Abstract
Broad-spectrum β-lactam antibiotic resistance in Staphylococcus aureus is a global healthcare burden1,2. In clinical strains, resistance is largely controlled by BlaR13, a receptor that senses β-lactams through the acylation of its sensor domain, inducing transmembrane signalling and activation of the cytoplasmic-facing metalloprotease domain4. The metalloprotease domain has a role in BlaI derepression, inducing blaZ (β-lactamase PC1) and mecA (β-lactam-resistant cell-wall transpeptidase PBP2a) expression3,4,5,6,7. Here, overcoming hurdles in isolation, we show that BlaR1 cleaves BlaI directly, as necessary for inactivation, with no requirement for additional components as suggested previously8. Cryo-electron microscopy structures of BlaR1—the wild type and an autocleavage-deficient F284A mutant, with or without β-lactam—reveal a domain-swapped dimer that we suggest is critical to the stabilization of the signalling loops within. BlaR1 undergoes spontaneous autocleavage in cis between Ser283 and Phe284 and we describe the catalytic mechanism and specificity underlying the self and BlaI cleavage. The structures suggest that allosteric signalling emanates from β-lactam-induced exclusion of the prominent extracellular loop bound competitively in the sensor-domain active site, driving subsequent dynamic motions, including a shift in the sensor towards the membrane and accompanying changes in the zinc metalloprotease domain. We propose that this enhances the expulsion of autocleaved products from the active site, shifting the equilibrium to a state that is permissive of efficient BlaI cleavage. Collectively, this study provides a structure of a two-component signalling receptor that mediates action—in this case, antibiotic resistance—through the direct cleavage of a repressor.
Similar content being viewed by others
Main
Staphylococcus aureus underlies one of the top three most deadly drug-resistant bacterial infections worldwide2. Strains of methicillin-resistant S. aureus (MRSA) are particularly problematic owing to their prevalence in nosocomial and community-acquired infections, increased rates of morbidity and death, as well as elevated healthcare costs1. MRSA is notorious for its ability to acquire resistance to not only β-lactams but also many other classes of antibiotics1,9. The drugs typically used to treat MRSA infections, such as vancomycin, are less effective, more toxic and are subject to established and growing resistance mechanisms as evidenced in vancomycin-resistant S. aureus strains9,10,11. Despite some progress in the development of new therapeutics1, S. aureus has recently been estimated to cause more than a quarter of all mortality attributed to antimicrobial resistance in economically developed countries2, demonstrating that improved treatment options are urgently required. A clearer understanding of resistance in this pathogen could facilitate the development of new cocktail therapies to preserve β-lactam action.
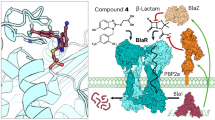
Inducible β-lactam resistance in S. aureus is mediated by the homologous bla or mec divergons. These encode regulatory systems that sense environmental β-lactams through a membrane-embedded sensor–inducer (BlaR1/MecR1; 35% sequence identity; Extended Data Fig. 1) and induce cleavage of a cytoplasmic transcriptional repressor (BlaI/MecI; 61% sequence identity)7. This results in the expression of blaZ and mecA, which encode β-lactamase PC15 (conferring resistance to penicillin antibiotics) and the altered penicillin-binding protein (enzymes involved in cell-wall synthesis) PBP2a (conferring insensitivity to almost all other β-lactams)6,12, respectively (Fig. 1a). The presence of mec, embedded in a mobile genetic element called the Staphylococcal cassette chromosome (SCCmec), and the production of PBP2a are hallmarks of MRSA13. However, there is a complex interplay between divergons. BlaR1–BlaI can control mecA expression3,14,15 (Fig. 1a), notably inducing PBP2a production in response to β-lactam in minutes compared with hours with MecI–MecR1, which can leave the bacterium susceptible to drug treatment16. Accordingly, many SCCmec types lack functional MecR1 and/or MecI17. One of the predominant MRSA strains in the United States is clonal complex 8 USA300 in which mecA is located on a type IV SCCmec cassette with a truncated mecR1 and a deleted mecI17,18 with expression under the control of plasmid-encoded BlaR1 and BlaI. More generally, type IV SCCmec or its closely related subtypes are frequently encountered owing to their presence in epidemic community MRSA strains worldwide. Thus, BlaR1 and BlaI are critically important in determining broad-spectrum antibiotic resistance in many MRSA strains.
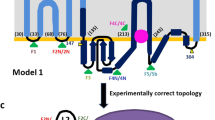
a, The bla pathway controlling β-lactam antibiotic resistance in S. aureus. Previously published models for PC1 (Protein Data Bank (PDB): 3BLM) and PBP2a (PDB: 1MWS) were used to generate the figure. b, Topology of BlaR1. The zinc metalloprotease and sensor domains are highlighted, as well as the dimerization interface. Residue numbers are indicated in parentheses. c, Ribbon representation of the BlaR1 dimer with the monomers shown in either sequence (N to C) rainbow or white. d, Dimeric BlaR1 embedded in a lipid bilayer (Methods). Each protomer of the domain-swapped dimer is shown either as a coulombic electrostatic surface (kcal (mol e−)−1 at 298 K) or a green ribbon. Positively charged residues (Arg, Lys, His) proximal to membrane are shown as sticks. SXXK (sensor) and HEXXH (protease) motif residues are shown in ball and stick representation. e, Ribbon model of the BlaR1 dimer coloured white for chain A and light green for chain B. BlaR1 EL1 is shown in dark grey and dark green for chains A and B, respectively. A superposed benzylpenicillin molecule is shown in orange from an aligned X-ray structure of the BlaR1 sensor domain (PDB: 1XA7). Inset: EL1 as both a ribbon and a transparent solvent-excluded surface, demonstrating that the modelled position of EL1 occupies the β-lactam-binding cleft and would be competitive with the superposed benzylpenicillin antibiotic. f, X-ray structure of the S. aureus BlaR1 sensor-domain active site with covalently bound benzylpenicillin (PDB: 1XA7). Residues of key catalytic motifs are shown as sticks.
BlaR1 and MecR1 are membrane proteins with an N-terminal zinc metalloprotease domain and a C-terminal extracellular β-lactam-binding sensor domain. Isolated structures of the β-lactam-binding sensor domain in complex with antibiotics have been determined for S. aureus BlaR1 and MecR1, showing near-identical architectures (Cα root mean squared deviation of around 2.9 Å) that are related to the class D serine β-lactamases15. No structures of the transmembrane and proteolytic regions have been determined and the signal transduction events linking β-lactam binding and BlaI cleavage on opposing sides of the membrane remain unclear19,20,21. To address these unknowns, we present structures of full-length S. aureus BlaR1. On the basis of conformational differences with and without the β-lactam ampicillin, we propose a mechanism of bla regulation that defines BlaR1 as a two-component signalling receptor. These insights provide a molecular foundation for the development of therapeutics for both MRSA and homologous pathways in other drug-resistant pathogens, including Mycobacterium tuberculosis22 and Clostridioides difficile23.
Cryo-EM analysis of BlaR1
To overcome the historical challenge of obtaining adequate quantities of active BlaR121,24, we used the nisin-controlled gene expression system in Lactococcus lactis25. Membranes containing expressed wild-type BlaR1 (BlaR1(WT)) were able to cleave BlaI, suggesting that the zinc metalloprotease domain is active (Extended Data Fig. 2a), as shown previously in Escherichia coli membranes containing expressed S. aureus BlaR126. Purification of BlaR1 was monitored by the covalently bound fluorescent β-lactam BOCILLIN FL27. As in S. aureus and E. coli28, BlaR1(WT) expressed in L. lactis undergoes spontaneous autocleavage (Extended Data Fig. 2c) between residues Ser283 and Phe284 (confirmed by Edman N-terminal sequencing), which was mitigated by mutation of Phe284 in the P1′ position to alanine (BlaR1(F284A)) (Extended Data Fig. 2e). Detergent-solubilized BlaR1(WT) and BlaR1(F284A) behave like oligomers (Extended Data Fig. 2f) and rate-zonal ultracentrifugation was used to enrich the sample for the intact oligomeric species (Extended Data Fig. 2b–e).
The structures of BlaR1(WT) and BlaR1(F284A) (the latter alone and in complex with ampicillin (Extended Data Fig. 2e)) were determined using cryo-electron microscopy (cryo-EM) to nominal resolutions of 3.8–4.6 Å (Fig. 1, Extended Data Figs. 3–5 and Extended Data Table 1). BlaR1 was consistently observed in a dimeric form (Extended Data Fig. 3b,c) with two-fold symmetry averaging applied in the overall reconstructions for each sample. More structural heterogeneity was detected in the BlaR1(WT) dataset compared with the uncleavable mutant BlaR1(F284A), suggesting enhanced dynamic motions. An asymmetric homodimer class was identified, resulting in a reconstruction of the previously uncharacterized BlaR1(WT) transmembrane and zinc metalloprotease region to 3.8 Å with a local resolution extending to about 3.3 Å (Extended Data Fig. 3e).
BlaR1 forms a domain-swapped dimer
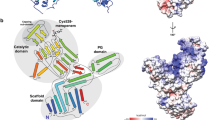
Notably, BlaR1 is observed as an extensive and intimate dimer mediated by an unambiguous domain swap (Fig. 1c,d, Extended Data Fig. 6e and Supplementary Video 1). The N-terminal zinc metalloprotease domain is well-ordered and includes four full transmembrane α-helices, a re-entrant loop (often implicated in meditating protein channel specificity, transport or signalling functions)29 and a cytosolic zinc metalloprotease active site defined by the characteristic H201EXXH and E242XXXD gluzincin signature motifs30 (Figs. 1b and 2 and Extended Data Fig. 1). The active site projects towards the cytosolic face, allowing a point of access for BlaI binding, while the soluble C-terminal β-lactam-sensing domain protrudes from the opposing extramembranal face of the bilayer (Figs. 1 and 2). The N terminus is also oriented to the extracellular side of the membrane leading to an Nout, Cout topology (Fig. 1b,c), which is rarely observed in bacterial membrane proteins31,32.
a, The BlaR1 dimer is shown as a ribbon and is coloured by monomer. The position of the bound catalytic zinc (dashed line) relative to the membrane leaflet (solid line) is indicated. The box out shows the labelled zinc metalloprotease active-site residues, highlighting the intermolecular composition (residues are shown in stick with CPK colouring, with carbons in light green, except for those of the domain-swapped contributing residue Arg294, which are shown in white). Autocleavage loop residues are shown in darker green. b, Alignment of BlaR1 zinc metalloprotease active-site residues (depicted as described in a) with thermolysin (PDB: 2TLX), shown in CPK with gold carbons. The proposed thermolysin catalytic water is represented as a cyan sphere. c,d, Electrostatic surface representation (kcal (mol e−)−1 at 298 K) of the BlaR1(WT) active-site cleft with intact (c) and cleaved (d) autocleavage loop residues, shown in CPK colouring with green and white carbons, respectively. The cryo-EM map is shown as a transparent grey surface. See also Extended Data Fig. 4b,c. e, A 2D representation of the proposed steps leading to the formation of a oxyanion tetrahedral transition state in the zinc metalloprotease active site.
