Abstract
The biology of breast cancer response to neoadjuvant therapy is underrepresented in the literature and provides a window-of-opportunity to explore the genomic and microenvironment modulation of tumours exposed to therapy. Here, we characterised the mutational, gene expression, pathway enrichment and tumour-infiltrating lymphocytes (TILs) dynamics across different timepoints of 35 HER2-negative primary breast cancer patients receiving neoadjuvant eribulin therapy (SOLTI-1007 NEOERIBULIN-NCT01669252). Whole-exome data (N = 88 samples) generated mutational profiles and candidate neoantigens and were analysed along with RNA-Nanostring 545-gene expression (N = 96 samples) and stromal TILs (N = 105 samples). Tumour mutation burden varied across patients at baseline but not across the sampling timepoints for each patient. Mutational signatures were not always conserved across tumours. There was a trend towards higher odds of response and less hazard to relapse when the percentage of subclonal mutations was low, suggesting that more homogenous tumours might have better responses to neoadjuvant therapy. Few driver mutations (5.1%) generated putative neoantigens. Mutation and neoantigen load were positively correlated (R2 = 0.94, p = <0.001); neoantigen load was weakly correlated with stromal TILs (R2 = 0.16, p = 0.02). An enrichment in pathways linked to immune infiltration and reduced programmed cell death expression were seen after 12 weeks of eribulin in good responders. VEGF was downregulated over time in the good responder group and FABP5, an inductor of epithelial mesenchymal transition (EMT), was upregulated in cases that recurred (p < 0.05). Mutational heterogeneity, subclonal architecture and the improvement of immune microenvironment along with remodelling of hypoxia and EMT may influence the response to neoadjuvant treatment.
Similar content being viewed by others
Introduction
Breast cancer is the most commonly diagnosed cancer and the leading cause of female cancer death worldwide1. It represents a heterogeneous group of tumours with characteristic molecular features, prognosis and responses to available therapy2,3.
In the early stage breast cancer setting, treatment decisions are guided by clinical subtypes, namely hormone receptor (HR) positive (HR+/HER2−), human epidermal growth factor receptor 2 amplified (HER2+) and triple-negative breast cancer (TNBC). This general classification does not take into account the complex genomic landscape and breast cancer evolution during therapy administration and disease recurrence or progression4,5.
Currently, the biology of the neoadjuvant response to therapy is underrepresented in the literature. Large-scale genomic studies have mostly focused on the analysis of single primary breast cancers2,3,6,7,8,9, which do not provide information on cancers over time. The analysis of the gene expression landscape of tumours has been shown to correlate with response to cytotoxic therapies10,11,12, though very limited work has been done to characterise the genomic and transcriptomic changes across breast cancer patients receiving neoadjuvant therapy13. Therefore, the neoadjuvant setting in breast cancer provides a window-of-opportunity to explore the genomic and microenvironment modulation of tumours exposed to therapy over time5,14.
In this study, we temporally characterised the mutational, gene expression, pathway enrichment and tumour-infiltrating lymphocytes (TILs) dynamics across different timepoints over a 12-week period in HER2-negative primary breast cancers enrolled in the single-arm SOLTI-1007 NEOERIBULIN phase II clinical trial (NCT01669252). We show that the mutational and immune tumour microenvironment remodelling of HER2-negative primary breast cancers provides a path forward for gathering biological insights from primary breast cancers.
Results
A clinical cohort of HER2-negative breast cancer patients
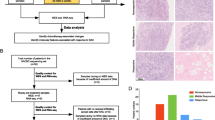
Primary breast cancer tumour specimens were obtained from the open-label, single-arm SOLTI-1007 NEOERIBULIN phase II clinical trial (NCT01669252). We included sequential primary tumour biopsies from 35 HER2-negative (22 HR-positive and 13 HR-negative) breast cancer patients (1–3 tumour tissue samples per patient) during eribulin administration. Whole-exome data (N = 88 samples) generated mutational profiles and candidate neoantigens and were analysed along with RNA-Nanostring 545-gene expression (N = 96 samples) and stromal TILs (N = 105 samples) from 35 patients with HER2-negative breast cancer (Fig. 1a).
a. Tumour tissue samples underwent (i) Whole-exome sequencing (WES) for mutation and clonality detection followed by neoantigen prediction; (ii) Nanostring gene expression profiling; and (iii) stromal TILs counting. Our goal was to select samples that passed quality control and perform the temporal characterisation of the mutational, gene expression and TILs in serial primary HER2-negative breast cancers that were good responders or poor responders to eribulin. DNA sequencing (WES) was performed in 88 primary invasive breast cancers and matched the normal DNA of each patient. Of these, 66 tumour samples were used for mutational and clonality analyses. RNA-Nanostring gene expression profiling was performed in 96 primary invasive breast cancers. From the DNA sequencing data, candidate neoantigens were predicted. Stromal TILs were counted from the H&E slides in 91 out of 105 tumour specimens. Clinical features and the PAM50 intrinsic molecular subtypes of each of the sequential primary tumour’s biopsies were examined. TMB tumour mutation burden, ORR overall response rate. b Schematics of the clinical trial. Temporal tumour sampling and a number of samples included in each analysis and time point are depicted. Distribution of PAM50 molecular intrinsic breast cancer subtypes at V1 (diagnostic biopsy), and ORR at V3 (surgery). E eribulin administration, V1 visit one, V2 visit two, V3 visit three, VR visit recurrence, CR complete response, PR partial response, SD stable disease, PD progressive disease.
