Abstract
Non-coding RNA (ncRNA) is abundant in plant genomes, but is poorly described with unknown functionality in most species. Using whole genome RNA sequencing, we identified 1885, 1910 and 1299 lncRNAs and 186, 157 and 161 miRNAs at the whole genome level in the three Brassica species B. napus, B. oleracea and B. rapa, respectively. The lncRNA sequences were divergent between the three Brassica species. One quarter of lncRNAs were located in tandem repeat (TR) region. The expression of both lncRNAs and miRNAs was strongly biased towards the A rather than the C subgenome in B. napus, unlike mRNA expression. miRNAs in genic regions had higher average expression than miRNAs in non-genic regions in B. napus and B. oleracea. We provide a comprehensive reference for the distribution, functionality and interactions of lncRNAs and miRNAs in Brassica.
Similar content being viewed by others
Introduction
Non-coding RNA (ncRNA) refers to RNAs such as rRNAs, tRNAs and snRNAs that can be transcribed, but which do not encode proteins. Non-coding RNAs play different roles in the cell and in gene expression: common types include ribosomal RNAs (rRNAs) and transfer RNAs (tRNAs), which function in mRNA translation, small nuclear RNAs (snRNAs) involved in splicing, small nucleolar RNAs (snoRNAs) that act to modify rRNAs, and microRNAs (miRNAs) and small interfering RNAs (siRNAs) that regulate the translation and/or stability of mRNA1. More recent discoveries include piwi-interacting RNAs (piRNAs), small RNAs which suppress transposon activity and regulate gene expression2, and long non-coding RNAs (lncRNAs).
lncRNAs are defined as RNAs more than 200 bp in length but lacking in protein-coding potential3. There is accumulating evidence for participation of lncRNAs in a broad range of processes4. For example, lncRNAs have been revealed to play major roles in transcriptional regulation, splicing, the organization of nuclear domains and chromatin modification5,6. Classification of lncRNAs is a complex task, particularly as functional knowledge is still missing for many identified lncRNAs4. Classifications can be based on features such as transcript length, association with annotated protein-coding genes, repeats or other DNA elements of known function, on resemblance to protein coding RNA, on association with a biochemical pathway or subcellular structure, on sequence and structural conservation or on functionality4.
LncRNAs have been studied in different tissues and under stress conditions in many plants, and have been shown to play a role in both plant development and various stress responses7. Some specific examples include stress-responses in Arabidopsis8 and in wheat9, regulation of photoperiod-sensitive male sterility in hybrid rice10, as well as in rice sexual reproduction11, where some lncRNAs act as competing endogenous RNAs (ceRNAs). lncRNAs appear poorly conserved between flowering plant species12,13,14, with highly tissue-specific expression7,12,14 and gene regulation through either cis or trans pathways12,14. Other genomic elements may also interact with lncRNAs. For example, Wang et al. (2015d)13 showed that transposable elements play a major role in the origin of Lycopersicon-specific lncRNAs in tomato. Wang et al. (2015a)15 also found lncRNAs functioning as competing endogenous target mimics (eTMs) for microRNAs in tomato response to tomato yellow leaf curl virus (TYLCV) infection.
MicroRNAs (miRNAs) are small noncoding RNAs with the length of 20–24 nt which play a major role in development and various stress responses through silencing of target mRNAs16. miRNA genes are transcribed into primary miRNA (pri-miRNA) by polymerase II, and fold into precursor miRNA (pre-miRNA) with a stem-loop structure17 before forming into mature miRNA. miRNAs recognize target transcripts through Argonaute (AGO) proteins, which repress mRNAs via binding with 3′ UTR or coding sequences. The degradation of mRNAs by miRNAs occurs through deadenylation, decapping and exonucleolytic digestion17. To date, miRNA genes have been identified and characterized in dozens of flowering plant species18,19,20,21, with ongoing investigations into the role of these small RNAs in regulation of gene expression in many different pathways22,23,24.
