Abstract
Astrocytic tumours are the most common type of primary malignant brain tumour. Most astrocytic tumours will recur at some point after surgery. Currently, the combination of radiotherapy and chemotherapy does not prevent the recurrence of astrocytic tumours. In this study, we investigated the consistency in isocitrate dehydrogenase 1 (IDH1), tumour protein p53 (TP53) and telomerase reverse transcriptase promoter (TERTp) mutations during astrocytic tumour recurrence. We also evaluated the protein loss of O-6-methylguanine-DNA methyltransferase (MGMT) and alpha-thalassemia/mental retardation, X-linked (ATRX) during disease recurrence. We then determined the prognostic significance of these findings in terms of progression-free survival (PFS) using Kaplan-Meier analysis and Cox regression models. Our results showed that in most cases, IDH1, TP53 and TERTp mutation status and MGMT and ATRX protein expression levels were stable during recurrence, which may indicate that these alterations occurred early in astrocytic tumour development. Furthermore, in IDH1 wild type group, the patients who were negative for MGMT and had a low Ki67 index showed a longer PFS. Therefore, we suggest that IDH1 mutation combined with MGMT expression level and Ki67 index might be an effective biomarker panel for evaluating the PFS of patients with astrocytic tumours.
Similar content being viewed by others
Introduction
Astrocytic tumours are the most common primary malignant tumours in the central nervous system1. The World Health Organization (WHO) classification system divides astrocytic tumours into four levels (I, II, III and IV) according to biological behaviour2. The majority of astrocytic tumours, except grade I astrocytomas, will recur at some point after surgery, and some recurrent cases evolve from a lower grade tumour (grade II or III) to a higher grade tumour (grade III or IV)3,4. At present, the prognosis of high-grade tumours is poor; for instance, the 5-year survival rate of patients with glioblastoma is less than 5%, and the average survival time is only 14 months5. To date, there is no effective way to prevent the recurrence of astrocytic tumours and no effective treatment for recurrent disease6. Recurrence is one of the most important reasons for the poor outcomes associated with astrocytic tumours.
In the last few years, considerable advances have been made in the sequencing of astrocytic tumours, which led to the discovery of key genetic alterations in gliomas, including mutations in the isocitrate dehydrogenase 1/2 (IDH1/2), homologue of Drosophila capicua (CIC), far-upstream binding protein 1 (FUBP1), and alpha-thalassemia/mental retardation, X-linked (ATRX) genes and in the TERT promoter (TERTp) region7,8,9,10. These biological molecules are strongly correlated with the traditional tumour classification and prognosis10,11,12. In 2012, Jiao et al.12 introduced a new way to classify gliomas and evaluate prognosis by integrating ATRX, CIC, FUBP1 and IDH1 mutations into the classification. The resulting three categories were as follows: I-A (IDH1/ATRX mutation), I-CF (IDH1/CIC/FUBP1 mutation), and I-X gliomas (not I-A or I-CF). The median overall survival of patients with I-CF gliomas (96 months) was longer compared to that of patients with I-A gliomas (51 months) or I-X gliomas (13 months)12. In the following years, several studies explored the classification of gliomas by integrating TERTp, IDH1/2, TP53, and ATRX mutations and 1p/19q co-deletions13,14,15,16,17,18. Therefore, the above biomarkers became important auxiliary tools for the diagnosis, prognosis and treatment of gliomas. At present, the WHO classification system includes these molecular parameters, in addition to histology, to distinguish among the different types of glioma18, opening the way to the era of molecular diagnosis of gliomas.
In terms of the abovementioned molecular parameters, key mutations in astrocytic tumours include classic mutations in the IDH1 and TP53 genes and mutations affecting telomere length in the ATRX gene and TERT promoter region. MGMT promoter hyper-methylation is another important epigenetic marker that might result in gene expression silencing and further impair the ability to repair DNA. Patients with astrocytic tumours might benefit from temozolomide because of the presence of MGMT gene silencing19. Previous studies on these gene alterations, especially those of TERTp and the ATRX gene, have mostly focused on primary tumours, with little investigation of paired recurrent tumours. Furthermore, studies on the function of IDH1, TP53, ATRX and TERTp as prognostic biomarkers for the progression-free survival (PFS) of patients with astrocytic tumours are controversial11,19,20,21,22,23,24,25,26,27,28,29. Therefore, the aim of this study was to evaluate the consistency of key genetic alterations in IDH1, TP53 and TERTp mutation status and ATRX and MGMT protein expression level in the course of astrocytic tumour recurrence; furthermore, the prognostic significance of these findings for PFS was discussed.
