Abstract
Red claw crayfish (Cherax quadricarinatus) is an aquatic crustacean with considerable potential for the commercial culture and an ideal model for studying the mechanism of sex determination. To provide better genomic resources, we assembled a chromosome-level genome with a size of 5.26 Gb and contig N50 of 144.33 kb. Nearly 90% of sequences were anchored to 100 chromosomes, which represents the high-quality crustacean genome with the largest number of chromosomes ever reported. The genome contained 78.69% repeat sequences and 20,460 protein-coding genes, of which 82.40% were functionally annotated. This chromosome-scale genome would be a valuable reference for assemblies of other complex genomes and studies of evolution in crustaceans.
Similar content being viewed by others
Background & Summary
Crustaceans are a diverse and ancient group of arthropods1, and are not only essential components of the marine and freshwater environments, but also an interesting model for the study of evolutionary biology and developmental biology. However, due to the high complexity, assembly of complete and exact crustacean genomes is difficult, let alone genomes at the chromosome level2.
Cherax quadricarinatus, also known as the red claw crayfish, is a large tropical freshwater crustacean with significant commercial interest for global aquaculture3. Intersexuality appears relatively widespread throughout gonochoristic crustaceans and has been reported in several crayfish species4. In red claw crayfish, the intersex individuals undergo a dramatic morphological and physiological sex shift, which makes it a fascinate model to study the mechanisms underlying sex determination and differentiation of crustacean. Although a genome of this species has been reported previously, with uncomplete and fragmental genome assembly (assembled genome size, 3.24 Gb and Contig N50, 33 kb), it still prevents many studies from going deep5. Here, we de novo assembled a chromosome-level genome of red claw crayfish with the assembled genome size of 5.26 Gb and contig N50 of 144,316 bp. This high-quality genome would enrich the genomic resources of crustaceans and provides basic data for further genome-wide selective breeding.
Methods
Sample collection and genomic sequencing
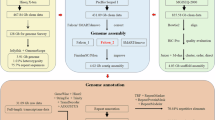
All samples used in this study were from a healthy male adult red claw crayfish farmed in Honghai Co., LTD., Zhejiang, China. Fresh muscle and haemolymph were used for whole genomic sequencing and Hi-C sequencing, respectively. Seven tissues including muscle, intestine, eyestalk, hepatopancreas, gills, stomach, and antennal gland were used for transcriptomic sequencing. Isolation of DNA/RNA, construction of libraries and genomic sequencing were carried out according to protocols from https://www.protocols.io/widgets/doi?uri=dx.doi.org/10.17504/protocols.io.bs8inhue.
For whole genomic sequencing (WGS), the genomic DNA was sonicated into ~250 bp fragments that used to build the 100 bp paired-end (PE100) sequencing library. The library was then sequenced on the BGISEQ-500 platform and generated 280.51 Gb raw data, which covered ~58X of the estimated genome (Table 1).
For PacBio Continuous Long Reads (CLR) sequencing, seven sequencing libraries were constructed using ~20Kb high-quality molecular DNA fragments. All libraries were sequenced on the PacBio Sequel II platform, which generated 568.55 Gb raw data with an N50 of 17,393 bp (Table 1).
For the construction of Hi-C library, DNA was fixed with formaldehyde solution and isolated from nuclei, and digested with MboI, the digested fragments were labeled with biotinylated nucleotides. Eight libraries were sequenced on the BGISEQ-500 platform and produced a total of 542.71 Gb raw data, which covered ~105X of the estimated genome (Table 1).
Seven RNA libraries were constructed according to the protocols and sequenced on the BGISEQ-500 platform, generating a total of 136.96 Gb raw data (Table 1).
Genome survey
Raw PE100 reads were firstly filtered by SOAPnuke (v1.6.5)6 with parameters of “–M 1 –d –A 0.4 –n 0.05 –l 10 –q 0.4 –Q 2 –G –5 0”, and 240 Gb clean data were retained (Table 1). Then Jellyfish (v2.2.6)7 was used to count k-17mers and GenomeScope8 was used to estimate the size, heterozygosity, and repetitive sequences of the genome at 4.74 Gb, 0.86% and 85.6%, respectively (Fig. 1a).
