Abstract
The prevalence and contribution of BRCA1/2 (BRCA) pathogenic variants (PVs) to the cancer burden in Latin America are not well understood. This study aims to address this disparity. BRCA analyses were performed on prospectively enrolled Latin American Clinical Cancer Genomics Community Research Network participants via a combination of methods: a Hispanic Mutation Panel (HISPANEL) on MassARRAY; semiconductor sequencing; and copy number variant (CNV) detection. BRCA PV probability was calculated using BRCAPRO. Among 1,627 participants (95.2% with cancer), we detected 236 (14.5%) BRCA PVs; 160 BRCA1 (31% CNVs); 76 BRCA2 PV frequency varied by country: 26% Brazil, 9% Colombia, 13% Peru, and 17% Mexico. Recurrent PVs (seen ≥3 times), some region-specific, represented 42.8% (101/236) of PVs. There was no ClinVar entry for 14% (17/125) of unique PVs, and 57% (111/196) of unique VUS. The area under the ROC curve for BRCAPRO was 0.76. In summary, we implemented a low-cost BRCA testing strategy and documented a significant burden of non-ClinVar reported BRCA PVs among Latin Americans. There are recurrent, population-specific PVs and CNVs, and we note that the BRCAPRO mutation probability model performs adequately. This study helps address the gap in our understanding of BRCA-associated cancer in Latin America.
Similar content being viewed by others
Introduction
Breast cancer is the most frequently diagnosed cancer and the leading cause of cancer-related mortality in Latin American women1. High mortality rates in the region are driven in part by advanced stage at diagnosis and limited access to cancer care2. Also, the mean age at diagnosis is younger than US non-Hispanic white (NHW) populations and there is a high prevalence of hormone receptor and Her2 negative breast cancer (TNBC), features common in hereditary disease. Thus, strategies aimed at preventing, or detecting breast cancer at an earlier stage, are crucial within the region.
Hereditary breast and ovarian cancer (HBOC) syndrome accounts for approximately 10% of all breast cancer and 15–20% of ovarian cancer cases, and the most commonly associated genes with HBOC are BRCA1 and BRCA2 (BRCA)3. Women with likely pathogenic and pathogenic variants (PVs) in either of these genes, have up to 80% lifetime risk of breast cancer and 20–50% risk of ovarian cancer4. The identification of PVs in at-risk individuals has significant clinical utility, as there are effective interventions aimed at both cancer prevention and early diagnosis5. We previously reported a high rate of BRCA PVs among 110 United States (U.S.) Hispanics with early-onset breast cancer6. However, Hispanics are not well represented in most studies7,8. For example, just 3.4% of participants were Hispanic in a recent study of female PV carriers from 69 centers in 49 countries on 6 continents9.
BRCA PVs are present in around 1/800 people in the general population. However, a higher rate of PVs has been reported in some founder populations such as the Ashkenazi Jewish, with a PV prevalence of 1/4010. From published high risk clinic series, approximately 10% of women meeting National Comprehensive Cancer Network (NCCN) genetic testing criteria will carry a BRCA PV5,11. Some studies suggest that individuals with Latin American or African ancestry have a higher prevalence of PVs7 while others demonstrated a lower yield among Hispanic immigrants in the United States (U.S.)12. Our group reported a high prevalence of BRCA PVs (25%) in a large study of Hispanics living in the Southwestern U.S.13.
The frequency of BRCA PVs has been reported to be between 1.2 and 15.6% among Latin American cancer patients14,15,16,17, and 15–28% among young breast or ovarian cancer patients in Mexico, unselected for family history of breast cancer18,19. However, the prevalence and contribution of BRCA PVs to the overall cancer burden in the region are not yet well understood.
Women with HBOC are usually younger at the time of breast cancer diagnosis, have a family history of breast and/or ovarian cancer and more frequently present with TNBC. Each of these characteristics is an indication for genetic testing according to international guidelines20. In Latin America, the proportion of young women with breast cancer is almost twice that seen in more developed regions of the world, and there also is a greater proportion of TNBC, meaning that a significant number of Latin American patients would meet criteria for gene testing21,22. Unfortunately, despite this great need, access to genetic cancer risk assessment and testing is limited across the region and is not covered by public insurance23.
Because genetic testing is expensive, pretest probability models are useful for selecting women at a higher risk of carrying a BRCA PV. However, because probability models are based mainly on data from NHW populations, their performance among Hispanics is limited. Some studies have found that BRCAPRO accuracy might be inferior in US minorities because the prevalence of carriers may differ by ethnicity and race24. BRCAPRO performance was reported as greater than 80% in NHW, 75% in African Americans and 58–75% in small samples of US Hispanics25,26.
BRCA PVs are detected by complex genetic technologies including multiple mutation scanning methods and Sanger DNA sequencing. Assays for the detection of large genomic rearrangements, herein referred to as copy number variants (CNVs), are also expensive. CNVs represented 11% of BRCA PVs among US Hispanics13, of which 62% were the BRCA1 exon 9–12del (c.548-?_4185 + ?del), a Mexican founder PV, seen most frequently among patients from central Mexico, and estimated to have originated ~1,440 years ago13,21,22,27.
