Abstract
Huntington’s disease (HD) is a severe inherited neurological disorder caused by a CAG repeat expansion in the huntingtin gene (HTT), leading to the accumulation of mutant huntingtin with polyglutamine repeats. Despite its severity, there is no cure for this debilitating disease. HTT lowering strategies, including antisense oligonucleotides (ASO) showed promising results very recently. Attempts to develop stem cell-based therapeutics have shown efficacy in preclinical HD models. Using an HD patient’s autologous cells, which have genetic defects, may hamper therapeutic efficacy due to mutant HTT. Pretreating these cells to reduce mutant HTT expression and transcription may improve the transplanted cells’ therapeutic efficacy. To investigate this, we targeted the SUPT4H1 gene that selectively supports the transcription of long trinucleotide repeats. Transplanting SUPT4H1-edited HD-induced pluripotent stem cell-derived neural precursor cells (iPSC-NPCs) into the YAC128 HD transgenic mouse model improved motor function compared to unedited HD iPSC-NPCs. Immunohistochemical analysis revealed reduced mutant HTT expression without compensating wild-type HTT expression. Further, SUPT4H1 editing increased neuronal and decreased reactive astrocyte differentiation in HD iPSC-NPCs compared to the unedited HD iPSC-NPCs. This suggests that ex vivo editing of SUPT4H1 can reduce mutant HTT expression and provide a therapeutic gene editing strategy for autologous stem cell transplantation in HD.
Similar content being viewed by others
Introduction
Huntington’s disease (HD) is a progressive neurodegenerative disorder caused by the abnormal expansion of CAG repeats (>40) in the huntingtin gene (HTT) exon 11. The expanded CAG trinucleotides encode a polyglutamine stretch that can accumulate into neurotoxic proteinaceous cytoplasmic and intranuclear aggregates2. HD patients display progressive brain atrophy with increases in lateral ventricle size. These changes are followed by cognitive deficits, motor control impairment, and psychological symptoms.
There are currently no disease-modifying therapies for HD, consequently resulting in a significant unmet medical need. Effective neurorestorative or neurodegenerative strategies based on human stem cells are potential therapeutics. Previous studies have shown that transplanting human embryo-derived neural stem cells3 or mouse-induced pluripotent stem cell (iPSC)-derived neural stem cells4 into HD transgenic (TG) mice promoted neuronal or astrocytic differentiation with functional benefits. Patient-specific iPSCs are a potentially renewable source of autologous cells for stem cell therapy that will not induce immune rejection5,6,7. However, iPSCs derived from patients with genetic diseases carry the mutation. Therefore, gene modification before transplantation may increase the therapeutic potential of autologous iPSC therapy.
HTT is an evident candidate gene to target in HD because its genetic knockout or knockdown using CRISPR/Cas9, siRNA, or antisense oligonucleotides (ASO) rescues neurotoxicity8,9,10,11,12. HTT lowering strategies, including ASO were considered as a promising approach13,14. However, these methods will likely knockdown the mutant HTT (mHTT) and normal HTT, thereby ablating the physiological role of normal HTT. Normal HTT is thought to be required for embryogenesis, since deletion of the HD gene in mice results in early embryonic lethality15. Allele-specific targeting of mHTT via a single-nucleotide polymorphism (SNP) has been suggested11,12,16. However, not all HD patients harbor SNPs to allow for allele-specific knockdown of mHTT. Therefore, identifying a universal genetic manipulation to reduce mHTT levels without decreasing wild-type HTT levels would be useful for all HD patients.
SPT4 is a transcription elongation factor that is encoded by SUPT4H1 and regulates RNA polymerase II processivity17,18,19. One case report indicated that SPT4 is required for specific expression of the trinucleotide and hexanucleotide repeat expansions in HD20. Therapeutically targeting SPT4 to specifically reduce the mutant products derived from repeat expansion mutations was investigated. In the case of HD, SUPT4H1 knockdown reduced mHTT expression in Q81 or Q111 neuronal cells without compensatory increases in wild-type HTT levels. It also decreased mHTT aggregation and toxicity by inhibiting repeated trinucleotides20. Furthermore, suppressing the mouse homologue of SUPT4H1, Supt4a, by administering a SUPT4H1-targeting antisense oligonucleotide in the Q175 mouse HD model decreased mHTT mRNA and protein expression21. Heterozygous deletion of the Supt4a gene in R6/2 HD mice also reduced mHTT mRNA and protein expression20.
For these reasons, we investigated whether targeted ablation of SUPT4H1 (SUPT4H1-edited) in iPSC-NPCs derived from human HD patients increases the therapeutic possibility of autologous stem cell therapy iPSC-NPCs using a mouse HD model. The transplanted SUPT4H1-edited HD iPSC-NPCs were better engrafted than the unedited HD iPSC-NPCs at 12 w post-transplantation. Moreover, differentiated neurons and astrocyte were improved similarly. The restoration of healthy neurons and astrocytes consequently increased the striatal intensity of NeuN-positive cells and recovered motor and cognitive functions. These results strongly suggest the potential to develop autologous cell therapy in HD patients through SUPT4H1-edited iPSCs that suppress the HTT mutation.
Results
SUPT4H1-edited Q57 HD iPSC-NPCs recovered developmental defects in neurons and astrocytes
To investigate the potential therapeutic effects of SUPT4H1 knockdown in iPSC-NPCs, we first established NPC differentiation from iPSCs derived from an HD patient with 57 CAG repeats (Cell line ID: ND41656, RUCDR cell line service). We used the in vitro embryoid body-based SFEBq method to differentiate these iPSCs into NPCs.
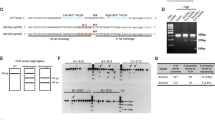
We screened and identified the optimal region of SUPT4H1 exon 1 for sgRNA targeting in NPCs using CRISPR/Cas9 (Supplementary Fig. 1a). The sgRNA-targeting noncoding region of the AAVS1 locus served as a control. Targeted deep sequencing analyses revealed efficient gene editing in the AAVS1 and SUPT4H1 loci reaching up to mean of 93.6 and 95.7% total indel efficiencies (mean of 29.5% in-frame, 64.1% out-of-frame mutation for Non-HD, and 27.9% in-frame, 67.9% out-of-frame mutation for Q57 HD iPSC-NPCs) in Non-HD and Q57 HD iPSC-NPCs subsequently (Supplementary Fig. 1a and Supplementary Table 1). To confirm SUPT4H1 mRNA knockdown following gene editing, we performed qRT-PCR analysis, which revealed robust knockdown of SUPT4H1 in both Non-HD and Q57 HD iPSC-NPCS with SUPT4H1 gene editing (Supplementary Fig. 1b). For NPC characterization, we performed immunostaining using NPC-positive markers (anti-SOX2, NESTIN, MUSASHI) and NPC-negative marker (anti-MAP2). No changes in NPC characteristics were observed after SUPT4H1 gene editing, judged by immunostaining using NPC markers (Supplementary Fig. 1c). IXMC analysis demonstrated that all treated groups showed over 90% positive staining for NPC markers (Supplementary Fig. 1d). Notably, SUPT4H1 gene editing did not produce changes in cell viability (Supplementary Fig. 1e). Furthermore, no karyotypic changes were detected after SUPT4H1 gene editing (Supplementary Fig. 2). To confirm SPT4 protein knockdown following gene editing, we performed immunostaining using anti-SPT4 and the NPC marker, anti-NESTIN. SPT4 expression in Q57 HD iPSC-NPCs increased compared to control NPCs (AAVS1- or SUPT4H1- edited control iPSC-NPCs). Conversely, SUPT4H1-edited Q57 HD iPSC-NPCs showed reduced SPT4 expression. Western blot analysis showed the same results (Supplementary Fig. 1f).
