–231 (2012).
Wu, G. et al. Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat. Genet 44, 251–253, (2012).
Harutyunyan, A. S. et al. H3K27M induces defective chromatin spread of PRC2-mediated repressive H3K27me2/me3 and is essential for glioma tumorigenesis. Nat. Commun. 10, 1262 (2019).
Lewis, P. W. et al. Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science 340, 857–861 (2013).
Jain, S. U. et al. Histone H3.3 G34 mutations promote aberrant PRC2 activity and drive tumor progression. Proc. Natl. Acad. Sci. USA 117, 27354–27364 (2020).
Chen, C. C. L. et al. Histone H3.3G34-mutant interneuron progenitors co-opt PDGFRA for gliomagenesis. Cell 183, 1617–1633 e1622 (2020).
Khazaei, S. et al. Single substitution in H3.3G34 alters DNMT3A recruitment to cause progressive neurodegeneration. Cell 186, 1162–1178.e1120 (2023).
Sampson, J. H., Gunn, M. D., Fecci, P. E. & Ashley, D. M. Brain immunology and immunotherapy in brain tumours. Nat. Rev. Cancer 20, 12–25 (2020).
Tawbi, H. A. et al. Relatlimab and nivolumab versus nivolumab in untreated advanced melanoma. N. Engl. J. Med. 386, 24–34 (2022).
Hendriks, L. E. L. et al. Outcome of patients with non-small cell lung cancer and brain metastases treated with checkpoint inhibitors. J. Thorac. Oncol. 14, 1244–1254 (2019).
Hwang, E. I. et al. The current landscape of immunotherapy for pediatric brain tumors. Nat. Cancer 3, 11–24 (2022).
Leruste, A. et al. Clonally expanded T cells reveal immunogenicity of rhabdoid tumors. Cancer Cell 36, 597–612.e598 (2019).
Lin, G. L. et al. Non-inflammatory tumor microenvironment of diffuse intrinsic pontine glioma. Acta Neuropathol. Commun. 6, 51 (2018).
Lieberman, N. A. P. et al. Characterization of the immune microenvironment of diffuse intrinsic pontine glioma: Implications for development of immunotherapy. Neuro Oncol. 21, 83–94 (2019).
Liu, I. et al. The landscape of tumor cell states and spatial organization in H3-K27M mutant diffuse midline glioma across age and location. Nat. Genet 54, 1881–1894 (2022).
Ross, J. L. et al. Platelet-derived growth factor beta is a potent inflammatory driver in paediatric high-grade glioma. Brain 144, 53–69 (2021).
McNicholas, M. et al. A compendium of syngeneic, transplantable pediatric high-grade glioma models reveals subtype-specific therapeutic vulnerabilities. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-23-0004 (2023).
Jessa, S. et al. K27M in canonical and noncanonical H3 variants occurs in distinct oligodendroglial cell lineages in brain midline gliomas. Nat. Genet 54, 1865–1880 (2022).
Pombo Antunes, A. R. et al. Single-cell profiling of myeloid cells in glioblastoma across species and disease stage reveals macrophage competition and specialization. Nat. Neurosci. 24, 595–610 (2021).
Slyper, M. et al. A single-cell and single-nucleus RNA-Seq toolbox for fresh and frozen human tumors. Nat. Med 26, 792–802 (2020).
Srakočić, S. et al. Proposed practical protocol for flow cytometry analysis of microglia from the healthy adult mouse brain: Systematic review and isolation methods’ evaluation. Front Cell Neurosci. 16, 1017976 (2022).
Karimi, E. et al. Single-cell spatial immune landscapes of primary and metastatic brain tumours. Nature, https://doi.org/10.1038/s41586-022-05680-3 (2023).
Klemm, F. et al. Interrogation of the microenvironmental landscape in brain tumors reveals disease-specific alterations of immune cells. Cell 181, 1643–1660.e1617 (2020).
Efremova, M., Vento-Tormo, M., Teichmann, S. A. & Vento-Tormo, R. CellPhoneDB: Inferring cell-cell communication from combined expression of multi-subunit ligand-receptor complexes. Nat. Protoc. 15, 1484–1506 (2020).
Pathania, M. et al. H3.3 K27M cooperates with Trp53 loss and PDGFRA gain in mouse embryonic neural progenitor cells to induce invasive high-grade gliomas. Cancer Cell 32, 684–700.e689 (2017).
Golbourn, B. J. et al. Loss of MAT2A compromises methionine metabolism and represents a vulnerability in H3K27M mutant glioma by modulating the epigenome. Nat. Cancer 3, 629–648 (2022).
Krug, B. et al. Pervasive H3K27 acetylation leads to ERV expression and a therapeutic vulnerability in H3K27M gliomas. Cancer Cell 35, 782–797 e788 (2019).
