Abstract
The human spaceflight environment is notable for the unique factor of microgravity, which exerts numerous physiologic effects on macroscopic organisms, but how this environment may affect single-celled microbes is less clear. In an effort to understand how the microbial transcriptome responds to the unique environment of spaceflight, the model Gram-positive bacterium Bacillus subtilis was flown on two separate missions to the International Space Station in experiments dubbed BRIC-21 and BRIC-23. Cells were grown to late-exponential/early stationary phase, frozen, then returned to Earth for RNA-seq analysis in parallel with matched ground control samples. A total of 91 genes were significantly differentially expressed in both experiments; 55 exhibiting higher transcript levels in flight samples and 36 showing higher transcript levels in ground control samples. Genes upregulated in flight samples notably included those involved in biofilm formation, biotin and arginine biosynthesis, siderophores, manganese transport, toxin production and resistance, and sporulation inhibition. Genes preferentially upregulated in ground control samples notably included those responding to oxygen limitation, e.g., fermentation, anaerobic respiration, subtilosin biosynthesis, and anaerobic regulatory genes. The results indicated differences in oxygen availability between flight and ground control samples, likely due to differences in cell sedimentation and the toroidal shape assumed by the liquid cultures in microgravity.
Similar content being viewed by others
Introduction
In certain respects, human spaceflight habitats resemble other confined built environments, such as submersible vehicles, aircraft, hospital isolation wards, or remote research installations.1 However, the spaceflight environment is unique because it contains two additional altered physical parameters: reduced (micro-)gravity and increased ionizing radiation from solar and galactic sources. Extensive investigations conducted in spaceflight on macroscopic organisms have resulted in a relatively good understanding of the biological effects of microgravity and radiation at levels ranging from the whole body down to the organ, cellular, and molecular level in humans,2 animals,3 and plants.4 While microorganisms have also been the subject of focused research in the spaceflight environment, it has proven more difficult to understand their responses to spaceflight stress.5,6,7 From a theoretical perspective, exposure to microgravity results in a number of alterations in a microbial cell’s immediate surroundings, such as loss of convective mass and heat transfer, reduction in mechanical shear forces, and alterations in the way liquids behave at air and solid interfaces. Changes in such fundamental physical forces alter the rates at which gases, nutrients, signaling molecules, and waste products are exchanged between microbes and their surroundings. It has been proposed that upon perception of these alterations in their environment, microbes mount a complex set of stress responses (the so-called “spaceflight syndrome”8).
Considerable effort has been expended to understand microbial responses to spaceflight and their underlying causes. In early studies, various phenotypic outputs from microbes grown in space were measured, such as: growth rate and yield; virulence; biofilm formation and architecture; and resistance to antibiotics or abiotic stresses.5,6,9 More recent efforts tended toward gene expression studies using genome-wide techniques such as microarrays to understand how the global pattern of RNA synthesis (i.e., the transcriptome) responds to the spaceflight environment. To date, microarray studies have reported a wide range of responses to spaceflight including increased transcription of genes encoding general metabolism,10,11 secondary metabolite biosynthesis,12 synthesis of ribosomal proteins,11,13 and virulence factors.13,14 Regardless of the output measured, it has proven difficult to derive consistent conclusions from these disparate studies due to several confounding factors.
First, until recently spaceflight transcriptome studies have been performed on only a small selection of Gram-negative bacteria (Salmonella enterica serovar Typhimurium, Pseudomonas aeruginosa, Rhodospirillum rubrum), limiting the ability to generalize conclusions to a broader range of microbes. Second, spaceflight experiments have been conducted under widely different: (i) culture conditions (e.g., media formulations, agar vs. liquid, aeration, temperature); (ii) growth stage of the cultures at harvest; (iii) spaceflight hardware employed; (iv) pre- and post-flight treatment of samples; and (v) types of assays conducted.6,7 Third, experimental variation derives from the measurements themselves (technical effects) or from the natural variation inherent in biological systems (biological effects).15 In an effort to control for variation, prior microbial spaceflight experiments have included multiple replicates; however, most experiments reported in the literature have been flown on a single mission only. Intense competition for limited cargo space destined to research platforms such as the International Space Station (ISS) generally results in the choice a new experiment taking precedence over a repetition of a previously flown experiment. Because of this, the intrinsic mission-to-mission variability in the response of microbiological systems to the spaceflight environment has remained largely unexplored.
In 2015, we were afforded the opportunity to send an experimental package to the ISS to test the responses of the Gram-positive bacterium Bacillus subtilis to the human spaceflight environment. This was the 21st mission to the ISS using Biological Research in Canister-Petri Dish Fixation Unit (BRIC-PDFU) hardware, and the experiment was dubbed BRIC-21. From the BRIC-21 experiment we have previously reported in detail measurements of the growth, antibiotic resistance, frequency and spectrum of mutagenesis exhibited by B. subtilis flight (FL) samples in comparison to matched ground control (GC) samples.16,17 In addition, we also performed RNA-seq analyses to compare the transcriptomes of BRIC-21 FL vs. GC samples, as we will report in this communication. In 2016 we had the good fortune, in collaboration with the NASA GeneLab group, to fly a second mission to the ISS (dubbed BRIC-23) using the same B. subtilis strain, media, and hardware, and again to perform RNA-seq analyses on the samples. Here we provide a comparative analysis of the B. subtilis transcriptome profiles from the BRIC-21 and BRIC-23 spaceflight missions. We report on the complete transcriptome profiling of a Gram-positive bacterium grown in the human spaceflight environment (a prior study focused on a subset of primary and secondary metabolite genes in Streptomyces coelicolor12). In this study, we show the effect of exposure to the human spaceflight environment on the B. subtilis transcriptome by identifying sets of genes expressed in common in both the BRIC-21 and BRIC-23 missions.
Results
RNA-seq was used to characterize the transcriptomic response of B. subtilis cultures exposed to the human spaceflight environment of the ISS (FL samples) vs. matched GC samples on two separate missions, BRIC-21 (n = 3) and BRIC-23 (n = 9). The data were analyzed using the bioinformatics pipeline described in the section 'Methods' and the results are presented below.
Overview of the datasets
In order to assess the quality and reproducibility of the datasets obtained, principal component analysis (PCA) was performed. In the PCA, the first and second principal components explained 52 and 21% of the variance, respectively. Four distinct population clusters were identified corresponding to the four environmental conditions tested (Fig. 1). In the BRIC-21 FL and GC samples, the three replicates were grouped rather tightly, indicating relatively good agreement. In the BRIC-23 FL and GC samples, the nine replicates were somewhat more disperse, but still formed distinct groups (Fig. 1). Examination of Principal Component 1 revealed that the major source of variation in the datasets derived from the differences in the two missions themselves, while variation in Principal Component 2 was due to differences between the FL and GC datasets in each experiment (Fig. 1).