Results
Frequency of IDH1, TP53, and TERTp mutation and loss of MGMT and ATRX in the evolution of astrocytic tumours
We compared sequence alterations in IDH1, TP53 and TERTp and changes in the protein level of MGMT and ATRX in 47 primary astrocytic tumours and 48 corresponding recurrences. In the primary tumours, the frequencies of IDH1 and TP53 mutations were higher in astrocytoma (A) and anaplastic astrocytoma (AA) than in pGBM (primary glioblastoma). In contrast, TERTp mutations were rarer in A and AA than in pGBM. In recurrent tumours, the frequencies of IDH1 and TP53 mutations were higher in A, AA and secondary glioblastoma (sGBM) than in rGBM (recurrence of pGBM), and the frequency of TERTp mutations was lower in A, AA and sGBM than in rGBM. Regarding protein loss (negative immunohistochemical staining results), ATRX loss in primary tumours was more common in A and AA than in pGBM, whereas MGMT loss was not significantly different among all tumour grades. In recurrent tumours, ATRX loss was more common in A, AA and sGBM than in rGBM; MGMT loss was not significantly different in A, AA, sGBM and rGBM. The TERTp mutation pattern showed a remarkable inverse correlation with IDH1 and TP53 mutations and ATRX protein loss. The specific genetic mutation frequencies and protein loss frequencies in primary and recurrent tumours are shown in Table 1.
IDH1, TP53 and TERTp mutation sites in astrocytic tumours
In primary tumours, 21 (21/47, 44.68%) IDH1 mutations were detected, all of which were the R132H mutation (CGT-CAT). In addition, 19 TP53 mutations were found in 17 (36.17%, 17/47) primary samples (Fig. 1a); these mutations were missense or truncation mutations and involved 8 previously identified loci30,31,32,33,34,35: R175H (CGC-CAC), H193R (CAT-CGT), I195T (ATC-ACC), V216M (GTG-ATG), R273C (CGT-TGT), R273H (CGT-CAT), C275F (TGT-TTT) and R306 (CGA-TGA). Among the 17 samples in which TP53 mutations were found, two contained two mutations each: R175H and R273C (No. 2) and I195T and R273C (No. 39). Fifteen (15/47, 31.91%) TERTp mutations were found in primary tumour tissue, including 12 cases of C228T (−124 bp) and 3 cases of C250T (−146 bp).
Mapview of TP53 gene mutations in 47 astrocytic tumours (drawn online using MutationMapper, http://www.cbioportal.org/mutation_mapper.jsp). The relationship between mutation type, mutation site and domains of the TP53 gene in primary astrocytic tumours (a) and recurrent astrocytic tumours (b). The sites with parentheses indicate those with a recurring mutation, and the number in brackets represents the mutation frequency of the site. AD (activation domain), DBD (DNA binding domain), TD (tetramerization domain).
In recurrent tumours, all 22 (22/48, 45.83%) of the detected IDH1 mutations were R132H (CGT-CAT). Eighteen TP53 mutations were detected in 17 (35.42%, 17/48) recurrent tumours; again, these mutations were missense or truncation mutations. In addition, two other previously reported mutation sites were detected: P177R (CCC-CGC)36 and R273Y (CGT-TAT)24 (Fig. 1b). Furthermore, patient No. 39 lost the mutation at codon 273 H, and only the I195T mutation was found in the recurrent tumour. In recurrent tumour tissue, 16 (16/48, 33.33%) TERTp mutations were found, including 14 cases of C228T (−124 bp) and 2 cases of C250T (−146 bp).
IDH1, TP53 and TERTp mutation status and MGMT and ATRX protein expression levels were consistent in primary and recurrent gliomas
The IDH1, TP53 and TERTp status in primary and corresponding recurrent tumours was analysed by McNemar’s test, and no significant differences were found (all p values were approximately equal to 1, Table 2). Furthermore, the consistency analysis results showed that all kappa values were quite high, and the recurrent tumours had consistent IDH1, TP53 and TERTp status with their matched primary tumours (κ = 1.00, κ = 0.724, κ = 0.856; Table 2). All three investigated biomarkers were consistent in primary tumours and matched recurrences in 38 cases. The other 9 cases showed inconsistencies in the TP53 or TERTp biomarker (Fig. 2). TP53 status changed in 6 cases, among which 3 cases gained TP53 mutations upon recurrence. In the other 3 cases, the TP53 mutations detected in the primary tumours were not found in the recurrent tumours. TERTp status of the recurrent tumour was different from that of the primary tumour in 3 cases. One primary tumour was TERTp mutant, while the matched recurrent tumour was wild type. In contrast, the other 2 primary tumours were TERTp wild type, while TERTp mutations were found in the matched recurrent tumours (Fig. 2).
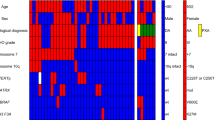
Mutation type in IDH1, TP53 and TERTp and protein expression level of MGMT and ATRX in 47 paired cases of astrocytic tumours (drawn online using OncoPrinter, http://www.cbioportal.org/oncoprinter.jsp). P (primary astrocytic tumours), R (recurrent astrocytic tumours).
