Abstract
Sinosolenaia oleivora (Bivalve, Unionida, Unionidae), is a near-endangered edible mussel. In 2022, it was selected by the Ministry of Agriculture and Rural Affairs as a top-ten aquatic germplasm resource, with potential for industrial development. Using Illumina, PacBio, and Hi-C technology, a high-quality chromosome-level genome of S. oleivora was assembled. The assembled S. oleivora genome spanned 2052.29 Mb with a contig N50 size of 20.36 Mb and a scaffold N50 size of 103.57 Mb. The 302 contigs, accounting for 98.41% of the total assembled genome, were anchored into 19 chromosomes using Hi-C scaffolding. A total of 1171.78 Mb repeat sequences were annotated and 22,971 protein-coding genes were predicted. Compared with the nearest ancestor, a total of 603 expanded and 1767 contracted gene families were found. This study provides important genomic resources for conservation, evolutionary research, and genetic improvements of many economic traits like growth performance.
Similar content being viewed by others
Background & Summary
Freshwater mussels (Unionoida) represent the most diverse order of freshwater bivalves1 and are found in all regions of the world except the Antarctic2. They not only play an important role in the food web structure and material cycle of ecosystems3,4 but also have high economic value, such as for food5, pearl cultivation6, and anti-tumor ingredients7. They also have been used as an indicator for biological monitoring and evaluation of heavy metal pollution8.
Freshwater mussels are benthic filter feeders9. Suitable substrate, water quality, and food are important factors for the survival and reproduction of mussels. In recent years, human activities, such as river diversion, chemical pollution, and overfishing have caused serious damage to mussel habitats10. The developmental life history of most mussels involves a parasitic larval stage (glochidia) that must attach to vertebrate hosts (primarily fish) to complete metamorphosis11 which increases their vulnerability2. The International Union for Conservation of Nature (IUCN) Red List reports that 173 species are extinct, endangered, or threatened, 99 are vulnerable or nearly threatened, and 84 are unclassified because data are deficient12.
There are 57 endemic species in China13, and eight species have now been listed as Grade II national protected animals14. The biodiversity and population size of freshwater mussels in large water bodies such as the Yangtze River15 and the Songhua River16 have shown a significant decline. S.oleivora is endemic to China. In 2022, S. oleivora was identified as one of the top ten characteristic aquatic germplasm resources by the Ministry of Agriculture and Rural Affairs. S. oleivora has fresh and tender meat, delicious taste, and high nutrient content17. In Fuyang of Anhui Province, Tianmen of Hubei Province, and other places, S. oleivora is a famous delicacy with a high economic value, and it is called “abalone in Huaihe River.” It once ranged an extensive distribution—in five freshwater lakes and the tributaries of the Yangtze and Huaihe Rivers18. Habitat fragmentation and other human activities (e.g., overfishing) have resulted in their endangerment19. Tianmen in Hubei Province and Fuyang in Anhui Province has established the S. oleivora Nature Reserve to support this ecologically and economically vital resource.
Genomic data is considered fundamental for revealing biological characteristics, inferring evolutionary mechanisms, and promoting effective conservation20. To date, only seven freshwater mussel species have had their genomes sequenced (Table S1, Supplementary File)21,22,23,24,25,26,27,28, and only one of these is a Chinese species27. The whole genome of S. oleivora is lacking. We applied multiple sequencing technologies, including Illumina Nova 6000 sequencing, PacBio long-read sequencing (PacBio), and high-throughput chromosome conformation capture (Hi-C) technology to complete genome sequencing and assembly. Three methods, including de novo gene prediction, homolog, and RNA-Seq-based prediction, were used to perform genomic annotation. In addition, the comparative genomics analysis of S. oleivora and 10 other distantly related species was performed. This study provides important genomic resources for conservation and evolutionary research and guides genetic trait improvements (e.g., growth).