Genome assembly of the red claw crayfish. (a) The 17-mer analysis of the genome. (b) The karyotypic analysis. The karyotype formula of the male is n = 100 = 36 m + 33 sm + 14 st + 17 t. (c) The linear regression analysis between sequence length and physical length of chromosomes. (d) Genomic features.
Chromosome karyotyping
The number and length of chromosomes in red claw crayfish were obtained by karyotyping experiment using 15 male adults, according to the published pipeline9. Chromosomes were measured using Adobe Photoshop CS6 measurement tools under a magnification of 600 × . The chromosome pairs were classified following the nomenclature of Levan (1964)10 into m = metacentric (long arm/short arm (r) = 1–1.7), sm = submetacentric (r = 1.7–3), st = subtelocentric (r = 3–7), and a = acrocentric (r > 7). The karyotype formula of the male red claw crayfish is n = 100 = 36 m + 33 sm + 14 st + 17 t (Fig. 1b), and the arm lengths data were listed in Supplementary Table 1.
Genome assembly
Reads longer than 5 kb were kept from raw Pacbio CLR reads and corrected by Canu (v1.5)11, based on which the draft genome was assembled by Wtdbg212 with parameters of “-p 21 -E 2 -S 4 -s 0.05 -L 5000 -X 40”. The draft genome was further polished by Pilon13 using clean PE100 reads with default parameters, giving an assembly with the size of 5.26 Gb and the contig N50 of 144.33 kb (Table 2).
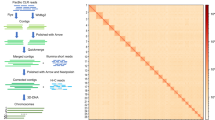
Based on the polished genome, 84.34 Gb Hi-C data were validated through quality control by Hi-C-Pro (v. 2.8.0)14, which were then applied for chromosomal reconstruction by Juicer (v1.5)15 and 3D-DNA (3D-de novo assembly)16. To get more precise chromosomes, we manually made some adjustments according to the chromosomal interaction heatmap by Juicebox17 (Fig. 2). Finally, a total of 4.70 Gb sequences were anchored to 100 chromosomes, of which the longest is 142.95 Mb and the shortest is 18.54 Mb (Supplementary Table 2). The linear regression analysis of karyotyping and assembly showed a high correlation (R2 = 0.9874) between the physical length and sequence length of 100 chromosomes (Fig. 1c), indicating the high-quality crustacean genome with the largest number of chromosomes ever reported.
Repeat annotation
Based on aligning the genome to the Repbase library by TRF (v.4.09)18, repetitive sequences were predicted by RepeatMasker (v. 3.3.0) and RepeatProteinMask (v. 3.3.0)19. In addition, transposable elements (TEs) were constructed and RepeatModeler (v1.0.8)20 (Table 3). All the above results together showed that red claw crayfish contains 78.69% repetitive sequences, among which TEs were most abundant (3,482 Mb) (Fig. 3, Table 4). Compared with other decapod crustaceans, the proportion of TES in crayfish was generally much higher.
Gene prediction
For homology-based gene prediction, the encoded protein sequences of six crustacean species include Cherax quadricarinatus (previous version), Eriocheir sinensis, Hyalella azteca, Macrobrachium nipponense, Penaeus vannamei, and Procambarus virginalis were aligned with the genomic sequence of red claw crayfish using BLAST20 and Genewise21 with default parameters. Augustus (v3.2.3)22 and Genscan23 were used for de novo gene prediction24. RNA reads were mapped to the genome by HISAT2 (v2.1.0)25 and gene structure were predicted by Stringtie (v1.2.2)26. Meanwhile, transcriptome was de novo assembled by Trinity (v2.1.1)27 and splicing variations were identified by PASApipeline (v2.4.1)28. EVidenceModeler (v1.1)29 was applied to integrate the above evidence and a total of 20,460 protein-coding genes were predicted, with average gene length and exon number per gene of 40,182.55 bp and 6.5, respectively (Tables 5, 6).
These genes were then functionally annotated through BLAST against NCBI non-redundant proteins (NR), TrEMBL, Gene Ontology (GO), SwissProt, and Kyoto Encyclopedia of Genes and Genomes (KEGG) protein databases. Finally, 16,859 genes accounting for 82.40% of the total were successfully annotated with at least one public functional database (Table 7).