The McNemar’s test did not show significant differences in ATRX and MGMT status between primary tumours and recurrences (all p values were approximately equal to 1, Table 2). The agreement test showed that ATRX and MGMT status in recurrent tumours and their primary counterparts was moderately consistent (κ = 0.452; κ = 0.487, Table 2). Neither ATRX nor MGMT protein expression changed in 31 cases of recurrent tumours. In the remaining 16 cases, 10 showed a change in ATRX or MGMT protein expression level, and 6 showed a change in both protein biomarkers (Fig. 2).
In all, the five studied biomarkers did not change in 25 cases upon recurrence (Fig. 2, Supplementary Table S1). The other 22 cases showed changes in one or two biomarkers upon comparison of recurrent tumours and primary tumours (Fig. 2, Supplementary Table S1). Clinicopathological features of primary and recurrent tumours and median PFS were not significantly different between the two groups (Supplementary Table S2).
Prognostic value of IDH1, TP53 and TERTp mutation and MGMT and ATRX expression level in astrocytic tumours
The value of IDH1, TP53, and TERTp mutations and MGMT and ATRX protein expression level in primary tumours for evaluating the PFS of patients with astrocytic tumours was analysed using the Kaplan-Meier survival estimator. PFS was significantly longer in patients harbouring IDH1 mutations and in MGMT-negative or ATRX-negative patients (p = 0.000069, Fig. 3a; p = 0.042, Fig. 3c; p = 0.004, Fig. 3d; Table 3). In contrast, TERTp mutation was associated with a shorter PFS in patients with astrocytic tumours (p = 0.024, Fig. 3b; Table 3). Combined with clinical features and immunophenotype, a multivariate model for PFS was established (Table 3). In this model, IDH1 mutation status, MGMT loss and low Ki67 index were significantly associated with a favourable influence on PFS (p = 0.04, p = 0.001, p = 0.009, Table 3). To further investigate the prognosis of the IDH1WT (IDH1 wild type) group, we combined MGMT expression level and the Ki67 index to evaluate PFS. We classified 47 astrocytic tumours into 3 groups: Group 1, patients with IDH1MUT (IDH1 mutant) regardless of Ki67 index or MGMT expression level; Group 2, patients with IDH1WT, low Ki67 index and MGMT protein loss; and Group 3, patients with IDH1WT but not classified as Group 2. Patients from Group 1 had the longest PFS, whereas patients from Group 3 had the shortest PFS. The PFS of Group 2 was between that of Groups 1 and 3. The median PFS of Groups 1, 2 and 3 was 23 months, 15 months and 7.5 months, respectively (p = 0.0000009, Fig. 3e). To investigate whether this new comprehensive classification is a better prognostic marker for PFS than the individual biomarkers (IDH1, Ki67 and MGMT), receiver-operating characteristic (ROC) curves of 1-year PFS were generated. The AUC (area under curve) of the comprehensive classifier for 1-year PFS was 0.792, which was greater than that for IDH1 (0.732), Ki67 (0.618) or MGMT (0.623) alone in our dataset (Supplementary Table S3).
The relationships between molecular markers and progression-free survival (PFS). (a) Patients with IDH mutations showed a significantly improved PFS. (b) Patients with TERTp mutations showed a shorter PFS compared to those with TERTp wild type. (c) MGMT-positive patients had a shorter PFS compared to MGMT-negative patients. (d) ATRX-negative patients had a significantly improved PFS. (e) Among IDH1 wild type patients, those with a low proliferative index and MGMT loss had a long PFS.
The prognostic significance of IDH1, TP53, and TERTp mutations and loss of MGMT and ATRX in primary tumours for overall survival (OS) was evaluated using Kaplan-Meier analysis. IDH1 and TP53 mutations and loss of ATRX were associated with a better OS (p = 0.014, Supplementary Figure S1a; p = 0.016, Supplementary Figure S1c; p = 0.022, Supplementary Figure S1g). OS was significantly shorter in patients harbouring TERTp mutation (p = 0.001, Supplementary Figure S1e). There was no difference in OS between patients with or without MGMT loss. IDH1, TP53 and TERTp mutation and ATRX loss at recurrence retained prognostic value for OS (p = 0.014, Supplementary Figure S1b; p = 0.005, Supplementary Figure S1d; p = 0.004, Supplementary Figure S1f; p = 0.03, Supplementary Figure S1h). Unfortunately, despite our best efforts, there were twenty-two patients lost to follow-up, which may lead to attrition bias. We did not do further analysis using Cox regression models in view of withdraw bias. Further study is required to confirm our observations.
