Abstract
Pseudomonas aeruginosa uses multiple protein regulators that work in tandem to control the production of a wide range of virulence factors and facilitate rapid adaptation to diverse environmental conditions. In this opportunistic pathogen, ToxR was known to positively regulate the production of the major virulence factor exotoxin A and now, through analysis of genetic changes between two sublines of P. aeruginosa PAO1 and functional complementation of swarming, we have identified a previously unknown role of ToxR in surface-associated motility in P. aeruginosa. Further analysis revealed that ToxR had an impact on swarming motility by regulating the Rhl quorum sensing system and subsequent production of rhamnolipid surfactants. Additionally, ToxR was found to tightly bind cyclic diguanylate (c-di-GMP) and negatively affect traits controlled by this second messenger including reducing biofilm formation and the expression of Psl and Pel exopolysaccharides, necessary for attachment and sessile communities matrix scaffolding, in P. aeruginosa. Moreover, a link between the post-transcriptional regulator RsmA and toxR expression via the alternative sigma factor PvdS, induced under iron-limiting conditions, is established. This study reveals the importance of ToxR in a sophisticated regulation of free-living and biofilm-associated lifestyles, appropriate for establishing acute or chronic P. aeruginosa infections.
Similar content being viewed by others
Introduction
Pseudomonas aeruginosa is an opportunistic pathogen able to adopt different free-living or biofilm-associated lifestyles, appropriate for establishing acute or chronic infections. This bacterium produces a wide range of virulence-associated factors including proteases, pyocyanin, exotoxin A, rhamnolipids and hydrogen cyanide1 in a population-dependent manner through quorum sensing (QS)-mediated mechanisms2. These include two N-acylhomoserine lactone (AHL) signal-dependent QS systems, the RhlR-RhlI and LasR-LasI systems. LasI synthesizes N-(3-oxododecanoyl)-l-homoserine lactone (3-oxo-C12-HSL) and RhlI, N-butanoyl-l-homoserine lactone (C4-HSL) which activate their cognate regulators, LasR and RhlR respectively, leading to the regulation of multiple virulence traits2.
In addition to QS, P. aeruginosa uses post-transcriptional regulators to facilitate rapid adaptation to diverse environmental conditions and regulate virulence3. One of these, the RNA binding protein RsmA, has a significant impact on ~9% of the gene transcripts from P. aeruginosa in both a negative and positive fashion4. This small RNA-binding protein belongs to the CsrA family of post-transcriptional regulators, initially described in Escherichia coli5, that interacts with ANGGA motifs situated within RNA hairpins6,7. As a result, RsmA represses the translation of genes involved in the establishment of chronic biofilm infections, including transcripts from genes coding for type VI secretion systems and exopolysaccharide (EPS) biosynthesis8,9. Additionally, RsmA positively influences phenotypes related to acute infections such as motility and type III secretion systems10,11.
Moreover, RsmA regulates lifestyles in P. aeruginosa via controlling the intracellular levels of cyclic diguanylate (c-di-GMP)12, a global modulator of complex cellular processes at different levels: transcriptional, post-transcriptional and post-translational13. C-di-GMP is synthesized by diguanylate cyclases (DGCs) and hydrolysed by phosphodiesterases (PDEs)14,15,16,17. Proteins containing a GGDEF domain (responsible for DGC activity), an EAL domain and/or an HD-GYP motif (both responsible for PDE activity) have been identified in a large number of bacterial genomes16,18,19,20. In particular, it is accepted that c-di-GMP levels control phenotypic changes related to the transition between motility and sessility, with high levels promoting biofilm formation and low levels leading to biofilm detachment and increased motility such as swarming15,21,–23.
Swarming is a characteristic mode of surface translocation used by a wide range of bacteria24. In P. aeruginosa this motility occurs on semisolid surfaces, for example low percentage agar plates, where this bacterium can form a distinctive pattern of radiating tendrils21. P. aeruginosa swarming is mediated by both flagellum and type IV pili, and requires rhamnolipids and QS activity as it is the result of a collective behaviour25,26,27. Rhamnolipids are believed to support migration initiation or seeding dispersal28 by lowering surface tension and thus allowing flagella-based propulsion of P. aeruginosa cells29. Rhamnolipid production is upregulated under iron-limited conditions and correlates with the formation of flat, unstructured biofilms30. It involves the activity of several enzymes coded by the rhl genes, including the acyltransferase activity of RhlA, the rhamnosyl transferase activities of RhlB and RhlC to form mono- and di-rhamnolipids respectively31,32,33,34. Expression of rhlA, rhlB and rhlC have all been shown to be under the control of the QS transcription factor RhlR31,32,34,35.
The major virulence factor exotoxin A (ETA, encoded by toxA), originally described by Liu et al.36,37, is a highly specific ADP-ribosyltransferase released by P. aeruginosa38. In the context of infection, ETA exerts its enzymatic activity within the target host cell by specific ADP-ribosylation of the highly conserved diphthamide residue in the eukaryotic elongation factor-239. As a result, protein biosynthesis is hindered leading to cell cycle arrest and apoptosis40. The regulation of toxA expression is complex and several studies have established a relation between ETA production and iron metabolism41,42,43. A regulatory element influencing ETA production was first discovered by the research group of Galloway44 and later designated toxR, for toxin regulation45. ToxR was found to positively regulate ETA production since overexpressing toxR in trans increased ETA yields 10-fold44.
In this study we sought to identify genetic elements responsible for differences in the swarming motility of two sublines of P. aeruginosa PAO1: PAO1-Nottingham (PAO1-N) and PAO1-Lausanne (PAO1-L). Genome sequence data revealed fundamental differences between these two sublines, including a 59-kb deletion containing toxR and a mutation in the c-di-GMP-degrading PDE BifA in PAO1-N. Among the genes from PAO1-L able to restore swarming deficiency to a PAO1-N ΔrsmA mutant, toxR was found to impact on swarming through regulation of the rhl QS system and subsequent rhamnolipid production. Interestingly, ToxR was found to bind c-di-GMP tightly and to negatively affect EPS production and biofilm formation in P. aeruginosa. Moreover, a link between RsmA regulation and toxR expression via the alternative sigma factor PvdS is established. Here we propose a model for the regulation of surface-associated behaviour via RsmA, PvdS and ToxR and demonstrate the previously unknown involvement of ToxR in the c-di-GMP regulated phenotypes of swarming and biofilm formation.
Results
Genetic elements from P. aeruginosa PAO1-Lausanne subline restore swarming deficiency in PAO1-Nottingham ΔrsmA
While investigating surface-associated motility in rsmA mutants of P. aeruginosa PAO1, clear differences in swarming motility between sublines originating from different laboratories were observed. Consistent with previous studies, swarming was abolished entirely in a ΔrsmA mutant made in the PAO1-N subline from Nottingham46, whereas the same deletion made in the PAO1-L subline from Lausanne had only a modest effect on motility (Fig. 1). Moreover, swarming motility was reduced in the wild type (WT) background of PAO1-N with respect to PAO1-L WT (Fig. 1), suggesting the existence of distinct genetic element(s) that contribute to this differential behaviour.
To discover component(s) within the PAO1-L genome able to restore swarming in the PAO1-N ΔrsmA mutant other than rsmA itself, random Sau3AI chromosomal DNA fragments from PAO1-L (2–4 kb) were cloned into pME6000, transformed into the ΔrsmA mutant of PAO1-N and the resulting clones (~4,000, 92% genomic coverage) screened for restoration of swarming motility. Thirteen clones partially complemented swarming motility in the PAO1-N ΔrsmA mutant (Supplementary Fig. 1), suggesting that their chromosomal DNA inserts incorporated genes absent or downregulated in PAO1-N. To isolate the genetic elements able to complement swarming when overexpressed, ORFs in the selected inserts were individually sub-cloned and seven genes were identified. Amongst these, several genes encoding hypothetical proteins (PA0285, PA2072, PA2567) were found. In addition, PA3825 and proE coding for c-di-GMP-specific phosphodiesterases47,48 were isolated. Other genes which restored swarming motility in the PAO1-N ΔrsmA mutant were rsmN encoding the small RNA-binding protein RsmN3, which was present in three of the thirteen clones indicating a good level of coverage of the PAO1-L genome, and unexpectedly toxR, a gene coding for the regulator of ETA production ToxR. Interestingly, six of the genes found to restore swarming in PAO1-N ΔrsmA mutant showed conserved domains from the family of proteins involved in c-di-GMP metabolism and hence would be expected to have an impact on swarming through the modulation of c-di-GMP levels (Table 1 and Supplementary Fig. 1).
