Abstract
Lung-function impairment underlies chronic obstructive pulmonary disease (COPD) and predicts mortality. In the largest multi-ancestry genome-wide association meta-analysis of lung function to date, comprising 588,452 participants, we identified 1,020 independent association signals implicating 559 genes supported by ≥2 criteria from a systematic variant-to-gene mapping framework. These genes were enriched in 29 pathways. Individual variants showed heterogeneity across ancestries, age and smoking groups, and collectively as a genetic risk score showed strong association with COPD across ancestry groups. We undertook phenome-wide association studies for selected associated variants as well as trait and pathway-specific genetic risk scores to infer possible consequences of intervening in pathways underlying lung function. We highlight new putative causal variants, genes, proteins and pathways, including those targeted by existing drugs. These findings bring us closer to understanding the mechanisms underlying lung function and COPD, and should inform functional genomics experiments and potentially future COPD therapies.
Similar content being viewed by others
Main
Lung-function abnormality predicts mortality and is a diagnostic criterion for chronic obstructive pulmonary disease (COPD)1, which is the most prevalent respiratory disease globally2 and lacks disease-modifying treatments. Although smoking and other environmental risk factors for COPD are well known and genetic susceptibility is recognized, the molecular pathways underlying COPD are incompletely understood. As with other complex traits there has been a lack of ancestral diversity in genome-wide association studies (GWAS)3 of lung function4,5,6. Multi-ancestry studies improve the power and fine-mapping resolution of GWAS and increase the prospects for prediction, prevention, diagnosis and treatment in diverse populations3,4,7.
Understanding of the genes, proteins and pathways involved in disease-related traits underpins modern drug development. A high yield of genetic-association signals, improved signal resolution and integration with functional evidence assist confident identification of causal genes as well as the variants and pathways that impact gene function and regulation. Although datasets and in silico tools to connect GWAS signals to causal genes are improving, the findings from different datasets and tools have lacked consensus8,9, highlighting a need for frameworks to integrate functional evidence types and compare findings10.
Aggregation of lung-function-associated genetic variants into a genetic risk score (GRS) provides a tool for COPD prediction5. When a GRS comprises many variants, partitioning the GRS according to the biological pathways the variants influence could provide a tool to explore their aggregated consequences across different traits through phenome-wide association studies (PheWAS). Just as PheWAS of individual genetic variants predicts the consequences of perturbations of specific protein targets, informing assessment of drug efficacy, drug safety and drug repurposing11, PheWAS of pathway-partitioned GRS could inform the understanding of the consequences of perturbations of specific pathways.
Through the largest global assembly of lung-function genomics studies to date we: (1) undertook a multi-ancestry GWAS meta-analysis of lung-function traits in 588,452 individuals to detect novel signals, improve fine mapping and estimate heterogeneity in allelic effects attributable to ancestry; (2) tested whether lung-function signals are age- or smoking-dependent, and assessed their relationship to height; (3) investigated cell-type and functional specificity of lung-function association signals; (4) fine-mapped signals through annotation-informed credible sets, integrating functional data such as respiratory cell-specific chromatin accessibility signatures; (5) applied a consensus-based framework to systematically investigate and identify putative causal genes, integrating eight locus-based or similarity-based criteria; (6) developed and applied a GRS for the ratio of forced expiratory volume in 1 s (FEV1) to forced vital capacity (FVC) in different ancestries in the UK Biobank and COPD case–control studies; and (7) applied PheWAS to individual variants, GRS for each lung-function trait and GRS partitioned by pathway. Through these approaches, we aimed to detect novel lung-function signals and putative causal genes as well as provide new insights into the mechanistic pathways underlying lung function, some of which may be amenable to drug therapy.
Results
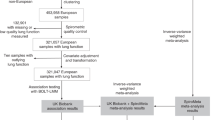
We undertook genome-wide association analyses of FEV1, FVC, FEV1/FVC and peak expiratory flow rate (PEF) from 49 cohorts (Methods and Supplementary Tables 1,2). Our study of up to 588,452 participants comprised individuals of African (AFR; n = 8,590), American/Hispanic (AMR; n = 14,668), East Asian (EAS; n = 85,279), South Asian (SAS; n = 4,270) and European ancestry (EUR; n = 475,645; Supplementary Fig. 1a,b). In cohort-specific analyses we adjusted for age, age squared, sex and height, accounting for population structure and relatedness (Methods and Supplementary Tables 2–4), and then applied genomic control using the linkage disequilibrium (LD) score regression intercept12. After filtering and meta-analysis across multi-ancestry cohorts, 66.8 million variants were available in each of four lung-function traits, with genomic inflation factors λ of 1.025, 1.022, 0.984 and 0.996 for FEV1, FVC, FEV1/FVC and PEF, respectively (Supplementary Figs. 2,3 and Supplementary Table 5).
1,020 signals for lung function
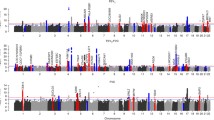
After excluding eight signals associated with smoking behavior (Supplementary Table 26) and combining signals that co-localized across traits, we identified 1,020 distinct signals for lung function using a stringent threshold of P < 5 × 10−9 (ref.13; Fig. 1a). Of these, 713 are novel with respect to the signals and studies described in the Supplementary Note (Supplementary Table 6). These 1,020 signals explain 33.0% of FEV1/FVC heritability (21.3% for FEV1, 17.3% for FVC and 21.4% for PEF; Methods).
To facilitate fine mapping, we included larger, more diverse populations than previous lung-function GWAS. We performed multi-ancestry meta-regression with MR-MEGA7, which incorporates axes of genetic ancestry as covariates to model heterogeneity (Methods). We then incorporated functional annotation for chromatin accessibility and transcription-factor binding sites in respiratory-relevant cells and tissues, and enriched genomic annotations14 to weight prior causal probabilities of association for putative causal variants (Methods). Overall reductions in credible set size and higher maximum posterior probabilities of association for the most likely causal variants were evident after multi-ancestry meta-regression and after functional annotations were incorporated (Supplementary Fig. 4). Following fine mapping, 438 (43%) signals had a single putative causal variant (posterior probability > 50%) and the median credible set size was nine variants (Supplementary Note).
Of the 960 sentinels represented in ≥7 cohorts, 109 signals showed heterogeneity attributable to ancestry (PHet < 0.05; Supplementary Fig. 5 and Supplementary Table 7), which was more than expected (binominal test, P = 3.93 × 10−15). Among these, five signals (rs9393688, rs28574670 (LTBP4), rs7183859 (THSD4), rs59985551 (EFEMP1) and rs78101726 (MECOM)) showed significant ancestry-correlated heterogeneity (Bonferroni correction for 960 signals tested, PHet < 5.21 × 10−5; Supplementary Fig. 6a–e). The intronic variant rs7183859 in THSD4, which we previously implicated in lung function15, showed larger effect-size estimates in non-EUR ancestries and in particular AFR ancestries (PHet = 3.33 × 10−5; Supplementary Fig. 6c).
We examined associations of lung-function-associated SNPs in children’s cohorts (Supplementary Table 8) and tested for differences in the estimated effect sizes of lung-function-associated SNPs between children and adults as well as between ever-smokers and never-smokers in EUR individuals (Methods). The effect-size estimates between children and adults were correlated (r from 0.51 for FEV1/FVC to 0.79 for FEV1; Supplementary Fig. 7), although 113 signals showed nominal evidence (P < 0.05) of age-dependent effects (more than expected, binomial P = 2.56 × 10−13). Three signals (rs7977418 (CCDC91), rs34712979 (NPNT) and rs931794 (HYKK) showed age-dependent effects (Bonferroni-corrected P < 4.64 × 10−5; Supplementary Table 9). We observed nominal evidence (P < 0.05) of smoking-dependent effects for 69 of 1,020 signals (Supplementary Fig. 8), more than expected (binomial P = 0.0079). The intronic SNP rs7733410 in HTR4, a signal we previously reported for lung function15, showed a 76.2% larger effect on FEV1 in ever-smokers compared with never-smokers (P = 4.09 × 10−5; Supplementary Table 10). As height is a determinant of lung growth, we compared height and lung-function associations, and tested the impact of additional height adjustments for sentinel SNPs. We found no correlation between estimated effect sizes for height and lung function (Supplementary Fig. 9), and the addition of height squared and height cubed covariates had little impact on effect-size estimates (Supplementary Fig. 10).
