Abstract
This study corresponds to the first large-scale genetic analysis of inherited eye diseases (IED) in Argentina and describes the comprehensive genetic profile of a large cohort of patients. Medical records of 22 ophthalmology and genetics services throughout 13 Argentinian provinces were analyzed retrospectively. Patients with a clinical diagnosis of an ophthalmic genetic disease and a history of genetic testing were included. Medical, ophthalmological and family history was collected. A total of 773 patients from 637 families were included, with 98% having inherited retinal disease. The most common phenotype was retinitis pigmentosa (RP, 62%). Causative variants were detected in 379 (59%) patients. USH2A, RPGR, and ABCA4 were the most common disease-associated genes. USH2A was the most frequent gene associated with RP, RDH12 early-onset severe retinal dystrophy, ABCA4 Stargardt disease, PROM1 cone-rod dystrophy, and BEST1 macular dystrophy. The most frequent variants were RPGR c.1345 C > T, p.(Arg449*) and USH2A c.15089 C > A, p.(Ser5030*). The study revealed 156/448 (35%) previously unreported pathogenic/likely pathogenic variants and 8 possible founder mutations. We present the genetic landscape of IED in Argentina and the largest cohort in South America. This data will serve as a reference for future genetic studies, aid diagnosis, inform counseling, and assist in addressing the largely unmet need for clinical trials to be conducted in the region.
Similar content being viewed by others
Introduction
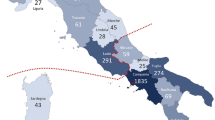
The Latino population has diverse genetic ancestry that includes Native American, Asian, European, West African, and other minorities such as Jewish1. Argentina has received multiple migratory currents from Europe (mostly Spain and Italy), who also brought enslaved peoples from West Africa. The Argentinian population is reported to have 67% European, 28% Native American, 4% West African, and 1% East Asian ancestry2. Given it is the second largest country in South America, the genetic heterogeneity between regions is statistically significant, with European ancestry being the largest in Buenos Aires (76%) and the lowest in the North–West (33%)3,4. African roots are highest in the center of the country (Mendoza, San Juan), and Native American ancestry prevails in the North–West & Chaco (Fig. 1).
The blue gradient represents the percentage of previously unreported/total variants in each province, with darker tones corresponding to higher percentage and lighter, lower. Of note, the province with the highest percentage (60%, Jujuy) contained only a few variants and cases, possibly representing a bias.
Genetics is one of the fastest-growing fields in healthcare, with substantial technological advancements during the last decades5. Initially, access to genetic testing further expanded the disparity between those with access to quality healthcare and those without6. As the cost of testing has decreased, worldwide access has improved, including being covered by the public national healthcare systems in countries such as the United Kingdom.
The sequencing of the first human genome was used to create a standard reference (currently GRCh38), based on 11 individuals from African and white backgrounds7,8. The previous version, GrCh37, is thought to have an ancestral make-up of 57% European, 37% African American and 6% East Asian8,9. Even though these constructs are mostly adequate for clinical and research purposes, the lack of diversity and the use of such reference for other ethnicities has been questioned10. Furthermore, the inequitable representation in genomic research leads to increased incidence of variants of uncertain significance (VUS) amongst individuals from ethnic minorities11,12,13. This disparity leads to difficulties in variant interpretation, genetic counseling, and the need of further exploration, all potentially more challenging in these often less affluent populations.
In the current era of thriving genetic therapies especially in the ophthalmic genetics field14,15, developing countries are starting to share data from their own regions, contributing with previously unreported disease-causing variants, atypical presentations, and detailed longitudinal information16,17,18,19.
In this study, we present the largest South American cohort of genetically confirmed families with inherited eye diseases (IED); an important and timely addition to the global IED genomic dataset.
Results
Demographics and clinical diagnosis
Seven hundred and seventy-three patients from 637 families were included in the study. Three hundred and eighty-six patients (50%) were female and 387 (50%) were male. Amongst those who declared ethnicity (96%), 371 (50%) were white, 299 (40%) were Hispanic or Latino, 66 (9%) were Native Americans, and 6 (1%) were mixed. Two hundred and fifty-eight patients (33%) declared a positive family history of similar eye disease.
The mean age of onset was reported to be 14.8 ± 13.1 years old (birth—82 years range), the mean age at diagnosis was 22.4 ± 15.7 years old (birth—82 years), and the mean age at genetic testing was 36.5 ± 18.9 (6 months old—83 years). The mean time between symptoms onset and diagnosis was 4.3 ± 9.6 years (0–81), between disease onset and genetic testing, 14.9 ± 17.1 (0–82), and between diagnosis and genetic testing, 11.5 ± 14.6 (0–75).
