Abstract
Zika virus (ZIKV), an arbovirus transmitted by mosquitoes, was identified as a cause of congenital disease during a major outbreak in the Americas in 2016. Vaccine design strategies relied on limited available isolate sequence information due to the rapid response necessary. The first-generation ZIKV mRNA vaccine, mRNA-1325, was initially generated and, as additional strain sequences became available, a second mRNA vaccine, mRNA-1893, was developed. Herein, we compared the immune responses following mRNA-1325 and mRNA-1893 vaccination and reported that mRNA-1893 generated comparable neutralizing antibody titers to mRNA-1325 at 1/20th of the dose and provided complete protection from ZIKV challenge in non-human primates. In-depth characterization of these vaccines indicated that the observed immunologic differences could be attributed to a single amino acid residue difference that compromised mRNA-1325 virus-like particle formation.
Similar content being viewed by others
Introduction
Zika virus (ZIKV), an arbovirus transmitted by mosquitoes, is a member of the Flaviviridae family, which includes yellow fever, dengue, and West Nile viruses1. ZIKV was first identified in 1947, and, until 2007, caused sporadic infections across Africa. In 2007, a large-scale epidemic occurred on Yap Island, followed by a larger outbreak in French Polynesia in 20132. Complications associated with ZIKV infection during pregnancy, including microcephaly and other congenital abnormalities in the developing fetus and newborn3, were reported during the 2015 outbreak of ZIKV in the Western Hemisphere and led the World Health Organization to declare ZIKV a Public Health Emergency of International Concern in 20164. In addition to maternal-fetal transmission5,6,7, sexual transmission of ZIKV has been reported8. Based upon prior genetic and phylogenetic analyses, 2 ZIKV lineages have been identified and classified as African and Asian, with the latter traced to recent outbreaks9,10,11.
ZIKV contains an 11-kilobase, non-segmented, positive-sense, single-stranded RNA genome. The genome contains 5′ and 3′ untranslated regions flanking a single open reading frame (ORF) that encodes a single polyprotein that is cleaved into three structural proteins (capsid [C], premembrane/membrane [prM], and envelope [E]) and 7 non-structural proteins (NS1, NS2A, NS2B, NS3, NS4A, NS4B, and NS5)12. Within the secretory pathway, prM is cleaved by the host furin protease during maturation to produce M and pr. When the virion exits the cell and enters the neutral pH of the extracellular environment, pr is released from the virion13,14. E is the major antigenic determinant of the virus and mediates receptor binding and virus-cell membrane fusion during entry15,16. Since E is the primary target for neutralizing antibodies (nAbs), it is the antigen targeted by most vaccines17. Expression of prME in mammalian cells produces non-infectious virus-like particles (VLPs) that share antigenic features with infectious virions18,19; VLPs retain key properties of viruses, including fusogenic activity20 and the induction of a nAb response.
As the ZIKV epidemic required a rapid public health response, vaccine design strategies relied on the limited isolate sequence information available at the time. The public bank of full-length human ZIKV sequences contained 8 sequences in 2014; this increased to 30 by March 201621. The first-generation mRNA vaccine, mRNA-1325 (Moderna, Inc., Cambridge, MA), encodes the prME ORF from a Micronesia 2007 ZIKV isolate22, with an upstream immunoglobulin E (IgE) leader sequence to drive secretion. As additional sequences became available, a second mRNA vaccine was generated, mRNA-1893, which encodes the contemporary RIO-U1 Zika prME ORF. RIO-U1 was isolated from the urine of a pregnant woman with symptoms of an acute phase ZIKV infection21,23. mRNA-1893 encodes a Japanese encephalitis virus (JEV)–derived leader sequence, which has been reported to improve host signalase cleavage18,24. Aside from the different leader sequence, mRNA-1325 and mRNA-1893 have 5 amino acid residue differences in the prME ORF (2 in the pr region; 3 in the E gene).
Here, we compared mRNA-1325 and mRNA-1893 and showed that mRNA-1893 elicited similar nAb levels at lower doses compared with mRNA-1325 and provided complete protection from ZIKV challenge in non-human primates (NHPs). Through further characterization of mRNA-1893 and mRNA-1325, we identified a single amino acid residue difference that compromises mRNA-1325 VLP formation and may underlie the immunologic differences observed between the vaccines.
Results
mRNA-1893 generates VLPs in vitro
mRNA-1893 and mRNA-1325 were formulated in ionizable lipid-based nanoparticles (LNPs) and tested for intracellular expression levels of prME and differences in VLP production following in vitro transfection. The antibody ZV-116 was used to detect E protein in cell lysates and VLPs purified from supernatant25. E protein was detected in cell lysates from mRNA-1325 and mRNA-1893, and in mRNA-1893 supernatant (Fig. 1a). Bands corresponding to the approximate molecular weight of uncleaved prM were detected for mRNA-1325 and mRNA-1893 but were more prominent for mRNA-1893 (Fig. 1b). In addition, prM was not detected in supernatant, but a band of the approximate size of cleaved M protein was present in mRNA-1893 supernatant, suggesting the production of mature VLPs. An additional band above 50 kD was present in all cell lysates, including the control, reflecting a non-specific band.
a Cell lysates and VLP pellets were blotted for E protein using ZV-116 antibody. Actin was used as a loading control for cell lysate samples. b Cell lysates and VLP pellets were blotted for prM and M proteins using an anti-Zika prM polyclonal antibody. a, b Molecular weights were estimated using Precision Plus Dual Color Standards (Bio-Rad). c VLP secretion was measured by sandwich ELISA. Data are representative of a single experiment, with error bars indicating the SD of technical repeats on the same plate (ie, triplicate wells). d For negative staining and immunogold labeling, ZV-116 at 30,000× magnification (left) or ZV-117 at 50,000× magnification (right) was used, followed by the use of a gold-labeled anti-human secondary antibody. Scale bar indicates 100 nm. E envelope, ELISA enzyme-linked immunosorbent assay, M membrane, prM premembrane/membrane, SD standard deviation, VLP virus-like particle.