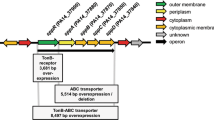
To determine which of the gene(s) identified was involved in the altered swarming motility of PAO1-N with respect to PAO1-L, the extent of the genetic differences between sublines was investigated using whole-genome sequencing, and the assembled chromosomes of both sublines were compared with the reference genome of PAO1-UW49. Candidate single nucleotide polymorphisms (SNPs) as well as small insertions/deletions (INDELs) between sublines were then selected (Supplementary Table 1). Additionally, optical restriction mapping was performed to detect large scale chromosomal rearrangements, deletions and insertions50,51. Although none of the previously identified genes was affected by the SNPs identified when comparing genomes from both sublines, sequencing analysis revealed a large deletion (~59 kb) in the PAO1-N genome that included the toxR gene (Fig. 2 and Supplementary Table 2). These results strongly suggested that ToxR might play a role in the control of swarming motility. Optical restriction mapping revealed that neither PAO1-N nor PAO1-L sublines have the large 2-Mb rrnA/rrnB chromosomal inversion reported for PAO1-UW49 (Supplementary Fig. 2), and sequencing results indicated that both carry the 12-kb RGP42 island initially found in PAO1-DSM52, excluding these as possible differences between the sublines.
A triple mutation of rsmA, toxR and bifA abolishes swarming in PAO1-L
To investigate the role of ToxR in swarming, the impact of toxR deletion and the double deletion ΔrsmA ΔtoxR on PAO1-L swarming motility was assessed. Although deletion of toxR resulted in a slight reduction in swarming motility, supporting a link with this phenotype, no swarming deficiency was recorded for the ΔrsmA ΔtoxR double mutant in PAO1-L in contrast to the PAO1-N ΔrsmA strain (ΔrsmA and ~59-kb deletion including toxR) (Fig. 3). We therefore hypothesised that another genetic element missing in PAO1-N could be responsible for the residual swarming activity in the PAO1-L ΔrsmA ΔtoxR mutant. Hence, alternative genomic modifications explaining this divergence were explored. Among the thirteen distinct SNPs/INDELs found when comparing the two PAO1 sublines, a non-synonymous point mutation was predicted in a PAO1-N gene encoding the c-di-GMP degrading PDE BifA (Supplementary Table 1). This mutation results in tyrosine to aspartate substitution at amino acid 442 (Y442D), a position immediately preceding a glutamine (Q443) involved in substrate binding53,54. This suggested that the Y442D substitution in PAO1-N might reduce BifA affinity for its substrate by introducing a negative charge next to the glutamine involved in c-di-GMP binding.
Interestingly, a PAO1 bifA mutant was previously shown to have increased intracellular c-di-GMP levels leading to a non-swarming phenotype22,55,56. Thus, to determine whether defective BifA activity contributed to the swarming motility defect in PAO1-N ΔrsmA and played a role in the differential swarming behaviour between sublines, the motility of strains generated by introducing bifAY442D or bifAD442Y point mutations in PAO1-L WT, PAO1-L ΔrsmA ΔtoxR and PAO1-N ΔrsmA were also studied. As shown in Fig. 3, the triple mutation ΔrsmA ΔtoxR bifAY442D resulted in swarming impairment in PAO1-L. At the same time, the reversion of Y442D to D442Y restored swarming motility in PAO1-N ΔrsmA, suggesting that bifA is the third genetic element involved in this phenotype which potentially explains motility differences between the two PAO1 sublines.
RsmA positively controls toxR expression via PvdS
Given that the alternative sigma factor PvdS is required for optimal toxR expression57 and that pvdS transcript levels were shown to be affected by RsmA4,58, we sought to determine whether RsmA regulates ToxR activity partly through the control of pvdS expression therefore establishing a link between both regulators and swarming motility. To this end, transcriptional fusions of the lux operon to the toxR promoter region were constructed using the mini-CTX system59 and introduced into the PAO1-L WT and the IPTG-inducible rsmAind, ΔpvdS and ΔpvdS rsmAind mutant chromosomes. Moreover, since toxR can be transcribed from two promoters (P1 and P260) we investigated the effect of a rsmA mutation on expression from the complete intergenic region upstream of toxR (PtoxR1,2-luxCDABE) as well as from each promoter separately (PtoxR1/PtoxR2 - luxCDABE) (Supplementary Fig. 3A). Additionally, since the transcription of pvdS is upregulated under iron-limiting conditions61, the toxR reporter strains were tested under iron deficiency using CAA medium.
As shown in Supplementary Fig. 3B, the non-induced PAO1-L rsmAind strain exhibited lower toxR expression levels than WT. Conversely, IPTG-induced expression of rsmA in PAO1-L rsmAind stimulated toxR expression to levels matching or above those of the WT, suggesting a positive impact of RsmA on ToxR production. Results from assays including ΔpvdS and ΔpvdS rsmAind strains showed a significant reduction of P2 promoter activity in the ΔpvdS mutant compared with WT strain. In agreement with previous studies, P2-driven expression was upregulated under low iron conditions57 (Supplementary Fig. 3C). In addition, in the absence of PvdS, RsmA could no longer activate toxR expression, suggesting that the RsmA control on toxR expression is at the transcriptional level via PvdS (Supplementary Fig. 3C).
ToxR impacts on the Rhl QS system expression and rhamnolipid biosynthesis
Previous studies have shown that iron depletion stimulates surface motility in P. aeruginosa62,63. Moreover, rhamnolipid biosynthesis is also upregulated under low iron conditions and is under the control of RhlR30,35,64. The correlation between surface motility, iron regulation and the Rhl QS system led us to hypothesise that the mechanism by which ToxR impacts on swarming could be linked to rhl expression and hence rhamnolipid production. To investigate this, we first compared expression of the rhamnosyltransferase gene rhlA using a PrhlA chromosomal transcriptional fusion to the lux operon in the PAO1-L WT and the toxR conditional IPTG-inducible mutant strain toxRind using the integrative vector mini-CTX. A significant reduction in rhlA transcription in the toxR mutant was observed with respect to the WT strain (Fig. 4a). Conversely, the IPTG-induced expression of toxR restored the expression of rhlA to WT levels (Fig. 4a), indicating that ToxR has a positive impact on rhlA expression. To determine whether the reduced transcription of rhlA observed PAO1-L toxRind resulted in decreased rhamnolipid production, extraction and quantification of these biosurfactants were performed in both strains. Results showed that rhamnolipid yields were reduced in the toxRind strain when compared to WT levels and addition of IPTG stimulated rhamnolipid production (Fig. 4b), confirming the positive impact of ToxR on rhamnolipid biosynthesis.
In all these experiments gene expression and phenotypic analysis were performed in the PAO1-L WT and toxRind mutant strains with and without 1 mM IPTG induction of toxR. a Transcriptional activity of the rhlA promoter assessed at 37 °C in LB by measuring the luminescence emitted by cells containing a PrhlA - luxCDABE transcriptional reporter integrated in the chromosome using the mini-CTX system. Values correspond to the area under the curve (AUC) derived from plotting relative light units normalized to culture density (RLU/OD600) over time (24 h). b Rhamnolipid production measured using the Orcinol method86 from the supernatant fraction of the three strains tested grown in LB at 37 °C for 24 h. c Effect of toxR mutation on the transcriptional activity of the rhlI promoter in LB at 37 °C for 24 h. Values correspond to the area under the curve (AUC) derived from plotting relative light units normalized to culture density (RLU/OD600) over time (24 h). d C4-HSL UV absorbance values (214 nm) normalised to PAO1-L WT and toxRind mutant growth (OD600) in LB at 37 °C for 6 h. C4-HSL was extracted from culture supernatant fractions using acidified ethyl acetate and analysed by LC-MS. Values given are averages from three different cultures ± standard deviation. Statistical differences between group means were determined by one-way ANOVA analysis using Tukey’s multiple comparisons test. (*p < 0.05, **p < 0.01, ***p < 0.005, ****p < 0.0001).
To establish whether ToxR modulates rhamnolipid production via the activation of the Rhl QS system, the effect of the toxR mutation on the expression of rhlI, coding for the C4-HSL autoinducer synthase RhlI, was assessed using a mini-CTXlux transcriptional fusion of PrhlI in PAO1-L toxRind. In the absence of IPTG (ToxR negative), toxRind showed ~25% reduction in rhlI expression compared with the WT. Conversely, the addition of IPTG led to an increase of ~25% in rhlI expression over the WT levels (Fig. 4c), suggesting a positive effect of ToxR on rhlI expression under the conditions tested. We then measured the levels of C4-HSL produced by the WT and toxRind strains using LC-MS. The toxRind strain showed reduced levels of C4-HSL and the addition of IPTG restored the levels of this molecule close to those of the WT strain (Fig. 4d), further supporting an impact of ToxR on the Rhl QS system. These results provide further insights into the mechanism by which ToxR affects swarming motility in PAO1 through the activation of rhamnolipid production via the modulation of the Rhl QS system.