The domain swap involves the cytosolic-facing helix α9 of the zinc metalloprotease domain and helix α10, the terminal transmembrane helix, resulting in an extensive interface with a buried surface area of about 3,165 Å2. Within the zinc metalloprotease domain, additional interface interactions involve helices α8 (same chain) and α9′ (domain swapped), which are oriented almost parallel to the membrane, and helix α7, the C-terminal re-entrant helix that houses the H201EXXH motif. Dimerization defines a central cavity, which in the native membrane is presumably filled with lipids (Fig. 1d). Densities consistent with phosphatidylglycerol headgroups are observed lining the electropositive surface formed at the base of the cavity. Additional diffuse density for fatty acid chains enabled modelling of phosphatidylglycerol(16:0/16:0), a common lipid found in L. lactis and S. aureus33,34,35,36 (Extended Data Fig. 6a–c).
The loop connecting helices α8 and swapped α9, housing the scissile bond between residues Ser283 and Phe284 (residues 271 to 289; here termed the autocleavage loop), enters the zinc metalloprotease active site of its own chain, directly demonstrating that autocleavage occurs in cis (Fig. 2a). The swapped position of α9 effectively lengthens the distance between the autocleavage loop tether points and creates a stabilizing set of hydrophobic core contacts at the dimeric interface involving a number of conserved side chains. In the BlaR1(F284A) mutant structures, the autocleavage loop is intact; however, in the BlaR1(WT) structure, we detected a mixture of uncleaved and cleaved forms, the latter with the central P1′ Phe284 slightly shifted in position (Fig. 2c,d, Extended Data Fig. 4b,c and Supplementary Video 2). This observation is consistent with the presence of both cleaved and uncleaved BlaR1(WT) observed using SDS–PAGE (Extended Data Fig. 2c). We propose that dimerization results in encapsulation of the autocleavage loops within the lumen of the dimer, potentially providing enhanced stability of these dynamic signalling regions to unbridled conformational freedom and/or random hydrolysis.
The extracellular BlaR1 sensor domain was less-well resolved, suggesting a more dynamic relative orientation. The reconstructions of BlaR1(WT) and BlaR1(F284A) without ampicillin provided sufficient secondary structure features (local resolution 6–6.5 Å) to enable confident placement of the sensor-domain X-ray crystal structure37 (Extended Data Figs. 6d,f and 7a–c), which is oriented with the β-lactam-binding grooves facing each other (Fig. 1c–e). In the BlaR1(WT) dataset, the sensor domains were observed to be dynamic and are able to adopt different orientations with respect to each other and the transmembrane domains (Fig. 3a, Extended Data Fig. 7 and Supplementary Video 3). A comparison with BlaR1(F284A) shows that the sensor domains are positioned at different distances to the transmembrane domain in order of BlaR1(WT), BlaR1(F284A) and BlaR1(F284A)–ampicillin (Fig. 3a–c and Extended Data Fig. 7a–c).
a–c, Structures of dimeric BlaR1(WT) (a; green/white ribbon), BlaR1(F284A) (b; orange/white ribbon) and BlaR1(F284A)–ampicillin (c; cyan/white ribbon). The dashed lines highlight the extreme positions of the sensor (extracellular) or zinc metalloprotease (cytosol) domains with respect to the membrane and are coloured according to the BlaR1(WT) or BlaR1(F284A)–ampicillin structures. The labelled arrows in c highlight the β-lactam-induced movement towards the outer and inner leaflets as expanded on in d–f. Unresolved EL1 residues 56–86 are indicated by dashed lines. The double asterisk indicates the location of the determined signal peptidase cleavage site Gly331. d, Benzylpenicillin from PDB 1XA7 (transparent grey) superposed onto b (BlaR1(F284A), orange), showing the structural overlap with the modelled EL1 loop (white). β-Lactam binding as described in c (BlaR1(F284A)–ampicillin, cyan) induces a shift in the sensor domain towards the outer leaflet of the membrane, we propose as a result of competitive displacement of EL1 from the sensor-domain active site. e, β-Lactam-induced motions of transmembrane helix α3, directly downstream of EL1, coloured as described in a–c. The distance between the sensor domain and the membrane correlates with increased displacement of the N-terminal region of α3. f, Superposed active-site structures from BlaR1 structures in a–c. The F284A mutation results in an unoccupied S1′ pocket and associated movement of Tyr261 on helix α8 towards His201 of the HEXXH motif in b (orange) and c (cyan; dashed line). Supporting data are provided in Extended Data Fig. 7. Superpositions in d–f are based on structural alignment of the re-entrant helices containing the catalytic HEXXH…E motif.
The extracellular loop (EL1; residues 56–96), which has long been hypothesized to be involved in BlaR1 signalling19,20, was observed to protrude from transmembrane helices α2 and α3 at the outer leaflet into the region adjacent to the sensor domain of the neighbouring monomer as a result of the domain swap (Fig. 1c–e). Like the sensor domains, the EL1 loops are less-well resolved, consistent with a more dynamic nature. EL1 was modelled using AlphaFold238 as a starting template and optimized using Rosetta density-guided refinement39 (Fig. 1e, Extended Data Fig. 6g–i and Supplementary Video 1). Conserved residues Ile87–Ile94 were specifically observed occupying the β-lactam-binding groove in trans. Notably, acidic Glu89 (consensus Asp) and hydrophobic Ile94 (consensus Val) occupy a similar space to the C3 β-lactam carboxylate and hydrophobic R1 groups in structures of isolated acylated sensor domains (Fig. 1e,f and Extended Data Fig. 6i). The β-lactam carboxylate forms a near universal favourable electrostatic interaction with the adjacent electropositive KTG motif (Lys526 in BlaR1) and the R1 group forms extensive van der Waals interactions with the edge strand in the active site (BlaR1 residues Gly530–Val532). In support of these competitive binding interactions, a peptide with the sequence of EL1 has previously been found to interact with the sensor domain in solution19,20,40. It is clear that acylation of the sensor-domain active site serine by β-lactam antibiotics will displace the EL1 loop from the active site, a structural hallmark of initiation of downstream signalling events now illuminated here.
The protease active site and cleavage mechanism
Helices α6, α7 and β-strand β3 together take on a thermolysin-like fold (the prototypical gluzincin), protruding from a transmembrane scaffold, with analogous positioning of several key active site residues (Fig. 2b). Readily identified by localization of the gluzincin catalytic metalloprotease HEXXH…E sequence motif41, BlaR1 helix α6a here positions His201 and His205, along with Glu242 from neighbouring helix α7, to coordinate the clearly observed catalytic zinc in the expected tetrahedral manner (Fig. 2a and Extended Data Figs. 4b,c and 7e) and present at a 1:1 molar ratio with BlaR1 (Supplemental Table 1). Helices α8 (specifically Ile265 in the S1′ subsite and Tyr261 approaching His201) and α9′ (Arg294′, which forms a stabilizing interchain salt bridge with Asp246) both contribute conserved residues to the active site, suggesting that their precise position is important for function and supporting the observed domain-swapped dimer42 (Fig. 2a,b).
The initial zinc metalloprotease substrate, the autocleavage loop captured here in uncleaved and cleaved states (Fig. 2c,d), provides insights into the molecular details of substrate specificity. The P4–P1′ residues of the autocleavage loop (Leu280, Ser281, His282, Ser283, Phe284) adopt the typical β-strand conformation of protease substrates43, which runs anti-parallel to β3 of the gluzincin domain β-sheet analogous to substrate binding in thermolysin related proteases (Fig. 2). In addition to main-chain β-strand hydrogen bonds between Thr172 and His282, the substrate is anchored by hydrophobic interactions of Leu280 into the S4 pocket at one end and P1′ Phe284 inserting into the S1′ subsite at the other, with the intervening P2 His282 within the S2 pocket the other important specificity determinant. The well-defined S1′ subsite, typically the major determinant for substrate specificity in endolytic metalloproteases44, has a large hydrophobic cavity that is suited for the binding of the aromatic phenylalanine side chain of P1′ Phe284 (Fig. 2c,d). Sequence conservation at this position shows that there is a preference for a bulky hydrophobic amino acid (Extended Data Fig. 8b). By contrast, phenylalanine at the P1′ of the BlaI repressor cleavage sequence is almost invariant (Extended Data Fig. 8c), hinting at a tighter affinity that may be important in displacing autocleaved products from the BlaR1 metalloprotease active site.
In the WT structure, one chain shows clear separation of Ser283(P1) and Phe284(P1′) supporting a post-autocleavage state (Fig. 2c,d and Extended Data Fig. 4b,c). By contrast, the density in the alternate chain is more uninterrupted, suggesting a pre-cleavage state, consistent with a population of both uncleaved and cleaved protein in the sample used for cryo-EM. The autocleavage loop is in the approximate position for cleavage, with the P1 Ser283 carbonyl orientated favourably towards the catalytic zinc, which canonically acts to polarize the scissile bond, promoting its electrophilicity and subsequent nucleophilic attack by an activated water to promote cleavage. Although we observed the same binding orientation in the autocleavage-deficient BlaR1(F284A) structure, the P1′ alanine mutant leads to the loss of the close van der Waals-driven socketing of the aromatic phenylalanine side chain into the S1′ pocket, preventing productive coordination of the carbonyl oxygen to the zinc. Although the hydrolytic water was not observed at this resolution, by comparison with thermolysin, we expect that Glu202 in BlaR1 coordinates the catalytic water and provides general base activation to the nucleophilic form for attack on the P1 carbonyl and subsequent hydrolysis (Fig. 2b,e).
Notably, both the intact and cleaved autocleavage loop forms occupy the catalytic cleft such that their presence is not sterically compatible with the binding and cleavage of BlaI. Thus, our structures show that movement of the autocleavage loop out of the active site, either in an intact or cleaved state, is needed for cleavage of BlaI to proceed. We suggest again a role for the observed dimerization in encompassing and stabilizing these loops in both forms at the dimeric interface, minimizing their susceptibility to random proteolysis that would presumably be enhanced in the more exposed, even less stable monomeric forms.
BlaR1 autocleavage is spontaneous
The role of autocleavage in the regulation of BlaR1 catalytic activity and in turn the BlaR1 divergon has been an important focus4,26,28. Here, BlaR1(WT) undergoes spontaneous autocleavage (Extended Data Fig. 2c), consistent with previous reports4,26,28. Autocleavage is time and temperature dependent with substantial albeit incomplete cleavage at 4 °C, which was used for cryo-EM sample preparation, and near-complete fragmentation overnight at room temperature or 37 °C (Extended Data Fig. 2g).
To further investigate the role of autocleavage, we compared the growth of BlaR1(WT) and BlaR1(F284A) in the common clinical variant USA300 MRSA strain SF8300, in which mecA expression is under control of a blaI-blaR1 plasmid, in the presence of the β-lactam antibiotic nafcillin (Extended Data Fig. 8a). In the absence of antibiotic, all of the strains displayed a similar growth pattern. Nafcillin treatment fully inhibited the growth of the strain containing blaR1(null)-blaI, presumably due to its inability to derepress BlaI. The strains containing an empty vector or a non-functional blaR1(null)-blaI(null) construct did not produce growth inhibition, suggesting constitutive mecA expression in absence of a functional BlaI repressor. The strain containing BlaR1(F284A) (blaR1(F284A)-blaI) displayed intermediate growth attenuation in presence of nafcillin compared with either wild-type BlaR1 (blaR1-blaI) or the functionally defective BlaR1 (null) (blaR1(null)-blaI) (Extended Data Fig. 8a), showing that the F284A mutant is partially functional.