Methods
Sample collection and sequencing
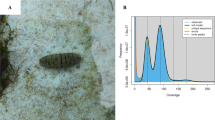
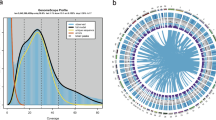
One female S. oleivora was sampled from the national-level protection zone of the aquatic germplasm resource of S. oleivora in the Fuyang Division of Huaihe River (32.428725°N, 115.600287°E). Total DNA was extracted from the adductor muscle of S. oleivora using the DNeasy Blood and Tissue Kit (Qiagen, Germany) for genome sequencing. For short-read sequencing, Covaris M220 was used to break DNA into 300–350 bp fragments. DNA library preparation was completed by terminal repair, an A-tail addition, sequencing junction addition, DNA purification, and bridge PCR. Based on a paired-end(PE) sequencing strategy. These libraries were sequenced on the Illumina NovaSeq Nova 6000 platform. For long-read sequencing, according to the PacBio standard protocol, a PacBio HiFi library was generated using an SMRTbell Template Prep Kit 2.0 (Pacific Biosciences, USA) and sequenced using the PacBio Sequel II platform. A Hi-C library was prepared following the Hi-C library protocol29 and sequenced using the Illumina Novaseq 6000 platform. Total RNA was extracted from the adductor muscle of S. oleivora using TRIzol reagent (Invitrogen, MA, USA) for transcriptome sequencing. The RNA-seq library was generated using NEBNext®UltraTM RNA Library Prep Kit (NEB, USA) for PE sequencing, and short reads were produced on the Illumina NovaSeq 6000 platform. A total of 192.1 Gb of Illumina data, 63.2 Gb of PacBio data, 191.8 Gb of Hi-C data, and 5.6 Gb RNA-Seq data were obtained (Fig. 1, Table 1).
Estimation of genome size
A K-mer-based method30 was applied to estimate the genome size, heterozygosity, and repeat content in S. oleivora. We performed a k-mer (k = 17) frequency distribution analysis using 192.1 Gb of Illumina clean data (Fig. 2). A total of 153,573,141,235 k-mers with a depth of 73 was obtained. The genome size was 2,025 Mb, the heterozygosity ratio was 0.78%, and the repeat sequence ratio was 61.37%.
Genome assembly
PacBio Hi-Fi reads were assembled using Hifiasm(v. 0.16.1-r375) software31 with the default parameters. Redundant sequences were filtered out using Purge_Haplotigs (v1.0.4) software32 with the parameter of cutoff “-a 70 -j 80 -d 200.” Based on PacBio sequencing data, the genome length was 2090.51 Mb. The number of contigs was 302 and N50 reached 23.99 Mb. The max length was 88.20 Mb and the GC content was 34.38% (Table 2).
Hi-C-assisted chromosome-level assembly
To assemble the chromosome-level genome, Hi-C sequencing data were mapped and sorted against the draft genome assembly with Juicer v1.6 software33. The contigs were linked to 19 distinct chromosomes by 3D-DNA (v. 180922)34. Based on chromosome interactions, the contig orientation was corrected and suspicious fragments were removed from the contigs in the Juicebox software35. The genome contigs were further anchored and oriented to chromosomes by Hi-C scaffolding. The Hi-C library generated 191.8.2 Gb of clean data, with 55.56% valid pairs. A total of 302 contigs, accounting for 98.41% of the total assembled genome, were anchored into 19 chromosomes. The 19 pseudo-chromosomes were clearly distinguished from the Hi-C heatmap with strong pseudo-chromosome interactions confirming high-quality Hi-C assembly (Figs. 3, 4). This resulted in a high-quality genome of 2052.30 Mb, with a contig N50 of 20.36 Mb and scaffold N50 of 103.57 Mb (Table 3).
Chromosomes Hi-C heatmap of Sinosolenaia oleivora. Blocks represent height pseudochromosomes. The color bar represents contact density from white (low) to red (high). The same applies to Fig. 4.