ToxR negatively regulates biofilm formation and Psl/Pel exopolysaccharide production
Swarming motility and biofilm formation are intimately interconnected and described as inversely regulated in P. aeruginosa21,22,23. Given that ToxR modulates swarming motility in PAO1, we monitored biofilm formation by the toxR conditional mutant toxRind and PAO1-L WT strains using Bioflux microfluidic chambers to assess whether ToxR could also affect this phenotype. Our results showed that biofilm growth was enhanced in the absence of toxR expression, while the IPTG-induced transcription of this gene decreased biofilm biomass compared to the parental PAO1-L WT strain (Fig. 5a, b), suggesting that ToxR has a negative impact on biofilm formation. To study whether changes in exopolysaccharide (EPS) biosynthetic genes expression mediated the effect of ToxR on biofilm formation, the activity of the EPS Psl and Pel biosynthetic operon promoters was determined by measuring the fluorescence from cells containing Ppsl-dsRed and Ppel-dsRed reporter fusions, respectively. Results showed that the inactivation of toxR significantly increased psl and pel gene expression especially for the pel promoter fusion. Conversely, toxR complementation through the addition of IPTG reduced the expression of both EPS biosynthetic gene clusters (Fig. 5c, d). These data suggest that ToxR partially represses biofilm formation through a negative impact on EPS production.
a Confocal images of mClover3-labelled P. aeruginosa PAO1-L WT and toxRind mutant ±1 mM IPTG biofilms obtained after 48 h incubation in microfluidic channels (Bioflux, Fluxion) in RPMI-1640 medium. Scale bar: 100 µM. b Quantification of biomass from confocal images of PAO1-L WT and toxRind mutant (±1 mM IPTG) biofilm cultures using Comstat289. c and d Effect of toxR mutation on EPS biosynthetic gene transcription. Expression was determined by measuring the fluorescence from wild type PAO1-L and the toxR inducible mutant toxRind cells (±1 mM IPTG) carrying the transcriptional fusions (c) Ppsl-dsRed and (d) Ppel - dsRed in their chromosomes and grown at 37 C for 24 h in LB. Reported values are averages from three different cultures ± SD and correspond to the area under the curve (AUC) derived from plotting relative fluorescent units normalized to culture density (RLU/OD600) over time. Statistical differences between group means were determined by one-way ANOVA analysis using Tukey’s multiple comparisons test. (*p < 0.05, **p < 0.01, ***p < 0.005, ****p < 0.0001).
ToxR binds c-di-GMP and negatively affects signalling by this second messenger
Our results show that ToxR, in addition to regulating the production of exotoxin A, influences swarming and biofilm formation in P. aeruginosa. Given that both phenotypes are known to be modulated by c-di-GMP signalling, we hypothesised that ToxR might influence c-di-GMP levels impacting those traits. To test this, the intracellular levels of c-di-GMP were semi-quantified in cells transitioning from motility to sessile lifestyles by monitoring the activity of the transcriptional fusion PcdrA-gfp(ASV)C, which responds to the levels of this signal65, in the PAO1-L WT and ΔtoxR mutant strain. As shown in Fig. 6 (and Supplementary Fig. 4), the toxR mutant displayed high levels of cdrA promoter activity, well above that of the WT, consistent with elevated c-di-GMP levels and supporting the hypothesis that ToxR impacts on motility/biofilm formation by reducing c-di-GMP signalling.
a Fluorescence output of PcdrA gfp-expressing PAO1-L and ΔtoxR cells as they transition from liquid culture to glass surface over time (3 h). Scale bar: 20 µm. b Representative combined Differential Interference Contrast (DIC) and widefield epifluorescence images of PAO1-L and ΔtoxR after 3 h exposure to a glass surface. Scale bar: 27 µm. c Graph shows the fraction of cells expressing PcdrA-gfp(ASV)C over time for the first 100 min after attachment to a glass surface. Dots represent mean ±1 SEM (n = 3).
An EAL domain was previously identified in ToxR (Table 1) however, given the absence of conserved residues critical for PDE activity in its sequence (Supplementary Fig. 5), this regulator was assumed to lack c-di-GMP catalytic activity66. To verify that the degenerate EAL domain in ToxR is an enzymatically inactive PDE sequence, a 6×His-fusion ToxR protein was expressed, purified and tested for c-di-GMP degradation activity by LC-MS analysis. No c-di-GMP cleavage was detected, indicating that ToxR does not negatively regulate c-di-GMP signalling through degradation of this second messenger. To test whether c-di-GMP is a ToxR ligand, a Surface Plasmon Resonance (SPR) binding assay using purified ToxR protein was performed as previously described67. As shown in Fig. 7, we were able to detect reproducible concentration-dependent binding of c-di-GMP by ToxR, with a calculated KD of 2.36 ± 0.7 μΜ. This indicates that ToxR is a c-di-GMP-binding protein and suggests that ToxR may impact on swarming and biofilm formation through direct stimulation of additional enzymes able to metabolise this signalling molecule explaining the altered c-di-GMP levels in the toxR mutant.
Purified ToxR was tested for dinucleotide binding by Surface Plasmon Resonance (SPR) as previously described67. a SPR sensorgrams showing affinity measurements of an increasing range of ToxR concentrations (0–5 µM) binding to biotinylated c-di-GMP. Protein binding and dissociation phases are shown. b Affinity fit for ToxR-c-di-GMP binding. The binding response for each concentration was recorded, and the KD values for ToxR binding to c-di-GMP (2.36 ± 0,7 µM) were calculated using Biacore T200 BiaEvaluation software version 1.0 (GE Healthcare) and confirmed by GraphPad Prism. The experiment was repeated three times independently.
Discussion
An analysis of the genetic differences between P. aeruginosa PAO1-Lausanne and PAO1-Nottingham sublines, combined with an exhaustive screening for genes able to complement swarming in PAO1-N ΔrsmA mutant using a plasmid library containing random chromosomal fragments of PAO1-L, allowed us to identify new genetic elements restoring this motility. The robustness of our approach is supported by the finding of genes previously described to be involved in swarming i.e. bifA and rsmN22,46, but also ORFs encoding proteins with conserved domains implicated in c-di-GMP metabolism and hence expected to have an impact on this surface-associated motility15,22,23,68. While seven potential genes able to restore a swarming deficiency in PAO1-N ΔrsmA were identified, we have not yet characterised these further and focused on toxR, currently known to encode a regulator of ETA biosynthesis in P. aeruginosa44.
Remarkably, the P. aeruginosa PAO1-N subline used in our laboratory, and which exhibits a reduction in swarming motility compared to PAO1-Lausanne, contains a substantial deletion in its chromosome including the regulatory gene toxR. This 59-kb deletion was found to be present in strain PANO67, the first P. aeruginosa mutant selected in our laboratory69, suggesting that it was already present in the PAO1 subline that we obtained over 30 years ago, but the exact origin of which can no longer be ascertained. Additionally, a non-synonymous point mutation in bifA, encoding a c-di-GMP PDE with known roles in motility and early biofilm formation22, was identified in PAO1-N that could potentially affect substrate binding by this enzyme. Although mutation of toxR or bifA causes only a partial swarming defect in PAO1-L, here we show that a ToxR deficiency in combination with that of the regulatory proteins RsmA and BifA leads to total swarming impairment in this PAO1 subline. Furthermore, RsmN was found to bind the bifA transcript before3 suggesting a link between these two factors influencing swarming.
Our results indicate that ToxR affects swarming by stimulating expression of the autoinducer synthase RhlI and subsequently increasing the levels of the signal C4-HSL and the rhamnolipid surfactants, which support migration by lowering surface tension thus facilitating flagella-based propulsion28. Interestingly, a previous study reported that low levels of c-di-GMP induce the expression of the Rhl QS operon, and this finding correlated with an increased production of Rhl-regulated virulence factors including rhamnolipids70. Moreover, our results show that ToxR has an impact on biofilm formation, a phenotype closely associated with chronic infections. This control could be exerted, at least partially, through repression of Psl and Pel production as we observed that ToxR negatively impacts on the expression of the biosynthetic genes for these exopolysaccharides. Regulation of Psl and Pel has been shown to be controlled by c-di-GMP through the activity of PDEs and DGCs47,71. Given these factors and the fact that c-di-GMP modulates the transition from motile to sessile (biofilm) lifestyles in P. aeruginosa71, it seems reasonable that ToxR could indirectly exert its effect on these processes through governing c-di-GMP levels in the cell.