Commercial costs for these tests range between $249 and $5,000 U.S. dollars (USD), which is not affordable as an out-of-pocket expense for most of the Latin American population28. Disparities in access to testing have a significant impact on affected populations, due in part to underrepresentation in surveys of regional cancer etiology and in genomic variant databases9,29,30,31,32. To address disparities in genetic testing, we have developed and implemented a low-cost testing strategy for at-risk individuals, as part of a larger dissemination and implementation intervention33.
Here we report our results employing a sequential genetic testing strategy to detect BRCA PVs in a large sample of Latin Americans meeting referral criteria, using a combination of Sequenom MassARRAY technology, Ion Torrent semiconductor sequencing, and PCR-based methodologies for CNVs. We also report on the performance of the BRCAPRO mutation prediction model among our Latin American cohort.
Results
Accrual and study population characteristics
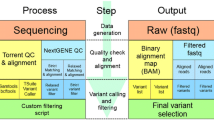
From December 2012 to August 2017, a total of 1627 participants were prospectively enrolled. Median age at enrollment was 44.6 years (range 14–96 years); and 1,607 (98.8%) were women. Of the 1627, 1375 (84.5%) had a primary breast cancer, 125 (7.7%) ovary cancer, and 49 (3.0%) other cancer types. Family history of breast cancer at ≤ 50 years in at least one first- or second-degree relative was reported in 412 (25.3%) participants, and of ovarian cancer at any age in 133 (8.2%) participants. Demographic characteristics are described in Table 1. The mean age at diagnosis of a first breast cancer in participants was 40.3 years (range 19–79); 15.0% had clinical stage 0-I and 85.0% had stage II-IV; 31.4% had TNBC (Table 2). A stepwise low-cost screening strategy (Fig. 1) was developed and implemented in our laboratory for the genotyping of a large Latin American cohort for mutations in the BRCA genes. Incremental genotyping costs by strategy was analyzed (Supplementary Table 1), and all strategies totaled approximately $100/case.
Frequency of BRCA variants
The overall frequency of BRCA PVs for the cohort was 14.5% (236/1627). The population with the highest frequency was Brazil (25.7%), followed by Mexico (17.4%), and Peru (12.6%), while the lowest frequencies were found in Colombia and Puerto Rico (9.3%).
There were 196 unique variants of unknown significance (VUS). The VUS rate by country varied significantly: 12.2% for Brazil, 9.8% for Colombia, 12.2% for Mexico, 17.8% for Peru and 9.3% for Puerto Rico (p = 0.006).
Of the 125 unique PVs (Supplementary Data 1), 17 (13.6%) were not represented in ClinVar, a freely accessible, public archive of reports of the relationships among human genetic variations and phenotypes, with supporting evidence (https://www.ncbi.nlm.nih.gov/clinvar/). Two of those 17 were recurrent. Of the 196 unique variants of unknown significance (VUS), 111 (56.6%) were not in ClinVar (Supplementary Data 1).
HISPANEL and ion torrent sequencing: sensitivity and performance
Primary screening with our custom designed recurrent Hispanic Mutation Panel (HISPANEL) comprised of 114 known PVs on the Sequenom MassARRAY platform, combined with a three-primer PCR assay (3-PA) for the Mexican founder PV (BRCA1 exon 9–12del), yielded 47.4% (112/236) of all positives (Table 3). The HISPANEL sensitivity was 47% overall; 57% (63/110) in Mexico; 57% (12/21) in Colombia; 53% (10/19) in Brazil; 50% (2/4) in Puerto Rico; and 30% (25/82) in Peru. BRCA sequencing yielded 41.9% (99/236) of all positives and other CNV methods yielded 11% (23/236).
Copy number variants
We found a total of 48 CNVs, all of them in BRCA1. No BRCA2 CNVs were detected. BRCA1 CNVs represented 20.3% (48/236) of all positives and 30% of BRCA1 PVs, with the Mexican founder PV, BRCA1 exon 9–12del (detected by investigator developed PCR assay27), accounting for 47.9% (23/48) of all CNVs. The remaining 52.1% of BRCA1 CNVs were detected by NGS and confirmed by multiplex ligation-dependent probe amplification (MLPA). In Mexico, 34.5% (38/110) of all PVs were CNVs, followed by 9.5% in Colombia, 8.5% in Peru, and 5.3% in Brazil. In Puerto Rico 1 of 4 PVs was a CNV. Within Mexico, CNVs represented 41.2% (7/17) of PVs in Guadalajara (including one BRCA1 exon 9–12del variant) and 33.3% (31/93) of PVs in Mexico City (including 22 BRCA1 exon 9–12del CNVs).
