Abstract
Synthetic hexaploid wheats (SHWs) are effective genetic resources for transferring agronomically important genes from wild relatives to common wheat (Triticum aestivum L.). Dozens of reference-quality pseudomolecule assemblies of hexaploid wheat have been generated, but none is reported for SHW-derived cultivars. Here, we generated a chromosome-scale assembly for the SHW-derived cultivar ‘Chuanmai 104’ based on PacBio HiFi reads and chromosome conformation capture sequencing. The total assembly size was 14.81 Gb with a contig N50 length of 58.25 Mb. A BUSCO analysis yielded a completeness score of 99.30%. In total, repetitive elements comprised 81.36% of the genome and 122,554 high-confidence protein-coding gene models were predicted. In summary, the first chromosome-level assembly for a SHW-derived cultivar presents a promising outlook for the study and utilization of SHWs in wheat improvement, which is essential to meet the global food demand.
Similar content being viewed by others
Background & Summary
Common wheat (Triticum aestivum L., 2n = 6x = 42, AABBDD) is a natural amphiploid derived from the intergeneric cross between T. turgidum subsp. durum (Desf.) Husn., a cultivated allotetraploid (2n = 4x = 28, AABB), and Aegilops tauschii Coss., a diploid goat grass (2n = 2x = 14, DD). The genetic diversity among common wheat cultivars has been drastically reduced owing to bottlenecks resulting from polyploidy, domestication, and modern plant breeding. The decline in genetic diversity can be counteracted by direct hybridization between common wheat and A. tauschii1,2, or through hybridization between common wheat and synthetic hexaploid wheats (SHWs) developed by crossing tetraploid wheat and A. tauschii3.
The primary objective of the direct hybridization method is to augment genetic diversity specifically for the D genome in common wheat, addressing a crucial concern in wheat breeding, because significantly lower genetic diversity values characterize this genome compared with the A and B genomes4. However, the diminished genetic diversity resulting from the bottlenecks also affects the A and B genomes. Consequently, the utilization of SHW lines enables the diversity of all three subgenomes of common wheat to be enhanced. This approach facilitates the direct transfer of genes/loci for traits of interest from diploid and tetraploid to hexaploid wheat.
To date, the International Maize and Wheat Improvement Centre (CIMMYT) has developed more than 1200 SHW lines3. Since the introduction of more than 200 SHW accessions from CIMMYT in 1995, four SHW-derived cultivars, namely, Chuanmai 38, 42, 43, and 47, have been raised and cultivated, which have been widely used in wheat breeding as elite parents in China. Subsequently, a number of secondary SHW-derived cultivars have been developed and released, including Chuanmai 104, developed from the cross of Chuanmai 42 and Chuannong 16. Chuanmai 104 is an important high-yielding wheat cultivar grown in Southwest China in recent years. The maximum yield of Chuanmai 104 attains 10,947 kg/ha under the humid and predominantly cloudy climate of the Sichuan Basin in Southwest China5. Chuanmai 104 is becoming a cornerstone breeding parent of wheat in China. Furthermore, China is among the main countries that are exploiting the advantages of SHW lines as genetic resources, especially in Southwest China3. The increasing utilization of SHW worldwide is indicative of the success of such an approach, which will gradually become an effective means of overcoming the bottleneck of wheat breeding.
Considering previous studies based on SHWs, a major potential limiting factor is the limited genetic resources and lack of reference-quality pseudomolecule assemblies (RQAs)6. Chapman et al. integrated whole-genome sequencing and genetic mapping to assemble and ordered contigs of the SHW cultivar W79847. However, given the short reads generated by next-generation sequencing (NGS), and the lack of chromosome conformation capture sequencing or chromosome isolation via flow sorting, the assembly was only 9.1 Gb, which was substantially less than the estimated 15 Gb size of the hexaploid wheat genome6. Although single-nucleotide polymorphism (SNP) genotyping arrays are relatively simple and inexpensive, a limitation is that only the variants pre-selected for inclusion on the array can be analyzed. Consequently, if the SNP panels were designed using common wheat genome assemblies, they would lack sufficient representation of variants in the target gene pools, and thus assessment of useful variation in SHW and derivative germplasm would be challenging. More recently, reduction in costs have meant that RQAs and large-scale whole-genome resequencing are feasible and affordable for SHWs.
