Abstract
Organelles do not act as autonomous discrete units but rather as interconnected hubs that engage in extensive communication by forming close contacts called “membrane contact sites (MCSs)”. And many proteins have been identified as residing in MCS and playing important roles in maintaining and fulfilling specific functions within these microdomains. However, a comprehensive compilation of these MCS proteins is still lacking. Therefore, we developed MCSdb, a manually curated resource of MCS proteins and complexes from publications. MCSdb documents 7010 MCS protein entries and 263 complexes, involving 24 organelles and 44 MCSs across 11 species. Additionally, MCSdb orchestrates all data into different categories with multitudinous information for presenting MCS proteins. In summary, MCSdb provides a valuable resource for accelerating MCS functional interpretation and interorganelle communication deciphering.
Similar content being viewed by others
Background & Summary
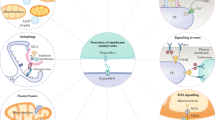
“All things are mutually woven together and therefore have an affinity for each other”—Marcus Aurelius, Meditations. Most biologists would agree with Aurelius’ statement because connectivity is observed at every level of biology, occurring between biomolecules, cells, tissues and organisms1,2,3,4,5. Therefore, it is becoming increasingly evident that organelles do not act as autonomous discrete units but rather as interconnected hubs that engage in extensive communication by forming close contacts called “membrane contact sites (MCSs)”6,7. The MCS is defined as an area of close apposition (from 10 to 80 nm) between the membranes of two organelles that are physically connected via proteinaceous tethers but do not fuse (Fig. 1)8,9. Current studies on MCSs are moving toward the central stage in cell biology9,10. Multiple MCSs have been identified between virtually all organelles in eukaryotic cells and participate in various biological processes and intracellular signaling, such as autophagy, lipid metabolism, calcium homeostasis and organelle trafficking and remodeling10,11,12,13. Moreover, aberrant loss or gain of function of MCSs can contribute to various diseases, such as cancer, metabolic diseases and neurodegenerative disorders14,15,16,17,18. In a sense, studies on spatiotemporal coordination among organelles indicate the existence of a hidden world of cellular interorganelle communication networks connected by MCS waiting to be explored19,20.
As a key component of MCSs, the proteins residing in membrane contacts play a crucial role in maintaining and fulfilling functions specific to MCSs9,21. Understanding how these protein tethers and membrane contacts coordinate organelle function will redefine our view of the cell14,22. Recently, a growing number of MCS proteins have been identified and functionally characterized. For example, three Aster proteins (Aster-A, -B, -C) can be recruited to the plasma membrane (PM)-endoplasmic reticulum (ER) and facilitate nonvesicular plasma membrane to ER cholesterol transport23. The Sel1 L-Hrd1 protein complex is involved in ER-mitochondria (MT) crosstalk and can affect mitochondrial dynamics in brown adipocytes24. The protein complex consisting of SLPD1, SLPD2 and LIPA mediates lipid droplet (LD)-PM tethering in plant cells25. Loewen et al., identified a short conserved determinant called the FFAT motif. This motif interacts with the VAP protein family, which are conserved integral membrane proteins located in the ER. These proteins play a pivotal role in the formation and function of various ER-related MCSs26,27,28. Subsequently, a series of proteins that contain the FFAT motif were recognized as MCS proteins29,30. Several studies have begun to screen MCS proteins by combining traditional biochemical approaches (subcellular fractionation and pull-down) with mass spectrometry (MS)-based proteomics31,32,33. However, the limitations (e.g., destabilized contacts, contamination by other components) of such traditional biochemical approaches may lead to a large number of false-positive proteins being detected9,34. Nonetheless, some proximity labeling approaches combined with high-throughput proteomic analysis, such as BioID, Contact-ID, and Split-TurboID, have recently been developed for global mapping of MCS proteins and are promising for MCS proteomics studies35,36,37,38,39.
Although MCSs have received increasing attention and the proteins residing in MCSs have been extensively identified in the past few years40,41,42, an appropriative database for storing, integrating and reorganizing MCS proteins is still lacking. Therefore, we developed MCSdb, a manually curated database of experimentally supported MCS proteins and complexes from publications. The current version of MCSdb documents approximately 7000 manually curated MCS protein entries and 263 complexes with experimental evidence, involving 24 organelles and 44 MCSs across 11 species. Furthermore, MCSdb grades all MCS protein entries into 3 categories according to the confidence level of experimental evidence. MCSdb also provides multitudinous information to help query and analyze MCS proteins and complexes. To our knowledge, MCSdb is the first database specifically focusing on proteins located in MCS. We believe that this database will be invaluable in accelerating MCS functional research and interorganelle communication deciphering. Dataset of the MCS proteins and complexes is free available in Figshare43.
