Abstract
The blackfin Icefish (Chaenocephalus aceratus) belongs to the family Channichthyidae and the suborder Notothenioidei which lives in the Antarctic. We corrected the mis-scaffolds in the previous linkage map results by Hi-C analysis to obtain improved results for chromosome-level genome assembly. The final assembly analysis resulted in a total of 3,135 scaffolds, a genome size of 1,065.72 Mb, and an N50 of 33.46 Mb. 820.24 Mb, representing 88.88% of the total genome, is anchored to 24 chromosomes. The final gene set of 38,024 genes, including AFGPs, was annotated using RNA evidence, proteins, and ab-initio predictions. The complete percentage of BUSCO analysis is 92.7%. In this study, we aim to contribute to the study of polar fishes by improving the genome sequences of the blackfin icefish with the AFGP genes belonging to the Notothenoidei.
Similar content being viewed by others
Background & Summary
The Antarctic Ocean is a very cold and difficult place for any species to survive. The seawater temperature is at subzero levels even in summer, and the intertidal ecosystem does not function because ice covers the shoreline and coastal waters to depths ≥30 m. However, some species can survive in these extreme environments. The Antarctic marine fish fauna consists of approximately 275 species, 95 of which belong to the perciform suborder Notothenioidei. Some species have unusual adaptations, such as the presence of antifreeze glycoprotein (AFGP) in their blood or the absence of hemoglobin, to survive under these frigid conditions1,2. The blackfin icefish is a species of crocodile icefish belonging to the family Channichthyidae and the suborder Notothenioidei. Its natural habitat ranges from Southern Georgia to the northern part of the Antarctic Peninsula in the Atlantic sector of the Southern Ocean and Bouvetøya Island. It is found in shelf waters to a depth of 450–770 m3. Blackfin icefish species have thin, highly vascularized, scaleless skin; elongated bodies; and a weaker skeleton in comparison with most red-blooded notothenioid species. Their body structure makes them extremely vulnerable to injury4. Icefish, also known as white-blooded fish, belong to a unique family in that they are the only known vertebrates to lack hemoglobin. Consequently, their blood oxygen-carrying capacity is just 10% of that of other teleosts. The blood of the blackfin icefish Chaenocephalus aceratus has significantly fewer erythrocytes. The blood sample of C. aceratus does not have a trace of red color. Instead, it has a translucent whitish color. The plasma is clear. The cell mass at the bottom of a centrifuged hematocrit tube has been reported to be creamy white, accounting for approximately 1% of the blood content5. The 15 known species of the notothenioid family Channichthyidae, including C. aceratus, have the same diploid number of chromosomes (2n = 48), predominantly acrocentric chromosomes6.
A previous study7 reported the genome assembly of the blackfin icefish and published its genetic linkage map. However, its chromosome-level genome assembly remains unknown. Here, we report the upgraded chromosome-level whole-genome assembly of the blackfin icefish using the Hi-C approach with the tissue of the same individual used in the previous study. The genome assembly was highly consistent with the genetic linkage map at the chromosome level, and some mis-scaffolding in the genetic linkage map was rectified. We compared the chromosome-level genome sequence with that of another icefish, the South Georgia icefish (Pseudochaenichthys georgianus), to verify chromosomal conformity. For assessing chromosomal stability, we compared the sequences with those of medaka (Oryzias latipes), torafugu (Takifugu rubripes), and stickleback (Gasterosteus aculeatus). Moreover, to perform gene prediction more accurately, we reconstructed the annotation process using the integrated process of GeneMark8 and PASA pipeline9 with EVidenceModeler10. Using the customized prediction process, we predicted the functions of 10 copies of trypsinogen genes, nine copies of antifreeze glycoprotein (AFGP) genes, and two copies of AFGP/trypsinogen-like protease chimeric genes, and a trypsinogen-like protease gene with high tandem duplication at intron and exon levels.