Despite the prediction that ToxR incorporates a single putative EAL domain, an ubiquitous motif linked to PDEs and c-di-GMP turnover in bacteria72, we could not detect c-di-GMP-specific PDE activity when testing the purified protein in vitro. Consistent with this result, ToxR was previously defined as a stand-alone EAL domain protein and expected to lack catalytic activity against c-di-GMP due to the absence of extra domains assisting dimerisation and lack of conserved residues essential for c-di-GMP metabolism54,66. Although catalytically inactive, EAL domain proteins can retain the ability to bind c-di-GMP and therefore additional interaction partners with these proteins are anticipated66,73. Indeed, our results show that ToxR binds c-di-GMP with high affinity and the intracellular levels of this second messenger increase significantly in a toxR mutant compared with the WT strain, suggesting that the ToxR regulon may contain additional c-di-GMP-metabolizing enzymes responsible for ultimately controlling the levels of this intracellular signal leading to the changes in gene expression and the phenotypes observed. Therefore, identification of the residues within ToxR that are involved in c-di-GMP binding could form part of future studies. In particular, it will be interesting to see if a toxR mutant in the EXLXR motif is still able to bind c-di-GMP and influence surface-associated behaviour.
Notably, iron depletion, an environmental parameter that stimulates toxR expression57, represses biofilm formation and promotes biosynthesis of the pyoverdine siderophore necessary for acute infections41 are conditions where low levels of c-di-GMP are expected. In addition, we show that RsmA exerts positive control over ToxR via PvdS, strengthening the established links between the Rsm and c-di-GMP regulatory networks74 (Fig. 8). The positive impact of RsmA on pvdS expression under iron limiting conditions found here supports the results of Brencic and Lory4 who showed a reduction in pvdS transcript levels in an rsmA mutant compared with the WT P. aeruginosa PAK strain and suggests that this regulation is conserved across different strains. The link between RsmA, PvdS, ToxR and c-di-GMP signalling is further supported by previous work from Jones et al.75 in which the authors demonstrated that mutants in AmrZ, a transcriptional factor important for P. aeruginosa virulence accumulates intracellular c-di-GMP, and AmrZ increased expression of RsmA (1.55 fold) and repressed many genes involved in iron acquisition including PvdS (1.65 fold) and ToxR (1.5 fold)75.
Previous attempts to determine the mechanism by which ToxR activates ETA production have found that ToxR can function with the RNA polymerase of E. coli and P. aeruginosa to efficiently transcribe toxA both in vitro and in vivo and the ToxR-mediated enhancement of toxA expression is improved in the presence of cytoplasmic extracts of Pseudomonas aeruginosa76,77. Moreover, ToxR has been shown to be required for open-complex formation at the toxA promoter77. Interestingly, several studies have documented that c-di-GMP can control the switch between acute and chronic infection modes in bacteria by facilitating protein-protein and protein-DNA interactions72. Therefore, it is tempting to hypothesise that c-di-GMP could facilitate ToxR regulation of transcription, for instance through modulation of ToxR stability, as it has been reported that c-di-GMP binding may lead to enhanced stability of PgaCD interaction in E. coli78. However, given that EAL proteins have not been shown to bind DNA to date but are known to bind other proteins, stimulation of another regulatory protein or a ToxR-RNA polymerase interaction seems to be a more likely model for ToxR function than direct transcriptional control.
Overall, the work presented here shows that ToxR, originally thought to exclusively activate ETA production, has a negative impact on biofilm formation through the modulation of Psl and Pel exopolysaccharides, and a positive impact on swarming through activation of the Rhl QS system and hence rhamnolipid production. This regulation is also modulated by the post-transcriptional regulator RsmA via PvdS and hence iron depletion. However, the molecular basis by which ToxR impacts on these regulatory networks through c-di-GMP binding remains to be elucidated.
Methods
Bacterial strains and culture conditions
Strains, plasmids and oligonucleotides used in this study are listed in Supplementary Table 3. E. coli and P. aeruginosa strains were routinely grown at 37 °C on lysogeny broth (LB) or LB agar supplemented with antibiotics as required. For studying toxR expression, toxR reporter strains were grown under iron deficiency using CAA medium79. IPTG was added to cultures at 1 mM final concentration to complement inducible mutant strains. Swarming assays were performed in 60 ×15 mm dishes with 8 g/L Nutrient Broth No.2 (Oxoid), 0.5% w/v Bacto agar (Difco) and 0.5% w/v D-glucose (Sigma). Plates were inoculated with 3 µL of OD600 1.0 ON cultures washed in fresh LB. Plates were then incubated for 16 h at 37 °C.
Biofilms were grown continuously in microfluidic channels using a Bioflux system (Fluxion). Briefly, mClover3-labelled80 P. aeruginosa PAO1-L WT and toxRind mutant strains were grown at 37 °C for 16 h in 2 mL Lysogeny broth (LB) broth. The cultures were diluted to OD600 0.05 in RPMI-1460 (Lonza) and used to seed the microfluidic channels. The biofilm was allowed to form at a flow rate of 16 μL/h (corresponding to a shear rate of 0.5 dyn/cm2) at a temperature of 37 °C for a period of 48 h in RPMI-1640 (Lonza Bioscience).
P. aeruginosa subline PAO1-L genomic library
To find genetic elements in PAO1-L restoring swarming motility in P. aeruginosa PAO1-N ΔrsmA, a genomic library was generated by partially digesting the PAO1-L chromosome with Sau3AI and 2–4 kb fragments cloned into pME6000 plasmid. The resulting plasmids were transformed into P. aeruginosa PAO1-N ΔrsmA and each clone screened for restoration of swarming46.
Sequencing of P. aeruginosa PAO1 sublines and optical restriction genome mapping
To catalogue single nucleotide polymorphisms (SNPs) and small insertions/deletions (INDELs) in PAO1-L and PAO1-N sublines, paired-end sequencing libraries were prepared using the Illumina Nextera kit according to the manufacturer’s protocols (Illumina), and whole-genome sequencing was performed on an Illumina MiSeq platform. Whole-genome sequence datasets were subjected to adaptor trimming using Trimmomatic version 0.3080 and hard trimming at the 5′ end using a custom Perl script. Trimmed reads were aligned to the modified PAO1-UW sequence3 using Smalt version 0.7.0.1 (sanger.ac.uk/tool/smalt-0/) with a kmer of 12 and a step size of 2. The alignment files were sorted, and PCR duplicates were removed using Samtools81. INDEL realignment was carried out with the Genome Analysis Toolkit82, and SNPs and INDELs were determined using Samtools mpileup, employing varying base quality thresholds (parameter-Q). SNPs were filtered to exclude those with low coverage (<5x) and those not supported by at least two reads on each strand using a custom Perl script. A custom Perl script was used to characterise SNPs in genes, and the resulting amino acid changes. Regions with zero read coverage were detected using the “genomeCoverageBed” functionality of bedtools with the -bga option (version 2.17.0)83. De novo assembly was carried out using Velvet version 1.2. with a kmer of 121 with no scaffolding and the expected coverage flag set to auto.
Large scale chromosomal rearrangements, deletions and insertions in sublines PAO1-L and PAO1-N were analysed by optical restriction mapping50,51. Briefly, high molecular weight (HMW) DNA was isolated from single colony cultures of P. aeruginosa PAO1-L and PAO1-N using the HMW DNA Isolation Kit (Opgen Inc.) according to the manufacturer’s instructions. The quality of HMW DNA was assessed using an Argus QCard Kit (Opgen Inc.), and the concentration of the HMW DNA was adjusted using dilution buffer. DNA molecules were stretched and linearised on charged glass coverslips using a high-density channel-forming device (CFD). The restriction endonuclease BamHI was used in an Argus MapCard (Opgen Inc), and the glass coverslips were incubated in a MapCard Processor to allow digestion of the DNA molecules on the glass surface. The fragmented DNA was then stained with intercalating fluorescent dye, and the DNA molecules were imaged under a high-resolution microscope. DNA molecules were assembled into a whole-genome map using the OpGen Genome-Builder software. Optical maps were aligned to the reference P. aeruginosa subline PAO1-UW49 (GenBank accession NC_002516) using the OpGen MapSolver software and visually inspected to locate regions of difference >2 kb, the lower limit for efficient optical mapping.