Seven hundred and fifty-six patients (98%) had inherited retinal diseases (IRD), and the remaining 17 (2%) had other etiologies, such as optic atrophy (6) and coloboma (4). Four hundred and eighty-three patients (62%) had a diagnosis of non-syndromic retinitis pigmentosa (RP), 41 (5%) of early-onset severe retinal dystrophy (EOSRD), 40 (5%) of Stargardt disease, 39 (5%) of Usher syndrome, 38 (5%) of macular dystrophy (MD), 19 (2%) of cone-rod dystrophy (CORD), 19 (2%) of choroideremia (CHM), and the remaining patients had less frequent conditions (Fig. 2 and Supplementary Table 1).
RP retinitis pigmentosa, EOSRD early-onset severe retinal dystrophy, MD macular dystrophy, CHM choroideremia, CORD cone-rod dystrophy, IRD inherited retinal dystrophy, BBS Bardet–Biedl syndrome, FEVR familial exudative vitreoretinopathy, XLRS X-linked retinoschisis, MAC microphthalmia-anophthalmia-coloboma spectrum, PHPV persistent hyperplastic primary vitreous, CSNB congenital stationary night blindness, PPRCA pigmented paravenous retinochoroidal atrophy, OA optic atrophy. Albinism & related conditions include oculo-cutaneous albinism, ocular albinism, and foveal hypoplasia.
Genetics
Cohort
Of 637 families, (i) 379 (59%) had a definitive genetic testing result and were considered genetically solved (“positive”), (ii) 178 (28%) had negative testing, (iii) 42 (7%) had only one disease-causing variant in a recessive gene, and (iv) 38 (6%) harbored one or more VUS.
There was no significant difference between the age of onset, age at diagnosis, and age at genetic testing between these four groups (ANOVA P= 0.2621, 0.0654, and 0.6613, respectively). Positive family history was declared by 40% of individuals in the positive genetic testing group, 29% in the negative group, 23% in the one candidate variant, and 29% in the VUS.
In the genetically solved group (n = 379), 220 had autosomal recessive inheritance (58%, 166 compound heterozygous and 54 homozygous), 82 (22%) had an autosomal dominant inheritance, and 77 (20%) were X-linked (Fig. 3A). The most common disease-causing genes were USH2A in 58 families, RPGR in 46, ABCA4 in 35, RHO in 25, PRPF31 and EYS in 14 each, CHM in 13, RDH12 in 11, CRB1 in 10, and the remaining cases appeared in less than 10 families nationwide (Fig. 3B). USH2A was the most common gene to cause RP, RDH12 EOSRD, ABCA4 Stargardt, PROM1 CORD, and BEST1 MD. In the pediatric cohort (under 18 years of age), the most common genes were RPGR (n = 10 families), RS1 (n = 8), RHO (n = 7), ABCA4 (n = 6), and RDH12 and CNGB3 (n = 5 each, Fig. 4A).
A Bar graph showing the most frequently found genes in patients under 18 years of age, ranked by the number of affected families. B Bar graph showing the most frequently found variants in the cohort, ranked by the number of alleles. The remaining variants were present in up to three families nationwide.
Sequence variants
Four hundred and forty-eight different sequence variants were detected in the entire cohort: 193 (43%) missense, 107 (24%) frameshift insertion and/or deletions, 70 (16%) nonsense, 30 (7%) splice site (−2 to +5), 25 (6%) copy number/structural variants, 11 (2%) deep intronic, 8 (2%) inframe insertions and/or deletions, and 4 (1%) synonymous changes.
The most common variants were RPGR c.1345 C > T, p.(Arg449*), present in 18 alleles of 18 unrelated families; USH2A c.15089 C > A, p.(Ser5030*) in 14 alleles of 14 families; USH2A c.2299del, p.(Glu767Serfs*21) and c.2276 G > T, p.(Cys759Phe) in 13 alleles of 13 families each; CERKL c.847 C > T, p(.Arg283*) in 12 alleles of 6 families; ABCA4 c.5882 G > A, p.(Gly1961Glu) in 11 alleles of 11 families, and USH2A c.9119 G > A, p.(Trp3040*) in 10 alleles of 7 families (Fig. 4B).
The most common variant combinations in recessively inherited genes were CERKL homozygous c.847 C > T, p(.Arg283*) in 6 families; USH2A c.15089 C > A, p.(Ser5030*) and c.2299del, p.(Glu767Serfs*21) in 4 each; and CDHR1 homozygous c.1219 C > T, p.(Arg407*), CNGB3 homozygous c.1148del p.(Thr383Ilefs*13), and USH2A homozygous c.9119 G > A, p.(Trp3040*) in 3 families each.