We further examined the ability of purified BlaR1—including both the variants used in our structural studies and that used in the growth assay above (BlaR1(USA300))—to directly cleave purified BlaI. BlaR1(WT) and BlaR1(USA300) cleave BlaI in a manner that is completely abrogated by EDTA but not serine/cysteine protease inhibitors, supporting specific metalloprotease activity (Extended Data Fig. 8d–f). Edman N-terminal sequencing confirmed cleavage between BlaI residues Asn101 (P1) and Phe102 (P1′), as previously identified in BlaR1-containing S. aureus membranes4. Autocleavage-deficient BlaR1(F284A) also shows BlaI cleavage, albeit at a reduced level compared with BlaR1(WT), suggesting that some level of BlaI binding and cleavage is still possible in the presence of an intact autocleavage loop (barring a minor population of autocleaved BlaR1(F284A) that is not detectable in our fluorescence-based gel assay (Extended Data Fig. 2e).
Collectively, these results suggest that the autocleavage event is spontaneous and is not part of the signal transduction mechanism itself, and it does not appear to be an absolute requirement for BlaI cleavage and activation of the bla divergon to occur, although it does permit a more efficient response to environmental β-lactams that may be required for resistance in vivo. These data also show that BlaR1 can recruit and cleave BlaI directly with no need for intermediary components, as was proposed previously8.
β-Lactam-induced allosteric changes
β-Lactam binding to the extracellular BlaR1 sensor domain regulates BlaI cleavage by the zinc metalloprotease domain in the cytoplasm. The three structures presented here reveal conformational changes linking the two BlaR1 active sites about 65 Å apart across the intervening membrane. The overall transmembrane and zinc metalloprotease domains are similar across structures suggesting that no major conformational rearrangements are induced on acylation, although subtle motions of select helices were observed that we believe are functionally important. By contrast, we observed that the sensor domain is more dynamic (Fig. 3a). Focused 3D classification shows that the BlaR1(WT) sensor domain samples multiple orientations, although none refine to a high resolution, which indicates a continuous dynamic trajectory. 3D variability analysis45 shows a general up/down translation of 4–5 Å relative to the membrane (Supplementary Video 2). The BlaR1(F284A) structure (refined with C2 symmetry) exhibited less variability and the sensor domains adopt a position slightly closer to the membrane compared with BlaR1(WT) (Fig. 3a–c and Extended Data Fig. 7a–c). However, in the presence of ampicillin, we see a pronounced rigid body shift of about 6 Å to a position immediately abutting the membrane. The magnitude of this shift is consistent with the expected displacement of EL1 induced by the competitive binding of ampicillin (Fig. 3d). Thus, β-lactam binding and the consequent extrusion of EL1 from the active site appears to trigger movement of the sensor domain towards the outer leaflet of the membrane.
Correlated motions are also observed in the transmembrane and zinc metalloprotease domains. Notably, a general compression parallel to the membrane normal was observed in the BlaR1(F284A)–ampicillin structure. Careful superposition of the reconstructions onto the re-entrant helices containing the catalytic HEXXH…E motif clearly demonstrates that structural elements on either side of the membrane are compressed towards each other in the presence of ampicillin (Fig. 3a–c and Extended Data Fig. 7). As above, a key determinant of the observed motions appears to be the steric displacement of EL1 by the shifted sensor domain. EL1 is situated immediately upstream of helix α3, which is observed kinked into the outer leaflet of the membrane as a result of the shift of EL1 (Fig. 3e and Extended Data Fig. 7d). This is coupled to a general movement of the distal region of the zinc metalloprotease domain towards the inner leaflet of the membrane (Fig. 3a–c and Supplementary Video 3).
Discussion
The structure of S. aureus BlaR1 has been unclear since the initial identification of the gene over 30 years ago3,46. This paper advances knowledge of BlaR1-mediated antibiotic resistance, providing insights into the mechanism of BlaR1 signal transduction that could facilitate the development of therapeutics to treat MRSA infections.
BlaR1 forms a domain-swapped dimer, with the zinc metalloprotease active site accessible from the cytoplasm. The gluzincin fold superposes well with the prototypical soluble representative thermolysin and the most closely related integral membrane zinc metalloproteases with known structures: eukaryotic nuclear membrane ZMPSTE2447 and Ste24p48. In addition to the conserved H201EXXH and E242XXXD motifs, the analysis further defines residues that are critical to catalytic function. Asp210 and Tyr261 are near strictly conserved and are positioned to stabilize core catalytic residues His205 or His201, respectively (Fig. 3f and Extended Data Fig. 9). Mutation of either to alanine abolished β-lactamase induction in B. licheniformis49. Asp246 forms a salt bridge with Arg294′, an interaction that is conserved in other gluzincins; however, in BlaR1, this is located on domain-swapped helix α9′ and forms an intermolecular interaction. In thermolysin, the arginine side chain binds to the substrate P1′ carbonyl42. Consistently, Arg294′ projects towards Phe284 of the autocleavage loop in BlaR1. Mutation of Asp246 abolished β-lactamase induction in B. licheniformis, whereas double mutation of Arg293 and Arg294 severely diminished activity49. We did not observe an equivalent of thermolysin His231, which is proposed to be involved in transition-state binding and stabilization50 (Fig. 2b). His231 is conserved in other gluzincins, including ZMPSTE24, in which it is located in a loop connecting helices equivalent to BlaR1 α8 and α9, which in BlaR1 is the autocleavage loop.
An outstanding question of BlaR1 function is how sensing environmental β-lactams outside the cell regulates BlaI cleavage inside. The structures here describe concerted motions that link the sensor and metalloprotease domains located on opposite sides of the membrane (Fig. 3 and Supplementary Video 3). We hypothesize that the extremes are representative of changes that are involved in the activation and inhibition of the metalloprotease. Although a range of states exists in all of the samples, the equilibrium is shifted—BlaR1(WT) is dominated by a more extended conformation and the BlaR1(F284A)–ampicillin structure is more compact, with the sensor directly in contact with the outer leaflet of the membrane. In the sensor domain, the substrate-mimicking interactions of EL1 within the β-lactam-binding site and the resulting induced domain movement after displacement explain its previously predicted role in signal transduction19,20. On the inner leaflet, concomitant motions are observed, with the metalloprotease domain drawn up to the membrane in the presence of ampicillin. Central to this is helix α3, which traverses the membrane directly linking EL1 and the metalloprotease domain. This helix is bent into the membrane at its N terminus in the BlaR1(F284A)–ampicillin structure, seemingly as a result of the force exerted on EL1 due to the acylation-induced sensor-domain movement (Fig. 3e and Extended Data Fig. 7d). The current resolutions, in particular for the BlaR1(F284A)–ampicillin data, limit analysis of the precise structural changes. However, we note that the overall compression results in a more compact metalloprotease active site with Tyr261 (discussed above) positioned at different distances to His201, moving closer in the ampicillin-bound state of BlaR1(F284A) or in the post-cleavage state of BlaR1(WT), in which the now-free P1′ Phe284 of the cleaved product is not as deeply socketed in the S1′ subsite (Fig. 3f and Extended Data Fig. 7e). In all of the structures, the protease domain is in an autoinhibited state with the autocleavage loop (both cleaved and uncleaved) occupying the active site, suggesting an equilibrium between active site occupied/autoinhibited and active site accessible states (Fig. 4). We hypothesize that the regulation of BlaI cleavage is dependent on the equilibrium between these conformations, with both spontaneous autocleavage and β-lactam acylation serving to modulate this structural equilibrium, pushing it to one favouring displacement of the cleaved autoinhibitory loop fragments from the active site to allow efficient BlaI repressor access and cleavage after the detection of environmental β-lactam (Fig. 4).
a, In the absence of β-lactams, BlaR1 primarily exists in the basal state with the sensor and metalloprotease domains in the most extended conformation away from the membrane outer and inner leaflets, respectively. The EL1 loop is bound within the sensor active site, the SPase cleavage site is exposed and the zinc metalloprotease domain is blocked by the autocleavage loop from efficiently cleaving BlaI. The dotted and dashed lines indicate the edge of the sensor and zinc metalloprotease domains in their extended or contracted (with β-lactam) states, respectively. b, In this basal BlaR1 state, elevated cytosolic BlaI concentrations favour repressor dimerization and binding to the operator sequence, minimizing the expression of genes encoding resistance proteins. In parallel, SPase cleavage of the sensor domain coupled with autocleavage-mediated destabilization of the transmembrane region promotes turnover of BlaR1 in the membrane in the absence of a β-lactam threat. c, Extracellular β-lactams are detected by the BlaR1 sensor through acylation of the SXXK motif, displacing EL1 and promoting movement of the sensor and the metalloprotease domains towards the membrane (Fig. 3), leading to allosteric activation. These events are thought to stabilize the active state of BlaR1 in the presence of antibiotics by blocking of SPase cleavage and promoting a more catalytically competent metalloprotease active site. Autocleavage loop proteolysis also favours BlaI cleavage, probably due to reduced steric hinderance of BlaI binding. However, autocleavage does not appear to be triggered by β-lactam, and we propose that autocleavage is instead needed for BlaR1 maturation and turnover. d, The activated metalloprotease domain cleaves BlaI, decreasing the concentration of BlaI dimer and reducing BlaI binding to the operator. Expression of the divergon genes mecA and blaZ leads to β-lactam protection by PBP2a (β-lactam insensitive; PDB 4WVJ shown) and PC1 (β-lactamase activity; PDB 3BLM shown), respectively.
S. aureus kinase (Stk1)-mediated phosphorylation of BlaR1 is involved in the activation of the bla pathway51. Tyrosine phosphorylation was specifically implicated51, and the only conserved cytoplasmic tyrosine residues are Tyr197, projecting from the C-terminal end of α6a, and the above mentioned Tyr261 on α8 that we observed in a spectrum of active-site-engaged positions (Fig. 3f and Extended Data Fig. 9b). Compellingly, these two side chains stack in an antiparallel manner, with Tyr197 projecting to the exterior and accessible for phosphorylation and Tyr261 towards His201. As such, it is possible that the observed conformational changes may facilitate phosphorylation, inducing added structural changes to destabilize the autocleavage loop products from the active site. Identification of phosphorylated residues and their structural effects could illuminate how phosphorylation contributes to the regulation of resistance and would lend further credence to existing efforts to develop inhibitors targeting Stk151.
Another interesting aspect of BlaR1 is the need for turnover of the basal-level-expressed receptor28. Signal-peptidase-mediated cleavage of Staphylococcus epidermidis BlaR152 and shedding of the sensor domain in S. aureus cultures at equivalent residues (between Gly331 and Gln332) have been documented28. We observed the presence of an analogous ~31 kDa band, which is acylated by BOCILLIN FL (Extended Data Fig. 2c). The S. aureus signal peptidase SpsB cleaves BlaR1(WT), with mutation of BlaR1 Gly331 to lysine abrogating this effect (Extended Data Fig. 8g). This cleavage position sits in the dynamic linker between the C-terminal transmembrane helix α10 and the sensor. In the more extended (minus β-lactam) positioning of the sensor domain away from the membrane (Fig. 3a), this loop would be more accessible for processing by membrane-anchored SpsB. Together with autocleavage, these processing events could promote destabilization of the dimeric interface of BlaR1 on both leaflets and promote inactivation/turnover to prevent potentially deleterious accumulation in the membrane and leaky BlaI cleavage/expression of the resistance genes in the absence of β-lactam. After sensing β-lactam, with reorientation of the sensor domain against the membrane (Fig. 3c), this loop is more protected from processing, maintaining BlaR1 integrity to combat the presence of the bactericidal antibiotic (Fig. 4). These observations provide further support for the ongoing interest in the development of signal peptidase inhibitors, of which the potential for synergistic effects on targets beyond the signal peptide processing roles, such as BlaR1 turnover here, is increasingly compelling.