In the current study, we first generated a chromosome-level assembly for Chuanmai 104 (Fig. 1), based on an integrated approach including PacBio HiFi sequencing reads and chromosome conformation capture sequencing. The final Chuanmai 104 genome assembly consisted of 14.81 Gb with a contig N50 of 58.25 Mb, a contig N90 of 8.41 Mb, and a longest contig of 422.27 Mb (Table 1). Among previously published hexaploidy wheat assemblies, seven of the 21 chromosomes in the Chuanmai 104 were the longest (Table 2). The long terminal repeat (LTR) Assembly Index (LAI)8 of the Chuanmai 104 genome assembly was 15.17, 14.64, and 10.85 for A subgenome, B subgenome, D subgenome respectively, and for each chromosome, the LAI values ranges from 10.21 to 15.71 (Table 3). Benchmarking universal single-copy orthologs (BUSCO) analysis yielded a completeness score of 99.30%, which was comparable with that of common wheat genomes and notably higher than that of the SHW cultivar W7984 (Table 1). Repeats comprised 81.36% of the sequences with a predominance of retrotransposons, which accounted for 62.96% of the sequences (Table 4). In total, 122,554 high-confidence and 136,431 low-confidence protein-coding gene models were predicted (Table 5); this number was similar to that for the common wheat Chinese Spring (Table 1). The high-quality Chuanmai 104 genome assembly generated in this study provides a reference genome for SHW-derived cultivars, and offers a promising outlook for the study and utilization of SHW genetic resources in wheat improvement, which is essential to meet the global food demand.
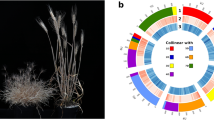
Overview of Chuanmai 104 chromosome-scale assembly. (a) Distribution of the A. tauschii clone A6-10 subtelomeric tandem repeat sequence (GenBank Accession AY249980.1). (b) Distribution of the A. tauschii clone 6C6-3 (GenBank Accession AY249981.1) and 6C6-4 (GenBank Accession AY249982.1) and T. monococcum ssp. aegilopoides clone BAC TbBAC5 (GenBank Accession DQ904440.1) and TbBAC30 (GenBank Accession EF624064.1) centromere-specific tandem repeat sequences. (c) Distribution of the non-coding gene density. (d) Distribution of the transposable elements’ density. (e) Distribution of the tandem repeat density. (f) Distribution of the long terminal repeat density. (g) Distribution of the high-confidence protein-coding gene density. (h) Distribution of the significant enrichment of subgenome-specific k-mers identified by SubPhaser (gold colour for A, blue for B, and orange for D). (i) density distribution of the D subgenome-specific k-mer set. (j) density distribution of the B subgenome-specific k-mer set. (k) density distribution of the A subgenome-specific k-mer set. Links between chromosomes are collinearity blocks, which are coloured according to the homologous chromosomes. All the densities were calculated using sliding windows (window size: 1Mbp, step size: 1Mbp), except the density distribution of the non-coding genes, which use a window size of 10Mbp and a step size of 1Mbp for smoother visualization.
Methods
Plant material, DNA extraction, and sequencing
The SHW-derived cultivar Chuanmai 104 was kindly provided by Wuyun Yang (Crop Research Institute, Sichuan Academy of Agricultural Sciences). The plants used for sequencing were grown in a growth chamber with a controlled environment of 20 degree Celsius under a 12 h light/12 h dark photoperiod for 2 weeks. Genomic DNA (gDNA) was extracted from seedling leaf tissues using the cetyltrimethylammonium bromide method. Three methods were applied for DNA quantification and quality testing, including (i) NanoDrop 2000 spectrophotometer (Thermo Fischer Scientific), (ii) gel electrophoresis and (iii) Qubit fluorometer (Invitrogen). Total DNA was purified by AMPure PB beads (Pacific Biosciences, CA, USA; PN 100-265-900). High-quality gDNA (≥10 μg, ≥100 ng/μl) was prepared for the next step of library construction. PacBio single-molecule real-time (SMRT) bell library preparation was performed using the SMRTbell® Express Template Prep Kit 2.0 (Pacific Biosciences, CA, USA; PN 101-853-100) in accordance with the manufacturer’s instructions. The library was prepared for sequencing with a 30 h movie on the Sequel IIe system (Pacific Biosciences) by the Berry Genomics Corporation (Beijing, China). Totally, we generated 668.43 Gb bases (~45X) with 40,999,150 CCS reads from 20 SMRT cells.