One hundred and ninety-four previously unreported variants (43%) were identified in our cohort; with 156 classified as pathogenic/likely pathogenic and 38 as VUS (Table 1 and Supplementary Table 2). Twenty-one were found in USH2A, 18 in RPGR, 13 in EYS, 8 in ABCA4, 9 in PRPF31, and others in less frequent genes (Table 1). RPGR c.2412_2418del (p.Gly805Lysfs*8), CDHR1 c.1219 C > T (p.Arg407*), CNGB1 c.382-3 C > G, and ARL6 c.506del (p.Gly169Alafs*6) were each present in 5 unrelated families; and PROM1 c.1956T > G (p.Tyr652*), EYS c.6131_6134del (p.Asn2044Thrfs*11), RPGR c.2405_2406del (p.Glu802Glyfs*32), and PRPF31 c.527 + 1 G > A were in three unrelated families, each; possibly representing founder mutations.
The variants appeared in similar proportion of individuals with Latino and white self-claimed identity (Supplementary Table 1). The distribution of these previously unreported variants in Argentina is depicted in Fig. 1, where we see that provinces with less mixing between different ethnic populations and European migration (Jujuy, Tucuman and Chaco) have 60%, 43 and 41% of their variants not formerly reported, respectively4,20. The transcripts used in this project are detailed on Supplementary Table 3.
Twenty-one percent diagnostic uplift
One hundred and fifty-six families (156/637) harboring one or more VUS were analyzed in detail, as described in “Methods”.
After such analysis (Supplementary Table 4), 46 families were confirmed as negative, 41 remained in the VUS group, 37 were classified as one (likely) disease-causing variant only, and 32 families (21%) were reclassified to genetically solved (positive).
Discussion
The disparity in healthcare access between populations and ethnicities is a huge global concern and arguably only increasing21. This inequality affects all aspects of medicine, with particular challenges in expensive fields such as advanced therapies and molecular genetics, which are unreachable to huge numbers of people around the world22,23. There is a need to advocate for and work towards equal access, not only for ethical purposes but also because more representative global data will increase our understanding of diseases and potentially how different environmental factors play a role.
The study herein is the first large-scale genetic analysis of IED in Argentina and the largest in South America, describing the genetic profile of this understudied population. The diagnostic rate (59%) was in keeping with other countries such as UK24, Spain25, Poland26, Korea27, China28, and USA29. It is noteworthy that next-generation sequencing (NGS)-based panels continue to be a key first-tier test worldwide, with constantly updated panels including complex regions such as RPGR-ORF15 and deep intronic areas24. These panel tests are currently not covered by most health insurances in Argentina, however, this study further reinforces their relevance as a standard-of-care assessment and their applicability to our region1.
The mean age at genetic testing in Argentina was similar to a large cohort in USA (36.5 years versus 37.3)30, and younger than other groups in Asia27,31. The percentage of individuals with positive family history was similar to other cohorts as well, with consistent no significant age differences between positive and negative family history subcohorts27,31. Still, there was an 8-year difference between mean age of onset and diagnosis, and a further 14-year gap until genetic testing. Of course, this represents a significant delay in genetic diagnosis (“the genetic odyssey”), emphasizing the critical need to improve access to affordable genetic testing at the point of clinical diagnosis, rather than several years/decades later.
The most common genes and variants mirrored other large IRD cohorts, with the caveat that our patients were primarily ascertained via a patient group for RP, hence ABCA4 was the third most common gene instead of the first25,30,31,32,33,34. USH2A was the most common gene to cause RP, in agreement with other reports;35,36 and RPGR, the most common gene to cause X-linked RP37, as second in prevalence. Interestingly, RDH12 was the most frequently identified gene causing EOSRD in our cohort, and not CEP290, GUCY2D, or CRB1, as described in Brazil, North Africa, and UK, respectively16,17,32. The large variability worldwide regarding EOSRD genes may be due to the small sample size and the potential misclassification of some cases as RP or other rod-cone dystrophies; or maybe a true reflection of genetic diversity globally. BEST1 appearing as the most frequent gene in MD (with four families), and not ABCA4 or PRPH2 (present in three families each), may relate to the selection bias of our sample and Stargardt being a separate clinical category25,30.
Our cohort has also provided additional evidence for rare genes with limited cases in the literature, supporting their pathogenicity in IED and their associated phenotypes. FRMD7 was found in a patient with X-linked congenital nystagmus, as previously reported38; biallelic RTN4IP1 changes were detected in a patient with concomitant RP and optic atrophy, a recently described phenotype39; ARHGEF18 in a patient with autosomal recessive RP40; PRPF6 in a patient with autosomal dominant RP41, and ARSG in a patient with Usher syndrome type 442. Of note, the variant ABCA4 p.Asn1868Ile was not reported by the clinical laboratory, hence its linkage with other variants could not be ascertained43.