Gutmann, D. H. & Kettenmann, H. Microglia/brain macrophages as central drivers of brain tumor pathobiology. Neuron 104, 442–449 (2019).
Hutter, G. et al. Microglia are effector cells of CD47-SIRPα antiphagocytic axis disruption against glioblastoma. Proc. Natl. Acad. Sci. USA 116, 997–1006 (2019).
Yan, D. et al. Inhibition of colony stimulating factor-1 receptor abrogates microenvironment-mediated therapeutic resistance in gliomas. Oncogene 36, 6049–6058 (2017).
Quail, D. F. et al. The tumor microenvironment underlies acquired resistance to CSF-1R inhibition in gliomas. Science 352, aad3018 (2016).
Pyonteck, S. M. et al. CSF-1R inhibition alters macrophage polarization and blocks glioma progression. Nat. Med 19, 1264–1272 (2013).
DeSisto, J. et al. Tumor and immune cell types interact to produce heterogeneous phenotypes of pediatric high-grade glioma. Neuro-Oncol. 26, 538–552 (2024).
Morimoto, K. & Nakajima, K. Role of the immune system in the development of the central nervous system. Front Neurosci. 13, 916 (2019).
Dardalhon, V. et al. Tim-3/galectin-9 pathway: regulation of Th1 immunity through promotion of CD11b+Ly-6G+ myeloid cells. J. Immunol. 185, 1383–1392 (2010).
Alban, T. J. et al. Glioblastoma myeloid-derived suppressor cell subsets express differential macrophage migration inhibitory factor receptor profiles that can be targeted to reduce immune suppression. Front Immunol. 11, 1191 (2020).
Hara, T. et al. Interactions between cancer cells and immune cells drive transitions to mesenchymal-like states in glioblastoma. Cancer Cell 39, 779–792.e711 (2021).
Gangoso, E. et al. Glioblastomas acquire myeloid-affiliated transcriptional programs via epigenetic immunoediting to elicit immune evasion. Cell 184, 2454–2470.e2426 (2021).
Przystal, J. M. et al. Targeting CSF1R alone or in combination with PD1 in experimental glioma. Cancers (Basel) 13, https://doi.org/10.3390/cancers13102400 (2021).
Stras, S. F. et al. Maturation of the human intestinal immune system occurs early in fetal development. Dev. Cell 51, 357–373.e355 (2019).
Zheng, G. X. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Cunningham, F. et al. Ensembl 2022. Nucleic Acids Res. 50, D988–D995 (2022).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587 e3529 (2021).
McInnes, L., Healy, J., Saul, N. & Großberger, L. UMAP: Uniform manifold approximation and projection. J. Open Source Softw. 3, 861 (2018).
Ma, F. & Pellegrini, M. ACTINN: automated identification of cell types in single cell RNA sequencing. Bioinformatics 36, 533–538 (2020).
Tan, Y. & Cahan, P. SingleCellNet: A computational tool to classify single cell RNA-Seq data across platforms and across species. Cell Syst. 9, 207–213 e202 (2019).
Li, C. et al. SciBet as a portable and fast single cell type identifier. Nat. Commun. 11, 1818 (2020).
Jessa, S. et al. Stalled developmental programs at the root of pediatric brain tumors. Nat. Genet 51, 1702–1713 (2019).
Alquicira-Hernandez, J., Sathe, A., Ji, H. P., Nguyen, Q. & Powell, J. E. scPred: accurate supervised method for cell-type classification from single-cell RNA-seq data. Genome Biol. 20, 264 (2019).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Michielsen, L., Reinders, M. J. T. & Mahfouz, A. Hierarchical progressive learning of cell identities in single-cell data. Nat. Commun. 12, 2799 (2021).
Duan, B. et al. Learning for single-cell assignment. Sci. Adv. 6, https://doi.org/10.1126/sciadv.abd0855 (2020).
Lin, Y. et al. scClassify: sample size estimation and multiscale classification of cells using single and multiple reference. Mol. Syst. Biol. 16, e9389 (2020).
Large, J., Lines, J. & Bagnall, A. A probabilistic classifier ensemble weighting scheme based on cross-validated accuracy estimates. Data Min. Knowl. Discov. 33, 1674–1709 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Liberzon, A. et al. The molecular signatures database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Barbie, D. A. et al. Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature 462, 108–112 (2009).
Borcherding, N. et al. Mapping the immune environment in clear cell renal carcinoma by single-cell genomics. Commun. Biol. 4, 122 (2021).
Keren-Shaul, H. et al. A unique microglia type associated with restricting development of Alzheimer’s disease. Cell 169, 1276–1290 e1217 (2017).