Methods
Data collection
The MCS proteins in the database were curated manually from the literature (before Jun. 2023). First, we retrieved literature from PubMed, bioRxiv, Web of Science and Google Scholar using the following keywords: ‘membrane contact site’, ‘organelle communication’, ‘organelle interaction’, ‘mitochondria-associated membranes’, ‘protein tether’, and ‘proximity labeling’. All binary phrases consist of two organelles: ‘endoplasmic reticulum-plasma membrane’, ‘endoplasmic reticulum-Golgi’, ‘endoplasmic reticulum–peroxisome’ and ‘endoplasmic reticulum-lipid droplet’ (Fig. 2). Then, all retrieved publications were preliminarily reviewed by expert curators to filter out false-positive papers. According to several review articles9,10, the MCS is defined as an area of close apposition (from 10 to 80 nm) between two bi- or mono-layer membrane-bound organelles that are physically connected via proteinaceous tethers but do not fuse. And to be included as an MCS protein in MCSdb, there must be experimental confirmation that the protein is located at the MCS, or evidence showing that it can be recruited to the MCS, contributing to its formation or to the functions associated with the MCS. Additionally, the protein complexes located and acting in MCSs are recorded in MCSdb.
Data organization
First, we distinguished MCSs by the connected organelles of an MCS (named ER-PM, ER-MT and MT-LD, etc.), and a total of 44 MCSs were defined. Then, we divided all MCS protein entries into different categories according to the MCS of the proteins located (Fig. 2). Meanwhile, we graded all documented MCS protein entries into 3 categories: low-throughput (LT) experimental-based methods, proximity labeling (PL)-based methods and mass spectrometric (MS)-based methods. LT-based methods represent the proteins identified and functionally characterized by low-throughput experimental methods, and there are two additional inclusion criteria for LT-based proteins: (1) Proteins cannot be solely identified through high-throughput experiments; (2) The number of MCS proteins identified by the literature source for a given protein is less than 10 (inspiration from the protein-protein interaction (PPI) databases’ criteria: MINT44, mentha45, InWeb_InBioMap46). PL methods represent the proteins identified by combining PL approaches with high-throughput proteomics. MS methods represent the proteins identified by combining traditional biochemical approaches with MS techniques. To enhance user ability to evaluate the reliability of MS-based data, we introduce a scoring system anchored in protein subcellular localization and protein-protein interaction (PPI) networks. We sourced interaction information for MS-based proteins from the String database and subcellular localization data from the Uniprot database47 to ascertain if MS-based proteins and their interacting partners are situated within the MCS organelle. This system stratifies MS-based data into five confidence levels (L1 to L5), with detailed rules outlined on the “Help” page.
Data annotation
To unify the proteins from multiple publications in authoritative reference databases, all MCS proteins were mapped to the NCBI gene database (Entrez ID)48 and UniProt (UniProt ID)47. Five compounds involved in MCS complexes were mapped to the PubChem database (PubChem CID)49. Information of subcellular localization, cell line/tissue and descriptions of MCS proteins was manually curated from the literature (Fig. 2). Human and mouse gene expression data across different tissues were collected from Human Protein Atlas (HPA) (62 human tissues)50 and the TISSUES 2.0 database (39 mouse tissues)51, respectively. Protein sequence data were collected from the UniProt database. Orthology information of MCS proteins was collected from five databases: EggNOG52, HOGENOM53, OrthoDB54, TreeFam55 and GeneTree56. The PPIs involved in MCS complexes were extracted from the bioGRID database57.
Data Records
Recorded datasets
MCSdb is free available at Figshare43. it provides four types of datasets. The first dataset consisted of detail information of all MCS proteins (xlsx file), including the Entrez ID, protein name, Synonyms, UniProt ID, species, MCS location, and the references (Experimental Method, Cell line/Tissue, PMID and Description and evidences). The second dataset consisted of detail information of all complexes (xlsx file), including complex name, subunit number, species, MCS location, and the information about all subunits (protein names and UniProt ID). And the detail information of the compound subunit was also provided (names, PubChem CID, Formula and SMILES). The third dataset consisted of list of 44 MCS locations and along with their corresponding organelles (xlsx file). The last dataset consisted of detail information of all literatures, including PMIDs, DOI, journal name, authors, title, abstract and published time.