To differentiate between monomeric E protein and assembled VLPs, a sandwich enzyme-linked immunosorbent assay (ELISA) was performed using the anti-E antibodies ZV-116 or ZV-117 (for capture) and ZV-67 (for detection). Both ZV-116 and ZV-67 recognize a linear epitope in the Domain III lateral ridge26 and recognize free E monomers or VLP-associated E protein. ZV-117 binds across an E protein dimer and preferentially recognizes E protein incorporated into VLPs, although it can also weakly bind to E protein monomer27. Supernatants collected 48 hours after transfection were added to plates coated with either ZV-116 or ZV-117 (Fig. 1c). E protein was detected in supernatant following mRNA-1893 transfection using ZV-116 or ZV-117. However, levels of supernatant E protein following mRNA-1325 transfection were comparable to the untransfected controls with both antibodies, suggesting that mRNA-1325 does not generate secreted VLPs in vitro.
Transmission electron microscopy and negative staining, used to characterize VLP composition generated after mRNA-1893 transfection, revealed that VLPs had a roughly spherical structure, with an average diameter of 50 to 100 nm. Particles were imaged by immunogold labeling using ZV-116 and ZV-117 antibodies. VLPs were reactive with both antibodies (Fig. 1d), consistent with the ELISA results. Collectively, the data demonstrate that mRNA-1893 transfection in vitro results in the formation and secretion of VLPs.
Intracellular E protein generated by mRNA-1325 has a short half-life and is rapidly degraded by the proteasome
In vitro experiments were conducted to examine the apparent absence of VLPs or E protein dimers after mRNA-1325 transfection despite detectable E protein expression, albeit at lower levels than those seen for mRNA-1893 (Fig. 1a). As rapid protein degradation could contribute to the lack of VLP generation, protein half-life was examined. Intracellular E protein was detected by ZV-116 (Fig. 2a) and a pan-flavivirus antibody 4G2 (Fig. 2b)16,28. While mRNA-1893 protein had a half-life of ≥6 hours, mRNA-1325 had a relatively short half-life of <2 hours (Fig. 2a). In addition, no E protein was detected at any time after transfection for mRNA-1325 using 4G2, suggesting that protein produced by mRNA-1325 could be misfolded (Fig. 2b). To further elucidate the pathway involved in the accelerated E protein degradation, 293T cells transfected with mRNA-1325 were treated with cycloheximide (CHX) plus chloroquine or CHX plus MG132 to inhibit lysosomal proteolysis or ubiquitin-mediated proteasomal degradation pathways, respectively. Chloroquine had no effect on E protein half-life (Fig. 2c), whereas MG132 extended E protein half-life ≥8 hours after transfection (Fig. 2d). These results suggest that mRNA-1325-expressed E protein is at least partially misfolded and undergoes rapid proteasomal degradation, thus leading to loss of VLP production.
a E protein expression was assessed in cell lysates from mRNA-transfected cells, following the addition of CHX and using antibody ZV-116. b E protein levels were also assayed, following the addition of CHX and using the anti-flavivirus fusion loop antibody 4G2, under non-reducing conditions. c 293T cells were transfected with mRNA-1325 and were treated with CHX or CHX + CQ, harvested at the indicated time points following treatment, and blotted for E using ZV-116. d 293T cells were transfected with mRNA-1325 and were treated with CHX or CHX + MG132 to block proteasomal degradation. a–d Molecular weights were estimated using Precision Plus Dual Color Standards (Bio-Rad). CHX, cycloheximide; CQ, chloroquine; E, envelope.
Alanine to threonine substitution at residue 40 of the E protein in mRNA-1325 rescues VLP production in vitro and enhances nAb titers in mice
mRNA-1325 and mRNA-1893 differ in signal peptides (IgE and JEV, respectively) and in 5 amino acid residues (valine, serine, alanine, valine, and threonine in mRNA-1325 substituted for alanine, asparagine, threonine, methionine, and methionine in mRNA-1893, respectively) in the prME ORF (Fig. 3a). To investigate whether these differences contributed to the VLP production defect of mRNA-1325, a panel of mRNA-1325 sequence variants was generated. Due to their proximity to the amino terminus and potential to affect VLP production, the residues pr1 and E40 were targeted to generate the following variants: IgEsp-pr(Val1Ala); IgEsp-E(Ala40Thr); and IgEsp-pr(Val1Ala) + E(Ala40Thr). To exclude the possibility of the leader sequence affecting VLP production, IgEsp-RIO-U1 was generated, which contains the entire prME ORF from mRNA-1893 but retains the IgEsp from mRNA-1325.
a mRNA-1325 and mRNA-1893 differ in signal sequences and 5 amino acid residues within the prME ORF. b Cell lysates and VLP pellets were blotted for E protein using ZV-116 antibody. GAPDH was used as a loading control. Molecular weights were estimated using Precision Plus Dual Color Standards (Bio-Rad). c VLP secretion from transfected cells was measured in a sandwich ELISA with ZV-117 antibody, followed by ZV-67 antibody, and an anti-mouse HRP-conjugated secondary antibody. d Colored bars represent GMT for the group, and black circles represent individual animals (n = 8 per group). Error bars indicate SD from the GMT. Dashed line indicates assay LOD. E, envelope; EC50, half-maximal effective concentration; ELISA, enzyme-linked immunosorbent assay; GMT, geometric mean titer; HRP, horseradish peroxidase; IgE, immunoglobulin E; JEV, Japanese encephalitis virus; LOD, limit of detection; ORF, open reading frame; prME, premembrane/membrane and envelope; RVP, reporter virus particle; SD, standard deviation; UTR, untranslated region; VLP, virus-like particle; ZIKV, Zika virus.
mRNA-1325, mRNA-1893, and the variants were transiently transfected into 293T cells, and lysates and supernatant were assayed for E protein expression and VLP production. All variants rescued VLP production to levels detectable by Western blot (Fig. 3b). A sandwich ELISA using ZV-117 and ZV-67 antibodies demonstrated that mRNA-1893, IgEsp-RIO-U1, and IgEsp-E(Ala40Thr) produced similar levels of secreted VLPs (Fig. 3c).