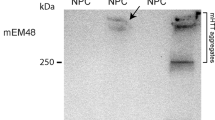
Having achieved highly efficient SUPT4H1 gene editing in HD iPSC-NPCs, we proceeded to spontaneously differentiate them into neural cells over 42 d (Fig. 1a). Before differentiation, we showed NPC characterization using immunofluorescence of NESTIN and SOX2 (Fig. 1b). After differentiation, we performed immunofluorescence and western blotting and found that SUPT4H1 gene editing reduced EM48 expression, a marker for mHTT protein, compared to AAVS1 gene editing in HD cells (Fig. 1c, d). To correlate with our western blot data, we performed RT-qPCR and found reduced expression of HTT in SUPT4H1 gene-edited Q57 HD NPCs (Supplementary Fig. 3).
a In vitro experimental scheme. The Q57 HD iPSC line was maintained for approximately 7 d (D-7). The dissociated cells were cultured in SFEBq media for 8 d (D-8). Embryoid bodies were dissociated in NPC media, then attempted Neon electroporation. The NPCs were maintained for several days and showed neuronal differentiation after 42 d (after iPSC culture, D-number; after gene editing, number day; blue color). b NPC immunostaining (positive marker: NESTIN, SOX2) following gene editing. c mHTT protein immunostaining (EM48) demonstrated a decreased HD phenotype in SUPT4H14-edited Q57 HD iPSC-NPCs (SUPT4H1 gRNA). d Western blot analysis of EM48 expression (n = 3, *p < 0.05, **p < 0.01). e MAP2 immunostaining for morphology exhibited neuronal dendrite maturation in SUPT4H1-edited Q57 HD iPSC-derived neurons. f. Western blot analysis of HMW and LMW MAP2 expression (n = 3, *p < 0.05, **p < 0.01). g The number of neurite and mean neurite length were increased in MAP2-positive cells of the SUPT4H1 gRNA group compared to the AAVS1-edited Q57 HD iPSC-derived neuron (AAVS1 gRNA) (each group: 100 cells, scale bar: 20 μm). h GFAP immunostaining. i Western blot analysis for GFAP and Kir4.1 revealed normal astrocytic function in the SUPT4H14 gRNA group. (n = 3, *p < 0.05, **p < 0.01). j Real-time PCR analysis for astrocytic development in AAVS1-edited Q57 HD iPSC-derived NPCs. The CTL (control) and SUPT4H1-edited groups showed significant increases in CDH2, a radial glial cell marker, and S100β, a maturate astrocyte marker, mRNA compared to the AAVS1-edited group (n = 3, *p < 0.05, **p < 0.01, and ***p < 0.001). Data were analyzed using two-way ANOVAs followed by Tukey’s post hoc tests with SPSS software or GraphPad Prism. The error bars on the bar charts represent the standard deviation.
Interestingly, we observed that AAVS1-edited Q57 HD iPSC-derived MAP2- or GFAP-positive cells exhibited different morphology compared to the control or SUPT4H1-edited group (Fig. 1e, h). MAP2-positive cell maturation was first identified by western blotting (Fig. 1f). Low molecular weight (LMW) MAP2 isoforms are the immature forms that result from the exclusion of the sequence encoded by exons E7-E9. They are downregulated after the early stages of neuronal development when E7-E9 exon-including high molecular weight (HMW) isoforms are favored22. Although HMW-MAP2 expression was observed in the AAVS1-edited group, the SUPT4H1-edited group showed significantly increased HMW-MAP2 expression compared to the AAVS1-edited group.
We next analyzed MAP2-positive cell morphology using MetaXpress for more reliable evidence. We found that the SUPT4H1-edited Q57 HD iPSC-derived MAP2-positive cells exhibited significantly more total outgrowths and increased maximum outgrowth length compared to the AAVS1-edited Q57 HD iPSC-derived MAP2-positive cells (Fig. 1g). Furthermore, western blotting revealed expression of GFAP and the potassium channel Kir4.1, functional astrocyte markers. The SUPT4H1-edited group showed increased GFAP and Kir4.1 expression compared to the AAVS1-edited group (Fig. 1i). To identify differentiated neurons and astrocytes, we examined the mRNA expression of GABA and DARPP-32 for mature neurons, and N-cadherin (CDH2) and S100β for radial glial cell and mature astrocytes, respectively. The SUPT4H1-edited group showed significant increases in GABA and DARPP-32 mRNA expression, compared to the AAVS1-edited group (Supplementary Fig. 4). In addition, the SUPT4H1-edited groups showed significant increases in CDH2 and S100β mRNA compared to the AAVS1-edited group (Fig. 1j). Therefore, the SUPT4H1-edited Q57 HD iPSC-NPCs with reduced mHTT expression underwent normal neural differentiation similar to the control iPSC-derived NPCs.
Although knocking down HTT using gene editing reduces mHTT, it also reduces normal wild-type HTT levels, which play a role in neuronal development15. To investigate this, we performed HTT gene editing using sgRNAs targeting the start codon of the HTT gene (Supplementary Fig. 5a). We achieved a similar gene-editing efficiency between AAVS1- and SUPT4H1-targeting sgRNAs. The HTT-edited Q57 HD iPSC-NPCs showed an incomplete differentiation compared to the SUPT4H1-edited group. Regarding MAP2-positive cell morphology, the number of total neurites and maximum neurite lengths were significantly reduced in the HTT-edited group compared to the control group or AAVS1- or SUPT4H1-edited groups (Supplementary Fig. 5b, c). Interestingly, the MAP2-positive cells of the HTT-edited group did not exhibit increased maturation compared to the AAVS1-edited group. As HTT-edited Q57 HD iPSC-NPCs showed defective neuronal development, we chose to focus on SUPT4H1-edited Q57 HD iPSC-NPCs.