Eighty per cent of cases had Ki67 greater than 14% at diagnosis. Disease recurrence after neoadjuvant therapy with eribulin was observed in six patients, despite the use of anthracyclines as adjuvant therapy. Although six patients presented clinic-radiologic recurrence, tumour material was available at the time of recurrence in one patient. The samples that passed quality control (i.e., tumour cellularity, sequencing quality, see Fig. 1a and “Methods”), were further processed and analysed.
Clinical features of the cohort are summarised in Table 1 and the schematics of the study design are shown in Fig. 1b. The 5-year relapse-free survival was 85.6% (95% CI: 74.7–98.1%) after breast cancer was diagnosed and the overall survival rate at 5 years was 91.3% (95% CI: 82.4–100%) for the patients analysed here.
The mutational and TILs profiling across temporal primary breast cancers
We characterised the mutational landscape of the tumours through an analysis of tumour mutational burden (TMB), breast cancer driver identification15, mutational signatures16 and mutational clonality17 at baseline (V1) (28 cases) and correlated these at visit 2 (V2) (24 cases) and with the response after 12 weeks of therapy at the time of surgery (V3) (13 cases).
TMB varied among patients at baseline (V1) (median 1.75 mutations per megabase (Mb) [range: 0.59–8.19], with a standard deviation of 1.48) but there was no evidence of variation across the three sampling time points (V2, median 1.62 mutations/Mb (range: 0.13–8.19), p = 0.62; V3, median 1.73 mutations/Mb (range: 0.38–7.11), p = 0.81) (mean coverage was ~40×) (Fig. 2a). At least one breast cancer driver gene18,19 was detected in 28 (80%) patients further analysed. TP53 (n = 15) and PIK3CA (n = 11) were the most prevalent mutated driver genes in the cohort (Supplementary Fig. 1, Supplementary Data). Mutations within these breast cancer drivers were detected at all time points as well as in the recurrent sample (case N021). In V1, TP53 mutation was more prevalent in basal cancers (N = 9), as compared to Luminal-A (N = 4) and Luminal-B (N = 2). Relapse free survival was not statistically significant when samples bearing TP53 mutations in V1 (N = 15) were compared to those without the mutation (N = 20) [HR = 2.07, 95% CI: 0.35–12.4, p value = 0.43]. Likewise, no difference was found in patients with PIK3CA mutation (n = 11) [HR = 1.46, 95% CI: 0.24–8.75, p value = 0.68] (Supplementary Fig. 1). However, this cohort was not adequately powered to detect an association between mutations in driver genes and clinical outcomes.
a Landscape of mutational alterations over time. Stacked plots display mutational burden (top), breast cancer drivers (tile plots, middle), PAM50 molecular intrinsic subtypes, clinical–pathological responses per patient and purity (tile plots, middle), mutation signatures (filled histogram). b Mutation clonality and subclonal distribution across different responses to eribulin. Left panels: odds for complete or partial response; right panels: a relative hazard for relapse-free survival (RFS). The analysis was performed for all comers (top panels) and for HR-positive and HR-negative patients (bottom panels). c Distribution of selected driver mutations generating neoantigens. Driver gene mutations are coloured whether the mutation is clonal or subclonal. d TILs across PAM50 intrinsic subtypes. (*) refer to p value < 0.05; ns nonsignificant. For each box plot, the centre line, the boundaries of the box, the ends of the whiskers and points beyond the whiskers represent the median value, the interquartile range, the minimum and maximum values, and the outliers, respectively. e Relationship between predicted neoantigen load (y-axis) and nonsynonymous mutational load (x-axis) and between predicted neoantigen load (y-axis) and mean stromal TILs per patient (x-axis).
We next investigated signatures of mutagenic biological processes to determine whether there were any detectable shifts during neoadjuvant therapy. A non-negative matrix factorisation technique to identify mutagenic processes in breast cancer including ageing, APOBEC cytidine deaminases, defective DNA repair, BRCA1/BRCA2 deficiency was used16. Mutational signatures were studied in each tumour sample across neoadjuvant therapy (Fig. 2a—bottom panel). Signature 1 (C > T transitions at CpG dinucleotides) contributed to a relatively higher proportion of mutations, but others were predominant in some patients, such as APOBEC (signatures 2 and 13), were more prevalent in luminal B patients, and BRCA-related (signature 3) in basal-like tumours. Signatures were not always conserved across tumour samples (defective DNA repair signature was conserved in 62.5% of patients (n = 8) while APOBEC and BRCA-related in 40% (n = 7 and n = 10, respectively). No statistical differences were observed between mutation signatures and overall response rate (ORR) (all p values > 0.05, Fisher’s test) (Supplementary Fig. 1), although the study was not initially powered to detect this association.
We computed tumour clonal composition and CCFs by applying the ABSOLUTE17 algorithm across temporal primary tumours from each patient to determine how clonal composition relates to outcomes. Mutations predicted to be clonal (found in every cancer cell) or subclonal (found in a subset of cancer cells) were identified: the proportion of clonal mutations was 44.7% on the first visit, 21.0% on the second visit, 28.9% at surgery and 5% at recurrence).
We studied the shape of the association of subclonal mutations at baseline (V1) and the patientsʼ odds to obtain a complete or partial response (PR) and also the hazard to relapse (Fig. 2b, see “Methods”). Overall, there was a trend towards higher odds of response and less hazard to relapse when the percentage of subclonal mutations was low (i.e., from 0 to 40%), suggesting that less heterogeneous tumours might have better responses to neoadjuvant therapy. Further increases above 40% in the percentage of subclonal did not seem to impact outcomes. The same analysis was carried out for HR-positive and HR-negative patients (Fig. 2b, bottom panels). Overall, HR-positive patients had a trend to have better outcomes.