To determine whether the rescue of VLP production in vitro was correlated with an increase in immunogenicity in vivo, C57BL/6 mice were immunized intramuscularly at days 1 and 29 with 2 µg or 0.4 µg LNPs containing mRNA-1325, mRNA-1893, or variants. Seven of 8 mice immunized with mRNA-1325 2 µg had nAb titers below the assay detection limit. Conversely, animals immunized with mRNA-1893 2 µg or 0.4 µg or sequence variants IgEsp-RIO-U1 or IgEsp-E(Ala40Thr) had half-maximal effective concentration (EC50) values of >1/103, with most animals having titers between 1/104 and 1/105 (Fig. 3d). These data indicated that the VLP production defects and lower immunogenicity observed with mRNA-1325 are not due to the IgEsp leader sequence, but instead can be mapped to the presence of alanine at E40 instead of threonine.
mRNA-1893 shows superior immunogenicity and efficacy compared with mRNA-1325 in rhesus macaques
To evaluate the immunogenicity and efficacy of mRNA-1325 and mRNA-1893 in a challenge model, both mRNA constructs were formulated in ionizable LNPs and administered intramuscularly to rhesus macaques. mRNA-1325 was tested at 3 concentrations (10, 50, and 200 µg) following a 2-dose regimen (days 1, 29). mRNA-1325 was also tested as a single dose, in which the highest concentration (200 µg) was administered only on day 1. In addition, mRNA-1893 was tested at 1 concentration (10 µg) following the 2-dose regimen (days 1, 29). A non-coding mRNA formulated in LNP (control) was administered at 200 µg on days 1 and 29. nAbs were measured using the ZIKV RVP neutralization assay29 at week 4 before boost and immediately before challenge at week 8.
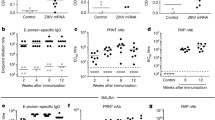
Among mRNA-1325 regimens at week 4, nAb titers were generally higher with the 200-μg regimens than with the 10-μg regimen (Fig. 4a; Supplementary Table 1). At week 8, a dose response was observed for the low- vs high-dose 2-dose regimens of mRNA-1325 (10 µg vs 200 µg; P < 0.001; one-way analysis of variance (ANOVA)); mean nAb titers for the 10-, 50-, and 200-µg doses were 1697, 4347, and 7659, respectively. When comparing the 1- vs 2-dose regimens at week 8, mRNA-1325 200 µg generated significantly higher nAb titers when administered as a 2-dose compared with a 1-dose regimen (P < 0.05; one-way ANOVA). Comparisons between constructs at the same dose at week 8 showed a significantly lower nAb response for mRNA-1325 compared with mRNA-1893 when the vaccines were administered as a 10-µg, 2-dose regimen (P < 0.05; one-way ANOVA). Surprisingly, nAb titers in mRNA-1893 10-µg, 2-dose regimen recipients were comparable to titers in mRNA-1325 200-µg, 2-dose regimen recipients (6479 vs 7659, respectively; P ≈ 0.97; one-way ANOVA), suggesting that mRNA-1893 generated similar levels of nAbs at 1/20th of the dose and was consistent with the increase in VLP formation observed in the in vitro experiments.
a nAb titers were measured using the ZIKV RVP assay on week 4 (day 28) and week 8 (day 56). Colored bars represent GMTs for each group (mean ± SD) and black circles represent individual animals (n = 5 per group). Dashed line indicates assay LOD. Statistical analysis was performed using one-way ANOVA. *P < 0.05, **P < 0.005, ***P < 0.001. b Viremia was detected via qPCR on days 3, 4, 5, and 7 following challenge. Dashed line indicates assay LOD. ANOVA, analysis of variance; EC50, half-maximal effective concentration; FFU, focus forming units; GMT, geometric mean titer; LOD, limit of detection; nAb, neutralizing antibody; NHP, non-human primate; qPCR, quantitative polymerase chain reaction; RVP, reporter virus particle; VLP, virus-like particle; ZIKV, Zika virus.
All groups were challenged with 1 × 103 plaque-forming units of ZIKV (strain PRVABC59) delivered subcutaneously on day 56 and assayed for viremia by quantitative reverse transcription polymerase chain reaction on days 3, 4, 5, and 7 thereafter. All animals in the non-coding mRNA control group and ≥1 animal from each of the mRNA-1325 groups had detectable viremia (Fig. 4b). No viremia was detected in animals in the mRNA-1893 group following challenge.
To evaluate whether mRNA-1893 vaccination generated sterilizing immunity, nAb titers were evaluated 14 and 28 days following challenge for individual animals in each dose group (Fig. 5). EC50 values for ≥1 animal in each of the mRNA-1325 groups had a ≥ 4-fold anamnestic response. All animals that had a ≥4-fold increase also had detectable viremia following challenge (Fig. 5). No animals vaccinated with mRNA-1893 had a 4-fold increase in neutralization activity samples analyzed post-challenge. Thus, immune responses elicited by mRNA-1893 rapidly control infection, thereby preventing an anamnestic response. Collectively, these data indicate that mRNA-1893 generates more robust protection at doses substantially lower than mRNA-1325 in NHPs.
Data are plotted as the EC50 fold change relative to the time of challenge (week 8), which was set to 1. Tables below each graph indicate whether the virus was detected on any day following the challenge and if the EC50 value had a 4-fold or greater increase following the challenge. n = 5 per group. EC50, half-maximal effective concentration; FC, fold change; nAb, neutralizing antibody; NHP, non-human primate; RVP, reporter virus particle; ZIKV, Zika virus.
ZIKV CprME–based mRNA constructs produce C-containing VLPs in vitro and are immunogenic in mice
The ZIKV ORF encodes the structural proteins C, prM, and E30,31,32. Expression of prME is sufficient for VLP formation, but it is possible that C-containing VLPs (CprME) are more immunogenic than prME-only VLPs32,33,34. Two CprME-encoding constructs were generated, including 1 encoding wild-type RIO-U1 Zika CprME ORF, and compared with mRNA-1893, both for VLP production in vitro and nAb responses in mice. As the production of CprME VLPs requires C-prM cleavage by NS2B3 protease30,31,32, we generated a second mRNA encoding RIO-U1 NS2B3. As an alternative method of generating cleaved C, a C-2A-prME construct was designed with a ribosome 2A stop-go sequence between C and prM. Insertion of this sequence should result in the release of C from the nascent polyprotein during translation, eliminating the need for an NS2B3-mediated cleavage step35.