Functional improvement and neuroprotective effects of SUPT4H1-edited Q57 HD iPSC-NPC transplantation in YAC128 HD mice
To compare the therapeutic potential of SUPT4H1-edited and unedited Q57 HD iPSC-NPCs, we transplanted these cells into YAC128 mice harboring mHTT with 128 CAG repeats. We investigated whether SUPT4H1-edited HD iPSC-NPCs rescued motor deficits on the rotarod test in the YAC128 mice at 3 m following transplantation (Fig. 2a). We found that mice injected with unedited Q57 HD iPSC-NPCs showed delays in motor deficits up to 1 m postinjection, whereas mice injected with SUPT4H1-edited Q57 HD iPSC-NPCs maintained motor function up to 3 m postinjection. (Fig. 2a; p < 0.01). YAC128 mice also typically show deficits in the fine motor control and grip strength. Therefore, we further observed that YAC128 mice transplanted with SUPT4H1-edited Q57 HD iPSC-NPCs demonstrated a significantly improved performance at 3 m after transplantation compared to the nonedited Q57 iPSC-NPCs transplanted group (Fig. 2b, p < 0.01). These results were comparable to those transplanted with control iPSC-NPCs (Supplementary Fig. 6).
a, b Functional motor recovery was monitored every month using the accelerating rotarod and grip strength tests. YAC128 mice transplanted with SUPT4H1-edited Q57 HD iPSC-NPCs showed significantly accelerated motor recovery compared to the Q57 HD iPSC-NPCs transplants. All transplant mice significantly recovered at 1 m post-transplantation and this recovery was maintained until 3 m post-transplantation in the SUPT4H1-edited Q57 HD iPSC-NPCs group (WT = 9, Media = 8, Q57 = 8, and Q57/SUPT4H1 = 8; Media vs. Q57 or Q57/SUPT4H1: *p < 0.05 and **p < 0.01; Q57 vs. Q57/SUPT4H1: +p < 0.05). c Immunostaining revealed an increased NeuN-positive area density in the striatum of YAC128 mice transplanted with SUPT4H1-edited Q57 HD iPSC-NPCs (scale bar: 100 μm). d Image J analysis showing a significant increase in the striatal density of mice that received SUPT4H1-edited Q57 HD iPSC-NPCs transplants (n = 3, *p < 0.05, **p < 0.01). Data were analyzed using two-way ANOVAs followed by Tukey’s post hoc tests with SPSS software. The error bars on the bar charts represent the standard deviation.
We next investigated whether transplanted SUPT4H1-edited Q57 HD iPSC-NPCs (Q57/SUPT4H1) provide a protective effect on neuronal survival in the brain of YAC128 mice. Three striatal sections from five mice per group were stained with neuronal nuclei (NeuN) antibody and the intensity of NeuN-positive neurons was quantified using ImageJ software 3 m after media or Q57 HD iPSC-NPC transplantation. The YAC128 mice exhibited significantly reduced NeuN-positive density compared to WT. However, the YAC128 mice transplanted with SUPT4H1-edited-Q57 HD iPSC-NPCs showed significantly increased NeuN-positive density (Fig. 2c, d).
SUPT4H1-edited Q57 HD iPSC-NPCs engrafted better than unedited cells in YAC128 mice
As SUPT4H1-edited Q57 HD iPSC-NPCs showed therapeutic efficacy compared to unedited cells, we next investigated the fate of these cells following transplantation. Immunofluorescence analysis for human-specific NESTIN (hNESTIN) confirmed the absence of SPT4 expression in transplanted SUPT4H1-edited Q57 HD iPSC-NPCs, indicative of transplanted human cells. However, notable SPT4 expression was found in nonedited cells 1 week following transplantation (Fig. 3a). We counted the pixel numbers of merged hNESTIN and SPT4 in the immunofluorescence images using colocalization analysis of Zen black software (Fig. 3b).
a Double staining for transplanted cell markers (hNESTIN) and Spt4 protein revealed that transplanted unedited or SUPT4H1-edited Q57 HD iPSC-NPCs decreased Spt4 expression (scale bar: 20 μm). b The number of pixels for Spt4 expression quantified using ZEN black software analysis were significantly decreased in transplanted SUPT4H1-edited Q57 HD iPSC-NPCs (n = 3, **p < 0.01). c Immunostaining for hNu 12 week post-transplantation showed human cell survival in the striatum of YAC128 mice transplanted with unedited or SUPT4H1-edited Q57 HD iPSC-NPCs (scale bar: 20 μm). d Quantification of hNu-positive cell numbers in the 6–7 sections included the striatal lesion site (n = 5). Data were analyzed using two-way ANOVAs followed by Tukey’s post hoc tests with SPSS software. The error bars on the bar charts represent standard deviation.
Human nuclei (hNu)-positive cells were observed in the striatum of SUPT4H1-edited and unedited cell transplantation groups 3 m after transplantation (Fig. 3c), indicating that the transplanted iPSC-NPCs survived in the striatum of YAC128 mice. However, the total number of hNu-positive cells in the striatum of the unedited cell transplantation group was substantially less than in the SUPT4H1-edited cell transplantation group (54,280 ± 823 or 128,676 ± 4356 in the striatum, corresponding to approximately 13.5 or 32.2% of the total number of transplanted cells; i.e., 4 × 105) (Fig. 3d). More GFAP-positive cells, co-localized with hNu-positive cells, were observed in YAC128 mice transplanted with the unedited Q57 HD iPSC-NPCs (Fig. 5a) compared to hMAP2-positive cells (Fig. 4c). In contrast, more hMAP2-positive neurons were detected in YAC128 mice transplanted with the SUPT4H1-edited Q57 HD iPSC-NPCs (Supplementary Fig. 7).
a Double staining for the transplanted cell marker (hNu) and MSN marker (DARPP-32) revealed that transplanted unedited or SUPT4H1-edited Q57 HD iPSC-NPCs differentiated into MSNs (scale bar: 20 μm). b Double staining for the human-specific MSN marker (hDARPP-32) and HD phenotype marker (EM48) in the striatum demonstrated decreased EM48 expression in transplanted SUPT4H1-edited Q57 HD iPSC-derived MSNs (scale bar: 20 μm). c Immunostaining to identify hMAP2-positive cells exhibited neuronal dendrite degeneration in the transplanted unedited Q57 HD iPSC-NPCs-derived neurons (scale bar: 20 μm). d Quantification of neurite numbers and max neurite length indicated more neuronal maturation in transplanted SUPT4H1-edited Q57 HD iPSC-NPCs-derived MAP2-positive cells than the unedited cells. (Photograph: same color is one cell for analysis, scale bar: 100 μm, for two regions in 6–7 brain sections of each group, ***p < 0.001). Data were analyzed using two-way ANOVAs followed by Tukey’s post hoc tests or Student’s t tests with GraphPad Prism. The error bars on the bar charts represent the standard deviation.