Cell-type and functional specificity
We assessed whether our association signals were enriched for regulatory or functional features in specific cell types. Using stratified LD-score regression16, we found enrichment of all histone marks tested (H3K27ac, H3K9ac, H3K4me3 and H3K4me1) in lung- and smooth-muscle-containing cell lines (Supplementary Table 11). Using GARFIELD17 we assessed for enrichment of our signals for DNase l hypersensitivity sites and chromatin accessibility peaks, showing enrichment in a wide variety of cell types, including higher enrichment in both fetal and adult lung and blood for FEV1, FEV1/FVC and PEF as well as fibroblast enrichment for FVC (Supplementary Fig. 11a). Our signals were enriched for transcription-factor footprints in fetal lung for FEV1, FEV1/FVC and PEF, for footprints in skin for FVC and also in blood for PEF (Supplementary Fig. 11b). Genic annotation enrichment patterns were similar across all traits, with enrichment mainly in exonic, 3′ UTR and 5′ UTR regions (Supplementary Fig. 11c). For all traits, we saw enrichment for transcription start sites, weak enhancers, enhancers and promoter flanks, with cell types for weak enhancer enrichment including endothelial cells for FEV1, FEV1/FVC and PEF (Supplementary Fig. 11d). For transcription-factor binding sites, we observed a similar enrichment pattern across all of the lung-function traits, with the largest fold enrichment observed for endothelial cells (Supplementary Fig. 11e). Our signals were enriched for assay for transposase-accessible chromatin using sequencing (ATAC–seq) peaks (Supplementary Note) in matrix fibroblast 1 for FVC, matrix fibroblast 2 for FEV1, myofibroblast for FEV1, FEV1/FVC and PEF, and alveolar type 1 cells in FEV1/FVC; furthermore, genic annotations showed enrichment of exon variants for FEV1, FEV1/FVC and 3′ UTR variants for FEV1 and FVC. We also found enrichment of transcription-factor binding sites in lung across all phenotypes and in bronchus for FEV1/FVC (Supplementary Table 12).
Identification of putative causal genes and variants
To identify putative causal genes, we systematically integrated orthogonal evidence using eight locus- or similarity-based criteria (Supplementary Note): (1) the nearest gene to the sentinel SNP, (2) co-localization of the GWAS signal and expression quantitative trait loci (eQTL) or (3) protein quantitative trait loci (pQTL) signals in relevant tissues (Methods), (4) rare variant association in whole-exome sequencing in the UK Biobank, (5) proximity to a gene for a Mendelian disease with a respiratory phenotype (±500 kb), (6) proximity to a human ortholog of a mouse-knockout gene with a respiratory phenotype (±500 kb), (7) an annotation-informed credible set14 containing a missense/deleterious/damaging variant with a posterior probability of association >50% and (8) the gene with the highest polygenic priority score (PoPS)9. We identified 559 putative causal genes satisfying at least two criteria, of which 135 were supported by at least three criteria (Figs. 1b, 2 and Supplementary Fig. 12). Among the 20 genes supported by ≥4 criteria (Supplementary Table 13), six previously implicated genes (TGFB2, NPNT, LTBP4, TNS1, SMAD3 and AP3B1)5,15,18,19,20 were supported by additional criteria compared with the original reports. Fourteen of the 20 genes supported by ≥4 criteria have not been previously confidently implicated in lung function (CYTL1, HMCN1, GATA5, ADAMTS10, IGHMBP2, SCMH1, GLI3, ABCA3, TIM1, CFH, FGFR1, LRBA, CLDN18 and IGF2BP2). These are involved in smooth-muscle function (FGFR1, GATA5 and STIM1), tissue organization (ADAMTS10), alveolar and epithelial function (ABCA3 and CLDN18), and inflammation and immune response to infection (CFH, CYTL1, HMCN1, LRBA and STIM1).
The number of variant-to-gene criteria implicating the gene is in brackets after the gene name. The gray in the first eight columns indicates that at least one variant implicates the gene as causal via the evidence for that column. The last four columns indicate the level of association of the most significant variant implicating the gene as causal with respect to the FEV1/FVC decreasing allele; red indicates that this association is in the same direction of effect as the FEV1/FVC decreasing allele and blue indicates the opposite direction with the shade indicating P < the corresponding value in the legend.
To supplement our understanding of the biological pathways and clinical phenotypes influenced by lung-function-associated variants, we undertook PheWAS of selected individual variants. We selected 27 putative causal genes implicated by ≥4 criteria (20 genes) or by a single putative causal missense variant that was deleterious (five genes: ACAN, ADGRG6, SCARF2, CACNA1S and HIST1H2BE) or rare (two genes: SOS2 and ADRB2; Supplementary Table 14). We interpreted the PheWAS findings (shown in full in Supplementary Fig. 13 and Supplementary Table 15) alongside literature reviews (Supplementary Table 16) and highlight examples below.
The putative causal deleterious missense ABCA3 rs149989682 (A allele; frequency of 0.6%) variant associated with reduced FEV1/FVC was reported to cause pediatric interstitial lung disease21. ABCA3, which is expressed in alveolar type II cells and localized to lamellar bodies, is involved in surfactant-phospholipid metabolism, and ABCA3 mutations cause severe neonatal surfactant deficiency22. The putative causal deleterious missense GATA5 rs200383755 (C allele, frequency of 0.6%) variant associated with lower FEV1 was associated with increased asthma risk, higher blood pressure and reduced risk of benign prostatic hyperplasia (Supplementary Fig. 13i). GATA5 associations have not been previously noted in asthma GWAS, although Gata5-deficient mice show airway hyperresponsiveness23. GATA5 encodes a transcription factor expressed in bronchial smooth muscle, bladder and prostate; a previous benign prostatic hyperplasia GWAS reported a GATA5 signal23,24. CLDN18 was implicated by four criteria, including a mouse knockout with abnormal pulmonary alveolar epithelium morphology25. Through calcium-independent cell adhesion, CLDN18 influences epithelial-barrier function through tight-junction-specific obliteration of the intercellular space26. Its splice variant, CLDN18.1, is predominantly expressed in the lung27. Reduced CLDN18 expression was reported in asthma26. However, our PheWAS showed no association with asthma susceptibility or other traits (CLDN18 rs182770 in Supplementary Table 15). LRBA was also implicated by four criteria. Mutations resulting in LRBA deficiency cause common variable immunodeficiency-8 with autoimmunity, which can include coughing, respiratory infections, bronchiectasis and interstitial lung disease28,29. The putative causal LRBA tolerated missense variant rs2290846 (posterior probability of 56.3%) was associated with 31 traits (false discovery rate (FDR) < 1%; Supplementary Fig. 13n and Supplementary Table 15); the G allele, associated with lower FVC and lower FEV1, was associated with lower neutrophils as well as lower risk of cholelithiasis, cholecystitis30 and diverticular disease.
FGFR1, encoding Fibroblast growth factor receptor 1, has roles in lung development and regeneration31. Loss-of-function FGFR1 mutations cause hypogonadotropic hypogonadism32. The T allele of rs881299, associated with lower FEV1/FVC and higher FVC, was strongly associated with higher testosterone (particularly in males) and higher sex-hormone-binding globulin (SHBG), lower body-mass index (BMI) as well as lower levels of alanine transaminase and urate (Supplementary Fig. 13w–y and Supplementary Table 15). The missense SOS2 variant rs72681869 also showed association with SHBG; in both sexes, the G allele, associated with lower FVC and lower FEV1, was associated with lower SHBG, higher alanine aminotransferase (ALT) and aspartate aminotransferase (AST), higher fat mass, HbA1c and higher systolic and diastolic blood pressure, higher urate and creatinine, and in males lower testosterone and reduced inguinal hernia risk (Supplementary Fig. 13z–bb). Mutations in SOS2 have been reported in individuals with Noonan syndrome. The A allele of rs7514261 implicating CFH, associated with lower FVC, was strongly associated with reduced risk of macular degeneration33 as well as raised albumin (Supplementary Fig. 13g).
CACNA1S is one of several putative causal genes encoding calcium voltage-gated channel subunits in skeletal muscle (CACNA1S, CACNA1D and CACNA2D3 supported by ≥2 criteria; CACNA1C was supported by PoPS). CACNA1S mutations have been reported in hypokalemic periodic paralysis34 and malignant hyperthermia35. CACNA1S is strongly expressed in skeletal muscle but at much lower levels in airway smooth muscle. The common CACNA1S missense variant rs3850625 (A allele, frequency of 11.8% in EUR and 21.4% in SAS) was associated with lower FVC, lower FEV1, lower whole body fat-free mass, reduced hand grip strength as well as lower AST and creatinine levels (Supplementary Fig. 13f). CACNA1S and CACNA1D are targeted by dihydropyridine calcium channel blockers, which previously produced small improvements in lung function in asthma36. For the low-frequency missense ADRB2 variant rs1800888 (T; 1.49% in EUR), associated with lower FEV1 and lower FEV1/FVC, the strongest PheWAS association was with increased eosinophil count (Supplementary Fig. 13d).