The B. subtilis strain 168 transcriptome consists of 4397 total genes, of which 4280 encode proteins.18 Analysis of the BRIC-21 RNA-seq data resulted in identification of 293 total genes whose expression differed significantly in FL vs. GC samples, representing ~6.8% of the protein-coding genome. Of these genes, 177 were significantly higher in FL samples, and 116 were significantly higher in GC samples. These data are summarized in Supplemental Table S1. Analysis of the BRIC-23 RNA-seq data resulted in identification of 255 total genes whose expression differed significantly in FL vs. GC samples, representing ~6.0% of the protein-coding transcriptome. Of these genes, 163 were significantly higher in FL samples, and 92 were significantly higher in GC samples. These data are summarized in Supplemental Table S2.
We reasoned that comparison of the transcriptome datasets from the BRIC-21 and BRIC-23 experiments would identify genes that were significantly up- or down-regulated in both missions, thus defining genes whose expression was consistently altered in response to spaceflight. A comparison of the datasets obtained from the BRIC-21 and BRIC-23 experiments is depicted graphically as Venn diagrams (Fig. 2). A total of 91 genes were significantly differentially expressed in both experiments. Fifty-five of the shared genes exhibited higher transcript levels in FL samples and 36 genes showed higher transcript levels in GC samples (Fig. 2).
The finding that only ~1/3 of the significantly differentially expressed transcripts were shared in both the BRIC-21 and BRIC-23 missions indicated a substantial amount of between-experiment variation. What could be the source of this variation? While we attempted to keep discrepancies between the two experiments to a minimum, one notable difference between the BRIC-21 and BRIC-23 experiments was the difference in incubation times (25 vs. 36 h), which may have contributed to the ~2/3 discordance in the two datasets. Unfortunately, BRIC-PDFU hardware does not allow direct measurement of growth to be determined in situ during spaceflight. However, because the transcription of genes encoding fundamental growth processes (e.g., replication, transcription, translation) exhibits a positive correlation with growth rate,19 as a proxy for growth rate we compared the fold changes for transcripts involved in DNA replication (dnaE), RNA transcription (rpoABCD), and protein synthesis (rps and rpl genes encoding small and large ribosomal subunit proteins). Neither limma or DESeq2 identified a significant difference in the transcript expression levels for these genes in either the BRIC-21 or BRIC-23 datasets, suggesting that the FL and GC samples were exhibiting similar growth rates at the time of harvest for RNA extraction.
In pre-flight ground-based experiments we determined that under ISS ambient temperature (~23 °C), 25 and 36 h of incubation corresponded to late-exponential phase and the “transition state” between exponential and stationary-phase growth, respectively.20 Entrance into the transition state in B. subtilis results in the transcriptional activation of nearly 300 genes which comprise a regulon under control of the AbrB protein;21 we reasoned that comparison of AbrB-controlled transcripts between the BRIC-21 and BRIC-23 datasets might provide insight into the growth phase of these two populations. We found that 60 and 57 AbrB-dependent transcripts were significantly altered in the BRIC-21 and BRIC-23 datasets, respectively, and that 27 transcripts were significantly altered in both experiments (Fig. 3). From this analysis it therefore appeared that cultures in both experiments were at or near the transition phase of growth.
Genes upregulated in FL samples
The 55 genes whose expression was significantly upregulated in FL samples from both BRIC-21 and BRIC-23 are listed in Table 1. They are arranged according to their BSU locus tag, i.e., in the order that they are located on the B. subtilis 168 chromosome map. Examination of the data revealed upregulation of blocks of genes associated with particular phenotypes in B. subtilis:
-
(i)
Biofilm formation. Under particular environmental conditions, various microorganisms can produce biofilms consisting of cells embedded in a matrix of extracellular polymeric substances (EPS) consisting of polysaccharides, proteins, nucleic acids, and lipids.22 It has been reported that spaceflight exposure promoted biofilm formation in Pseudomonas aeruginosa23 and Escherichia coli,24 and promoted invasive growth of the yeasts Saccharomyces cerevisiae25 and Candida albicans.26 Bacterial biofilms have been implicated in contamination and biofouling of potable water systems in long-duration space habitats, and data from spaceflight have documented biofilms containing Bacillus spp. in the Space Shuttle water system.27 Laboratory strains of B. subtilis such as strain 168 and its descendants do not form robust biofilms due to mutations that have accumulated during their domestication,28 and biofilm formation was not noted in FL or GC samples from BRIC-21 or BRIC-23. Nonetheless, numerous biofilm-related genes were observed to be significantly upregulated in both BRIC-21 and BRIC-23 FL samples, including genes of the srfAA-AB-AC-AD-ycxA operon, which encodes the pathway for production of the cyclic lipopeptide surfactin29,30 and genes of the tapA-sipW-tasA operon encoding the major protein matrix component of biofilms31 (Table 1). B. subtilis biofilms also contain as a major component the exopolysaccharide poly-N-acetylglucosamine,32 produced in a biosynthetic pathway encoded by the epsABCDEFGHIJKLMNO operon. Our analysis found that only the epsB, epsC, and epsI genes were significantly upregulated in FL samples of both BRIC-21 and BRIC-23 (Table 1). However, closer examination of the datasets revealed that in the BRIC-23 experiment, 12 of the 15 eps genes (epsABCDEFGHIJKL) were significantly upregulated in FL samples (Supplemental Table S2). In the BRIC-21 experiment, these 12 genes also displayed a significant upregulation in FL samples (p < 0.01), but only three of these genes (epsB, espC, and epsI) met our >2-fold change cutoff, while the remaining nine genes exhibited fold changes slightly below the cutoff (Supplemental Table S1). The last three genes in the eps operon, epsM, epsN, and epsO, were found not to be significantly differentially expressed in FL samples from either mission. In addition, examination of the BRIC-21 dataset revealed that the bslA, ycdA, and luxS genes, also involved in swarming motility and biofilm formation,33,34 were significantly upregulated in FL samples (Supplemental Table S1).
-
(ii)
Biotin biosynthesis. Biotin (vitamin H) is an essential cofactor for enzymes such as acetyl-CoA carboxylase and pyruvate carboxylase, which are important in fatty acid metabolism and central metabolism, respectively.35 Production of biotin from pimelic acid is accomplished by the gene products of the bioWAFDBI biosynthetic operon. Examination of the data from Table 1 revealed significant upregulation in FL samples of the bioW, bioA, bioD, and bioB genes in both the BRIC-21 and BRIC-23 missions, as well as upregulation of the bioY gene encoding the energy coupling factor (ECF) transporter biotin-specific S-protein (Table 1). In addition, inspection of the BRIC-21 dataset showed significant upregulation in FL samples of the bioF and bioI biotin biosynthetic genes, the yuiG (bioYB) gene encoding a putative second biotin-specific ECF transporter S-protein, the yhfT gene whose product is involved in surfactin production, and the yhfS encoding a putative acetyl-CoA C-acetyltransferase (Supplemental Table S1); along with the bio genes, these additional genes belong to a regulon under control of a repressor named BirA,36 thus are likely upregulated as a block.