Recurrent PVs
Of the 236 PVs detected in this cohort, 101 (42.8%) were recurrent (observed 3 or more times) in the cohort (Table 4). The most frequent PV was the BRCA1 exon 9–12del (c.548-?_4185 + ?del) PV, observed 23 times in Mexico. The Ashkenazi Jewish founder PV, BRCA1 c.66_67delAG, was next (n = 13; 8 Peru, 5 Mexico), followed by BRCA1 c.5123 C > A (n = 11), c.815_824dup (n = 8; 5 Peru, 3 Mexico), and another Ashkenazi Jewish founder PV, c.5266dupC (n = 8; 6 Brazil, 2 Mexico). PVs found to be frequent in Colombia include: BRCA1 c.3331_3334del (n = 3; 2 Colombia, 1 Brazil) and c.5123 C > A (n = 11; 4 Colombia, 4 Mexico, 3 Peru). Regarding BRCA2, only a Colombian founder variant was recurrent: c.2808_2811del (n = 6: 3 in Brazil, 2 in Mexico and 1 in Colombia). In Brazil 3 recurrent PVs accounted for 52.6% (10/19) of the total; in Mexico 11 recurrent PVs accounted for 52.7% (58/110); in Peru 6 recurrent PVs accounted for 31.7% (26/82); in Colombia 3 recurrent PVs accounted for 33.3% (7/21).
BRCAPRO probability model
Sensitivity, specificity, positive predictive value (PPV) and negative predictive value (NPV) were calculated for the BRCAPRO model at a 10% threshold. The area under the receiver operating characteristic (AUROC) curve was 0.76 for BRCAPRO to predict the probability that a participant in this cohort carried a BRCA PV (Fig. 2). The sensitivity, specificity, PPV, NPV and accuracy were 37.6%, 92.1%, 44.1%, 89.9% and 84.3%, respectively.
Discussion
This is the largest study of BRCA PVs among Latin Americans to date, and represents a cross-section including Brazil, Colombia, Peru, Puerto Rico and Mexico. Among 1,627 prospectively accrued participants, most affected with cancer, we detected 236 (14.5%) BRCA PVs using a low-cost BRCA genotyping workflow. Most previous studies were population specific and/or had a small sample size (Supplementary Data)17.
The prevalence of PVs was 9.3–25.7% across the countries included in this study, similar to the range (3.0–47.8%) reported in previous studies (Supplementary Data 2). There were wide variations in reported prevalence within each country. For example, a study of 327 patients in Mexico only reported 7.3% PVs34 and did not test for CNVs other than the Mexican founder PV. Another study of 101 patients in Mexico at high-risk and/or with breast cancer at or under the age of 40 years, which included CNV analysis, reported 13.9% PVs35, which more closely approximated our observation of 17% of the 632 patients in Mexico.
This study highlights the extent to which BRCA CNVs play an important role in the etiology of cancer in Latin America, and particularly in Mexico where CNVs represented 34.5% of PVs, nearly half of which were the Mexican founder, BRCA1 exon 9–12del. The burden of this PV has previously been documented by us and others in independent Mexican cohorts18,19,34,36, and among Mexican Americans13,27,37. CNVs represented < 10% of PVs in Colombia, Peru, and Brazil. The sample size was too small to estimate for Puerto Rico.
The absence of BRCA2 CNVs was notable. The Consortium of Investigators of Modifiers of BRCA1/2 (CIMBA) reported a ratio of 3.5:1 excess of BRCA1 CNVs compared to BRCA2, among 18,435 breast cancer cases with BRCA1 PVs and 11,351 with BRCA2, ascertained from 69 centers in 49 countries on 6 continents. However, Hispanics only represented 3.4% of the total which was primarily NHW9. We confirmed all CNVs by MLPA or long-range PCR, and the CNV detection algorithm incorporated into the Ion Reporter workflow for NGS performs well38. Two of the three studies in Mexico that included CNV analyses (Supplementary Data 2) did not detect any BRCA2 CNVs, though 14 BRCA1 CNVs were detected (11 of which were the Mexican founder PV)35,36. The remaining study reported 3 BRCA1 CNVs and 11 BRCA2 CNVs. However, 9 of 11 were single exon deletions and were not confirmed by a secondary MLPA kit or long-range PCR39, and confirmation is recommended due to a high false-positive rate for single exon CNVs on MLPA. Consequently, we believe the weight of evidence suggests a paucity of BRCA2 CNVs among the populations studied here.
Although limited by high-risk clinic ascertainment, the observed frequency of BRCA PVs highlights the burden of hereditary cancer in every country we examined. We found the highest frequency of PVs in Brazil, where the inclusion criteria required a greater clinical burden of breast/ovarian cancer, followed by Mexico and Peru, while the lowest frequency was found in Colombia and Puerto Rico. Another limitation is known heterogeneity in genetic ancestral composition of among Hispanics from the Americas40,41. We found that the founder Jewish PVs (BRCA1 c.66_67del and c.5266dupC) were recurrent in our cohort. We were among the first to document a shared genotype between Ashkenazi Jewish individuals and Mexican Americans with no known Jewish ancestry who carried BRCA1 c.66_67del6. Subsequent publications have reinforced the observation, and suggest they are likely descendants of conversos or crypto-Jews who emigrated to the Americas in the late 15th century and over generations assimilated into the larger Hispanic society, representing an underappreciated diaspora to the new world13,42,43.