Methods
Hi-C sequencing
Tissue sample of blackfin Icefish from the same individuals used in the previous study7 were used for Hi-C analysis. The DovetailTM Hi-C library was prepared using the DovetailTM Hi-C Library Kit (Dovetail Genomics, Santa Crus, CA, USA), according to the manufacturer’s instructions. Ground tissue (250 mg) was crosslinked with PBS/formaldehyde; the chromatin sample was then prepared with SDS and wash buffer. After normalizing the chromatin sample, 800 ng of chromatin was used to prepare the library. The chromatin was picked up using chromatin capture beads and then digested using a restriction enzyme. The end was labeled with biotin and ligated to form intra-aggregated DNA. After cross-link reversal, 200 ng of DNA was sheared using the Covaris system (Covaris Inc., Woburn, MA, USA). Sheared DNA fragments were end-repaired and ligated using an Illumina adapter. Ligated DNA was purified using streptavidin magnetic beads. Purified DNA was then amplified via PCR to enrich the fragments. Capillary electrophoresis verified the amplified libraries’ quality (Bioanalyzer System, Agilent Technologies, Palo Alto, CA, USA). Sequencing was performed using the Illumina NovaSeq 6000 system (Illumina Inc., San Diego, CA, USA), according to the protocols provided for 2 × 150 sequencing11.
Hi-C analysis with previous draft assembly
HiRise software, a pipeline for performing scaffolding analysis using proximity ligation data produced using the draft genome assembly, and Dovetail Hi-C technology were used for chromosome-level genome assembly12. The Hi-C reads were aligned to the draft assembly using SNAP. The positions of the mapped read pairs were used to construct a likelihood model of the genomic distance between read pairs. Genomic linking information between contigs was generated using the model and misjoins were corrected to construct a pseudomolecule-level scaffold genome. Juicer v.1.5.713,14 was used to generate a hic file containing contact matrices with duplicate removal from the linking data. The Hi-C raw sequence data were aligned using BWA-MEM15. A contact map plot was drawn in detail using Juicebox v.1.513, with the Juicer output being a hic file. DovetailTM HiRise allowed the upgrade from draft genome assembly to chromosome-level genome assembly within 24 chromosomal sequences (Fig. 1a). The longest scaffold length was 48 Mb, and the scaffold N50 value was 33 Mb (Table 1). We confirmed that there were 24 scaffolds of ≥10 Mb, consistent with the number of chromosomes in the blackfin icefish (2n = 48). Moreover, the total size of unplaced scaffolds was 262.76 Mb (Table 2).
Summary of the final genome assembly results. (a) Contact map plot of the blackfin icefish genome. The Hi-C raw read pairs were aligned with the genome sequences. The x and y axes indicate their positions. The red dots indicate the position of the read pairs, and a high density of red dots denotes that they are located on the same chromosome. (b) Correction of mis-scaffolding of the linkage group in the blackfin icefish genome by Hi-C analysis. Mis-scaffolding of the LG14 linkage group was confirmed by Hi-C analysis. The 3.26M-sized sequence of LG14 was located on part of LG7, and the high density of linkage (red dot) was confirmed on the contact map at the position. (c) Overview of the blackfin icefish genome. The features are arranged in the order of gene density, repeat density, GC contents, and GC skew from outside to inside at 1-Mb intervals across the 24 chromosomes.
Comparative genomics analysis
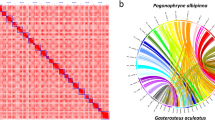
To compare genome sequences at the chromosome level, nucmer in the MUMmer software package v.4.02b16 was used with the parameters -c 1000 -l 1000 and add--mum for unique matching and avoiding repeat regions. For a clear chromosome comparison, only long sequences corresponding to chromosomes were extracted and compared; unordered contig or scaffold sequences were excluded. Circos17 is a useful tool for comparing genome sequences based on homogeneous coordinates. In our study, a custom script was used to convert the coordinate data obtained through nucmer into a readable format in Circos. The results of chromosome comparison between two genomes were diagrammed using Circos. For visualizing detailed structural variation, GenomeRibbon18 was used to assess the coordinate data obtained through nucmer. To confirm the chromosomal stability of the Hi-C assembly, 24 chromosomes of the South Georgia icefish (P. georgianus)19 and medaka (O. latipes)20 genomes were compared with 24 chromosomes of the Hi-C assembly to assess their similarity. Each chromosome of the blackfin icefish was exclusively linked to each chromosome of the South Georgia icefish and medaka, thereby reconfirming the chromosomal stability of the scaffolds from the Hi-C assembly and verifying the integrity of the analysis (Fig. 2a,b). Antarctic fishes, including icefish species, diverged from the stickleback lineage approximately 77 million years ago7. For comparison with the chromosomes of the blackfin icefish, 21 stickleback chromosomes were aligned with the chromosome-level assembly. The results revealed that three chromosomes of the stickleback (G. aculeatus)21 were split into six chromosomes of the blackfin icefish (Fig. 2c). Moreover, 22 chromosomes of the pufferfish (T. rubripes)22 were compared with 24 chromosomes of the blackfin icefish. Pufferfish diverged from the Antarctic fish and stickleback lineages approximately 122 million years ago. Four chromosomes of the blackfin icefish (CAv2_00041, CAv2_00320, CAv2_00011, and CAv2_00012) were found to align with two chromosomes (chromosome 1:NC_042285.1 and chromosome 8: NC_042292.1) of pufferfish (Fig. 2d).