Discussion
With the exception of grade I astrocytic tumours, astrocytic tumours show diffuse infiltration of brain tissue and cannot be completely removed by surgical resection, which invariably leads to recurrent episodes. Recurrence remains one of the most important limitations in the cure of astrocytic tumours. Astrocytic tumours follow Darwin’s theory of evolution and acquire novel mutations, which account in part for treatment failure and recurrence37. IDH1, TP53 and TERTp play key roles in tumourigenesis. However, few studies have investigated their mutation state in recurrent gliomas, particularly regarding TERTp mutation. In our study, similar to previously reported studies22,38,39, the IDH1 status of the primary tumour was consistent with that of the corresponding recurrent tumour in all cases. However, previous studies reported that the IDH1 mutation status changed in a minority of recurrent tumours4,40. The TP53 status remained unchanged in 85.11% of the recurrent tumours in our group, which is close to the percentage reported by Groenendijk et al. (85.19%)41 and Kraus et al. (72.73%)24. To the best of our knowledge, the articles by Heidenreich et al.42 and Nonoguchi et al.10 are the only two describing TERTp mutations in adult malignant gliomas and matched recurrences. Two (2/20)42 and three (3/21)10 primary/recurrent tumour pairs with TERTp mutation in the primary lesion but not the recurrent lesion have been described. Our research confirmed that TERTp status changed in recurrent tumours in only a small percentage of cases (3/47). In all cases, IDH1, TP53, and TERTp status in the primary tumour always predicted the status in the recurrent tumour. These results suggest that alterations in IDH1, TP53, and TERTp occur early during gliomagenesis and remain stable in tumour recurrence.
ATRX mutation leads to a truncated protein and is highly associated with negative staining by immunohistochemistry, which can be used as an alternative indicator for ATRX mutation43. In contrast to the pattern of TERTp mutations, which were predominately detected in pGBM, ATRX protein loss was mostly detected in A, AA and sGBM. These results suggest that astrocytomas and pGBM maintain telomere length while cells divide by altering different genes. In our samples, 11 of 47 cases (23.40%) showed a change in ATRX expression status, which was in agreement with the work by Johnson et al.38.
MGMT greatly improves tumour cell resistance to alkylating nitrosoureas and methylating agents by repairing alkylating lesions in DNA. Loss of MGMT expression in glioma is rarely caused by deletion, mutation, or rearrangement of the MGMT gene, but is mainly caused by methylation of the upstream promoter44. A significant inverse correlation was observed between MGMT protein expression and MGMT promoter methylation44,45,46. Notably, regulation of MGMT expression is a complicated process in which promoter hypermethylation is not the sole determinant27. Thus, an inconsistent correlation between abnormal promoter methylation and protein expression loss has been observed in some studies47,48. To a certain extent, the limitations of promoter methylation and protein expression detection methods interfere with the evaluation of correlations. Although promoter methylation can be directly and objectively detected, the methods for analysing MGMT promoter methylation status are more complicated than those for detecting MGMT protein expression by immunohistochemistry, which is a technique that is available in most laboratories. In our study, MGMT protein expression status by immunohistochemistry was changed in 11 of 47 cases (23.40%) during recurrence. The probability of change in negative tumours was equal to that in positive tumours; this observation is different from that reported by Brandes et al.49. Previous studies reported that MGMT methylation status changed after recurrence in 10% to 58.3% of the patients19,41,49,50. In brief, MGMT and ATRX protein expression levels were stable during recurrence in most cases.
All five biomarkers (IDH1, TP53 and TERTp mutation status and MGMT and ATRX protein expression levels) remained unchanged in 25 cases during recurrence, suggesting that these recurrences might represent direct expansion of the primary residual tumour and may stem from linear clonal evolution38. Another possibility is that these alterations occurred early in tumour development, prior to branched clonal evolution. The other 22 cases showed diversity in one or two biomarkers. Among these 22 cases, some acquired novel gene mutation or protein loss during recurrence, perhaps following Darwin’s theory of evolution. The other cases in which gene mutation or protein loss detected in the primary tumour was no longer found in the recurrent tumour could be partly explained by considering that the recurrent tumours in these patients were seeded by cells derived from the primary tumour at an early stage of evolution39. The genotype changes during recurrence and may follow the pattern of branched clonal evolution38.