In summary, the data presented here provide insights into the structure and function of the BlaR1 master controller of β-lactam resistance in S. aureus. In addition to clinically important Staphylococcal pathogens, analogous signalling systems have been identified in C. difficile23 and M. tuberculosis22. C. difficile—a notoriously persistent spore-forming pathogen—has equivalent components to the bla pathway (BlaR1, around 30% sequence identity with S. aureus; Extended Data Fig. 1) controlling the expression of a β-lactamase23. The M. tuberculosis BlaR1 homologue has a similar zinc metalloprotease domain but lacks an equivalent sensor domain making it unclear how β-lactams are detected. One possible functional substitution could involve phosphorylation by PknB, a kinase with a β-lactam-binding PASTA domain that is essential to M. tuberculosis22. We expect that these BlaR1 homologues have a similar domain-swapped architecture and proteolytic derepression of broad-spectrum resistance mechanisms, making the findings here broadly relevant in terms of mechanistic understanding and structure-guided inhibition for the preservation of an important drug class for fighting infections—the β-lactam antibiotics.
Methods
Plasmid construction and site-directed mutagenesis
S. aureus BlaR1 (UniProt: Q00419); residues 1–585) or BlaR1 from MRSA strain USA300 (RefSeq: WP_001096381.1; residues 1–585) were cloned into the Lactococcus lactis expression vector pNZ804853 with a C-terminal hexahistidine tag, using restriction-free cloning54. Cloning and site-directed mutagenesis of pNZ8048 was performed in E. coli MC1061 grown in LB medium supplemented with 10 µg ml−1 chloramphenicol. After verification of the DNA sequence, plasmids were transformed into electrocompetent L. lactis using standard techniques55.
BlaR1 protein expression
BlaR1 constructs were expressed in L. lactis PNZ9000 using the nisin-controlled gene expression system25. Expression strain seed culture was grown in Difco M17 broth (BD)56 supplemented with 10 µg ml−1 chloramphenicol and 10 g l−1 glucose. Large-scale protein expression was performed in a fermenter (Applicon) at 30 °C in M17 BlaR1 Expression Media (M17BEM) and inoculated with seed culture in a 1:100 (v/v) ratio. M17BEM was developed for BlaR1 protein expression based on M17 medium56 (Supplementary Table 2). After sterilization, the medium was supplemented with 10 µg ml−1 chloramphenicol and 100 µM zinc sulfate. The medium was maintained at pH 7 during fermentation with the addition of ammonium hydroxide. Cells were grown to an optical density at 600 nm of around 1.5 before nisin was added to a final concentration of 5 ng ml−1. Protein was expressed for 2–3 h, before collection by centrifugation, then the cells were frozen in liquid nitrogen and stored at −80 °C.
BlaR1 purification
Frozen cell pellets were resuspended in lysis buffer (150 mM sodium chloride; 50 mM HEPES, pH 7.5) with 5 mg ml−1 lysozyme (Bio Basic) and 20 µg ml−1 DNase I (Roche) and incubated at 37 °C for 1 h with stirring. The cells were then cooled in an ice bath before being lysed with a single pass using a French press (Thermo Fisher Scientific) at ~25,000 psi. All of the subsequent steps were carried out at 4 °C. The lysate was centrifuged at 18,600g for 30 min. The supernatant was collected and centrifuged at 40,000 rpm in a Type 45 Ti rotor (Beckman Coulter) for 1 h to pellet the membranes. The membranes were resuspended in lysis buffer using a glass Teflon Dounce homogenizer before being frozen in liquid nitrogen and stored at −80 °C until needed.
Membranes were thawed and resuspended in lysis buffer with 1% (w/v) n-dodecyl-β-d-maltopyranoside (DDM) for 1 h to extract the membranes. All detergents used here were acquired from Anatrace. The extracted membranes were centrifuged at 40,000 rpm in a Type 45 Ti rotor (Beckman Coulter) for 45 min to pellet insolubilized membranes. The supernatant was then filtered through a 0.45 µm membrane and imidazole at pH 7.5 was added to 20 mM before being loaded onto a 1 ml HisTrap HP (GE healthcare) equilibrated in buffer A (150 mM sodium chloride; 20 mM HEPES, pH 7.5; 0.016% (w/v) Anagrade DDM or 0.01% (w/v) lauryl maltose neopentyl glycol (LMNG)) supplemented with 20 mM imidazole at 0.5 ml min−1. The column was then sequentially washed with 15 ml buffer A with 20 mM imidazole, 15 ml buffer A with 100 mM imidazole, and eluted in buffer A with 325 mM imidazole. The protein was then immediately desalted into buffer A with a XK-26 column (Amersham Biosciences) packed with Sephadex G-25 medium resin (GE healthcare). Protein was then concentrated using a stirred-cell concentrator with a 100 kDa molecular mass cut-off cellulose membrane (Merck Millipore) before continued purification or flash-freezing in liquid nitrogen followed by storage at −80 °C. BlaR1 concentration was determined by absorbance at 280 nm using the theoretical extinction coefficient, 106,245 M−1 cm−1, calculated using ExPASy57.
During purification of BlaR1, a band directly below the main full-length band was observed on the SDS–PAGE gel (Extended Data Fig. 2e). Edman N-terminal sequencing revealed cleavage was occurring between BlaR1 residues Ser72 and Lys73, corresponding to an exposed region of EL1 and loss of transmembrane helices 1 and 2. As this minor cleavage product has not to our knowledge been observed in BlaR1 expressed in S. aureus we expect that this cleavage is due to proteases originating in the L. lactis expression system used. We did not see any evidence of this cleavage product in our cryo-EM subclasses or reconstructions.
For cryo-EM, BlaR1 samples in LMNG were further purified using the GraDeR58 rate zonal ultracentrifugation method. Specifically, 5–25% (v/v) glycerol gradients (light buffer: 5% (v/v) glycerol, 150 mM sodium chloride, 20 mM HEPES, pH 7.5 with or without 0.01% LMNG detergent; heavy buffer: 25% (v/v) glycerol, 150 mM sodium chloride, 20 mM HEPES, pH 7.5) were made using a Gradient Master (BioComp Instruments)59. For samples with ampicillin, BlaR1(F284A) was incubated for around 15 min on ice with 1 mM ampicillin and then all subsequent buffers were made with 250 µM ampicillin. After overlaying 150–250 µl of protein sample onto the glycerol gradients, they were immediately centrifuged for the equivalent of 8 h at 55,000 rpm in a SW 55 Ti rotor (Beckman Coulter) at 4 °C. The gradients were then fractionated using a gradient fractionator (BioComp Instruments). The relevant fractions were concentrated using an Amicon Ultra 100 kDa molecular mass cut-off centrifugal filter (Merck Millipore) at 6,000g and 4 °C, before being desalted using a Micro Bio-Spin P-30 Chromatography Column (Bio-Rad Laboratories) into buffer B (150 mM sodium chloride and 20 mM HEPES, pH 7.5). The purity was checked on SDS–PAGE gels. For some gels, BlaR1 was incubated with a final concentration of 50 µM BOCILLIN FL27 for approximately 5 min at room temperature before the addition of loading dye to fluorescently label the sensor domain.
Edman N-terminal sequencing
Samples for Edman N-terminal sequencing were separated on an SDS–PAGE gel before being transferred to a polyvinylidene fluoride membrane and stained using Coomassie. Edman N-terminal sequencing was performed at the SPARK BioCentre.
BlaR1 activity assay with BlaI substrate
Purification of T7-tagged BlaI
The full-length blaI gene (UniProt: P0A042) from S. aureus was cloned into pET21a with an N-terminal T7-tag using restriction-free cloning54. This plasmid was transformed into electrocompetent E. coli Rosetta (DE3) cells. Transformed cells were grown at 37 °C in LB medium supplemented with 100 µg ml−1 ampicillin. At an OD600 of around 0.6–0.8 protein expression was induced by the addition of isopropyl β-d-1-thiogalactopyranoside (IPTG) to 1 mM and the temperature was lowered to 20 °C. Cells were allowed to express protein overnight before collection, flash-freezing in liquid nitrogen and storage at −80 °C.
Protein purification was performed at 4 °C with adaptations from a previous protocol60. Cells were resuspended in buffer 2A (50 mM HEPES, pH 7.0) with 1 cOmplete EDTA-free protease inhibitor cocktail tablet (Roche) and DNase I to ~20 µg ml−1 and lysed in a French press at 16,000–25,000 psi. BlaI was then purified by ammonium sulfate precipitation with sequential addition of ammonium sulfate to 50%, 60%, 70% and 80% saturation. After each addition of ammonium sulfate, the lysate was incubated for 15 min with inversion followed by centrifugation to pellet the precipitated protein. Protein pellets from 70% and 80% saturated ammonium sulfate solutions were resuspended in buffer 2A and dialysed overnight in buffer 2B (50 mM HEPES, pH 7.0; 5% glycerol). The protein was then loaded on a 5 ml Heparin column (GE Healthcare) equilibrated in buffer 2B at 1 ml min−1. The column was washed with buffer 2B until the absorbance at 280 nm stabilized. Protein was eluted from the column with a linear gradient of buffer 2B to buffer 2C (50 mM HEPES, pH 7.0; 5% glycerol; 2 M sodium chloride) over 60 min at a flow rate of 2 ml min−1. Fractions of interest were concentrated on an Amicon Ultra 10 kDa molecular mass cut-off centrifugal filter (Merck Millipore) and loaded onto a Superdex 75 10/300 column (GE healthcare) equilibrated in buffer 2D (20 mM HEPES, pH 7.0; 500 mM sodium chloride; 5% glycerol). Purified protein was concentrated as before to 4 mg ml−1 (as determined by absorbance at 280 nm and correction using the theoretical extinction coefficient 19,940 M−1 cm−1 calculated using the ExPASy server)57 before being frozen in liquid nitrogen and stored at −80 °C until needed.
Assaying BlaR1 activity with purified, T7-tagged BlaI
BlaR1(WT), BlaR1(F284A) or BlaR1(USA300) was incubated with BlaI for 4 h at room temperature in assay buffer (150 mM sodium chloride, 20 mM HEPES, pH 7.5, 0.016% DDM). After the incubation period, the reaction was quenched with SDS loading buffer and the samples were analysed using western blotting. The western blot was probed with an anti-T7-tag antibody HRP conjugate (Novagen) diluted 1:15,000 in TTBS (20 mM Tris, 150 mM NaCl, 0.1 % Tween-20) and 3% non-fat skimmed milk powder) for 1 h at room temperature. The blot was developed using Western Chemiluminescent HRP Substrate (Immobilon) and imaged using the Chemi Genius 2 Bio Imaging System (Syngene).
Determining the BlaI cleavage site
To aid Edman N-terminal sequencing, a BlaI–maltose-binding protein (MBP) fusion construct was cloned to the C terminus of BlaI with a Gly-Ser-Ser linker. Initial attempts to transfer the 3 kDa fragment onto the polyvinylidene fluoride membrane failed, but when fused to MBP, the transfer was trivial. The BlaI-MBP construct was designed with an N-terminal decahistidine tag and was cloned into a pGEX-6P-1 expression plasmid with restriction-free cloning54.