-
(iii)
Siderophores. Iron is an important cofactor for several enzymes, but is only sparingly soluble in most aerobic environments.37 To acquire iron, most microorganisms produce and excrete siderophores which bind iron with high affinity for subsequent import by specific transport systems. B. subtilis produces the siderophore bacillibactin (2,3-dihydroxybenzoate) encoded by the besA-dhbACEBF operon.38 FL samples from both BRIC-21 and BRIC-23 were found to significantly upregulate the besA and dhbC transcripts (Table 1), and subsequent examination of the BRIC-21 data revealed that the entire besA-dhbACEBF operon was significantly upregulated in FL samples (Supplementary Table S1). Another siderophore produced by B. subtilis called pulcherriminic acid or pulcherrimin is encoded by two small operons, yvmC-cypX and pchR-yvmA.39 The yvmC and pchR genes were found to be significantly upregulated in FL samples of both BRIC-21 and BRIC-23 (Table 1), and further inspection revealed that both the yvmC-cypX and pchR-yvmA operons were significantly upregulated in FL samples from BRIC-21 (Supplemental Table S1).
-
(iv)
Arginine biosynthesis. The amino acid arginine is produced in a pathway encoded by the argCJBD-carAB-argF operon which converts glutamate to citrulline, and the argGH operon, which converts citrulline to arginine.40 The argGH operon was found to be significantly upregulated in both BRIC-21 and BRIC-23 flight samples. Closer examination of the datasets revealed significant upregulation of argCJBD-carAB-argF operon transcripts in BRIC-21 FL samples (Supplemental Table S1) and significant upregulation of the arginyl-tRNA genes trnJ-Arg and trnE-Arg in BRIC-23 FL samples (Supplemental Table S2).
-
(v)
Manganese transport. The major manganese ABC-type transporter in B. subtilis, encoded by the mntABCD operon, was found to be strongly upregulated in both BRIC-21 and BRIC-23 FL samples (Table 1). In addition, closer examination of the datasets revealed that the manganese-proton symporter mntH was significantly upregulated in BRIC-23 FL samples (Supplementary Table S2). Paradoxically, in BRIC-21 FL samples we also observed significant upregulation of the ydfM gene, encoding a putative Mn(II) efflux pump (Supplementary Table S1).
-
(vi)
Resistance and toxin genes. A number of genes encoding resistance and toxic functions were observed to be upregulated in FL samples of both BRIC-21 and BRIC-23, including: ybfO, encoding a putative erythromycin esterase; the bmrCD operon, encoding a multidrug efflux transporter; yvqJ, encoding a putative macrolide-efflux protein; cadA, encoding a cadmium efflux pump; and sdpC, which encodes a lytic toxin (Table 1). At first glance, increase in the aforementioned transcripts might suggest that FL samples could be exhibiting a higher degree of resistance to antibiotics or toxic compounds. However, in a separate publication we reported that replicate samples from the BRIC-21 experiment were exposed post-flight to a battery of 72 antibiotics and growth inhibitors, and no significant difference in resistance levels was found in FL vs. GC samples.16
-
(vii)
Inhibitors of sporulation initiation. Sporulation in B. subtilis is triggered when cells sense the depletion of nutrients in their environment. Three genes [sivA, sivB (bslA), and sivC; siv for sporulation-inhibitory vegetative genes] have recently been described which actively inhibit the initiation of sporulation when B. subtilis is growing in the presence of sufficient nutrients.41 Both the sivA and sivC genes were found to be significantly upregulated in FL samples of both BRIC-21 and BRIC-23 (Table 1), and as mentioned above, the sivB (bslA) gene associated with biofilm formation was upregulated in BRIC-21 FL samples (Supplemental Table S1).
Additional genes upregulated in FL samples
Examination of Table 1 revealed a number of additional genes upregulated in both BRIC-21 and BRIC-23 FL samples for which no clear phenotypic consequence could be discerned. First, a number of genes encoding products of unknown function were upregulated, including ybdO, yczC, ycxA, the yrkEFHIJ operon (Table 1). Second, genes encoding transporters for glycerol-3-phosphate (glpT), mannose (manP), and oligopeptides (appA/1 and appA/2) were upregulated, as were genes encoding an extracellular lipase (lipA), proline dehydrogenase (putB), aconitase (citB), a single methyl-accepting chemotaxis protein (tlpA), urate oxidase (pucL), and a secreted peptide controlling LiaR-LiaS activity (yydF) (Table 1). Each of these belong to its own cohort of genes devoted to different functions in B. subtilis, but expression of the other members of the cohort were not significantly altered. As an example, the threonyl transfer RNA gene trnD-Thr is transcribed as part of a 16-tRNA gene operon located just downstream from the ribosomal RNA rrnD gene cluster in B. subtilis,42 but it is unclear why only this transcript, and not the entire operon, was significantly upregulated. Third, two genes encoded by the prophage SPβ (yomK and bhlA) were upregulated in FL samples (Table 1), but numerous SPβ-related genes were also upregulated in GC samples (Table 2), so no coherent pattern of gene expression could be ascertained.
Genes upregulated in GC samples
The 36 genes whose expression was significantly upregulated in GC samples from both BRIC-21 and BRIC-23 are listed in Table 2. They are arranged according to their BSU locus tag, i.e., in the order that they are located on the B. subtilis 168 chromosome. Examination of the data revealed upregulation of several blocks of genes associated with the response of B. subtilis to oxygen limitation. A previous study43 reported that the global response of the B. subtilis transcriptome to strict anaerobiosis resulted in the induction or repression of hundreds of genes involved in a variety of cell functions including carbon metabolism, electron transport, iron uptake, antibiotic production, and stress responses. Our data revealed that only a subset of the entire anaerobic regulon was activated, indicating that cells in GC samples underwent only partial oxygen deprivation, and these genes are described below.
-
(i)
Fermentation. TSYG medium contains the fermentable sugar glucose, and B. subtilis is capable of mixed-acid fermentation using separate pathways for production of the end products acetate, lactate, acetoin, 2,3-butanediol, and ethanol.44 In GC samples from both the BRIC-21 and BRIC-23 missions we observed significant upregulation of the ldh-lctP operon for fermentation of lactate from pyruvate,45,46 as well as the alsSD operon for fermentation of acetoin from pyruvate47 and the bdhA gene for fermentation of 2,3-butanediol from acetoin48 (Table 2). Interestingly, we did not note any significant changes in the expression of genes involved in acetate or ethanol fermentation in our samples (Supplemental Tables S1 and S2).