Chromosome conformation capture (Hi-C) sequencing of Chuanmai 104 was performed using the protocol of Peng et al.9. In brief, 2–4 g tender leaves from the plants used for genome sequencing were harvested and stored in liquid nitrogen, and then the Hi-C libraries were prepared and sequenced on the MGISEQ-2000 platform by BGI (Wuhan, China). Samples were cut into pieces of ca. 2 cm2, and transferred to 50 ml tubes containing 15 ml of ice-cold nuclear isolation buffer (NBE) with 2% formaldehyde, followed by vacuum infiltration (400 mbar) and incubation with a supplemented cross-linking agent for 1 h. Cross-linking was quenched by adding 2 M glycine to a final concentration of 0.125 M with incubation for 5 min under vacuum, followed by fixation on ice. Then, the fixed leaf pieces were washed three times with sterile Milli-Q water, ground in liquid nitrogen and subjected to nucleus isolation. The isolated nuclei were purified, checked for quality and quantity and digested with 100 units of DpnII. The next steps were Hi-C specific, including marking the DNA ends with biotin-14-dATP and performing blunt-end ligation of the cross-linked fragments. After ligation, cross-linking was reversed by overnight incubation with proteinase K at 65 °C. Biotin-14-dATP was further removed from non-ligated DNA ends using the exonuclease activity of T4 DNA polymerase. DNA was purified by phenol:chloroform (1:1) extraction, precipitated and washed as previously described. The purified DNA was physically sheared to a size of 300–600 bp by sonication and was size-fractionated using standard 2% agarose gel electrophoresis to obtain fragments in the range of 300–600 bp. The fragmented ends were blunt-end repaired and A-tailed, followed by purification through biotin-streptavidin-mediated pulldown. PCR amplification was conducted using 12–15 cycles to enrich the ligation products. Totally, we generated more than 2 Tb bases (>135 X) with 6.69 Gb read pairs.
For full-length transcriptome sequencing, we collected pooled sample for Chuanmai 104, which comprised whole plant organs except for roots from seed germination to the three-leaf stage, shoots at the seedling stage, and leaves, stems, ears, and seeds from the heading to the late-filling stages. Total RNA was isolated using TRIzol Reagent in accordance with the manufacturer’s instructions (Thermofisher). The RNA purity and raw contamination were first assed by Nanodrop 2000 (Thermo Fischer Scientific), and then the RNA Integrity Number (RIN) and concentration were further assessed by an Agilent 4200 (Agilent Technologies). High-quality RNA (2 μg, 300 ng/μl) was prepared for the next step of library construction. PacBio SMRT bell library preparation was performed using the SMRTbell® Express Template Prep Kit 2.0 (Pacific Biosciences) in accordance with the manufacturer’s instructions. The library was prepared for sequencing with a 30 h movie on the Sequel IIe system (Pacific Biosciences) by the Berry Genomics Corporation. Totally, we generated 186.35 Gb bases with 2,283,790 polymerase reads from one SMRT cell. The final 46,130,981 subreads range from 51 bp to 241,082 bp, with a mean and N50 value of 4,039.55 bp and 4,561 bp respectively.
Genome assembly
The PacBio HiFi CCS reads were assembled using hifiasm10 (v0.16.1, with default parameters). The Hi-C reads were incorporated using Juicer tools11 (v1.6) and EndHiC12. In brief, preprocessing of the Hi-C reads was performed with juicer.sh11 (parameter: -s DpnII). The output file corresponding to the Hi-C contacts with duplicates removed and mapping quality values larger than 30 was generated as input for EndHiC12. These result files were plotted to visualize the Hi-C map and for manual curation, and were used to generate the final assembly (21 pseudomolecules and one unanchored pseudomolecule). The NCBI Foreign Contamination Screen (FCS)13 was used to identify and remove contaminant sequences (adaptors and organelles) in genome assemblies. Totally, the FCS identified total 754 contaminant fragments, including one adaptor fragment and 753 mitochondrial fragments, and all these contaminants are located on the unanchored pseudomolecule and were masked.