Expression was performed as described for the T7-tagged BlaI construct and cells were lysed as before in buffer 3A (25 mM Tris, pH 8, and 100 mM sodium chloride) with 1 cOmplete EDTA-free protease inhibitor cocktail tablet (Roche) and DNase I (Roche) to ~20 µg ml−1. The cell lysate was centrifuged at 40,000 rpm in a type 45 Ti rotor (Beckman Coulter) for 45 min at 4 °C. The supernatant was filtered through a 0.45 µm filter and imidazole was added to 10 mM before the sample was loaded onto a 1 ml HisTrap HP (GE Healthcare) equilibrated in buffer 3A at 0.75 ml min−1. The column was then washed with 10 cv of 2% buffer 3B (50 mM Tris pH 8, 500 mM imidazole and 100 mM sodium chloride), before being eluted on a gradient of 0–100% over 40 min at flow rate of 1 ml min−1. Fractions containing the protein of interest were concentrated on an Amicon Ultra 30 kDa molecular mass cut-off centrifugal filter (Merck Millipore). The concentrated protein was loaded onto the Superdex 75 10/300 column (GE healthcare) equilibrated in buffer 3C (25 mM Tris, pH 8.0, 100 mM sodium chloride). Purified protein was concentrated as before, frozen in liquid nitrogen and stored at −80 °C until needed.
Inductively coupled plasma mass spectrometry
BlaR1(F284A) was purified in LMNG using immobilized metal affinity chromatography before being concentrated and purified by SEC in buffer B. Protein and buffer samples were prepared in 1% HNO3 with 45Sc (100 ppb) as an internal standard. Inductively coupled plasma mass spectrometry to determine zinc concentration was performed using the NexION 300D (Perkin Elmer) system equipped with a SC-2 DX autosampler, DXi-FAST micro-peristaltic pump, a cyclonic spray chamber, a triple cone interface, a quadrupole ion deflector and Universal Cell Technology. Calibration was performed using IV-Stock-4 calibration standard (Inorganic Ventures). All elements were run in reaction mode (using Dynamic Reaction Cell technology) using ammonia as a reaction gas to remove potential polyatomic interferences. The detection limit for 66Zn was determined to be 0.765 ppb.
Cryo-electron microscopy of BlaR1
For all cryo-EM datasets, C-flat (1.2/1.3) 300 mesh grids (Protochips) were glow-discharged before 3 µl of protein sample at an absorbance at 280 nm of 0.27–0.35 was applied to the grid. The grids were blotted for 1–2 s using the Vitrobot Mark IV (Thermo Fisher Scientific) at 4 °C and 100% humidity before being plunged into liquid ethane. The grids were transferred to liquid nitrogen for storage until imaging.
Cryo-EM grids were screened at the High-resolution Macromolecular Cryo-Electron Microscopy facility at the University of British Columbia on a Glacios (Thermo Fisher Scientific) transmission cryogenic electron microscope with a 200 kV accelerating voltage and a Falcon 3 camera using the EPU (Thermo Fisher Scientific) software package.
The BlaR1(WT) dataset was collected at the High-resolution Macromolecular Cryo-Electron Microscopy facility at the University of British Columbia using the 300 kV Titan Krios (Thermo Fisher Scientific) transmission cryogenic electron microscope equipped with a Falcon 4i (Thermo Fisher Scientific) direct electron detector and a Selectris (Thermo Fisher Scientific) energy filter. Images were gathered in electron-counting mode with a calibrated magnification of ×165,000, corresponding to 0.77 Å per physical pixel. The total dose delivered to the grids was 50 e− Å−2 over 875 frames in the EER video format. Fully automated data collection was performed using EPU (Thermo Fisher Scientific) with a nominal defocus range set from 0.5 to 3.0 μm.
The BlaR1(F284A) ligand-free dataset was collected at the Pacific Northwest Cryo-EM Centre using a 300 kV Titan Krios (Thermo Fisher Scientific) transmission cryogenic electron microscope equipped with the GIF Quantum energy filter (Gatan; operated at 20 keV) and a K3 Summit (Gatan) direct electron detector. Images were gathered in super-resolution mode at a calibrated magnification of 81,000x, corresponding to 1.059 Å per physical pixel (0.5295 Å per super-resolution pixel). A total dose of 33 e− Å−2 was delivered to the grids over 51 frames. Fully automated data collection was performed using SerialEM with a nominal defocus range set from 1.5 to 3.5 μm (ref. 61).
The BlaR1(F284A)–ampicillin dataset was collected at the HHMI Janelia Research Campus using the 300 kV Titan Krios (Thermo Fisher Scientific) transmission cryogenic electron microscope equipped with a spherical aberration corrector, a GIF Quantum energy filter (Gatan) operated at 20 keV and a K3 Summit (Gatan) direct electron detector. Images were gathered in super-resolution mode at a calibrated magnification of ×64,000, corresponding to 1.065 Å per physical pixel (0.5325 Å per super-resolution pixel). The total dose delivered to the grids was 60 e− Å−2 over 60 frames. Fully automated data collection was performed using SerialEM with a nominal defocus range set from 1.0 to 2.0 μm (ref. 61).
Cryo-EM data processing
For the BlaR1(WT) sample, 16,599 videos were collected at a pixel size of 0.77 Å per pixel. Motion correction was performed using MotionCor262 and the contrast transfer functions (CTFs) of the summed and dose-weighted micrographs were determined using CTFFIND463. Approximately 2,000 particles were manually selected to generate reference-free 2D-class averages in RELION64. Representative 2D classes were used as templates for automated particle picking in RELION (v.3.1 and v.4) with 7,508,828 particles picked. 2D classification was performed using CryoSPARC (v.3.2 and v.3.3.2)65, resulting in 4,286,427 particles taken forward for further processing. Three initial volumes were generated and processed for multiple rounds of heterogeneous refinement. The best volume was further refined with NU-refinement with C2 symmetry imposed and used as the initial model for additional rounds of heterogenous refinement to recover good particles that were previously discarded in the 2D classification steps. A particle dataset of 286,692 particles was transferred back to RELION with cs2star in the pyem66 software package and processed for multiple rounds of 3D classification. The particles associated with the best volumes were grouped, refined and used for Bayesian polishing67 and CTF refinement68. Partial signal subtraction was used to remove the micelle density. A reconstruction with C2 symmetry applied was generated from 143,227 particles to a resolution of 4.2 Å as determined using the gold-standard refinement procedure with an FSC of 0.143 and high-resolution noise substitution to correct soft-mask effects69. Symmetry expansion followed by further 3D classification without alignment using a mask covering the sensor domains identified different sensor domain states. A representative class with the most resolved EL1 region contained 23,797 unique particles and refined to 4.9 Å resolution with no symmetry applied (C1). Signal subtraction to remove the sensor domain density was carried out on the 143,227 particles again followed by symmetry expansion and 3D classification without alignment resulting in 66,347 unique particles that refined to 3.8 Å resolution with no symmetry applied (C1) (Extended Data Table 1 and Extended Data Figs. 3 and 4).
For the BlaR1(F284A) sample, a total of 11,844 video series was collected in super-resolution mode at 0.5295 Å per pixel and binned to 1.059 Å per pixel during motion correction performed with MotionCor262. The CTFs of the summed and dose-weighted micrographs were determined using CTFFIND463. Particle picking was carried out as described above with 4,181,855 particles automatically picked. Reference-free 2D classification was performed in CryoSPARC65, resulting in 534,610 good particles taken for further processing. Further particle sorting was performed as described above and 221,007 particles were transferred back to RELION and processed for multiple rounds of 3D classification, and the particles associated with the best volumes were grouped, refined, and used for Bayesian polishing67 and CTF refinement68, and the micelle density was removed through partial signal subtraction. One round of 3D classification without alignment was performed and the particles belonging to the best volumes were grouped and refined. The resulting volume after postprocessing reached a resolution of 4.3 Å with C2 symmetry averaging applied (Extended Data Fig. 3). An example data-processing overview is included in Extended Data Fig. 5.
For the BlaR1(F284A)–ampicillin sample, 10,872 video series were collected in super-resolution mode at 0.5325 Å per pixel and binned to 1.065 Å per pixel during motion correction performed using MotionCor262. The CTFs of the summed and dose-weighted micrographs were determined using CTFFIND463. Particle picking was performed as described above with 5,613,933 particles automatically picked. Reference-free 2D classification was performed in CryoSPARC65, resulting in 216,832 good particles. Further particle sorting was carried out using heterogeneous refinement as described above and 245,700 particles were transferred back to RELION. Multiple rounds of 3D classification were carried out with the particles associated with the best volumes grouped, refined, used for Bayesian polishing67 and CTF refinement68, and the micelle density was removed through partial signal subtraction. One round of 3D classification without alignment was performed and the particles belonging to the best volumes were grouped and refined. The resulting volume after post-processing reached a resolution of 4.6 Å with C2 symmetry averaging applied (Extended Data Fig. 3).
Model building and refinement
A prediction of the transmembrane and catalytic regions of the BlaR1 structure from HHPred70 and Modeller71 was used as a starting point for model building using the BlaR1(WT) transmembrane reconstruction to 3.8 Å resolution. Incorrect regions were deleted and iterative model building and refinement carried out in Coot72, Phenix real.space refine73 in the Phenix software package74, and density-guided Rosetta refinement with symmetry39. For the sensor domain, the X-ray crystal structure of the S. aureus sensor domain (PDB: 1XA7) was automatically fit into the 4.2 Å C2 reconstruction using Phenix dock_in_map. Although this region was of lower local resolution (Extended Data Fig. 3), clear secondary structure features enabled confident placement (Extended Data Fig. 7a–c). Residues 52–96, forming the extracellular facing loop EL1, were not clearly resolved in entirety. AlphaFold2 (not available at the time of initial model building) reproducibly and confidently predicted the C-terminal region of this loop as bound to the sensor-domain β-lactam binding groove, which was supported by low-resolution features in the reconstruction (Extended Data Fig. 6g–i). The EL1 loop conformation for residues 87–97 from AlphaFold2 was incorporated into the final model and subject to extensive density restrained refinement in Rosetta and Phenix real.space refine. Sensor-domain loop 413–427 was omitted due to interchain clashes. For the BlaR1(WT) C1 (4.9 Å) reconstruction with shifted sensor-domain positions, residues 413–427 did not clash and were included. For the BlaR1(F284A) (4.3 Å) and BlaR1(F284A)–ampicillin (4.6 Å) reconstructions, the transmembrane/catalytic regions and sensor domains were docked into the density. The relative distance between sensor and transmembrane domains was different between BlaR1(WT) (furthest), BlaR1(F284A) and BlaR1(F284A)–ampicillin (closest) with the loop (residues 332–345) joining the two adopting different relative positions. For BlaR1(F284A), the final model consists of residues 1–55 and 87–585 and, for BlaR1(F284A)–ampicillin, residues 1–55, 97–412 and 428–585 (EL1 residues 87–96 and sensor-domain residues 413–427 were omitted). Two active site zincs were included in both cases (Extended Data Table 1 and Extended Data Figs. 3, 4 and 6).