To a certain extent, the histological and clinical features of astrocytic tumours can be prognostic indicators. However, to date, it is unclear why the interval between first surgery and recurrence is long in some patients but relatively short in others. Although Thon et al.21 reported that patients with IDH1 mutation grade II A had a short PFS, other studies have shown that A and AA22 and pGBM11 patients with IDH1 mutations have a significantly longer PFS. Our data also showed that patients with astrocytic tumours harbouring IDH1 mutations had a significantly longer PFS than those with wildtype IDH1. Furthermore, Hartmann et al.11 found that patients with IDH1-mutant glioblastoma have a more favourable PFS than those with IDH1-WT AA. All together, these observations indicate that IDH1 mutation can be considered a prognostic marker of PFS in patients with astrocytic tumours. Together with IDH1 gene mutation, MGMT protein expression level and Ki67 index were identified as independent prognostic factors for PFS in multivariate Cox regression analysis. The value of Ki67 and MGMT as independent prognostic factors in glioma was reported previously51,52,53,54,55. We classified 47 astrocytic tumours into 3 groups based on IDH1 mutation status and MGMT and Ki67 expression level without considering histological grade. Group 1 patients (IDH1MUT regardless of Ki67 index or MGMT expression level) had the longest PFS, whereas Group 3 patients (IDH1WT but not in Group 2) had the shortest PFS. Group 2 patients (IDH1WT, low Ki67 index and MGMT protein loss) had an intermediate PFS between that of Groups 1 and 3. The AUC of the comprehensive classifier at the first year was greater than IDH1, Ki67 or MGMT alone in our dataset, suggesting that IDH1 mutation combined with MGMT protein expression level and Ki67 index is a better prognostic factor for PFS in patients with astrocytic tumours. Therefore, we propose this new molecular classification for evaluating the PFS of patients with astrocytic tumours in future clinical trials. We focused on determining the prognostic significance of certain markers. In the future, further clinical studies are needed to clarify whether these markers are predictive for treatment response56.
In conclusion, our study reveals that in most cases, IDH1, TP53 and TERTp mutation status and MGMT and ATRX expression levels are stable during disease recurrence, perhaps indicating that those investigated biomarker alterations occurred early in the astrocytic tumour development. Meanwhile, in IDH1WT group, patients who were negative for MGMT and a low Ki67 index had a longer PFS than those who were positive for MGMT or had a high Ki67 index. The combination of IDH1 mutation, MGMT protein expression level and Ki67 index is a better indicator of the survival of patients with astrocytic tumours than any of these indicators alone. Therefore, IDH1 mutation combined with MGMT protein expression level and Ki67 index is a potential novel biomarker for evaluating the PFS of patients with astrocytic tumours. A larger study is required to confirm our observations.
Methods
Patients
Forty-seven paired astrocytic tumour samples were obtained from Xijing Hospital. Forty-six patients had a primary tumour and one recurrent tumour, and one patient had a primary tumour and two subsequent recurrent tumours. Tumour specimens were obtained by surgical resection between January 2006 and March 2015. Pathological diagnosis was reconfirmed by two neuropathologists in the Department of Pathology of Xijing Hospital under the 2007 WHO Classification of Tumours of the Central Nervous System. All primary tumours were untreated prior to operation and underwent total gross resection. The clinical data for each patient are shown in Table 4. All the study protocols were approved by the ethics committee of Xijing Hospital, Fourth Military Medical University. All study procedures were performed in accordance with the approved guidelines and regulations of the Fourth Military Medical University. Informed consent was obtained from all the patients, and all patient records were anonymised according to ethical and legal standards.
Molecular Genetic Analysis
Whole resected tumour tissues from individual patients were sampled to extract tumour DNA. Genomic DNA was extracted from formalin-fixed, paraffin-embedded tissues using the Qiagen GeneReadTM FFPE DNA Extraction Kit. Before the tumour DNA was isolated, the proportion of tumour in the consecutive haematoxylin and eosin-stained section was evaluated. If the proportion of tumour was more than ninety percent, five 10-μm-thick tissue sections were placed directly in a sterile 1.5-ml Eppendorf tube for DNA extraction. If the proportion of tumour was less than ninety percent, tumour-containing regions were removed from micro-dissected 10-μm sections to avoid contamination with normal tissue. After the tumour DNA was isolated, DNA quantity and quality were analysed using a Nanodrop 2000 spectrophotometer and 0.5% agarose gels.
Detection of TERTp, TP53 and IDH1 Hotshot Mutations
Mutations were analysed by PCR and direct sequencing. Primer sequences and amplification conditions for exons 5–8 of TP53 and IDH1 codon 132 mutations were previously reported57,58. Primer sequences for TERTp were designed using the Primer Premier software package (version 5, Premier Inc.). Primer sequences and amplification conditions for IDH1, TP53 and TERTp are shown in detail in Supplementary Table S4.