-
(ii)
Anaerobic respiration. Growth under oxygen-limiting conditions results in two major modifications of the B. subtilis respiratory electron transport chain. First, oxygen depletion activates synthesis of the high-affinity cytochrome bd ubiquinol oxidase encoded by the cydABCD operon.49 We observed that the cydC and cydD genes were significantly upregulated in GC samples of BRIC-21 and BRIC-23 (Table 2), and further examination of the datasets revealed that the entire cydABCD operon was strongly upregulated in GC samples from BRIC-21 (Supplemental Table S1). Second, anaerobiosis results in activation of genes responsible for utilizing nitrate and nitrite as alternative terminal electron acceptors.44 In B. subtilis GC samples we noted significant upregulation of the narGHJI operon encoding nitrate reductase, the nasDE operon encoding nitrite reductase, and the narK-fnr operon encoding the NarK nitrite extrusion protein and the Fnr regulator of anaerobic gene expression (Table 2). It is interesting to note that the ywcJ transcript, encoding a putative nitrite transporter previously identified as a member of the B. subtilis anaerobic regulon,43 was also upregulated in GC samples. Because TSYG medium does not provide a significant source of nitrate or nitrite, it is likely that although expression of the nar and nas genes was induced by oxygen limitation, they did not serve a useful physiological function for cells in the GC samples.
-
(iii)
Subtilosin production. Further indication for oxygen limitation in BRIC-21 and BRIC-23 GC samples was evidenced by induction of the genes encoding the antilisterial antibiotic subtilosin A, which was previously shown to be induced by anaerobiosis43 (Table 2). The pathway for subtilosin A biosynthesis is encoded by the sboAXalbABCDEFG operon,50 and in both BRIC-21 and BRIC-23 GC samples the sboA, sboX, albE, albF, and albG transcripts were significantly upregulated (Table 2). Further inspection of the datasets revealed that the entire sboAXalbABCDEFG operon was significantly upregulated in BRIC-21 GC samples (Supplemental Table S1).
-
(iv)
Other genes belonging to the anaerobic regulon. Two genes encoding products of unknown or only putatively annotated function (ydhE and ytkA) were seen to be upregulated in GC samples of both BRIC-21 and BRIC-23 (Table 2). Activation of transcripts for these genes under anaerobic conditions was noted previously.43
-
(v)
Genes regulating anaerobiosis. The response to oxygen limitation in B. subtilis has been well studied and is controlled by a complex regulatory hierarchy.51 Oxygen limitation is sensed by the membrane sensor kinase ResE and transmitted to the ResD response regulator, encoded by the last two genes of the resABCDE operon. ResD activates expression of a large set of genes and operons including its own (resABCDE), cydABCD, nasBCDEF, and sboAXalbABCDEFG. One of the genes activated by ResD is fnr, itself a regulator of anaerobic gene expression. Fnr activates its own expression (narK-fnr), as well as that of narGHJI and another regulator encoded by the arfM gene. In addition, changes in cell physiology associated with the switch to oxygen limitation activate a transcriptional regulator called AlsR, which activates expression of alsSD directly47,52 and bdhA indirectly.53 Meanwhile, the regulatory protein Rex senses changes in the NAD+/NADH ratio brought on by oxygen limitation and responds by activating a number of genes including the cydABCD and ldh-lctP operons.51 We were prompted by these observations to search for regulators of anaerobic gene expression in the datasets, and found significant upregulation of the arfM gene in BRIC-21 (Supplemental Table S1), the resD and resE genes in BRIC-23 (Supplemental Table S2), and the narK-fnr operon in both BRIC-21 and BRIC-23 GC samples (Table 2).
-
(vi)
Phospholipid desaturase. Most prokaryotes regulate membrane fluidity in part by controlling the degree of saturation of membrane phospholipids. B. subtilis accomplishes this using a fatty acid desaturase encoded by des, the first gene in the des-desKR operon.54,55 Expression of des is activated by exposure to low temperature, mediated through a two-component system composed of a membrane-bound sensor kinase and a response regulator encoded by the desK and desR genes, respectively.55 Transcription of des and desK were seen to be significantly upregulated in GC samples of both BRIC-21 and BRIC-23 (Table 2), and closer examination of the datasets revealed that the entire des-desKR operon was upregulated in BRIC-23 GC samples (Supplemental Table S2). While the biological significance of this observation is at present unclear, it is interesting to note that in previous work we reported that des-desKR transcription was also activated by exposure of B. subtilis to low atmospheric pressure.56
-
(vii)
Metabolism of a cysteine analog. A pathway in B. subtilis was recently described which can convert the cysteine analog S-methyl-cysteine directly to cysteine as a sulfur source, encoded by the snaA-tcyJKLMN-cmoOIJ-ribR-sndA-ytnM operon.57 We observed that three genes from this operon (cmoJ, tcyL, and tcyN) were upregulated in GC samples from both BRIC-21 and BRIC-23 (Table 2). Further examination of the datasets revealed additional genes of the operon significantly upregulated in BRIC-21 (snaA, tcyJ, tcyK, tcyM, cmoI, and ribR) and BRIC-23 (cmoO, sndA, and ytnM) GC samples (Supplemental Tables S1 and S2).
Additional genes upregulated in GC samples
Examination of Table 2 revealed a number of genes significantly up-regulated in GC samples from both BRIC-21 and BRIC-23 for which no clear phenotypic consequence could be discerned. First, three genes encoding products of unknown function (yhfH, ytkA, and ywhL) were upregulated in GC samples (Table 2). Second, upregulation of genes encoding a putative macrolide glycosyltransferase (ydhE), a component of the twin-arginine translocase (tatAC), and a transcriptional antiterminator (sacT) (Table 2), but these genes form parts of larger cohorts of genes which were not themselves significantly upregulated. Third, transcripts for a number of genes encoded by the SPβ prophage (yoqM, yopU, yopT, yonN, yomL, and youB) were induced, the function of most of which are unknown.
Discussion
In the present study, we analyzed the effects of spaceflight on the Bacillus subtilis transcriptome in two separate spaceflight experiments designated BRIC-21 and BRIC-23. This is the first spaceflight bacterial transcriptome reported from a Gram-positive bacterium, and the first transcriptome study of two separate spaceflight missions using the same bacterial strain, growth media, and hardware. The results uncovered several differences between FL and GC samples that were exhibited in both BRIC-21 and BRIC-23 missions.