Chromosomal comparison with the blackfin icefish. P. georgianus (a) and O. latipes (b) which have the same number of chromosomes (2n = 48) were compared with the blackfin icefish. Chromosomal comparison of the blackfin icefish with G. aculeatus (c, 2n = 42) and T. rubripes (d, 2n = 44) which have less than the number of chromosomes.
Repeat analysis
A de novo repeat library was constructed using RepeatModeler v.1.0.323, including RECON and RepeatScout v.1.0.524, with default parameters. Moreover, Tandem Repeats Finder25 was used to predict consensus sequences and classification data for each repeat. All repeats collected by RepeatModeler were searched against the UniProt/SwissProt database26; transposons were excluded. To identify highly accurate long terminal repeat retrotransposons (LTR-RTs), an LTR library was constructed using LTR_retriever v.2.9.027 with combined raw LTR data from LTRharvest28 with parameters ‘-minlenltr 100 -maxlenltr 7000 -mintsd 4 -maxtsd 6 -motif TGCA -motifmis 1 -similar 85 -vic 10 -seed 20’ and LTR_FINDER29 with parameters ‘-harvest_out -size 1000000 -time 300’. Repetitive elements were identified using RepeatMasker v.4.0.9 with a de novo repeat library with parameters ‘-no_is -norna’. Various TE subfamilies were detected in the genome, accounting for 52.88% of the analyzed genome. Their distribution was as follows: DNA transposons, 15.74%; long interspersed nuclear elements, 7.73%; short interspersed nuclear elements, 0.43%; LTRs, 16.92%; and unknown elements, 9.72% (Table 3).
Gene prediction and annotation
Genome prediction was performed using EVidenceModeler (EVM) v.1.1.110, which integrates the results of multiple gene predictions. Repeat masked genomes were used for ab initio gene prediction using GeneMark-ES v.4.6830 and Augustus v.3.4.031. Then, the hints for protein and ab initio predictions were extracted with massive protein sequences from Actinopterygii, a clade of bony fishes, in the UniProt/SwissProt protein database32 using ProtHint v.2.6.08. The hints were used to perform protein predictions using GeneMark-EP + v.4.688 and ab initio predictions using Augustus. To obtain transcriptome-level evidence, the PASA pipeline v.2.3.39 with Iso-Seq data was used. EVM was used to integrate the ab initio, transcriptome, and protein prediction results to obtain the final gene prediction with weight parameters ‘ABINITIO_PREDICTION = 1, PROTEIN = 50, TRANSCRIPT = 50’. Finally, to predict changes in exons by the addition of untranslated regions (UTRs), the PASA pipeline with Iso-Seq data was used again. Genome Annotation Generator v.2.0.133 was used for adding start/stop codon data and generating a well-formed gff file. Other noncoding RNAs were identified using v.0.9. Putative tRNA genes were identified using tRNAscan-SE v.2.0.534. The predicted genes were annotated by aligning them to the NCBI non-redundant protein (nr) database35 using NCBI BLAST v.2.9.036 with a maximum e-value of 1e-5. To obtain protein domain information, InterProScan v.5.44.7937 was used with a protein sequence translated from a transcript. Moreover, Trinotate38 was used for the comprehensive annotation of transcriptome sequences, and TransDecoder v.5.5 with eggNOG (evolutionary genealogy of genes: Non-supervised Orthologous Groups) and KEGG (Kyoto Encyclopedia of Genes and Genomes) were used for decoded peptide sequences. Protein signal peptide prediction was performed using SignalP v.5.039, and transmembrane domain prediction was performed using TMHMM v2.040. Gene Ontology (GO) terms26 were assigned to the genes using the BLAST2GO pipeline v.4.041. A total of 38,024 genes and 39,889 coding sequences (CDSs) were analyzed in the C. aceratus genome. The average length of CDSs was 1,248 bp, and the average number of exons per gene was 7.9 (Table 4). Consequently, a total of 39,889 CDSs were annotated from a minimum of 17.51% to a maximum of 90.31% in seven databases for functional annotation. In one or more databases, 79.03% of CDSs were annotated (Table 5). To confirm the gene prediction results, BUSCO was used in transcriptome mode with CDSs. The percentage of complete BUSCOs was 80.7%, while that of missing was 13.4% (Table 6).