In addition, we evaluated the BRCAPRO model performance for the first time in a Latin American cohort and found that it performed well (AUROC 0.76) and was comparable to that reported for genetic risk clinics in England (0.76)44. BRCAPRO can be used in clinical practice to select patients at risk for carrying BRCA PVs, though it was developed using rates derived from primarily European populations. Similar to our findings, a small study of U.S. Hispanics reported an AUROC of 0.7026. The BRCAPRO results from our study of native Latin American populations, reinforces our observations of prevalence and relationship to clinical phenotype, and also serve as a surrogate indication that our genetic testing methods were sensitive. Further, our study indicates that BRCAPRO can be used as a tool to select individuals at the highest risk of carrying BRCA PVs, which could enable prioritized allocation of limited resources in Latin America.
Worldwide, BRCA testing in the past was commonly performed by capillary DNA (Sanger) sequencing, which was considered the gold standard, albeit expensive. Our study spans 2012–2017, during which time we developed and applied inexpensive approaches, including the HISPANEL on a Sequenom MassARRAY spectroscopy platform, and subsequently, multiplex amplicon library NGS on the Ion Torrent PGM platform that could be deployed in countries with limited resources. The HISPANEL served well in early studies, with an analytic sensitivity compared to full sequencing of BRCA1 and BRCA2 of up to 70% in Mexico, largely because recurrent PVs constituted a large proportion of the overall burden. HISPANEL also performed well in Colombia, due to the prevalence of founder PVs in this country. For example, the three prevalent PVs seen in Colombia, BRCA1 c.5123 C > A, BRCA1 c.3331_3334del, and BRCA2 c.4889 C > G, accounted for a third of all PVs found in this region. In Mexico, the BRCA1 exon 9–12del PV accounted for a fifth of all mutations detected and was not seen outside of Mexico45. In Peru, the observed sensitivity of HISPANEL was 30%, approximately half of what was observed in Colombia and Mexico. This is consistent with the isolation of the Indigenous American Peruvian population, while other populations show more migration between regions46. Despite limitations, this report shows that using a low-cost strategy for broad access to BRCA genotyping in Latin America is feasible. The most recent iteration in our laboratory is a multiplex amplicon multigene panel, validated in the CARRIERS project47.
There are 125 unique PVs among the 236 BRCA PVs reported here; 17/125 (13.6%) were not represented in ClinVar, highlighting the underrepresentation of Latin American ancestry. This lack of representation may influence the effectiveness of variant curation by clinicians. Data from this ongoing study also are valuable to construct population-specific variant frequencies, and to increase representation in public databases48. We also found that 111/196 (56.6%) unique VUS were not reported previously in ClinVar. We recently demonstrated that sequence data well controlled for racial/ethnic composition can yield important observations about pathogenicity of uncertain variants29. The proportion of VUS reported varied by study and gene. The proportion of VUS that we report here (14%) is higher than that in NHW women and comparable to other studies including Hispanics7,49,50. In a meta-analysis of largely European populations, BRCA1 VUS were reported in 1.23% of cases (95% CI 0.75–2.04) and for BRCA2 in 3.29% of cases (CI 95% 0.75–2.42)51. Similarly, the frequency of VUS is elevated in other underrepresented populations, such as in African American and Asiatic49,52, and we demonstrated varying rates of variant reclassification by ancestry over time50. Again, increased representation in public databases is important for clinical translation in diverse populations. Over half of the VUS were not seen in ClinVar, suggesting there may be a registration bias towards deposit of PVs in ClinVar.
In conclusion, strategies to develop affordable BRCA testing in low- and middle-income countries with limited resources are important to extend the reach of lifesaving genetic cancer risk assessment (GCRA). The burden of hereditary breast and ovarian cancer in the respective countries is evident from this and other studies. More than 80% of the breast cancers in this series were diagnosed at Stage II or greater, and 268 cases had multiple primary tumors, including 27 ovarian cancers that might have been prevented if genetic testing was available at breast cancer diagnosis. Dissemination and implementation of GCRA, supported by clinical training and accessible genetic testing33, could result in measurable prevention of breast cancer and its related mortality in Latin America and other low- and middle-income countries globally.
Methods
Study population
Patients seen for GCRA through Clinical Cancer Genomics Community Research Network (CCGCRN)13,53 sites in Latin America between December 2012 and August 2017 were prospectively enrolled after informed consent on an IRB-approved protocol and offered genetic testing. The overall protocol was approved by the City of Hope IRB (#96144; ClinicalTrials.gov Identifier: NCT04185935) data coordinating center, and approved separately, as a Federated consortium at each participating center: (1) Instituto Nacional de Cancerlogía (INCan) in Mexico City, Mexico; (2) Instituto Jalisciense de Cancerologia, in Guadalajara, Mexico; (3) Instituto des Enfermedades Neoplásicas (INEN) in Lima, Peru; (4) Clínica del Country, Oncology Center, in Bogota, Colombia; (5) The University of Puerto Rico and MD Anderson Cancer Center in San Juan, Puerto Rico; and (6) Hospital de Clínicas de Porto Alegre in Porto Alegre, Brazil. Patients met the National Comprehensive Cancer Network (NCCN) guidelines for genetic/familial high-risk assessment: breast and ovarian20. Demographic characteristics, clinical variables and four-generation pedigrees focused on family cancer history were obtained. When more than one person was enrolled and tested in a family, the first person tested was selected for inclusion in study.
DNA extraction and BRCA testing
Genomic DNA was obtained from peripheral blood drawn into ACD-A collection tubes using the Qiagen (Germantown, MD) FlexiGene DNA Kit.