Variant interpretation is key to providing an accurate diagnosis to patients and families, facilitating the best possible clinical management, family counseling/planning, and enabling access to potential gene-based therapies. Particularly in the discipline of rare diseases, every contribution is helpful to better understand the pathophysiology of these conditions. One hundred and fifty-six previously unreported (likely) disease-causing variants were identified, representing 35% of all the variants in the cohort (Table 1). Perhaps unsurprisingly, this is a larger proportion than that reported in well-characterized populations such as those in North America and Europe44,45,46, and closer to values reported in Asian projects31,47. A further 38 previously unreported variants were classified as VUS, with more data needed to reclassify them as benign or pathogenic. Similar to proposed disease- and gene-specific guidelines to classify variants48, it would be valuable to also introduce population or minority-specific criteria, to be able to recognize population-associated evidence in large-scale genome-based studies.
Certain variants reported herein were not only seen in European alleles (i.e., USH2A c.12575 G > A in Spain49, USH2A c.1751G > T in Italy50, PRPF31 c.371_375del in Germany51, TYR c.996 G > A in Denmark52, USH2A c.11864 G > A in UK)53, or other American countries (COL2A1 c.3574 C > T in Brazil54, RPGR c.1345 C > T in North America55), but also in populations from all around the world (USH2A c.5329 C > T in Japan56, EYS c.5450 G > A in a Bedouin tribe in Israel57, FAM161A c.1003 C > T in Palestine58, CNGB3 c.1148del in Pingelapese islanders of Micronesia59, and USH2A c.5858 C > G in Tunisia, among others)60.
Remarkably, 100% of CERKL-associated retinopathy in Argentina was due to c.847 C > T, p(.Arg283*), a variant enriched in European populations, not characterized as prevalent amongst Latinos (https://gnomad.broadinstitute.org/variant/2-182423344-G-A?dataset=gnomad_r2_1)61. This is possibly due to European migration to Argentina.
This study’s limitations include its retrospective nature, and that there was a predominant representation of patients with RP compared to other IEDs. Expanding the analysis to include a broader spectrum of disease in the future would benefit patients and scientists alike. Segregation data and detailed clinical information were also limited. There is a restricted testing capacity of NGS-based panels, such as intronic regions remaining untested and the inability to interrogate new genes; tests with larger coverage, such as whole genome sequencing, would be required to uncover a larger proportion of pathogenic variants, although this introduces additional complexities and challenges62. Access to testing in this study is likely to have not been uniform across the country and so there may be regions and provinces that are not/underrepresented. Furthermore, patients from rural areas may have traveled to nearby cities to get tested, hence large provinces such as Cordoba and Buenos Aires may include inhabitants from neighboring provinces. There is also limited funding for further required research, such as trio analysis, particularly relevant due to the high incidence of VUS.
In summary, this is the first comprehensive study of the genetic landscape of IRD in Argentina, describing over 150 previously unreported disease-associated variants, and 8 possible founder mutations. RPGR and two USH2A-exon 13 variants (c.2299del and c.2276 G > T) are frequent in our cohort, in keeping with previous reports30,63, and relevant for directed gene therapy clinical trials (NCT04671433, NCT05158296 and NCT05176717). Two unrelated patients with RPE65-EOSRD have been treated with Luxturna for the first time in Argentina in 2022, paving the way for more to come. We believe this data improves the understanding of IED genetics in Argentina and will support access to the best possible clinical care for patients, as well as contribute to worldwide registries, and the development of public health policies towards a more equitable access to healthcare.
Moreover, reporting this Argentinian variome for the first time in a cohort this large will contribute to improving the understanding of disease-causing variants, delineating future large-scale population genome projects in South America and, along with other efforts worldwide64,65,66, bring us closer to map human diversity67,68.
Methods
Medical records review
Medical records of 22 ophthalmology and genetics services throughout 13 provinces in Argentina were reviewed for this retrospective study (Fig. 1). Patients with a clinical diagnosis of an ophthalmic genetic disease and a history of genetic testing were included. The diagnoses were made by trained ophthalmologists and the diagnostic algorithm varied amongst the regions, with a clinical diagnosis based on history and retinal examination in rural areas, and additional multimodal imaging and retinal functional assessments in urban environments. Medical, ophthalmological, and family history was collected.
To reach a diagnostic consensus across centers, RP was defined as a rod-cone dystrophy with onset after 5 years of age; EOSRD, a severe retinal dystrophy presenting before 5 years old;69 and Stargardt disease was a category on its own70.