Mutant strain construction
In-frame deletion mutants were obtained by two-step allelic exchange using pDM4 or pME3087 vectors. To construct P. aeruginosa PAO1-L ΔrsmA, ΔtoxR and ΔpvdS mutants, two PCR products amplifying each gene’s upstream and downstream nucleotide regions were generated using the primer pairs D1FW/D1RV and D2FW/D2RV, respectively (Supplementary Table 3). PCR products were ligated together in pBluescript II KS+ to create a deletion in the corresponding gene and subsequently cloned into the suicide vector pDM4 or pME3087 resulting in plasmids pZH13, pJD22 and pJD93, respectively. Following transformation into the target strain by conjugation, single crossovers were selected on chloramphenicol (Cm, 375 μg mL−1) or tetracycline (Tc, 125 μg mL−1). To achieve rsmA and toxR deletion, recombinants resistant to Cm were grown in LB supplemented with 10% sucrose overnight and plated on LB-10% sucrose. The carbenicillin enrichment method selected the pvdS double-crossover mutants84. The resulting P. aeruginosa colonies were screened for the loss of antibiotic resistance by plating on LB supplemented with or without Cm/Tc. PCR and sequence analysis confirmed the in-frame deletions.
The conditional mutant IPTG-inducible toxR (toxRind) strain was constructed by introducing the lacIQ repressor gene and the tac promoter transcribing the lacZ 5′ untranslated region and its ribosome binding site (RBS) directly upstream of the toxR open reading frame, resulting in constitutive transcription and translation only in the presence of IPTG. The suicide plasmid pAMP5 was constructed in several stages and is a pDM4 derivative that carries, in sequential order, the following elements: (a) a 365 bp HindIII-EcoRI fragment of the upstream region of toxR obtained by PCR using primers toxRpFW/RV (Supplementary Table 3); (b) the 2.0 kb BamHI Sm/Spc integron from pHP45Ω; (c) the 1.5 kb BamHI-EcoRI lacIQ Ptac inducible promoter fragment of pME6032 and (d) an 0.5 kb EcoRI-XhoI fragment carrying the toxR open reading frame (obtained from pBluescript II KS+-based vector constructed for the ΔtoxR mutant). The mutant was obtained by double crossover of the fragment carried by pAMP5 in P. aeruginosa PAO1-L and selection of sucrose and streptomycin-resistant clones.
To generate the bifAY442D mutant in PAO1-L and derivative mutants as well as to revert the bifA point mutation in PAO1-N, the bifA gene was amplified from PAO1-N and PAO1-L genomes using the primer pair BifAFW/BifARS (Supplementary Table 3). The resulting PCR products were ligated with pBlueScript II KS+ and subsequently cloned as HindIII-EcoRI fragments into the suicide plasmid pME3087, generating the plasmids pJL103 and pJL108 and used for double crossover recombination. The resulting P. aeruginosa colonies were screened for the loss of antibiotic resistance by plating on LB supplemented with or without Tc. PCR and sequence analysis confirmed the mutations.
Transcriptional reporter fusions
The promoter regions of toxR, rhlA and rhlI were amplified from the PAO1-L genome using the primer pairs ToxRp1FW/RV, ToxRp2FW/RV, ToxRp12FW/RV, RhlApFW/RV and RhlIpFW/RV (Supplementary Table 3). The resulting PCR products were ligated into the mini-CTXlux vector using the appropriate restriction enzyme to generate the transcriptional reporters pAMP11 (mini-CTX::PtoxR1 - luxCDABE), pAMP12 (mini-CTX::PtoxR2 - luxCDABE), pAMP13 (mini-CTX::PtoxR1,2 - luxCDABE), pSH30 (mini-CTX::PrhlA - luxCDABE) and pSH32 (mini-CTX::PrhlI - luxCDABE). Finally, the transcriptional reporters were integrated into the chromosome of P. aeruginosa PAO1 strains by conjugation with E. coli S17.1 λpir followed by selection with Tc.
For Ppel - dsRed and Ppsl-dsRed transcriptional reporter construction, the promoter regions of pelA and pslA genes were amplified by PCR from P. aeruginosa PAO1-L chromosomal DNA using primer pairs PpelAdsRedFW/RV and PpslAdsRedFW/RV, respectively. The reporter dsRed-express2 gene was amplified using primer pairs dsRedFW/RW from pCMVDsRed-Express2 vector and cloned into pBluescript II KS+, resulting in plasmid pSC581. Then dsRed-express2 cut from pSC581 with HindIII/EcoRI, and the KpnI/HindIII cut pelA or pslA promoter regions were ligated into pUCP18, resulting in plasmids pSC855 and pSC856.
Clones with active fusions were selected and analysed for bioluminescence or fluorescence output activity over growth in CAA or LB media at 37 °C using a 96-well plate TECAN Genios Pro multifunction microplate reader.
C4-HSL and rhamnolipid quantification
P. aeruginosa strains were grown for 6 h (early stationary phase) in LB broth at 37 °C with vigorous shaking (200 rpm). QS molecules were extracted from 5 mL culture by adding 5 mL of acidified ethyl acetate85. Sample separation was performed on an Agilent 1200 series HPLC with an Ascentis Express C18 150 ×2.1 mm internal diameter, 2.7 mm particle size, maintained at 50 °C. The mobile phase consisted of formic acid 0.1% (v/v) in water and formic acid 0.1% (v/v) in acetonitrile run as a gradient over 20 min at a flow rate of 0.3 ml min−1. C4-HSL identification was performed using a Bruker HCT Plus ion trap LC-mass spectrometer and Hystar software (Bruker). Retention times and MS/MS peak spectra were matched to 10 mM synthetic C4-HSL standard. For rhamnolipids quantification strains were grown at 37 °C in 50-ml Erlenmeyer flasks containing 10 ml of M9 medium supplemented with glycerol (2% vol/vol), glutamate (0.05%), and Triton X-100 (0.05%) with shaking for 18 h. Rhamnolipids were extracted with three volumes of diethyl ether from culture supernatants filtered through a 0.22-μm-pore-size membrane. Extracts were extracted once with 20 mM HCl, and the ether phase was evaporated to dryness. The residue was dissolved in water. Rhamnose content in each sample was determined by duplicate oricinol assays compared to rhamnose standards. Rhamnolipid was determined by the relation that 1.0 mg of rhamnose corresponds to 2.5 mg of rhamnolipid11,86.
ToxR purification
The pBAD TOPO TA expression system (Invitrogen) was used to produce His-tagged ToxR protein within host E. coli BL21 (DE3) cells. The toxR gene was amplified from PAO1-L genome using primers ToxRtopoFW/RV and cloned into pBAD TOPO vector according to manufacturer’s instructions. E. coli BL21 (DE3) cells carrying the resulting pAMP14 vector were cultured overnight in LB at 37 °C with vigorous shaking (200 rpm), diluted 1:100 in Terrific Broth (TB) and grown to OD600 0.4 at 37 °C before protein expression was induced overnight with 0.2% L-Arabinose at 18 °C. Cells were then lysed by sonication and centrifuged at 15,000 × g for 1 h. ToxR was purified from the supernatant by NTA-Ni chromatography. HiTrap chelating HP columns of 1 mL (GE healthcare, life sciences) were equilibrated with 10 volumes of washing buffer (20 mM HEPES pH 7.5, 250 mM NaCl, 2 mM MgCl2, and 2.5% (v/v) glycerol pH 6.8) and loaded with cell lysate. Following protein immobilisation, the column was washed with 10 volumes of buffer containing 50 mM imidazole before the protein was eluted using 500 mM imidazole buffer in a single step elution. Due to ToxR instability, it was required that later assays were performed within a few hours after protein purification.
ToxR phosphodiesterase activity and c-di-GMP binding detection methods
To assess cleavage of c-di-GMP by a possible PDE activity of ToxR, purified ToxR (0.7 mg) was incubated with c-di-GMP (100 μM) (Biolog Life Science Institute) for 2 h at 30 °C. The remaining c-di-GMP in the samples was detected by liquid chromatography-mass spectrometry (LC-MS) analysis using the TSQ Quantum Access Max system (Thermo Scientific) in positive-ion mode, as previously described87. The areas of the major MS-MS fragments of c-di-GMP (m/z 152 and 540) were measured, and a standard curve was established using defined concentrations of c-di-GMP to allow sample quantification. Moreover, the main degradation product of c-di-GMP cleavage, 5′-phosphoguanylyl-(3′→5′)-guanosine (pGpG), was also quantified in the samples by measuring the area of the major fragment (m/z = 152) of the isolated precursor ion (m/z = 709) corresponding to pGpG as determined using a standard (Biolog).