The MCS proteins documented in the database were identified by various experimental methods and thus have different confidence levels. For example, some collected MCS proteins are high-confidence because they were well identified and functionally characterized by multiple low-throughput experimental methods, whereas some other proteins were only screened by the high-throughput method and require further experimental validation9,34. Therefore, after careful consideration of common perspectives from multiple review articles and the characteristics of the data9,10,35,36,37,38,58,59, we graded all documented MCS protein entries into 3 categories according to the confidence level of experimental evidence: LT experimental-based methods, PL-based methods and MS-based methods (confidence: LT-based methods > PL-based methods > MS-based methods). Please note that the data from MS-based methods have a high false positive rate. You are advised to use it with caution.
Data statistics
This current version of MCSdb documents 7010 manually curated MCS protein entries with experimental evidence (including 5985 entries detected by MS-based methods, 616 entries detected by LT-based methods and 409 entries detected by PL-based methods), referring to 24 organelles and 44 MCSs across 11 species; 263 complexes residing in MCSs are also included (Fig. 3a). The MCS category distributions of protein are shown in Fig. 3b–d. The protein entries detected by LT-based methods are distributed in multiple MCSs (Fig. 3b), most of which are located in ER-related MCSs (ER-MT: 213, ER-PM: 88 and ER-Endosome: 52, etc.). The protein entries detected by PL-based methods are divided into MCSs of ER-MT (277 proteins), ER-PM (66 proteins) and ER-Peroxisome (66 proteins). Over 95% protein entries detected by MS-based methods are located in the MCS of ER-MT (5729 proteins). All complexes, which were detected by LT-based methods, are distributed in multiple MCSs (data not shown). The organismal distribution of MCS proteins and complexes is shown in Fig. 3e–g. The protein entries detected by LT-based methods are distributed in 11 species, mostly human (281 proteins) and mouse (150 proteins) proteins (Fig. 3e). A total of 2253 human, 3476 mouse and 256 yeast protein entries were detected by MS-based methods (Fig. 3f). All the protein entries detected by PL-based methods are human proteins (data not shown). All the complexes are distributed in 11 species (Fig. 3g). The subunit number distribution of the complexes is shown in Fig. 3h, and most of the complexes are binary (153/263).
Data statistics of MCSdb. (a) Overview of MCS protein entries and complexes. (b) Category distributions of protein entries detected by LT-based methods. (c) Category distributions of protein entries detected by PL-based methods. (d) Category distributions of protein entries detected by MS-based methods. (e) Organismal distribution of protein entries detected by LT-based methods. (f) Organismal distribution of protein entries detected by MS-based methods. (g) Organismal distribution of complexes. (h) Subunit number distribution of complexes.
Data submission
To acknowledge that the MCSdb collection may not include all proteins residing in MCSs, we offer a ‘Submit’ interface (https://cellknowledge.com.cn/mcsdb/submit.html) for researchers to submit new MCS proteins that have not yet been documented in the database. We will thoroughly review and update all submitted data in a timely manner.
Technical Validation
The MCS protein entries in our database were carefully curated from peer-reviewed literature through manual selection, and we only included experimentally supported MCS proteins. In addition, all collected entries were evaluated and double-checked by at least two expert curators separately. Any discrepancies were resolved by consensus through discussion with the third expert curator.
To ensure the accuracy of our data, we collected detailed information on wet lab experiments used to identify MCS proteins, such as experimental methods and cell lines/tissues, from original articles. Additionally, we extracted original sentences from literature that explicitly described a protein’s role and residence in the MCS, providing further evidence for the accuracy of our data. All of these supporting data from the literature can be easily accessed on the website of our database: https://cellknowledge.com.cn/mcsdb/.
Usage Notes
In addition to accessing the datasets via the Figshare repository43, MCSdb is also free available at https://cellknowledge.com.cn/mcsdb/. Moreover, the website provides a user-friendly ‘Help’ page that presents a step-by-step tutorial to assist users in manipulating, querying, and browsing the MCSdb database. On this ‘Help’ page, we not only offer guidance on maintaining data quality but also provide specific instances of errors as examples for users to reference (refer to the file of all revised entries.xlsx available on the ‘download’ page).