The mRNA constructs encoding wild-type CprME, either alone or with NS2B3 mRNA, and C-2A-prME (alone) were transfected into 293T cells side-by-side with mRNA-1893 and variants IgEsp-RIO-U1 and IgEsp-E(Ala40Thr). VLP production was compared across constructs by purifying VLPs from supernatant and Western blotting for E and prM proteins (Fig. 6a, b). Both C-containing constructs produced VLPs, though as expected, the CprME construct required the co-transfection of NS2B3. The CprME/NS2B3 combination resulted in a higher amount of detectable E protein than any of the prME-based constructs or C-2A-prME. In addition, the presence of the low molecular weight band representing cleaved M protein indicates that the VLPs produced by these C constructs are mature (Fig. 6a). To investigate whether the VLPs did in fact contain C, cell lysates and purified VLPs from transfected cells were blotted for the C protein (Fig. 6b). The C protein was detected in the lysates of both constructs, with C-2A-prME producing a slightly higher molecular weight protein due to the 19-amino acid residue tail generated by the 2A sequence. The C protein was also detected in the purified VLPs from CprME/NS2B3 co-transfection, but not transfection of C-2A-prME. As the E protein Western blot on purified VLPs indicates that the C-2A-prME construct produces VLPs, the lack of signal could be due to an overall lower level of expression and VLP production of C-2A-prME compared with CprME/NS2B3, or to the lack of packaging of C protein, resulting in prME-only VLPs (Fig. 6a).
a The VLP pellet was blotted for E protein using ZV-116 (top) and prM and M proteins were blotted using an anti-Zika prM polyclonal antibody (bottom). Arrows indicate prM and M. Molecular weights were estimated using a mixture of MagicMark XP Western Protein Standard (Invitrogen) and Precision Plus Dual Color Standards (Bio-Rad, cat. no. 1610374) to obtain reference bands every 5 to 10 kDa. b Purified VLPs (top) and lysates from transfected cells (bottom) were blotted for C protein using an anti-Zika capsid polyclonal antibody. (c) VLP secretion was measured by sandwich ELISA, with either ZV-116 (left) or ZV-117 (right), followed by the addition of the anti-E antibody ZV-67 and an anti-mouse HRP-conjugated secondary antibody. Data are representative of a single experiment, with error bars indicating the SD of technical repeats on the same plate (ie, triplicate wells). d Cells were transfected with CprME + NS2B3 mRNAs. Concentrated VLPs were stained with uranyl acetate and labeled with ZV-116 at 30,000× magnification, followed by a gold-labeled anti-human secondary antibody. Scale bar indicates 100 nm. e nAb titers were measured using the ZIKV RVP neutralization assay on day 56 following 2 doses of mRNA administered on day 1 and day 29. Colored bars represent GMT for each group (mean ± SD), and black circles represent individual animals (n = 8 per group). Dashed line indicates assay LOD. Statistical analysis was performed using two-way ANOVA. *P < 0.05, **P < 0.005, ***P < 0.001. ANOVA, analysis of variance; C, capsid; CprME, capsid, premembrane/membrane, and envelope; E, envelope; EC50, half-maximal effective concentration; ELISA, enzyme-linked immunosorbent assay; GMT, geometric mean titer; HRP, horseradish peroxidase; LOD, limit of detection; M, membrane; nAb, neutralizing antibody; prM, premembrane/membrane; prME, premembrane/membrane and envelope; RVP, reporter virus particle; SD, standard deviation; VLP, virus-like particle; ZIKV, Zika virus.
To differentiate between monomeric E protein and assembled VLPs, supernatant was run in the ZV-116 and ZV-117 sandwich ELISAs using the anti-E antibodies ZV-116 or ZV-117 (for capture) and ZV-67 (for detection) (Fig. 6c). Both ZV-116 and ZV-117 ELISAs detected supernatant E protein from transfection of CprME/NS2B3 and C-2A-prME, but not CprME alone. VLPs were detected for both C-containing designs and prME-only constructs. The relative quantity of CprME/NS2B3 VLPs in relation to prME VLPs varied between the ELISA and Western blot results, with SN VLP levels relatively similar between CprME/NS2B3 and prME constructs in the ELISA, in contrast to the predominance of CprME/NS2B3 E and M bands in the Western blot (Fig. 6a). As C protein stability is important for the assembly of infectious flavivirus virions36, the inclusion of C may allow CprME VLPs to better withstand the purification process than prME-only particles, and therefore, give the appearance of increased VLP production. However, as the same purification conditions were used for all constructs, we do not exclude the possibility that the purification conditions were suboptimal for prME VLPs and that the lower yield of prME VLPs observed post-purification is due to the experimental protocol rather than inherent stability differences between CprME and prME VLPs. Electron microscopy confirmed the presence of VLPs from cells transfected with CprME + NS2B3 mRNAs (Fig. 6d).
To determine whether the CprME VLPs were better immunogens than prME-only VLPs, C57BL/6 mice were immunized intramuscularly with LNPs containing mRNA-1893/non-coding control, CprME/NS2B3, or C-2A-prME/non-coding control. Each combination contained a mass ratio of 2 parts structural protein mRNA to 1 part non-coding or NS2B3 mRNA; 3 or 0.6 µg of total mRNA were administered. Animals received 2 doses 28 days apart and were phlebotomized at day 56 for serum nAb analysis using ZIKV RVPs. All constructs at the high dose (3 µg) generated robust and similar nAb titers (P > 0.05; two-way ANOVA; Fig. 6e; Supplementary Table 2). When comparing constructs at the low dose (0.6 µg), CprME/NS2B3 elicited higher nAb titers than the C2AprME construct (P < 0.005; two-way ANOVA). Titers were similar between mRNA-1893 and the other constructs when comparing within the 3- and 0.6-µg dose levels (P > 0.05; two-way ANOVA). Within each construct, both mRNA-1893 and C-2A-prME produced significant dose-dependent effects (P < 0.01 and <0.001, respectively; two-way ANOVA).
Discussion
VLP-based vaccines offer several advantages compared with other modalities, such as monomeric proteins or whole-inactivated virus; these advantages include safety due to a lack of genetic material and an inability to replicate, and elicitation of strong immune responses due to native multi-array antigen presentation and efficient B-cell receptor engagement37,38. Delivery of VLPs by mRNA or other gene-based vectors adds the element of CD8 T cell induction not afforded by protein vaccines. ZIKV VLPs consisting of prME or CprME have been employed as the vaccine antigen across multiple vaccine platforms, including DNA, adenovirus-vectored, and purified particles18,19,39,40,41,42,43,44. Herein, we assessed the immune response elicited by mRNA-1325, which encodes the prME ORF from a Micronesia 2007 ZIKV isolate, and mRNA-1893, which encodes the contemporary RIO-U1 Zika prME ORF and differs from mRNA-1325 by 5 residues and the leader sequence.