Druggable targets
Using the Drug Gene Interaction Database, we surveyed 559 genes supported by ≥2 criteria. CheMBL interactions identified 292 drugs mapping to 55 genes (Supplementary Table 17), including ITGA2, which encodes integrin subunit alpha 2. The reduced expression of ITGA2 in lung tissue associated with the C allele of rs12522114 mimics vatelizumab-induced ITGA2 inhibition; this allele is associated with higher FEV1 and FEV1/FVC, indicating the potential to repurpose vatelizumab, which increases T regulatory cell populations37, for COPD treatment.
Pathway analysis
Using ConsensusPathDB38, we tested biological pathway enrichment for 559 causal genes supported by ≥2 criteria, highlighting pathways relevant for development, tissue integrity and remodeling (Supplementary Table 18). These include pathways not previously implicated in pathway enrichment analyses for lung function—such as PI3K–Akt signaling, integrin pathways, endochondral ossification, calcium signaling, hypertrophic cardiomyopathy and dilated cardiomyopathy—as well as those previously implicated via individual genes5 such as TNF signaling, actin cytoskeleton, AGE–RAGE signaling, Hedgehog signaling and cancers. We found strengthened enrichment through newly identified genes in previously described pathways, such as extracellular matrix organization (34 new genes), elastic fiber formation (eight new genes) and TGF–Core (four new genes). Consistent with our ConsensusPathDB findings, Ingenuity Pathway Analysis (https://digitalinsights.qiagen.com/IPA)39 highlighted enrichment of cardiac hypertrophy signaling and osteoarthritis pathways and also implicated pulmonary and hepatic fibrosis signaling pathways, axonal guidance and PTEN signaling as well as the upstream regulators TGFB1 and IGF-1 (Supplementary Table 19).
Multi-ancestry GRS for FEV1/FVC and COPD
We built multi-ancestry and ancestry-specific GRSs weighted by FEV1/FVC effect sizes and tested association with FEV1/FVC and COPD (GOLD stage 2–4) within groups of individuals of different ancestries in the UK Biobank (Methods). Our new GRS improved lung-function and COPD prediction compared with a previous GRS based only on individuals of EUR ancestry5 (Fig. 3a,b and Supplementary Table 20), and the multi-ancestry GRS outperformed the ancestry-specific GRS in all UK Biobank ancestries. We then tested the multi-ancestry GRS in five independent COPD case–control studies (Supplementary Table 21 and Methods). Stronger COPD susceptibility associations were observed across five EUR-ancestry studies compared with a previous GRS5 (Fig. 3c and Supplementary Table 22). In the meta-analysis of these EUR studies, the odds ratio for COPD per s.d. of GRS increase was 1.63 (95% confidence interval (CI), 1.56–1.71; P = 7.1 × 10−93); members of the highest GRS decile had a 5.16-fold higher COPD risk than the lowest decile (95% CI, 4.14–6.42; P = 1.0 × 10−48; Fig. 3d and Supplementary Table 23). The results for individuals in the SPIROMICS study of AFR ancestry were comparable to individuals from the UK Biobank with AFR ancestry but lower in magnitude compared with the COPDGene AFR population (Fig. 3c).
a, Prediction performance of three GRSs across ancestry groups for FEV1/FVC shown as the s.d. change in FEV1/FVC per s.d. increase in GRS for individuals in the UK Biobank grouped according to ancestry. Sample sizes: AFR, n = 4,227; AMR, n = 2,798; EAS, n = 1,564; and EUR, n = 320,656. b, Prediction performance of three GRSs for COPD shown as COPD odds ratio per s.d. increase in GRS. Sample sizes: AFR, 250 cases and 3,977 controls; AMR, n = 151 cases and 2,647 controls; EUR, 24,062 cases and 296,594 controls. UKB, UK Biobank. c, Odds ratio for COPD per s.d. change in GRS in COPD case–control studies. P values were calculated from a logistic regression adjusted for age, age squared, sex, height and principal components, followed by fixed-effect meta-analysis. d, Decile analysis meta-analyzed across five EUR studies shown as the COPD odds ratio compared between members of each decile and the reference decile. n = 11,074 (4,328 cases and 6,746 controls). Statistical tests were two-sided, the height of the bars show the point estimate of the effect and whiskers show the 95% CI. OR, odds ratio.
PheWAS of trait-specific GRSs
To study the aggregate effects of lung-function-associated genetic variants on a wide range of diseases and disease-relevant traits, we created GRSs for FEV1, FVC, FEV1/FVC and PEF, each comprising sentinel variants (P < 5 × 10−9) with weights estimated from the multi-ancestry meta-regression (Methods), and tested these in PheWAS. These GRS values showed distinct patterns of associations with respiratory and non-respiratory phenotypes (Supplementary Fig. 14 and Supplementary Table 24). A GRS for lower FEV1 was most strongly associated with increased risk of asthma and COPD, family history of chronic bronchitis/emphysema, lower hand grip strength, increased fat mass, increased HbA1c and type 2 diabetes risk, and elevated C-reactive protein. In addition, associations were observed with increased asthma exacerbations and lower age of onset for COPD (Supplementary Fig. 14a). The GRS for lower FEV1/FVC was associated with key respiratory phenotypes: increased risk of COPD and asthma, family history of chronic bronchitis/emphysema, increased emphysema risk, increased risk of respiratory insufficiency or respiratory failure and younger age of onset for COPD but a slightly lower risk of COPD exacerbations (Supplementary Fig. 14b). In contrast, the GRS for lower FVC was strongly associated with many traits—among the strongest associations were high C-reactive protein, increased fat mass, raised HbA1c and type 2 diabetes, raised systolic blood pressure, lower hand grip strength and raised ALT as well as increased risk of clinical codes for asthma and COPD (Supplementary Fig. 14c). Although the GRS for lower FEV1/FVC was associated with increased standing and sitting height, the GRSs for lower FEV1 and FVC were associated with increased standing height but reduced sitting height. Broadly similar phenome-wide associations were seen for the PEF and the FEV1 GRS (Supplementary Fig. 14d).
PheWAS of GRSs partitioned by pathway
Finally, we hypothesized that partitioning our lung-function GRS into pathway-specific GRSs according to the biological pathways the variants influence could inform understanding of mechanisms underlying impaired lung function, and the probable consequences of perturbing specific pathways. Informed by the above prioritization of putative causal genes and classification of these genes by pathway (‘Pathway analysis’ section), we undertook PheWAS for FEV1/FVC-weighted GRSs partitioned by each of the 29 pathways enriched (FDR < 10−5) for the 559 genes implicated by ≥2 criteria (Methods). Partitioning of GRSs in this way highlighted markedly different patterns of phenome-wide associations (Supplementary Fig. 15 and Supplementary Table 25). Figures 4–7 highlight four pathway-specific GRS examples; all demonstrated association with COPD clinical codes and family history of chronic bronchitis/emphysema, although the associations with other traits varied. The GRS for lower FEV1/FVC specific to elastic fiber formation was associated with increased risk of inguinal, abdominal, diaphragmatic and femoral hernia; diverticulosis; arthropathies; hallux valgus as well as genital prolapse; reduced carpal tunnel syndrome risk and BMI; and increased asthma risk (Fig. 4). In contrast, the GRS for lower FEV1/FVC specific to PI3K–Akt signaling was associated with increased asthma risk, lower IGF-1, lower liver enzymes (ALT, AST and gamma glutamyltransferase (GGT)), lower lymphocyte counts, raised eosinophils, lower fat mass and BMI, and reduced diabetes risk (Fig. 5). The GRS for lower FEV1/FVC specific to the hypertrophic cardiomyopathy pathway was associated with reduced liver enzymes (ALT and GGT) as well as lower apolipoprotein B, LDL, IGF-1 and mean platelet volume (Fig. 6). The GRS associations for lower FEV1/FVC partitioned to signal transduction were specific to respiratory traits, including asthma and emphysema (Fig. 7). Variable height associations were evident: the GRS for lower FEV1/FVC showed association with increased height when partitioned to elastic fiber formation or hypertrophic cardiomyopathy (Figs. 4 and 6), reduced height when partitioned to ESC pluripotency (Supplementary Fig. 15g) and no height association when partitioned to PI3K–Akt signaling or signal transduction (Figs. 5 and 7).