Repeat annotation, gene prediction, and gene functional annotation
Combined homologous and de novo prediction methods, repeat elements of the S. oleivora genome, were annotated. For homologous alignment, we used RepeatMasker (v4.1.2-p1)36 and Repeat-proteinmask (v4.1.0)37 to annotate the transposable elements (TEs) by comparing sequences to the Repbase database38. For de novo prediction, Tandem Repeat Finder (TRF) (version 4.09)39 was executed to detect the tandem repeat elements based on sequence features. LTR_FINDER (v. 1.07)40 and RepeatModeler (v. 2.0.3)36 were used to construct a repeat library. The library was then used to detect repetitive sequences by RepeatMasker (v. 4.1.2-p1)36. After eliminating redundancy, we obtained the final annotated repeat sets. A total of 1171.79 Mb repeat sequences were annotated accounting for 56.05% of the total genome sequence (Table 4). The major repetitive elements were DNA (15.74%), long interspersed nuclear elements (LINEs, 8.95%), and long terminal repeats (LTRs, 4.98%) (Table 5).
The genome sequence was soft-masked based on repetitive element predictions and then used for protein-coding gene prediction. We employed three methods for gene prediction. For homology-based annotation, the protein sequences of Mizuhopecten yessoensis, Crassostrea gigas, Crassostrea virginica, and Mytilus galloprovincialis were downloaded from NCBI and aligned to the genome sequence using BLAST(E-value: 1e-5)41. Homologous sequences were then aligned to corresponding matching proteins using GeneWise (v. wise2-4-1)42. For the RNA-seq-based annotation, transcriptomic data were assembled using Trinity v2.1143, and BLAST(E-value: 1e-5)41 to align transcriptome to the genome. For de novo prediction, Augustus(v3.4.0)44, and Genscan (version1.0)45 were used to generate de novo-predicted gene sets. Maker (v2.31.10)46 was used to integrate the results from these methods to produce the final gene set. The genome sequence was also aligned to the homologous single-copy gene database of Benchmarking Universal Single-Copy Orthologs(BUSCO)47. MAKER (version 2.31.10)48 and HiCESAP (Wuhan Gooalgene Co., Ltd., https://www.gooalgene.com/) were employed to merge all the data and filter out redundancies. The combination of de novo and homolog-based methods predicted 22,971 protein-coding genes (Table 6). The predicted genes were functionally annotated based on exogenous protein databases including SwissProt, InterPro, TrEMBL, Kyoto Encyclopedia of Genes and Genomes (KEGG), and Gene Ontology (GO). A total of 19,229 genes, accounting for 87.52% of all predicted genes, were annotated using public databases (Table 7).
Based on Rfam49 and miRbase50 databases, we used tRNAscan-SE (v1.3.1)51 to identify transfer RNAs (tRNAs), and Infernal(v1.1.2)52 to annotate other ncRNAs, including microRNAs (miRNAs) and small nuclear RNAs (snRNAs), and BLAST(E-value: 1e-5)41 was used to obtain ribosomal RNA (rRNA) to predict noncoding RNA (ncRNA) in the genome of S. oleivora. For non-coding RNA predictions, we successfully annotated 119 miRNAs, 2643 tRNAs, 366 rRNAs, and 867 snRNAs, with average lengths of 98, 74, 254, and 168 bp, respectively (Table 8).