BRCA1 exon 9–12del three-primer PCR assay (3-PA)
For all individuals, 25 ng of genomic DNA was screened for the presence of the copy number variant (CNV), BRCA1 exon 9–12del (c.548-?_4185 + ?del), by a three-primer PCR assay (3-PA)27.
Sequenom MassARRAY genotyping
As described previously, a panel of 114 known pathogenic recurrent BRCA point6,27,13,54 and insertion/deletion PVs among Hispanics was used to genotype all individuals on the Sequenom (San Diego, CA) MassARRAY platform, and together with the BRCA1 exon 9–12del 3-PA, comprises our Hispanic mutation panel (HISPANEL)19.
BRCA CNV
Using Ion Torrent PGM semiconductor sequencing (see below) a CNV baseline was established within the Ion Reporter workflow to enable detection of BRCA1 and BRCA2 CNV. Cases with uninformative HISPANEL and BRCA sequencing were further analyzed for CNVs in BRCA1 by MLPA (MRC-Holland, Amsterdam, the Netherlands). For each MLPA reaction, 125 ng of DNA was PCR amplified with probemix P002 according to the manufacturer’s instructions. PCR fragments were size separated on an ABI (Applied Biosystems, Carlsbad, CA) 3130xl capillary sequencer. Dosage calculations and determination of CNV were performed with Coffalyser.Net software v.140701 (MRC-Holland) and/or GeneMarker software (SoftGenetics, State College, PA). CNVs detected by NGS or MLPA, were confirmed with MLPA probe mixes P087 and P045, respectively. BRCA2 was only evaluated for CNV by NGS, in part due to the known 5-fold greater rate of occurrence in BRCA1 relative to BRCA2 of CNVs37.
Ion torrent PGM semiconductor sequencing
HISPANEL negative cases that met NCCN criteria20 received full sequencing of all BRCA coding exons and exon-intron junctions on the Ion Torrent PGM platform. 10 ng of Qubit (Thermo Fisher Scientific, Waltham, MA) quantified DNA was PCR amplified using the commercially available Ion AmpliSeq BRCA1 and BRCA2 Community Panel (https://www.thermofisher.com/us/en/home/life-science/sequencing/next-generation-sequencing/ion-torrent-next-generation-sequencing-workflow/ion-torrent-next-generation-sequencing-select-targets/ampliseq-target-selection/community-panels.html) - pilot tested by our laboratory for the AmpliSeq BRCA Global Consortium55 – to generate targeted, barcoded, amplicon libraries. Following template preparation and enrichment, 36 individual sample libraries were loaded on an Ion 318 Chip v2 and run for 500 cycles (250 bp single end reads).
Ion reporter cloud
Base calling, call qualification, sequence alignment (genome reference hg19), and coverage analysis were performed in Ion Torrent Suite 5.0.4 (Thermo Fisher Scientific). The average coverage was ≥ 500x across the 171 primers in the panel. After signal processing with Torrent Variant Caller, sequence aligned files were uploaded to Ion Reporter v5.0 Cloud (https://ionreporter.thermofisher.com/ir/) for additional variant calling, annotation and reporting. Using a custom designed workflow, with a CNV baseline established by sequencing 10 BRCA CNV negative control cases, individual cases were examined for all variants.
Variant classification
All variants annotated in Ion Reporter were visualized in ClinVar (https://www.ncbi.nlm.nih.gov/clinvar/) for classification (last time accessed May 2019). Only entries from established commercial vendors (Ambry Genetics, Color Genomics, Counsyl, GeneDx, Invitae, and/or Myriad Genetics (through the Sharing Clinical Reports Project (SCRP), http://clinicalgenome.org/data-sharing/sharing-clinical-reports-project-scrp/), were considered. If a BRCA variant had been evaluated by the Evidence-based Network for Interpretation of Germline Mutant Alleles (ENIGMA) consortium (https://enigmaconsortium.org/), recognized as an expert reviewer in ClinVar, that classification was used. Variant call format files for variants without ClinVar entrees were also uploaded into Ingenuity Variant Analysis (IVA) (Qiagen, Germantown, MD) which applies American College of Medical Genetics and Genomics (ACMG) guidelines to variant classification.
Variant confirmation
Small sequence variants (point, in/del PVs) were confirmed with Sanger sequencing using the BigDye Terminator v3.1 Cycle Sequencing Kit (Thermo Fisher Scientific) and run on an ABI (Applied Biosystems) 3130xl capillary sequencer. CNV variants detected by Ion Torrent sequencing were confirmed with MLPA (see above).
Testing cost
The cost for each test was calculated in USD and based on laboratory reagents and materials as follows: BRCA1 exon 9–12del 3-PA ($0.25/case), Sequenom MassARRAY BRCA Panel ($10/case), Ion Torrent PGM AmpliSeq BRCA Panel ($82/case) and BRCA1 MLPA ($25/case).
