Abstract
Cynomolgus macaque (Macaca fascicularis) and common marmoset (Callithrix jacchus) have been widely used in human biomedical research. Long-standing primate genome assemblies used the human genome as a reference for ordering and orienting the assembled fragments into chromosomes. Here we performed de novo genome assembly of these two species without any human genome-based bias observed in the genome assemblies released earlier. We assembled PacBio long reads, and the resultant contigs were scaffolded with Hi-C data, which were further refined based on Hi-C contact maps and alternate de novo assemblies. The assemblies achieved scaffold N50 lengths of 149 Mb and 137 Mb for cynomolgus macaque and common marmoset, respectively. The high fidelity of our assembly is also ascertained by BAC-end concordance in common marmoset. Our assembly of cynomolgus macaque outperformed all the available assemblies of this species in terms of contiguity. The chromosome-scale genome assemblies produced in this study are valuable resources for non-human primate models and provide an important baseline in human biomedical research.
Measurement(s) | sequence_assembly • genome • DNA |
Technology Type(s) | sequence assembly process • DNA sequencing |
Sample Characteristic - Organism | Macaca fascicularis • Callithrix jacchus |
Machine-accessible metadata file describing the reported data: https://doi.org/10.6084/m9.figshare.14665314
Similar content being viewed by others
Background & Summary
Cynomolgus macaque (or crab-eating macaque, Macaca fascicularis) and common marmoset (Callithrix jacchus), belonging to old world monkey and new world monkey respectively, have been widely used in human biomedical research and drug developments with expectations that they recapitulate human physiology and pathology1. Their genomes, consisting of 42 and 46 chromosomes in diploids2,3,4, respectively, were assembled initially using first- and second-generation sequencing technologies. Short-read de novo assembly was not able to resolve complex repetitive genomic regions, and the resulting contigs tended to remain fragmentary. Techniques such as mate-pair sequencing were commonly used to join the contigs into longer scaffolds, albeit with sequence gaps in between. Long-standing non-human primate (NHP) genome assemblies released earlier used the human genome for ordering and orienting the assemblies into chromosomes, which prevents the observation of intrinsic structural differences between the primate genomes5. For example, a large inversion of around 20 Mb was observed in chromosome 16 of the earlier marmoset genome assembly, which should have been the result of the ‘humanization’ bias6.
Recent technological advancements allow us to obtain chromosome-scale assemblies without relying on existing genome assemblies, where such errors or bias can be avoided. Single-molecule long-read sequencing (Pacific Biosciences [PacBio] and Oxford Nanopore Technologies) have drastically increased the contiguity of assemblies, and chromatin contact profiling with Hi-C and other techniques such as optical mapping have paved the way to reconstructing chromosome-scale sequences. Taking advantage of these recent advancements, the genome sequences of some non-human primates (NHP) including gorilla, orangutan, and chimpanzee were largely improved5,7, followed by the ones for bonobo (Bioproject accession: PRJNA526933), and Northern white-cheeked gibbon (PRJNA369439). High-quality genome assemblies of old world monkeys were also recently reported, such as the ones for Rhesus macaque by three different research groups (PRJNA4764748, PRJNA5094459, and PRJNA51419610), the ones for olive baboon (PRJNA52787411), golden snub-nosed monkey (PRJNA52494912), and Francois’s langur (PRJNA48853013). We had also previously produced pseudo-chromosome assembly by using PacBio long-reads for common marmoset6, where ‘humanized’ sequences were still used as a reference.
In this study, we focus on cynomolgus macaque and common marmoset to establish a solid baseline for human biomedical research. We performed genome sequencing and de novo assembly for both species by using PacBio long-reads, along with Hi-C for chromosome-scale scaffolding through optimized Hi-C data acquisition14. We corrected misjoins of the scaffolds through examination of the Hi-C contact maps. We further investigated misjoins through cross-checking alternate contigs made from PacBio data and corrected them. Lastly, we assessed the quality of the resultant assemblies based on the contiguity of assembled sequences (N50), completeness of conserved protein-coding genes, and consistency to BAC clone sequences. The results indicate that our genome assemblies are of chromosomal-scale contiguity, with nearly complete coverage of gene space, and highly concordant to the conventional genomic resource obtained independently to our data set. Above all, for cynomolgus macaque, our genome assembly achieved the optimal quality in comparison with other resources available for this species.