To test whether c-di-GMP is in fact a ToxR ligand, a Surface Plasmon Resonance (SPR) binding assay using purified ToxR protein was performed67. Briefly, a Biacore T200 system (GE Healthcare) equipped with a Streptavidin SA sensor chip (GE healthcare), containing four flow cells with a SA pre-immobilized to a carboxymethylated dextran matrix each, was used. Flow cell one (FC1) and flow cell three (FC3) were kept blank for reference subtraction. The chip was washed three times with 1 M NaCl and 50 mM NaOH to remove any unconjugated streptavidin. Biotinylated c-di-GMP (100 nM) (BioLog) was immobilised on FC2 and FC4 of the streptavidin chip at a 50 RU immobilisation level with a flow rate of 5 μL/min. Soluble ToxR protein was prepared in SPR buffer (10 mM HEPES, 500 mM NaCl, 0.1% (v/v) Tween 20, 2 mM MgCl2, pH 6.8). Samples were injected with a flow rate of 5 μL/min over the reference and c-di-GMP cells for 60 s, followed by buffer flow for 60 s. The chip was washed at the end of each cycle with 1 M NaCl. An increasing range of protein concentrations (78.125 nM, 156.25 nM, 312.5 nM, 625 nM, 1.25 µM, 2.5 µM, 5.0 µM) was used, with replicates for each protein concentration included as appropriate. All sensorgrams were analysed using Biacore T200 BiaEvaluation software version 1.0 (GE Healthcare). The experiment was repeated three times independently.
Microscopy and image analysis methods
P. aeruginosa PAO1-L WT and ΔtoxR mutant strains carrying the plasmid PcdrA-gfp(ASV)C 65 were used to report c-di-GMP levels in cells transitioning from liquid culture to a glass surface over time (3 h). For time-lapse imaging, a bespoke multimode microscope (Cairn Ltd)88, based in an inverted Eclipse Ti microscope (Nikon), was used. The microscope was fitted with an environmental chamber (Okolab) to regulate temperature and relative humidity. Differential Interference Contrast (DIC) and widefield epifluorescence imaging were carried out using a single channel white MonoLED light source (Cairn Ltd) and a 40× objective (Nikon, CFI Plan Fluor 40×/1.3). Images were acquired using an Orca-Flash 4.0 digital CMOS camera (Hamamatsu) every 30 sec per hour. Experiments were conducted using a 35 mm glass-bottom dish (Cellvis). The fraction of cells expressing PcdrA-gfp(ASV)C was calculated by counting objects in each frame in both DIC and epifluorescence mode. Images were processed in ImageJ (NIH), and an automatic threshold was applied using the Yen method to each data set. Bacterial tracks were processed through a custom single-cell tracking algorithm88 to determine the average fluorescence (F values) of tracks corresponding to PcdrA-gfp(ASV)C expression levels.
Biofilms expressing the fluorescent protein mClover3 were visualised under a Confocal Laser Scanning Microscope (LSM700, Carl Zeiss) using 488 nm laser. At least 5 replicate Z-stack biofilm images were taken randomly, and biomass was quantified using Comstat2 plugin89 in ImageJ software.
Statistical analysis
One-way ANOVA analysis using Tukey’s posthoc multiple comparisons tests were applied to determine whether mutants’ responses differed significantly from that of the parental strain (p < 0.05) when compared with the variations within the replicates (n = 3) using GraphPad Prism 8.0 (GraphPad Software, Inc., San Diego, CA).
Data availability
Whole genome sequencing results reported in this paper have been deposited at the National Center for Biotechnology Information (NCBI), under BioProject No. PRJNA830884. Assembled genome sequences for sublines PAO1-L and PAO1-N are available from GenBank, Accession Nos. CP099798 and CP099797, respectively. The data that support the findings of this study are available from the corresponding author, MC, upon reasonable request.
References
Moradali, M. F., Ghods, S. & Rehm, B. H. Pseudomonas aeruginosa lifestyle: a paradigm for adaptation, survival, and persistence. Front. Cell. Infect. Microbiol. 7, 39 (2017).
Schuster, M. & Greenberg, E. P. A network of networks: quorum-sensing gene regulation in Pseudomonas aeruginosa. Int. J. Med. Microbiol.: IJMM 296, 73–81 (2006).
Romero, M. et al. Genome-wide mapping of the RNA targets of the Pseudomonas aeruginosa riboregulatory protein RsmN. Nucleic acids Res. 46, 6823–6840 (2018).
Brencic, A. & Lory, S. Determination of the regulon and identification of novel mRNA targets of Pseudomonas aeruginosa RsmA. Mol. Microbiol. 72, 612–632 (2009).
Romeo, T., Gong, M., Liu, M. Y. & Brun-Zinkernagel, A. M. Identification and molecular characterization of csrA, a pleiotropic gene from Escherichia coli that affects glycogen biosynthesis, gluconeogenesis, cell size, and surface properties. J. Bacteriol. 175, 4744–4755 (1993).
Schubert, M. et al. Molecular basis of messenger RNA recognition by the specific bacterial repressing clamp RsmA/CsrA. Nat. Struct. Mol. Biol. 14, 807–813 (2007).
Schulmeyer, K. H. et al. Primary and secondary sequence structure requirements for recognition and discrimination of target RNAs by Pseudomonas aeruginosa RsmA and RsmF. J. Bacteriol. 198, 2458–2469 (2016).
Allsopp, L. P. et al. RsmA and AmrZ orchestrate the assembly of all three type VI secretion systems in Pseudomonas aeruginosa. Proc. Natl Acad. Sci. USA 114, 7707–7712 (2017).
Irie, Y. et al. Pseudomonas aeruginosa biofilm matrix polysaccharide Psl is regulated transcriptionally by RpoS and post-transcriptionally by RsmA. Mol. Microbiol. 78, 158–172 (2010).
Goodman, A. L. et al. A signaling network reciprocally regulates genes associated with acute infection and chronic persistence in Pseudomonas aeruginosa. Dev. Cell 7, 745–754 (2004).
Heurlier, K. et al. Positive control of swarming, rhamnolipid synthesis, and lipase production by the posttranscriptional RsmA/RsmZ system in Pseudomonas aeruginosa PAO1. J. Bacteriol. 186, 2936–2945 (2004).
Moscoso, J. A., Mikkelsen, H., Heeb, S., Williams, P. & Filloux, A. The Pseudomonas aeruginosa sensor RetS switches type III and type VI secretion via c-di-GMP signalling. Environ. Microbiol. 13, 3128–3138 (2011).
Valentini, M. & Filloux, A. Biofilms and cyclic di-GMP (c-di-GMP) signaling: lessons from Pseudomonas aeruginosa and other bacteria. J. Biol. Chem. 291, 12547–12555 (2016).
Chang, A. L. et al. Phosphodiesterase A1, a regulator of cellulose synthesis in Acetobacter xylinum, is a heme-based sensor. Biochemistry 40, 3420–3426 (2001).
Simm, R., Morr, M., Kader, A., Nimtz, M. & Romling, U. GGDEF and EAL domains inversely regulate cyclic di-GMP levels and transition from sessility to motility. Mol. Microbiol. 53, 1123–1134 (2004).
Sun, S. & Pandelia, M.-E. HD-[HD-GYP] phosphodiesterases: activities and evolutionary diversification within the HD-GYP family. Biochemistry 59, 2340–2350 (2020).
Tal, R. et al. Three cdg operons control cellular turnover of cyclic di-GMP in Acetobacter xylinum: genetic organization and occurrence of conserved domains in isoenzymes. J. Bacteriol. 180, 4416–4425 (1998).
Römling, U., Gomelsky, M. & Galperin, M. Y. C-di-GMP: the dawning of a novel bacterial signalling system. Mol. Microbiol. 57, 629–639 (2005).
Ryjenkov, D. A., Tarutina, M., Moskvin, O. V. & Gomelsky, M. Cyclic diguanylate is a ubiquitous signaling molecule in bacteria: insights into biochemistry of the GGDEF protein domain. J. Bacteriol. 187, 1792–1798 (2005).
Schmidt, A. J., Ryjenkov, D. A. & Gomelsky, M. The ubiquitous protein domain EAL is a cyclic diguanylate-specific phosphodiesterase: enzymatically active and inactive EAL domains. J. Bacteriol. 187, 4774–4781 (2005).