Biofilm-associated transcripts
The observation that transcripts associated with biofilm formation were significantly upregulated both in BRIC-21 and BRIC-23 FL samples is in agreement with the results from prior spaceflight experiments using several different microbes indicating enhanced biofilm production in spaceflight.23,24,25,26,27 Although the domesticated laboratory strain B. subtilis 168 does not form robust biofilms, we propose that biofilm formation in spaceflight-grown B. subtilis could readily be studied in detail by using undomesticated biofilm-producing strains such as NCIB 3610 and its derivatives.58,59
Oxygen limitation-associated transcripts
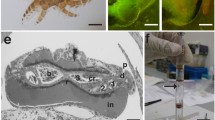
We noted significant upregulation in GC samples of transcripts associated with fermentation, anaerobic respiration, and subtilosin production, and upregulation in FL samples of transcripts associated with siderophore production. These observations indicate that FL and GC samples were experiencing different degrees of oxygen availability in the two experiments. How could exposure to microgravity vs. 1 xg result in different levels of oxygen available to the liquid cultures? First, we noted that during deintegration of frozen BRIC-21 samples from PDFUs, FL samples incubated in microgravity had assumed a toroidal shape (Fig. 4), while GC samples formed a disk-shaped layer in the bottom of the Petri dish as expected. By assuming a toroidal configuration in microgravity, the liquid FL cultures may present a greater surface area to the air phase than in GC samples. Second, under the influence of gravity, cells in GC samples would tend to sediment toward the bottom of the Petri dish, away from the liquid/air interface, thus limiting their access to oxygen. Third, under the influence of gravity, dissolved gases, nutrients, and waste products are transported in liquids by convection and diffusion; in microgravity, convection is negated and transport becomes dominated by diffusive processes.9 Oxygen transfer by convection would be expected to be greater in GC than FL cultures, but this effect appears to be outweighed by the first two factors. Unfortunately, because B. subtilis prefers an oxygen-rich environment for optimum growth, inclusion of an air-containing headspace in the PDFU cultures is unavoidable. In order to address this hardware limitation, we are currently working on the design and construction of PDFU inserts that will maintain a constant geometry of liquid and air space both in microgravity and in 1-xg controls.
Nutrient utilization-associated transcripts
As stated above, convection ceases in microgravity and the transport of nutrients or waste products through liquid media becomes diffusion-limited.9 We observed significantly upregulated transcripts for biotin and arginine biosynthetic genes in FL samples, indicating that availability of these two nutrients differed in FL vs. GC cultures. However, we did not note significant differences between FL and GC samples in the expression of any other biosynthetic pathways, nor was there a significant difference in the final growth yield of cells between FL and GC cultures.17 Therefore, it does not appear that exposure to microgravity led to a generalized nutrient deficiency in FL samples.
In our analysis we treated the BRIC-21 and BRIC-23 missions as replicate experiments, but it should be noted that the two experiments differed in incubation times, thus at harvest the BRIC-21 and BRIC-23 samples were at slightly different growth phases (late-exponential and transition-phase, respectively). It is therefore to be expected that the resulting transcriptome profiles would differ somewhat, due to the substantial reorganization of global gene expression which occurs during the transition from exponential to stationary-phase growth in B. subtilis.60 In addition, it should be kept in mind that measuring the transcriptome captures only one aspect of physiology; it does not take into account regulatory controls exerted at the level of numerous posttranscriptional processes (translation, protein processing and modification, metabolic regulation of enzyme activity, assembly of subcellular structures, etc.) which must take place in order for a microbe to manifest its final phenotype. With these caveats in mind, combining the datasets from the two experiments allowed us to perform a more robust analysis and led to the identification of a common set of genes that were consistently differentially expressed between FL and GC samples in both experiments. These genes, particularly those involved in biofilm formation, will be interesting candidates for future study.
Methods
Bacterial strain, media, and growth conditions
The strain used in this study was Bacillus subtilis subsp. subtilis strain 168 (trpC2) from our laboratory stock collection. Medium used throughout was Trypticase Soy Yeast Extract (TSY) medium consisting of (g/L): tryptone, 15; soytone, 5; NaCl, 5; yeast extract, 3; K2HPO4, 2.5; glucose, 2.5; final pH 7. For semisolid plates, agar was added to TSY to a final concentration of 15.0 g/L. Glycerol was added to TSY liquid medium to 10% (v/v) final concentration, resulting in TSYG medium. B. subtilis spores were routinely prepared by cultivation in liquid Schaeffer sporulation medium61 at 37 °C with vigorous aeration. The culture was harvested when phase-contrast microscopic examination revealed that it consisted of >90% free spores, usually after 3–4 days of incubation. Spores were purified by lysozyme treatment, buffer washing, and heat shock (80 °C, 10 min) as described previously,62 determined by phase-contrast microscopy to be >99% free of cell debris and unsporulated cells, and stored at 4 °C in deionized water. The spore suspension was heat-activated (65 °C, 20 min) before use. From a working suspension (108/mL) of spores in water, aliquots of 0.1 mL (~107 CFU) were applied to the bottoms of sterile 60-mm diameter Petri dishes (Falcon Cat. No. 1007, Fisher Scientific) and air-dried for 48–72 h at room temperature protected from light. Samples were integrated into Biological Research in Canister Dual-Chamber Petri Dish Fixation Units (BRIC-PDFU) spaceflight hardware using aseptic technique as described in detail previously.20
BRIC-PDFU hardware
Biological Research in Canister (BRIC)-Petri Dish Fixation Unit (PDFU) hardware has been described in detail previously.63,64 Each BRIC canister enclosed 5 PDFUs, and each PDFU contained a space to accommodate the bottom half of a single 60-mm diameter Petri dish to which air-dried spores were deposited. A separate syringe compartment contained TSYG liquid medium. Growth is initiated by injection of 8.5 mL of TSYG medium into the Petri dish compartment, leaving a headspace of ~18 mL of ambient air which exchanges with the static culture via passive diffusion. The headspace is the only source of additional oxygen, and because the system is hermetically sealed, O2 is consumed and CO2 accumulates in the culture. Once initiated, there is no mechanism to access cultures for removal of samples during flight.
Experimental timeline
The schedules of pre-flight, flight, and post-flight activities are described in detail elsewhere for the BRIC-2116,17 and BRIC-2320 experiments. Experimental details for the two FL experiments were essentially identical in terms of hardware (BRIC-PDFU), inoculum size (107 spores), medium used (TSYG), and media volumes injected (8.5 mL). Slight differences in ambient ISS temperature were recorded inside the BRIC-21 (average ~22.8 °C) and BRIC-23 (average ~22.3 °C) canisters. The only notable difference in the two experiments was the time of incubation for BRIC-21 and BRIC-23 samples, 25 h and 36 h, respectively. In both missions, growth was terminated by transferring BRIC canisters to the −80 °C freezer onboard the ISS. In both cases, samples remained solidly frozen until returned to the lab. Asynchronous GC samples were treated identically to FL samples in terms of hardware, configuration, schedule, temperature profiles, pre- and post-flight handling.16,17,20
BRIC-21 RNA isolation, processing, and sequencing
Frozen FL and GC samples (n = 3) were partially thawed at room temperature. Partially thawed samples were transferred into 15-mL conical tubes, and placed on ice until completely thawed. Cells were recovered by centrifugation (7000 × g, 20 min, 0 °C) in a benchtop centrifuge. Supernatants were transferred into sterile 15-mL conical tubes and stored at −70 °C. Cell pellets were immediately processed for total RNA extraction and treatment with RNase-free DNase using the RiboPureTM RNA Purification Kit (Thermo Fisher Scientific Inc, Waltham, MA) following the manufacture’s protocols. RNA samples were quantified using a Qubit fluorometer (Invitrogen, Thermo Fisher Scientific Inc, Waltham, MA) and quality evaluated using the RNA 6000 Nano Chip on an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA). RNA Integrity Number (RIN) values ranged from 9.8 to 10.0, indicating high-quality total RNA preparations suitable for further processing. Samples were sent to Hudson Alpha Institute for Biotechnology (Huntsville, AL, USA) for ribosomal RNA depletion, library preparation, and 100-nt paired-end sequencing on an Illumina 2500 instrument.