We generated HLA-class I binding predictions for somatic mutations that may result in protein changes. Totally, 492 out of 2009 (24.50%) unique mutations were predicted to encode for at least one strong binder (binding rank < 0.5%) of HLA-class I. We analysed candidate neoantigens generated from nonsynonymous mutations. A total of 1002 putative neoantigens (average of 36 per patient, [range: 2–153]) were present in 27 patients (Supplementary Data).
We inferred candidate neoantigens derived from breast cancer driver mutations (5.19% of total neoantigens) and whether they were clonal or subclonal. Overall, 52 neoantigens (average of 3.11 per patient [range: 1–11]) were generated by 22 (unique) driver mutations in 17 patients. Driver mutations including TP53 (p.R110P, p.R209Kfs*6), PIK3CA (p.H1047L, p.K111E), MAP3K1 (p.R961Sfs*44, p.E504Vfs*36) predicted neoantigens in more than one patient, and in general, they were clonal and present across all samples of each patient (Fig. 2c). Each candidate neoantigen was predicted to be bound to one or more HLA class-I molecule.
Finally, stromal TILs were counted in 91 samples of 34 patients and had a median of 9.3% [range: 0–40] among patients. Among the PAM50 intrinsic subtypes, basal-like tumours had significantly more TILs than luminal A (p value = 0.02) but not luminal B (p value = 0.11) (Fig. 2d, Supplementary Fig. 1).
We observed that the higher nonsynonymous mutation load was strongly correlated with neoantigen load and the latter was weakly correlated with stromal TILs. A significant positive correlation was observed in both analyses (R2 = 0.94, p = <0.001 and R2 = 0.16, p = 0.02), respectively (Fig. 2e).
Therefore, although TMB and driver mutations were temporally conserved across neoadjuvant therapy, cases with a lower percentage of subclonal mutations, thus more homogeneous breast cancers, had a trend for better responses. TILs were weakly correlated with neoantigen load, and a small fraction of neoantigens were derived from driver gene mutations.
Multiple expressed genes across temporal primary breast cancers
To identify differentially expressed gene candidates during the administration of neoadjuvant chemotherapy, we performed targeted gene expression profiling using a custom 545-gene Nanostring panel composed of breast cancer-related genes among 35 patients (Supplementary Data).
We observed a total of 155 genes differentially expressed in patients responding to eribulin at surgery (good responders, CR or PR [N = 19]) vs. those with poor responses (PD or SD [N = 16]). After a false discovery rate (FDR) adjustment p value < 0.05, ALDH1A1, EVI2A, ADRA2A, LHFP, LOC400043 and PTGER4, were upregulated while VEGFA, FANCA, ORC6, KIFC1 and ANGPTL4 were downregulated in the good responder group (Fig. 3a, Supplementary Data).
a ORR at surgery (V3) (patients with good response to eribulin vs. poor response) (Top). Genes identified have a change corresponding to eribulin treatment. Volcano plots with the strength of the association on the y-axis (−log10 p values) and the effect size on the x-axis (log 2-fold change (FC)). Differentially expressed genes were highlighted during different timepoints across neoadjuvant therapy (V1–V3). Genes above the red dotted line represent those whose expression levels were significantly different (p value < 0.01). A full list of the most up and downregulated genes can be found on Supplementary Data. Boxplots of good and poor responders on eribulin over time, colour-coded by their corresponding poor response (red) or good response (blue) in eribulin from baseline to surgery (Bottom). b Same as in (a). Patients with later clinical recurrence vs. no recurrence (Top). Boxplots of individual patients on eribulin over time, colour-coded by their corresponding recurrence (purple) or nonrecurrence (grey) in eribulin from baseline to surgery (Bottom).
To assess variability within and between individuals, gene expression levels were analysed over a 12-week period for each patient (Fig. 3a bottom panels, Supplementary Fig. 2). Consistent with the general trend in the previous analysis, increased levels of ALDH1A1, EVI2A, ADRA2A, LHFP, LOC400043 and PTGER4 were modulated over time in patients responding to eribulin.
Overall and across all time points, we analysed patients that developed disease recurrence after curative therapy (n = 6) vs. those that did not recur (n = 29). We observed 58 upregulated genes in the group of patients with clinical recurrence as compared to those who did not have a recurrence. Among those were FABP5, YBX1, TUBB6, PLOD1 and CXCL8 genes that were upregulated and analysed over 12 weeks during neoadjuvant therapy.
Our analyses evidenced fatty acid-binding protein 5 (FABP5) overexpression at surgery (V3) was the only statically significant after FDR adjustment p < 0.05 in cases that recurred, as compared to those that did not recur (FDR = 0.04) (Fig. 3b, Supplementary Fig. 2). FABP5 is an inductor of the epithelial-mesenchymal transition (EMT) transition process20. EMT has been reported to contribute to tumour aggressiveness and eribulin resistance21,22.
Finally, we used pathway enrichment analysis to identify the activity of the immune system and common cancer signalling pathways before and after neoadjuvant therapy (see “Methods”, Table 2). At the baseline V1 timepoint, it was observed an increase in pathways associated with angiogenesis, hypoxia and epithelial features, in patients that had poor responses to therapy (SD/PD). In the good responder group, eribulin treatment was associated with statistically significant enrichment of pathways associated to lymphocyte and T cell activation in the end of therapy (V3 or surgery), and with reduction in expression of the immunosuppressive programmed cell death pathway. In contrast, there was an increase in cell proliferation and reduction in cytotoxic T cell pathways for the poor responder group at V3 or surgery timepoint. This suggests that eribulin apparent reversal of EMT and restoration of hypoxia might lead to improvement of immunosuppressive features in the primary tumours of good responders, and an increase of proliferation and reduction of immune cytotoxic activity in poor responders.