Annotation of AFGP genes
The regions containing AFGP and trypsinogen genes were extracted from the whole-genome sequence using NCBI BLAST v.2.9.036 against transcript and protein sequences of the Antarctic toothfish42. AFGP genes were predicted using Exonerate v.2.4 with the following specific parameters:--model protein2genome--minintron 20--maxintron 10000--score 250--percent 60 from the extracted region sequence. The final AFGP gene set was identified based on identity, similarity, and alignment length and was integrated into the final gene prediction data. The sequence encoding AFGP, which is similar to the long repetition of simple sequences, is very repetitive and is not assembled in the short sequence of the next-generation sequence despite their high throughput sequences. We identified that genes encoding AFGP were tandemly duplicated in the Cav2_00055 scaffold from 34,915,108 bp to 35,620,009 bp. The AFGP–trypsinogen locus was located between genes encoding mitochondrial 39 S ribosomal protein L17 (mrpl17) and E3 ubiquitin-protein ligase CBL-C isoform X1 (cbl), as reported in a previous study. However, in this study, 10 copies of trypsinogen genes, nine copies of AFGP genes, two copies of AFGP/trypsinogen-like protease chimeric genes, and a trypsinogen-like protease gene were predicted at the exon/CDS level (Fig. 3). AFGP genes evolved from trypsinogen genes in Antarctic fishes43. The prediction of gene features of AFGP genes is too difficult by the normal automated prediction method because the AFGP gene sequence has a high incidence of tandem repeats. We developed a customized process to predict complete AFGP gene features and analyzed exons and CDSs of AFGP and trypsinogen genes. Our results were consistent with previous results, except in the case of one AFGP gene. Moreover, we obtained tandemly duplicated AFGP gene sequences. Using our developed method, further analysis of the AFGP genes of other Antarctic fishes can be performed.
Technical Validation
We assessed the completeness of genome assembly using Benchmarking Universal Single-Copy Orthologs (BUSCO)45 v.5.4.4 with the Actinopterygii lineage dataset with default parameters. A total of 3,375 (92.7%) BUSCOs were identified as complete. Of these, 3,241 (89.0%) were single-copy and 134 (3.7%) were duplicated. The numbers of partially matched and missing were 48 (1.3%) and 217 (6.0%), respectively (Table 7). The k-mer completeness and quality value (QV) were evaluated by Merqury v1.346. Merqury analysis were QV of 29.96 and completeness of 88.29 (Table 8). On comparing the Hi-C scaffolds and linkage groups, high concordance was noted; however, some inconsistencies remained. In particular, mis-scaffolding was noted between LG14 and LG17. Assessment of the Hi-C scaffold confirmed that the 3.26M-sized sequence located in LG14 (LG14: 6,694,531–9,957,046) was transferred to the middle of LG7 (CaV2_00044: 12,160,643–15,423,158). Moreover, the CaV2_00044 scaffold, which was consistent with LG7, was completely scaffolded on the Hi-C contact map (Fig. 1b). These results confirmed that the mis-scaffold on the linkage group was corrected through Hi-C analysis. Moreover, the Hi-C scaffold was verified with the contact map. Many linkage group-based genome assembly results have been improved or finalized for several years through Hi-C analysis47,48.