This study was performed in accordance with the ethical standards of the Declaration of Helsinki and was approved by the ethics committee of the Argentine Society of Ophthalmology. Written informed consent was obtained in all cases prior to genetic testing. Most of the patients (96%) had genetic testing through a sponsored program by Invitae laboratory (San Francisco, CA, USA), which took place between July 2021 and August 2022. It included an NGS-based IRD panel of 330 genes (https://www.invitae.com/en/providers/test-catalog/test-72100). Twenty patients were tested with an NGS IRD panel of 224 genes (https://mendelics.com.br/en/especialidades/oftalmologia-en/hereditary-retinopathy-panel/), and nine had an older NGS IRD panel of 39 genes (https://dbgen.com/ 2017). Most patients were referred to testing by the RP Argentina Foundation (FARP, www.retinosisargentina.com), hence the sample had a selection bias towards RP.
Genetic testing analysis
Invitae uses Illumina sequencing technology, with a minimal read depth ≥50x, and aligns the reads to the reference sequence GRCh37. Variants reported as pathogenic and likely pathogenic by the accredited diagnostic laboratory were interpreted as such and not queried. VUS were analyzed by MDV and GA when deemed as possibly disease-causing, based on family history, phenotype, and/or if concurrent with a pathogenic/likely pathogenic change in a candidate recessive gene. This analysis considered the VUS protein effect, familial segregation when available, pathognomonic retinal phenotype when applicable, frequency in the general population (https://gnomad.broadinstitute.org/)71, American College of Medical Genetics (ACMG) classification72, in silico prediction tools (Revel, MutationTaster, and SpliceAI)73,74,75,76, conservation score (PhyloP100way)77, and their presence in genetic databases (HGMD and ClinVar, Supplementary Table 4). Cases were uplifted to positive when the VUS could be reclassified as likely pathogenic or pathogenic, categorized as negative when no sufficient evidence was found, classified into a “one candidate variant” category if they carried only one pathogenic or likely pathogenic variant in a candidate recessive gene, or placed into a VUS category if the case remained uncertain after analysis. In the exceptional case where the phenotype was pathognomonic of one gene only, and the family history was consistent with the inheritance pattern (Supplementary Table 4, ID 5), PP4 was uplifted to moderate evidence to classify this variant.
GraphPad Prism 8.0.2 (GraphPad Software, San Diego, CA, USA) was implemented for statistical analysis. The threshold of significance was set at P < 0.05.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request and upon Data Usage Agreement.
References
Daich Varela, M. et al. Ophthalmic genetics in South America. Am. J. Med Genet. C. Semin Med Genet. 184, 753–761 (2020).
Homburger, J. R. et al. Genomic insights into the ancestry and demographic history of South America. PLoS Genet. 11, e1005602 (2015).
Avena, S. et al. Heterogeneity in genetic admixture across different regions of Argentina. PLoS ONE 7, e34695 (2012).
Muzzio, M. et al. Population structure in Argentina. PLoS ONE 13, e0196325 (2018).
Durmaz, A. A. et al. Evolution of genetic techniques: past, present, and beyond. Biomed. Res. Int. 2015, 461524 (2015).
Hall, M. & Olopade, O. I. Confronting genetic testing disparities knowledge is power. J. Am. Med. Assoc. 293, 1783–1785 (2005).
Reich, D. et al. Reduced neutrophil count in people of African descent is due to a regulatory variant in the Duffy antigen receptor for chemokines gene. PLoS Genet. 5, e1000360 (2009).
Green, R. E. et al. A draft sequence of the Neandertal genome. Science 328, 710–722 (2010).
Sherman, R. M. & Salzberg, S. L. Pan-genomics in the human genome era. Nat. Rev. Genet. 21, 243–254 (2020).
Lek, M. & Mardis, E. R. Envisioning the next human genome reference. Dis. Model Mech. 14, dmm049426 (2021).
Soewito S. et al. Abstract P2-09-06: Increased rates of genetic variants of unknown significance in Latino and African American populations of south Texas. Cancer Res. 82(4_Supplement), P2-09-06-P2-09-06 (2022).
Opatt, D. M., Morrow, M. & Daly, M. The incidence of BRCA1 and BRCA2 variants of unknown significance varies in different ethnic populations. J. Clin. Oncol. 24, 10002 (2006).
Chern, J. Y., Lee, S. S., Frey, M. K., Lee, J. & Blank, S. V. The influence of BRCA variants of unknown significance on cancer risk management decision-making. jgo 30, 0 (2019).
Varela, M. D., de Guimaraes, T. A. C., Georgiou, M. & Michaelides, M. Leber congenital amaurosis/early-onset severe retinal dystrophy: current management and clinical trials. Br. J. Ophthalmol. 106, 445–451 (2022).