Reactome pathway database. CP, composite phenotype and DFP, Data-Field ID phenotype (Methods). The peach-colored line means FDR 1%.
Kyoto Encyclopedia of Genes and Genomes. CP, composite phenotype; DFP, Data-Field ID phenotype (Methods). The peach-colored line means FDR 1%.
Kyoto Encyclopedia of Genes and Genomes. CP, composite phenotype; DFP, Data-Field ID phenotype (Methods). The peach-colored line means FDR 1%.
Reactome pathway database. CP, composite phenotype (Methods). The peach-colored line means FDR 1%.
We hypothesized that individuals may have high GRS for ≥1 pathways and low GRS for other pathways. Comparisons of the GRSs of individuals across pairs of pathways for each of the 29 pathways (Supplementary Fig. 16a) and in detail for the elastic fiber, PI3K–Akt signaling, hypertrophic cardiomyopathy and signal transduction pathways (Supplementary Fig. 16b) demonstrated how GRS profiles may be concordant or discordant across pathways, which could have implications for the choice of therapy.
Discussion
We present a large ancestrally diverse lung-function GWAS and a comprehensive initiative to relate lung-function- and COPD-associated variants to functional annotations, cell types, genes and pathways. It is the first to investigate possible consequences of intervening in relevant pathways through PheWAS studies, utilizing pathway-partitioned GRS.
The 1,020 signals identified were enriched in functionally active regions in alveolar type 1 cells, fibroblasts, myofibroblasts, bronchial epithelial cells, and adult and fetal lung. We showed effect heterogeneity attributable to ancestry for 109 signals (including LTBP4, THSD4, EFEMP1 and MECOM), between ever-smokers and never-smokers (HTR4), and differences in effects between adults and children (including CCDC91 and NPNT). We mapped lung-function signals to 559 putatively causal genes meeting ≥2 independent criteria. Exemplar genes supported by ≥4 criteria or by deleterious or rare putative causal missense variants implicated surfactant-phospholipid metabolism, smooth-muscle function, epithelial morphology and barrier function, innate immunity, calcium signaling, adrenoceptor signaling, and lung development and regeneration. Among the pathways enriched for putative causal genes were PI3K–Akt signaling, integrin pathways, endochondral ossification, calcium signaling, hypertrophic cardiomyopathy and dilated cardiomyopathy. These pathways have not been previously implicated in lung function using GWAS.
Combined as a GRS weighted by FEV1/FVC effect size, the 1,020 variants strongly predicted COPD in the UK Biobank and in COPD case–control studies, with a more than fivefold change in risk between the highest and lowest GRS deciles. This GRS more strongly predicted FEV1/FVC and COPD across all ancestries than a previous GRS5. Partitioning the FEV1/FVC GRS by the pathways defined by specific variants, informed by detailed, systematic variant-to-gene mapping and pathway analyses, and using our new Deep-PheWAS platform40, illustrated unique patterns of phenotype associations for each pathway GRS. These patterns of PheWAS findings are relevant to the potential efficacy and side effects of intervention in these pathways. As a proof-of-concept, the GRS associated with lower FEV1/FVC specific to PI3K–Akt signaling was associated with increased risk of COPD but a lower risk of diabetes; PI3K inhibition impairs glucose uptake in muscle and increases hepatic gluconeogenesis, contributing to glucose intolerance and diabetes41. The PheWAS and druggability analyses we conducted have the potential to identify drug repurposing opportunities for COPD.
The patterns of pleiotropy we show through PheWAS for individual variants, trait-specific GRS and pathway-partitioned GRS may help explain variants and pathways that increase susceptibility to more than one disease and thereby predispose to particular patterns of multimorbidity. For example, the elastic fiber pathway GRS was associated with increased risk of muscular (for example, hernias) and musculoskeletal conditions related to connective-tissue laxity. Our findings also further inform the complex relationship between height, BMI and obesity, and lung function and their genetic determinants5,42. Lung-function and height associations were uncorrelated, and height relationships differed between GRS for different lung-function traits, and even between sitting and standing height for the same trait. The pathway-partitioned GRS analyses indicate that the relationship between genetic variants, height and lung-function traits depends on the pathways through which the variants act.
The last comprehensive attempt to map lung-function-associated variants to genes identified 107 putative causal genes, mostly through eQTLs only, and only eight genes were then implicated by ≥2 criteria5. In contrast, we implicated 559 causal genes meeting ≥2 criteria by drawing on new data and methodologies, such as single-cell epigenome data, rare variant associations identified in sequencing data in the UK Biobank and similarity-based approach PoPS9. Nevertheless, our study has limitations. We focused on multi-ancestry rather than ancestry-specific signals, as the sample sizes for lung-function genomics studies in all non-EUR ancestry groups were limited, particularly for the AFR ancestries4. Non-EUR ancestries are under-represented in genomic studies3, constraining GWAS and PheWAS studies in these populations. Correcting this will require substantial global investment in suitably phenotyped and genotyped studies, with appropriate community participation and workforce development. Improved sample sizes across all ancestries would improve power in ancestry-specific studies42 and fine mapping of multi-ancestry meta-analysis signals.
Strategies for in silico mapping of association signals to causal genes are evolving and difficult to evaluate without a reference set of fully functionally characterized lung-function-associated variants and causal genes. Our variant-to-gene mapping framework parallels one that was recently adopted10 and could help prioritization of genes for functional experiments such as gene editing in relevant organoids with appropriate readouts to confirm mechanism. An additional limitation is that classifications of pathways may be imperfect; we used multiple pathway classifications as it is unclear which is superior across all component pathways and we present the pathway-partitioned PheWAS results as a resource to others.
In summary, our multi-ancestry study highlights new putative causal variants, genes and pathways, some of which are targeted by existing drug compounds. These findings bring us closer to understanding mechanisms underlying lung function and COPD and will inform functional genomics experiments to confirm mechanisms and consequently guide the development of therapies for impaired lung function and COPD.
Methods
GWAS in each cohort
Following cohort-level quality control of the lung-function phenotypes (Supplementary Note), all phenotypes were rank inverse-normal transformed after adjustment for age, sex, height, smoking, ancestry principal components and relatedness (mixed models in BOLT-LMM or SAIGE). Quality control of the imputation and association summary statistics in each cohort was performed by the central analysis team (Supplementary Note). We assigned each cohort to one of the five 1000 Genomes super-populations—EUR, AFR, AMR, EAS or SAS—based on self-reported ancestry, apart from the UK Biobank (57.4% of the total sample size), where we used ADMIXTURE v1.3.0 (ref. 43) to determine ancestry (Supplementary Note and Supplementary Table 4). We also acquired lung-function-association results from each cohort using untransformed phenotypes for analysis using MR-MEGA.
Meta-analysis
Before meta-analysis, association statistics in each cohort were adjusted by the LD-score regression intercept calculated in each cohort to adjust for any residual confounding (Supplementary Table 5); the appropriate ancestry-specific LD reference was used for each cohort (10,000 UK Biobank samples for EUR and 1000 Genomes Project samples for AFR, AMR, SAS and EAS). Before meta-analysis, variants with imputation INFO < 0.5 or minor-allele counts (MAC) < 3 were excluded. As transformed effects were not on comparable scales, we meta-analyzed across cohorts using sample-size weighted Z-score meta-analysis with METAL (released version 28 August 2018)44. No genomic control was applied post meta-analysis. Following meta-analysis, variants with MAC < 20 were excluded.
Signal selection and conditional analysis
We chose a genome-wide significance threshold of P < 5 × 10−9, as recommended from sequencing studies13. We selected 2-Mb regions centered on the most significant variant for all regions containing a variant with P < 5 × 10−9. Regions within 500 kb of each other were merged for conditional analysis. Stepwise conditional analysis was run in each region in each cohort using GCTA v1.93.2beta45 with an ancestry-specific LD reference for each cohort (Supplementary Note), and then the conditional results were meta-analyzed across cohorts and any new conditionally independent signals with P < 5 × 10−9 were added to our list of signals. We used moloc v0.1.0 (ref. 46) to co-localize signals across the four lung-function traits to obtain a set of distinct signals, which were then co-localized with previously reported signals to obtain a set of novel lung-function signals (Supplementary Note).