Code availability
The bioinformatics analysis software used in this study was analyzed using the standard parameters provided by the software developers. If manually adjusted parameters were used, the software version and method used are described in the Methods.
Change history
08 January 2024
A Correction to this paper has been published: https://doi.org/10.1038/s41597-023-02861-1
References
Policansky, D. (American Association for the Advancement of Science, 1994).
Eastman, J. T. Antarctic fish biology: evolution in a unique environment. (Academic Press, 2013).
Reid, W. D., Clarke, S., Collins, M. A. & Belchier, M. Distribution and ecology of Chaenocephalus aceratus (Channichthyidae) around South Georgia and Shag Rocks (Southern Ocean). Polar Biology 30, 1523–1533 (2007).
Le François, N. R. et al. Characterization and husbandry of wild broodstock of the blackfin icefish Chaenocephalus aceratus (Lönnberg 1906) from the Palmer Archipelago (Southern Ocean) for breeding purposes. Polar Biology 40, 2499–2516 (2017).
Holeton, G. F. Oxygen uptake and circulation by a hemoglobinless Antarctic fish (Chaenocephalus aceratus lonnberg) compared with three red-blooded Antartic fish. Comparative biochemistry and physiology 34, 457–471 (1970).
Morescalchi, A. et al. A multiple sex-chromosome system in Antarctic ice-fishes. Polar Biology 11, 655–661 (1992).
Kim, B. M. et al. Antarctic blackfin icefish genome reveals adaptations to extreme environments. Nat Ecol Evol 3, 469–478, https://doi.org/10.1038/s41559-019-0812-7 (2019).
Brůna, T., Lomsadze, A. & Borodovsky, M. GeneMark-EP+: eukaryotic gene prediction with self-training in the space of genes and proteins. NAR genomics and bioinformatics 2, lqaa026 (2020).
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic acids research 31, 5654–5666 (2003).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome biology 9, 1–22 (2008).
NCBI Sequence Read Archive https://identifiers.org/insdc.sra:SRX20493982 (2023).
Putnam, N. H. et al. Chromosome-scale shotgun assembly using an in vitro method for long-range linkage. Genome Res 26, 342–350, https://doi.org/10.1101/gr.193474.115 (2016).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell systems 3, 99–101 (2016).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell systems 3, 95–98 (2016).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv preprint arXiv:1303.3997 (2013).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome biology 5, R12 (2004).
Krzywinski, M. I. et al. Circos: An information aesthetic for comparative genomics. Genome Research https://doi.org/10.1101/gr.092759.109 (2009).
Nattestad, M., Chin, C.-S. & Schatz, M. C. Ribbon: visualizing complex genome alignments and structural variation. BioRxiv, 082123 (2016).
NCBI Assembly https://identifiers.org/insdc.gca:GCA_902827115.2 (2020).
NCBI Assembly https://identifiers.org/insdc.gca:GCA_002234715.1 (2017).
Ensembl Database https://ensembl.org/Gasterosteus_aculeatus (2006).
NCBI Assembly https://identifiers.org/insdc.gca:GCA_901000725.3 (2019).
Bao, Z. & Eddy, S. R. Automated de novo identification of repeat sequence families in sequenced genomes. Genome Res 12, 1269–1276, https://doi.org/10.1101/gr.88502 (2002).
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21(Suppl 1), i351–358, https://doi.org/10.1093/bioinformatics/bti1018 (2005).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res 27, 573–580, https://doi.org/10.1093/nar/27.2.573 (1999).
Dimmer, E. C. et al. The UniProt-GO annotation database in 2011. Nucleic acids research 40, D565–D570 (2012).
Ou, S. & Jiang, N. LTR_retriever: A Highly Accurate and Sensitive Program for Identification of Long Terminal Repeat Retrotransposons. Plant Physiology 176, 1410–1422, https://doi.org/10.1104/pp.17.01310 (2018).