Model validation was performed using MolProbity as implemented in Phenix75,76. Figures were created using PyMol (Schrödinger), UCSF Chimera77, UCSF ChimeraX78 and Adobe Illustrator 2021 (Adobe). BlaR1 was inserted into a membrane generated with the CHARM-GUI membrane builder79. Protein interface analysis was carried out with the PISA server using the default parameters80. Sequence conservation was analysed using the ConSurf server81.
BlaR1 functional assays carried out in S. aureus
The S. aureus strains used in the assay (Supplementary Table 3) were cultured in tryptic soy broth (TSB) at 37 °C at 180 rpm. The natural plasmid of SF8300 (a USA300 strain) that contains the blaR1-blaI regulatory genes (pUSA300HOUMR) was used as a template for PCR amplification. Different variants of the blaR1-blaI constructs were generated by splice overlap PCR and cloned into a constitutively expressing vector, pTXΔ82. The plasmids were then transformed to RN4220 through electroporation. After sequence verification, the plasmids were introduced to SF8300ermS (erythromycin sensitive SF8300) through transduction using the phage, Φ11. SF8300ermS was obtained by evicting pUSA300HOUMR through serial passaging of SF8300 in TSB and by selecting for an erythromycin-sensitive colony83. pUSA300HOUMR, in addition to containing blaZ and its control elements (blaR1-blaI), also contains an erythromycin-resistance gene84. Thus, the SF8300ermS lacked β-lactamase and its control elements (blaZ-blaR1-blaI). Lists of the plasmids and primers used in this assay are provided in Supplementary Tables 4 and 5, respectively. Strains containing the pTXΔ plasmid were grown and maintained in 12.5 µg ml−1 tetracycline. blaR1 and blaI are expressed from a plasmid and in turn control the expression of chromosomally encoded mecA. For this purpose, blaR1(F284A)-blaI was cloned into a constitutively expressed plasmid and transduced into SF8300ermS that lacked the natural plasmid (pUSA300HOUMR) containing blaZ and its regulatory elements (blaR1-blaI). SF8300ermS transduced with an identical plasmid with either an empty vector, a fully functional blaR1-blaI or a functionally attenuated blaR1(null)-blaI and blaR1(null)-blaI(null) constructs were used as controls. The null mutants were created by mutating the methionine start codon of blaR1 and blaI to threonine, leading to strains deficient of these proteins. The isogenic strains thus created were assessed for their growth properties either in absence or in presence of nafcillin, an easily available and prototype β-lactam antibiotic that is structurally similar to methicillin.
Growth assays were performed using the Bioscreen C Pro system (Growth Curves USA). The initial bacterial OD600 was adjusted to 0.1 in TSB containing 2 µg ml−1 nafcillin or no nafcillin and 200 µl of this mixture was added to a Honeycomb microplate (Growth Curves USA) in triplicates. Wells containing TSB without any antibiotics were included as controls. The readings were recorded every 30 min for 24 h at 600 nm. The temperature was set to 37 °C. The settling time was set to 30 s and the shake duration was 15 s. DNA sequence analysis was performed using DNAstar software. All experiments except for sequencing were repeated at least twice. The fidelity of all the plasmid constructs and point mutations described in this study were validated through Sanger sequencing.
BlaR1 cleavage with S. aureus signal peptidase SpsB
The gene for S. aureus MRSA252 SpsB (UniProt: Q6GIC3) was cloned into the expression vector pET28a+ (Novagen) using the NdeI and XhoI sites. The expressed protein has an N-terminal hexahistidine tag, followed by a thrombin cleavage site (MGSSHHHHHHSSGLVPRGSHM). Plasmid was transformed into E. coli BL21(DE3). One litre of LB medium supplemented with kanamycin (0.05 mg ml−1) was inoculated with 50 ml of overnight culture. Cells were grown at 37 °C with shaking (250 rpm) until an OD600 of 0.6, at which point 1 mM of IPTG was added to the culture to induce protein expression. Cells were grown for 4 h after induction, then pelleted by centrifugation (6,000g, for 5 min at 4 °C). Then, 5 g of the cell pellet was resuspended in 25 ml of buffer A (20 mM Tris pH 8.0, 100 mM NaCl) and then sonicated using a Sonic Dismembrator Model 500 (Thermo Fisher Scientific), with a 15 s pulse and 30 s rest cycle three times at 30% amplitude, on ice and lysed by passing through an EmulsiFlex-C3 cell disruptor (Avestin) for 5 min. The resulting cell lysate was clarified by centrifugation (35,000g for 30 min at 4 °C). The pellet was resuspended in 25 ml of buffer A with 0.1% Triton X-100 and dounced 50 times followed by centrifugation (35,000g for 30 min at 4 °C). The clarified supernatant was then subjected to Ni2+-NTA affinity chromatography purification. The clarified supernatant, containing the His-tagged SpsB, was poured onto a column containing 3 ml of Ni2+-charged resin (Qiagen) pre-equilibrated with kinetic buffer (buffer A with 0.01% DDM). The column was washed twice with 30 ml of kinetic buffer containing first 10 mM, and then 20 mM imidazole. Next, His-tagged SpsB was eluted using a step gradient, created by 6 ml of kinetic buffer containing imidazole in increasing concentrations of 100 to 500 mM imidazole. Fractions containing purified SpsB were concentrated to 1 ml using an Amicon ultracentrifugal filter (Millipore) with a 3 kDa cut-off and dialysed against 1 l of kinetic buffer at 4 °C overnight. Protein concentration was determined by BCA assay (Thermo Fisher Scientific) according to the manufacturer’s protocol.
To assay for SpsB-mediated BlaR1 cleavage, BlaR1(WT) or BlaR1(G331K) in reaction buffer (150 mM sodium chloride; 20 mM HEPES, pH 7.5, 0.01% (w/v) LMNG) was incubated overnight at room temperature with an excess molar ratio of SpsB. The resulting sample was incubated with BOCILLIN-FL27 for 5 min before SDS–PAGE.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
References
Turner, N. A. et al. Methicillin-resistant Staphylococcus aureus: an overview of basic and clinical research. Nat. Rev. Microbiol. 17, 203–218 (2019).
Murray, C. J. et al. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 399, 629–655 (2022).
Hackbarth, C. J. & Chambers, H. F. blaI and blaR1 regulate β-lactamase and PBP 2a production in methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 37, 1144–1149 (1993).
Zhang, H. Z., Hackbarth, C. J., Chansky, K. M. & Chambers, H. F. A proteolytic transmembrane signaling pathway and resistance to β-lactams in staphylococci. Science 291, 1962–1965 (2001).
Herzberg, O. & Moult, J. Bacterial resistance to β-lactam antibiotics: crystal structure of β-lactamase from Staphylococcus aureus PC1 at 2.5 Å resolution. Science 236, 694–701 (1987).
Lim, D. & Strynadka, N. C. J. Structural basis for the β-lactam resistance of PBP2a from methicillin-resistant Staphylococcus aureus. Nat. Struct. Biol. 9, 870–876 (2002).
Blázquez, B. et al. Regulation of the expression of the β-lactam antibiotic-resistance determinants in methicillin-resistant Staphylococcus aureus (MRSA). Biochemistry 53, 1548–1550 (2014).
Amoroso, A. et al. A peptidoglycan fragment triggers β-lactam resistance in Bacillus licheniformis. PLoS Pathog. 8, e1002571 (2012).
Rodvold, K. A. & Mcconeghy, K. W. Methicillin-resistant Staphylococcus aureus therapy: past, present, and future. Clin. Infect. Dis. 58, S20–S27 (2014).
Wu, Q. et al. Systematic review and meta-analysis of the epidemiology of vancomycin-resistance Staphylococcus aureus isolates. Antimicrob. Resist. Infect. Control 10, 101 (2021).
Rossi, F. et al. Transferable vancomycin resistance in a community-associated MRSA lineage. N. Engl. J. Med. 370, 1524–1531 (2014).
Otero, L. H. et al. How allosteric control of Staphylococcus aureus penicillin binding protein 2a enables methicillin resistance and physiological function. Proc. Natl Acad. Sci. USA 110, 16808–16813 (2013).
Katayama, Y., Ito, T. & Hiramatsu, K. A new class of genetic element, Staphylococcus cassette chromosome mec, encodes methicillin resistance in Staphylococcus aureus. Antimicrob. Agents Chemother. 44, 1549–1555 (2000).
Arêde, P., Ministro, J. & Oliveira, D. C. Redefining the role of the β-lactamase locus in methicillin-resistant Staphylococcus aureus: β-lactamase regulators disrupt the MecI-mediated strong repression on mecA and optimize the phenotypic expression of resistance in strains with constitutive mecA. Antimicrob. Agents Chemother. 57, 3037–3045 (2013).
Alexander, J. A. N. et al. Structural analysis of avibactam-mediated activation of the bla and mec divergons in methicillin-resistant Staphylococcus aureus. J. Biol. Chem. 295, 10870–10884 (2020).
McKinney, T. K., Sharma, V. K., Craig, W. A. & Archer, G. L. Transcription of the gene mediating methicillin resistance in Staphylococcus aureus (mecA) is corepressed but not coinduced by cognate mecA and β-lactamase regulators transcription of the gene mediating methicillin resistance in Staphylococcus aureus. J. Bacteriol. 183, 6862–6868 (2001).
Lakhundi, S. & Zhang, K. Methicillin-resistant Staphylococcus aureus: molecular characterization, evolution, and epidemiology. Clin Microbiol. Rev. 31, e00020-18 (2018).
Liu, C. et al. A population-based study of the incidence and molecular epidemiology of methicillin-resistant Staphylococcus aureus disease in San Francisco, 2004–2005. Clin. Infect. Dis. 46, 1637–1646 (2008).
Hanique, S. et al. Evidence of an intramolecular interaction between the two domains of the BlaR1 penicillin receptor during the signal transduction. J. Biol. Chem. 279, 14264–14272 (2004).
Frederick, T. E., Wilson, B. D., Cha, J., Mobashery, S. & Peng, J. W. Revealing cell-surface intramolecular interactions in the BlaR1 protein of methicillin-resistant Staphylococcus aureus by NMR spectroscopy. Biochemistry 53, 10–12 (2014).
Belluzo, B. S. et al. An experiment-informed signal transduction model for the role of the Staphylococcus aureus MecR1 protein in β-lactam resistance. Sci. Rep. 9, 19558 (2019).
Sala, C. et al. Genome-wide regulon and crystal structure of BlaI (Rv1846c) from Mycobacterium tuberculosis. Mol. Microbiol. 71, 1102–1116 (2009).
Sandhu, B. K., Edwards, A. N., Anderson, S. E., Woods, E. C. & McBride, S. M. Regulation and anaerobic function of the Clostridioides difficile β-lactamase. Antimicrob. Agents Chemother. 64, e01496-19 (2019).
Sobhanifar, S., Prehna, G. & Strynadka, N. in Handbook of Proteolytic Enzymes (eds Barrett, A. et al.) 1237–1243 (Academic, 2012).
Mierau, I. & Kleerebezem, M. 10 years of the nisin-controlled gene expression system (NICE) in Lactococcus lactis. Appl. Microbiol. Biotechnol. 68, 705–717 (2005).
Llarrull, L. I. & Mobashery, S. Dissection of events in the resistance to β-lactam antibiotics mediated by the protein BlaR1 from Staphylococcus aureus. Biochemistry 51, 4642–4649 (2012).
