Abstract
Hibiscus syriacus L. is a renowned ornamental plant. We constructed 95 chloroplast genomes of H. syriacus L. cultivars using a short-read sequencing platform (Illumina) and a long-read sequencing platform (Oxford Nanopore Technology). The following genome assembly, we delineate quadripartite structures encompassing large single-copy, small single-copy, and inverted repeat (IRa and IRb) regions, from 160,231 bp to 161,041 bp. Our comprehensive analyses confirmed the presence of 79 protein-coding genes, 30 tRNA genes, and 4 rRNA genes in the pan-chloroplast genome, consistent with prior research on the H. syriacus chloroplast genome. Subsequent pangenome analysis unveiled widespread genome sequence conservation alongside unique cultivar-specific variant patterns consisting of 193 single-nucleotide polymorphisms and 61 insertions or deletions. The region containing intra-species variant patterns, as identified in this study, has the potential to develop accession-specific molecular markers, enhancing precision in cultivar classification. These findings are anticipated to drive advancements in breeding strategies, augment biodiversity, and unlock the agricultural potential inherent in H. syriacus.
Similar content being viewed by others
Background & Summary
H. syriacus, commonly known as rose of Sharon, is a fast-growing deciduous shrub belonging to the Malvaceae family and is renowned for its diverse applications, including culinary, ornamental, and medicinal uses1. Its wide range of flower colors makes it an attractive choice for decorative landscaping2,3. In North American countries, it has gained immense popularity as a garden tree due to its versatile properties4. However, breeding H. syriacus presents significant challenges due to its self-incompatibility, resulting in most landraces being natural hybrids5. Consequently, there have been limited reports of breeding trials aimed at developing polyploidy plants4,6. In Korea, breeding advancements have been achieved through methods such as the propagation of naturally occurring mutants, inter-generic crossings, and the induction of mutations using gamma-ray irradiation6,7,8,9. The complexities of breeding H. syriacus highlights the importance of elucidating the phylogenetic relationships among its cultivars to establish a breeding system capable of generating F1 hybrids.
Given the challenges in H. syriacus breeding, the utilization of chloroplast genomes represents a strategic approach due to their unique features. These organelles are typically maternally inherited, except in some gymnosperms where inheritance is paternally directed. Chloroplast genomes contain non-recombinant sequences and are usually inherited in a uniparental manner, allowing for lineage tracing through the maternal line and minimizing uncertainties associated with biparentally inherited nuclear genomes10,11,12,13. Furthermore, the high conservation of the chloroplast genome, including gene repertories and structures, enables comparative analyses that offer clear insights into the evolutionary trajectories and phylogenetic relationships among cultivars14,15,16. Previous studies on Atractylodes species and Panax ginseng demonstrated that even with low divergence, unique polymorphic chloroplast-derived markers could be developed to distinguish inter- and intra-species differences, respectively11,17,18,19,20,21,22. This highlights the potential applications of chloroplast genomes in the development of highly species-specific molecular markers, even at the intra-species level, thereby overcoming challenges posed by minimal genetic divergence. Nevertheless, the majority of studies on Hibiscus chloroplast genomes have predominantly focused on the taxonomic level of genus, leaving in-depth intra-species studies relatively unexplored10,23,24,25. Given the breeding challenges of H. syriacus outlined earlier, comparative studies at the intra-species level are not only crucial but indispensable. Developing more molecular markers at the intra-species level is essential to gain unparalleled insights into the evolutionary trajectory and contribute to the precise taxonomic classification of H. syriacus26,27,28.
In this study, we generated 94 H. syriacus chloroplast genomes using a short-read sequencing platform (Illumina) and 1 genome using a long-read sequencing platform (Oxford Nanopore Technology). Subsequent pangenome analysis of these 95 H. syriacus chloroplast genomes revealed a high degree of conservation in the majority of genome sequences, while also identifies unique cultivar-specific variant patterns. A total of 193 single-nucleotide polymorphisms (SNPs) and 61 insertions or deletions (Indels) were identified, highlighting their potential applications as intra-species molecular markers29. The development of molecular markers utilizing these regions will play a pivotal role in achieving precise classification among H. syriacus cultivars and establishing refined breeding strategies. Moreover, these results will offer essential insights for species conservation, biodiversity enhancement, and the exploration of the agricultural and ornamental potentials of H. syriacus.
