Abstract
The human mannose-binding lectin (MBL) gene encodes a polymorphic protein that plays a crucial role in the innate immune response. Human MBL deficiency is associated with immunodeficiencies, and its variants have been linked to autoimmune and infectious diseases. Despite this significance, gene studies concerning MBL sequencing are uncommon in Malaysia. Therefore, we aimed to preliminary described the human MBL sequencing dataset based on the Kelantan population. Blood samples were collected from 30 unrelated individuals and underwent DNA extraction, genotyping, and sequencing. The sequencing data generated 886 bp, which were deposited in GenBank (ON619541-ON619546). Allelic variants were identified and translated into six MBL haplotypes: HYPA, HYPB, LYPB, LXPB, HXPA, and LXPA. An evolutionary tree was constructed using the haplotype sequences. These findings contribute to the expansion of MBL information within the country, providing a valuable baseline for future research exploring the association between the gene and targeted diseases.
Similar content being viewed by others
Background & Summary
Human mannose-binding lectin (MBL) is a C-type serum lectin produced in the liver that plays a crucial role in innate immunity1. MBL may mediate phagocytosis by binding to specific carbohydrate moieties on various pathogens, thereby utilising identified phagocyte receptors2. Additionally, MBL employs MBL-associated serine proteases (MASP)-1 and −2 to activate the MBL pathway of complement3. The MBL protein is encoded by the polymorphic MBL gene, which consists of four exons interrupted by three introns and is located on chromosome 10 (10q11.2-q21)4.
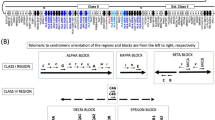
Single nucleotide polymorphisms (SNPs) at specific nucleotide positions have been identified: −550 (G/C or H/L), −221 (C/G or X/Y), +4 (C/T or P/Q), +223 (C/T or A/D), +230 (G/A or A/B) and +239 (G/A or A/C) within the promoter/5′ untranslated and exon 1 regions5. Functional characterisation of SNPs in the promoter region, altering MBL transcription, underscores the significance of these genetic differences for MBL circulating levels and expression6. In the exon 1 region, the wild-type (normal) allele is denoted as A, while the B (codon 52), C (codon 54) and D (codon 57) allelic variants are collectively referred to as O. Changes in amino acid resulting from exon 1 variations are believed to influence the functional properties of the MBL protein collagenous region7.
Theoretically, 64 unique haplotypes could be generated by combining those six polymorphisms. However, only eight common haplotypes are typically observed in population studies due to significant linkage disequilibrium (LD) between the promoter and exon 1 SNPs5,6. Different haplotypes have been associated with high MBL levels, such as HYPA, LYQA, and LYPA; an intermediate MBL level represented by LXPA; and haplotypes linked to low MBL levels, including HYPD, LYPB, LYQC, and LYPD8,9.
Within the promoter and exon 1 regions, at least 30 SNPs and six deletion sites have been previously identified10. However, SNPs at positions −550, −221, +4, +223, +230, and +239 are commonly investigated and recognised as common point mutation secretors. These SNPs are frequently studied to characterise MBL variants and determine serum MBL protein levels. This strategy can be implemented using either serum level measurements or genotyping method such as polymerase chain reaction-sequence-specific amplification (PCR-SSP), -sequence-based typing (PCR-SBT), -restriction fragment length polymorphism (PCR-RFLP) or direct sequencing. Previous studies have shown that these MBL SNPs are associated with autoimmune diseases such as rheumatoid arthritis (RA)11,12, Sjögren syndrome (SS)13,14 and systemic lupus erythematosus (SLE)15,16; as well as infectious diseases including pulmonary tuberculosis (TB)17,18, acute respiratory infection (ARI)19,20 and vulvovaginal candidiasis (VVC)21,22.