Caiazza, N. C., Shanks, R. M. Q. & O’Toole, G. A. Rhamnolipids modulate swarming motility patterns of Pseudomonas aeruginosa. J. Bacteriol. 187, 7351–7361 (2005).
Kuchma, S. L. et al. BifA, a cyclic-Di-GMP phosphodiesterase, inversely regulates biofilm formation and swarming motility by Pseudomonas aeruginosa PA14. J. Bacteriol. 189, 8165–8178 (2007).
Römling, U., Galperin, M. Y. & Gomelsky, M. Cyclic di-GMP: the first 25 years of a universal bacterial second messenger. Micro Mol. Biol. R. 77, 1–52 (2013).
Kearns, D. B. A field guide to bacterial swarming motility. Nat. Rev. Microbiol 8, 634 (2010).
Daniels, R., Vanderleyden, J. & Michiels, J. Quorum sensing and swarming migration in bacteria. FEMS Microbiol. Rev. 28, 261–289 (2004).
Morris, J. D. et al. Imaging and analysis of Pseudomonas aeruginosa swarming and rhamnolipid production. Appl Environ. Micro. 77, 8310–8317 (2011).
Overhage, J., Lewenza, S., Marr, A. K. & Hancock, R. E. Identification of genes involved in swarming motility using a Pseudomonas aeruginosa PAO1 mini-Tn5-lux mutant library. J. Bacteriol. 189, 2164–2169 (2007).
Wang, J., Yu, B., Tian, D. & Ni, M. Rhamnolipid but not motility is associated with the initiation of biofilm seeding dispersal of Pseudomonas aeruginosa strain PA17. J. Biosci. 38, 149–156 (2013).
Kohler, T., Curty, L. K., Barja, F., van Delden, C. & Pechere, J. C. Swarming of Pseudomonas aeruginosa is dependent on cell-to-cell signaling and requires flagella and pili. J. Bacteriol. 182, 5990–5996 (2000).
Glick, R. et al. Increase in rhamnolipid synthesis under iron-limiting conditions influences surface motility and biofilm formation in Pseudomonas aeruginosa. J. Bacteriol. 192, 2973–2980 (2010).
Campos-García, J. et al. The Pseudomonas aeruginosa rhlG gene encodes an NADPH-dependent beta-ketoacyl reductase which is specifically involved in rhamnolipid synthesis. J. Bacteriol. 180, 4442–4451 (1998).
Deziel, E., Lepine, F., Milot, S. & Villemur, R. rhlA is required for the production of a novel biosurfactant promoting swarming motility in Pseudomonas aeruginosa: 3-(3-hydroxyalkanoyloxy)alkanoic acids (HAAs), the precursors of rhamnolipids. Microbiol. (Read., Engl.) 149, 2005–2013 (2003).
Ochsner, U. A., Fiechter, A. & Reiser, J. Isolation, characterization, and expression in Escherichia coli of the Pseudomonas aeruginosa rhlAB genes encoding a rhamnosyltransferase involved in rhamnolipid biosurfactant synthesis. J. Biol. Chem. 269, 19787–19795 (1994).
Rahim, R. et al. Cloning and functional characterization of the Pseudomonas aeruginosa rhlC gene that encodes rhamnosyltransferase 2, an enzyme responsible for di-rhamnolipid biosynthesis. Mol. Microbiol. 40, 708–718 (2001).
Ochsner, U. A., Koch, A. K., Fiechter, A. & Reiser, J. Isolation and characterization of a regulatory gene affecting rhamnolipid biosurfactant synthesis in Pseudomonas aeruginosa. J. Bacteriol. 176, 2044–2054 (1994).
Liu, P. V. Exotoxins of Pseudomonas aeruginosa. I. Factors that influence the production of exotoxin A. J. Infect. Dis. 128, 506–513 (1973).
Liu, P. V., Yoshii, S. & Hsieh, H. Exotoxins of Pseudomonas aeruginosa. II. Concentration, purification, and characterization of exotoxin A. J. Infect. Dis. 128, 514–519 (1973).
Michalska, M. & Wolf, P. Pseudomonas exotoxin A: optimized by evolution for effective killing. Front Microbiol 6, 963–963 (2015).
Iglewski, B. H., Liu, P. V. & Kabat, D. Mechanism of action of Pseudomonas aeruginosa exotoxin Aiadenosine diphosphate-ribosylation of mammalian elongation factor 2 in vitro and in vivo. Infect. Immun. 15, 138–144 (1977).
Chang, J. H. & Kwon, H. Y. Expression of 14-3-3delta, cdc2 and cyclin B proteins related to exotoxin A-induced apoptosis in HeLa S3 cells. Int. Immunopharmacol. 7, 1185–1191 (2007).
Cornelis, P. & Dingemans, J. Pseudomonas aeruginosa adapts its iron uptake strategies in function of the type of infections. Front. Cell. Infect. Microbiol. 3, 75 (2013).
Hunt, T. A., Peng, W. T., Loubens, I. & Storey, D. G. The Pseudomonas aeruginosa alternative sigma factor PvdS controls exotoxin A expression and is expressed in lung infections associated with cystic fibrosis. Microbiol. (Read., Engl.) 148, 3183–3193 (2002).
Lamont, I. L., Beare, P. A., Ochsner, U., Vasil, A. I. & Vasil, M. L. Siderophore-mediated signaling regulates virulence factor production in Pseudomonas aeruginosa. Proc. Natl Acad. Sci. USA 99, 7072–7077 (2002).
Hedstrom, R. C., Funk, C. R., Kaper, J. B., Pavlovskis, O. R. & Galloway, D. R. Cloning of a gene involved in regulation of exotoxin A expression in Pseudomonas aeruginosa. Infect. Immun. 51, 37–42 (1986).
Wozniak, D. J., Cram, D. C., Daniels, C. J. & Galloway, D. R. Nucleotide sequence and characterization of toxR: a gene involved in exotoxin A regulation in Pseudomonas aeruginosa. Nucleic Acids Res. 15, 2123–2135 (1987).
Morris, E. R. et al. Structural rearrangement in an RsmA/CsrA ortholog of Pseudomonas aeruginosa creates a dimeric RNA-binding protein, RsmN. Structure 21, 1659–1671 (2013).
Feng, Q. et al. Regulation of exopolysaccharide production by ProE, a cyclic-Di-GMP phosphodiesterase in Pseudomonas aeruginosa PAO1. Front. Microbiol. https://doi.org/10.3389/fmicb.2020.01226 (2020).
Kulesekara, H. et al. Analysis of Pseudomonas aeruginosa diguanylate cyclases and phosphodiesterases reveals a role for bis-(3′-5′)-cyclic-GMP in virulence. Proc. Natl. Acad. Sci. USA 103, 2839–2844 (2006).
Stover, C. K. et al. Complete genome sequence of Pseudomonas aeruginosa PAO1, an opportunistic pathogen. Nature 406, 959–964 (2000).
Jing, J. et al. Automated high resolution optical mapping using arrayed, fluid-fixed DNA molecules. PNAS 95, 8046–8051 (1998).
Lim, A. et al. Shotgun optical maps of the whole Escherichia coli O157:H7 genome. Genome Res 11, 1584–1593 (2001).
Klockgether, J. et al. Genome diversity of Pseudomonas aeruginosa PAO1 laboratory strains. J. Bacteriol. 192, 1113–1121 (2010).
Minasov, G. et al. Crystal structures of YkuI and its complex with second messenger cyclic Di-GMP suggest catalytic mechanism of phosphodiester bond cleavage by EAL domains. J. Biol. Chem. 284, 13174–13184 (2009).
Rao, F. et al. The Functional Role of a Conserved Loop in EAL Domain-Based Cyclic di-GMP-Specific Phosphodiesterase. J. Bacteriol. 191, 4722–4731 (2009).
Kuchma, S. L. et al. Cyclic-di-GMP-mediated repression of swarming motility by Pseudomonas aeruginosa: the pilY1 gene and its impact on surface-associated behaviors. J. Bacteriol. 192, 2950–2964 (2010).
Kuchma, S. L., Griffin, E. F. & O’Toole, G. A. Minor pilins of the type IV pilus system participate in the negative regulation of swarming motility. J. Bacteriol. 194, 5388–5403 (2012).
Gaines, J. M. et al. Regulation of the Pseudomonas aeruginosa toxA, regA and ptxR genes by the iron-starvation sigma factor PvdS under reduced levels of oxygen. Microbiol. (Read., Engl.) 153, 4219–4233 (2007).