The tRNAscan-SE30 was used to annotate the tRNAs based on annotated features such as isotype, anticodon, and tRNAscan-SE bit score. The rRNA sequences were annotated from homologous references in close species. MiRNAs and snRNAs were predicted by the INFERNAL31 based on the covariance model of the Rfam database. Totally 6,954 non-coding RNAs were predicted, including 25 miRNA, 1,448 rRNA, 5,023 tRNA and 458 snRNA genes (Table 8).
Data Records
The genomic WGS sequencing data were deposited in the SRA at NCBI SRR2241264932, SRR2241264133.
The genomic PacBio sequencing data were deposited in the SRA at NCBI SRR2241265434.
The transcriptomic sequencing data were deposited in the SRA at NCBI SRR2241265135, SRR2241265236, SRR2241265337, SRR2241263738, SRR2241263839, SRR2241263940, SRR2241264041.
The Hi-C sequencing data were deposited in the SRA at NCBI SRR2241264242, SRR2241264343, SRR2241264444, SRR2241264545, SRR2241264646, SRR2241264747, SRR2241264848, SRR2241265049.
The final chromosome assembly was deposited in GenBank at NCBI JAPQEV00000000050.
The genome annotation file is available in figshare51.
Technical Validation
The quality and quantity of total DNA was checked using agarose gel electrophoresis, and the concentration was determined using a NanoDrop 2000 spectrophotometer. RNA integrity was evaluated using an Agilent 2100 Bioanalyzer (Agilent Technologies, CA, USA). The sample used in our study had an RNA integrity number (RIN) larger than 8. To further assess the quality of the genome, clean PE100 reads were aligned back to the genome by BWA52, showing the mapping rate as high as 99.03%. The depth and GC content were also statistically analyzed within a 10Kb sliding window. Moreover, 85.7% completed and 6.2% fragmented BUSCOs53 (Benchmarking Universal Single-Copy Orthologs, v4.0) in arthropoda_odb9 database were identified, which showed a noticeable improvement than the previous version (81.3%).
Code availability
No specific code was developed in this work. The parameters of all commands and pipelines used for data processing are described in the Methods section. If no detailed parameters are mentioned for a software, the default parameters were used, as suggested by the developer.
Change history
23 May 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41597-023-02186-z
References
Stillman, J. H. et al. Recent advances in crustacean genomics. Integr. Comp. Biol. 48(6), 852–868 (2008).
Meehan, D., Xu, Z., Zuniga, G. & Alcivar-Warren, A. High frequency and large number of polymorphic microsatellites in cultured shrimp, Penaeus (Litopenaeus) vannamei [Crustacea: Decapoda]. Mar. Biotechnol. 5(4), 311–330 (2003).
Saoud, I. P., Ghanawi, J., Thompson, K. R. & Webster, C. D. A review of the culture and diseases of redclaw crayfish Cherax quadricarinatus (von Martens 1868). J WORLD AQUACULT SOC. 44(1), 1–29 (2013).
Ford, A. T., Fernandes, T. F., Read, P. A., Robinson, C. D. & Davies, I. M. The costs of intersexuality: a crustacean perspective. Mar. Biol. 145(5), 951–957 (2004).
Tan, M. H. et al. A giant genome for a giant crayfish (Cherax quadricarinatus) with insights into cox1 pseudogenes in decapod genomes. Front. Genet. 11, 201 (2020).
Chen, Y. et al. SOAPnuke: a MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. GigaScience. 7(1), gix120 (2018).
Li, R. et al. The sequence and de novo assembly of the giant panda genome. Nature. 463(7279), 311–317 (2010).
Vurture, G. W. et al. GenomeScope: fast reference-free genome profiling from short reads. Bioinformatics. 33(14), 2202–2204 (2017).
Shi, L. L., Xu, X. H., Zhang, L. &Li, Y. H. Comparative analysis of karyotype in female and male Procambarus clarkii. Journal of Anhui Agricultural University. 46(2), 234–241 (in Chinese) (2019).