Methods
Sample preparation, sequencing, and de novo assembly
A cynomolgus monkey was purchased by Shiga University of Medical Science from Shin Nippon Biomedical Laboratories, Ltd through Angkor Primates Center Inc in Kingdom of Cambodia. The identification number is CE1976F in Shiga University of Medical Science and K150090 in Shin Nippon Biomedical Laboratories, Ltd. We followed the Reporting in Vivo Experiments (ARRIVE) guidelines developed by the National Centre for the Replacement, Refinement & Reduction of Animals in Research (NC3Rs). All animal experimental procedures were approved by the Animal Care and Use Committee of Shiga University of Medical Science (approval number: 2017-10-2 (H1)). Genomic DNA (gDNA) was extracted from the kidney of the 6-year-8-month-old female cynomolgus macaque with MagAttract HMW DNA kit (48) [Qiagen 5067–5583; Cat No 67563] according to the manufacturer’s instruction. After measurement of concentration with Qubit3 - Qubit dsDNA (BR) [Thermo Fisher Scientific] and optical density (OD) with nanoDrop 2000 [Thermo Fisher Scientific], ethanol precipitation of the extracted gDNA was performed with DNA Clean & Concentrator-10 (25) [Zymo research] to condense it. Then, concentration and OD of the gDNA were again measured with NanoDrop 2000, Qubit3 - Qubit dsDNA (BR): the amount, 70.6 μg (307 ng/μL × 230 μL); 260/280, 1.87; 260/230, 2.28. Also, the successful extraction of long gDNA was confirmed by electrophoresis with TapeStation ScreenTape GenomicDNA on TapeStation [Agilent]. Using PacBio Sequel II, approximately 81x sequence data of the cynomolgus macaque was sequenced with a read N50 length of 12.05 kbp. For common marmoset, the 43x PacBio RSII read dataset from our earlier sequencing effort (PRJDB8242), was used to reassemble the genome (Fig. 1a; Table 1).
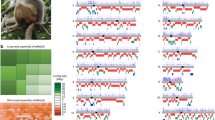
Quality assessment of the experiments. (a) Read length distribution of the common marmoset and cynomolgus macaque genomes, with a vertical line representing the N50 length. (b) Length distribution of the Hi-C DNA analyzed with Agilent TapeStation using the Genomic DNA ScreenTape. (c) Length distribution of the Hi-C library analyzed with Agilent TapeStation using the High Sensitivity D1000 ScreenTape.
To assemble both the NHP genomes, we employed a series of assembly programs (supplementary fig. S1). Based on the performances from the earlier marmoset genome assembly study, we chose the assemblers Flye15, Redbean (wtdbg2)16, SMARTdenovo17, and miniasm18,19. After de novo assembly, all the assemblies were processed by PacBio’s polishing tool, arrow, to eliminate possible sequencing errors. After considering the results from multiple de novo assembly tools, Redbean was chosen as the base assembler, as it produced the most contiguous assemblies for both the common marmoset (contig N50 of 8.04 Mb; Table 2) and the cynomolgus macaque (contig N50 of 5.84 Mb) genomes (Table 3).
Hi-C data acquisition and scaffolding
Hi-C libraries were constructed with the restriction enzyme DpnII following the iconHi-C protocol14, using the skeletal muscle tissues of an adult female marmoset and a cynomolgus macaque that were kept frozen in −80 °C after dissection and snap-freezing in liquid nitrogen. Fixed tissue materials containing 2 μg of DNA were used for the preparation of Hi-C DNA by in situ restriction digestion and ligation. Library preparation was performed using 1 μg of Hi-C DNA with 5 cycles of PCR for the marmoset and 6 cycles of PCR for the cynomolgus macaque. Quality control of the Hi-C DNA and the Hi-C library was performed as described in the iconHi-C protocol14. Quality control of the Hi-C DNA showed an expected pattern of a shift in size – shortening after digestion and elongation after ligation, indicating successful preparation of the Hi-C DNA (Fig. 1b). Quality control of the Hi-C library by restriction digestion confirmed the existence of expected ligation junction sequences inside the library molecules at a high proportion and successful generation of the library (Fig. 1c).