Despite a wide coverage of studies and a flux of research findings, studies on the MBL gene in Malaysia are uncommon. Here we preliminary described the human MBL sequencing datasets, which include six common point mutation secretors and five additional polymorphic sites in the promoter region among Kelantan individuals (Table 1, Fig. 1). A diagram illustrating the workflow in this study is presented in Fig. 2. Genomic DNA was extracted from blood samples and subjected to direct PCR and Sanger sequencing. The resulting 886 bp sequence products were assembled using molecular genetic software to identify MBL haplotypes, which were later used to construct an evolutionary tree. This preliminary study provides baseline information for future research on the association between MBL gene polymorphisms and targeted diseases. Additionally, the generated data will be representative of the Malaysian population and may encourage broader population studies in the future.
Methods
Ethical statement
This study was approved by the Human Research Ethics Committee of Universiti Sains Malaysia (USM/JEPeM/19090533) and was performed in accordance with the guidelines set forth by the National Blood Centre, Ministry of Health Malaysia. Signed informed consent and demographic background information were obtained from each participant who agreed to take part in the study, and the data will be published without revealing their identities.
Sample collection and DNA extraction
Genomic samples were collected from 30 unrelated individuals from Kelantan. Approximately, 10 cc of blood was drawn from the peripheral vein and stored in the ethylenediaminetetraacetic acid (EDTA) tube. The total DNA was extracted using the gSYNCTM DNA Extraction Kit (Geneaid, Taiwan), following protocol provided by the manufacturer.
MBL genotyping
The isolated genomic DNA was amplified using a set of forward reverse primers specific for MBL (Table 2)19. The PCR amplification was carried out using Veriti TM 96-well fast thermal cycler. Subsequently, the amplified PCR products were purified using the GeneJET PCR Purification Kit (Thermo Scientific, United States). DNA sequencing was performed using ABI 3100 DNA Sequencer at First Base Laboratories Sdn Bhd (Malaysia).
Sequencing analysis
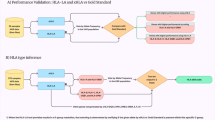
The raw sequences were visualised and analysed using SnapGene version 6.1 (Fig. 3). Subsequently, all sequences were aligned with the Homo sapiens MBL2 RefSeqGene (LRG_154) of chromosome 10, utilising Sequencher version 5.4.6. The identification of MBL haplotypes was based on the allelic mutations (Table 1)5. Additionally, a Neighbor-joining tree23 was constructed using the Molecular Evolutionary Genetics Analysis (MEGA) version 1124, with Pan troglodytes (chimpanzee) sequences (AY970679 and AY970685)25 as an outgroup (Fig. 4).
Data Records
The sequencing data are accessible in the National Centre for Biotechnology Information (NCBI), assigned with the reference numbers ON619541-ON619546 (Table 3)26,27,28,29,30,31. These sequencing information details pertain to a subset of individuals.
Technical Validation
The MBL-PCR products underwent assessment to confirm the absence of contamination through 2% gel electrophoresis. Subsequently, the raw sequences were aligned with RefSeqGene (NG_008196), and each sequence was manually inspected to ensure the absence of three stop codons: UAA, UAG and UGA. The evolutionary history was inferred using the NJ method23. The percentage of replicate trees (>50%) in which the associated taxa clustered together in the bootstrap test (1,000 replicates) I indicated next to the branches32.
Code availability
The analyses were conducted using the version and parameters as described below:
1) SnapGene, version 6.1, parameters used: interrupted circle, show polymorphism cite locations, unique 6+ cutters
2) Sequencher, version 5.4.6, parameters used: minimum match percentage 85, minimum overlap 20.
3) MEGA, version 1124, parameters used: NJ tree with Kimura2 parameter model33.
References
Turner, M. W. Mannose-binding lectin: the pluripotent molecule of the innate immune system. Immunol Today 17, 532–540, https://doi.org/10.1016/0167-5699(96)10062-1 (1996).
Nepomuceno, R. R., Henschen-Edman, A. H., Burgess, W. H. & Tenner, A. J. cDNA cloning and primary structure analysis of C1qR(P), the human C1q/MBL/SPA receptor that mediates enhanced phagocytosis in vitro. Immunity 6, 119–129, https://doi.org/10.1016/s1074-7613(00)80419-7 (1997).