Varela, M. D., Georgiadis, T. & Michaelides, M. Genetic treatment for autosomal dominant inherited retinal dystrophies: approaches, challenges and targeted genotypes. Br. J. Ophthalmol. https://doi.org/10.1136/bjo-2022-321903 (2022).
Bouzidi, A. et al. Clinical and genetic spectrums of 413 North African families with inherited retinal dystrophies and optic neuropathies. Orphanet J. Rare Dis. 17, 197 (2022).
Sallum, J. M. F. et al. Clinical and molecular findings in a cohort of 152 Brazilian severe early onset inherited retinal dystrophy patients. Am. J. Med Genet. C. Semin Med Genet. 184, 728–752 (2020).
Villanueva, A. et al. Identification of the genetic determinants responsible for retinal degeneration in families of Mexican descent. Ophthalmic Genet. 39, 73–79 (2018).
Pandova, M. G. et al. Inherited retinal dystrophies in a Kuwaiti tribe. Ophthalmic Genet. 43, 438–445 (2022).
Luisi, P. et al. Fine-scale genomic analyses of admixed individuals reveal unrecognized genetic ancestry components in Argentina. PLoS ONE 15, e0233808 (2020).
Pérez-Stable, E. J., Alvidrez, J. & Hill, C. V. Definitions, principles, and concepts for minority health and health disparities research. Sci. Health Disparities Res. 1–12 https://doi.org/10.1002/9781119374855.ch1 (2021).
Elam, A. R. et al. Disparities in vision health and eye care. Ophthalmology 129, e89-e113 (2022).
Suther, S. & Kiros, G. E. Barriers to the use of genetic testing: a study of racial and ethnic disparities. Genet. Med J. Am. Coll. Med. Genet. 11, 655–662 (2009).
Sheck, L. H. N. et al. Panel-based genetic testing for inherited retinal disease screening 176 genes. Mol. Genet Genom. Med. 9, e1663 (2021).
Perea-Romero, I. et al. Genetic landscape of 6089 inherited retinal dystrophies affected cases in Spain and their therapeutic and extended epidemiological implications. Sci. Rep. 11, 1526 (2021).
Tracewska, A. M. et al. Non-syndromic inherited retinal diseases in Poland: genes, mutations, and phenotypes. Mol. Vis. 27, 457–465 (2021).
Ma, D. J. et al. Whole-exome sequencing in 168 Korean patients with inherited retinal degeneration. BMC Med. Genom. 14, 74 (2021).
Huang, X. F. et al. Genotype–phenotype correlation and mutation spectrum in a large cohort of patients with inherited retinal dystrophy revealed by next-generation sequencing. Genet Med. 17, 271–278 (2015).
Goetz, K. E. et al. Genetic testing for inherited eye conditions in over 6,000 individuals through the eyeGENE network. Am. J. Med. Genet. C. Semin Med. Genet. 184, 828–837 (2020).
Stone, E. M. et al. Clinically focused molecular investigation of 1000 consecutive families with inherited retinal disease. Ophthalmology 124, 1314–1331 (2017).
Chen, T. C. et al. Genetic characteristics and epidemiology of inherited retinal degeneration in Taiwan. NPJ Genom. Med. 6, 16 (2021).
Pontikos, N. et al. Genetic basis of inherited retinal disease in a molecularly characterized cohort of more than 3000 families from the United Kingdom. Ophthalmology 127, 1384–1394 (2020).
Yeung, K. Y. et al. Molecular diagnostics for retinitis pigmentosa. Clin. Chim. Acta 313, 209–215 (2001).
Tuson, M., Marfany, G. & Gonzàlez-Duarte, R. Mutation of CERKL, a novel human ceramide kinase gene, causes autosomal recessive retinitis pigmentosa (RP26). Am. J. Hum. Genet. 74, 128–138 (2004).
McGee, T. L., Seyedahmadi, B. J., Sweeney, M. O., Dryja, T. P. & Berson, E. L. Novel mutations in the long isoform of the USH2A gene in patients with Usher syndrome type II or non-syndromic retinitis pigmentosa. J. Med Genet. 47, 499–506 (2010).
Rivolta, C., Sweklo, E. A., Berson, E. L. & Dryja, T. P. Missense mutation in the USH2A gene: association with recessive retinitis pigmentosa without hearing loss. Am. J. Hum. Genet. 66, 1975–1978 (2000).
Yang, L. et al. Novel mutations of RPGR in Chinese retinitis pigmentosa patients and the genotype-phenotype correlation. PLoS ONE 9, e85752 (2014).