Methods
Plant materials and sequencing
H. syriacus cv. Gangneung was used for long-read-based chloroplast genome assembly30. A core collection of H. syriacus from the National Institute of Forest Science was utilized for short-read-based chloroplast genome assembly. Genomic DNA was extracted from fresh leaf tissues of H. syriacus plants using the standard cetyltrimethylammonium bromide method31.
The quantity and quality of genomic DNA were assessed using a Nanodrop spectrophotometer with a quality cut-off at an OD260/280 ratio of 1.8–2.0 and a Qubit dsDNA HS Assay Kit (Thermo Fisher Scientific, Massachusetts, USA). Following quality assessment, the DNA was used to generate libraries with an average insert size of 550 bp. Paired-end sequencing was performed to obtain 150-bp sequences at both ends using an Illumina NovaSeq. 6000 platform (Illumina Inc., San Diego, CA, USA).
Genome assembly and annotation
For long-read assembly, the generated reads30 were aligned to a reference chloroplast sequence obtained from prior research32, using minimap2 (v2.22) with default parameters33. Reads with a mapping coverage exceeding 80 were extracted using Seqtk (https://github.com/lh3/seqtk) v1.3. These extracted reads were then assembled into a pseudo-molecule using Flye (v.2.9)34 and subsequently polished using NextPolish (v1.4)35 to correct base errors arising from noisy long reads.
For short-read assembly, Trimmomatic (v0.39)36 was used to trim adapters and eliminate low-quality sequences from the raw reads to enhance read quality. The trimmed reads were then aligned to the reference chloroplast genomes obtained from prior studies37,38,39,40,41,42,43,44, using the Burrows–Wheeler alignment (v0.7.17) tool45 (Table 1). The mapped reads were assembled using NOVOPlasty (v4.3.1)46, which employed a 39 k-mer and default RUBP sequences as seeds for chloroplast assembly47,48. The contigs generated by NOVOPlasty were ordered and merged into a single pseudo-molecule according to the reference chloroplast genome sequence.
Genome annotation was performed using the GeSeq platform, which provides rapid and accurate annotation of organellar genomes49. We employed BLAT50, Chloë (v0.1.0), and HMMER51 to annotate coding sequences and rRNA, and ARAGORN (v1.2.38)52 and tRNAscan-SE (v2.0.7)53 to annotate tRNA. Annotation accuracy was validated against H. syriacus var. Baekdansim30, and any discrepancies were manually curated. The circular map representation of the chloroplast genome was generated using OGDRAW (v1.3.1)54 (Fig. 1).
Circular map of the chloroplast genome in H. syriacus var. Gangneung. The center of the plot displays the cultivar name and genome length. The inner grey circle represents the GC content proportion in each region, with the line representing 50%. Genes located outside the outer circle are transcribed counterclockwise, and those inside the circle are transcribed clockwise. Genes with different functional annotations are differentiated by color.
Chloroplast genome alignment and pan-chloroplast genome-graph construction
To validate the genome assembly, we employed the chloroplast genome of H. syriacus var. Gangneung, constructed using long-read sequencing, as a reference for multiple sequence alignment. Sequence alignment was performed using MAFFT55 with default parameters. Subsequently, pairwise alignments of the chloroplast genomes were generated using MUMmer456.
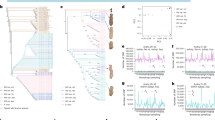
To construct a pan-chloroplast genome-graph encompassing 95 H. syriacus genomes, we utilized the Minigraph-Cactus Pangenome Pipeline (v2.6.8)57. The integration process involved the iterative addition of the remaining 94 genomes with the reference chloroplast genome. Precise base-level alignments were achieved with the Cactus-pangenome tool using the parameters “--giraffe --fa --bz --viz.” From this comprehensive graph, we employed the Cactus-graphmap (v2.6.8) tool to map the graph utilizing the default parameters. We identified a total of 193 SNPs and 61 Indels across the entire genomes, observations that offer significant potential for the future development of intra-species molecular markers29. Overall, H. syriacus cultivars exhibit similarity across all genomic regions. However, for H. syriacus var. Russian Violet, a notable divergence in similarity was observed in the regions spanning 59,000 bp to 62,000 bp (Fig. 2).