Sanin, D. E. et al. A common framework of monocyte-derived macrophage activation. Sci. Immunol. 7, eabl7482 (2022).
Szabo, P. A. et al. Single-cell transcriptomics of human T cells reveals tissue and activation signatures in health and disease. Nat. Commun. 10, 4706 (2019).
Acknowledgements
This work was supported by a Large-Scale Applied Research Project grant from Génome Quebec, Genome Canada, the Government of Canada, and Ministère de l’Économie et de l’Innovation du Québec, with the support of the Ontario Research Fund through funding provided by the Government of Ontario to N.J. and C.L.K. Additional funding support was provided by the We Love You Connie Foundation, Fondation Charles-Bruneau (to N.J), National Institutes of Health (NIH) grant P01-CA196539 (to N.J); Canadian Institutes of Health Research (CIHR) grants FDN-154307 (to N.J.) and PJT-156086 (to C.L.K.); Canadian Cancer Society (CCSRI) grant 705182 and a Fonds de Recherche du Québec–Santé (FRQS) salary award (to C.L.K.); the Fonds de recherche du Québec–Santé, Génome Québec, and the Cancer Research Society (to C.L.K.); NSERC RGPIN-2016-04911 (to C.L.K.); CFI Leaders Opportunity Fund 33902 (to C.L.K.); and Digital Research Alliance of Canada Resource Allocation Projects WST-164-AB and MJD-574-AC. Data analyses were enabled by computing and storage resources provided by Digital Research Alliance of Canada and Calcul Québec. N.J. is a member of the Penny Cole Laboratory and holds a Canada Research Chair (CRC) Tier 1 in Pediatric Oncology from CIHR. This work was supported in part by the Stand Up To Cancer grant “Immuno-modulation to Treat Poor-Prognosis Pediatric Brain Tumors”. M.P. is supported by start-up funding from the Cancer Research UK Children’s Brain Tumor Center of Excellence; Brain Research UK Project Grant 202021-28; Brain Tumor Charity Quest for Cures Collaborative Discovery Teams Award GN-000728; Great Ormond Street Hospital Children’s Charity Project Grant V4020; a donation from the family of Emily Parsons; and a Cancer Research UK Career Establishment Award RCCCEA-May22\100003. L.K. is supported by Yale University start-up funds, Yale Program for the Promotion of Interdisciplinary Science, Binational Science Foundation award number 2019075 and NIH grants R21TR002639, R21HD102565, and R01AI171980. Figure 3A-B; 4A; 5C and 6E-F, created with BioRender.com, released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license. We would like to thank Bristol-Meyer Squibb for kindly providing mouse PD1 inhibitor (4H2) used in this study. The authors are thankful for the technical support from Single Cell and Imaging Mass Cytometry Platform (SCIMAP) and Histology core facilities from the Rosalind and Morris Goodman Cancer Institute (GCI), Life Sciences Complex at McGill University, McGill Genome Center, RI-MUHC Histology Platform and the Segal Cancer Center Research Pathology Facility, Jewish General Hospital.
Author information
Authors and Affiliations
Contributions
A.F.A, A.A., E.K., D.G.T., M.R., Y.L., E.G.G.S., S.J., B.C., L.K., L.A.W., C.L.K. and N.Jabado. conceived and designed the analysis. A.F.A, A.A., E.K., D.G.T., M.R., Y.L., M.M., E.G.G.S., D.L., A.G., S.J., A.D.C., C.R., G.D., E.P. and Y.W. collected the data. A.F.A., A.A., E.K., D.G.T., M.R., Y.L., M.M., E.G.G.S., D.L., A.G., S.J., A.D.C., B.C., D.F., G.D., E.P., Y.W., B.K., M.Z., Q.W., S.H., V.L., A.G.W., R.W.R.D., J.K., S.A., D.F.Q., B.E., L.K., L.A.W., M.P., C.L.K., N.Jabado contributed data or analysis tools. A.F.A, A.A., E.K., D.G.T., M.R., Y.L., M.M., E.G.G.S., D.L., A.G., S.J., A.D.C., B.C., D.F., G.D., E.P., Y.W., L.K., L.A.W., M.P., C.L.K. performed the analysis. E.M.N. was responsible for biobank management, N.Juretic was project and fund manager. A.F.A. and N.Jabado wrote the paper with the input of all co-authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Gregor Hutter, Siddhartha Mitra and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Andrade, A.F., Annett, A., Karimi, E. et al. Immune landscape of oncohistone-mutant gliomas reveals diverse myeloid populations and tumor-promoting function. Nat Commun 15, 7769 (2024). https://doi.org/10.1038/s41467-024-52096-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-52096-w
- Springer Nature Limited