Mutation probability model analysis
The probabilities of carrying a BRCA PV were estimated using BRCAPRO (version 2.1.4) in 1,535 cases. BRCAPRO risk estimates are derived through a Bayesian probability model, and accounts for the patient’s first- and second-degree relatives, age of diagnosis of breast and/or ovarian cancer, and ages of unaffected family members56. Pedigrees were created electronically using Progeny (Progeny Software, Delray Beach, FL)57. Sensitivity, specificity, PPV and NPV were calculated for the BRCAPRO model at a 10% threshold. Calibration of the model was conducted by comparing observed versus predicted numbers of PV carriers. The AUROC was calculated to evaluate model discrimination.
Statistical analysis
Variables were summarized using descriptive statistics, including mean, range and frequency. Frequencies of BRCA variants (pathogenic/VUS) were described for the whole cohort, and individually for each country. We also describe the type of mutation (CNV, recurrent and unique variants). Differences between countries of origin were compared and expressed as percentages. The sensitivity of HISPANEL testing was calculated using observed versus detectable, with full BRCA sequencing as the country-specific reference for testing. SAS version 9.4 analytic software (SAS Institute, Cary, NC) was used to carry out the statistical analysis.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data generated and analyzed during this study are described in the following data record: https://doi.org/10.6084/m9.figshare.1491009648. The pathogenic BRCA variants and VUS data are included as separate tabs in ‘Supplementary tables 2 and 3.xlsx’, which is available via the online version of the related article as well as in Excel format as part of the data record. Unique variants have been deposited in ClinVar under accessions https://identifiers.org/clinvar.submission:SCV001739435 to https://identifiers.org/clinvar.submission:SCV00173944458. The minimum dataset necessary to interpret, replicate and/or build upon our methods will be made available on request to the corresponding author.
Code availability
Not applicable.
References
World Health Organization. Global Cancer Observatory, https://gco.iarc.fr/ (2019).
Justo, N., Wilking, N., Jonsson, B., Luciani, S. & Cazap, E. A review of breast cancer care and outcomes in Latin America. Oncologist 18, 248–256 (2013).
Ford, D. et al. Genetic heterogeneity and penetrance analysis of the BRCA1 and BRCA2 genes in breast cancer families. Am. J. Hum. Genet. 62, 676–689 (1998).
Mavaddat, N. et al. Cancer risks for BRCA1 and BRCA2 mutation carriers: results from prospective analysis of EMBRACE. J. Natl. Cancer Inst. 105, 812–822 (2013).
Weitzel, J. N., Blazer, K. R., MacDonald, D. J., Culver, J. O. & Offit, K. Genetics, genomics and cancer risk assessment: state of the art and future directions in the era of personalized medicine. Cancer J. Clin. 61, 327–359 (2011).
Weitzel, J. N. et al. Prevalence of BRCA mutations and founder effect in high-risk Hispanic families. Cancer Epidemiol. Biomark. Prev. 14, 1666–1671 (2005).
Hall, M. J. et al. BRCA1 and BRCA2 mutations in women of different ethnicities undergoing testing for hereditary breast-ovarian cancer. Cancer 115, 2222–2233 (2009).
Neuhausen, S. et al. BRCA1 and BRCA2 mutation carriers in the Breast Cancer Family Registry: an open resource for collaborative research. Breast Cancer Res. Treat. 116, 379–386 (2009).
Rebbeck, T. R. et al. Mutational spectrum in a worldwide study of 29,700 families with BRCA1 or BRCA2 mutations. Hum. Mutat. 39, 593–620 (2018).
Struewing, J. P. et al. The risk of cancer associated with specific mutations of BRCA1 and BRCA2 among Ashkenazi Jews. N. Engl. J. Med. 336, 1401–1408 (1997).
Beck, A. C. et al. Rate of BRCA mutation in patients tested under NCCN genetic testing criteria. Am. J. Surg. 219, 145–149 (2020).
John, E. M., Phipps, A. I., Davis, A. & Koo, J. Migration history, acculturation, and breast cancer risk in Hispanic women. Cancer Epidemiol. Biomark. Prev. 14, 2905–2913 (2005).
Weitzel, J. N. et al. Prevalence and type of BRCA mutations in Hispanics undergoing genetic cancer risk assessment in the southwestern United States: a report from the Clinical Cancer Genetics Community Research Network. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 31, 210–216 (2013).
Hernandez, J. E. et al. Prevalence of BRCA1 and BRCA2 mutations in unselected breast cancer patients from Medellin, Colombia. Hereditary Cancer Clin. Pract. 12, 11 (2014).
Abugattas, J. et al. Prevalence of BRCA1 and BRCA2 mutations in unselected breast cancer patients from Peru. Clin. Genet. 88, 371–375 (2015).
Rodríguez, A. O. et al. BRCA1 and BRCA2 mutations among ovarian cancer patients from Colombia. Gynecol. Oncol. 124, 236–243 (2012).
Dutil, J. et al. The spectrum of BRCA1 and BRCA2 alleles in Latin America and the Caribbean: a clinical perspective. Breast Cancer Res. Treat. 154, 441–453 (2015).
Villarreal-Garza, C. et al. Significant clinical impact of recurrent BRCA1 and BRCA2 mutations in Mexico. Cancer 121, 372–378 (2015).
Villarreal-Garza, C. et al. The prevalence of BRCA1 and BRCA2 mutations among young Mexican women with triple-negative breast cancer. Breast Cancer Res. Treat. 150, 389–394 (2015).