Comparative genomic analyses
To clarify the evolutionary position of S. oleivora, OrthoMCL (Verison v2.0.9)53 with the parameter “-l 1.5” was used to detect orthologous groups by retrieving the protein sequences of Mizuhopecten yessoensis, Biomphalaria glabrata, Crassostrea gigas, C. virginica, Lingula anatina, Lottia gigantea, Mercenaria mercenaria, Ostrea edulis, Pecten maximus, and Pomacea canaliculate. Sequence alignment was performed by MUSCLE(v5)54 for single-copy orthologous genes. Basing on this result, KaKs Calculator(v2.0)55 was utilized to fetch Kolmogorov-Smirnov(Ks) with default parameters. The S. oleivora genome shared 82,067 gene families and 17,699 single-copy genes with ten other mollusk species. The S. oleivora genome contained 21971 genes clustered into 18,312 gene families and 2,273 unique families (Table 9). The phylogenetic tree was constructed using the “-f a -N 100 -m GTRGAMMA” parameter of RAxML (version 8.2.12)56 based on multiple sequence alignment. Divergence times were estimated using the MCMCtree (v4.9) program in PAML (v4.9)57 with clock = 3 and model = 0 parameters. The divergence time of L. anatina and C. gigas 619.3 (582.0–689.2 MYA); B. glabrata and C. gigas 544.1 (520.2–567.9 MYA); P. canaliculata and B. glabrata 444.6 (377.0–490.4 MYA) from TimeTree database58 (http://www.timetree.org/) were used for calibration. Divergence time analysis showed that S. oleivora was closely related to M. mercenaria, with a divergence time of 516.7 (486.9–541.0) Mya (Fig. 5).
CAFE59,60 was applied for gene expansion and contraction analysis. Compared with the nearest ancestor, a total of 603 expanded and 1767 contracted gene families were found in S. oleivora (Fig. 6). There were 69 significantly expanded (984 genes) and 83 significantly contracted (118 genes) gene families (p < 0.05). We then performed GO and KEGG enrichment analysis and terms with enrichment-adjusted p-values ≤ 0.05 were chosen for further analysis. The program CODEML (v4.9)57 of PAML was used for positive selection gene (PSG) identification. PSGs were also chosen for enrichment analysis. A total of 552 protein-coding genes were positively selected in S. oleivora (FDR < 0.05, Table 10). GO and KEGG enrichment of positively selected genes focused on the DNA binding, nucleolus, and protein processing in the endoplasmic reticulum, ribosome, and mTOR signaling pathway (Figs. 7, 8).
Data Records
All sequencing data from three sequencing platforms have been uploaded to the NCBI SRA database (transcriptomic sequencing data: SRR2835217161, genomic Illumina sequencing data: SRR2655134462, genomic PacBio sequencing data: SRR2840605563, Hi-C sequencing data: SRR2840626464). The final chromosome-level assembled genome file has been uploaded to the GenBank database under the accession JBDPLI00000000065. Genome annotation files have been uploaded to the Figshare database66.
Technical Validation
Evaluating the quality of the DNA and RNA
The quality and concentration of extracted DNA/RNA were assessed using NanoDrop 2000 Spectrophotometer (Thermo Fisher Scientific, San Jose, CA, USA) and Qubit 3.0 Fluorometer (Thermo Fisher Scientific, San Jose, CA, USA)(OD260/280 and OD260/230) before the genome sequencing and their integrity was further evaluated on 1% agarose gel stained with ethidium bromide.
Evaluating the quality of the genome assembly
We evaluated the genome assembly quality through the following measures: (i) Confirmation that the assembly result belongs to the target species was made by software BLAST(E-value: 1e-5)26 comparison to the NCBI nucleotide database (NT library)(Table S2, S3, Supplementary File);(ii) Illumina short reads and PacBio reads were mapped onto the assembled genome using BWA (v. 0.7.17-r1188)67 and Minimap268 to evaluate the completeness and accuracy of the genome. The read-mapping rates were 99.27% and 99.74%, and genome coverage rates were 99.7% and 99.98% for the Illumina and PacBio reads, respectively (Table 11), indicating high mapping efficiency and comprehensive coverage. (iii) BUSCO (v5.2.3)32 analysis was conducted to evaluate the assembly quality based on the mollusca_odb10 database. Using BUSCO analysis, 100% (5295/5295) of complete BUSCO genes were found in the assembly, including 88.6% complete BUSCOs, 85.8% complete and single-copy BUSCOs, and 2.8% complete and duplicated BUSCOs (Table 12).