Introduction to Revised Data
During the data collection process, we continuously identified and corrected errors. To better safeguard users against encountering similar issues during literature searches and data collection, we have presented the errors found and their details during the revision process in the form of data tables on our database website (all revised entries.xlsx). In addition, to enhance user awareness and prevent similar mistakes, we have preserved all modified and obsolete entries in the database for user reference. Specifically, for modified entries, we offer two versions on the website: Version 1 and Version 2, with hyperlinks provided at the top of the detail pages for each version (Fig. 4). Version 1 is the original version, in which we have highlighted the specific modifications for user comparison and reference, while Version 2 presents the latest modified data. This approach allows users to easily see the changes made to the data. Moreover, we have separately displayed all obsolete entries in a list format on a dedicated “obsolete list” page, which includes three tables: the obsolete entries list, listing all obsolete protein entries; the table of obsolete Complex Entries list, listing all deleted complex entries; and the obsolete literatures list, listing all removed references.
Code availability
MCSdb is free available at Figshare43 and https://cellknowledge.com.cn/mcsdb/ to all users. The source code for the MCSdb website has been uploaded to GitHub: https://github.com/ZhangCellab/MCSdb.
References
Gottschling, D. E. & Nyström, T. The Upsides and Downsides of Organelle Interconnectivity. Cell 169, 24–34 (2017).
Zhang, Y. et al. CellCall: integrating paired ligand–receptor and transcription factor activities for cell–cell communication. Nucleic Acids Research 49, 8520–8534 (2021).
Huh, J. R. & Veiga-Fernandes, H. Neuroimmune circuits in inter-organ communication. Nat Rev Immunol 20, 217–228 (2020).
Leitão, A. L., Costa, M. C., Gabriel, A. F. & Enguita, F. J. Interspecies Communication in Holobionts by Non-Coding RNA Exchange. Int J Mol Sci 21, 2333 (2020).
Zhang, Y. et al. Cellinker: a platform of ligand-receptor interactions for intercellular communication analysis. Bioinformatics 37, 2025–2032 (2021).
Petkovic, M., Oses-Prieto, J., Burlingame, A., Jan, L. Y. & Jan, Y. N. TMEM16K is an interorganelle regulator of endosomal sorting. Nat Commun 11, 3298 (2020).
Elbaz, Y. & Schuldiner, M. Staying in touch: the molecular era of organelle contact sites. Trends Biochem Sci 36, 616–623 (2011).
Wu, H., Carvalho, P. & Voeltz, G. K. Here, there, and everywhere: The importance of ER membrane contact sites. 361, eaan5835 (2018).
Scorrano, L. et al. Coming together to define membrane contact sites. Nat Commun 10, 1287 (2019).
Prinz, W. A., Toulmay, A. & Balla, T. The functional universe of membrane contact sites. Nat Rev Mol Cell Biol 21, 7–24 (2020).
Venditti, R. & Wilson, C. & De Matteis, M. A. Regulation and physiology of membrane contact sites. Current Opinion in Cell Biology 71, 148–157 (2021).
Chu, Q. et al. Regulation of the ER stress response by a mitochondrial microprotein. Nat Commun 10, 4883 (2019).
Ravindran, R. et al. Endosomal-associated RFFL facilitates mitochondrial clearance by enhancing PRKN/parkin recruitment to mitochondria. Autophagy, 1–14 (2022).
Petkovic, M., O’Brien, C. E. & Jan, Y. N. Interorganelle communication, aging, and neurodegeneration. Genes Dev 35, 449–469 (2021).
Herker, E., Vieyres, G., Beller, M., Krahmer, N. & Bohnert, M. Lipid Droplet Contact Sites in Health and Disease. Trends Cell Biol 31, 345–358 (2021).
Lim, C. Y. et al. ER-lysosome contacts enable cholesterol sensing by mTORC1 and drive aberrant growth signalling in Niemann-Pick type C. Nat Cell Biol 21, 1206–1218 (2019).
Hirabayashi, Y. et al. ER-mitochondria tethering by PDZD8 regulates Ca(2+) dynamics in mammalian neurons. Science 358, 623–630 (2017).