Zhao, G., Meier, T. I., Kahl, S. D., Gee, K. R. & Blaszczak, L. C. BOCILLIN FL, a sensitive and commercially available reagent for detection of penicillin-binding proteins. Antimicrob. Agents Chemother. 43, 1124–1128 (1999).
Llarrull, L. I., Toth, M., Champion, M. M. & Mobashery, S. Activation of BlaR1 protein of methicillin-resistant Staphylococcus aureus, its proteolytic processing, and recovery from induction of resistance. J. Biol. Chem. 286, 38148–38158 (2011).
Viklund, H., Granseth, E. & Elofsson, A. Structural classification and prediction of reentrant regions in α-helical transmembrane proteins: application to complete genomes. J. Mol. Biol. 361, 591–603 (2006).
Jongeneel, C. V., Bouvier, J. & Bairoch, A. A unique signature identifies a family of zinc-dependent metallopeptidases. FEBS Lett. 242, 211–214 (1989).
Granseth, E., Daley, D. O., Rapp, M., Melén, K. & von Heijne, G. Experimentally constrained topology models for 51,208 bacterial inner membrane proteins. J. Mol. Biol. 352, 489–494 (2005).
Daley, D. O. et al. Global topology analysis of the Escherichia coli inner membrane proteome. Science 308, 1321–1323 (2005).
White, D. C. & Frerman, F. E. Extraction, characterization, and cellular localization of the lipids of Staphylococcus aureus. J. Bacteriol. 94, 1854–1867 (1967).
Young, S. A., Desbois, A. P. & Coote, P. J. Characterisation of Staphylococcus aureus lipids by nanoelectrospray ionisation tandem mass spectrometry (nESI-MS/MS). Preprint at bioRxiv https://doi.org/10.1101/593483 (2019).
Schleifer, K. H. et al. Transfer of Streptococcus lactis and related Streptococci to the genus Lactococcus gen. nov. Syst. Appl. Microbiol. 6, 183–195 (1985).
Driessen, A. J. M., Zheng, T., Veld, G. I. T., Op Den Kamp, J. A. F. & Konings, W. N. Lipid requirement of the branched-chain amino acid transport system of Streptococcus cremoris. Biochemistry 27, 865–872 (1988).
Wilke, M. S., Hills, T. L., Zhang, H.-Z., Chambers, H. F. & Strynadka, N. C. J. Crystal structures of the Apo and penicillin-acylated forms of the BlaR1 β-lactam sensor of Staphylococcus aureus. J. Biol. Chem. 279, 47278–47287 (2004).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Dimaio, F. et al. Atomic-accuracy models from 4.5-Å cryo-electron microscopy data with density-guided iterative local refinement. Nat. Methods 12, 361–365 (2015).
Staude, M. W. et al. Investigation of signal transduction routes within the sensor/transducer protein BlaR1 of Staphylococcus aureus. Biochemistry 54, 1600–1610 (2015).
Hooper, N. M. Families of zinc metalloproteases. FEBS Lett. 354, 1–6 (1994).
Marie-Claire, C. et al. Evidence by site-directed mutagenesis that arginine 203 of thermolysin and arginine 717 of neprilysin (neutral endopeptidase) play equivalent critical roles in substrate hydrolysis and inhibitor binding. Biochemistry 36, 13938–13945 (1997).
Tyndall, J. D. A., Nall, T. & Fairlie, D. P. Proteases universally recognize beta strands in their active sites. Chem. Rev. 105, 973–999 (2005).
Gomis-Rüth, F. X., Botelho, T. O. & Bode, W. A standard orientation for metallopeptidases. Biochim. Biophys. Acta 1824, 157–163 (2012).
Punjani, A. & Fleet, D. J. 3D variability analysis: resolving continuous flexibility and discrete heterogeneity from single particle cryo-EM. J. Struct. Biol. 213, 107702 (2021).
Kobayashi, T., Zhu, Y. F., Nicholls, N. J. & Lampen, J. O. A second regulatory gene, blaR1, encoding a potential penicillin-binding protein required for induction of β-lactamase in Bacillus licheniformis. J. Bacteriol. 169, 3873–3878 (1987).
Quigley, A. et al. The structural basis of ZMPSTE24-dependent laminopathies. Science 339, 1604–1607 (2013).
Pryor, E. E. et al. Structure of the integral membrane protein CAAX protease Ste24p. Science 339, 1600–1604 (2013).
Berzigotti, S., Benlafya, K., Sépulchre, J., Amoroso, A. & Joris, B. Bacillus licheniformis BlaR1 L3 loop is a zinc metalloprotease activated by self-proteolysis. PLoS ONE 7, e36400 (2012).
Beaumont, A. et al. The role of histidine 231 in thermolysin-like enzymes: a site-directed mutagenesis study. J. Biol. Chem. 270, 16803–16808 (1995).
Boudreau, M. A., Fishovitz, J., Llarrull, L. I., Xiao, Q. & Mobashery, S. Phosphorylation of BlaR1 in manifestation of antibiotic resistance in methicillin-resistant Staphylococcus aureus and its abrogation by small molecules. ACS Infect. Dis. 1, 454–459 (2015).
Powers, M. E. et al. Type I signal peptidase and protein secretion in Staphylococcus epidermidis. J. Bacteriol. 193, 2677–2686 (2011).
Kuipers, O. P., de Ruyter, P. G. G. A., Kleerebezem, M. & de Vos, W. M. Quorum sensing-controlled gene expression in lactic acid bacteria. J. Biotechnol. 64, 15–21 (1998).
van den Ent, F. & Löwe, J. RF cloning: a restriction-free method for inserting target genes into plasmids. J. Biochem. Biophys. Methods 67, 67–74 (2006).
Geertsma, E. R. & Poolman, B. High-throughput cloning and expression in recalcitrant bacteria. Nat. Methods 4, 705–707 (2007).
Terzaghi, B. & Sandine, W. Improved medium for lactic streptococci and their bacteriophages. Appl. Microbiol. 29, 807–813 (1975).
Gasteiger, E. et al. in The Proteomics Protocols Handbook (ed. Walker, J. M.) 571–607 (Springer, 2005); https://doi.org/10.1385/1-59259-890-0:571
Hauer, F. et al. GraDeR: membrane protein complex preparation for single-particle cryo-EM. Structure 23, 1769–1775 (2015).
Coombs, D. H. & Watts, N. R. M. Generating sucrose gradients in three minutes by tilted tube rotation. Anal. Biochem. 148, 254–259 (1985).
Grossman, M. J. & Lampen, J. O. Purification and DNA binding properties of the blal gene product, repressor for the (3-lactamase gene, blaP, of Bacillus licheniformis. Nucleic Acids Res. 15, 6049–6062 (1987).
Mastronarde, D. N. Automated electron microscope tomography using robust prediction of specimen movements. J. Struct. Biol. 152, 36–51 (2005).
Zheng, S. Q. et al. MotionCor2—anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. W. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. CryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Asarnow, D., Palovcak, E. & Cheng, Y. UCSF pyem v0.5. Zenodo https://doi.org/10.5281/zenodo.3576630 (2019).
Zivanov, J., Nakane, T. & Scheres, S. H. W. A Bayesian approach to beam-induced motion correction in cryo-EM single-particle analysis. IUCrJ 6, 5–17 (2019).
Zivanov, J., Nakane, T. & Scheres, S. H. W. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Chen, S. et al. High-resolution noise substitution to measure overfitting and validate resolution in 3D structure determination by single particle electron cryomicroscopy. Ultramicroscopy 135, 24–35 (2013).
Zimmermann, L. et al. A completely reimplemented MPI bioinformatics toolkit with a new HHpred server at its core. J. Mol. Biol. 430, 2237–2243 (2018).
Webb, B. & Sali, A. Comparative protein structure modeling using MODELLER. Curr. Protoc. Bioinform. 54, 5.6.1–5.6.37 (2016).
Casañal, A., Lohkamp, B. & Emsley, P. Current developments in Coot for macromolecular model building of electron cryo-microscopy and crystallographic data. Protein Sci. 29, 1069–1078 (2020).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D 74, 531–544 (2018).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D 75, 861–877 (2019).
Williams, C. J. et al. MolProbity: more and better reference data for improved all‐atom structure validation. Protein Sci. 27, 293 (2018).
Afonine, P. V. et al. New tools for the analysis and validation of cryo-EM maps and atomic models. Acta Crystallogr. D 74, 814–840 (2018).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Pettersen, E. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Jo, S., Kim, T., Iyer, V. G. & Im, W. CHARMM-GUI: a web-based graphical user interface for CHARMM. J. Comput. Chem. 29, 1859–1865 (2008).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Ashkenazy, H. et al. ConSurf 2016: an improved methodology to estimate and visualize evolutionary conservation in macromolecules. Nucleic Acids Res. 44, W344–W350 (2016).
Chan, L. C. et al. PBP 4 mediates high-level resistance to new-generation cephalosporins in Staphylococcus aureus. Antimicrob. Agents Chemother. 60, 3934–3941 (2016).
Chan, L. C. et al. Ceftobiprole- and Ceftaroline-resistant methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 59, 2960–2963 (2015).
Lowy, F. D. Antimicrobial resistance: the example of Staphylococcus aureus. J. Clin. Invest. 111, 1265–1273 (2003).
Acknowledgements
We thank the Canadian Foundation of Innovation and BC Knowledge Development Fund for infrastructure at the High Resolution Macromolecular EM facility at UBC and the members of the operational team C. Atkinson, F. Rossmann and J. Felt; X. Zhao and Z. Yu for help with earlier cryo-EM data collection at the Howard Hughes Medical Institute CryoEM Shared Resources facility and, more recently, H. Scott for assistance with cryo-EM data collection at the Pacific Northwest Cryo-EM Center (PNCC); and T. Richardson-Sanchez in the laboratory of M. Murphy for running the inductively coupled plasma mass spectrometry samples. A portion of this research was supported by NIH grant U24GM129547 and performed at the PNCC at OHSU and accessed through EMSL (grid.436923.9); a DOE Office of Science User Facility sponsored by the Office of Biological and Environmental Research. Molecular graphics and analyses were performed using UCSF ChimeraX, developed by the Resource for Biocomputing, Visualization and Informatics at the University of California, San Francisco, with support from National Institutes of Health R01-GM129325 and the Office of Cyber Infrastructure and Computational Biology, National Institute of Allergy and Infectious Diseases. This work was funded by the Canadian Institute of Health Research (to N.C.J.S.) and the National Institutes of Health (2R01AI100291 and R21AI142501, to S.S.C.). N.C.J.S. is a Canada Research Chair Tier 1. J.A.N.A. thanks the Vanier Graduate Scholarship Program for salary funding during this work.
Author information
Authors and Affiliations
Contributions
J.A.N.A., M.V., D.C., M.P., N.S. and R.P. performed cloning. J.A.N.A., M.V., F.I.R., J.J., D.C. and M.P. performed protein purification. J.H., L.J.W. and J.A.N.A. performed cryo-EM data processing. L.J.W. carried out model building and refinement. L.J.W., J.A.N.A., N.C.J.S. and M.P. performed structure analyses. J.A.N.A., M.V., W.A.M. and S.S. carried out in vitro activity assays. N.S., R.P. and S.S.C. carried out in vivo activity assays. J.A.N.A., L.J.W. and N.C.J.S. wrote the manuscript with input from H.F.C. and S.S. All of the authors read the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks John Rubinstein and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Amino acid sequence alignment of S. aureus BlaR1 homologues: S. aureus BlaR1 (UniProt ID: Q00419), BlaR1 from MRSA strain USA300 (Refseq ID: WP_001096381.1), S. aureus MecR1 (UniProt ID: P0A0B1), and Clostridioides difficile BlaR (UniProt ID: Q188R3).