Evaluating the quality of the genome annotation
BUSCO (v5.2.2)32 was used to evaluate the completeness of the genome annotation. The reference BUSCO database was mollusca_odb10. Among the 5295 BUSCO groups searched, 4575 (86.4%) of the complete BUSCOs were detected in the genome annotations (Table 12).
Code availability
The manuscript did not use custom code to generate or process the data described.
References
Bieler, R., Carter, J. G. & Coan, E. V. Classification of bivalve families. Pp. 113–133, in: Bouchet, P. & Rocroi, J.P. (2010), Nomenclator of Bivalve Families. Malacologia. 52, 1–184 (2010).
Bogan, A. E. Global diversity of freshwater mussels (Mollusca, Bivalvia) in freshwater. Hydrobiologia. 595, 139–147 (2008).
Nedeau, E. J. et al. Freshwater Mussels of the Pacific Northwest (second edition) (The Xerces Society in Portland, 2009).
Aldridge, D. C. The morphology, growth and reproduction of Unionidae (Bivalvia) in a Fenland waterway. J Mollus Stud. 65, 47–60 (1999).
Wen, H. B. Study on basic biological characteristics and metamorphosis and development of hook larva of purple black winged mussel. Nanjing Agricultural University (2016).
Fukushima, E. et al. A xenograft mantle transplantation technique for producing a novel pearl in an akoya oyster host. Mar Biotechnol. 16, 10–16 (2014).
Liu, J. et al. Antitumor activities of liposome incorporated aqueous extracts of Anodonta woodiana (Lea, 1834). Eur Food Res Technol. 227, 919–924 (2008).
Yang, J., Harino, H., Liu, H. B. & Miyazaki, N. Monitoring the organotin contamination in the Taihu Lake of China by Bivalve mussel Anodonta woodiana. B Environ Contam Tox. 81, 164–168 (2008).
Fogelman, K. J., Stoeckel, J. A., Miller, J. M. & Helms, B. S. Feeding ecology of three freshwater mussel species (Family: Unionidae) in a North American lentic system. Hydrobiologia. 850, 385–397 (2022).
Lopes-Lima, M. et al. Conservation of freshwater bivalves at the global scale: diversity, threats and research needs. Hydrobiologia. 810, 1–14 (2018).
Barnhart, M. C., Haag, W. R. & Roston, W. N. Adaptations to host infection and larval parasitism in unionoida. J N Am Benthol Soc. 27, 370–394 (2008).
International Union for Conservation of Nature (IUCN). The IUCN Ed List of Threatened Specie. Version 2023-1. https://www.iucnredlist.org.
Hu, Z. Q. Geographical distribution of endemic species of Chinese freshwater bivalves. Chin J Zool. 40, 80–83 (2005).
Ministry of Forestry and Ministry of Agriculture, China. List of National Key Wildlife Species. China (2021).
Shu, F. Y., Wang, H. J., Pan, B. Z., Liu, X. Q. & Wang, H. Z. Assessment of species status of Mollusca in the mid-lower Yangtze Lakes. Acta Hydrobiol Sinica. 33, 1051–1058 (2009).
Zhang, J. & Yu, H. X. Study on zoobenthos community structure and water quality assessment in Songhua River along Harbin city. J Aquacult. 22, 40–45 (2009).
Ma, X. Y. et al. Seasonal Variations of Nutrients and Mineral Elements in oleivora from Huaihe River. J Agron. 11, 90–94, 119. (2021).
Liu, Y.Y., Zhang, W.Z. & Wang, Y.X. Economic Fauna of China: Freshwater Mollusks (Beijing Science Press, 1979).
Wen, H. B. Study of Germplasm of Major Economic Freshwater Mollusks of China (Nanjing Agricultural University, 2009).
Han, Z. et al. Chromosome-level genome assembly of burbot (Lota lota) provides insights into the evolutionary adaptations in freshwater. Mol Ecol Resour. 21, 2022–2033 (2021).