Discussion
Our study describes a step forward in gathering insights related to response to neoadjuvant therapy for HER2-negative primary breast cancers. Whole-exome sequencing, gene expression, pathway enrichment and TILs analyses of temporal primary breast tumours of 35 HER2-negative breast cancer patients show cancer remodelling during neoadjuvant chemotherapy.
The strengths of our analysis include the prospective design of the study, and the orthogonal genomic and immune infiltration data for each primary breast cancer systematically obtained from the SOLTI-1007 NEOERIBULIN phase II clinical trial (NCT01669252). Clonal driver mutations were maintained over time, but intratumoral genomic heterogeneity, measured as a fraction of subclonal mutations may affect response to therapy. Heterogeneity in mutational signatures across patients was considerably greater than at the intrapatient level, which was, in general, not conserved across temporal samples of each patient in 12 weeks of therapy administration. It should be noted, however, that eribulin is a weak chemotherapeutic agent which might explain the minimal change within the genomic landscape21. Its efficacy might be related to the fact that that it might change tumour phenotype (based on the EMT hypothesis)21 rather than changing tumour genotypes.
Neoantigens have been predicted in a few primary breast cancer datasets23,24. There is usually a discrepancy of neoantigens predicted computationally and those actually shown to leverage robust T-cell responses25,26. An immunosuppressive tumour microenvironment might be associated with this fact. A recent analysis of The Cancer Genome Atlas (TCGA) dataset of invasive breast cancers, predicted HLA class I-binding neoepitopes for 870 breast cancer samples24. About 40% of the nonsynonymous mutations led to the generation of candidate neoepitopes and the neoepitope load was also highly correlated with the mutational burden. Here, 21% of the nonsynonymous mutations led to the generation of putative neoantigens. Candidate neoantigens and stromal TILs were positively correlated and were more frequent in basal-like primary tumours. Driver mutations such as frameshifts mutations in TP53 and MAP3K1 genes or missense mutations in the PIK3CA gene can generate tumour neoantigens, suggesting that a T cell-mediated immune response, if present and properly validated, would target all cancer cells15,27,28. The relevance of neoantigens derived from clonal driver mutations, particularly those arising from truncating mutations, is the possibility of being incorporated as targets for adoptive T cell therapies and cancer vaccines28,29.
The evaluation of TILs has been shown to represent a reliable surrogate of the immune anti-tumour activity and a robust independent prognostic biomarker in breast cancer patients, especially in the TNBC and HER2-positive breast cancer subtypes30,31. Retrospective reports analysed TILs as prognostic and predictive markers in metastatic TNBC patients that received eribulin32. In a previous analysis, the high TILs group vs low TILs within the metastatic TNBC cases had better outcomes and suggested TILs to predict treatment response to eribulin in the TNBC metastatic setting32. In our early stage breast cancer cohort, stromal TILs detected in H&E slides were more frequent in basal-like primary tumours. A statistically significant higher percentage of stromal TILs after the end of neoadjuvant therapy was not observed possibly due to the small sample size and other biological factors including immunosuppressive TME. However, we observed an enrichment in pathways associated with immune infiltration and reduced programmed cell death after 12 weeks of eribulin administration corroborating findings observed in metastatic breast cancer. In a recent study33, lymphocyte infiltration increase and PDL1 expression turning to negative values were suggested to be improvements in the immunosuppressed TME of patients receiving eribulin for advanced breast cancer. Our findings add to current evidence for potentially combining checkpoint inhibitors to eribulin in the early breast cancer setting34.
There is the emerging rationale for eribulin to activate the immune system in pre-clinical and retrospective studies, through EMT suppression, and vascular remodelling and improvement of the tumour immune microenvironment21. Our analyses indicate that temporal gene expression responses to eribulin might be associated with the process of restoration of hypoxia, and the EMT process, which should be contextualised with the immune TME. Eribulin treatment has shown to suppress genes that are known to be involved in hypoxic signalling cascades, including VEGF, contributing to TME remodelling and restoring the scenario of normoxia35,36. We observed that VEGF was downregulated overtime during eribulin therapy in the good responder group. At diagnosis, poor responders presented an overexpression of angiopoietin-like 4 (ANGPTL4), a pro-angiogenic factor that is modulated by hypoxia and associated with poor prognosis, metastasis, cell differentiation and vascular permeability37. Previously, aldehyde dehydrogenase 1A1 (ALDH1A1) was reported as a cancer stem cell marker and suggested to have a favourable prognostic role for cervical cancer38,39. Although we observed ALDH1A to be upregulated in good responders, its role in breast cancer is undefined yet. Furthermore, FABP5 promotes tumour cell growth in numerous cell types and is a negative prognostic marker in renal cell carcinoma20,40. FABP5 was upregulated in cases that presented further clinic-radiological recurrence. However, identification of FABP5 as a biomarker in HER2-negative breast cancer requires substantially increased cohort size and mechanistic validation for robust interpretation.
Our work has limitations which are mostly secondary to the small sample size and therefore lack of power to detect specific associations. Nonetheless, this HER2-negative breast cancer cohort gives a glimpse of the changes in cancer biology during neoadjuvant therapy administration. Another limitation is the 545 targeted gene expression panel used did contemplate only a few immune markers including IDO1, LAG3 and STAT1. Furthermore, the primary endpoint of the present clinical trial is pathologic complete response (pCR) in the breast. We used ORR for analyses of responses after the neoadjuvant therapy since there are only a few pCR cases; residual cancer burden was used but was not informative in the dataset; thus, the association between biomarkers and response is challenging.