Chao, W. & Quan, Z. O. U. A Machine Learning Method for Differentiating and Predicting Human-Infective Coronavirus Based on Physicochemical Features and Composition of the Spike Protein. Chinese Journal of Electronics 30, 815–823 (2021).
Wu, Y. et al. Contacts between the endoplasmic reticulum and other membranes in neurons. Proc Natl Acad Sci USA 114, 4859–4867 (2017).
Lujan, P., Angulo-Capel, J., Chabanon, M. & Campelo, F. Interorganelle communication and membrane shaping in the early secretory pathway. Current Opinion in Cell Biology 71, 95–102 (2021).
Hoyer, M. J. et al. A Novel Class of ER Membrane Proteins Regulates ER-Associated Endosome Fission. Cell 175, 254–265 (2018).
Jain, A. & Zoncu, R. Organelle transporters and inter-organelle communication as drivers of metabolic regulation and cellular homeostasis. Mol Metab 60, 101481 (2022).
Sandhu, J. et al. Aster Proteins Facilitate Nonvesicular Plasma Membrane to ER Cholesterol Transport in Mammalian Cells. Cell 175, 514–529.e520 (2018).
Zhou, Z. et al. Endoplasmic reticulum-associated degradation regulates mitochondrial dynamics in brown adipocytes. Science 368, 54–60 (2020).
Krawczyk, H. E. et al. SEED LIPID DROPLET PROTEIN1, SEED LIPID DROPLET PROTEIN2, and LIPID DROPLET PLASMA MEMBRANE ADAPTOR mediate lipid droplet-plasma membrane tethering. Plant Cell 34, 2424–2448 (2022).
Loewen, C. J., Roy, A. & Levine, T. P. A conserved ER targeting motif in three families of lipid binding proteins and in Opi1p binds VAP. Embo j 22, 2025–2035 (2003).
Loewen, C. J. & Levine, T. P. A highly conserved binding site in vesicle-associated membrane protein-associated protein (VAP) for the FFAT motif of lipid-binding proteins. J Biol Chem 280, 14097–14104 (2005).
Loewen, C. J., Young, B. P., Tavassoli, S. & Levine, T. P. Inheritance of cortical ER in yeast is required for normal septin organization. J Cell Biol 179, 467–483 (2007).
Chao, J. T. et al. Polarization of the endoplasmic reticulum by ER-septin tethering. Cell 158, 620–632 (2014).
Murray, R., Flora, E., Bayne, C. & Derré, I. IncV, a FFAT motif-containing Chlamydia protein, tethers the endoplasmic reticulum to the pathogen-containing vacuole. Proc Natl Acad Sci USA 114, 12039–12044 (2017).
Wang, X., Wen, Y., Dong, J., Cao, C. & Yuan, S. Systematic In-Depth Proteomic Analysis of Mitochondria-Associated Endoplasmic Reticulum Membranes in Mouse and Human Testes. Proteomics 18, e1700478 (2018).
Horner, S. M., Wilkins, C., Badil, S., Iskarpatyoti, J. & Gale, M. Jr. Proteomic analysis of mitochondrial-associated ER membranes (MAM) during RNA virus infection reveals dynamic changes in protein and organelle trafficking. PLoS One 10, e0117963 (2015).
Ma, J. H. et al. Comparative Proteomic Analysis of the Mitochondria-associated ER Membrane (MAM) in a Long-term Type 2 Diabetic Rodent Model. Sci Rep 7, 2062 (2017).
Huang, X., Jiang, C., Yu, L. & Yang, A. Current and Emerging Approaches for Studying Inter-Organelle Membrane Contact Sites. Front Cell Dev Biol 8, 195 (2020).
Kwak, C. et al. Contact-ID, a tool for profiling organelle contact sites, reveals regulatory proteins of mitochondrial-associated membrane formation. Proc Natl Acad Sci USA 117, 12109–12120 (2020).
Cho, K. F. et al. Split-TurboID enables contact-dependent proximity labeling in cells. Proc Natl Acad Sci USA 117, 12143–12154 (2020).
Go, C. D. et al. A proximity-dependent biotinylation map of a human cell. Nature 595, 120–124 (2021).
Jing, J. et al. Proteomic mapping of ER-PM junctions identifies STIMATE as a regulator of Ca2+ influx. Nat Cell Biol 17, 1339–1347 (2015).