BRIC-23 RNA isolation, processing, and sequencing
BRIC-23 sample processing was performed by the GeneLab Sample Processing Lab (NASA Ames Research Center, Mountain View, CA); detailed protocols are described in the BRIC-23 GeneLab dataset GLDS-138 (https://genelab-data.ndc.nasa.gov/genelab/accession/GLDS-138/). Briefly, thawed FL and GC samples (n = 9) were centrifuged at 16,000 × g for 5 min at 0 °C. Total RNA extraction and RNase-free DNase treatment was also performed using the RiboPureTM RNA Purification Kit (Thermo Fisher Scientific Inc, Waltham, MA) following the manufacture’s protocol. RNA concentrations were measured using a Qubit 3.0 Fluorometer (Thermo Fisher Scientific, Waltham, MA), and RNA quality was assessed on an Agilent 2100 Bioanalyzer (Agilent, Santa Clara, CA). Ribosomal RNA depletion was performed using the Ribo-Zero rRNA Removal Kit for Gram-positive bacteria (Illumina) and sample cDNA libraries were synthesized using KAPA RNA HyperPrep reagents (Roche) and amplified with 10 PCR cycles. Library sequencing was performed on an Illumina HiSeq 4000 platform.
Read processing, alignment, and quantification
Quality control, mapping, and gene level quantification of Illumina sequences were performed using the Galaxy suite available through the University of Florida’s High-Performance Research Computing Center. The first 12 bases were trimmed off all reads using FASTQ Trimmer v0.01465 to remove random hexamer primers, and read quality of the resulting sequences were checked using the FastQC program.66 Corresponding paired-end read files were mapped to the Bacillus subtilis strain 168 genome [National Center for Biotechnology Information (NCBI) RefSeq accession number NC_000964.3] using Bowtie2 v2.3.2.67 Mapping quality was evaluated using SAMStat68 followed by gene level quantification using htseq-count v0.6.1.69 Gene counts for BRIC-21 and BRIC-23 were separated into separate count matrices for differential expression analysis.
Differential expression and functional analyses
To reduce false positive results, two Bioconductor packages, limma v3.32.1070,71 and DESeq2 v1.16.1,72 were used to determine differential expression. These two packages have been shown to exhibit consistent detection precision in comparisons using a large or very small number of replicates73 making these packages suitable for our BRIC-21 (n = 3) and BRIC-23 (n = 9) comparisons. Per limma recommendation, genes with a sum of less than nine counts across all samples were removed before analysis. For limma analyses, filtered genes were normalized between samples using TMM normalization and differential expression analysis was conducted using the built-in voom transformation.74 Because DESeq2 requires non-normalized count data, analysis was performed on the raw filtered genes. P values from both analyses were corrected for multiple testing using the Benjamini–Hochberg method.75 To be considered differentially expressed, genes had to have a >2-fold difference with a P value < 0.01 between FL and GC expression using both limma and DESeq2 methods. Genes meeting these criteria were annotated using the NCBI database and the Subtiwiki web server.76 Functional analysis was conducted using the Search Tool for the Retrieval of Interacting Genes/Proteins (STRING) protein network database.77 STRING protein–protein interaction networks for up- and downregulated genes were generated separately, and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways were found using the built-in enrichment tools. To check the variability between all the BRIC-21 and BRIC-23 datasets, a PCA was performed in the statistical environment R v3.4.4. Gene loading scores and sample eigenvalues for each principal component were calculated using the princomp function in the R package stats v3.4.4. Sample eigenvalues for the first two principal components were plotted in R using the default plotting function.
Data availability
The RNA-seq datasets have been deposited in the NASA GeneLab Data System under the accession numbers GLDS-185 (BRIC-21) and GLDS-138 (BRIC-23).
References
Mora, M. et al. Microorganisms in confined habitats: microbial monitoring and control of intensive care units, operating rooms, cleanrooms and the International Space Station. Front. Microbiol. 7, 20 (2016).
Pietsch, J. et al. The effects of weightlessness on the human organism and mammalian cells. Curr. Mol. Med. 11, 350–364 (2011).
Lychakov, D. V. Behavioural and functional vestibular disorders after space flight: 2. Fish, amphibians and birds. J. Evol. Biochem. Physiol. 52, 1–16 (2016).
Vandenbrink, J. P. & Kiss, J. Z. Space, the final frontier: a critical review of recent experiments performed in microgravity. Plant Sci. 243, 115–119 (2016).
Rosenzweig, J. A. et al. Spaceflight and modeled microgravity effects on microbial growth and virulence. Appl. Microbiol Biotechnol. 85, 885–891 (2010).
Rosenzweig, J. A., Ahmed, S., Eunson, J. & Chopra, A. K. Low-shear force associated with modeled microgravity and spaceflight does not similarly impact the virulence of notable bacterial pathogens. Appl. Microbiol Biotechnol. https://doi.org/10.1007/s00253-014-6025-8 (2014).
Taylor, P. W. Impact of space flight on bacterial virulence and antibiotic susceptibility. Infect. Drug Resist 8, 249–262 (2015).
National Research Council. Recapturing a Future for Space Exploration: Life and Physical Sciences for a New Era, 442 (National Academies Press: Washington, DC, 2011).
Horneck, G., Klaus, D. M. & Mancinelli, R. L. Space microbiology. Microbiol Mol. Biol. Rev. 74, 121–156 (2010).
Zea, L. et al. A molecular genetic basis explaining altered bacterial behavior in space. PLoS One 11, e0164359 (2016).
Mastroleo, F. et al. Experimental design and environmental parameters affect Rhodospirillum rubrum S1H response to space flight. ISME J. 3, 1402–1419 (2009).
Huang, B., Liu, N., Rong, X., Ruan, J. & Huang, Y. Effects of simulated microgravity and spaceflight on morphological differentiation and secondary metabolism of Streptomyces coelicolor A3(2). Appl. Microbiol. Biotechnol. 99, 4409–4422 (2015).
Crabbé, A. et al. Transcriptional and proteomic responses of Pseudomonas aeruginosa PAO1 to spaceflight conditions involve Hfq regulation and reveal a role for oxygen. Appl. Environ. Microbiol 77, 1221–1230 (2011).
Wilson, J. W. et al. Space flight alters bacterial gene expression and virulence and reveals a role for global regulator Hfq. Proc. Natl Acad. Sci. USA 104, 16299–16304 (2007).