Taken together, these results suggest that mutational heterogeneity, subclonal architecture and the immune microenvironment along with remodelling of hypoxia and EMT may influence the response to neoadjuvant treatment in early stage disease, with possible implications for clinical decision-making and monitoring of treatment efficacy.
Methods
Patients and tissues
Primary breast cancer tumour specimens were obtained from the phase II study, open-label, single-arm SOLTI-1007 NEOERIBULIN (NCT01669252). The patients reported here were treated at the Vall Hebron Institute of Oncology, Barcelona. In brief, eribulin was administered as neoadjuvant treatment for stage I-II HER2-negative breast cancer with a dose of 1.4 mg/m2 intravenously on Days 1 and 8 every 21-day cycle, for 4 cycles, and the pathological CR was defined as the absence of residual invasive tumour in breast tumour specimens. All patients were negative for HER2 overexpression on clinical assays. The Ethics Committee of the Vall Hebron University Hospital, Barcelona, Spain approved the study. All patients gave informed consent for DNA and RNA sequencing.
Clinic-pathological response analyses
We adopted a conservative method of defining ORR after neoadjuvant therapy as CR or PR by response evaluation criteria in solid tumours (RECIST) (major decrease in tumour burden following treatment) and poor response as progressive disease (PD) or stable disease (SD) by RECIST (major increase or stability in tumour burden following treatment) after eribulin therapy.
Patients were classified into two groups depending on their clinical response to therapy at surgery: good responders—patients that had CR or PR; poor responders—patients that had PD or SD.
Tissue processing and DNA and RNA extraction
Frozen biopsies were embedded in a frozen tissue matrix (OCT; Sakura Finetek, Torrance, CA) and cut at the cryostat for tumour cellularity assessment by a pathologist. Genomic DNA was isolated from 10 × 10um sections using the DNeasy Kit (Qiagen, Hilden, Germany). Samples with less than 10% tumour material or that produced low DNA yield were excluded from the analysis.
A section of the FFPE breast tissue was first examined with haematoxylin and eosin staining to confirm the presence of invasive tumour cells (≥10%).
For RNA purification (High Pure Formalin-Fixed Paraffin-embedded RNA Isolation Kit, Roche Diagnostics Limited, West Sussex, UK), one to five 10 μm FFPE slides were used for each tumour specimen (at diagnosis, cycle 2 and surgery).
Gene expression and Intrinsic subtype analyses
A minimum of ∼125 ng of total RNA was used to measure the expression of 545 genes involved in breast cancer, including 5 housekeeping genes (ACTB, MRPL19, PSMC4, RPLP0 and SF3A1), using the Prosigna assay (NanoString Technologies, Seattle, USA). Samples with 20 or fewer counts in at least 70% of the genes were removed. Data were log base 2 transformed and normalised using the housekeeping genes.
The same RNA was used to measure the expression of 50 genes of the PAM50 intrinsic subtype predictor assay. For each sample, we calculated the PAM50 signature scores (Basal-like, HER2-E, Luminal A and B) and the proliferation signature score41. Differential gene expression analysis fold change of each gene was calculated as the ratio of average gene expression intensity of (i) the good responder group (n = 19 [5 CR, 14 PR]) to that of the poor responder group (n = 16 [12 SD, 4 PD]) or (ii) the patients that recurred (n = 6) to that of the non-recurrent group (n = 29). A two-sample t test was used to compare gene expression intensities between groups.
A gene was claimed to be differentially expressed if it showed a fold change of >1 (increased in good responders, or non-recurrent) or ≤ −1 (increased in poor responders, or in the later recurred) and further adjustment FDR ≤ 0.05 was applied. Volcano plots were used to visualise log 2-fold change on the x-axis and −log10 p values on the y-axis.
Pathway enrichment analysis was performed using the gprofiler toolkit42 comparing good responder vs. poor responder groups using two ontology databases as reference: (i) Gene Ontology and (ii) Kyoto Encyclopaedia of Genes and Genomes.
DNA sequencing
WES was performed to the breast cancer tumour and matched normal DNA obtained from the buffy coat of each HER2-negative breast cancer patient.
Libraries for Illumina sequencing were prepared using Illumina Nextera Rapid Capture Exome kit (cat. FC-140-1003, Illumina) as we reported previously. Prior to library preparation DNA concentrations for each sample were quantified using a fluorescence-based method (Quant-IT dsDNA BR, cat. Q33130, Thermo Fisher Scientific) and 50 ng of genomic DNA was used for library preparation.
Samples were processed following the manufacturer’s instructions (part# 15037436 Rev. J, Illumina) for WES. Prior to the first hybridisation, all libraries were quantified using quantitative polymerase chain reaction (qPCR). KAPA Library Quantification Kit (cat. KK4873, KAPA Biosystems) as used as per manufacturer’s recommendations. A subset of libraries was analysed using DNA 1000 Kit (cat. 5067-1504, Agilent).
Whole-genome libraries and exome libraries were normalised and pooled in equal volumes to create balanced pools. Each pool was normalised to molarity of 4 nM and used for sequencing with clustering concentration 20 pM with 1% spike-in of PhiX control. Sequencing was performed on an Illumina HiSeq2500 using v4 chemistry and 50 cycles single-end for s-WGS and 75 cycles paired-end for WES. Demultiplexing was performed using Illumina’s bcl2fastq2 v.2.17 software using default options. FASTQ files were used for subsequent data analysis.