This study suggested that the lower immunogenicity of mRNA-1325 compared with mRNA-1893 may arise from a defect in VLP production and secretion. In vitro characterization suggested that, although mRNA-1325 and mRNA-1893 expressed E protein intracellularly, E protein was only detectable in supernatant with transfection of mRNA-1893. Supernatant ELISAs and negative staining of purified particles, both using the VLP-specific antibody ZV-117, suggested that the secreted E protein generated by mRNA-1893 was in the form of a VLP. Protein half-life experiments demonstrated that E protein produced by mRNA-1325 has a shorter half-life than that produced by mRNA-1893 and is rapidly degraded by the proteasome. This rapid degradation may be due to partial misfolding, as the conformationally dependent antibody 4G2 did not detect E protein expression in cells transfected with mRNA-1325. While these in vitro defects may appear severe, the NHP data suggest that mRNA-1325 generates immunogens in vivo that elicit an antibody response.
The IgEsp-E(Ala40Thr) construct rescued VLP production to levels comparable to mRNA-1893 in vitro and generated nAb titers equivalent to mRNA-1893 in vaccinated mice, suggesting that the lower protection from ZIKV challenge afforded by mRNA-1325 compared with mRNA-1893 may be due to the Ala40Thr substitution in the E protein ORF. Of note, of the 1,509 ZIKV E protein sequences with data for position 40 in the Virus Pathogen Database and Analysis Resource (as of July 2022), most sequences contained Thr40 and only 1 sequence contained Ala40 (accession number EU545988); the latter sequence was used for the design of mRNA-1325. Ala40 may be detrimental to normal E protein function, explaining the lower protection afforded by mRNA-1325. Although it is unclear why this substitution would be deleterious to VLP production, this may be due to C-terminal serine endopeptidase cleavage between Lys41 and Pro42. When Pro is located at the C-terminal side of Lys, its ring structure restricts the freedom of rotation, making the polypeptide backbone more rigid. Lys-Pro bonds are typically cleavage-resistant but can be cleaved if the amino acid residues flanking the Lys-Pro sequence have small side chains that allow flexibility of the peptide bonds45. The Lys-Pro-Ala sequence of mRNA-1325 may make the Lys-Pro bond accessible to proteases, whereas in the Lys-Pro-Thr sequence of mRNA-1893, the bulky Thr side chain compared with Ala may prevent the torsion necessary for protease accessibility and cleavage.
Previous studies using a purified VLP vaccine platform reported that ZIKV C-containing VLPs are immunogenically superior to prME-only VLPs39,40. Here, we compared mRNA-1893 with 2 alternative designs that produce C-containing VLPs. The CprME ORF was encoded on a single mRNA, with cleavage of C from the downstream prME polyprotein achieved by the addition of a second mRNA encoding the NS2B3 protease or a 2A ribosome stop-go sequence between C and prME. Transfection of either CprME design resulted in production of secreted, C-containing VLPs that were detectable by the VLP-specific antibody ZV-117. Interestingly, although ELISAs conducted with neat supernatant did not reveal differences in the level of VLP production, CprME/NS2B3 transfection repeatedly resulted in a higher level of VLPs in any assay involving ultracentrifugation. Potential explanations for this observation include variation in particle size, centrifugation conditions, and/or the ability of particles to withstand purification. However, based exclusively on nAb titers, the inclusion of C in the VLP had a marginal effect on nAb titers in mice, as vaccination with equivalent doses of prME-only mRNA-1893 versus either of the 2 C-encoding mRNA constructs resulted in similar nAb titers; additional differences may bear out in challenge studies. These in vivo results, coupled with the observation that CprME-containing VLPs may be more resilient to ultracentrifugation, suggest that the differences between CprME and prME-only VLPs as immunogens may be influenced by vaccine platform. Of note, the addition of C to mRNA constructs may have practical implications, such as reduced efficiency in mRNA vaccine manufacturing due to the increased number of nucleotides46.
Here, we provided evidence that mRNA-1893 is superior to mRNA-1325 in the rhesus macaque challenge model. Prime and boost vaccination with mRNA-1893 10 µg generated nAb titers comparable to those elicited by mRNA-1325 200 µg. In addition, no animals immunized with mRNA-1893 had detectable viral load at any time following challenge, whereas ≥1 animal in each of the mRNA-1325 dose groups exhibited viral breakthrough. Finally, no animals vaccinated with mRNA-1893 showed a boost in nAb titers following challenge, while ≥1 animal in each dose group of mRNA-1325 had a ≥ 4-fold increase in nAb titers at either 2 or 4 weeks following challenge, indicating induction of an anamnestic response. The lack of an anamnestic response and absence of detectable viremia in the animals receiving mRNA-1893 10 µg is indicative of rapid viral control. The viral breakthrough in animals vaccinated with mRNA-1325 despite comparable nAb titers between mRNA-1325 and mRNA-1893 could be due to differences in immune mediators like T-cell responses, antibody-dependent cell-mediated cytotoxicity, antibody-dependent cellular phagocytosis that are not captured by neutralization assays, or other qualitative differences in the total antibody response. Indeed, previous reports have noted that antibody responses to DNA vaccine candidates expressing ZIKV prME are variably sensitive to prM content47; future studies should investigate antibodies against mature forms of the virion, as this may be more predictive than other neutralization assays.
In summary, results indicated that mRNA-1893 is superior to mRNA-1325. In addition to providing complete protection from ZIKV challenge in NHPs, mRNA-1893 generates higher nAb titers in mice than mRNA-1325 when given at an equivalent dose and elicits comparable titers at 1/20th of the dose in NHPs. This enhanced immunogenicity may arise from a single amino acid residue difference in the E protein ORF of mRNA-1893, given that an Ala40Thr substitution rescues VLP secretion and restores immunogenicity of mRNA-1325 to levels comparable to mRNA-1893. Finally, CprME-encoding mRNA constructs do not outperform mRNA-1893 in mouse immunogenicity studies, suggesting that the prME-only VLPs produced by mRNA-1893 are not immunogenically inferior to C-containing VLPs. The data generated through preclinical animal models and in vitro characterization experiments support the advancement of mRNA-1893 into clinical trials. In a phase 1 trial (NCT04064905), mRNA-1893 had an acceptable safety profile and elicited strong anti-ZIKV serum nAb and binding antibody responses that persisted up to 1 year after vaccination48. Results from a phase 1 trial (NCT03014089) showed that mRNA-1325 elicited poor immune responses; the preclinical data generated here may explain these clinical findings48. Together, these preclinical and clinical results support the continued development of mRNA-1893.