NCCN. NCCN clinical practice guidelines in oncology version 3.2019 genetic/familial high-risk assessment: breast and ovarian. NCCN Clinical Practice Guidelines (2019).
Villarreal-Garza, C. et al. Breast cancer in young women in Latin America: an unmet, growing burden. Oncologist 18, 1298–1306 (2013).
Villarreal-Garza, C. et al. Real-world outcomes in young women with breast cancer treated with neoadjuvant chemotherapy. Breast Cancer Res. Treat. 157, 385–394 (2016).
Cruz-Correa, M. et al. Clinical cancer genetics disparities among Latinos. J. Genet. counseling 26, 379–386 (2017).
Kurian, A. W. et al. Performance of prediction models for BRCA mutation carriage in three racial/ethnic groups: findings from the Northern California Breast Cancer Family Registry. Cancer Epidemiol. Biomark. Prev. 18, 1084–1091 (2009).
Huo, D. et al. Prediction of BRCA mutations using the BRCAPRO model in clinic-based African American, Hispanic, and other minority families in the United States. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 27, 1184–1190 (2009).
Vogel, K. J. et al. BRCA1 and BRCA2 genetic testing in Hispanic patients: mutation prevalence and evaluation of the BRCAPRO risk assessment model. J. Clin. Oncol. Off. J. Am. Soc. Clin. Oncol. 25, 4635–4641 (2007).
Weitzel, J. N. et al. Evidence for common ancestral origin of a recurring BRCA1 genomic rearrangement identified in high-risk Hispanic families. Cancer Epidemiol. Biomark. Prev. 16, 1615–1620 (2007).
Chavarri-Guerra, Y., Blazer, K. R. & Weitzel, J. N. Genetic cancer risk assessment for breast cancer in Latin America. Rev. Invest. Clin. 69, 94–102 (2017).
Weitzel, J. N. et al. Pathogenic and likely pathogenic variants in PALB2, CHEK2, and other known breast cancer susceptibility genes among 1054 BRCA-negative Hispanics with breast cancer. Cancer 125, 2829–2836 (2019).
Popejoy, A. B. & Fullerton, S. M. Genomics is failing on diversity. Nature 538, 161–164 (2016).
Spratt, D. E. et al. Racial/Ethnic disparities in genomic sequencing. JAMA Oncol. 2, 1070–1074 (2016).
Petrovski, S. & Goldstein, D. B. Unequal representation of genetic variation across ancestry groups creates healthcare inequality in the application of precision medicine. Genome Biol. 17, 157 (2016).
Blazer, K. R. et al. Development and pilot implementation of the Genomic risk assessment for cancer implementation and sustainment (GRACIAS) intervention in Mexico. JCO Global Oncology 7, 992–1002 (2021).
Quezada Urban, R. et al. Comprehensive analysis of germline variants in Mexican patients with hereditary breast and ovarian cancer susceptibility. Cancers 10, https://doi.org/10.3390/cancers10100361 (2018).
Zayas-Villanueva, O. A. et al. Analysis of the pathogenic variants of BRCA1 and BRCA2 using next-generation sequencing in women with familial breast cancer: a case-control study. BMC Cancer 19, 722 (2019).
Fragoso-Ontiveros, V. et al. Mexican BRCA1 founder mutation: Shortening the gap in genetic assessment for hereditary breast and ovarian cancer patients. PLoS ONE 14, e0222709 (2019).
Judkins, T. et al. Clinical significance of large rearrangements in BRCA1 and BRCA2. Cancer 118, 5210–5216 (2012).
Germani, A. et al. Rapid detection of copy number variations and point mutations in BRCA1/2 genes using a single workflow by ion semiconductor sequencing pipeline. Oncotarget 9, 33648–33655 (2018).
Fernández-Lopez, J. C. et al. Population and breast cancer patients’ analysis reveals the diversity of genomic variation of the BRCA genes in the Mexican population. Hum. Genomics 13, 3 (2019).
Adhikari, K., Chacon-Duque, J. C., Mendoza-Revilla, J., Fuentes-Guajardo, M. & Ruiz-Linares, A. The genetic diversity of the Americas. Annu. Rev. Genomics Hum. Genet. 18, 277–296 (2017).
Ruiz-Linares, A. et al. Admixture in Latin America: geographic structure, phenotypic diversity and self-perception of ancestry based on 7,342 individuals. PLoS Genet. 10, e1004572 (2014).
Atzmon, G. et al. Abraham’s children in the genome era: major Jewish diaspora populations comprise distinct genetic clusters with shared Middle Eastern Ancestry. Am. J. Hum. Genet. 86, 850–859 (2010).
Velez, C. et al. The impact of Converso Jews on the genomes of modern Latin Americans. Hum. Genet. 131, 251–263 (2012).
Amir, E., Freedman, O. C., Seruga, B. & Evans, D. G. Assessing women at high risk of breast cancer: a review of risk assessment models. J. Natl. Cancer Inst. 102, 680–691 (2010).
Torres, D. et al. Absence of the BRCA1 del (exons 9-12) mutation in breast/ovarian cancer families outside of Mexican Hispanics. Breast Cancer Res. Treat. 117, 679–681 (2009).