Hua, R. et al. VAPs and ACBD5 tether peroxisomes to the ER for peroxisome maintenance and lipid homeostasis. J Cell Biol 216, 367–377 (2017).
Li, S. et al. A new type of ERGIC-ERES membrane contact mediated by TMED9 and SEC. 12 is required for autophagosome biogenesis. Cell Res 32, 119–138 (2022).
Boutry, M. & Kim, P. K. ORP1L mediated PI(4)P signaling at ER-lysosome-mitochondrion three-way contact contributes to mitochondrial division. Nat Commun 12, 5354 (2021).
Wang, C. et al. FUNDC1-dependent mitochondria-associated endoplasmic reticulum membranes are involved in angiogenesis and neoangiogenesis. Nat Commun 12, 2616 (2021).
Pan, X. et al. MCSdb, a database of proteins residing in membrane contact sites, figshare, https://doi.org/10.6084/m9.figshare.c.6634166.v1 (2023).
Licata, L. et al. MINT, the molecular interaction database: 2012 update. Nucleic Acids Research 40, 857–861 (2012).
Calderone, A., Castagnoli, L. & Cesareni, G. mentha: a resource for browsing integrated protein-interaction networks. Nat Methods 10, 690–691 (2013).
Li, T. et al. A scored human protein-protein interaction network to catalyze genomic interpretation. Nat Methods 14, 61–64 (2017).
UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Research 49, 480–489 (2021).
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Research 49, 10–17 (2021).
Kim, S. et al. PubChem in 2021: new data content and improved web interfaces. Nucleic Acids Research 49, 1388–1395 (2021).
Uhlén, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Palasca, O., Santos, A., Stolte, C., Gorodkin, J. & Jensen, L. J. TISSUES 2.0: an integrative web resource on mammalian tissue expression. Database 2018, bay028 (2018).
Huerta-Cepas, J. et al. eggNOG 5.0: a hierarchical, functionally and phylogenetically annotated orthology resource based on 5090 organisms and 2502 viruses. Nucleic Acids Research 47, 309–314 (2019).
Penel, S. et al. Databases of homologous gene families for comparative genomics. BMC Bioinformatics 10, S3 (2009).
Zdobnov, E. M. et al. OrthoDB in 2020: evolutionary and functional annotations of orthologs. Nucleic Acids Research 49, 389–393 (2020).
Schreiber, F., Patricio, M., Muffato, M., Pignatelli, M. & Bateman, A. TreeFam v9: a new website, more species and orthology-on-the-fly. Nucleic Acids Research 42, 922–925 (2014).
Howe, K. L. et al. Ensembl Genomes 2020-enabling non-vertebrate genomic research. Nucleic Acids Research 48, 689–695 (2020).
Oughtred, R. et al. The BioGRID database: A comprehensive biomedical resource of curated protein, genetic, and chemical interactions. Protein Sci 30, 187–200 (2021).
Liao, P. C. et al. Touch and Go: Membrane Contact Sites Between Lipid Droplets and Other Organelles. Front Cell Dev Biol 10, 852021 (2022).
Li, C. et al. Endoplasmic Reticulum-Plasma Membrane Contact Sites: Regulators, Mechanisms, and Physiological Functions. Front Cell Dev Biol 9, 627700 (2021).
Acknowledgements
We thank Prof Hao Lin for constructive comments of this research. We thank Springer Nature Author Services for the language editing of this manuscript. This study was supported by the National Natural Science Foundation of China (62131004, 62202069), the Natural Science Foundation of Sichuan Province (2022NSFSC1610, 2023NSFSC0678, 2023NSFSC0569), the China Postdoctoral Science Foundation (2023M730507) and the Sichuan Province Postdoctoral Research Project Special Support Foundation (TB2023012).
Author information
Authors and Affiliations
Contributions
Y.Z., H.C.L. and Q.Z. directed the project and wrote the manuscript. X.R.P., Y.Y., and L.P.R. collected the data. X.R.P. and Y.X. constructed the database. Y.Y., L.P.R., L.N., H.C.L. and Y.B.Z. checked the data and debug the website. All named authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pan, X., Ren, L., Yang, Y. et al. MCSdb, a database of proteins residing in membrane contact sites. Sci Data 11, 281 (2024). https://doi.org/10.1038/s41597-024-03104-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03104-7
- Springer Nature Limited