Pan-chloroplast genome-graph for 95 H. syriacus cultivars. (a) The pan-chloroplast genome-graph represents all 95 H. syriacus cultivars with the total chloroplast genome scale. (b) An enlarged view of the pan-chloroplast genome graph highlighting a region of the largest variation identified in H. syriacus var. Russian violet, indicated by red bars. (c) Multiple sequence alignment for the largest variation site among the 95 H. syriacus varieties.
Comparative genomic analysis in 95 H. syriacus chloroplast genomes
Structural similarity and gene distribution among the 95 chloroplast genomes were analyzed using mVISTA software in LAGAN mode with the default settings, with H. syriacus var. Baekdansim used as the reference58,59,60,61 (Fig. 3). This observation was consistent with the results from the pan-chloroplast genome analysis, where H. syriacus var. Russian Violet exhibits a significant deletion in specific regions.
The 95 H. syriacus accessions mVISTA map, with the Gangneung chloroplast genome as the reference. The vertical scale represents the percentage of identity, ranging from 50% to 100%. The horizontal axis corresponds to the base sequence region. Red indicates non-coding sequences(CNS), blue indicates the exons of protein-coding genes and light green indicates untranslated regions(UTR) including tRNA or rRNA.
Hypervariable regions within the chloroplast genome of H. syriacus were identified using DnaSP version 6 software62. A total of 95 H. syriacus chloroplast genomes were aligned using MAFFT55 with default parameters. Nucleotide diversity was calculated through sliding window analysis, with the window size set at 600 bp with a step size of 100 bp22 (Fig. 4). The inverted repeat regions tend to be more conservative than the single copy regions. The highest nucleotide diversity was identified in the trnS-psbZ region. This region has the potential for use as a DNA barcode to facilitate distinction among the H. syriacus cultivars.
Nucleotide diversity in 95 H. syriacus chloroplast genomes. Sliding window analysis was performed with a window length of 600 bp and a step size of 100 bp. The x-axis represents nucleotide position, while the y-axis represents nucleotide diversity (Pi). Genes within the most hypervariable regions are highlighted in red.
Data Records
A total of 94 raw reads obtained through Illumina sequencing have been deposited in the NCBI Sequence Read Archive under the accession number SRP46454163. The assembled chloroplast genome sequences, accompanied by their corresponding gene annotations for the 94 cultivars have been submitted to NCBI GenBank64,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114,115,116,117,118,119,120,121,122,123,124,125,126,127,128,129,130,131,132,133,134,135,136,137,138,139,140,141,142,143,144,145,146,147,148,149,150,151,152,153,154,155,156,157 and are detailed in Table S1. Additionally, H. syriacus var. Gangneung has been deposited in the NCBI GenBank with the accession number OR619829158.
Technical Validation
Evaluation of chloroplast genome assembly
To evaluate the completeness of the chloroplast genome assembly, chloroplast reads were aligned to the chloroplast genome as described in the “Genome assembly and annotation” section. The lengths of the 95 assembled pseudo-molecules ranged from 160,231 bp to 161,041 bp, which is consistent with the observed chloroplast genome length in other members of the Malvaceae family23,24,25,26,27. Synteny analyses were conducted using MUMmer159 with the previously reported chloroplast genome of H. syriacus var. Baekdansim as the reference30. The dot plot revealed that the assembled genomes align cohesively with no major rearrangements observed (Fig. 5). Instead, the plot displayed inversions, represented by a blue line, corresponding to the chloroplast-specific inverted region.
Pairwise comparative analysis of chloroplast genomes in various H. syriacus cultivars with H. syriacus var. Baekdansim using MUMmer plots. (a) Comparison of chloroplast genomes constructed using ONT and PacBio long-read sequencing platforms. (b) Comparison of chloroplast genomes constructed using Illumina short-read and PacBio long-read sequencing platforms. The red lines represent collinear sequences and the blue lines represent inverted sequences.
Evaluation of gene annotation
The accuracy of the gene annotations was meticulously evaluated by comparing them to the H. syriacus var. Baekdansim61 chloroplast genome. Any discrepancies identified were refined through manual curation. In total, 113 distinct genes were identified, including 79 protein-coding genes, 30 tRNA genes, and 4 rRNA genes (Table 2).