Immunohistochemistry analysis
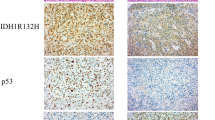
Immunohistochemistry was performed to detect the protein expression of MGMT and ATRX in the tissue samples. Briefly, 3-µm-thick tissue sections were prepared from formalin-fixed, paraffin-embedded tissue. Then, sections were deparaffinised in xylene and rehydrated in a series of ethanol washes. The sections were treated with 0.01 M sodium citrate buffer (pH 6.0) for 1 minute using a steam pressure cooker for antigen retrieval. The sections were immersed in 3% H2O2 for 15 minutes at room temperature to block endogenous peroxidase activity. After washing with phosphate-buffered saline (PBS) three times, sections were blocked in 5% bovine serum albumin (BSA) for 30 minutes. The sections were incubated at 4 °C overnight with anti-ATRX antibody (cat# HPA001906, lot J104918, Sigma-Aldrich, St. Louis, USA; dilution 1:500) or anti-MGMT antibody (clone MT3.1, MAB-0361, MAIXIN, Fuzhou, China; dilution 1:100). The next day, the sections were washed with PBS three times and then incubated with secondary antibody (REALTM EnVisionTM HRP anti-rabbit/mouse K5007, lot 20015510, Dako, USA) at room temperature for 30 minutes. The positive signal was visualised using diaminobenzidine as the chromogen, and nuclei were counterstained with haematoxylin. Blood vessel endothelium was used as an internal positive control, and PBS was used as a negative control. The immunoreactivity to ATRX and MGMT protein was evaluated by estimating the proportion of positive cells: ≤ 10% was regarded as negative, and > 10% was regarded as positive59,60. The stained sections were scored by two pathologists. Discrepancies between the two pathologists were settled by additional observation of the specimens and discussion until consensus was achieved. The pathologists were blinded to the clinical and molecular information during the immunohistochemical analysis.
Statistical Analysis
The agreement and difference in IDH1, TP53 and TERTp mutation status and MGMT and ATRX expression level in primary and recurrent tumours were estimated using the agreement test and McNemar’s test, respectively. The relation between molecular markers and clinical data was analysed using the chi-square test. PFS was defined as the interval between first surgical treatment and tumour recurrence. The PFS of different groups was estimated by Kaplan-Meier analysis, and the prognostic difference was evaluated with the log-rank test. Then, multivariate Cox regression models were used to evaluate potential prognostic factors. The new comprehensive classification was validated using ROC curves. P values were all two-tailed, and p values less than 0.05 indicated statistical significance. All data were analysed using SPSS software package (version 23, SPSS Inc.).
Data Availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
References
Ostrom, Q. T. et al. CBTRUS Statistical Report: Primary Brain and Central Nervous System Tumors Diagnosed in the United States in 2008–2012. Neuro Oncol 17(Suppl 4), v1 (2015).
Louis, D. N. et al. The2007 WHO classification of tumours of the central nervous system. Acta Neuropathol 114, 97 (2007).
Choi, S. H. et al. Impact of Including Peritumoral Edema in Radiotherapy Target Volume on Patterns of Failure in Glioblastoma following Temozolomide-based Chemoradiotherapy. Sci Rep 7, 42148 (2017).
Cai, J. et al. Detection of ATRX and IDH1-R132H immunohistochemistry in the progression of 211 paired gliomas. Oncotarget 7, 16384 (2016).
Oike, T. et al. Radiotherapy plus concomitant adjuvant temozolomide for glioblastoma: Japanese mono-institutional results. Plos One 8, e78943 (2013).
Wen, P. Y. & Kesari, S. Malignant gliomas in adults. N Engl J Med 359, 492 (2008).
Balss, J. et al. Analysis of the IDH1 codon 132 mutation in brain tumors. Acta Neuropathol 116, 597 (2008).
Bettegowda, C. et al. Mutations in CIC and FUBP1 contribute to human oligodendroglioma. Science 333, 1453 (2011).
Heaphy, C. M. et al. Altered telomeres in tumors with ATRX and DAXX mutations. Science 333, 425 (2011).
Nonoguchi, N. et al. TERT promoter mutations in primary and secondary glioblastomas. Acta Neuropathol 126, 931 (2013).
Hartmann, C. et al. Patients with IDH1 wild type anaplastic astrocytomas exhibit worse prognosis than IDH1-mutated glioblastomas, and IDH1 mutation status accounts for the unfavorable prognostic effect of higher age: implications for classification of gliomas. Acta Neuropathol, 120(6), p. 707-18. (2010).
Jiao, Y. et al. Frequent ATRX, CIC, FUBP1 and IDH1 mutations refine the classification of malignant gliomas. Oncotarget 3, 709 (2012).
Yang, P. et al. Classification based on mutations of TERT promoter and IDH characterizes subtypes in grade II/III gliomas. Neuro Oncol 18, 1099 (2016).
Eckel-Passow, J. E. et al. Glioma Groups Based on 1p/19q, IDH, and TERT Promoter Mutations in Tumors. N Engl J Med 372, 2499 (2015).
Brat, D. J. et al. Comprehensive, Integrative Genomic Analysis of Diffuse Lower-Grade Gliomas. N Engl J Med 372, 2481 (2015).
Arita, H. et al. A combination of TERT promoter mutation and MGMT methylation status predicts clinically relevant subgroups of newly diagnosed glioblastomas. Acta Neuropathologica Communications 4 (2016).
Killela, P. J. et al. Mutations in IDH1, IDH2, and in the TERT promoter define clinically distinct subgroups of adult malignant gliomas. ONCOTARGET 5, 1515 (2014).