Renaut, S. et al. Genome survey of the freshwater mussel Venustaconcha ellipsiformis (Bivalvia: Unionida) using a hybrid de novo assembly approach. Genome Biol Evol. 10, 1637–1646 (2018).
Gomes-dos-Santos, A. et al. The Crown Pearl: a draft genome assembly of the European freshwater pearl mussel Margaritifera margaritifera (Linnaeus, 1758). DNA Res. 28, dsab002 (2021).
Gomes-dos-Santos, A. et al. The Crown Pearl V2: an improved genome assembly of the European freshwater pearl mussel Margaritifera margaritifera (Linnaeus, 1758). GigaByte. 1–14 (2023a).
Gomes-dos-Santos, A. et al. PacBio Hi-Fi genome assembly of the Iberian dolphin freshwater mussel Unio delphinus Spengler, 1793. Sci Data. 10, 340 (2023b).
Rogers, R. L. et al. Gene family amplification facilitates adaptation in freshwater unionid bivalve Megalonaias nervosa. Mol Ecol. 30, 1155–1173 (2021).
Smith, C. H. A high-quality reference genome for a parasitic bivalve with doubly Uniparental Inheritance (Bivalvia: Unionida). Genome Biol Evol. 13, evab029 (2021).
Bai, Z. et al. Chromosome-level genome assembly of freshwater pearl mussel, Hyriopsis cumingii, provides insights into outstanding biomineralization ability. Authorea Preprints (2022).
Gomes-dos-Santos, A. et al. A PacBio Hi-Fi genome assembly of the Painter’s Mussel Unio pictorum (Linnaeus, 1758). Genome Biol Evol. 15, evad116 (2023c).
Belton, J. M. et al. Hi-C: a comprehensive technique to capture the conformation of genomes. Methods. 58, 268–276 (2012).
Liu, B. H. et al. Estimation of genomic characteristics by analyzing k-mer frequency in de novo genome projects. Quant Biol. 35, 62–67 (2013).
Cheng, H. Y., Concepcion, G. T., Feng, X. W., Zhang, H. W. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat Methods. 18, 170–175 (2021).
Roach, M. J., Schmidt, S. A. & Borneman, A. R. Purge Haplotigs: allelic contig reassignment for third- gen diploid genome assemblies. BMC Bioinformatics. 19, 460 (2018).
Durand, N. C. et al. Juicer provides a one-click system for analyzing LoopResolution Hi-c experiments. Cell Syst. 3, 95–98 (2016a).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science. 356, 92–95 (2017).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016b).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr Protoc Bioinformatics. Chapter 4, 4.10.11–14.10.14 (2009).
Bairoch, A. & Apweiler, R. Te swiss-prot protein sequence database and its supplement TrEMBL in 2000. Nucleic Acids Res. 28, 45–48 (2000).
Bao, W., Kojima, K. K. & Kohany, O. Repbase update, a database of repetitive elements in eukaryotic genomes. Mob DNA. 6, 11 (2015).
Benson, G. Tandem repeats finder: A program to analyze DNA sequences. Nucleic Acids Res. 27, 573–580 (1999).
Xu, Z. & Wang, H. LTR_FINDER: An efficient tool for the prediction of full-length LTR retrotransposons. Nucl Acids Res. 35, 265–268 (2007).
Mount, D. W. Using the Basic Local Alignment Search Tool (BLAST). CSH Protoc. pdb.top17 (2007).
Birney, E., Clamp, M. & Durbin, R. GeneWise and genomewise. Genome Res. 14, 988–995 (2004).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol. 29, 644–652 (2011).
Mario, S., Rasmus, S., Stephan, W. & Burkhard, M. AUGUSTUS: A web server for gene finding in eukaryotes. Nucleic Acids Res. 32, 309–312 (2004).
Burge, C. & Karlin, S. Prediction of complete gene structures in human genomic DNA. J Mol Biol. 268, 78–94 (1997).