Ellinghaus, D., Kurtz, S. & Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC bioinformatics 9, 18 (2008).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic acids research 35, W265–W268 (2007).
Lomsadze, A., Ter-Hovhannisyan, V., Chernoff, Y. O. & Borodovsky, M. Gene identification in novel eukaryotic genomes by self-training algorithm. Nucleic acids research 33, 6494–6506 (2005).
Stanke, M., Schoffmann, O., Morgenstern, B. & Waack, S. Gene prediction in eukaryotes with a generalized hidden Markov model that uses hints from external sources. Bmc Bioinformatics 7, 62 (2006).
Consortium, U. UniProt: a worldwide hub of protein knowledge. Nucleic acids research 47, D506–D515 (2019).
Geib, S. M. et al. Genome Annotation Generator: a simple tool for generating and correcting WGS annotation tables for NCBI submission. Gigascience 7, giy018 (2018).
Chan, P. P. & Lowe, T. M. tRNAscan-SE: Searching for tRNA Genes in Genomic Sequences. Methods Mol Biol 1962, 1–14, https://doi.org/10.1007/978-1-4939-9173-0_1 (2019).
Marchler-Bauer, A. et al. CDD: a Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Research 39, D225–D229, https://doi.org/10.1093/nar/gkq1189 (2010).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. Journal of molecular biology 215, 403–410 (1990).
Jones, P. et al. InterProScan 5: genome-scale protein function classification. Bioinformatics 30, 1236–1240, https://doi.org/10.1093/bioinformatics/btu031 (2014).
Bryant, D. M. et al. A tissue-mapped axolotl de novo transcriptome enables identification of limb regeneration factors. Cell reports 18, 762–776 (2017).
Armenteros, J. J. A. et al. SignalP 5.0 improves signal peptide predictions using deep neural networks. Nature biotechnology 37, 420–423 (2019).
Möller, S., Croning, M. D. & Apweiler, R. Evaluation of methods for the prediction of membrane spanning regions. Bioinformatics 17, 646–653 (2001).
Conesa, A. et al. Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676 (2005).
Nicodemus-Johnson, J., Silic, S., Ghigliotti, L., Pisano, E. & Cheng, C.-H. C. Assembly of the antifreeze glycoprotein/trypsinogen-like protease genomic locus in the Antarctic toothfish Dissostichus mawsoni (Norman). Genomics 98, 194–201 (2011).
Chen, L., DeVries, A. L. & Cheng, C.-H. C. Evolution of antifreeze glycoprotein gene from a trypsinogen gene in Antarctic notothenioid fish. Proceedings of the National Academy of Sciences 94, 3811–3816 (1997).
NCBI Assembly https://identifiers.org/insdc.gca:GCA_023974075.1 (2022).
Simao, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212, https://doi.org/10.1093/bioinformatics/btv351 (2015).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome biology 21, 1–27 (2020).
Peichel, C. L., Sullivan, S. T., Liachko, I. & White, M. A. Improvement of the threespine stickleback genome using a Hi-C-based proximity-guided assembly. Journal of Heredity 108, 693–700 (2017).
Lee, S. J. et al. Chromosomal assembly of the Antarctic toothfish (Dissostichus mawsoni) genome using third-generation DNA sequencing and Hi-C technology. Zoological research 42, 124 (2021).
Acknowledgements
This work was supported by a Korea Institute of Marine Science & Technology Promotion (KIMST) grant funded by the Ministry of Oceans and Fisheries (KIMST 20220547), the National Institute of Fisheries Science (NIFS; R2023003), and by a grant from the Korea University.
Author information
Authors and Affiliations
Contributions
H.P. and J.-H.K. conceived the study. S.J.L., E.K.C., E.J. and M.C. performed genome sequencing and assembly. S.J.L. and H.P. wrote the manuscript. All the authors contributed to writing and editing the manuscript and preparing the figures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, S.J., Kim, J., Choi, E.K. et al. A chromosome-level reference genome of the Antarctic blackfin icefish Chaenocephalus aceratus. Sci Data 10, 657 (2023). https://doi.org/10.1038/s41597-023-02561-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-023-02561-w
- Springer Nature Limited