Louis, D. N. et al. The2016 World Health Organization Classification of Tumors of the Central Nervous System: a summary. Acta Neuropathol 131, 803 (2016).
Laffaire, J. et al. Methylation profiling identifies 2 groups of gliomas according to their tumorigenesis. Neuro Oncol 13, 84 (2011).
Ishii, N. et al. Cells with TP53 mutations in low grade astrocytic tumors evolve clonally to malignancy and are an unfavorable prognostic factor. Oncogene 18, 5870 (1999).
Thon, N. et al. IDH1 mutations in grade II astrocytomas are associated with unfavorable progression-free survival and prolonged postrecurrence survival. Cancer-Am Cancer Soc 118, 452 (2012).
Yao, Y. et al. Mutation Analysis of IDH1 in Paired Gliomas Revealed IDH1 Mutation Was Not Associated with Malignant Progression but Predicted Longer Survival. Plos One 8, e67421 (2013).
Watanabe, K. et al. Incidence and timing of p53 mutations during astrocytoma progression in patients with multiple biopsies. Clin Cancer Res 3, 523 (1997).
Kraus, J. A. et al. TP53 alterations and clinical outcome in low grade astrocytomas. Genes Chromosomes Cancer 10, 143 (1994).
Park, C. K. et al. Expression level of hTERT is regulated by somatic mutation and common single nucleotide polymorphism at promoter region in glioblastoma. Oncotarget 5, 3399 (2014).
Labussiere, M. et al. TERT promoter mutations in gliomas, genetic associations and clinico-pathological correlations. Br J Cancer 111, 2024 (2014).
Younis, S. G., Khedr, R. A. & El-Shorbagy, S. H. Immunohistochemical analysis of O6-methylguanine-DNA methyltransferase (MGMT) protein expression as prognostic marker in glioblastoma patients treated with radiation therapy with concomitant and adjuvant Temozolomide. J Egypt Natl Canc Inst 28, 23 (2016).
Shen, D. et al. MGMT promoter methylation correlates with an overall survival benefit in Chinese high-grade glioblastoma patients treated with radiotherapy and alkylating agent-based chemotherapy: a single-institution study. Plos One 9, e107558 (2014).
Reifenberger, G. et al. Predictive impact of MGMT promoter methylation in glioblastoma of the elderly. Int J Cancer 131, 1342 (2012).
Gretarsdottir, S. et al. TP53 mutation analyses on breast carcinomas: a study of paraffin-embedded archival material. Br J Cancer 74, 555 (1996).
Murphy, M., McManus, D. T., Toner, P. G. & Russell, S. E. TP53 mutation in ovarian carcinoma. Eur J Cancer 33, 1281 (1997).
Krypuy, M. et al. High resolution melting for mutation scanning of TP53 exons 5–8. BMC Cancer 7, 168 (2007).
Nigro, J. M. et al. Mutations in the p53 gene occur in diverse human tumour types. Nature 342, 705 (1989).
Sugawara, W. et al. Association of germline or somatic TP53 missense mutation with oncogene amplification in tumors developed in patients with Li-Fraumeni or Li-Fraumeni-like syndrome. Genes Chromosomes Cancer 50, 535 (2011).
Saft, L. et al. p53 protein expression independently predicts outcome in patients with lower-risk myelodysplastic syndromes with del(5q). Haematologica 99, 1041 (2014).
Caron, D. F. C. & Soussi, T. TP53 tumor suppressor gene: a model for investigating human mutagenesis. Genes Chromosomes Cancer 4, 1 (1992).
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306 (2012).
Johnson, B. E. et al. Mutational Analysis Reveals the Origin and Therapy-Driven Evolution of Recurrent Glioma. Science 343, 189 (2014).
Ichimura, K. et al. IDH1 mutations are present in the majority of common adult gliomas but rare in primary glioblastomas. Neuro Oncol 11, 341 (2009).
Lass, U. et al. Clonal analysis in recurrent astrocytic, oligoastrocytic and oligodendroglial tumors implicates IDH1- mutation as common tumor initiating event. PLOS ONE 7, e41298 (2012).
Groenendijk, F. H. et al. MGMT promoter hypermethylation is a frequent, early, and consistent event in astrocytoma progression, and not correlated with TP53 mutation. J Neuro-Oncol 101, 405 (2011).
Heidenreich, B. et al. TERT promoter mutations and telomere length in adult malignant gliomas and recurrences. Oncotarget 6, 10617 (2015).
Liu, X. Y. et al. Frequent ATRX mutations and loss of expression in adult diffuse astrocytic tumors carrying IDH1/IDH2 and TP53 mutations. Acta Neuropathol 124, 615 (2012).
Esteller, M., Hamilton, S. R., Burger, P. C., Baylin, S. B. & Herman, J. G. Inactivation of the DNA repair gene O6-methylguanine-DNA methyltransferase by promoter hypermethylation is a common event in primary human neoplasia. Cancer Res 59, 793 (1999).