Levan, A., Fredga, K. & Sandberg, A. A. Nomenclature for centromeric position on chromosomes. Hereditas. 52(2), 201–220 (1964).
Koren, S. et al. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res, 27(5), 722–736 (2017).
Ruan, J. & Li, H. Fast and accurate long-read assembly with wtdbg2. Nat. Methods. 17(2), 155–158 (2020).
Walker, B. J. et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PloS one. 9(11), e112963 (2014).
Servant, N. et al. HiC-Pro: an optimized and flexible pipeline for Hi-C data processing. Genome Biol. 16(1), 1–11 (2015).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst. 3(1), 95–98 (2016).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science. 356(6333), 92–95 (2017).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3(1), 99–101 (2016).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27(2), 573–580 (1999).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinformatics. 4, 1–4 (2009).
McGinnis, S. & Madden, T. L. BLAST: at the core of a powerful and diverse set of sequence analysis tools. Nucleic Acids Res. 32 (suppl_2), W20–W25 (2004).
Birney, E., Clamp, M. & Durbin, R. GeneWise and genomewise. Genome Res. 14(5), 988–995 (2004).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439 (2006).
Burge, C. & Karlin, S. Prediction of complete gene structures inhuman genomic DNA. Journal of J. Mol. Biol. 268(1), 78–94 (1997).
Majoros, W. H., Pertea, M. & Salzberg, S. L. TigrScan and GlimmerHMM: Two open source ab initio eukaryotic gene-finders. Bioinformatics. 20(16), 2878–2879 (2004).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods. 12(4), 357–360 (2015).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11(9), 1650–1667 (2016).
Grabherr, M. G. et al. Trinity: reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nat. Biotechnol. 29(7), 644 (2011).
Roberts, A., Pimentel, H., Trapnell, C. & Pachter, L. Identification of novel transcripts in annotated genomes using RNA-Seq. Bioinformatics. 27(17), 2325–2329 (2011).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. GENOME BIOL EVOL. 9(1), 1–22 (2008).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25(5), 955–964 (1997).
Nawrocki, E. P., Kolbe, D. L. & Eddy, S. R. Infernal 1.0: inference of RNA alignments. Bioinformatics. 25(10), 1335–1337 (2009).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412649 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412641 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412654 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412651 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412652 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412653 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412637 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412638 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412639 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412640 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412642 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412643 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412644 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412645 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412646 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412647 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412648 (2022).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRR22412650 (2022).
Chen, H. L. Genbank https://identifiers.org/ncbi/insdc.gca:GCA_026875155.1 (2022).
Chen, H. L. Cqu.final.last.gff. figshare https://doi.org/10.6084/m9.figshare.21599397 (2022).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics. 25(14), 1754–1760 (2009).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics. 31(19), 3210–3212 (2015).
Acknowledgements
The work was supported by the Zhejiang Science and Technology Major Program (2021C02069-4). We thank Dr. Jun Wang for his kindly help in manuscript polishing.
Author information
Authors and Affiliations
Contributions
Haipeng Liu, Bao Lou and Changwei Shao conceived the study and supervised the project. Honglin Chen collected the samples and wrote the manuscript. Rui Zhang performed the data analysis, figures drawing and manuscript writing. Feng Liu, Fangfang Liu, Jindong Ren and Baolong Niu supported in chromosome karyotyping and data uploading. Weidong Li assisted in data analysis.
Corresponding authors
Ethics declarations
Competing interests
Here we declare together, all data of this project were obtained from our independent research. There’s no conflict of interest in this manuscript.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, H., Zhang, R., Liu, F. et al. The chromosome-level genome of Cherax quadricarinatus. Sci Data 10, 215 (2023). https://doi.org/10.1038/s41597-023-02124-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02124-z
- Springer Nature Limited
This article is cited by
-
Genome assembly of redclaw crayfish (Cherax quadricarinatus) provides insights into its immune adaptation and hypoxia tolerance
BMC Genomics (2024)
-
CrustyBase v.2.0: new features and enhanced utilities to support open science
BMC Genomics (2024)
-
The chromosome-level genome assembly of the red swamp crayfish Procambarus clarkii
Scientific Data (2024)