Methods
Generation of modified mRNA and LNPs
Modified mRNA was synthesized using in vitro T7 RNA polymerase-mediated transcription from a linearized DNA template, with N1-methyl-pseudouridine triphosphate and cap 1 to improve translation efficiency49. Formulations of mRNA-1325 and mRNA-1893 were prepared by dissolving ionizable lipids, distearoylphosphatidylcholine, cholesterol, and polyethylene glycol-lipid in ethanol (molar ratios of 50:10:38:5:1:5). This lipid mixture was combined with mRNA in a citrate buffer (50 mM, pH 4.0) at a ratio of 3:1 (aqueous:ethanol). The formulations were dialyzed against phosphate-buffered saline (pH 7.4) for <18 hours, concentrated, and passed through a 0.22-µm filter before storage at 4 °C. Final particle size and encapsulation were <100 nm and >80%, respectively, with endotoxin below 10 EU/mL50. The preclinical designations for mRNA-1325 and mRNA-1893 are CX-000171 and CX-005809, respectively; the studies described herein were conducted with preclinical material.
Cell lines and antibodies
293T and Vero cells were obtained from the cell bank (Moderna, Inc.) and were cultured in Dulbecco’s modified Eagle’s medium supplemented with 10% fetal bovine serum. Human monoclonal antibodies (mAbs) ZV-116 and ZV-117 were provided by Dr. James Crowe (Vanderbilt University Medical Center, Nashville, TN). Mouse mAb ZV-67 was provided by Dr. Michael Diamond (Washington University School of Medicine, St. Louis, MO). Commercially available flavivirus E protein-specific mAb 4G2 (EMD Millipore, Darmstadt, Germany), rabbit anti-ZIKV prM and C antibodies (GeneTex, Irvine, CA), and goat horseradish peroxidase (HRP)-conjugated anti-human, anti-mouse, and anti-rabbit immunoglobulin G secondary antibodies (Southern Biotech, Birmingham, AL) were used (Table 1).
Collection of cell lysates and VLPs for use in Western blot, ELISA, and electron microscopy analyses
Using the TransIT-mRNA Transfection Kit (Mirus, Madison, WI), 1 × 107 293T cells were transiently transfected with mRNA (20 µg). The supernatant was collected 48 hours after transfection, centrifuged (500 × g) for 5 minutes, layered on a 25% glycerol/Tris/NaCl/EDTA cushion, and centrifuged (110,000 × g) for 2 hours at 4 °C. Supernatants were aspirated and VLP-containing pellets were resuspended in phosphate-buffered saline. Isolated VLPs were analyzed by electron microscopy or Western blot. The corresponding 293T cells were lysed with RIPA buffer 48 hours after transfection. Protein concentrations were measured using a BCA assay kit (ThermoFisher, Waltham, MA). Western blot, ELISA, and electron microscopy experiments were performed using standard methodologies. For Western blots presented in Figs. 1–3, the molecular weight was estimated using Precision Plus Dual Color Standards (Bio-Rad, cat. no. 1610374, Philadelphia, PA). For Western blots presented in Fig. 6, both MagicMark XP Western Protein Standard (Invitrogen, cat. no. LC5602, Waltham, MA) and Precision Plus Dual Color Standards (Bio-Rad, cat. no. 1610374) were used to obtain reference bands approximately every 5 to 10 kDa to determine the molecular weights of the Western blot bands (Supplementary Figure).
Protein half-life experiments
293T cells (6.25 × 105) were transfected with 1 µg of mRNA for 24 hours, followed by treatment with CHX (100 µg/mL) ± chloroquine (50 µM) or MG132 (20 µM, Sigma Aldrich, St. Louis, MO) for varying amounts of time. Cells were lysed with RIPA buffer and blotted using ZV-116 or 4G2 antibodies. When 4G2 was used for Western blot, reducing reagent was omitted.
Immunization studies
Experiments involving animals were carried out in compliance with approval from the Animal Care and Use Committee of Moderna, Inc., and the NIH Vaccine Research Center and BIOQUAL, Inc. (Rockville, MD). The design of the mouse studies followed historical precedent of mRNA vaccines51,52,53. Based on data from mouse experiments, the sample size for NHPs was calculated to be more than 10-fold between groups. A sample size of 5 animals/group provided 88% power to detect a 10-fold difference between groups and 99% power to detect a 15-fold difference.
C57BL/6 female mice (weighing ~20 g) aged 8 weeks (Charles River Laboratories, Wilmington, MA) were inoculated intramuscularly with 50 μL of mRNA/LNP vaccine on day 1 and were immunized again on day 29 (n = 8 per dosing group). Blood, collected on day 56, was analyzed for serum nAbs.
Naive Indian-origin rhesus macaques (weighing 3–5 kg; aged 2–3 years) were inoculated intramuscularly with 500 µL of vaccine LNP constructs and challenged on day 56 with 1 × 103 plaque-forming units of ZIKV (strain PRVABC59; n = 5 per dosing group; 2 male/3 female). Blood was collected at week 4 post-dose 1 and at time of challenge and at indicated time points after the challenge to measure viral load using quantitative reverse transcription polymerase chain reaction and serum nAb titers using the green fluorescent protein-expressing RVP neutralization assay18,29. The limit of detection (LOD) was set as the reciprocal initial dilution of sera (1:60 for NHPs, and 1:400 for mice). Titers measured below the initial dilution were assigned a titer of one-half the LOD (30 for NHPs, and 200 for mice).