Exclusion of smoking signals from follow-up
We checked our sentinels for association with the smoking quantitative traits ‘age of initiation’ (n = 262,990) and ‘cigarettes per day’ (n = 263,954), and the binary traits ‘smoking cessation’ (n = 139,453 cases and n = 407,766 controls) and ‘smoking initiation’ (n = 557,337 cases and n = 674,754 controls) in the GWAS and Sequencing Consortium of Alcohol and Nicotine use (GSCAN) consortium47 (proxies with a squared correlation coefficient (r2) > 0.8 were checked for sentinels not present in GSCAN). We excluded eight lung-function signals from further analysis, which we determined to be primarily driven by smoking behavior (Supplementary Table 26), according to the following criteria: (1) P < 4.86 × 10−5 (Bonferroni-corrected 5% threshold for 1,028 signals) for association with any smoking trait and (2) the same ‘risk’ allele that increases smoking exposure behavior and decreases lung function.
Heritability estimate
We calculated the proportion of variance explained by the sentinels reported for each trait using the formula
where n is the number of variants, fi and βi are the frequency and effect estimates of the ith variant from the UK Biobank European ancestry untransformed results, respectively, and V is the phenotypic variance (always one as our phenotypes were inverse-normal transformed). We assumed a heritability of 40% (refs. 48,49) to estimate the proportion of additive polygenic variance.
Ancestry-adjusted trans-ethnic meta-analysis using MR-MEGA
To improve the fine-mapping resolution using LD differences between ancestries and to estimate the heterogeneity of variant associations attributable to ancestry, we undertook multi-ancestry meta-regression using MR-MEGA v0.2 (ref. 7), which incorporates axes of genetic ancestry as covariates. MR-MEGA uses multidimensional scaling of allele frequencies across cohorts to derive principal axes of genetic variation to use for ancestry adjustment (Supplementary Note). The location of the cohorts on the first two multidimensional scaling-derived principal components, plotted in Supplementary Fig. 17, shows clustering in accordance with the assigned ancestry groups. We used four principal components for ancestry adjustment, as this captured most of the variance. MR-MEGA implements genomic control at study level; therefore, no further genomic control was applied. We ran MR-MEGA at each locus containing ≥1 signals; in the loci with multiple signals, we ran MR-MEGA multiple times, each time conditioning on all except one signal at the locus. For each sentinel, we obtained an estimated ancestry-associated (P-value_ancestry_het) and residual (P-value_residual_het) heterogeneity. In addition, MR-MEGA reports the log-transformed Bayes factor, which can be used for the construction of credible sets.
Effects in children
To obtain unbiased effect estimates for comparison between adults and children, we first redefined 1,077 lead SNPs for lung function in the UK Biobank EUR population (n = 320,656) by selecting 1-Mb regions centered on the most significant variant for regions containing a variant with P < 5 × 10−8. For these SNPs, we then took the untransformed effect estimates from the meta-analysis of the non-UK Biobank EUR cohorts (34 cohorts for FEV1 and FVC, n = 128,071; 33 cohorts for FEV1/FVC, n = 123,429; 15 cohorts for PEF, n = 60,122). Next, we meta-analyzed two EUR-ancestry children’s cohorts—ALSPAC and Raine Study (age, 13–15 yr, n = 6,070)—to obtain effect estimates in children at the new lead SNPs. To investigate the age-dependent effects of genetic variants on lung function, we compared the effect sizes estimated in adults and children using a Welch’s t-test; a Bonferroni significance threshold for 1,077 tests was applied (P < 4.64 × 10−5).
Cell-type and functional specificity
Stratified LD-score regression
We tested for enrichment of regulatory features at variants overlapping four histone marks (H3K27ac, H3K9ac, H3K4me3 and H3K4me1) that are specific to adult lung, fetal lung, and peripheral blood mononuclear primary and smooth-muscle-containing cell lines (colon and stomach) using stratified LD-score regression12. We only considered EUR-specific meta-analysis with 39 cohorts for FVC, FEV1 and FEV1/FVC (17 cohorts for PEF). For the analysis of cell-type-specific annotations, we assessed statistical significance at the 0.05 level after Bonferroni correction for 60 hypotheses tested. Given that these annotations are not independent, a Bonferroni correction is conservative. We also report results with FDR < 0.05 using the Benjamini–Hochberg method.
Regulatory and functional enrichment using GARFIELD
We tested enrichment of SNPs at functionally annotated regions (DNase I hypersensitivity hotspots, open chromatin peaks, transcription-factor footprints and formaldehyde-assisted isolation of regulatory elements, histone modifications, chromatin segmentation states, genic annotations and transcription-factor binding sites) using GARFIELD17. We used the EUR meta-analysis with 17 cohorts for PEF and 39 cohorts for FVC, FEV1 and FEV1/FVC. We applied GARFIELD to DNase I hypersensitivity hotspot annotation in 424 cell lines and primary cell types from ENCODE and Roadmap Epigenomics and derived enrichment estimates at trait-genotype association P-value thresholds of P < 5 × 10−5 and P < 5 × 10−9.
Enrichment of annotations in respiratory-relevant cell types and tissues
We curated annotations from assays of respiratory-relevant cells and tissues—that is, (1) single-cell genome ATAC–seq data50 from 19 cell types (myofibroblast, pericyte, ciliated, T cell, club, capillary endothelial 1 and 2, basal, matrix fibroblast 1 and 2, arterial endothelial, pulmonary neuroendocrine, natural killer cell, macrophage, B cell, erythrocyte, lymphatic endothelial, alveolar type 1 and 2 (downloaded from https://www.lungepigenome.org/)), (2) ATAC–seq data for five human primary lung-cell types implicated in COPD pathobiology51 (large and small airway epithelial cells, alveolar type 2, pneumocytes and lung fibroblasts (downloaded from http://www.copdconsortium.org/)) and (3) tissue-specific transcription-factor binding sites from DNase-seq footprinting of 589 human transcription factors in lung and bronchus52. We tested for cell- and tissue-specific enrichment of these annotations at our lung-function signals using functional GWAS (fGWAS)14 (Supplementary Note).
Identification of putative causal genes and variants
eQTL and pQTL co-localization
Three eQTL resources were used for co-localization of lung-function signals with gene expression signals: (1) GTEx V8 (downloaded from https://www.gtexportal.org/, July 2020; tissues: stomach, small-intestine terminal ileum, lung, esophagus muscularis, esophagus gastroesophageal junction, colon transverse, colon sigmoid, artery tibial, artery coronary and artery aorta), (2) eQTLgen53 blood eQTLs and (3) UBC lung eQTL54. Two blood pQTL resources were used to co-localize with associations with protein levels, that is, INTERVAL pQTL55 and SCALLOP pQTL. The coloc_susie method56 was used to test eQTL and pQTL co-localization (Supplementary Note).
Rare variants from exome sequencing
We checked for rare (MAF < 1%) exonic associations near (±500 kb) our lung-function sentinels using both single-variant and gene-based collapsing tests from (1) 281,104 UK Biobank exomes from the AstraZeneca PheWAS Portal57 (https://azphewas.com/), (2) loss-of-function and missense variants in 454,787 UK Biobank participants58 and (3) gene-based tests on whole-exome imputation in 500,000 UK Biobank participants59. We used a threshold of P < 5 × 10−6 for both single-variant and gene-based tests (Supplementary Note).
Nearby Mendelian respiratory-disease genes
We selected rare Mendelian-disease genes from ORPHANET (https://www.orpha.net/) within ±500 kb of a lung-function sentinel that were associated with respiratory terms matching regular expression—that is, respir, lung, pulm, asthma, COPD, pneum, eosin, immunodef, cili, autoimm, leukopenia, neutropenia and Alagille syndrome. We implicated the gene if it had a corresponding respiratory term match in the disease name or if it occurs frequently in human phenotype ontology terms for that disease (Supplementary Note).
Nearby mouse-knockout orthologs with a respiratory phenotype
We selected human orthologs of mouse-knockout genes with phenotypes in the ‘respiratory’ category, as listed in the International Mouse Phenotyping consortium (https://www.mousephenotype.org/), within ±500 kb of a lung-function sentinel (Supplementary Note).
PoPS
We calculated a gene-level PoPS9 based on the assumption that if the associations enriched in genes share functional characteristics with a gene near to a lung-function signal, then that gene is more likely to be causal. The full set of gene features used in the analysis included 57,543 total features—40,546 derived from gene expression data, 8,718 extracted from a protein–protein interaction network and 8,479 based on pathway membership. In this study we prioritized genes for all autosomal lung-function signals within a 500-kb (±250 kb) window of the sentinel and reported the top prioritized genes in the region. For the signals that did not have prioritized genes within the 500-kb window, we looked for prioritized genes using a 1-Mb (±500 kb) window (Supplementary Note).