Tarpey, P. et al. Mutations in FRMD7, a newly identified member of the FERM family, cause X-linked idiopathic congenital nystagmus. Nat. Genet. 38, 1242–1244 (2006).
Rajabian, F. et al. Combined optic atrophy and rod–cone dystrophy expands the RTN4IP1 (Optic Atrophy 10) phenotype. J. Neuro-Ophthalmol. 41, e290-e292 (2021).
Arno, G. et al. Biallelic mutation of ARHGEF18, involved in the determination of epithelial apicobasal polarity, causes adult-onset retinal degeneration. Am. J. Hum. Genet. 100, 334–342 (2017).
Tanackovic, G. et al. A missense mutation in PRPF6 causes impairment of pre-mRNA splicing and autosomal-dominant retinitis pigmentosa. Am. J. Hum. Genet. 88, 643–649 (2011).
Velde, H. M. et al. Usher syndrome type IV: clinically and molecularly confirmed by novel ARSG variants. Hum. Genet. 141, 1723–1738 (2022).
Aguirre-Lamban, J. et al. Further associations between mutations and polymorphisms in the ABCA4 gene: clinical implication of allelic variants and their role as protector/risk factors. Invest Ophthalmol. Vis. Sci. 52, 6206–6212 (2011).
Villanueva-Mendoza, C. et al. The genetic landscape of inherited retinal diseases in a Mexican Cohort: genes, mutations and phenotypes. Genes 12 https://doi.org/10.3390/genes12111824 (2021).
Whelan, L. et al. Findings from a genotyping study of over 1000 people with inherited retinal disorders in Ireland. Genes 11 https://doi.org/10.3390/genes11010105 (2020).
Ellingford, J. M. et al. Molecular findings from 537 individuals with inherited retinal disease. J. Med. Genet. 53, 761 LP–767 (2016).
Wang, L. et al. Application of whole exome and targeted panel sequencing in the clinical molecular diagnosis of 319 Chinese families with Inherited retinal dystrophy and comparison study. Genes 9 https://doi.org/10.3390/genes9070360 (2018).
Hoskinson, D. C., Dubuc, A. M. & Mason-Suares, H. The current state of clinical interpretation of sequence variants. Curr. Opin. Genet. Dev. 42, 33–39 (2017).
Ávila-Fernández, A. et al. Mutation analysis of 272 Spanish families affected by autosomal recessive retinitis pigmentosa using a genotyping microarray. Mol. Vis. 16, 2550–2558 (2010).
Colombo, L. et al. Molecular epidemiology in 591 Italian probands with nonsyndromic retinitis pigmentosa and usher syndrome. Invest Ophthalmol. Vis. Sci. 62, 13 (2021).
Weisschuh, N. et al. Genetic architecture of inherited retinal degeneration in Germany: a large cohort study from a single diagnostic center over a 9-year period. Hum. Mutat. 41, 1514–1527 (2020).
Grønskov, K. et al. Birth prevalence and mutation spectrum in danish patients with autosomal recessive albinism. Invest. Ophthalmol. Vis. Sci. 50, 1058–1064 (2009).
Le Quesne Stabej, P. et al. Comprehensive sequence analysis of nine Usher syndrome genes in the UK National Collaborative Usher Study. J. Med Genet. 49, 27–36 (2012).
Zechi-Ceide, R. M. et al. Clinical evaluation and COL2A1 gene analysis in 21 Brazilian families with Stickler syndrome: identification of novel mutations, further genotype/phenotype correlation, and its implications for the diagnosis. Eur. J. Med. Genet. 51, 183–196 (2008).
Breuer, D. K. et al. A comprehensive mutation analysis of RP2 and RPGR in a North American cohort of families with X-linked retinitis pigmentosa. Am. J. Hum. Genet. 70, 1545–1554 (2002).
Nakanishi, H. et al. Novel USH2A mutations in Japanese Usher syndrome type 2 patients: marked differences in the mutation spectrum between the Japanese and other populations. J. Hum. Genet. 56, 484–490 (2011).
Wormser, O. et al. Combined CNV, haplotyping and whole exome sequencing enable identification of two distinct novel EYS mutations causing RP in a single inbred tribe. Am. J. Med. Genet. A 176, 2695–2703 (2018).
Zobor, D., Balousha, G., Baumann, B. & Wissinger, B. Homozygosity mapping reveals new nonsense mutation in the FAM161A gene causing autosomal recessive retinitis pigmentosa in a Palestinian family. Mol. Vis. 20, 178–182 (2014).
Sundin, O. H. et al. Genetic basis of total colourblindness among the Pingelapese islanders. Nat. Genet. 25, 289–293 (2000).