The gene repertoires were consistent across all 95 cultivars, with the only observed differences being related to specific gene loci details. Our results indicate that the gene repertoire was congruent with annotations commonly observed within the Malvaceae family23,160,161, with minor variations detected in the pafI (ycf3), pafII (ycf4), and pbf1 (psbN) genes162,163.
Code availability
All software used for data processing was implemented following the manual provided by the bioinformatic software cited in the method section. When specific parameters for the software were not detailed, the default settings were utilized.
References
Lee, S. J. et al. An antioxidant lignan and other constituents from the root bark of Hibiscus syriacus. Planta Med. 65, 658–660 (1999).
Akpan, G. A. Cytogenetic characteristics and the breeding system in six Hibiscus species. Theor. Appl. Genet. 100, 315–318 (2000).
Zhang, P. et al. Identification and quantitative analysis of anthocyanins composition and their stability from different strains of Hibiscus syriacus L. flowers. Ind. Crops Prod. 177, 114457 (2022).
Contreras, R. N., Friddle, M. & Lattier, J. D. Relative fertility and ploidy levels of selected rose of sharon cultivars. SNA Research Conference 58, 232–236 (2013).
Song, H.-S., Kim, J.-K., Lee, K.-U. & Lim, Y.-T. Studies on mutation breeding of Hibiscus syriacus. Report No. KAERI/RR-1669/96 (Korea Atomic Energy Research Institute, 1997).
Van Laere, K., Van Huylenbroeck, J. & Van Bockstaele, E. Breeding strategies to increase genetic variability within Hibiscus syriacus. Acta Hortic. 714, 75–82 (2006).
Park, H.-S., Cho, Y.-J., Chung, H.-G., Jang, Y.-S. & Chung, D.-J. Variation in flower color among hybrids of jeoktanshim Hibiscus syriacus L. Korean Journal of Plant Resources 8, 250–257 (2005).
Park, Y. M., Lee, S.-W., Kwon, S.-H. & Kwon, H. Y. ‘Haeoreum’, A new Hibiscus syriacus cultivar for street trees. HortScience 55, 1159–1161 (2020).
Kim, S. H., Sim, S. J., Nam, J. I. & Kwon, H. Y. Korean Hibiscus Variety Catalog, (The National Institute of Forest Science (NIFoS) Press, 2014).
Abdullah et al. Chloroplast genome of Hibiscus rosa-sinensis (Malvaceae): comparative analyses and identification of mutational hotspots. Genomics 112, 581–591 (2020).
Wang, Y. et al. Chloroplast genome variation and phylogenetic relationships of Atractylodes species. BMC Genomics 22, 103 (2021).
Dong, W., Xu, C., Cheng, T., Lin, K. & Zhou, S. Sequencing angiosperm plastid genomes made easy: a complete set of universal primers and a case study on the phylogeny of saxifragales. Genome Biol. Evol. 5, 989–997 (2013).
Park, H. S. et al. Inheritance of chloroplast and mitochondrial genomes in cucumber revealed by four reciprocal F(1) hybrid combinations. Sci. Rep. 11, 2506 (2021).
Shinozaki, K. et al. The complete nucleotide sequence of the tobacco chloroplast genome: its gene organization and expression. The EMBO Journal 5, 2043–2049 (1986).
Zhang, F., Li, W., Gao, C. W., Zhang, D. & Gao, L. Z. Deciphering tea tree chloroplast and mitochondrial genomes of Camellia sinensis var. assamica. Sci. Data 6, 209 (2019).
Huang, Y. et al. Comprehensive analysis of complete chloroplast genome and phylogenetic aspects of ten Ficus species. BMC Plant Biol. 22, 253 (2022).
Hultquist, S. J. et al. and nuclear DNA content variations among cultivars of switchgrass. Crop Sci. 36, 1049–1052 (1996).
Li, R. et al. Genetic diversity in Chinese sorghum landraces revealed by chloroplast simple sequence repeats. Genet. Resour. and Crop Evol. 57, 1–15 (2009).
Cho, K. S. et al. Determination of cytoplasmic male sterile factors in onion plants (Allium cepa L.) using PCR-RFLP and SNP markers. Mol. Cells 21, 411–417 (2006).