S. aureus BlaR1 secondary structure is indicated above the sequence alignment.
Extended Data Fig. 2 Characterization of S. aureus BlaR1 by glycerol gradient centrifugation, SDS-PAGE and size exclusion chromatography.
a, WT BlaR1 in membranes cleaves T7-tagged BlaI as shown by an anti-T7-tag antibody probed Western blot. Representative gel shown from three independent replicates. b, Glycerol gradient chromatogram of BlaR1(WT). c, Coomassie stained (left) and fluorescent (BOCILLIN FL) imaged (right) SDS-PAGE gel of BlaR1(WT) fractions from a glycerol gradient. Samples were incubated with BOCILLIN FL, a fluorescently labelled β-lactam antibiotic, prior to SDS-PAGE. BlaR1(WT) undergoes spontaneous autocleavage. BlaR1(WT) full-length (FL), the BlaR1(WT) C-terminal autocleavage product including residues 284-585 (C-CL), the BlaR1(WT) N-terminal autocleavage product including residues 1-283 (N-CL) and BlaR1(WT) sensor domain including residues 332-585 (SD) bands are indicated. Representative gel shown from multiple (>10) independent purifications. d, Glycerol gradient chromatogram of BlaR1(F284A) with and without ampicillin. e, Coomassie stained (left) and fluorescent (BOCILLIN FL) imaged (right) SDS-PAGE gel of BlaR1(F284A) fractions from the glycerol gradients with and without ampicillin as indicated. Acylation with ampicillin confirmed by inhibition of BOCILLIN binding. Representative gel shown from multiple (>5) independent purifications. f, Size exclusion chromatography using a Superdex 200, 5/150 column shows BlaR1(WT) and BlaR1(F284A) elute as single peaks. The later elution profile of the BlaR1(F284A) sample suggests it may form a more compact dimer. g, Purified BlaR1(WT) undergoes spontaneous cleavage. Samples were loaded on an SDS-PAGE gel following incubation for 1 day following purification at the temperature indicated. Representative gel shown from two independent replicates. h, Schematic of the BlaR1 cleavage products. Molecular weights of the protein ladder components are indicated in kDa for all gels. Gels in panels C and F were scanned for fluorescence using a Chemi Genius 2 Bio Imaging System (Syngene). For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 3 Single particle cryo-EM data processing of BlaR1.
a, Representative micrographs for BlaR1(WT), BlaR1(F284A) and BlaR1(F284A)-ampicillin, respectively. b, Representative 2D classes for BlaR1(WT), BlaR1(F284A) and BlaR1(F284A)-ampicillin, respectively. c, 3D reconstructions BlaR1(WT) C2, BlaR1(WT) C1, BlaR1(WT) TM, BlaR1(F284A) and BlaR1(F284A)-ampicillin, respectively. d, Gold-standard half-map FSC curves as in C (black line) and model-map FSC curves (grey line). Dashed line at 0.143 FSC. e, As c coloured according to local resolution.
Extended Data Fig. 4 Example cryo-EM density displayed for the BlaR1(WT) map.
a, Density for BlaR1(WT) model secondary structure elements shown in mesh. b and c, Stereo views of active site with an intact autocleavage loop (b) or cleaved autocleavage loop (c).
Extended Data Fig 6 Cryo-EM model building for BlaR1.
a, Model of the BlaR1 TM region built using the BlaR(WT) TM map at 3.8 Å resolution. b, Side view as in a. c, Boxed region in a showing two phosphatidylglycerol (16:0/16:0) lipids and three phosphates (presumably corresponding to lipid headgroups) modelled in the central chamber between TM regions. Positively charged residues (arginine, lysine, histidine) are shown as sticks. d, Model of BlaR1 including sensor domain built using the BlaR1(WT) C2 reconstruction at 4.2 Å resolution. Views in e and f indicated. e, View indicated in d showing domain swap mediated dimerization. The domain swap occurs between helices α8 and α9 (*). An AlphaFold2 monomeric model is shown as blue cylinders. f, Sensor domain view as indicated in d showing fit of X-ray structure (1XA7) in density. g, Five AlphaFold2 BlaR1 monomeric models docked in the BlaR1(WT) C2 reconstruction based on the TM region. The different EL1 loop predictions (residues 57-85; low confidence) are indicated with *. h, AlphaFold2 models fit in map based on sensor domain. The C-terminal residues 86-96 of EL1 are consistently predicted bound in the sensor domain β-lactam binding groove with increased confidence (pLDDT of 60-75; >70 indicates a confident prediction). i, Benzylpenicillin bound to sensor domain (1XA7) overlaid with representative AlphaFold2 model.
Extended Data Fig. 7 Conformational changes associated with autocleavage and β-lactam binding.
a–c, Sensor domains modelled into BlaR1(WT) C2 (a), BlaR1(F284A) (b) and BlaR1(F284A)-ampicillin (c) reconstructions. Top front view, bottom side view (indicated with arrow in top panel). Putative signal peptidase cleavage site between Gly330 and Gln331 indicated (**). d, Helix α3 position in BlaR1(WT) (green) and BlaR1(F284A)-ampicillin (cyan) representing the two conformational extremes. For comparison of conformational changes, models were first superposed based on residues 200-250 (HEXXH…E motif including re-entrant helix) and reconstructions superposed on these models. e, Active site density showing approximate distance between Tyr261 and His201/Asp246. Tyr261 moves closer in the post-cleavage state of the BlaR1(WT) TM reconstruction and in BlaR1(F284A) where the phenylalanine to alanine mutation results in an empty S1′ pocket. Cryo-EM map shown as mesh contoured at levels to best display protein (grey) and catalytic zinc position (green).
Extended Data Fig. 8 Growth curves for MRSA USA300 in the absence and presence of 2 µg ml−1 nafcillin and BlaR1 and BlaI cleavage.
a, The F284A mutation in BlaR1 resulted in an attenuated bacterial survival in presence of nafcillin. Growth assay of SF8300ermS containing pTXΔ with variants of blaR1 and blaI in absence of antibiotics showed similar growth rates among the selected strains. Growth assay of SF8300ermS containing pTXΔ with variants of blaR1 and blaI in presence of 2 µg ml−1 nafcillin showed that the F284A functional mutation in blaR1 (red diamonds) caused an obvious growth defect when compared to the wild type (yellow squares). For each condition three technical replicates and three biological replicates were performed. Data from each biological replicate is graphed separately on each row with growth under 0 µg ml−1 nafcillin shown in the left column and growth under 2 µg ml−1 shown in the right column. The mean and standard deviation of the technical replicates (n = 3) are shown for each treatment. b, Sequence conservation plot generated with the WebLogo server (https://weblogo.berkeley.edu/) of residues flanking the BlaR1 autocleavage site between residues Ser283 (P1) and Phe284 (P1′). c, Sequence conservation plot of residues flanking the BlaI cleavage site between residues Asn101 (P1) and Phe102 (P1′). d, BlaR1(WT), e, BlaR1(F284A) and f, BlaR1(USA300) mediate cleavage of BlaI as shown by Western blot. A 2× working concentration of cOmplete protease inhibitor cocktail (Roche) did not prevent BlaI cleavage. However, cleavage was abrogated with the addition of 100 mM EDTA. For panels c, d and e full-length (FLBlaI) and cleaved (CLBlaI) T7-tagged BlaI bands are indicated. BlaI cleavage experiments in d – f were carried out a minimum of three times with similar results, representative blots shown. g, S. aureus signal peptidase SpsB mediates cleavage of the BlaR1(WT) sensor domain as demonstrated by SDS-PAGE stained with Coomassie and BOCILLIN FL fluorescently labelled BlaR1 sensor domain. BlaR1 full-length (FL), the BlaR1 C-terminal autocleavage product including residues 284-585 (C-CL), the BlaR1 N-terminal autocleavage product including residues 1-283 (N-CL) and BlaR1WT sensor domain including residues 332-585 (SDBlaR1) bands are indicated. The BlaR1 G331K (BlaR1(G331K)) mutation in the P1 position of the SpsB cleavage site protects against SpsB mediated cleavage. We predict SpsB mediated cleavage is important for BlaR1 turnover in the absence of β-lactam antibiotic. The molecular weights of selected bands of the protein ladder are indicated in kDa for panels C-F. SpsB cleavage was carried out two times with similar results. For gel source data, see Supplementary Fig. 1.
Extended Data Fig. 9 Mapping of BlaR1 sequence conservation.
Chain A of the BlaR1 dimer is coloured by sequence conservation81 while chain B is coloured grey. Areas where sequence conservation is uncertain are coloured light yellow. a, Overall sequence conservation of the modelled BlaR1 residues. Focus in b, the BlaR1 zinc metalloprotease active site and in c, the BlaR1 domain swap interface with selected conserved residues shown as sticks.
Supplementary information
Supplementary Information
Supplementary Fig. 1 and Supplementary Tables 1–5.
Supplementary Video 1
Overview of BlaR1 dimer structure. One protomer is shown in green ribbon and the other in surface coloured according to electrostatics (Fig. 1d). Residues important to signalling are shown as ball and stick. Positively charged residues close to the membrane are shown as sticks. 00:00–00:14, 360 degree rotation. 00:18–00:34, close up of the active site. The autocleavage loop is coloured in darker green. 00:40–01:00, sensor-binding site showing EL1 bound in the sensor domain β-lactam-binding groove.
Supplementary Video 2
Structural variation in the BlaR1(WT) sensor domain and zinc metalloprotease active site. Morph between different sensor domain states observed for BlaR(WT). The low-pass filtered BlaR1(WT) C1reconstruction is shown in grey (solid and transparent), with the model in ribbon with one protomer coloured green and the other coloured grey. 00:11–00:25, close up morph between the two zinc metalloprotease active site states observed in the 3.8 Å BlaR1(WT) reconstruction representing pre-cleavage, with near continuous density along autocleavage loop, and post-cleavage, with clear interrupted density. The distance between Tyr261 and His231 is indicated by a blue dashed line.
Supplementary Video 3
β-lactam-induced conformational changes in BlaR1. Front view morph between BlaR1(F284A) structures without (orange) and with (cyan) ampicillin bound in the sensor domain. Maps are coloured according to Fig. 3 and are shown as solid and transparent surfaces with the structural model shown in ribbon and the protomers coloured green and grey. 00:11–00:15, side view. 00:18–00:22, slabbed (mid-point) side view. 00:25–00:31, animation of β-lactam binding and induced sensor domain movement.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alexander, J.A.N., Worrall, L.J., Hu, J. et al. Structural basis of broad-spectrum β-lactam resistance in Staphylococcus aureus. Nature 613, 375–382 (2023). https://doi.org/10.1038/s41586-022-05583-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-022-05583-3
- Springer Nature Limited
This article is cited by
-
Restoring susceptibility to β-lactam antibiotics in methicillin-resistant Staphylococcus aureus
Nature Chemical Biology (2024)
-
Surfactin and its Antibacterial Mechanism on Staphylococcus Aureus and Application in Pork Preservation
Food and Bioprocess Technology (2024)
-
Hydrogen bond-bridged phosphorene flexible film for photodynamic inhibiting Staphylococcus aureus
Nano Research (2024)