Neurons differentiated from SUPT4H1-edited Q57 HD iPSC-NPCs showed improved morphology in YAC128 mice
As more cells survived, we further investigated whether transplanted SUPT4H1-edited Q57 HD iPSC-NPCs were likely to differentiate into medium spiny neurons (MSNs) or show other morphology. For this purpose, we double stained slices with hNu and DARPP-32 antibodies (Fig. 4a). The merged hNu/DARPP-32-positive cells in YAC128 mice transplanted with unedited Q57 HD iPSC-NPCs exhibited an unhealthy morphology, such as the defective morphology of DARPP-32-positive neurons, compared to the SUPT4H1-edited group (Fig. 4a). The cells in the unedited group had EM48 aggregates that were not present in the SUPT4H1-edited group (Fig. 4b). To identify the neuronal maturation morphology, like in the in vitro study, we performed immunofluorescence and MetaXpress software analysis using a human-specific MAP2 (hMAP2) antibody. Interestingly, we found that hMAP2-positive cells in YAC128 mice transplanted with unedited Q57 HD iPSC-NPCs had shorter dendrites and no straight morphology compared to the SUPT4H1-edited group (Fig. 4c). The MetaXpress software analysis revealed that the outgrowth numbers and maximum length of hMAP2-positive cells in the SUPT4H1-edited group were significantly increased compared to the unedited group (Fig. 4d).
Astrocytes differentiated from SUPT4H1-edited Q57 HD iPSC-NPCs reduced reactive astrocytes in YAC128 mice
We next examined whether transplanted SUPT4H1-edited-Q57 HD iPSC-NPCs had the potential to differentiate into astrocytes (GFAP). Slices were double stained with hNu and GFAP antibodies (Fig. 5a). The hNu/GFAP-positive cells in the YAC128 mice transplanted with unedited Q57 HD iPSC-NPCs showed a hypertrophic morphology of reactive astrocytes, compared to the SUPT4H1-edited group (Fig. 5a). Furthermore, the cells in the unedited group had HTT aggregates that were not present in the SUPT4H1-edited group (Fig. 5b). To identify reactive astrocytes, we double stained slices with antibodies for human GFAP (hGFAP) and complement component 3 (C3), considered a reactive astrocyte marker, and analyzed them using immunofluorescence. We observed increased hGFAP/C3-positive cell expression in YAC128 mice transplanted with unedited Q57 HD iPSC-NPCs compared to the SUPT4H1-edited group (Fig. 5c). The MetaXpress software analysis revealed that the area and intensity of GFAP-positive cells in the AAVS1-edited group increased significantly and showed reactive astrocyte morphology, but the SUPT4H1-edited group did not show reactive astrocyte morphology (Fig. 5d). Therefore, we propose that the YAC128 mouse brain environment, in which mHTT proteins are highly accumulated, converts normal astrocytes into reactive astrocytes.
a Double staining for the transplanted cell marker (hNu) and astrocyte marker (GFAP) revealed that transplanted unedited or SUPT4H1-edited Q57 HD iPSC-NPCs differentiated into astrocytes (scale bar: 20 μm). b Double staining for the human-specific astrocyte marker (hGFAP) and HD phenotype marker (EM48) in the striatum revealed decreased EM48 expression in transplanted SUPT4H1-edited Q57 HD iPSC-derived astrocytes (scale bar: 20 μm). c Immunostaining to identify C3 expression revealed reactive astrocytes in transplanted unedited Q57 HD iPSC-NPCs-derived astrocytes (scale bar: 20 μm). d Quantification of the GFAP-positive area and intensity indicated more activation in the transplanted SUPT4H1-edited Q57 HD iPSC-NPCs-derived astrocytes than the unedited cells. (Photograph: same color is one cell for analysis, scale bar: 100 μm, for two regions in 6–7 brain sections of each group, ***p < 0.001). Data were analyzed using two-way ANOVAs followed by Tukey’s post hoc tests or Student’s t tests using GraphPad Prism. The error bars on the bar charts represent standard deviation.
Discussion
This study demonstrated that SUPT4H1-edited Q57 HD iPSC-NPC transplantation stabilized motor function and improved neuroprotection in the damaged brain of YAC128 mice. We show that SUPT4H1 gene editing in HD iPSC-NPCs can facilitate normal neural differentiation and functional recovery while avoiding the risks of immunosuppression associated with autologous stem cell therapy. For genetic diseases like HD, ex vivo approaches that combine gene and cell therapy will be highly useful for developing autologous stem cell therapies.
Experimental and preclinical studies supported the commencement of the first neural transplantation clinical trial in HD patients in the 1990s23. From human fetal tissue to porcine striatal cells, several cell sources have been used for clinical trials in HD patients24,25,26,27,28. However, the results have been highly variable and the clinical benefits were modest for motor26 and neuropsychological outcomes27. Graft survival and neuronal differentiation have been studied in patients using magnetic resonance imaging29. By contrast, iPSCs may produce an unlimited source of patient’s autologous cells that can be used in regenerative medicine30. However, mutated genes in autologous cells may cause abnormal neural differentiation and disrupt normal physiological functions, necessitating cell editing for stem cell therapy in HD. In this study, we showed that ablating SPT4 can contribute to the improvement of neurological defects associated with mHTT in YAC128 mice.
First, we compared HTT- and SUPT4H1-edited Q57 HD iPSC-NPCs. mHTT causes dysregulation of global gene expression and cellular dysfunction. Therefore, mHTT gene knockout or knockdown may be a focus of gene therapy in HD. However, normal and expanded alleles of the endogenous HTT gene are not readily distinguished by siRNA or antisense oligonucleotides11,12. HTT loss and mutation in mouse embryonic stem cells impairs progenitor specification and maturation31,32 and targeted genetic modification results in fewer neuronal cells and decreased neurite length33. The current study similarly edited the HTT gene, which is present in normal and expanded alleles. Our results showed that HTT-edited Q57 HD iPSC-NPCs reduced neuronal differentiation and neurite length, indicating that HTT gene ablation reduced neural differentiation and the maintenance of neuronal survival in vitro. This might be because our CRISPR/Cas9 HTT strategy did not distinguish between the wild-type and mutant alleles. In addition, complete HTT knockout might negatively affect neuronal development. Allele-specific targeting of mHTT with CRISPR/Cas9 based on SNP associations has recently been developed16. However, this approach is likely to be limited by the various associative patterns between SNPs and mHTT in patients. In addition, one base pair difference in target sites for CRISPR/Cas9 would likely result in quantitative, but not qualitative, preference in cleavage, increasing the chances of residual cleavage on wild-type HTT alleles.
Therefore, we considered that SPT4, a transcription elongation factor, would be the ideal candidate gene for editing HD iPSC-NPCs. Yeast and animal cells with defective SPT4 show reduced synthesis of proteins containing long polyQ stretches, such as mHTT17,20. In contrast, SPT4 gene defects do not alter cellular mRNA synthesis. Furthermore, HD Q175 mice injected with SPT4 antisense 2′-O-methoxyethyl oligonucleotide and SPT4-one-copy-deleted R6/2 mice showed selective decreases in mHTT mRNA and protein expression. Furthermore, SPT4-one-copy-deleted R6/2 mice showed a prolonged lifespan and delayed motor impairments17.
For these reasons, we hypothesized that SPT4 knockout or knockdown would be the best approach to selectively reduce mHTT. The present study investigated the in vivo and in vitro functions of SUPT4H14-edited Q57 HD iPSC-NPCs. Our first result demonstrated that SUPT4H1-edited Q57 HD iPSC-NPCs showed decreased SPT4 protein levels. Compared to dual HTT-edited Q57 HD iPSC-NPCs, the SUPT4H1-edited cells were more likely to differentiate into neuronal cells with increased neurite length and decreased expression of mHTT protein.