Sequencing to obtain Hi-C reads was performed on an Illumina HiSeq X in paired-ends with 151 cycles. The obtained reads were processed using Trim Galore to remove low-quality regions and adapter sequences. Post-sequencing quality control of the Hi-C libraries was performed as described previously14. Hi-C read statistics obtained by HiC-Pro20, using one million subsampled read pairs from the large scale sequencing data mapped to the PacBio contigs, confirmed high quality of the libraries while showing a high proportion of valid interaction read pairs, a low proportion of invalid ligation products (dangling end pairs), and a low proportion of contiguous restriction fragments (re-ligation pairs) (Supplementary table S1). Approximately 408 and 377 million Hi-C reads for the marmoset and the cynomolgus macaque were mapped to the Redbean contig sequences of the common marmoset and the cynomolgus macaque respectively using Juicer21. With the Juicer output files, Hi-C scaffolding was performed using 3d-dna22. Inversions and misjoins in the assemblies that occurred during the Hi-C scaffolding process were corrected by using Juicebox based on the frequency of Hi-C contacts23.
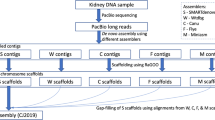
As a result of the genome assembly workflow involving de novo assembly followed by Hi-C scaffolding (Fig. 2a), the N50 lengths increased from approximately 8 to 135 Mb for the marmoset (Table 2) and from 5.8 to 149 Mb for the cynomolgus macaque (Table 3), and the number of scaffolds longer than 10 Mb presumed to be at the chromosome level became close to the actual number of chromosomes (Fig. 3). For the cynomolgus macaque, two pairs of the scaffold sequences longer than 10 Mb were merged into two pseudo-chromosomes, referred to as chromosomes 2 and 8 later, based on alignment concordance with the 3d-dna scaffold sequences of alternate assemblies, as well as the previous reference sequence.
Misjoin detection and gap-filling
All the alternate assemblies were also independently scaffolded using 3d-dna. These were later aligned against the Hi-C scaffolds from Redbean assembly to observe whether any misjoins were introduced by Hi-C scaffolding. Because the contiguity profiles of the assemblies are different across the employed assemblers, contigs which are broken into two or more sequences in one assembly could have been assembled into a single contiguous piece in another assembler (Supplementary fig. S1). This contiguity information from alternate assemblies was used to identify misjoins in the scaffold sequences from the Redbean assembly (Fig. 2b). When the alignments of the alternate assemblies against Redbean scaffolds were visualized using D-genies dot plots24, occasionally the scaffolds did not exhibit consistency in chromosomal structure among the 3d-dna scaffolds. These indicated the presence of misjoins introduced by Hi-C scaffolding (Supplementary fig. S2).
In case that such discrepancies in contig orders were observed in the alignments, a two-step check was performed, A) whether gaps are present in the Redbean scaffolds, and B) whether at least two of the alternate assemblies are consistent in those regions and discrepant against the Redbean scaffolds. If the above case is true, instead of re-ordering the misjoined scaffolds, we replaced the alignment block with one of the alternate assemblies making it a hybrid assembly. A similar procedure was also performed for gap-filling in regions without misjoins also. When a gap-containing region in Redbean scaffolds aligned to a gap-free region of contigs from alternate assemblies, those gap-containing regions were replaced with the corresponding blocks from alternate assemblies (Fig. 2b). When two or more alternate assemblies can be used to fill the gaps, one of them was randomly chosen to fill the gaps, followed by additional two rounds of polishing using arrow. This gap-filling procedure filled 1,147 gaps of length 573.94 kb in cynomolgus macaque, and 1,061 gaps of length 522.16 kb in common marmoset respectively.
Data Records
The N50 length of the contigs increased from 8.04 Mb to 24.82 Mb, and from 5.84 Mb to 26.27 Mb for the final assemblies of common marmoset and cynomolgus macaque genomes, respectively. The final genome assemblies displayed chromosome-sized sequences, designated here as ‘pseudo-chromosomes’, which exhibited scaffold N50 lengths of 132.27 Mb and 149.88 Mb, for the cynomolgus macaque and the common marmoset genomes, respectively (Table 2; Table 3).
The raw genome sequence data of the cynomolgus macaque are deposited at the DDBJ under the accession, DRA00958425. The Hi-C raw sequence data are deposited under the accessions, DRA00964126, and DRA00998727, for the cynomolgus macaque and the common marmoset genomes, respectively. The assembled genome sequences are deposited under the accessions, BLPH01000001-BLPH0100040028, and BLSI01000001-BLSI0100187229, for the cynomolgus macaque and the common marmoset genomes, respectively.