Altman, N. & Krzywinski, M. Sources of variation. Nat. Methods 12, 5–6 (2015).
Morrison, M. D., Fajardo-Cavazos, P. & Nicholson, W. L. Cultivation in space flight produces minimal alterations in the susceptibility of Bacillus subtilis cells to 72 different antibiotics and growth-inhibiting compounds. Appl. Environ. Microbiol. 83, e01584–01517 (2017).
Fajardo-Cavazos, P., Leehan, J. D. & Nicholson, W. L. Alterations in the spectrum of spontaneous rifampicin-resistance mutations in the Bacillus subtilis rpoB gene after cultivation in the human spaceflight environment. Front. Microbiol. 9, 192 (2018).
Markowitz, V. M. et al. IMG: the integrated microbial genomes database and comparative analysis system. Nucleic Acids Res. 40, D115–D122 (2012).
Matsumoto, Y., Murakami, Y., Tsuru, S., Ying, B. W. & Yomo, T. Growth rate-coordinated transcriptome reorganization in bacteria. BMC Genom. 14, 10 (2013).
Fajardo-Cavazos, P. & Nicholson, W. L. Establishing standard protocols for bacterial culture in Biological Research in Canister (BRIC) hardware. Gravit. Space Res. 4, 58–69 (2016).
Banse, A. V., Chastanet, A., Rahn-Lee, L., Hobbs, E. C. & Losick, R. Parallel pathways of repression and antirepression governing the transition to stationary phase in Bacillus subtilis. Proc. Natl Acad. Sci. USA 105, 15547–15552 (2008).
Flemming, H. C. & Wingender, J. The biofilm matrix. Nat. Rev. Microbiol. 8, 623–633 (2010).
Kim, W. et al. Spaceflight promotes biofilm formation by Pseudomonas aeruginosa. PLoS One 8, e62437 (2013).
Zea, L. et al. Phenotypic changes exhibited by E. coli cultured in space. Front. Microbiol. 8, 12 (2017).
Van Mulders, S. E. et al. The influence of microgravity on invasive growth in Saccharomyces cerevisiae. Astrobiology 11, 45–55 (2011).
Crabbé, A. et al. Spaceflight enhances cell aggregation and random budding in Candida albicans. Plos One 8, 24 (2013).
Koenig, D. W. & Pierson, D. L. Microbiology of the space shuttle water system. Water Sci. Technol. 35, 59–64 (1997).
McLoon, A. L., Guttenplan, S. B., Kearns, D. B., Kolter, R. & Losick, R. Tracing the domestication of a biofilm-forming bacterium. J. Bacteriol. 193, 2027–2034 (2011).
Vlamakis, H., Chai, Y. R., Beauregard, P., Losick, R. & Kolter, R. Sticking together: building a biofilm the Bacillus subtilis way. Nat. Rev. Microbiol. 11, 157–168 (2013).
Lopez, D., Fischbach, M. A., Chu, F., Losick, R. & Kolter, R. Structurally diverse natural products that cause potassium leakage trigger multicellularity in Bacillus subtilis. Proc. Natl Acad. Sci. USA 106, 280–285 (2009).
Driks, A. Tapping into the biofilm: insights into assembly and disassembly of a novel amyloid fibre in Bacillus subtilis. Mol. Microbiol 80, 1133–1136 (2011).
Roux, D. et al. Identification of poly-N-acetylglucosamine as a major polysaccharide component of the Bacillus subtilis biofilm matrix. J. Biol. Chem. 290, 19261–19272 (2015).
Kobayashi, K. & Iwano, M. BslA(YuaB) forms a hydrophobic layer on the surface of Bacillus subtilis biofilms. Mol. Microbiol 85, 51–66 (2012).
Lombardia, E., Rovetto, A. J., Arabolaza, A. L. & Grau, R. R. A LuxS-dependent cell-to-cell language regulates social behavior and development in Bacillus subtilis. J. Bacteriol. 188, 4442–4452 (2006).
Perkins, J. B. & Pero, J. in Bacillus subtilis and its Closest Relatives: from Genes to Cells (eds Sonenshein, A. L., Hoch, J. A., & Losick, R.), 271–286 (ASM Press, 2002).
Rodionov, D. A., Mironov, A. A. & Gelfand, M. S. Conservation of the biotin regulon and the BirA regulatory signal in Eubacteria and Archaea. Genome Res 12, 1507–1516 (2002).
Miethke, M. Molecular strategies of microbial iron assimilation: from high-affinity complexes to cofactor assembly systems. Metallomics 5, 15–28 (2013).
May, J. J., Wendrich, T. M. & Marahiel, M. A. The dhb operon of Bacillus subtilis encodes the biosynthetic template for the catecholic siderophore 2,3-dihydroxybenzoate-glycine-threonine trimeric ester bacillibactin. J. Biol. Chem. 276, 7209–7217 (2001).
Randazzo, P., Aubert-Frambourg, A., Guillot, A. & Auger, S. The MarR-like protein PchR (YvmB) regulates expression of genes involved in pulcherriminic acid biosynthesis and in the initiation of sporulation in Bacillus subtilis. BMC Microbiol. 16, 16 (2016).
Belitsky, B. R. in Bacillus subtilis and Its Closest Relatives: from Genes to Cells (eds Sonenshein, A. L., Hoch, J. A. & Losick, R.), 203–223 (ASM Press, 2002).
Garti-Levi, S., Eswara, A., Smith, Y., Fujita, M. & Ben-Yehuda, S. Novel modulators controlling entry into sporulation in Bacillus subtilis. J. Bacteriol. 195, 1475–1483 (2013).
Rudner, R. et al. Two transfer RNA gene clusters associated with ribosomal RNA operons rrnD and rrnE. Bacillus subtilis. J. Bacteriol. 175, 503–509 (1993).
Ye, R. et al. Global gene expression profiles of Bacillus subtilis grown under anaerobic conditions. J. Bacteriol. 182, 4458–4465 (2000).
Nakano, M. & Zuber, P. in Bacillus subtilis and Its Closest Relatives: from Genes to Cells (eds Sonenshein, A. L., Hoch, J. A., & Losick, R.), 393–404 (ASM Press, 2002).
Cruz Ramos, H. et al. Fermentative metabolism of Bacillus subtilis: physiology and regulation of gene expression. J. Bacteriol. 182, 3072–3080 (2000).
Marino, M., Hoffmann, T., Schmid, R., Mobitz, H. & Jahn, D. Changes in protein synthesis during the adaptation of Bacillus subtilis to anaerobic growth conditions. Microbiol. SGM 146, 97–105 (2000).
Renna, M. C., Najimudin, N., Winik, L. R. & Zahler, S. A. Regulation of the Bacillus subtilis alsS, alsD, and alsR genes involved in post-exponential-phase production of acetoin. J. Bacteriol. 175, 3863–3875 (1993).