Second, our results showed that the unedited Q57 HD iPSC-NPCs were more likely to differentiate into astrocytes than neurons. It has been reported that HD Q140 knock-in NSCs or human HD iPSC (60, 109, and 180Q)-NPCs resulted in fewer NPCs, fewer mature neurons, and more astrocytes due to defects in BDNF expression, which modulates neuronal maturation34. Our transplantation results demonstrated that unedited Q57 HD iPSC-NPCs were more likely to differentiate into astrocytes, whereas SUPT4H1-edited Q57 HD iPSC-NPCs were more likely to differentiate into mature neurons. Therefore, although further study is required, these results suggest that reducing mHTT via SUPT4H1 gene editing may enhance neuronal differentiation following in vivo transplantation.
Third, we found that the SUPT4H1-edited Q57 HD iPSC-NPCs improved astrocytic function, which is as important as neuronal replacement. HD patients’ brains contain reactive astrocytes with thicker processes, a larger soma, and abnormal function that is associated with reduced glutamate transporter35. In addition, abnormal astrocytes are implicated in HD pathophysiology35,36,37. Our results demonstrated that the SUPT4H1-edited Q57 HD iPSC-NPCs formed less reactive astrocytes due to reduced mHTT expression. Therefore, SUPT4H1-edited Q57 HD iPSC-derived astrocytes may improve the HD brain environment.
In addition, we demonstrated interesting iPSC characterization results. Several studies have demonstrated that HD patients’ iPSCs or their derived NPCs function normally without mHTT expression, based on ubiquitin-protein system activity38,39,40. For example, human embryonic stem cells with mHTT exhibit the associated pathology approximately 2 m following differentiation41. Patient-derived iPSCs from juveniles and adults with HD showed mHTT expression in neurons after 6 m of culture42 or transplantation43. The ubiquitin-protein system loses its normal function over the course of neural differentiation40. In the present study, YAC128 mice transplanted with unedited Q57 HD iPSC-NPCs exhibited motor recovery at 1 m post-transplantation. However, these mice also had mHTT, which was associated with reduced neural differentiation. Therefore, our results demonstrate that mHTT in the unedited Q57 HD iPSC-derived neurons or astrocytes of YAC128 mice may hamper iPSC-NPCs from engrafting and differentiating.
Stem cell therapy is an important therapeutic strategy for several diseases. Numerous clinical and non-clinical studies are actively underway to achieve more effective outcomes. Specifically, some research groups are attempting gene editing to develop stem cells that will not undergo immune rejection. A sufficient supply of autologous stem cells could produce the best treatment if they do not suffer from immune rejection. However, patients with gene mutations, such as in HD, should only have their iPSCs transplanted after gene correction or editing. An easier and more convenient “universal” gene editing approach would be to target the regulatory gene instead of the specific gene in question, which would require more laborious procedures, like gene correction. In the case of HD, we suggest SUPT4H1 gene editing as an appropriate and efficient strategy to develop autologous stem cell treatment. Furthermore, ex vivo approaches based on combining SUPT4H1 gene editing and cell therapy may warrant further study for autologous stem cell therapy in other repeat expansion mutation-associated diseases, such as C9FTD/ALS44.
Methods
Lead contact and material availability
Further information and requests for resources and reagents should be directed to and will be fulfilled by the Lead Contact Jihwan Song (jsong5873@gmail.com). This study did not generate new unique reagents.
iPSC cell lines
The HD patient iPSC line is the Q57 HD iPSC line (Cell line ID: ND41656) purchased from the RUCDR cell line service. The control iPSC line used in this study was CHAi001-A, which was established from the frozen cord blood of a healthy donor using the episomal method45.
Cell culture and NPC induction
The control and Q57 HD iPSC lines were maintained in Stemfit Basic02 media (Ajinomoto, Japan) supplemented with 100 ng/ml of basic fibroblast growth factor (bFGF, Peprotech) and 10 µM Y27632 (ROCK inhibitor, Peprotech) for approximately 7 d before they were treated with TrypLE solution (GIBCO) for 5 min at 37 °C in a CO2 incubator. The dissociated cells were cultured in SFEBq media consisting of DMEM/F12 (Invitrogen) supplemented with 1% antimycotic-antibiotics, 1% nonessential amino acids (NEAA, GIBCO), 0.1% beta-MeOH, 20% KnockoutTM serum replacement, 10 µM SB431542, 100 nM LDN193189, and 3× ROCK inhibitor at 37 °C in a CO2 incubator for neural induction. The cells were maintained in SFEBq media for 8 days. Embryoid bodies were dissociated in NPC media consisting of DMEM/F12 supplemented with 1:100 antimycotic-antibiotics (Welgene), 1:100 NEAA, sodium pyruvate (GIBCO), d-glucose (Sigma-Aldrich), l-glutamine (Welgene), 1:1000 beta-MeOH, 1:50 B-27 (without vitamin A, GIBCO), and 20 ng/ml bFGF in a dish coated with poly-l-ornithine (Sigma-Aldrich) and laminin (Sigma-Aldrich). Accutase (Stem Cell Technologies) was used to split the cells.
Preparation of sgRNA
sgRNAs were generated by in vitro transcription using T7 polymerase (New England Biolabs) according to the manufacturer’s protocol. sgRNAs used in this study are listed in Table 1.
Gene editing of iPSC-NPCs
For gene editing, ribonucleoprotein (RNP) complex-mediated electroporation was performed as previously described46. Briefly, a Neon electroporator (Thermo Fisher Scientific) was used to transfect 1 × 105 iPSC-NPCs with RNP complexes comprised of 1 μg of sgRNA and 4 μg of Cas9 protein (ToolGen). For targeted deep sequencing, genomic DNA (gDNA) was collected from cells 72 h after transfection.
Targeted deep sequencing
Gene-editing efficiency was verified using targeted deep sequencing as previously described46. Briefly, the on-target region was PCR amplified from the gDNA extracted from transfected cells using Phusion polymerase (New England Biolabs). The resulting PCR amplicons were then subjected to paired-end deep sequencing using Mi-Seq (Illumina). The deep sequencing data were analyzed using the online Cas-Analyzer tool (www.rgenome.net). Indels in the region 3 bp upstream from the PAM sequence were considered to be Cas9-induced mutations. The primers used in this study are listed in Table 2.
Neural differentiation
Unedited, SUPT4H1-, or HTT dual-edited Q57 HD iPSCs-NPCs were seeded at a density of 2 × 104 cells on poly-L-ornithine and fibronectin double-coated glass (24 × 60 mm). The medium was changed to neural differentiation medium (NDM) on the following day and changed every 2 days for 6 weeks. The NDM was based on a 1:1 mixed medium of DMEM F12: Neurobasal (GIBCO, Waltham, USA) and composed of 1× Glutamax (GIBCO) and 1% B27 supplement without vitamin A (GIBCO). The cells were fixed for immunocytochemistry and western blot 6 w following differentiation.