Validation of genome assemblies
Genome sizes were estimated using three algorithms (gce14, GenomeScope215, and findGSE16) with different k-mer sizes. The quality and completeness of the genome assemblies were assessed by merqury17, which uses a reference-free, k-mer-based approach, and BUSCO18 (v5, poales_odb10), which is based on evolutionarily informed expectations of the near-universal single-copy orthologous gene content. LTR assembly index (LAI)8 that evaluates assembly continuity using LTR-RTs were calculated.
Subgenome assignment, validation, and nomenclature
To assign each chromosome to each linkage group and apply the corresponding nomenclature in Chinese Spring, SubPhaser19, a robust allopolyploid subgenome phasing method based on subgenome-specific k-mers, was used. To validate the correctness of the subgenome assignment, a reference-guided strategy based on subgenome homology was also used to distinguish the subgenomes. We mapped the Chuanmai 104 genome to the Chinese Spring genome using mashmap20 (-f map–perc_identity 90 -s 1000000). Then, the alignments were plotted and manually checked. This procedure successfully categorized the 21 chromosomes into three homologous groups. The nomenclature system for Chinese Spring chromosomes was adopted for naming of the homologous groups (1–7) of the Chuanmai 104 genome.
Repeat annotation
Tandem repeats of all lengths were annotated with TandemRepeatsFinder21 v4.09 using the default parameters (Match Mismatch Delta PM PI Minscore MaxPeriod: 2 7 7 80 10 50 500). LTR_FINDER22 (v1.05) and LTR_harvest23 (v1.5.10) were used for long terminal repeat (LTR) identification, and the results were processed with LTR_retriever24 (v2.8) to generate a species-specific LTR library. The species-specific LTR libraries, wheat transposable element (TE) sequences from ClariTeRep (https://github.com/jdaron/CLARI-TE), and plant TE sequences from Repbase25 were merged to generate the TE library. Transposons were detected and classified by a homology search against the combined TE library. The program Vmatch (http://www.vmatch.de/), a fast and efficient matching tool suitable for large and highly repetitive genomes, was used for this computationally intensive task with the following parameters: identity ≥ 70%, minimal hit length 75 bp, and seed length 12 bp (exact command line: -d -p -l 75 -identity 70 -seedlength 12 -exdrop 5).
Non-coding gene annotation
Noncoding RNAs (ncRNAs), including miRNAs, small nuclear RNAs, rRNAs and regulatory elements, were identified using the Infernal26 (version 1.1.2) program to search against the Rfam27 database (v14.8). The rRNAs, and tRNAs were further identified using RNAmmer28 (version 1.2) and tRNAscan-SE29 (v1.3.1) respectively.
Protein-coding gene annotation
Gene model prediction was performed following the method described by Mascher et al.30, with minor modifications, which integrated transcriptomic data, protein homology, and ab initio prediction. (1) First, isoform sequencing (Iso-Seq) data were mapped to the genome using minimap231 (v2.17-r941; parameters: -ax splice -uf –secondary = no -C5). The redundant isoforms were further collapsed into transcript loci using cDNA_Cupcake (https://github.com/Magdoll/cDNA_Cupcake) (parameter: –dun-merge-5-shorter). TransDecoder (v5.5.0, https://github.com/TransDecoder/TransDecoder) was used to predict protein sequences among the transcripts. (2) For protein homology evidence, we projected the gene structures from Triticeae species, comprising Ae. tauschii, T. turgidum subsp. dicoccoides, T. turgidum subsp. durum, T. aestivum Chinese Spring, T. urartu, Ae. speltoides, and Hordeum vulgare Morex, onto the Chuanmai 104 genome using liftoff32 with default parameters. (3) We produced ab initio gene predictions using AUGUSTUS33 (v3.4.0), GeneMark-ET34 (v4.38), and GeneID35 (v1.4). In brief, AUGUSTUS33 gene prediction was performed using a model specifically trained from the software and a hints file generated using the previously mentioned Iso-Seq predictions. GeneMark-ET34 was used with the option -ET, and the intron coordinates were calculated using the above-mentioned Iso-Seq alignments. GeneID35 was run with a model specifically trained from the software (-GP taestivum.param). We used EVidenceModeler36 (EVM; v1.1.1) to integrate all of the gene evidence from transcriptomics, protein alignments, and ab initio predictions.