Technical Validation
Gene space completeness assessment
The web server gVolante30, in which two established pipelines CEGMA31 and BUSCO32 are implemented, was used to assess the sequence length distributions and gene space completeness in a uniform environment. The latter was based on the coverage of one-to-one reference orthologues with the ortholog search pipeline CEGMA and the gene set CVG that is specifically optimized to assess vertebrate genome sequences33. CEGMA identified pre-selected conserved protein-coding genes by using a combination of tblastn, and GeneWise for sequence alignment34,35, and GeneID for gene prediction36, in addition to HMMER3 for further homology search37, which are used to assess the completeness of the genome assemblies. The analysis of gene space completeness revealed a smaller number of missing CVG genes for both species, in comparison with the scores for the genome assemblies released earlier (Table 2; Table 3).
BAC-end alignment
BAC-end read pairs of the common marmoset from our earlier sequencing effort4, were aligned against the constructed assembly using bowtie238. After the misjoin detection and gap-filling procedure, the number of concordantly aligned read pairs increased by a count of 684, while the number of discordantly aligned read pairs decreased by a count of 401 (Table 2).
Comparison of common marmoset assembly against previous assemblies
For the common marmoset, NCBI hosts nine genome assemblies (https://www.ncbi.nlm.nih.gov/assembly/organism/9483/latest/), with cj1700 being the representative reference genome. The assembled genome in this study has a total scaffold length of 2.81 Gb with a contig N50 length of 24.82 Mb, in comparison to the total scaffold length of 2.79 Gb with the contig N50 length of 6.38 Mb from our earlier effort (CJ2019). Our newly assembled contigs have slightly lesser contiguity than the recently submitted assembly cj1700 which has a total scaffold length of 2.9 Gb with a contig N50 length of 26.62 Mb (Fig. 4a). Notably, our assembly is 23 Mb longer than cj1700 after excluding gap regions, and more than half of the sequence gaps in the pseudo-chromosomal sequences of the cj1700 assembly were identified as contiguous regions in our assembly. These results imply that these assemblies are similarly useful and, in some difficult regions, complementary to each other (Supplementary table S2).
Contiguity plots of the existing and obtained assemblies. N(X) plots comparing the contiguity profiles of existing and obtained genome assemblies of (a) common marmoset and (b) cynomolgus macaque. The contig lengths of MacFas_Jun2011 are well below 10 kb and did not fit into the scale of the plotted graph, and hence MacFas_Jun2011 was omitted from the figure.
Comparison of cynomolgus macaque assembly against previous assemblies
NCBI hosts five cynomolgus macaque genome assemblies (https://www.ncbi.nlm.nih.gov/assembly/organism/9541/latest/), with the macFas5 genome assembly being the representative reference genome (Table 3). Three of the assemblies including the representative genome have contig N50 lengths shorter than 100 kbp. In contrast, the recently submitted Macaca_Fascicularis_6.0 assembly had produced a much larger contig N50 length of 21.34 Mb. It has a total scaffold length 2.91 Gb, which is smaller than that of the representative reference genome’s 2.95 Gb. The assembly from our present study has a contig N50 length of 26.27 Mb and a total scaffold length 2.91 Gb. It is 65 Mb shorter than Macaca_Fascularis_6.0 even after excluding gap regions, and the ideal total length of the assembly is not yet conclusive. Notably, our assembly outperformed all the assemblies in terms of contiguity (Fig. 4b) and its coverage of orthologous genes in our assembly achieved the highest ratio, 93%, whereas the one in Macaca_Fascicularis_6.0 was 90% (Table 3). It indicates that the assembly produced here outperformed the rest not only in terms of contiguity, but also in base accuracy.
Comparison against other non-human primate genome assemblies
The contiguity of the two genomes assembled in this study was compared against other non-human primate genomes such as those of Rhesus macaque, Francois langur, chimpanzee, gorilla, orangutan, white-cheeked gibbon, golden snub-nosed monkey, bonobo, and olive baboon. Although there were three different sub-species genome assemblies for Rhesus macaque, we herein considered only the best assembly out of them in terms of contiguity. In comparison to the other non-primate genome assemblies, it was evident that the common marmoset and cynomolgus macaque assemblies produced the second and the third-best contiguities, with the Rhesus macaque genome assembly8, being the best among all in terms of contiguity (Supplementary fig. S3).
Code availability
The versions of the tools used and their parameters are described as follows.
Flye v2.3.6-release:
flye --pacbio-raw -g 2.9 g
Redbean (Wtdbg2) v2.3:
wtdbg2 -x rsII -g 2.7 g -L 5000
wtpoa-cns -i ctg.lay.gz
For cynomolgus macaque, rsII was replaced by sq.