Nikiforova, S. V., Cavalieri, D., Velasco, R. & Goremykin, V. Phylogenetic analysis of 47 chloroplast genomes clarifies the contribution of wild species to the domesticated apple maternal line. Mol. Biol. Evol. 30, 1751–1760 (2013).
Kim, K. et al. Comprehensive survey of genetic diversity in chloroplast genomes and 45S nrDNAs within Panax ginseng species. PLoS One 10, e0117159 (2015).
Joh, H. J. et al. Authentication of Golden-Berry P. ginseng cultivar ‘Gumpoong’ from a landrace ‘Hwangsook’ based on pooling method using chloroplast-derived markers. Plant Breed. and Biotechnol. 5, 16–24 (2017).
Cheng, Y., Zhang, L., Qi, J. & Zhang, L. Complete chloroplast genome sequence of Hibiscus cannabinus and comparative analysis of the Malvaceae family. Front. Genet. 11, 227 (2020).
Xu, X. R., Zhou, S. D. & Shi, X. Q. The complete chloroplast genome of Hibiscus Taiwanensis (Malvaceae). Mitochondrial DNA B: Resour. 4, 2533–2534 (2019).
Kwon, S.-H., Park, Y., Jang, Y. L. & Kwon, H.-Y. The complete chloroplast genome sequence of Hibiscus sabdariffa (Malvaceae). Korean Journal of Plant Taxonomy 52, 123–126 (2022).
Koo, H., Shin, A. Y., Hong, S. & Kim, Y. M. The complete chloroplast genome of Hibiscus syriacus using long-read sequencing: comparative analysis to examine the evolution of the tribe Hibisceae. Front. Plant Sci. 14, 1111968 (2023).
Kim, Y. S. et al. The complete chloroplast genome sequence of Hibiscus syriacus L. ‘Mamonde’ (Malvaceae). Mitochondrial DNA Part B 4, 558–559 (2019).
Wang, R., Park, S. Y., Park, S. W., Puja, A. M. & Kim, Y.-J. Development of a molecular marker based on chloroplast gene for specific identification of Korean Hibiscus (Hibiscus syriacus ‘Simbaek’). Appl. Biol. Chem. 64, 96 (2021).
Kim, Y.-M. SNPs and Indels in pan-chloroplast genomes of 95 Hibiscus syriacus cultivars. figshare https://doi.org/10.6084/m9.figshare.24320947.v1 (2023).
Koo, H. et al. Two long read-based genome assembly and annotation of polyploidy woody plants, Hibiscus syriacus L. using PacBio and Nanopore platforms. Sci. Data 10, 713 (2023).
Allen, G. C., Flores-Vergara, M. A., Krasynanski, S., Kumar, S. & Thompson, W. F. A modified protocol for rapid DNA isolation from plant tissues using cetyltrimethylammonium bromide. Nat. Protoc. 1, 2320–2325 (2006).
Kim, J.-H., Lee, H., Kwon, H.-Y. & Kim, S.-H. Hibiscus syriacus chloroplast, complete genome. GenBank https://identifiers.org/ncbi/insdc:KP688069.1 (2015).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Kolmogorov, M., Yuan, J., Lin, Y. & Pevzner, P. A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 37, 540–546 (2019).
Hu, J., Fan, J., Sun, Z. & Liu, S. NextPolish: a fast and efficient genome polishing tool for long-read assembly. Bioinformatics 36, 2253–2255 (2020).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Xu, Q. et al. Gossypium arboreum chloroplast, complete genome. GenBank https://www.ncbi.nlm.nih.gov/nuccore/NC_016712.1/ (2023).
Lee, S. B. et al. Gossypium hirsutum chloroplast, complete genome. GenBank https://www.ncbi.nlm.nih.gov/nuccore/NC_007944.1/ (2023).
Xu, Q. et al. Gossypium raimondii chloroplast, complete genome. GenBank https://www.ncbi.nlm.nih.gov/nuccore/NC_016668.1/ (2023).
Cheng, Y., Zhang, L., Qi, J. & Zhang, L. Hibiscus cannabinus chloroplast, complete genome. GenBank https://www.ncbi.nlm.nih.gov/nuccore/NC_045873.1 (2023).
Abdullah et al. Gossypium raimondii chloroplast, complete genome. GenBank https://www.ncbi.nlm.nih.gov/nuccore/NC_042239.1 (2023).