Burrowes, E., Baysse, C., Adams, C. & O’Gara, F. Influence of the regulatory protein RsmA on cellular functions in Pseudomonas aeruginosa PAO1, as revealed by transcriptome analysis. Microbiol. (Read., Engl.) 152, 405–418 (2006).
Hoang, T. T., Kutchma, A. J., Becher, A. & Schweizer, H. P. Integration-proficient plasmids for Pseudomonas aeruginosa: site-specific integration and use for engineering of reporter and expression strains. Plasmid 43, 59–72 (2000).
Storey, D. G., Frank, D. W., Farinha, M. A., Kropinski, A. M. & Iglewski, B. H. Multiple promoters control the regulation of the Pseudomonas aeruginosa regA gene. Mol. Microbiol. 4, 499–503 (1990).
Reinhart, A. A. & Oglesby-Sherrouse, A. G. Regulation of Pseudomonas aeruginosa virulence by distinct iron sources. Genes (Basel) 7, 126 (2016).
Patriquin, G. M. et al. Influence of quorum sensing and iron on twitching motility and biofilm formation in Pseudomonas aeruginosa. J. Bacteriol. 190, 662–671 (2008).
Singh, P. K. Iron sequestration by human lactoferrin stimulates P. aeruginosa surface motility and blocks biofilm formation. Biometals 17, 267–270 (2004).
Ochsner, U. A. & Reiser, J. Autoinducer-mediated regulation of rhamnolipid biosurfactant synthesis in Pseudomonas aeruginosa. PNAS 92, 6424–6428 (1995).
Rybtke, M. T. et al. Fluorescence-based reporter for gauging cyclic di-GMP levels in Pseudomonas aeruginosa. Appl Environ. Microbiol 78, 5060–5069 (2012).
El Mouali, Y. et al. Stand-alone EAL domain proteins form a distinct subclass of EAL proteins involved in regulation of cell motility and biofilm formation in enterobacteria. J. Bacteriol. https://doi.org/10.1128/jb.00179-17 (2017).
Trampari, E. et al. Bacterial rotary export ATPases are allosterically regulated by the nucleotide second messenger cyclic-di-GMP. J. Biol. Chem. 290, 24470–24483 (2015).
Caiazza, N. C., Merritt, J. H., Brothers, K. M. & O’Toole, G. A. Inverse regulation of biofilm formation and swarming motility by Pseudomonas aeruginosa PA14. J. Bacteriol. 189, 3603–3612 (2007).
Jones, S. et al. The lux autoinducer regulates the production of exoenzyme virulence determinants in Erwinia carotovora and Pseudomonas aeruginosa. Embo j. 12, 2477–2482 (1993).
Lin Chua, S. et al. Reduced intracellular c-di-GMP content increases expression of quorum sensing-regulated genes in Pseudomonas aeruginosa. Front. Cell. Infect. Microbiol. 7, 451–451 (2017).
Jenal, U., Reinders, A. & Lori, C. Cyclic di-GMP: second messenger extraordinaire. Nat. Rev. Microbiol 15, 271–284 (2017).
Chou, S. H. & Galperin, M. Y. Diversity of Cyclic Di-GMP-Binding Proteins and Mechanisms. J. Bacteriol. 198, 32–46 (2016).
Huang, B., Whitchurch, C. B. & Mattick, J. S. FimX, a multidomain protein connecting environmental signals to twitching motility in Pseudomonas aeruginosa. J. Bacteriol. 185, 7068–7076 (2003).
Frangipani, E. et al. The Gac/Rsm and cyclic-di-GMP signalling networks coordinately regulate iron uptake in Pseudomonas aeruginosa. Environ. Microbiol. 16, 676–688 (2014).
Jones, C. J. et al. ChIP-seq and RNA-seq reveal an AmrZ-mediated mechanism for cyclic di-GMP synthesis and biofilm development by Pseudomonas aeruginosa. PLOS Pathog. 10, e1003984 (2014).
Hamood, A. N. & Iglewski, B. H. Expression of the Pseudomonas aeruginosa toxA positive regulatory gene (regA) in Escherichia coli. J. Bacteriol. 172, 589–594 (1990).
Walker, S. L., Hiremath, L. S. & Galloway, D. R. ToxR (RegA) activates Escherichia coli RNA polymerase to initiate transcription of Pseudomonas aeruginosa toxA. Gene 154, 15–21 (1995).
Steiner, S., Lori, C., Boehm, A. & Jenal, U. Allosteric activation of exopolysaccharide synthesis through cyclic di-GMP-stimulated protein-protein interaction. Embo j. 32, 354–368 (2013).
Munsch, P., Geoffroy, V. A., Alatossava, T. & Meyer, J.-M. Application of siderotyping for characterization of Pseudomonas tolaasii and & “Pseudomonas reactans”; isolates associated with brown blotch disease of cultivated mushrooms. Appl Environ. Micro. 66, 4834–4841 (2000).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
McKenna, A. et al. The genome analysis toolkit: a mapreduce framework for analyzing next-generation DNA sequencing data. Genome Res 20, 1297–1303 (2010).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Malone, J. G. et al. The YfiBNR signal transduction mechanism reveals novel targets for the evolution of persistent Pseudomonas aeruginosa in Cystic Fibrosis airways. PLOS Pathog. 8, e1002760 (2012).
Diggle, S. P. et al. The Pseudomonas aeruginosa quinolone signal molecule overcomes the cell density-dependency of the quorum sensing hierarchy, regulates rhl-dependent genes at the onset of stationary phase and can be produced in the absence of LasR. Mol. Microbiol. 50, 29–43 (2003).
Pearson, J. P., Pesci, E. C. & Iglewski, B. H. Roles of Pseudomonas aeruginosa las and rhl quorum-sensing systems in control of elastase and rhamnolipid biosynthesis genes. J. Bacteriol. 179, 5756–5767 (1997).
Barraud, N. et al. Nitric oxide signaling in Pseudomonas aeruginosa biofilms mediates phosphodiesterase activity, decreased cyclic di-GMP levels, and enhanced dispersal. J. Bacteriol. 191, 7333–7342 (2009).
Hook, A. L. et al. Simultaneous tracking of Pseudomonas aeruginosa motility in liquid and at the solid-liquid interface reveals differential roles for the flagellar stators. mSystems 4, e00390–00319 (2019).
Heydorn, A. et al. Quantification of biofilm structures by the novel computer program comstat. Microbiol. (Read., Engl.) 146, 2395–2407 (2000).
Acknowledgements
This research was supported by the European commission (Grant FP6-2004-Mobility-2, contract number MEST-CT-2005-020278, Grant FP7-PEOPLE-2007-IEF, contract number PIEF-GA-2008-219200; Grant NABATIVI, EU-FP7-HEALTH-2007-B, contract number 223670), Wellcome Trust (Grant no 103884), and the National Biofilms Innovation Centre (NBIC) which is an Innovation and Knowledge Centre funded by the Biotechnology and Biological Sciences Research Council, InnovateUK and Hartree Centre [Award Number BB/R012415/1]. K-G Chan thanks University of Malaya for financial support (PPP grants: RMF1252-2021 and HIR grant: H50001-A-000027). Work in the J.G.M. lab was supported by ISP grant BB/J004553/1 (Biotic Interactions) to the John Innes Centre. We thank Alex Truman (University of Nottingham) for C4-HSL synthesis and Nigel Halliday (University of Nottingham) for C4-HSL and rhamnolipid quantification.
Author information
Authors and Affiliations
Contributions
M.C., S.H., P.W. and J.G.M. conceived the project. A.M.-P., J.-F.D. and M.R. designed and conducted the experiments. M.M. constructed and screened the genomic library. E.T. and N.B. conducted the SPR and L.C.-M.S. analysis, respectively. K.-G.C. and HNvG contributed to genome sequencing and in silico analysis of PAO1 genomes, respectively. A.M.C. acquired and analysed the bacterial single-cell tracking data. J.L. and Y.C. contributed to reporters and mutants construction and phenotypic testing. M.R., J.-F.D. and A.M.-P. wrote the manuscript with input from all other authors. S.R. contributed to manuscript reviewing/discussion. J.-F.D. and M.R. are considered “co-first author”.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dubern, JF., Romero, M., Mai-Prochnow, A. et al. ToxR is a c-di-GMP binding protein that modulates surface-associated behaviour in Pseudomonas aeruginosa. npj Biofilms Microbiomes 8, 64 (2022). https://doi.org/10.1038/s41522-022-00325-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41522-022-00325-9
- Springer Nature Limited