Protein-coding gene models from EVM were classified as high-confidence or low-confidence according to criteria used by the International Wheat Genome Sequencing Consortium, with minor modifications37. In brief, protein-coding gene models were considered as ‘complete’ when start and stop codons were present. A comparison with PTREP38 (the database of hypothetical proteins deduced from the nonredundant database of TEs within the TREP database), UniPoa39 (Poaceae database of annotated proteins from the UniProt database), and UniMag40 (validated Magnoliophyta proteins from SwissProt) was performed using DIAMOND41 (v2.0.9; parameters: -e 1e-10 –query-cover 80–subject-cover 80). Gene candidates were classified using the following criteria: a high-confidence gene model was ‘complete’ with a hit in the UniMag40 database and/or in UniPoa39 but not PTREP38; the remaining gene models were classified as low-confidence genes.
Functional assignments of the predicted protein-coding genes were obtained with BLAST42 by aligning the coding regions to sequences in public protein databases, including the trEMBL40, RefSeq43., and SwissProt40 databases. The putative domains and GO44 terms of the predicted proteins were identified using the InterProScan45 program. The putative orthologs in the KEGG46 database were identified using KoFamScan47.
Data Records
The HiFi reads, Iso-seq reads, and Hi-C reads that were used for the Chuanmai 104 genome assembly have been deposited in the NCBI Sequence Read Archive with accession number SRP488123 and under BioProject number PRJNA107040948. The HiFi reads, Iso-seq reads, and Hi-C reads were also deposited in the National Genomics Data Centre (NGDC) with BioProject ID PRJCA022052 (https://ngdc.cncb.ac.cn/bioproject/browse/PRJCA022052). The genome assembly has been deposited at GenBank under the accession JBBIFV00000000049. The genome assemblies and annotations have also been deposited at FigShare50 with doi number https://doi.org/10.6084/m9.figshare.25282654.
Technical Validation
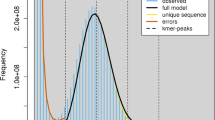
The assembled genome size is similar to the size estimated by different algorithms14,15,16 (Fig. 2a–c), and is significantly larger than that published previously for the SHW cultivar W7984 (Table 1). The base-level accuracy QV (consensus quality value) and k-mer completeness scores evaluated with merqury17 are 65.86 and 97.59%, respectively. The long terminal repeat (LTR) Assembly Index (LAI)8 of the Chuanmai 104 genome assembly was 15.17, 14.64, and 10.85 for A subgenome, B subgenome, D subgenome respectively, which are higher than the LAI values obtained for Chinese Spring (11.88, 12.51 and 9.97 for A subgenome, B subgenome, D subgenome respectively). The BUSCO18 score is 99.3% and only 0.7% BUSCO genes are missing (Fig. 2d). These results indicate a high completeness of the Chuanmai 104 assembly. Comparison with other common wheat genome assemblies revealed that the Chuanmai 104 NG50 value was significantly larger, implying high connectivity (Fig. 2e). The GC-depth plot (Fig. 2f) of the Chuanmai 104 genome across every 2 kb nonoverlapping sliding window showed no distinct secondary peaks, indicating that haplotype homology was adequately recognized during assembly.
Validations of Chuanmai 104 genome assemblies. (a) Genome sizes estimated using different algorithms with different K-mer sizes. (b,c) examples of genome size estimated by findGCE (K = 181, b) and GenomeScope2 (K = 181, c) respectively. (d) Gene completeness assessed by BUSCO using the Poales dataset with a total of 4896 groups. (e) NGx plots for the Chuanmai 104 and other common wheat genomes. (f) GC content and average sequencing depth (GC-depth) plot of the Chuanmai 104 genome across every 2-kb nonoverlapping sliding window.
The Hi-C contact map was manually curated and assessed with Juicebox and revealed a dense pattern along the diagonal, indicating no potential mis-assemblies (Fig. 3). The anti-diagonals are typical for Triticeae genomes51 (Fig. 3). The distribution of the A. tauschii subtelomeric tandem repeat sequences (NCBI GenBank accessions: AY249980.1, AY249981.1, and AY249982.1) and T. monococcum subsp. aegilopoides centromere-specific tandem repeat sequences (NCBI GenBank accessions: DQ904440.1 and EF624064.1) indicate the completeness in these complex regions (Fig. 1a–c).