Kim, J.-H., Lee, H., Kwon, H.-Y. & Kim, S.-H. Hibiscus syriacus chloroplast, complete genome. GenBank https://www.ncbi.nlm.nih.gov/nuccore/NC_026909.1 (2023).
Wang, J.-H., Moore, M. J., Wang, H., Zhu, Z.-X. & Wang, H.-F. Hibiscus taiwanensis chloroplast, complete genome. GenBank https://www.ncbi.nlm.nih.gov/nuccore/NC_054167.1 (2023).
Kwon, S.-H., Park, Y. M., Jang, Y. L. & Kwon, H.-Y. Hibiscus trionum chloroplast, complete genome. GenBank https://www.ncbi.nlm.nih.gov/nuccore/NC_060636.1 (2023).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Dierckxsens, N., Mardulyn, P. & Smits, G. NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res. 45, e18 (2017).
Jin, J.-J. et al. GetOrganelle: a fast and versatile toolkit for accurate de novo assembly of organelle genomes. Genome Biol. 21, 241 (2020).
Yun, S. & Kim, H. Insight into the phylogenetic relationships and evolutionary history of pepper cultivars (Capsicum annuum L.) through comparative analyses of plastomes. Horticulturae 9, 1092 (2023).
Tillich, M. et al. GeSeq - versatile and accurate annotation of organelle genomes. Nucleic Acids Res. 45, W6–W11 (2017).
Kent, W. J. BLAT–the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Finn, R. D., Clements, J. & Eddy, S. R. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 39, W29–37 (2011).
Laslett, D. & Canback, B. ARAGORN, a program to detect tRNA genes and tmRNA genes in nucleotide sequences. Nucleic Acids Res. 32, 11–6 (2004).
Chan, P. P., Lin, B. Y., Mak, A. J. & Lowe, T. M. tRNAscan-SE 2.0: improved detection and functional classification of transfer RNA genes. Nucleic Acids Res. 49, 9077–9096 (2021).
Greiner, S., Lehwark, P. & Bock, R. OrganellarGenomeDRAW (OGDRAW) version 1.3.1: expanded toolkit for the graphical visualization of organellar genomes. Nucleic Acids Res. 47, W59–W64 (2019).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Bio. and Evol. 30, 772–780 (2013).
Marçais, G. et al. MUMmer4: a fast and versatile genome alignment system. PLOS Comput. Biol. 14, e1005944 (2018).
Hickey, G. et al. Pangenome graph construction from genome alignments with Minigraph-Cactus. Nat. Biotechnol. (2023).
Brudno, M. et al. Glocal alignment: finding rearrangements during alignment. Bioinformatics. 19, i54–i62 (2023).
Frazer, K. A., Pachter, L., Poliakov, A., Rubin, E. M. & Dubchak, I. VISTA: computational tools for comparative genomics. Nucleic Acids Res. 32, W273–W279 (2004).
Song, W. et al. Comparative analysis the complete chloroplast genomes of nine Musa species: genomic features, comparative analysis, and phylogenetic implications. Plant Sci. 13, 832884 (2022).
Koo, H. et al. Hibiscus syriacus cultivar Baekdansim chloroplast, complete genome. GenBank https://identifiers.org/ncbi/insdc:OP874596.1 (2022).
Rozas, J. et al. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. 34, 3299–3302 (2017).