WES analyses
For WES analysis, reads were mapped to the human genome (GRCh37) and base quality recalibration were performed using Novoalign v 3.02 (Novocraft). Coordinate sorting of reads and PCR-duplicate marking was performed using Novosort (v 3.02). The resulting bam files for all samples for the same patient were locally realigned using the Genome Analysis Toolkit (GATK, v 3.4.46)43. MuTect (version 2) was run using default parameters44. Strelka (version 1.0.14) was run with recommended starting parameters for BWA and default parameters45. The mean coverage per sample was calculated with CollectWgsMetrics (Picard). A joint calling was done and filtered allelic fraction >10% in at least one sample of each patient. Somatic mutations were annotated using Variant Effect Predictor (VEP, http://grch37.ensembl.org/) and visualised using IGV. Output with reads > 15 and purity > 20% were included in further analyses.
ABSOLUTE
ABSOLUTE (v1.0.6) was used to infer the cancer cell fraction (CCF) of mutations and the mutations were classified as clonal or subclonal as previously described17,46. A mutation was classified as clonal if its probability of being clonal was >50% or if the lower bound of the 95% confidence interval of its CCF was >90%. Mutations that did not meet the above criteria were considered subclonal.
ABSOLUTE software was used to calculate tumour clonality, purity and ploidy. For running the ABSOLUTE we obtained the mutation annotation file by running vcf2maf script with VCF files for the corresponding tumour and normal control samples, annotation was performed by VEP. To find the segmentation we ran the CNVkit batch command with the.bam files from tumour and normal samples47. For running ABSOLUTE we subset the.cns files from the CNVkit output file for variant coordinates as well as probes and log2 values. We used the options min.ploidy = 0.95, max.ploidy = 10. Probability of a mutation to be clonal was defined essentially as described in ref. 48.
Mutation signature
Decomposition of the mutational signature was performed using deconstructSigs16, based on the set of 30 mutational signatures. These signatures are largely defined by the relative frequency of the six possible base substitutions (C > A, C > G, C > T, T > A, T > C, and T > G) in the sequence context of their adjacent 5′ and 3′ base. Of these, COSMIC signatures 1, 2, 3, 6, 13, 9, 10, 15, have been associated with breast cancer. For clarity, we explored the following mutational signatures in this cohort: ageing/clock-like signatures49,50 (signatures 1 and 5), APOBEC (signatures 2 and 13), BRCA-related (signature 3), defective DNA repair (signatures 6 and 15), polymerase n (signature 9), POLE mutations (signature 10), unknown breast (signature 30) and other signatures.
Neoantigen prediction
The 4-digit HLA type was obtained from matched normal WES data and the OptiType 1.3.1 analysis software package was used51. WES and HLA typing was integrated, and NeoPredPipe pipeline52 was applied with minor modifications for the neoantigen prediction. First, non-synonymous cancer-specific mutations, i.e., present in all tumour cells and absent in all normal cells, were used to generate a comprehensive list of peptides (9–11 amino acids in length) with the mutated amino acid represented at each peptide position and used as input was previous described. Prioritisation of neoantigens: the selection of candidate neoantigens were based on a %Rank < 0.5; epitopes already existing in the reference proteome indicated by the Novelty parameter were excluded.
Tumour infiltrating lymphocytes
Evaluations of stromal TILs were performed on haematoxylin and eosin-stained sections by an experienced board-certified pathologist (R.F.) according to the 2014 recommendations of the international TILs working group53.
Statistical analysis
Continuous variables were expressed as median and range, while categorical variables were expressed as absolute values or percentages. For statistical comparison, we used Mann–Whitney test for independent continuous variables, Wilcoxon signed-rank test for paired continuous data and Fisher’s exact test for categorical data. To study if there was a significant variation of TMB across sampling timepoints the Wilcoxon signed-rank test was used to compare pairwise timepoints. In addition, the range and standard deviation of TMB in each time point were calculated to estimate intra-period TMB heterogeneity. To study the association between mutations in driver genes and relapse-free survival the Cox proportional hazard model was fitted. Furthermore, to analyse the association between the percentage of subclonal mutations at baseline and clinical outcomes, we used the logistic and Cox models after relaxed linearity assumption using restricted cubic splines by means of rms R package. Data analyses were carried out using R version statistical software 3.6.3.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data generated and analysed during this study are described in the following data record: https://doi.org/10.6084/m9.figshare.1445426154. The whole-exome sequencing bam files have been deposited at the European Genome-phenome Archive (EGA), which is hosted by the EBI and the CRG, under the study accession number https://identifiers.org/ega.study:EGAS00001004953 and dataset accession number https://identifiers.org/ega.dataset:EGAD0000100698055. The decreased and increased pathway enrichment analyses are available via GitHub at https://github.com/NeoVaCan/NPJBCANCER_DeMattos_2021/tree/main/Data_Supp_Table2. The supplementary tables are also available in Excel format as part of the figshare data record. The patient metadata and patient tumour-infiltrating lymphocytes data are not publicly available for the following reason: data contain information that could compromise research participant privacy. However, the data can be made available upon reasonable request to the corresponding author.
Code availability
Custom R code used to analyse tumour whole-exome sequencing data and Nanostring gene expression data are available at https://github.com/NeoVaCan/NPJBCANCER_DeMattos_2021.
References
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA. Cancer J. Clin. https://doi.org/10.3322/caac.21492 (2018).
Banerji, S. et al. Sequence analysis of mutations and translocations across breast cancer subtypes. Nature 486, 405–409 (2012).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature https://doi.org/10.1038/nature10983 (2012).