Real-time PCR for GFAP development analysis
The gene-edited cells were washed with RNase-free ice-cold PBS, then total RNA was extracted using TRIzol 6 w following differentiation. RT was conducted using Superscript reverse transcription. Total RNA (1 μg) was used as a template for RT reaction. Quantitative RT-PCR was performed using the SYBRTM Green system (Thermo Fisher Scientific). Amplification was monitored and analyzed by measuring SYBR green binding. For each cDNA, a triplicate amplification was carried out using 2.5 μL (10–100 ng) cDNA, 12.5 mL of Universal PCR Master Mix (Thermo Fisher Scientific), 1 μM each of the primers for CDH2 (immature astrocyte) or S100β, and 1 μl of fluorescent probe (Table 3). The data were recorded as cycle threshold on a Step One Plus from Applied Biosystems.
YAC128 mice and transplantation
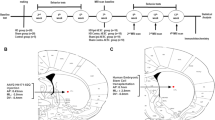
All experiments were performed in YAC128 mice maintained on a FVB/N background47. Mice were bred in the animal facility at CHA Bio Complex (Pangyo, Korea). Experiments were performed with the approval of the Institutional Animal Care and Use Committee (IACUC 200019) of CHA University. Six-month-old mice were used in the transplantation experiments. We stereotaxically injected a total of 10 mice with 1 µl of unedited or SUPT4H1-edited Q57 HD iPSC-NPCs (100,000 cells/µl), respectively into two sites of each hemisphere (a total volume of 4 µl in each mice) using the following coordinates: AP+0.5 mm, ML:±1.8 mm, and DV:−3 and −4 mm from bregma. In the sham group (n = 10), 1 µl of suspension medium (DMEM) was injected into two sites of each hemisphere using the same coordinates. No brain edema etc. was observed with hNu staining in the transplanted brains (Supplementary Fig. 8). Both cell-transplanted and medium-injected animals were intraperitoneally injected with cyclosporine A (5 mg/kg, Sigma) 3 d before the transplantation and daily for up to 3 m until they were sacrificed.
Behavioral tests
Rotarod and grip strength tests were performed over 3 months to evaluate the improvement of motor functions by transplanted cells in 6-m-old YAC128 mice.
Accelerating rotarod test
We used an accelerating rotarod protocol (San Diego Instruments) to assess motor coordination and gait changes. Accelerations ranged from 0 to 45 rpm over a period of 2 min. Mice were trained for two trials/d for 3 days. Following the training period, mice were tested for 3 consecutive trials in a single day and allowed 1.5 h of rest time between the trials. The rotarod was wiped clean with ethanol between each subject and trial.
Grip strength test
Grip strength was used as an additional assessment of motor function48. The apparatus (San Diego Instruments) consisted of an adjustable grip (6 cm wide, 0°–45°) connected to a digital gauge. For this measure, the mouse was lifted by the tail so that its forepaws could grasp the grip. The mouse was then gently pulled backward by the tail until the wire was released. The maximal force exerted before the mouse lost its grip was recorded. Each mouse was tested in 9 replicates, and the average of the 3 highest scores was used for subsequent analyses.
Western blot analysis
Brain tissues were lysed with RIPA buffer (150 mM NaCl, 1% Nonidet P-40, 0.5% deoxycholic acid, 0.1% SDS, and 50 mM Tris-HCl, pH 7.4) containing a protease inhibitor (Roche), sonicated for 1 min, and incubated on ice for 20 min. The supernatant was collected and the protein concentrations were determined using the BCA assay (Thermo Fisher Scientific) after centrifugation at 13,200 g for 15 min at 4 °C. Equal amounts of proteins were then loaded on ~8–12% SDS polyacrylamide gels. The separated samples were transferred to a PVDF membrane, incubated with primary antibodies against the target proteins, and subsequently incubated with HRP-conjugated secondary antibodies. The protein bands were visualized by enhanced chemiluminescence (Millipore) using a bioimaging analyzer (Bio-Rad). The relative intensity of each band was measured using ImageJ software (rsb.info.nih.gov, by W. Rasband). The following primary antibodies were used for western blotting: Spt4 (Cell Signaling 1:1000), EM48 (Millipore, 1:500), MAP2 (Abcam, 1:1000), GFAP (DAKO, 1:1000), and Kir4.1 (Millipore, 1:1000). All blots were derived from the same experiment and were processed in parallel. Un-cropped images of all Western blot results are provided (Supplementary Fig. 9).
Immunohistochemistry
The YAC128 mice used in this experiment were sacrificed 3 m after cell transplantation following completion of the behavioral analyses. All mice were perfused with saline containing heparin at 5 µl/ml then 4% paraformaldehyde solution. Mouse brains were fixed in 4% paraformaldehyde solution overnight and left to soak in 30% sucrose solution for approximately 3 d. Subsequently, the brain tissues were frozen using an OCT compound and sectioned into 30-µm-thick slices using a cryotome (CM3050, Leica, Germany). The cryosectioned brain tissues were stored in a cryoprotective solution (40% glycerol, 40% ethylene glycol, and 20% 0.2 M phosphate buffer solution) at −20 °C. The slices from each mouse were incubated in a blocking buffer (10% donkey serum and 0.3% Triton X-100 in PBS) for 30 min at room temperature. Thereafter, the samples were incubated overnight with primary antibodies at 4 °C, and then incubated with the appropriate fluorescent probe-conjugated secondary antibodies for 1 h at room temperature protected from light. Nuclei were stained with 4,6-diamidino-2-phenylindole (DAPI, Thermo Fisher Scientific) at a 1:5000 dilution. Images were captured using a confocal microscope (LSM880, Zeiss). The specific primary antibodies used in this study included SPT4 (Biorbyt, 1:200), hNESTIN (R&D Systems, 1:200), SOX2 (Millipore, 1:200), EM48 (Millipore, 1:100), MAP2 (Abcam, 1:200), GFAP (DAKO, 1:200), NeuN (Millipore, 1:200), hNu (Millipore, 1:200), DARPP-32 (Cell Signaling, 1:200), hMAP2 (Thermo Fisher Scientific, 1:200), hGFAP (R&D Systems, 1:200), and C3 (Abcam, 1:100).
Quantification of hNu-positive cells and striatal density
Unbiased stereological estimation of the total number of hNu-, DARPP-32-, and Iba-1 positive cells, as well as double stained GFAP/hNu-positive cells in the striatum was made using a ImageXpress microconfocal high-content imaging system (Molecular device). The sections used for counting covered the entire striatum over 2 mm, generally requiring 6–7 sections in a series. The positive cells were counted across all regions of the 6–7 sections and analyzed using ImageXpress microconfocal high-content imaging system (Molecular Device) connected to the stage that fed the Z-axis distance information to the computer. The striatum was delineated under a 20x objective. Only the DAPI-merged hNu-stained cells that came into focus within the counting volume (depth of 10 μm) were counted by the scoring module in MetaXpress (Molecular Devices). The total number of cells was calculated according to the optical fractionator formula49.