SMARTdenovo (git commit 3d9c22e25bdf4caf6c08ea1acb41ee58e52f61a8):
Default parameters with consensus generation.
Minimap2 v2.10-r761 and miniasm (git commit 17d5bd12290e0e8a48a5df5afaeaef4d171aa133):
minimap2 -x ava-pb | gzip -1 > m.paf.gz
miniasm -f reads.fastq m.paf.gz > tigs.gfa
For cynomolgus macaque, only reads longer than 10 kb were considered for the assembly.
Hi-C scaffolding:
Trim Galore v0.4.5: --paired --phred33 -e 0.1 -q 30
HiC-Pro v2.11.1: default parameters
Juicer v20180805: default parameters
3d-dna v20180929: -m haploid -i 5000 -r 2
Juicebox v1.3.6
Polishing:
Pbmm2 v0.12.0:
pbmm2 align assembly.referenceset.xml reads.subreadset.xml aln.alignmentset.xml --sort -j 18 -J 18 -m 5000 M
Variant Caller v2.3.2:
The consensus sequence was split into 50 parts and the arrow algorithm was executed using default parameters.
Arrow polishing was iteratively executed twice for flye and SMARTdenovo assemblies, and thrice for miniasm and redbean assemblies.
Validation:
gVolante v1.2.1: (CEGMA v2.5, CVG v10042017, NCBI BLAST v2.2.31, GeneWise v2.2.3-rc7, GeneID v1.4, HMMER v3.1b2)
cegma.mod.pl --interlen 100000 --boundaries 10000 --ext --protein CVG/CVGs.fa --hmm_profiles CVG/hmm_profiles --cutoff_file CVG/profiles_CVG_cutoff.tbl --complete_file CVG/completeness_CVG_cutoff.tbl
References
Harding, J. D. Genomic tools for the use of nonhuman primates in translational research. ILAR J. 58, 59–68 (2017).
Yan, G. et al. Genome sequencing and comparison of two nonhuman primate animal models, the cynomolgus and Chinese rhesus macaques. Nat. Biotechnol. 29, 1019–1023 (2011).
The Marmoset Genome Sequencing and Analysis Consortium. The common marmoset genome provides insight into primate biology and evolution. Nat. Genet. 46, 850–857 (2014).
Sato, K. et al. Resequencing of the common marmoset genome improves genome assemblies and gene-coding sequence analysis. Sci. Rep. 5, 16894 (2015).
Kronenberg, Z. N. et al. High-resolution comparative analysis of great ape genomes. Science 360, eaar6343 (2018).
Jayakumar, V. et al. An improved de novo genome assembly of the common marmoset genome yields improved contiguity and increased mapping rates of sequence data. BMC Genomics 21, 243 (2020).
Gordon, D. et al. Long-read sequence assembly of the gorilla genome. Science 352, aae0344–aae0344 (2016).
Warren, W. C. et al. Sequence diversity analyses of an improved rhesus macaque genome enhance its biomedical utility. Science 370, (2020).
Cirelli, K. M. et al. Slow delivery immunization enhances HIV neutralizing antibody and germinal center responses via modulation of immunodominance. Cell 177, 1153–1171.e28 (2019).
He, Y. et al. Long-read assembly of the Chinese rhesus macaque genome and identification of ape-specific structural variants. Nat. Commun. 10, 4233 (2019).
Batra, S. S. et al. Accurate assembly of the olive baboon (Papio anubis) genome using long-read and Hi-C data. GigaScience 9, giaa134 (2020).
Wang, L. et al. A high-quality genome assembly for the endangered golden snub-nosed monkey (Rhinopithecus roxellana). GigaScience 8, giz098 (2019).
Liu, Z. et al. Genomic mechanisms of physiological and morphological adaptations of limestone langurs to Karst habitats. Mol. Biol. Evol. msz301 (2019).
Kadota, M. et al. Multifaceted Hi-C benchmarking: what makes a difference in chromosome-scale genome scaffolding? GigaScience 9, giz158 (2020).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 37, 540–546 (2019).
Ruan, J. & Li, H. Fast and accurate long-read assembly with wtdbg2. Nat. Methods 17, 155–158 (2020).
Liu, H., Wu, S., Li, A. & Ruan, J. SMARTdenovo: a de novo assembler using long noisy reads. Gigabyte 2021, 1–9 (2021).