Deb, P., Mani, N. S., Sudumbrekar, S. M., Taneja, N. & Patrikar, S. Correlation of histomorphologic prognostic markers and proliferative index with loss of heterozygosity 1p/19q and MGMT status in diffusely infiltrating gliomas. Medical Journal Armed Forces India 69, 228 (2013).
Sonoda, Y. et al. O(6)-Methylguanine DNA methyltransferase determined by promoter hypermethylation and immunohistochemical expression is correlated with progression-free survival in patients with glioblastoma. Int J Clin Oncol 15, 352 (2010).
Mellai, M. et al. MGMT promoter hypermethylation in a series of 104 glioblastomas. Cancer Genomics Proteomics 6, 219 (2009).
Preusser, M. et al. Anti-O6-methylguanine-methyltransferase (MGMT) immunohistochemistry in glioblastoma multiforme: observer variability and lack of association with patient survival impede its use as clinical biomarker. Brain Pathol 18, 520 (2008).
Brandes, A. A. et al. O(6)-methylguanine DNA-methyltransferase methylation status can change between first surgery for newly diagnosed glioblastoma and second surgery for recurrence: clinical implications. Neuro Oncol 12, 283 (2010).
Nakamura, M., Watanabe, T., Yonekawa, Y., Kleihues, P. & Ohgaki, H. Promoter methylation of the DNA repair gene MGMT in astrocytomas is frequently associated with G:C–> A:T mutations of the TP53 tumor suppressor gene. carcinogenesis 22, 1715 (2001).
Ahmed, S., Rashed, H., Hegazy, A., Mohamed, A. M. & Elmesallamy, W. Prognostic Value of ALDH1, EZH2 and Ki-67 in Astrocytic Gliomas. Turk Patoloji Derg 32, 70 (2016).
Miconi, G. et al. Immunophenotypic characterization of human glioblastoma stem cells: correlation with clinical outcome. J Cell Biochem 116, 864 (2015).
Wiewrodt, D. et al. MGMT in primary and recurrent human glioblastomas after radiation and chemotherapy and comparison with p53 status and clinical outcome. Int J Cancer 122, 1391 (2008).
Ogura, R. et al. Immunohistochemical profiles of IDH1, MGMT and P53: Practical significance for prognostication of patients with diffuse gliomas. Neuropathology 35, 324 (2015).
Capper, D., Mittelbronn, M., Meyermann, R. & Schittenhelm, J. Pitfalls in the assessment of MGMT expression and in its correlation with survival in diffuse astrocytomas: proposal of a feasible immunohistochemical approach. Acta Neuropathol 115, 249 (2008).
Ballman, K. V. Biomarker: Predictive or Prognostic? J Clin Oncol 33, 3968 (2015).
Huang, C. C. et al. Different p53 mutation patterns in colorectal tumors from smokers and nonsmokers. Environ Mol Mutagen 47, 527 (2006).
Korshunov, A. et al. Combined molecular analysis of BRAF and IDH1 distinguishes pilocytic astrocytoma from diffuse astrocytoma. Acta Neuropathol 118, 401 (2009).
Wiestler, B. et al. ATRX loss refines the classification of anaplastic gliomas and identifies a subgroup of IDH mutant astrocytic tumors with better prognosis. Acta Neuropathol 126, 443 (2013).
Lotfi, M. et al. Immunohistochemical assessment of MGMT expression and p53 mutation in glioblastoma multiforme. Tumori 97, 104 (2011).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (81272651 and 81570180 to Zhe W).
Author information
Authors and Affiliations
Contributions
Wang Zhe designed the project. Jie Wei, Yixiong Liu, Linni Fan, Peifeng Li, Mingyang Li collected the tissue samples, clinical data. Yingmei Wang and Xia Li. performed the IHC. Xia Li, Danhui Zhao, Zhou Yu participated in the detection of TERTp, TP53 and IDH1 Hotshot Mutations. Ying Guo, Qingguo Yan evaluated the IHC analysis. Shuangping Guo helped analyze the data. Xia Li wrote the manuscript. Jing Ye revised the manuscript. All authors provided editorial input. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
41598_2017_13272_MOESM1_ESM.doc
Primary Astrocytic Tumours and Paired Recurrences have Similar Biological Features in IDH1, TP53 and TERTp Mutation and MGMT, ATRX Loss
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, X., Wei, J., Liu, Y. et al. Primary Astrocytic Tumours and Paired Recurrences have Similar Biological Features in IDH1, TP53 and TERTp Mutation and MGMT, ATRX Loss. Sci Rep 7, 13038 (2017). https://doi.org/10.1038/s41598-017-13272-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-13272-9
- Springer Nature Limited
This article is cited by
-
Assessment of genetic variant burden in epilepsy-associated brain lesions
European Journal of Human Genetics (2019)