Matsushita, M. & Fujita, T. Activation of the classical complement pathway by mannose-binding protein in association with a novel C1s-like serine protease. J Exp Med 176, 1497–1502, https://doi.org/10.1084/jem.176.6.1497 (1992).
Sastry, K. et al. The human mannose-binding protein gene. Exon structure reveals its evolutionary relationship to a human pulmonary surfactant gene and localization to chromosome 10. J Exp Med 170, 1175–1189, https://doi.org/10.1084/jem.170.4.1175 (1989).
Madsen, H. O., Satz, M. L., Hogh, B., Svejgaard, A. & Garred, P. Different molecular events result in low protein levels of mannan-binding lectin in populations from southeast Africa and South America. J Immunol 161, 3169–3175 (1998).
Madsen, H. O. et al. Interplay between promoter and structural gene variants control basal serum level of mannan-binding protein. J Immunol 155, 3013–3020 (1995).
Larsen, F., Madsen, H. O., Sim, R. B., Koch, C. & Garred, P. Disease-associated mutations in human mannose-binding lectin compromise oligomerization and activity of the final protein. J Biol Chem 279, 21302–21311, https://doi.org/10.1074/jbc.M400520200 (2004).
Naito, H. et al. Characterization of human serum mannan-binding protein promoter. J Biochem 126, 1004–1012, https://doi.org/10.1093/oxfordjournals.jbchem.a022543 (1999).
Minchinton, R. M., Dean, M. M., Clark, T. R., Heatley, S. & Mullighan, C. G. Analysis of the relationship between mannose-binding lectin (MBL) genotype, MBL levels and function in an Australian blood donor population. Scand J Immunol 56, 630–641, https://doi.org/10.1046/j.1365-3083.2002.01167.x (2002).
Bernig, T. et al. Sequence analysis of the mannose-binding lectin (MBL2) gene reveals a high degree of heterozygosity with evidence of selection. Genes Immun 5, 461–476, https://doi.org/10.1038/sj.gene.6364116 (2004).
Gupta, B. et al. Association of mannose-binding lectin gene (MBL2) polymorphisms with rheumatoid arthritis in an Indian cohort of case-control samples. Journal of Human Genetic 50, 583–591, https://doi.org/10.1007/s10038-005-0299-8 (2005).
Martiny, F. L. et al. Mannose-binding lectin gene polymorphisms in Brazilian patients with rheumatoid arthritis. J Rheumatol 39, 6–9, https://doi.org/10.3899/jrheum.110052 (2012).
Wang, Z. Y., Morinobu, A., Kanagawa, S. & Kumagai, S. Polymorphisms of the mannose binding lectin gene in patients with Sjögren’s syndrome. Ann Rheum Dis 60, 483–486, https://doi.org/10.1136/ard.60.5.483 (2001).
Ramos-Casals, M. et al. Mannose-binding lectin-low genotypes are associated with milder systemic and immunological disease expression in primary Sjögren’s syndrome. Rheumatology (Oxford) 48, 65–69, https://doi.org/10.1093/rheumatology/ken411 (2009).
Øhlenschlaeger, T., Garred, P., Madsen, H. O. & Jacobsen, S. Mannose-binding lectin variant alleles and the risk of arterial thrombosis in systemic lupus erythematosus. N Engl J Med 351, 260–267, https://doi.org/10.1056/NEJMoa033122 (2004).
Font, J. et al. Association of mannose-binding lectin gene polymorphisms with antiphospholipid syndrome, cardiovascular disease and chronic damage in patients with systemic lupus erythematosus. Rheumatology (Oxford) 46, 76–80, https://doi.org/10.1093/rheumatology/kel199 (2007).
Liu, C. et al. Association of Mannose-binding Lectin Polymorphisms with Tuberculosis Susceptibility among Chinese. Sci Rep 6, 36488, https://doi.org/10.1038/srep36488 (2016).
Guo, Y. L., Liu, Y., Ban, W. J., Sun, Q. & Shi, G. L. Association of mannose-binding lectin gene polymorphisms with the development of pulmonary tuberculosis in China. BMC Infectious Diseases 17, 1–7, https://doi.org/10.1186/s12879-017-2310-3 (2017).