Chebil, A. et al. [Genotype-phenotype correlation in ten Tunisian families with non-syndromic retinitis pigmentosa]. J. Fr. Ophtalmol. 39, 277–286 (2016).
Avila-Fernandez, A. et al. CERKL mutations and associated phenotypes in seven Spanish families with autosomal recessive retinitis pigmentosa. Invest. Ophthalmol. Vis. Sci. 49, 2709–2713 (2008).
Ellingford, J. M. et al. Whole genome sequencing increases molecular diagnostic yield compared with current diagnostic testing for inherited retinal disease. Ophthalmology 123, 1143–1150 (2016).
Carss, K. J. et al. Comprehensive rare variant analysis via whole-genome sequencing to determine the molecular pathology of inherited retinal disease. Am. J. Hum. Genet. 100, 75–90 (2017).
Francioli, L. C. et al. Whole-genome sequence variation, population structure and demographic history of the Dutch population. Nat. Genet. 46, 818–825 (2014).
100 000 Genomes Project Pilot Investigators. 100,000 genomes pilot on rare-disease diagnosis in health care—preliminary report. N. Engl. J. Med. 385, 1868–1880 (2021).
Auton, A. et al. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Cho, Y. S. et al. An ethnically relevant consensus Korean reference genome is a step towards personal reference genomes. Nat. Commun. 7, 13637 (2016).
Hindorff, L. A. et al. Prioritizing diversity in human genomics research. Nat. Rev. Genet. 19, 175–185 (2018).
Kumaran, N., Moore, A. T., Weleber, R. G. & Michaelides, M. Leber congenital amaurosis/early-onset severe retinal dystrophy: clinical features, molecular genetics and therapeutic interventions. Br. J. Ophthalmol. 101, 1147–1154 (2017).
Tanna, P., Strauss, R. W., Fujinami, K. & Michaelides, M. Stargardt disease: clinical features, molecular genetics, animal models and therapeutic options. Br. J. Ophthalmol. 101, 25–30 (2017).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Rowlands, C. et al. Comparison of in silico strategies to prioritize rare genomic variants impacting RNA splicing for the diagnosis of genomic disorders. Sci. Rep. 11, 20607 (2021).
Jaganathan, K. et al. Predicting splicing from primary sequence with deep learning. Cell 176, 535–548.e24 (2019).
Ioannidis, N. M. et al. REVEL: an ensemble method for predicting the pathogenicity of rare missense variants. Am. J. Hum. Genet. 99, 877–885 (2016).
Schwarz, J. M., Cooper, D. N., Schuelke, M. & Seelow, D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat. Methods 11, 361–362 (2014).
Pollard, K. S., Hubisz, M. J., Rosenbloom, K. R. & Siepel, A. Detection of nonneutral substitution rates on mammalian phylogenies. Genome Res. 20, 110–121 (2010).
Acknowledgements
G.A. is funded by a Fight For Sight UK Early Career Investigator Award (5045/46), the National Institute for Health Research Biomedical Research Centre at Great Ormond Street Hospital Institute for Child Health and at Moorfields Eye Hospital NHS Foundation Trust and UCL Institute of Ophthalmology, and Moorfields Eye Charity. O.M. is supported by The Wellcome Trust (206619/Z/17/Z). M.M. is supported by The Wellcome Trust (099173/Z/12/Z), by the National Institute for Health Research Biomedical Research Centre at Moorfields Eye Hospital NHS Foundation Trust and UCL Institute of Ophthalmology, Moorfields Eye Charity, Retina UK, and the Foundation Fighting Blindness. This work would not have been possible without the collaboration of Invitae, Francisco Albarracin, the president and members of RP Argentina Foundation, Alberto Inga, the support staff of all involved centers, and the trust and engagement of the patients and families.
Author information
Authors and Affiliations
Contributions
M.D.V., P.G.S., and G.A. contributed to the design of this study. J.L., N.L., M.B.Y., S.B., E.E., A.I., E.I.B., A.A.M., C.P.P., A.C.M., L.A., J.G., L.C., M.V., J.E.D., E.A., L.F.G., S.A., M.E.A., D.M.M., G.J.O., M.E.I., A.A.F., M.C., T.Z., P.J.P., V.L., P.J.N., and I.B.C. evaluated patients and collected their personal and medical information. O.M. and M.M. reviewed the manuscript and provided valuable feedback. All authors approved the submission of this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schlottmann, P.G., Luna, J.D., Labat, N. et al. Nationwide genetic analysis of more than 600 families with inherited eye diseases in Argentina. npj Genom. Med. 8, 8 (2023). https://doi.org/10.1038/s41525-023-00352-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41525-023-00352-1
- Springer Nature Limited