Statistical methods
One-way ANOVA was used to compare the effects of mRNA-1325 and mRNA-1893 and the different dosing regimens (1 vs 2 doses) on nAb titers in NHPs. This included testing of the dosing regimen as a single main effect and test article combinations. Two-way ANOVA was used to assess the dose response of CprME VLPs and prME-only VLPs and the effect of dose levels (3 vs 0.6 µg) on immunogenicity, including testing main effects of test articles and the dose levels and their interactions. Šidák correction was used to adjust for Type I error rate of multiple comparisons at an alpha level of 0.05, unless noted otherwise.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Upon request, and subject to review, Moderna, Inc., will provide the data that support the findings of this study.
References
Weaver, S. C. et al. Zika virus: History, emergence, biology, and prospects for control. Antiviral Res. 130, 69–80 (2016).
Cao-Lormeau, V. M. et al. Zika virus, French polynesia, South pacific, 2013. Emerg. Infect. Dis. 20, 1085–1086 (2014).
Broutet, N. et al. Zika virus as a cause of neurologic disorders. N. Engl. J. Med. 374, 1506–1509 (2016).
WHO. The history of Zika virus, https://www.who.int/news-room/feature-stories/detail/the-history-of-zika-virus (2016).
Brasil, P. et al. Zika virus infection in pregnant women in Rio de Janeiro. N. Engl. J. Med. 375, 2321–2334 (2016).
Calvet, G. et al. Detection and sequencing of Zika virus from amniotic fluid of fetuses with microcephaly in Brazil: a case study. Lancet Infect. Dis. 16, 653–660 (2016).
Sarno, M. et al. Zika virus infection and stillbirths: a case of hydrops fetalis, hydranencephaly and fetal demise. PLoS Negl. Trop. Dis. 10, e0004517 (2016).
Hills, S. L. et al. Transmission of Zika virus through sexual contact with travelers to areas of ongoing transmission—Continental United States, 2016. MMWR Morb. Mortal Wkly. Rep. 65, 215–216 (2016).
Faye, O. et al. Molecular evolution of Zika virus during its emergence in the 20(th) century. PLoS Negl. Trop. Dis. 8, e2636 (2014).
Haddow, A. D. et al. Genetic characterization of Zika virus strains: geographic expansion of the Asian lineage. PLoS Negl. Trop. Dis. 6, e1477 (2012).
Lanciotti, R. S., Lambert, A. J., Holodniy, M., Saavedra, S. & Signor Ldel, C. Phylogeny of Zika virus in Western Hemisphere, 2015. Emerg. Infect. Dis. 22, 933–935 (2016).
Liu, L. Fields virology, 6th Edition. Clin. Infect. Dis. 59, 613 (2014).
Pierson, T. C. & Diamond, M. S. Degrees of maturity: the complex structure and biology of flaviviruses. Curr. Opin. Virol. 2, 168–175 (2012).
Yu, I. M. et al. Structure of the immature dengue virus at low pH primes proteolytic maturation. Science 319, 1834–1837 (2008).
Boigard, H. et al. Zika virus-like particle (VLP) based vaccine. PLoS Negl. Trop. Dis. 11, e0005608 (2017).
Crill, W. D. & Chang, G. J. Localization and characterization of flavivirus envelope glycoprotein cross-reactive epitopes. J. Virol. 78, 13975–13986 (2004).
Lindenbach, B. D. & Rice, C. M. Molecular biology of flaviviruses. Adv. Virus Res. 59, 23–61 (2003).
Dowd, K. A. et al. Rapid development of a DNA vaccine for Zika virus. Science 354, 237–240 (2016).
Muthumani, K. et al. In vivo protection against ZIKV infection and pathogenesis through passive antibody transfer and active immunisation with a prMEnv DNA vaccine. NPJ Vaccines 1, 16021 (2016).
Nour, A. M., Li, Y., Wolenski, J. & Modis, Y. Viral membrane fusion and nucleocapsid delivery into the cytoplasm are distinct events in some flaviviruses. PLoS Pathog. 9, e1003585 (2013).
Wang, L. et al. From mosquitos to humans: genetic evolution of Zika virus. Cell Host Microbe 19, 561–565 (2016).
Lanciotti, R. S. et al. Genetic and serologic properties of Zika virus associated with an epidemic, Yap State, Micronesia, 2007. Emerg. Infect. Dis. 14, 1232–1239 (2008).
Bonaldo, M. C. et al. Isolation of infective zika virus from urine and saliva of patients in Brazil. PLoS Negl. Trop. Dis. 10, e0004816 (2016).
Zhang, R. et al. A CRISPR screen defines a signal peptide processing pathway required by flaviviruses. Nature 535, 164–168 (2016).
Sapparapu, G. et al. Neutralizing human antibodies prevent Zika virus replication and fetal disease in mice. Nature 540, 443–447 (2016).
Zhao, H. et al. Structural basis of zika virus-specific antibody protection. Cell 166, 1016–1027 (2016).
Hasan, S. S. et al. A human antibody against Zika virus crosslinks the E protein to prevent infection. Nat. Commun. 8, 14722 (2017).
Summers, P. L., Cohen, W. H., Ruiz, M. M., Hase, T. & Eckels, K. H. Flaviviruses can mediate fusion from without in Aedes albopictus mosquito cell cultures. Virus Res. 12, 383–392 (1989).
Dowd, K. A., DeMaso, C. R. & Pierson, T. C. Genotypic differences in Dengue virus neutralization are explained by a single amino acid mutation that modulates virus breathing. mBio 6, e01559–01515 (2015).
Stocks, C. E. & Lobigs, M. Signal peptidase cleavage at the flavivirus C-prM junction: dependence on the viral NS2B-3 protease for efficient processing requires determinants in C, the signal peptide, and prM. J. Virol. 72, 2141–2149 (1998).
Lobigs, M. Flavivirus premembrane protein cleavage and spike heterodimer secretion require the function of the viral proteinase NS3. Proc. Natl Acad. Sci. USA 90, 6218–6222 (1993).
Yamshchikov, V. F. & Compans, R. W. Formation of the flavivirus envelope: role of the viral NS2B-NS3 protease. J. Virol. 69, 1995–2003 (1995).
Oliveira, E. R. A., Mohana-Borges, R., de Alencastro, R. B. & Horta, B. A. C. The flavivirus capsid protein: structure, function and perspectives towards drug design. Virus Res. 227, 115–123 (2017).
Weaver, S. C. & Reisen, W. K. Present and future arboviral threats. Antiviral Res. 85, 328–345 (2010).