Tao, R., Hua, C. Z., Hu, Y. Z. & Shang, S. Q. Genetic polymorphisms and serum levels of mannose-binding lectin in Chinese pediatric patients with common infectious diseases. International Journal of Infectious Diseases 16, e403–e407, https://doi.org/10.1016/j.ijid.2012.01.014 (2012).
Toivonen, L. et al. Polymorphisms of Mannose-binding Lectin and Toll-like Receptors 2, 3, 4, 7 and 8 and the Risk of Respiratory Infections and Acute Otitis Media in Children. Pediatr Infect Dis J 36, e114–e122, https://doi.org/10.1097/inf.0000000000001479 (2017).
Hammad, N. M., El Badawy, N. E., Nasr, A. M., Ghramh, H. A. & Al Kady, L. M. Mannose-Binding Lectin Gene Polymorphism and Its Association with Susceptibility to Recurrent Vulvovaginal Candidiasis. Biomed Res Int 2018, 7648152, https://doi.org/10.1155/2018/7648152 (2018).
Kalia, N., Singh, J., Sharma, S. & Kaur, M. Impact of SNPs interplay across the locus of MBL2, between MBL and Dectin-1 gene, on women’s risk of developing recurrent vulvovaginal infections. Cell Biosci 9, 35, https://doi.org/10.1186/s13578-019-0300-4 (2019).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4, 406–425, https://doi.org/10.1093/oxfordjournals.molbev.a040454 (1987).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol Biol Evol 38, 3022–3027, https://doi.org/10.1093/molbev/msab120 (2021).
Seyfarth, J., Garred, P. & Madsen, H. O. The ‘involution’ of mannose-binding lectin. Hum Mol Genet 14, 2859–2869, https://doi.org/10.1093/hmg/ddi318 (2005).
Zahidin, M. A. et al. Homo sapiens mannan/mannose-binding protein variant LXPA gene, partial cds. Genbank https://identifiers.org/nucleotide:ON619541 (2023).
Zahidin, M. A. et al. Homo sapiens mannan/mannose-binding protein variant LYPB gene, partial cds. Genbank, https://identifiers.org/nucleotide:ON619546 (2023).
Zahidin, M. A. et al. Homo sapiens mannan/mannose-binding protein variant HXPA gene, partial cds. Genbank https://identifiers.org/nucleotide:ON619545 (2023).
Zahidin, M. A. et al. Homo sapiens mannan/mannose-binding protein variant LXPB gene, partial cds. Genbank https://identifiers.org/nucleotide:ON619544 (2023).
Zahidin, M. A. et al. Homo sapiens mannan/mannose-binding protein variant HYPB gene, partial cds. Genbank https://identifiers.org/nucleotide:ON619543 (2023).
Zahidin, M. A. et al. Homo sapiens mannan/mannose-binding protein variant HYPA gene, partial cds. Genbank https://identifiers.org/nucleotide:ON619542 (2023).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–791, https://doi.org/10.1111/j.1558-5646.1985.tb00420.x (1985).
Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16, 111–120, https://doi.org/10.1007/bf01731581 (1980).
Acknowledgements
We would like to thank the Hospital Universiti Sains Malaysia and Central Research Laboratory, School of Medical Sciences, Universiti Sains Malaysia (Health Campus) for the administration and logistic supports. This study was financially supported by the Ministry of Higher Education of Malaysia (FRGS/203/PPSP/6171241).
Author information
Authors and Affiliations
Contributions
M.A.Z., N.H.M.N. and H.A.E. designed the experiments. M.A.Z. collected blood samples and the whole processed were supervised by N.H.M.N., M.F.J., A.D.A. and H.A.E. M.A.Z. analysed the data and was evaluated by N.H.M.N., Z.Z., M.F.J. and H.A.E. The manuscript was written by M.A.Z., N.H.M.N. and Z.Z. with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zahidin, M.A., Mohd Noor, N.H., Johan, M.F. et al. Mannose-binding lectin gene sequence data in Kelantan population. Sci Data 11, 435 (2024). https://doi.org/10.1038/s41597-024-03274-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03274-4
- Springer Nature Limited