We measured the striatal density of NeuN (Millipore, 1:200)-positive areas to investigate the neuroprotective effects of the transplanted cells. NeuN immunostaining in the striatum was estimated using an optical fractionator and unbiased stereology of stained cells. To measure the striatal density, NeuN immunostaining images were captured in 3 sections near the middle of the striatum from a total of 3 mice. The images were delineated under a 4x objective and measured using ImageJ software (rsb.info.nih.gov, by W. Rasband).
Morphology analysis and number of MAP2- or GFAP-positive cells
Using an in vitro system, the number of neurites and mean neurite length of MAP2-positive cells were measured in 1000 cells from each culture group. An in vivo system was also used to quantify the number of processes, the maximal process length, the outgrowth intensity, and the body area of MAP2- or GFAP-positive cells in five striatal regions from 6–7 sections of brains from each experimental group (n = 3). The outgrowth was measured using the outgrowth module of MetaXpress software (Molecular Device).
For ratio analysis of differentiated transplanted cells in the mice, merged human hNu- and MAP2- or GFAP-positive cells were measured in two whole striatal regions from 6–7 sections of brains from each experimental group (n = 3). The number of merged positive cells was measured using the cell sorting module of MetaXpress software (Molecular Devices).
Real-time PCR (TaqMan probe-based)
For TaqMan probe-based qRT-PCR, 100 ng of cDNA was utilized for each reaction using TaqMan Gene expression master mix according to the manufacturer’s protocol (Thermo Fisher Scientific) on Quantstudio 3. The level of each gene expression was calculated using the CT value, and GAPDH was used as an endogenous control (Table 4).
NPC characterization analysis
For NPC characterization analysis, merged human hNu- and NESTIN- or SOX2- or MUSASHI-positive cells were measured in 96 well plate (Table 5). The number of merged positive cells was measured using the cell sorting module of MetaXpress software (Molecular Devices).
Cell viability analysis
Upon gene editing, cells were washed with PBS and detached by TrypLE Select (ThermoFisher). Cells were then centrifuged for 5 mins at 1500 rpm. These were then stained with 0.4% Tryphan Blue and viabilities were measured using automated cell counter FACSSCOPE B (Curiosis).
Karyotype analysis
Karyotyping was performed using a GTG-banding analysis (Korea Research of Animal Chromosomes, Korea).
Statistical analysis
Data were analyzed using two-way ANOVAs followed by Tukey’s post hoc tests or Student’s t tests, using SPSS software (version 10.0; Chicago, IL) or GraphPad Prism (version 5.0; San Diego). Significance was accepted at the 95% probability level. Data are presented as mean ± SEM.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Change history
21 April 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41536-022-00219-6
References
Bates, G. P. et al. Huntington disease. Nat. Rev. Dis. Prim. 1, 15005 (2015).
Ross, C. A. Polyglutamine pathogenesis: emergence of unifying mechanisms for Huntington’s disease and related disorders. Neuron 35, 819–822 (2002).
Reidling, J. C. et al. Human Neural Stem Cell Transplantation Rescues Functional Deficits in R6/2 and Q140 Huntington’s Disease Mice. Stem Cell Rep. 10, 58–72 (2018).
Al-Gharaibeh, A. et al. Induced Pluripotent Stem Cell-Derived Neural Stem Cell Transplantations Reduced Behavioral Deficits and Ameliorated Neuropathological Changes in YAC128 Mouse Model of Huntington’s Disease. Front Neurosci. 11, 628 (2017).
Ilic, D. & Ogilvie, C. Concise Review: Human Embryonic Stem Cells-What Have We Done? What Are We Doing? Where Are We Going? Stem Cells 35, 17–25 (2017).
Sebban, S. & Buganim, Y. Nuclear Reprogramming by Defined Factors: Quantity Versus Quality. Trends Cell Biol. 26, 65–75 (2016).
Yoshihara, M., Hayashizaki, Y. & Murakawa, Y. Genomic Instability of iPSCs: Challenges Towards Their Clinical Applications. Stem Cell Rev. Rep. 13, 7–16 (2017).
Gagnon, K. T. et al. Allele-selective inhibition of mutant huntingtin expression with antisense oligonucleotides targeting the expanded CAG repeat. Biochemistry 49, 10166–10178 (2010).
Lombardi, M. S. et al. A majority of Huntington’s disease patients may be treatable by individualized allele-specific RNA interference. Exp. Neurol. 217, 312–319 (2009).
Pfister, E. L. et al. Five siRNAs targeting three SNPs may provide therapy for three-quarters of Huntington’s disease patients. Curr. Biol. 19, 774–778 (2009).
van Bilsen, P. H. et al. Identification and allele-specific silencing of the mutant huntingtin allele in Huntington’s disease patient-derived fibroblasts. Hum. Gene Ther. 19, 710–719 (2008).
Zhang, Y., Engelman, J. & Friedlander, R. M. Allele-specific silencing of mutant Huntington’s disease gene. J. Neurochem 108, 82–90 (2009).
Kordasiewicz, H. B. et al. Sustained therapeutic reversal of Huntington’s disease by transient repression of huntingtin synthesis. Neuron 74, 1031–1044 (2012).
Tabrizi, S. J., Ghosh, R. & Leavitt, B. R. Huntingtin Lowering Strategies for Disease Modification in Huntington’s Disease. Neuron 102, 899 (2019).
Reiner, A., Dragatsis, I., Zeitlin, S. & Goldowitz, D. Wild-type huntingtin plays a role in brain development and neuronal survival. Mol. Neurobiol. 28, 259–276 (2003).
Shin, J. W. et al. Permanent inactivation of Huntington’s disease mutation by personalized allele-specific CRISPR/Cas9. Hum. Mol. Genet 25, 4566–4576 (2016).
Hirtreiter, A. et al. Spt4/5 stimulates transcription elongation through the RNA polymerase clamp coiled-coil motif. Nucleic Acids Res 38, 4040–4051 (2010).
Mason, P. B. & Struhl, K. Distinction and relationship between elongation rate and processivity of RNA polymerase II in vivo. Mol. Cell 17, 831–840 (2005).
Rondon, A. G., Garcia-Rubio, M., Gonzalez-Barrera, S. & Aguilera, A. Molecular evidence for a positive role of Spt4 in transcription elongation. EMBO J. 22, 612–620 (2003).
Liu, C. R. et al. Spt4 is selectively required for transcription of extended trinucleotide repeats. Cell 148, 690–701 (2012).
Cheng, H. M. et al. Effects on murine behavior and lifespan of selectively decreasing expression of mutant huntingtin allele by supt4h knockdown. PLoS Genet 11, e1005043 (2015).
Cabrera, J. R. & Lucas, J. J. MAP2 Splicing is Altered in Huntington’s Disease. Brain Pathol. 27, 181–189 (2017).
Dunnett, S. B. & Rosser, A. E. Cell therapy in Huntington’s disease. NeuroRx 1, 394–405 (2004).