Harris, D. N. et al. Evolutionary genomic dynamics of Peruvians before, during, and after the Inca Empire. Proc. Natl. Acad. Sci. USA 115, E6526–E6535 (2018).
Hu, C. et al. A population-based study of genes previously implicated in breast cancer. N. Engl. J. Med 384, 440–451 (2021).
Herzog, J. S. & Weitzel, J. N. Metadata record for the article: genetic epidemiology of BRCA1- and BRCA2-associated cancer across Latin America. figshare, https://doi.org/10.6084/m9.figshare (2021).
Caswell-Jin, J. L. et al. Racial/ethnic differences in multiple-gene sequencing results for hereditary cancer risk. Genet. Med. https://doi.org/10.1038/gim.2017.96 (2017).
Slavin, T. P. et al. Prospective study of cancer genetic variants: variation in rate of reclassification by ancestry. J. Natl Cancer Inst. https://doi.org/10.1093/jnci/djy027 (2018).
van Marcke, C., Collard, A., Vikkula, M. & Duhoux, F. P. Prevalence of pathogenic variants and variants of unknown significance in patients at high risk of breast cancer: a systematic review and meta-analysis of gene-panel data. Crit. Rev. Oncol. Hematol. 132, 138–144 (2018).
Ricks-Santi, L. et al. Next generation sequencing reveals high prevalence of BRCA1 and BRCA2 variants of unknown significance in early-onset breast cancer in African American women. Ethn. Dis. 27, 169–178 (2017).
Slavin, T. et al. Genetic gastric cancer susceptibility in the International Clinical Cancer Genomics Community Research Network. Cancer Genet. 216-217, 111–119 (2017).
Villarreal-Garza, C. M. et al. in ASCO 50th Annual Meeting 2014. (ASCO 50th Annual Meeting, 2014).
Herzog, J. et al. in American Society of Human Genetics. (American Journal of Human Genetics).
Parmigiani, G. et al. Validity of models for predicting BRCA1 and BRCA2 mutations. Ann. Intern. Med. 147, 441–450 (2007).
Sand, S. R., DeRam, D. S., MacDonald, D. J., Blazer, K. R. & Weitzel, J. N. Linkage of a pedigree drawing program and database to a program for determining BRCA mutation carrier probability. Fam. Cancer 4, 313–316 (2005).
Herzog, J. & Weitzel, J. N. ClinVar https://identifiers.org/clinvar.submission:SCV001739435 (2021).
Acknowledgements
Funding for this work was kindly provided by the Breast Cancer Research Foundation BCRF (BCRF-19–172; 20-172); AVON Foundation for Women (Grant No. 02–2013–044); the American Cancer Society (ACS) (ID No. RSGT-10–203–01-CPPB); Conquer Cancer Research Professorship in Breast Cancer Disparities (JNW), R25CA171998 (co-PIs: K. Blazer and J. Weitzel), R25CA085771 (support for J. Clague). City of Hope Clinical Cancer Genomics Community Research Network was also supported in part by Award Number RC4CA153828 (PI: JNW) from the National Cancer Institute and the Office of the Director, National Institutes of Health. The research reported in this publication included work performed in the Integrated Genomics Core at City of Hope supported by the National Cancer Institute of the National Institutes of Health under award number P30CA033572, as well as a supplement award titled: Strengthening capacity to promote genomic cancer risk assessment dissemination and implementation research and translation among underserved populations in the Americas. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. CNPq Brazil Grant No. 408313/2016–1 (Patricia Ashton-Prolla) and CNPq Brazil doctoral fellowship granted to Barbara Alemar. We acknowledge all of our CCGCRN collaborators from Latin America, especially Luis Herrera, Jazmin Arteaga, Dione Aguilar, and Sandra Franco. We thank Thomas Slavin for his collaboration on variant curation.
Author information
Authors and Affiliations
Contributions
Conception and design: S.S. and J.N.W.; data acquisition: J.S.H., Y.C.G., D.C., S.S., C.V.G., J.A., J.C.D., R.M.A.G., T.G.O., A.M.B., P.M., A.D.T.V., Y.R., M.C.C., P.A.P., B.A., L.G. and J.N.W.; data coordinator: R.M.; data management: K.Y.; data analysis: J.S.H., J.C.D., P.A.P. and J.N.W.; data interpretation: J.S.H., S.N. and J.N.W.; financial and administrative support: J.N.W.; and writing, review, and editing: all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Herzog, J.S., Chavarri-Guerra, Y., Castillo, D. et al. Genetic epidemiology of BRCA1- and BRCA2-associated cancer across Latin America. npj Breast Cancer 7, 107 (2021). https://doi.org/10.1038/s41523-021-00317-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41523-021-00317-6
- Springer Nature Limited
This article is cited by
-
Genomic ancestry and cancer among Latin Americans
Clinical and Translational Oncology (2024)
-
Access disparities and underutilization of germline genetic testing in Chilean breast cancer patients
Breast Cancer Research and Treatment (2023)
-
Frequency of germline pathogenic variants in breast cancer predisposition genes among young Turkish breast cancer patients
Breast Cancer Research and Treatment (2023)