Li, H. Minimap and miniasm: fast mapping and de novo assembly for noisy long sequences. Bioinformatics 32, 2103–2110 (2016).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16, 259 (2015).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3, 95–98 (2016).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95 (2017).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
Cabanettes, F. & Klopp, C. D-GENIES: dot plot large genomes in an interactive, efficient and simple way. PeerJ 6, e4958 (2018).
DNA Databank of Japan https://ddbj.nig.ac.jp/DRASearch/submission?acc=DRA009584 (2020).
DNA Databank of Japan https://ddbj.nig.ac.jp/DRASearch/submission?acc=DRA009641 (2020).
DNA Databank of Japan https://ddbj.nig.ac.jp/DRASearch/submission?acc=DRA009987 (2020).
DNA Databank of Japan https://ddbj.nig.ac.jp/public/ddbj_database/wgs/BL/BLPH.gz (2020).
DNA Databank of Japan https://ddbj.nig.ac.jp/public/ddbj_database/wgs/BL/BLSI.gz (2020).
Nishimura, O., Hara, Y. & Kuraku, S. gVolante for standardizing completeness assessment of genome and transcriptome assemblies. Bioinformatics 33, 3635–3637 (2017).
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Hara, Y. et al. Optimizing and benchmarking de novo transcriptome sequencing: from library preparation to assembly evaluation. BMC Genomics 16, 977 (2015).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Birney, E., Clamp, M. & Durbin, R. GeneWise and Genomewise. Genome Res. 14, 988–995 (2004).
Parra, G., Blanco, E. & Guigó, R. GeneID in Drosophila. Genome Res. 10, 511–515 (2000).
Eddy, S. R. Accelerated profile HMM searches. PLoS Comput. Biol. 7, e1002195 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Acknowledgements
This study was supported Drug Discovery & Development program in the Japan Agency for Medical Research and Development (AMED) under grant number JP17kk0305008, and research grants to RIKEN Preventive Medicine and Diagnosis Innovation Program and RIKEN Center for Integrative Medical Sciences from The Ministry of Education, Culture, Sports, Science and Technology (MEXT). In addition, Y.S. was supported by JSPS KAKENHI Grant Numbers 18H04127, and a Grant-in-Aid for Scientific Research on Innovative Areas “Frontier Research on Chemical Communications” [no. 17H06410] from the Ministry of Education, Culture, Sports, Science and Technology of Japan. Non-Clinical Evaluation Expert Committee in Drug Evaluation Committee in Japan Pharmaceutical Manufacturers Association (JPMA) gave us constructive comments and warm encouragements. The authors have had a lot of supports and encouragements of discussion with Dr. Takao Inoue in the National Institute of Health Sciences. Special thanks to Dr. Yoshihide Hayashizaki in RIKEN Preventive Medicine and Diagnosis Innovation Program and Dr. Yasushi Okazaki in RIKEN Center for Integrative Medical Sciences.
Author information
Authors and Affiliations
Contributions
Y.Sa., J.K., H.K., Y.M. conceived, designed and planned the project; Y.Y. managed the project; V.J., Y.Sa. assembled and analyzed the genome; M.K. constructed the Hi-C library; O.N., V.J., performed Hi-C scaffolding; O.N., M.K., V.J., S.K. analyzed Hi-C scaffolding results; V.J., O.N. resolved misjoins in the Hi-C scaffolds; V.J., O.N., M.K., S.K., N.K., H.K. analyzed the data and performed technical validation; M.N., T.T., Y.Se., E.S. dissected tissues for genomic DNA; S.N., M.E., E.S. contributed to the health qualification of the monkeys; V.J. handled data submission; V.J., O.N., M.K., S.K., H.K., Y.S. wrote the manuscript; All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
The Creative Commons Public Domain Dedication waiver http://creativecommons.org/publicdomain/zero/1.0/ applies to the metadata files associated with this article.
About this article
Cite this article
Jayakumar, V., Nishimura, O., Kadota, M. et al. Chromosomal-scale de novo genome assemblies of Cynomolgus Macaque and Common Marmoset. Sci Data 8, 159 (2021). https://doi.org/10.1038/s41597-021-00935-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-021-00935-6
- Springer Nature Limited
This article is cited by
-
Reduction of bitter taste receptor gene family in folivorous colobine primates relative to omnivorous cercopithecine primates
Primates (2024)
-
Long-read assembly of major histocompatibility complex and killer cell immunoglobulin-like receptor genome regions in cynomolgus macaque
Biology Direct (2022)
-
Identification and characterization of the T cell receptor (TCR) repertoire of the cynomolgus macaque (Macaca Fascicularis)
BMC Genomics (2022)