Carson, H. & Mark, Y. MAKER2: An annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinform. 12, 491 (2011).
Manni, M., Berkeley, M. R., Mathieu, S., Simo, F. A. & Zdobnov, E. M. BUSCO update: Novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Mol Biol Evol. 38, 4647–4654 (2021).
Cantarel, B. L., Korf, I., Robb, S. M. C., Parra, G. & Ross, E. MAKER: an easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res. 18, 188–196 (2008).
Griffiths-Jones, S., Bateman, A., Marshall, M. & Khanna, A. Rfam: an RNA family database. Nucleic Acids Res. 31, 439–441 (2002).
Kozomara, A. & Griffiths-Jones, S. miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res. 39, 152–157 (2010).
Chan, P. P. & Lowe, T. M. Trnascan-SE: Searching for tRNA genes in genomic sequences. Methods Mol Biol. 1962, 1–14 (2019).
Nawrocki, E. P. & Eddy, S. R. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics. 29, 2933–2935 (2013).
Li, L., Stoeckert, C. J. & Roos, D. S. OrthoMCL: Identification of ortholog groups for eukaryotic genomes. Genome Res. 13, 2178–2189 (2003).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Wang, D., Zhang, Y., Zhang, Z., Zhu, J. & Yu, J. KaKs_Calculator 2.0: a toolkit incorporating gamma-series methods and sliding window strategies. GPB. 8, 77–80 (2010).
Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 30, 1312–1313 (2014).
Yang, Z. PAML 4: Phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007).
Kumar, S., Stecher, G., Suleski, M. & Hedges, S. B. TimeTree: a resource for timelines, timetrees, and divergence times. Mol. Biol. Evol. 34, 1812–1819 (2017).
DeBie, T., Cristianini, N., Demuth, J. P. & Hahn, M. W. CAFE: a computational tool for the study of gene family evolution. Bioinformatics. 22, 1269–1271 (2006).
Mendes, F. K., Vanderpool, D., Fulton, B. & Hahn, M. W. CAFE 5 models variation in evolutionary rates among gene families. Bioinformatics. 36, 5516–5518 (2020).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR28352171 (2024).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR26551344 (2024).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR28406055 (2024).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRR28406264 (2024).
Ma, X. Y. Solenia oleivora genome. GenBank https://identifiers.org/ncbi/insdc:JBDPLI000000000 (2024).
Ma, X. Y. Chromosomal-scale genome assembly and annotation of the Sinosolenaia oleivora. figshare https://doi.org/10.6084/m9.figshare.25458940 (2024).
Li, H. & Durbin, R. Fast and accurate short read alignment with burrows-wheeler transform. Bioinformatics. 25, 1754–1760 (2009).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics. 34, 3094–3100 (2018).
Acknowledgements
This work was supported by the Central Public-interest Scientific Institution Basal Research Fund, CAFS [2023TD66; 2023TD65]; the Sinosolenaia oleivora Ecological Compensation Program (2023); the Central Public-interest Scientific Institution Basal Research Fund [2023JBFM12] and Anhui Fuyang City Seed Breeding, Propagation and Promotion Base Construction Project. The authors would like to thank Qi Liu in Wuhan OneMore-tech Co.,Ltd. for his assistance with bioinformatic analyses.
Author information
Authors and Affiliations
Contributions
X.Y.M. and H.B.W. designed the study. W.J., D.P.X., Q.L., W.W.C., H.Z.J., Y.F.Z. and P.X. collected the sequencing samples. X.Y.M. drafted the manuscript. D.P.X. and H.B.W. contributed to the revision of the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ma, X., Jin, W., Chen, W. et al. Chromosome-level genome assembly of the freshwater mussel Sinosolenaia oleivora (Heude, 1877). Sci Data 11, 606 (2024). https://doi.org/10.1038/s41597-024-03451-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03451-5
- Springer Nature Limited