Nicholson, W. L. The Bacillus subtilis ydjL (bdhA) gene encodes acetoin reductase/2,3-butanediol dehydrogenase. Appl. Environ. Microbiol 74, 6832–6838 (2008).
von Wachenfeldt, C. & Hederstedt, L. in Bacillus subtilis and Its Closest Relatives: from Genes to Cells (eds Sonenshein, A. L., Hoch, J. A., & Losick, R.), 163–179 (ASM Press, 2002).
Zheng, G. L., Yan, L. Z., Vederas, J. C. & Zuber, P. Genes of the sbo-alb locus of Bacillus subtilis are required for production of the antilisterial bacteriocin subtilosin. J. Bacteriol. 181, 7346–7355 (1999).
Härtig, E. & Jahn, D. Regulation of the anaerobic metabolism in Bacillus subtilis. Adv. Microb. Physiol. 61, 195–216 (2012).
Frädrich, C. et al. The transcription factor AlsR binds and regulates the promoter of the alsSD operon responsible for acetoin formation in Bacillus subtilis. J. Bacteriol. 194, 1100–1112 (2012).
Oliveira, R. R. & Nicholson, W. L. The LysR-type transcriptional regulator (LTTR) AlsR indirectly regulates expression of the Bacillus subtilis bdhA gene encoding 2,3-butanediol dehydrogenase. Appl. Microbiol Biotechnol. 97, 7307–7316 (2013).
Mansilla, M. C. & de Mendoza, D. The Bacillus subtilis desaturase: a model to understand phospholipid modification and temperature sensing. Arch. Microbiol. 183, 229–235 (2005).
Mendoza, D. Temperature sensing by membranes. Annu Rev. Microbiol 68, 101–116 (2014).
Fajardo-Cavazos, P. et al. Evolution of Bacillus subtilis to enhanced growth at low pressure: up-regulated transcription of des-desKR encoding the fatty acid desaturase system. Astrobiology 12, 258–270 (2012).
Chan, C.-M., Danchin, A., Marliere, P. & Sekowska, A. Paralogous metabolism: S-alkyl-cysteine degradation in Bacillus subtilis. Environ. Microbiol 16, 101–117 (2014).
Branda, S. S., Gonzalez-Pastor, J. E., Ben-Yehuda, S., Losick, R. & Kolter, R. Fruiting body formation by Bacillus subtilis. Proc. Natl Acad. Sci. USA 98, 11621–11626 (2001).
Konkol, M. A., Blair, K. M. & Kearns, D. B. Plasmid-encoded ComI inhibits competence in the ancestral 3610 strain of Bacillus subtilis. J. Bacteriol. 195, 4085–4093 (2013).
Phillips, Z. E. V. & Strauch, M. A. Bacillus subtilis sporulation and stationary phase gene expression. Cell Mol. Life Sci. 59, 392–402 (2002).
Schaeffer, P., Millet, J. & Aubert, J. P. Catabolic repression of bacterial sporulation. Proc. Natl Acad. Sci. USA 54, 704–711 (1965).
Nicholson, W. L. & Setlow, P. in Molecular Biological Methods for Bacillus (eds Harwood, C. R. & Cutting, S. M.), 391–450 (J. Wiley & Sons, 1990).
Paul, A.-L. et al. Spaceflight transcriptomes: unique responses to a novel environment. Astrobiology 12, 40–56 (2012).
Wells, B., McCray, R. H., Best, M. D. & Levine, H. G. A flight-rated Petri dish apparatus providing two stage fluid injection for aseptic biological investigations in space. in 31st International Conference on Environmental Systems 2001-01-2286 (SAE International, 2001).
Blankenberg, D. et al. Manipulation of FASTQ data with Galaxy. Bioinformatics 26, 1783–1785 (2010).
FastQC: a quality control tool for high throughput sequence data. v. 0.11.5. http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (Babraham Bioinformatics, 2010).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Lassmann, T., Hayashizaki, Y. & Daub, C. O. SAMStat: monitoring biases in next generation sequencing data. Bioinformatics 27, 130–131 (2011).
Anders, S., Pyl, P. T. & Huber, W. HTSeq–a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Liu, R. et al. Why weight? Modelling sample and observational level variability improves power in RNA-seq analyses. Nucleic Acids Res. 43, e97 (2015).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Seyednasrollah, F., Laiho, A. & Elo, L. L. Comparison of software packages for detecting differential expression in RNA-seq studies. Brief. Bioinform. 16, 59–70 (2015).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. Voom: precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Michna, R. H., Zhu, B., Mäder, U. & Stülke, J. SubtiWiki 2.0–an integrated database for the model organism Bacillus subtilis. Nucleic Acids Res. 44, D654–662 (2016).
Szklarczyk, D. et al. STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 43, D447–452 (2015).
Zhu, B. Y. & Stülke, J. SubtiWiki in 2018: from genes and proteins to functional network annotation of the model organism Bacillus subtilis. Nucleic Acids Res. 46, D743–D748 (2018).
Acknowledgements
The authors wish to thank the BRIC-21 and BRIC-23 payload development teams at NASA Kennedy Space Center (D. Dimapilis, A.D. Flowers, C. Grosse, J. Harp, D. Houze, H. Levine, G. Newsham, S. Manning-Roach, J. Richards, S. Richards, J. Smodell, and G. Washington) and the GeneLab Process Verification Team (V. Boyko, K. Chakravarty, S. Costes, H. Fogle, M. Dinh, J. Galazka, S. Gebre, S. Lai Polo, and S. Reinsch) at NASA Ames Research Center for their excellent technical assistance. This work was supported by the NASA Research Opportunities in Space Biology grant NNX14AT38G to W.L.N. and P.F.-C.
Author information
Authors and Affiliations
Contributions
W.L.N. and P.F.-C. conceived and designed the study. All authors performed the spaceflight experiments and RNA processing. M.D.M. and W.L.N. performed the data analysis. All authors contributed to writing the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Morrison, M.D., Fajardo-Cavazos, P. & Nicholson, W.L. Comparison of Bacillus subtilis transcriptome profiles from two separate missions to the International Space Station. npj Microgravity 5, 1 (2019). https://doi.org/10.1038/s41526-018-0061-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41526-018-0061-0
- Springer Nature Limited
This article is cited by
-
Whole transcriptome analysis highlights nutrient limitation of nitrogen cycle bacteria in simulated microgravity
npj Microgravity (2024)
-
Whole genome-scale assessment of gene fitness of Novosphingobium aromaticavorans during spaceflight
BMC Genomics (2023)
-
Impact of convection on the damping of an oscillating droplet during viscosity measurement using the ISS-EML facility
npj Microgravity (2021)
-
Molecular response of Deinococcus radiodurans to simulated microgravity explored by proteometabolomic approach
Scientific Reports (2019)
-
Medications in Space: In Search of a Pharmacologist’s Guide to the Galaxy
Pharmaceutical Research (2019)