Using SubPhaser19, a robust allopolyploid subgenome phasing method based on subgenome-specific k-mers, the 21 chromosomes of the Chuanmai 104 genome were aggregated into three linkage groups (Fig. 1h–k). These groups show high synteny to chromosomes of Chinese Spring at both the nucleotide and protein levels (Fig. 4), indicating the correctness of the chromosome assembly. Moreover, these synteny results show the relative conservation of the common wheat and SHW genomes, although the sources of the subgenomes and their evolutionary history differ.
Code availability
All software and pipelines were executed according to the manual and protocol of published tools. No custom code was generated for these analyses.
References
Gill, B. S. et al. Wheat Genetics Resource Center: The First 25 Years. in Advances in Agronomy vol. 89 73–136 (Academic Press, 2006).
Gill, B. S. & Raupp, W. J. Direct Genetic Transfers from Aegilops squarrosa L. to Hexaploid Wheat1. Crop Science 27, cropsci1987.0011183X002700030004x (1987).
Li, A., Liu, D., Yang, W., Kishii, M. & Mao, L. Synthetic Hexaploid Wheat: Yesterday, Today, and Tomorrow. Engineering 4, 552–558 (2018).
Mazzucotelli, E. et al. Gene Flow Between Tetraploid and Hexaploid Wheat for Breeding Innovation. in The Wheat Genome (eds. Appels, R., Eversole, K., Feuillet, C. & Gallagher, D.) 135–163. https://doi.org/10.1007/978-3-031-38294-9_8 (Springer International Publishing, Cham, 2024).
Li, J., Wan, H.-S. & Yang, W.-Y. Synthetic hexaploid wheat enhances variation and adaptive evolution of bread wheat in breeding processes. Journal of Systematics and Evolution 52, 735–742 (2014).
Walkowiak, S. et al. Multiple wheat genomes reveal global variation in modern breeding. Nature 588, 277–283 (2020).
Chapman, J. A. et al. A whole-genome shotgun approach for assembling and anchoring the hexaploid bread wheat genome. Genome Biology 16, 26 (2015).
Ou, S., Chen, J. & Jiang, N. Assessing genome assembly quality using the LTR Assembly Index (LAI). Nucleic Acids Res 46, e126 (2018).
Peng, Y. et al. Reference genome assemblies reveal the origin and evolution of allohexaploid oat. Nat Genet 54, 1248–1258 (2022).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat Methods 18, 170–175 (2021).
Durand, N. C. et al. Juicer provides a one-click system for analyzing loop-resolution Hi-C experiments. Cell Syst 3, 95–98 (2016).
Wang, S. et al. EndHiC: assemble large contigs into chromosome-level scaffolds using the Hi-C links from contig ends. BMC Bioinformatics 23, 528 (2022).
Astashyn, A. et al. Rapid and sensitive detection of genome contamination at scale with FCS-GX. Genome Biology 25, 60 (2024).
Liu, B. et al. Estimation of genomic characteristics by analyzing k-mer frequency in de novo genome projects. Preprint at https://www.semanticscholar.org/paper/Estimation-of-genomic-characteristics-by-analyzing-Liu-Shi/716199abb13c0cab875f3dfe6302cce857685385 (2013).
Ranallo-Benavidez, T. R., Jaron, K. S. & Schatz, M. C. GenomeScope 2.0 and Smudgeplot for reference-free profiling of polyploid genomes. Nat Commun 11, 1432 (2020).
Sun, H., Ding, J., Piednoël, M. & Schneeberger, K. findGSE: estimating genome size variation within human and Arabidopsis using k-mer frequencies. Bioinformatics 34, 550–557 (2018).
Rhie, A., Walenz, B. P., Koren, S. & Phillippy, A. M. Merqury: reference-free quality, completeness, and phasing assessment for genome assemblies. Genome Biology 21, 245 (2020).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Jia, K.-H. et al. SubPhaser: a robust allopolyploid subgenome phasing method based on subgenome-specific k-mers. New Phytologist 235, 801–809 (2022).
Jain, C., Koren, S., Dilthey, A., Phillippy, A. M. & Aluru, S. A fast adaptive algorithm for computing whole-genome homology maps. Bioinformatics 34, i748–i756 (2018).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res. 27, 573 (1999).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 35, W265–W268 (2007).