Rivenbark, A. G., O’Connor, S. M. & Coleman, W. B. Molecular and cellular heterogeneity in breast cancer: challenges for personalized medicine. Am. J. Pathol. https://doi.org/10.1016/j.ajpath.2013.08.002 (2013).
De Mattos-Arruda, L., Shen, R., Reis-Filho, J. S. & Cortés, J. Translating neoadjuvant therapy into survival benefits: one size does not fit all. Nat. Rev. Clin. Oncol. 13, 566–579 (2016).
Koboldt, D. C. et al. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Nik-Zainal, S. et al. Landscape of somatic mutations in 560 breast cancer whole-genome sequences. Nature https://doi.org/10.1038/nature17676 (2016).
Stephens, P. J. et al. The landscape of cancer genes and mutational processes in breast cancer. Nature 486, 400–404 (2012).
Yates, L. R. et al. Subclonal diversification of primary breast cancer revealed by multiregion sequencing. Nat. Med. 21, 751–759 (2015).
Fox, N. S., Haider, S., Harris, A. L. & Boutros, P. C. Landscape of transcriptomic interactions between breast cancer and its microenvironment. Nat. Commun. 10, 3116 (2019).
Paik, S. et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N. Engl. J. Med. 351, 2817–2826 (2004).
Wallden, B. et al. Development and verification of the PAM50-based Prosigna breast cancer gene signature assay. BMC Med. Genomics 8, 54 (2015).
Höglander, E. K. et al. Time series analysis of neoadjuvant chemotherapy and bevacizumab-treated breast carcinomas reveals a systemic shift in genomic aberrations. Genome Med. 10, 92 (2018).
Angelova, M. et al. Evolution of metastases in space and time under immune selection. Cell 175, 751–765.e16 (2018).
De Mattos-Arruda, L. et al. The genomic and immune landscapes of lethal metastatic breast cancer. Cell Rep. 27, 2690–2708.e10 (2019).
Alexandrov, L. B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
De Mattos-Arruda, L. et al. The genomic and immune landscapes of lethal metastatic breast cancer. Cell Rep. 27(2690-2708), e10 (2019).
Reiter, J. G. et al. Minimal functional driver gene heterogeneity among untreated metastases. Science 361, 1033–1037 (2018).
Ohata, T. et al. Fatty acid-binding protein 5 function in hepatocellular carcinoma through induction of epithelial-mesenchymal transition. Cancer Med. 6, 1049–1061 (2017).
Cortes, J., Schöffski, P. & Littlefield, B. A. Multiple modes of action of eribulin mesylate: emerging data and clinical implications. Cancer Treat. Rev. 70, 190–198 (2018).
Yoshida, T. et al. Eribulin mesilate suppresses experimental metastasis of breast cancer cells by reversing phenotype from epithelial–mesenchymal transition (EMT) to mesenchymal–epithelial transition (MET) states. Br. J. Cancer 110, 1497–1505 (2014).
Ren, Y. et al. HLA class-I and class-II restricted neoantigen loads predict overall survival in breast cancer. Oncoimmunology https://doi.org/10.1080/2162402X.2020.1744947 (2020).
Narang, P., Chen, M., Sharma, A. A., Anderson, K. S. & Wilson, M. A. The neoepitope landscape of breast cancer: implications for immunotherapy. BMC Cancer 19, 200 (2019).
Saini, S. K., Rekers, N. & Hadrup, S. R. Novel tools to assist neoepitope targeting in personalized cancer immunotherapy. Ann. Oncol. https://doi.org/10.1093/annonc/mdx544 (2017).
De Mattos-Arruda, L., Blanco-Heredia, J., Aguilar-Gurrieri, C., Carrillo, J. & Blanco, J. New emerging targets in cancer immunotherapy: the role of neoantigens. ESMO Open 4, e000684 (2020).
Turajlic, S. et al. Insertion-and-deletion-derived tumour-specific neoantigens and the immunogenic phenotype: a pan-cancer analysis. Lancet Oncol. 18, 1009–1021 (2017).
McGranahan, N. et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351, 1463–1469 (2016).
Robertson, J., Salm, M. & Dangl, M. Adoptive cell therapy with tumour-infiltrating lymphocytes: the emerging importance of clonal neoantigen targets for next-generation products in non-small cell lung cancer. Immuno-Oncol. Technol. 3, 1–7 (2019).
Pruneri, G., Vingiani, A. & Denkert, C. Tumor infiltrating lymphocytes in early breast cancer. Breast 37, 207–214 (2018).
Dushyanthen, S. et al. Relevance of tumor-infiltrating lymphocytes in breast cancer. BMC Med. 13, 202 (2015).
Kashiwagi, S. et al. Use of Tumor-infiltrating lymphocytes (TILs) to predict the treatment response to eribulin chemotherapy in breast cancer. PLoS ONE https://doi.org/10.1371/journal.pone.0170634 (2017).
Goto, W. et al. Eribulin promotes antitumor immune responses in patients with locally advanced or metastatic breast cancer. Anticancer Res. 38, 2929–2938 (2018).
Pizzuti, L. et al. Eribulin in triple negative metastatic breast cancer: critic interpretation of current evidence and projection for future scenarios. J. Cancer 10, 5903–5914 (2019).
Funahashi, Y. et al. Eribulin mesylate reduces tumor microenvironment abnormality by vascular remodeling in preclinical human breast cancer models. Cancer Sci. 105, 1334–1342 (2014).