Szymczak, A. L. & Vignali, D. A. Development of 2A peptide-based strategies in the design of multicistronic vectors. Expert. Opin. Biol. Ther. 5, 627–638 (2005).
Scaturro, P. et al. Characterization of the mode of action of a potent dengue virus capsid inhibitor. J. Virol. 88, 11540–11555 (2014).
Noad, R. & Roy, P. Virus-like particles as immunogens. Trends Microbiol. 11, 438–444 (2003).
Frietze, K. M., Peabody, D. S. & Chackerian, B. Engineering virus-like particles as vaccine platforms. Curr. Opin. Virol. 18, 44–49 (2016).
Garg, H., Sedano, M., Plata, G., Punke, E. B. & Joshi, A. Development of virus-like-particle vaccine and reporter assay for Zika virus. J. Virol. 91, https://doi.org/10.1128/JVI.00834-17 (2017).
Garg, H., Mehmetoglu-Gurbuz, T. & Joshi, A. Virus like particles (VLP) as multivalent vaccine candidate against Chikungunya, Japanese Encephalitis, Yellow Fever and Zika Virus. Sci. Rep. 10, 4017 (2020).
Cox, F. et al. Adenoviral vector type 26 encoding Zika virus (ZIKV) M-Env antigen induces humoral and cellular immune responses and protects mice and nonhuman primates against ZIKV challenge. PLoS ONE 13, e0202820 (2018).
Abbink, P. et al. Protective efficacy of multiple vaccine platforms against Zika virus challenge in rhesus monkeys. Science 353, 1129–1132 (2016).
Larocca, R. A. et al. Vaccine protection against Zika virus from Brazil. Nature 536, 474–478 (2016).
Pardi, N. & Weissman, D. Nucleoside modified mRNA vaccines for infectious diseases. Methods Mol. Biol. 1499, 109–121 (2017).
Manea, M., Mezo, G., Hudecz, F. & Przybylski, M. Mass spectrometric identification of the trypsin cleavage pathway in lysyl-proline containing oligotuftsin peptides. J. Pept. Sci. 13, 227–236 (2007).
Pardi, N., Hogan, M. J., Porter, F. W. & Weissman, D. mRNA vaccines—a new era in vaccinology. Nat. Rev. Drug Discov. 17, 261–279 (2018).
Maciejewski, S. et al. Distinct neutralizing antibody correlates of protection among related Zika virus vaccines identify a role for antibody quality. Sci. Transl. Med. 12, https://doi.org/10.1126/scitranslmed.aaw9066 (2020).
Essink, B. et al. The safety and immunogenicity of two Zika virus mRNA vaccine candidates in healthy flavivirus baseline seropositive and seronegative adults: the results of two randomised, placebo-controlled, dose-ranging, phase 1 clinical trials. Lancet Infect. Dis., https://doi.org/10.1016/S1473-3099(22)00764-2 (2023).
Nelson, J. et al. Impact of mRNA chemistry and manufacturing process on innate immune activation. Sci. Adv. 6, eaaz6893 (2020).
Sabnis, S. et al. A Novel Amino Lipid Series for mRNA delivery: improved endosomal escape and sustained pharmacology and safety in non-human primates. Mol. Ther. 26, 1509–1519 (2018).
Wu, K. et al. Variant SARS-CoV-2 mRNA vaccines confer broad neutralization as primary or booster series in mice. Vaccine 39, 7394–7400 (2021).
Walls, A. C. et al. Distinct sensitivities to SARS-CoV-2 variants in vaccinated humans and mice. Cell Rep. 40, 111299 (2022).
Scheaffer, S. M. et al. Bivalent SARS-CoV-2 mRNA vaccines increase breadth of neutralization and protect against the BA.5 Omicron variant in mice. Nat. Med., https://doi.org/10.1038/s41591-022-02092-8 (2022).
Acknowledgements
Medical writing and editorial assistance were provided by Kate Russin, PhD, and Jessica Nepomuceno, PhD, of MEDiSTRAVA in accordance with Good Publication Practice (GPP3) guidelines, funded by Moderna, Inc., and under the direction of the authors. The authors would like to acknowledge James Crowe, MD (Vanderbilt University Medical Center) and Mike Diamond, MD, PhD (Washington University School of Medicine) for providing the antibodies used in this study; Arshan Nasir (Moderna, Inc.) for bioinformatics analysis of the sequence database; and Mario Roederer (NIAID, NIH) and Martha Nason (NIAID, NIH) for involvement in the NHP studies conducted by the Vaccinees Research Center at BIOQUAL, Inc. This work was supported in part by the Intramural Research Program of the NIAID. This project has been funded in whole or in part with federal funds from the Department of Health and Human Services; Office of the Assistant Secretary for Preparedness and Response; Biomedical Advanced Research and Development Authority, under Contract No. HHSO100201600029C.
Author information
Authors and Affiliations
Contributions
B.B., H.B., S.B., W.-P.K., B.S.G., K.M.M., T.C.P., and A.C. contributed to the study concept and design. Data collection was performed by B.B., N.N., K.E.B., M.A., T.J.R., K.A.D., and T.C.P.; data analysis and interpretation was performed by B.B., N.N., K.B., C.J.H., H.B., B.F., K.E.B., T.J.R., K.A.D., T.C.P., and A.C. All authors contributed to the preparation of the manuscript and approved the final draft.
Corresponding author
Ethics declarations
Competing interests
B.B., N.N., K.B., C.J.H., H.B., and A.C. are employees of Moderna, Inc., and hold stock/stock options in the company. T.C.P., K.A.D., B.S.G., and W.-P.K. are inventors on patent applications describing the VRC5283 DNA vaccine (U.S. patent application number 16/334,099 and PCT/2018/018809). TCP is supported by the Division of Intramural Research, NIAID, NIH. S.B., B.F., K.E.B., M.A., B.E.F., T.J.R., and K.M.M. have no conflicts.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bollman, B., Nunna, N., Bahl, K. et al. An optimized messenger RNA vaccine candidate protects non-human primates from Zika virus infection. npj Vaccines 8, 58 (2023). https://doi.org/10.1038/s41541-023-00656-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41541-023-00656-4
- Springer Nature Limited
We’re sorry, something doesn't seem to be working properly.
Please try refreshing the page. If that doesn't work, please contact support so we can address the problem.