Boer, G. J. Ethical guidelines for the use of human embryonic or fetal tissue for experimental and clinical neurotransplantation and research. Network of European CNS Transplantation and Restoration (NECTAR). J. Neurol. 242, 1–13 (1994).
Hurelbrink, C. B. et al. Long-term hibernation of human fetal striatal tissue does not adversely affect its differentiation in vitro or graft survival: implications for clinical trials in Huntington’s disease. Cell Transpl. 12, 687–695 (2003).
Kopyov, O. V., Jacques, S., Lieberman, A., Duma, C. M. & Eagle, K. S. Safety of intrastriatal neurotransplantation for Huntington’s disease patients. Exp. Neurol. 149, 97–108 (1998).
Philpott, L. M. et al. Neuropsychological functioning following fetal striatal transplantation in Huntington’s chorea: three case presentations. Cell Transpl. 6, 203–212 (1997).
Sauer, H. & Brundin, P. Effects of cool storage on survival and function of intrastriatal ventral mesencephalic grafts. Restor. Neurol. Neurosci. 2, 123–135 (1991).
Hoang, T. Q. et al. Quantitative proton-decoupled 31P MRS and 1H MRS in the evaluation of Huntington’s and Parkinson’s diseases. Neurology 50, 1033–1040 (1998).
Tabar, V. & Studer, L. Pluripotent stem cells in regenerative medicine: challenges and recent progress. Nat. Rev. Genet 15, 82–92 (2014).
Nguyen, G. D., Gokhan, S., Molero, A. E. & Mehler, M. F. Selective roles of normal and mutant huntingtin in neural induction and early neurogenesis. PLoS One 8, e64368 (2013).
Nguyen, G. D., Molero, A. E., Gokhan, S. & Mehler, M. F. Functions of huntingtin in germ layer specification and organogenesis. PLoS One 8, e72698 (2013).
Conforti, P. et al. Lack of huntingtin promotes neural stem cells differentiation into glial cells while neurons expressing huntingtin with expanded polyglutamine tracts undergo cell death. Neurobiol. Dis. 50, 160–170 (2013).
Ritch, J. J. et al. Multiple phenotypes in Huntington disease mouse neural stem cells. Mol. Cell Neurosci. 50, 70–81 (2012).
Faideau, M. et al. In vivo expression of polyglutamine-expanded huntingtin by mouse striatal astrocytes impairs glutamate transport: a correlation with Huntington’s disease subjects. Hum. Mol. Genet 19, 3053–3067 (2010).
Jiang, R., Diaz-Castro, B., Looger, L. L. & Khakh, B. S. Dysfunctional Calcium and Glutamate Signaling in Striatal Astrocytes from Huntington’s Disease Model Mice. J. Neurosci. 36, 3453–3470 (2016).
Bradford, J. et al. Expression of mutant huntingtin in mouse brain astrocytes causes age-dependent neurological symptoms. Proc. Natl Acad. Sci. USA 106, 22480–22485 (2009).
Juopperi, T. A. et al. Astrocytes generated from patient induced pluripotent stem cells recapitulate features of Huntington’s disease patient cells. Mol. Brain 5, 17 (2012).
Szlachcic, W. J., Switonski, P. M., Krzyzosiak, W. J., Figlerowicz, M. & Figiel, M. Huntington disease iPSCs show early molecular changes in intracellular signaling, the expression of oxidative stress proteins and the p53 pathway. Dis. Model Mech. 8, 1047–1057 (2015).
Victor, M. B. et al. Striatal neurons directly converted from Huntington’s disease patient fibroblasts recapitulate age-associated disease phenotypes. Nat. Neurosci. 21, 341–352 (2018).
Lu, B. & Palacino, J. A novel human embryonic stem cell-derived Huntington’s disease neuronal model exhibits mutant huntingtin (mHTT) aggregates and soluble mHTT-dependent neurodegeneration. FASEB J. 27, 1820–1829 (2013).
Nekrasov, E. D. et al. Manifestation of Huntington’s disease pathology in human induced pluripotent stem cell-derived neurons. Mol. Neurodegener. 11, 27 (2016).
Jeon, I. et al. Neuronal properties, in vivo effects, and pathology of a Huntington’s disease patient-derived induced pluripotent. Stem Cells Stem Cells 30, 2054–2062 (2012).
Kramer, N. J. et al. Spt4 selectively regulates the expression of C9orf72 sense and antisense mutant transcripts. Science 353, 708–712 (2016).
Lee, S. et al. Repurposing the Cord Blood Bank for Haplobanking of HLA-Homozygous iPSCs and Their Usefulness to Multiple Populations. Stem Cells 36, 1552–1566 (2018).
Lee, J. S. et al. Evaluation of residual toxicity of hypochlorite-treated water using bioluminescent microbes and microalgae: Implications for ballast water management. Ecotoxicol. Environ. Saf. 167, 130–137 (2019).
Slow, E. J. et al. Selective striatal neuronal loss in a YAC128 mouse model of Huntington disease. Hum. Mol. Genet 12, 1555–1567 (2003).
Menalled, L. et al. Systematic behavioral evaluation of Huntington’s disease transgenic and knock-in mouse models. Neurobiol. Dis. 35, 319–336 (2009).
West, M. J., Slomianka, L. & Gundersen, H. J. Unbiased stereological estimation of the total number of neurons in thesubdivisions of the rat hippocampus using the optical fractionator. Anat. Rec. 231, 482–497 (1991).
Acknowledgements
This work was supported by Korean Health Technology R&D Project (NRF-2017M3A9B4061407, NRF-4792018R1C1B6008671, 2018M3A9H3020844) from the National Research Foundation of Korea.
Author information
Authors and Affiliations
Contributions
H.J.P. designed the study, performed the majority of experiments, analyzed the data, wrote the manuscript, and provided financial support; A.H. prepared unedited or SUPT4H1-edited Q57 HD iPSC-NPCs for the transplantation experiment and performed some in vitro experiments; J.C. and J.Y.K. helped with the animal experiments and discussed the results; S.J.K. discussed and interpreted the results; H.S.B., G.C., H.S., E.J.S., and K.L. constructed gRNA and analyzed the gene-editing experiments; J.Y.L. and J.S. supervised the entire study, provided financial support, wrote the manuscript, and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.S. is the founder and CEO of iPS Bio, Inc. H.S.B., G.C., H.S., E.J.S., K.L, S.J.K. and J.Y.L. are the employees of ToolGen, Inc. The other authors declare that they have no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Park, H.J., Han, A., Kim, J.Y. et al. SUPT4H1-edited stem cell therapy rescues neuronal dysfunction in a mouse model for Huntington’s disease. npj Regen Med 7, 8 (2022). https://doi.org/10.1038/s41536-021-00198-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41536-021-00198-0
- Springer Nature Limited
This article is cited by
-
Genetic correction of induced pluripotent stem cells from a DFNA36 patient results in morphologic and functional recovery of derived hair cell-like cells
Stem Cell Research & Therapy (2024)
-
Lowering mutant huntingtin by small molecules relieves Huntington’s disease symptoms and progression
EMBO Molecular Medicine (2024)