Zhao, S. et al. Elimination of tumor hypoxia by eribulin demonstrated by 18F-FMISO hypoxia imaging in human tumor xenograft models. EJNMMI Res. https://doi.org/10.1186/s13550-019-0521-x (2019).
Carbone, C. et al. Angiopoietin-like proteins in angiogenesis, inflammation and cancer. Int. J. Mol. Sci. 19, 431 (2018).
Charafe-Jauffret, E. et al. Aldehyde dehydrogenase 1-positive cancer stem cells mediate metastasis and poor clinical outcome in inflammatory breast cancer. Clin. Cancer Res. 16, 45–55 (2010).
Vassalli, G. Aldehyde dehydrogenases: not just markers, but functional regulators of stem cells. Stem Cells Int. 2019, 1–15 (2019).
Lv, Q. et al. FABP5 regulates the proliferation of clear cell renal cell carcinoma cells via the PI3K/AKT signaling pathway. Int. J. Oncol. https://doi.org/10.3892/ijo.2019.4721 (2019).
Parker, J. S. et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J. Clin. Oncol. 27, 1160–1167 (2009).
Raudvere, U. et al. g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47, W191–W198 (2019).
Depristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. https://doi.org/10.1038/ng.806 (2011).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. https://doi.org/10.1038/nbt.2514 (2013).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor–normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
De Mattos-Arruda, L. et al. Genetic heterogeneity and actionable mutations in HER2-positive primary breast cancers and their brain metastases. Oncotarget 9, 20617–20630 (2018).
Talevich, E., Shain, A. H., Botton, T. & Bastian, B. C. CNVkit: genome-wide copy number detection and visualization from targeted DNA sequencing. PLOS Comput. Biol. 12, e1004873 (2016).
Miao, D. et al. Genomic correlates of response to immune checkpoint blockade in microsatellite-stable solid tumors. Nat. Genet. 50, 1271–1281 (2018).
Alexandrov, L. B. et al. Clock-like mutational processes in human somatic cells. Nat. Genet. 47, 1402–1407 (2015).
Kim, Y. A. et al. Network-based approaches elucidate differences within APOBEC and clock-like signatures in breast cancer. Genome Med. https://doi.org/10.1186/s13073-020-00745-2 (2020).
Szolek, A. et al. OptiType: precision HLA typing from next-generation sequencing data. Bioinformatics 30, 3310–3316 (2014).
Schenck, R. O., Lakatos, E., Gatenbee, C., Graham, T. A. & Anderson, A. R. A. NeoPredPipe: high-throughput neoantigen prediction and recognition potential pipeline. BMC Bioinform. 20, 264 (2019).
Salgado, R. et al. The evaluation of tumor-infiltrating lymphocytes (TILs) in breast cancer: recommendations by an International TILs Working Group 2014. Ann. Oncol. 26, 259–271 (2015).
De Mattos-Arruda, L. Metadata record for the article: The temporal mutational and immune tumour microenvironment remodelling of HER2-negative primary breast cancers. figshare https://doi.org/10.6084/m9.figshare.14454261 (2021).
European Genome-phenome Archive https://identifiers.org/ega.dataset:EGAD00001006980 (2021).
Acknowledgements
The authors acknowledge the patients who participated in this study, the Vall d´Hebron Institute of Oncology and the Cellex foundation. This work was supported by Cancer Research UK. L.D.M.A. was partly funded by Spanish Association against cancer.
Author information
Authors and Affiliations
Contributions
L.D.M.A wrote the paper with the assistance of co-authors. L.D.M.A., J.C., D.T. and C.C. contributed to the overall design of this study. Data were collected by V.O. and J.P.G. Data were analysed and interpreted by J.B.H., D.T., G.V., S.G.R., L.P., C.A.S., S.J.S., P.C., R.F. and S.F.C. All authors have read and approved the final version of the paper.
Corresponding author
Ethics declarations
Competing interests
L.D.M.A. has received honoraria for participation in a speaker’s bureau/consultancy from Roche, and reports research collaboration and support from NanoString Technologies. J.C. reports consulting from Roche, Celgene, Cellestia, AstraZeneca, Biothera Pharmaceutical, Merus, Seattle Genetics, Daiichi Sankyo, Erytech, Athenex, Polyphor, Lilly, Servier, Merck Sharp&Dohme, GSK, Leuko, Bioasis, Clovis Oncology, Boehringer Ingelheim, Honoraria: Roche, Novartis, Celgene, Eisai, Pfizer, Samsung Bioepis, Lilly, Merck Sharp&Dohme, Daiichi Sankyo. Research funding to the Institution: Roche, Ariad pharmaceuticals, AstraZeneca, Baxalta GMBH/Servier Affaires, Bayer healthcare, Eisai, F.Hoffman-La Roche, Guardanth health, Merck Sharp&Dohme, Pfizer, Piqur Therapeutics, Puma C., Queen Mary University of London. Stock, patents and intellectual property: MedSIR. Travel, accommodation, expenses: Roche, Novartis, Eisai, pfizer, Daiichi Sankyo. A.P. reports consulting fees from Nanostring Technologies Roche, Pfizer, Novartis and Daiichi Sankyo outside the submitted work. C.C. is a member of the AstraZeneca External Science Panel and has research grants from Roche, Genentech, AstraZeneca, and Servier that are administered by the University of Cambridge. The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
De Mattos-Arruda, L., Cortes, J., Blanco-Heredia, J. et al. The temporal mutational and immune tumour microenvironment remodelling of HER2-negative primary breast cancers. npj Breast Cancer 7, 73 (2021). https://doi.org/10.1038/s41523-021-00282-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41523-021-00282-0
- Springer Nature Limited