Annotation-informed credible sets
We used the enriched annotations in respiratory-relevant cell types and tissues and enriched genic annotations (Supplementary Table 12) to create annotation-informed 95% credible sets using fGWAS based on the MR-MEGA ancestry-adjusted meta-regression results (Supplementary Note). We implicated a putative causal missense variant if it accounted for >50% of the posterior probability in the credible set and annotated these using Ensembl Variant Effect Predictor60 to check for a deleterious effect by the SIFT, PolyPhen or CADD metrics.
Allocation of genes prioritized with ≥3 variant-to-gene to lung-function biology categories
We allocated prioritized genes with ≥3 criteria to different lung-function roles (epithelial, inflammatory, peripheral lung (including alveolus and endothelial), lung remodeling (including connective tissue), chest-wall movement and lung development) based on literature reviews, including GeneCards (https://www.genecards.org) and PubMed (https://pubmed.ncbi.nlm.nih.gov). Eighteen of the genes were difficult to assign to a specific category on this basis, mainly because they were involved in generic processes such as transcriptional control in a wide variety of cell types; these are not shown in Supplementary Fig. 12 but are included in Supplementary Table 13.
Interaction with smoking
Association testing for lung-function traits (FEV1, FVC, FEV1/FVC and PEF) was calculated separately in ever- and never-smoker subgroups and meta-analyzed across EUR-ancestry cohorts. We included untransformed phenotypes with ever- and never-smoking summary statistics (n = 28 cohorts) comprising 206,162 ever-smokers and 229,046 never-smokers. A z-test was used to compare genetic effect between the untransformed association results for the ever- and never-smokers:
where se is the standard error of the effect β. We considered a significant interaction any signal with a P < 4.9 × 10−5 (5% Bonferroni-corrected for 1,020 signals tested).
GRS
We selected four ancestry groups in the UK Biobank (UKB) as test datasets (SAS was excluded from GRS analyses because UKB SAS was the only cohort in the multi-ancestry analysis for SAS): UKB EUR, UKB AMR, UKB EAS and UKB AFR. All of the other cohorts except UKB SAS and Qatar Biobank were used as discovery datasets.
We repeated the multi-ancestry meta-regression (MR-MEGA), after excluding the four test GWAS, incorporating the same four axes of genetic variation as covariates to account for ancestry. Autosomal signals for each lung-function trait that were reported in the target ancestry population were included in downstream analysis for each ancestry. For ancestry j (j = EUR, AMR, EAS or AFR), we estimated ancestry-specific predicted allelic effects for the ith SNP to be used as weights in the multi-ancestry GRS by
where \(\bar x_{\mathrm{kj}}\) is the averaged position of discovery studies with ancestry j on the kth axis of genetic variation from multi-ancestry meta-regression, and \(\alpha _{0{\mathrm{i}}}\) and \(\alpha _{\mathrm{ki}}\) denote the intercept and effect of the kth axis of genetic variation for the ith SNP from the multi-ancestry meta-regression.
We ran each of the ancestry-specific fixed-effect meta-analyses after excluding the test GWAS from the ancestry group using METAL using the inverse-variance weighting method. For comparison, SNPs used as weights in multi-ancestry GRS were selected to build ancestry-specific GRS for each ancestry.
Testing GRS in independent COPD case–control cohorts
We tested the association of multi-ancestry GRS with COPD susceptibility in five EUR-ancestry COPD case–control studies: COPDGene (non-Hispanic white), ECLIPSE, GenKOLS, NETT/NAS and SPIROMICS (non-Hispanic EUR) (Supplementary Table 21). We also tested the association in two AFR ancestry COPD case–control studies: COPDGene (African American) and SPIROMICS (African American) (Supplementary Table 21). Associations were tested using logistic regression models, adjusted for age, age squared, sex, height and principal components. In each COPD case–control study, we divided individuals into deciles according to their weighted GRS. For each decile, logistic models were fitted to compare the risk of COPD for members of the test decile with those with the lowest decile (that is, those with the lowest genetic risk). The results were meta-analyzed by ancestry-specific study groups using the fixed-effect model.
PheWAS
We used Deep-PheWAS40, which addresses both phenotype matrix generation and efficient association testing while incorporating the following developments that are not yet available in current platforms and online resources: (1) clinically curated composite phenotypes for selected health conditions that integrate different data types (including primary and secondary care data) to study phenotypes that are not well captured by current classification trees; (2) integration of quantitative phenotypes from primary care data, such as pathology records and clinical measures; (3) clinically curated phenotype selection for traits that are extremely highly correlated and (4) GRSs. The platform includes 2,421 phenotypes in the UK Biobank, with a subset of 2,243 recommended for association testing—some phenotypes that are generated are used solely in the definition of other phenotypes. We removed the four measures of lung function and added seven phenotypes defined in-house (P4002-6) to give 2,246 as our final maximum number of phenotypes for association. Deep-PheWAS then filters these, requiring a minimum case number; we chose to keep the default settings of a 50-case minimum for binary phenotypes and a 100-case minimum for quantitative phenotypes. After limiting to EUR ancestry and filtering for case numbers, 1,909 phenotypes were left for association analysis (Supplementary Table 27). No additional phenotypes were removed when removing pairs related up to second degree (KING kinship coefficient ≥ 0.0884).
There are five types of phenotypes within Deep-PheWAS categorized according to the data and methods used to create them. Composite phenotypes are made using linked hospital and primary care data, including in some cases primary care prescription data, alongside any of the UK Biobank field-IDs (DFP), including self-reported non-cancer diagnosis and self-reported operations. Phecodes are defined using only linked hospital data (https://phewascatalog.org/phecodes_icd10). Formula phenotypes combine available data using bespoke R code per phenotype rather than the in-built functions of phenotype development available in Deep-PheWAS. Added phenotypes are lists of cases and controls that have been added to the PheWAS and not developed by the Deep-PheWAS phenotype matrix generation pipeline. More complete definitions for all none-added phenotypes can be found in the Deep-PheWAS description40. All phenotypes were adjusted for age, sex and the first ten principal components.
Single-variant PheWAS
We ran 28 single-variant PheWAS across 1,909 traits (Supplementary Table 27) in up to 430,402 unrelated EUR individuals in the UK Biobank. We selected the variant with the most significant P value for each of the 20 genes with ≥4 lines of evidence for being causal (Supplementary Table 13). A further seven variants were included in single-variant PheWAS that were putatively causal (accounted for >50% posterior probability in the credible set and had a deleterious annotation; Supplementary Table 14) but in a gene that was implicated by fewer than four lines of evidence. The single-variant PheWAS was aligned to the lung-function-trait decreasing allele. Where we noted associations with testosterone and SHBG, we also undertook sex-stratified PheWAS.
Association with trait-specific GRS
We created four GRSs for the UK Biobank EUR samples, one for each trait FEV1, FVC, FEV1/FVC and PEF, including all conditionally independent sentinel variants for the trait that were associated with P < 5 × 10−9, yielding 425, 372, 442 and 194 variants in each trait-specific GRS, respectively. Each of the four GRS were weighted by the effect sizes from the multi-ancestry meta-regression for the relevant trait and then checked for association with 1,909 traits in the PheWAS.
Association with pathway-specific GRS
We selected 29 pathways that were enriched at FDR < 10−5 for our 559 genes implicated by ≥2 lines of evidence (Supplementary Table 18). We created a weighted GRS (weights estimated from multi-ancestry meta-regression for FEV1/FVC) for each of the 29 pathways by including for each gene in the pathway (as for ‘Single-variant PheWAS’) the variant with the most significant P value for the trait that implicates the gene in our variant-to-gene mapping (Supplementary Table 13). Each of the 29 GRSs were then checked for association with 1,909 traits in the PheWAS.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Genome-wide summary statistics for the multi-ancestry meta-analysis are available at the GWAS Catalog (https://www.ebi.ac.uk/gwas/) under the accession codes GCST90244092, GCST90244093, GCST90244094 and GCST90244095.
Change history
25 September 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41588-023-01531-7
References
Young, R. P., Hopkins, R. & Eaton, T. E. Forced expiratory volume in one second: not just a lung function test but a marker of premature death from all causes. Eur. Respir. J. 30, 616–622 (2007).
GBD Chronic Respiratory Disease Collaborators. Prevalence and attributable health burden of chronic respiratory diseases, 1990–2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet Respir. Med. 8, 585–596 (2020).
Sirugo, G., Williams, S. M. & Tishkoff, S. A. The missing diversity in human genetic studies. Cell 177, 26–31 (2019).
Tobin, M. D. & Izquierdo, A. G. Improving ethnic diversity in respiratory genomics research. Eur. Respir. J. 58, 2101615 (2021).
Shrine, N. et al. New genetic signals for lung function highlight pathways and chronic obstructive pulmonary disease associations across multiple ancestries. Nat. Genet. 51, 481–493 (2019).