Ellinghaus, D., Kurtz, S. & Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinformatics 9, 18 (2008).
Ou, S. & Jiang, N. LTR_retriever: A Highly Accurate and Sensitive Program for Identification of Long Terminal Repeat Retrotransposons. Plant Physiol 176, 1410–1422 (2018).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467 (2005).
Nawrocki, E. P., Kolbe, D. L. & Eddy, S. R. Infernal 1.0: inference of RNA alignments. Bioinformatics 25, 1335–1337 (2009).
Griffiths-Jones, S. et al. Rfam: annotating non-coding RNAs in complete genomes. Nucleic Acids Res 33, D121–124 (2005).
Lagesen, K. et al. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 35, 3100–3108 (2007).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 25, 955–964 (1997).
Mascher, M. et al. Long-read sequence assembly: a technical evaluation in barley. The Plant Cell 33, 1888–1906 (2021).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Shumate, A. & Salzberg, S. L. Liftoff: accurate mapping of gene annotations. Bioinformatics 37, 1639–1643 (2021).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Res. 34, W435–W439 (2006).
Besemer, J. & Borodovsky, M. GeneMark: web software for gene finding in prokaryotes, eukaryotes and viruses. Nucleic Acids Res 33, W451–454 (2005).
Blanco, E., Parra, G. & Guigó, R. Using geneid to identify genes. Curr Protoc Bioinformatics Chapter 4, Unit 4.3 (2007).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biology 9, R7 (2008).
The International Wheat Genome Sequencing Consortium (IWGSC). et al. Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361, eaar7191 (2018).
The TREP platform: A curated database of transposable elements. https://trep-db.uzh.ch/.
UniProt. https://www.uniprot.org/.
Boeckmann, B. et al. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Res. 31, 365–370 (2003).
Buchfink, B., Reuter, K. & Drost, H.-G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat Methods 18, 366–368 (2021).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Pruitt, K. D. et al. RefSeq: an update on mammalian reference sequences. Nucleic Acids Res. 42, D756–D763 (2014).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Quevillon, E. et al. InterProScan: protein domains identifier. Nucleic Acids Res 33, W116–120 (2005).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 40, D109–D114 (2012).
Aramaki, T. et al. KofamKOALA: KEGG Ortholog assignment based on profile HMM and adaptive score threshold. Bioinformatics 36, 2251–2252 (2020).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP488123 (2024).
NCBI Assembly https://identifiers.org/ncbi/insdc.gca:GCA_039655515.1 (2024).
Liu, Z. Genome assembly and annotation of the synthetic hexaploid wheat-derived cultivar Chuanmai 104. figshare https://doi.org/10.6084/m9.figshare.25282654 (2024).
Abrouk, M. et al. Chromosome-scale assembly of the wild wheat relative Aegilops umbellulata. Sci Data 10, 739 (2023).
Acknowledgements
This publication is based on work supported by the Accurate Identification Project of Crop Germplasm from Sichuan Provincial Finance Department (1 + 3 ZYGG001), the Science and Technology Department of Sichuan Province (2022NSFSC0601, 2021YFYZ0020, 2023NSFSC1925), the Program of Chinese Agriculture Research System (CARS-03), Strategic Scientist Studio of Sichuan Academy of Agricultural Sciences. We thank Robert McKenzie, PhD, from Liwen Bianji (Edanz) (www.liwenbianji.cn) for editing a draft of this manuscript.
Author information
Authors and Affiliations
Contributions
W.Y. and J.L. designed the study and provided funding. Z.L., F.Y. and C.D. collected and prepared the plant tissues samples for sequencing. Z.L., F.Y. and C.D. led the bioinformatics analyses and genome analyses. Z.L. and C.D. drafted the manuscript. W. Y., J. L., F.Y., H.W. H.T. J.F., Q.W. N.Y. revised the manuscript. Z.L., F.Y. and C.D. contributed equally. All authors discussed the results and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, Z., Yang, F., Deng, C. et al. Chromosome-level assembly of the synthetic hexaploid wheat-derived cultivar Chuanmai 104. Sci Data 11, 670 (2024). https://doi.org/10.1038/s41597-024-03527-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03527-2
- Springer Nature Limited