NCBI Seqeunce Read Archive https://identifiers.org/ncbi/insdc.sra:SRP464541 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Adenseu plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR350231.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar An-dong plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR164399.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Arang plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397843.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Asadal plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397844.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Asanyo plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397845.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Bicolor plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397846.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Bluebird plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397847.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Bulsae plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397848.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Chilbo plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397849.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Chini plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397850.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Chossarang plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397851.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Coelestis plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397852.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Daisengionmamori plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397853.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Daitokujihanagasa plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397854.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Gaeryangdansim plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397855.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Geunhyeong plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397856.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Gojumong plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397857.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Gwangmyeong plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397858.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Gyeongbuk1 plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397859.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Gyewolhyang plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397860.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Hanbit plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397861.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Hanboram plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397862.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Haneol plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397863.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Hanmaum plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397864.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Hanseo plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397865.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Hanyang plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397866.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Hwahap plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397867.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Hwahong plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397868.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Jaok plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397869.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Kkoma plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397870.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Kyungki plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397871.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Large_White plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397872.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Mujigae plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397873.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Murasakisaiben plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397874.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Naesarang plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397875.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Nanpa plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397876.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Nunmoe plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397877.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Oknyo plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397878.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Oktokki plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397879.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Paedal plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397880.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Paeksol plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397881.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Parangsae plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397882.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Pheasant_Eye plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397883.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Pompon_Rouge plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397884.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Pulkkot plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397885.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Purpureus plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397886.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Pyeli plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397887.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Pyeonghwa plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397888.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Rubis plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397889.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Russian_Violet plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397890.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Saeasadal plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397891.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Saehan plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397892.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Salmabaek plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397893.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Samchulri plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397894.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Sanchonye plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397895.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Seondeok plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397896.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Seonnyeo plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397897.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Seorak plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397898.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Serenade plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397899.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Shintaeyang plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397900.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Shirohanagasa plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397901.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Single_Red plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397902.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Snowdrift plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397903.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Sobong plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397904.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Soltanshim plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397905.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Sonde plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397906.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Soyang plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397907.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Suchihanagasa plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397908.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Tamna plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397909.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar The_Banner plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397910.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Wonhwa plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR397911.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Baekgeunip plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625140.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Baekgiwonsu plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625141.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Baeksoryun plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625142.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar bredon_spring plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625143.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Daedeoksaback plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625144.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Daedeoksailjung plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625145.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Diana_Baekjo plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625148.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar dorothycranc plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625149.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Gakchang plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625150.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Jeogiljung plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625153.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Jeokgiwonsu plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625154.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar lenny plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625155.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Rakchanghwarip plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625157.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Sangbon plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625160.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar WR_smith plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625161.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Hwarang plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625151.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Saimdang plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625159.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Daejabae plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625146.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Deungnang plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625147.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Jabae plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625152.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Pyeongseong_ps80-1 plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625156.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar red_heart plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR625158.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Simsan plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR619828.1 (2023).
Koo, H. et al. Hibiscus syriacus cultivar Gangneung plastid, complete genome. GenBank https://identifiers.org/ncbi/insdc:OR619829.1 (2023).
Delcher, A. L., Salzberg, S. L. & Phillippy, A. M. Using MUMmer to identify similar regions in large sequence sets. Curr. Protoc. in Bioinform. 00, 10.3.1–10.3.18 (2003).
Abdullah et al. Correlations among oligonucleotide repeats, nucleotide substitutions, and insertion–deletion mutations in chloroplast genomes of plant family Malvaceae. J. of Syst. and Evol. 59, 388–402 (2020).
Yu, J., Jin, W., Wang, Y. & Yuan, Q. The complete chloroplast genome of a medicinal plant, Abutilon theophrasti Medik. (Malvaceae). Mitochondrial DNA B: Resour. 5, 3777–3778 (2020).
Wicke, S., Schneeweiss, G. M., dePamphilis, C. W., Müller, K. F. & Quandt, D. The evolution of the plastid chromosome in land plants: gene content, gene order, gene function. Plant Mol. Biol. 76, 273–297 (2011).
Zupok, A. et al. A photosynthesis operon in the chloroplast genome drives speciation in evening primroses. The Plant Cell 33, 2583–2601 (2021).
Acknowledgements
This work was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (NRF-2021R1I1A2044678) to Y.-M.K.
Author information
Authors and Affiliations
Contributions
Y.-M.K. designed and organized the project. S.G., H.K., M.J., S.H. and G.Y. conducted the data analysis. S.G., H.K. and Y.-M.K. wrote the draft of the manuscript. All authors reviewed, revised, and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Go, S., Koo, H., Jung, M. et al. Pan-chloroplast genomes for accession-specific marker development in Hibiscus syriacus. Sci Data 11, 246 (2024). https://doi.org/10.1038/s41597-024-03077-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03077-7
- Springer Nature Limited