Wain, L. V. et al. Genome-wide association analyses for lung function and chronic obstructive pulmonary disease identify new loci and potential druggable targets. Nat. Genet. 49, 416–425 (2017).
Mägi, R. et al. Trans-ethnic meta-regression of genome-wide association studies accounting for ancestry increases power for discovery and improves fine-mapping resolution. Hum. Mol. Genet. 26, 3639–3650 (2017).
Barbeira, A. N. et al. Exploiting the GTEx resources to decipher the mechanisms at GWAS loci. Genome Biol. 22, 49 (2021).
Weeks, E.M. et al. Leveraging polygenic enrichments of gene features to predict genes underlying complex traits and diseases. Preprint at medRxiv https://doi.org/10.1101/2020.09.08.20190561 (2020).
Aragam, K. G. et al. Discovery and systematic characterization of risk variants and genes for coronary artery disease in over a million participants. Nat. Genet. 54, 1803–1815 (2022).
Wang, L. et al. Methodology in phenome-wide association studies: a systematic review. J. Med. Genet. 58, 720–728 (2021).
Finucane, H. K. et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat. Genet. 47, 1228–1235 (2015).
Pulit, S. L., de With, S. A. & de Bakker, P. I. Resetting the bar: statistical significance in whole-genome sequencing-based association studies of global populations. Genet. Epidemiol. 41, 145–151 (2017).
Pickrell, J. K. Joint analysis of functional genomic data and genome-wide association studies of 18 human traits. Am. J. Hum. Genet. 94, 559–573 (2014).
Repapi, E. et al. Genome-wide association study identifies five loci associated with lung function. Nat. Genet. 42, 36–44 (2010).
Bulik-Sullivan, B. K. et al. LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Iotchkova, V. et al. GARFIELD classifies disease-relevant genomic features through integration of functional annotations with association signals. Nat. Genet. 51, 343–353 (2019).
Cho, M. H. et al. Risk loci for chronic obstructive pulmonary disease: a genome-wide association study and meta-analysis. Lancet Respir. Med. 2, 214–225 (2014).
Soler Artigas, M. et al. Sixteen new lung function signals identified through 1000 Genomes Project reference panel imputation. Nat. Commun. 6, 8658 (2015).
Wyss, A. B. et al. Multiethnic meta-analysis identifies ancestry-specific and cross-ancestry loci for pulmonary function. Nat. Commun. 9, 2976 (2018).
Bullard, J. E., Wert, S. E., Whitsett, J. A., Dean, M. & Nogee, L. M. ABCA3 mutations associated with pediatric interstitial lung disease. Am. J. Respir. Crit. Care Med. 172, 1026–1031 (2005).
Yengo, L. et al. Meta-analysis of genome-wide association studies for height and body mass index in ∼700000 individuals of European ancestry. Hum. Mol. Genet. 27, 3641–3649 (2018).
Gudmundsson, J. et al. Genome-wide associations for benign prostatic hyperplasia reveal a genetic correlation with serum levels of PSA. Nat. Commun. 9, 4568 (2018).
Morrisey, E. E., Ip, H. S., Tang, Z., Lu, M. M. & Parmacek, M. S. GATA-5: a transcriptional activator expressed in a novel temporally and spatially-restricted pattern during embryonic development. Dev. Biol. 183, 21–36 (1997).
LaFemina, M. J. et al. Claudin-18 deficiency results in alveolar barrier dysfunction and impaired alveologenesis in mice. Am. J. Respir. Cell Mol. Biol. 51, 550–558 (2014).
Sweerus, K. et al. Claudin-18 deficiency is associated with airway epithelial barrier dysfunction and asthma. J. Allergy Clin. Immunol. 139, 72–81.e1 (2017).
Türeci, Ӧ, Mitnacht-Kraus, R., Wöll, S., Yamada, T. & Sahin, U. Characterization of zolbetuximab in pancreatic cancer models. Oncoimmunology 8, e1523096 (2019).
Krone, K. A. et al. Pulmonary manifestations of immune dysregulation in CTLA‐4 haploinsufficiency and LRBA deficiency. Pediatr. Pulmonol. 56, 2232–2241 (2021).
Shamriz, O. et al. Respiratory manifestations in LPS-responsive beige-like anchor (LRBA) protein-deficient patients. Eur. J. Pediatr. 177, 1163–1172 (2018).
Ferkingstad, E. et al. Genome-wide association meta-analysis yields 20 loci associated with gallstone disease. Nat. Commun. 9, 5101 (2018).
Yuan, T. et al. FGF10–FGFR2B signaling generates basal cells and drives alveolar epithelial regeneration by bronchial epithelial stem cells after lung injury. Stem Cell Rep. 12, 1041–1055 (2019).
Akkuş, G. et al. Hypogonadotropic hypogonadism due to novel FGFR1 mutations. J. Clin. Res. Pediatr. Endocrinol. 9, 95–100 (2017).
Klein, R. J. et al. Complement factor H polymorphism in age-related macular degeneration. Science 308, 385–389 (2005).
Miller, T. M. et al. Correlating phenotype and genotype in the periodic paralyses. Neurology 63, 1647–1655 (2004).
Monnier, N., Procaccio, V., Stieglitz, P. & Lunardi, J. Malignant-hyperthermia susceptibility is associated with a mutation of the ɑ1-subunit of the human dihydropyridine-sensitive L-type voltage-dependent calcium-channel receptor in skeletal muscle. Am. J. Hum. Genet. 60, 1316–1325 (1997).
Chiu, K. Y., Li, J. G. & Lin, Y. Calcium channel blockers for lung function improvement in asthma: a systematic review and meta-analysis. Ann. Allergy Asthma Immunol. 119, 518–523.e3 (2017).
Breuer, J. et al. VLA-2 blockade in vivo by vatelizumab induces CD4+FoxP3+ regulatory T cells. Int. Immunol. 31, 407–412 (2019).
Herwig, R., Hardt, C., Lienhard, M. & Kamburov, A. Analyzing and interpreting genome data at the network level with ConsensusPathDB. Nat. Protoc. 11, 1889–1907 (2016).
Krämer, A., Green, J., Pollard, J. Jr & Tugendreich, S. Causal analysis approaches in ingenuity pathway analysis. Bioinformatics 30, 523–530 (2014).
Packer, R.J. et al. Deep-PheWAS: a pipeline for phenotype generation and association analysis for phenome-wide association studies. Preprint at medRxiv https://doi.org/10.1101/2022.05.05.22274419 (2022).
Sahakian, N. et al. SGLT2 inhibitors as potentially helpful drugs in PI3K inhibitor-induced diabetes: a case report. Clin. Diabetes Endocrinol. 7, 17 (2021).
Zhu, Z. et al. A large-scale genome-wide association analysis of lung function in the Chinese population identifies novel loci and highlights shared genetic aetiology with obesity. Eur. Respiratory J. 58, 2100199 (2021).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Giambartolomei, C. et al. A Bayesian framework for multiple trait colocalization from summary association statistics. Bioinformatics 34, 2538–2545 (2018).
Liu, M. et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat. Genet. 51, 237–244 (2019).
Palmer, L. J. et al. Familial aggregation and heritability of adult lung function: results from the Busselton Health Study. Eur. Respir. J. 17, 696–702 (2001).
Wilk, J. B. et al. Evidence for major genes influencing pulmonary function in the NHLBI family heart study. Genet. Epidemiol. 19, 81–94 (2000).
Wang, A. et al. Single-cell multiomic profiling of human lungs reveals cell-type-specific and age-dynamic control of SARS-CoV2 host genes. eLife 9, e62522 (2020).
Benway, C. J. et al. Chromatin landscapes of human lung cells predict potentially functional chronic obstructive pulmonary disease genome-wide association study variants. Am. J. Respir. Cell Mol. Biol. 65, 92–102 (2021).
Funk, C. C. et al. Atlas of transcription factor binding sites from ENCODE DNase hypersensitivity data across 27 tissue types. Cell Rep. 32, 108029 (2020).
Võsa, U. et al. Large-scale cis- and trans-eQTL analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat. Genet. 53, 1300–1310 (2021).
Hao, K. et al. Lung eQTLs to help reveal the molecular underpinnings of asthma. PLoS Genet. 8, e1003029 (2012).
Sun, B. B. et al. Genomic atlas of the human plasma proteome. Nature 558, 73–79 (2018).
Wallace, C. A more accurate method for colocalisation analysis allowing for multiple causal variants. PLoS Genet. 17, e1009440 (2021).
Wang, Q. et al. Rare variant contribution to human disease in 281,104 UK Biobank exomes. Nature 597, 527–532 (2021).
Backman, J. D. et al. Exome sequencing and analysis of 454,787 UK Biobank participants. Nature 599, 628–634 (2021).
Barton, A. R., Sherman, M. A., Mukamel, R. E. & Loh, P.-R. Whole-exome imputation within UK Biobank powers rare coding variant association and fine-mapping analyses. Nat. Genet. 53, 1260–1269 (2021).
McLaren, W. et al. The Ensembl Variant Effect Predictor. Genome Biol. 17, 122 (2016).
Acknowledgements
This research has been conducted using the UK Biobank Resource under Application Number 648. We used the ALICE High Performance Computing Facility at the University of Leicester. This study was supported by BREATHE–The Health Data Research Hub for Respiratory Health (grant no. MC_PC_19004). This study was partially supported by the NIHR Leicester Biomedical Research Centre and the NIHR Nottingham Biomedical Research Centre; views expressed are those of the author(s) and not necessarily those of the NHS, the NIHR or the Department of Health. The funders had no role in the design of the study. This research was funded in part by the Wellcome Trust. For the purpose of open access, the authors have applied a CC BY public copyright license to any author accepted paper version arising from this submission. Cohort- and study-group-specific acknowledgements and funding information are in the Supplementary Note. This study was also supported by specific personal funding from the following: Wellcome Trust Institutional Strategic Support Fund (grant no. 204801/Z/16/Z) and BHF Accelerator Award AA/18/3/34220 to A.L.G.; NIH K08 HL136928, U01 HL089856, R01 HL155749 and a Research Grant from the Alpha-1 Foundation to B.D.H., B.D.H. also receives grant support from Bayer; Medical Research Council Clinical Research Training Fellowship (grant no. MR/P00167X/1) to C.J.; National Health and Medical Research Council (NHMRC) Ideas grant no. 2003629 and Department of Health Western Australia Merit Award 1186046 to B.H.M.; BBSRC CASE studentship with GSK to A.T.W.; GSK/British Lung Foundation Chair in Respiratory Research (L.V.W.); Wellcome Trust Investigator Award (WT202849/Z/16/Z) and Wellcome Trust Discovery Award (WT225221/Z/22/Z) to M.D.T.; MRC grant no. MR/N011317/1 to M.D.T. and L.V.W.; NIHR Senior Investigator Award to M.D.T. and I.P.H.; MRC grant no. G1000861 to I.S.; Wellcome Trust grant nos. WT098017 and WT064890 to A.P.M.; UKRI Innovation Fellowship at Health Data Research UK grant no. MR/S003762/1 to C.B.; MRC Human Genetics Unit program grant ‘Quantitative traits in health and disease’ (grant no. U. MC_UU_00007/10) to C.H., V.V. and S.M.K.; Academy of Finland (grant no. 336823) and Sigrid Juselius Foundation to J.K.; and MRC grant no. MR/P009581/1 to E.L.R.N.J.T. is a Wellcome Trust Investigator (202802/Z/16/Z), is the PI of the Avon Longitudinal Study of Parents and Children (MRC & WT 217065/Z/19/Z), is supported by the University of Bristol NIHR Biomedical Research Centre (BRC-1215-2001) and the MRC Integrative Epidemiology Unit (MC_UU_00011/1) and works within the CRUK Integrative Cancer Epidemiology Programme (C18281/A29019).
Author information
Authors and Affiliations
Consortia
Contributions
M.D.T. and I.P.H. supervised the study. M.D.T., I.P.H., N.S. and A.L.G. designed the study. N.S., A.G.I., J.C., R.P., R.J.H., R.J.T., C. Batini, K.A.F., C.J., K.C., N.N.P. and A.T.W. did the central analysis. F.D., A.P.M. and L.V.W. provided methodological and statistical advice. J.-H.L, I.S. and E.L.R. provided functional analyses. M.D.T., N.S., A.G.I., J.C., R.P. and I.P.H. wrote the paper. The following authors supervised or ran the analysis in the cohorts listed (full cohort names in Supplementary Note): ALHS cohort, A.B.W., J.J., S.S., M.L. and S.J.L.; ALSPAC cohort, R.G. and N.J.T.; ARIC cohort, M.R.B. and A.C.M.; B58C cohort, D.P.S.; BHS cohort, B.H.M., J.H., A. James and J.B.; Boston cohort, C.P., V.M., B.D.H., M.M., W.K., R.T.-S., P.B., E.K.S. and M.H.C.; CHS cohort, T.M.B. and S.A.G.; CKB cohort, A. Pozarickij, K.L., I.Y.M., Zhengming Chen, L. Li and R.G.W.; Croatia cohort, S.M.K., V.V., O.P. and I.R.; EPIC cohort, J. Luan, J.-H.Z. and C.L.; EXCEED cohort, A.T.W., R.C.F., X.W. and C. Brightling; FHS cohort, E.L., H.X., G.T.O. and J.D.; FinnTwin cohort, T.Palviainen, J.K., T.R. and K.H.P.; GS cohort, D.P., B.H.S., A.C. and C.H.; H2000 cohort, J.T.K., T.V., T. Laitinen and V.S.; HCHS cohort, T.S., J. Lee, M.L.D. and B.Y.; HUNT cohort, A.F.H., A.L., C.W., L.B. and B.B.; KORA cohort, C.F., S.K., C.G. and A. Peters; LBC1936 cohort, S.E.H., A.T., P.R. and S.R.C.; MESA cohort, A. Manichaikul, E.C.O., S.S.R. and R.G.B.; NEO cohort, T.O.F., P.S.H., A.M.S. and D.O.M.-K.; NFBC cohort, M.W. and M.-R.J.; ORCADES cohort, S.M.-W., N.P. and J.F.W.; PIVUS cohort, J.P.C., A. Mahajan, L. Lind and A.P.M.; QBB cohort, G.T., O.M.E.A. and K.S.; Raine Study, C.A.W., R.E.F., C.E.P. and G.L.H.; Rotterdam cohort, S.R.A.W., L. Lahousse, G.B. and A.G.U.; SAPALDIA cohort, M.I., A. Jeong and N.P.-H.; SHIP cohort, B.S., S.W., G.H. and C.O.S.; SPIROMICS cohort, X.H., V.E.O., D.A.M., E.R.B., S.B.G., N.G. and A.V.S.; TwinsUK cohort, K.P. and P.H.; UKHLS cohort, E.Z.; USC cohort, J.L.M., F.D.G., Zhanghua Chen and W.J.G.; VIKING cohort, K.A.K., P.K.J. and P.R.H.J.T.; and YFS cohort, T. Lehtimäki and O.T.R. All co-authors critically reviewed the paper.
Corresponding authors
Ethics declarations
Competing interests
M.D.T. and L.V.W. have previously received funding from GSK for collaborative research projects outside of the submitted work. I.P.H. has funded research collaborations with GSK, Boehringer Ingelheim and Orion. M.H.C. has received grant funding from GSK and Bayer, and speaking or consulting fees from AstraZeneca, Illumina and Genentech. B.D.H. has received grant funding from Bayer and speaking or consulting fees from AstraZeneca. I.S. has funded research collaborations with GSK, Boehringer Ingelheim and Orion outside of the submitted work. R.J.P., M.D.T., C.J. and L.V.W. have a funded research collaboration with Orion for collaborative research projects outside of the submitted work. The other authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Note, Figs. 1–17 and Tables 2–4, 9, 11, 12, 16 and 20–23.
Supplementary Table
Supplementary Tables 1, 5–8, 10, 13–15, 17–19 and 24–33.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shrine, N., Izquierdo, A.G., Chen, J. et al. Multi-ancestry genome-wide association analyses improve resolution of genes and pathways influencing lung function and chronic obstructive pulmonary disease risk. Nat Genet 55, 410–422 (2023). https://doi.org/10.1038/s41588-023-01314-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-023-01314-0
- Springer Nature America, Inc.
This article is cited by
-
Genome-wide association study identifies novel susceptible loci and evaluation of polygenic risk score for chronic obstructive pulmonary disease in a Taiwanese population
BMC Genomics (2024)
-
Human genetic associations of the airway microbiome in chronic obstructive pulmonary disease
Respiratory Research (2024)
-
Proteomic networks and related genetic variants associated with smoking and chronic obstructive pulmonary disease
BMC Genomics (2024)
-
Genetics of chronic respiratory disease
Nature Reviews Genetics (2024)
-
Genetic association of inflammatory marker GlycA with lung function and respiratory diseases
Nature Communications (2024)