Brassica napus (AACC; rapeseed) is a young allopolyploid species derived from hybridization of diploid species B. oleracea (CC; cabbage) and B. rapa (AA; turnip) <7500 years ago25,26. This extremely short evolutionary history makes B. napus an intriguing model for studies of hybrid and polyploid formation27, particularly for rapidly-evolving genomic elements like small RNAs. Recently, the availability of the B. napus, B. rapa and B. oleracea genome sequences28,29,30 has allowed investigation of small RNAs in Brassica. A total of 969 miRNAs from 680 miRNA genes have been annotated in B. rapa, and were found to be more commonly retained than genes during post-polyploidization genome fractionation31. As well, 76% of miRNAs were found to be conserved between B. napus and its progenitor species B. rapa and B. oleracea, with recent MIRNA expansion and loss events detected in B. napus32. In B. rapa, 2237 candidate lncRNAs with an average length of 497 bp were identified, and the functions of neighboring genes were analyzed33. In addition, a total of 3183 lncRNAs were found to be responsive to pathogen Sclerotinia sclerotiorum in B. napus34. However, very little is yet known about function or evolution of lncRNAs in Brassica. In this study, we aimed to determine how lncRNAs have evolved and diverged in B. napus relative to progenitor species B. rapa and B. oleracea, and how miRNAs and lncRNAs may functionally interact in each of these species.
Results
High-throughput sequencing of B. napus, B. oleracea and B. rapa
To explore the effect of lncRNAs in Brassica, lncRNA sequencing of young leaves of B. napus, B. oleracea and B. rapa was performed in two biological replicates. Six strand-specific cDNA libraries were constructed. We removed low quality, adapter and uncertain reads, and obtained a total of 93–102 million clean data reads. About 51–69% of reads mapped to the reference genome (Supplementary Table 1).
In addition, in order to understand the expression of small RNAs in Brassica, six small RNA libraries from leaves of Brassica species were constructed. After removing low quality, 5′-adapter containing, 3′-adapter null or insert null reads and ploy-N-containing reads, a total of 7.5–10.1 million clean reads were generated. Of these reads, 77–90% were mapped to the genome, of which a further 57–78% mapped to a unique position on the reference genome sequences (Supplementary Table 2). To estimate the reproducibility of the data, correlations between replicated samples were made (Supplementary Figure 1), obtaining correlation coefficients of 0.988, 0.993 and 0.987 for lncRNA sequencing and 0.879, 0.889 and 0.823 for small RNA sequencing in B. napus, B. oleracea and B. rapa, respectively, suggesting high reproducibility between the biological replicates.
Identification of lncRNAs in B. napus, B. oleracea and B. rapa
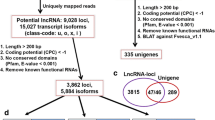
To identify lncRNAs in the three Brassica species, we developed a pipeline “RNAseq-Brassica” for the RNA-seq data (Fig. 1). After basic filtering and analysis of coding potential, 1885, 1910 and 1299 lncRNAs were found in B. napus, B. oleracea and B. rapa, respectively (Supplementary Table 3). BLASTn analysis was carried out to identify lncRNAs in B. napus in our study based on all lncRNAs previously published in this species34. No previous publications list locations of B. rapa or B. oleracea lncRNAs. We found 1483 of 1885 lncRNAs were novel lncRNAs in B. napus.
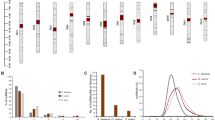
The lengths and distributions of the lncRNAs in the Brassica species genomes were analyzed. lncRNAs were evenly distributed across all Brassica chromosomes (Fig. 2A). The density of lncRNAs in the B. rapa and B. oleracea diploid genomes was 4.17 and 4.31 lncRNAs per Mb, respectively, with 2.00 lncRNAs per Mb in B. napus. The majority of lncRNAs (91.2%, 92.6% and 85.6%) were located in intergenic regions in B. napus, B. oleracea and B. rapa, respectively (Fig. 2B). The rest of the RNAs were anti-sense lncRNAs (7.27%, 5.50% and 9.39%) and intronic lncRNAs. lncRNA lengths were mostly less than 2 kb, with an average of 1163,1523 and 974 bp in B. napus, B. oleracea and B. rapa, respectively (Fig. 2C). The number of exons in lncRNAs in our study was 4–6 in all three Brassica species (although lncRNAs with only one exon were excluded). The structure and organization of lncRNAs was similar across the three Brassica species.
Identification of miRNAs in B. napus, B. oleracea and B. rapa
A total of 117, 102 and 123 conserved miRNAs found in B. napus, B. oleracea and B. rapa belonged to 63, 50 and 67 miRNA families, respectively (Supplementary Table 4). Of these 74 miRNA families, 43 miRNA families (75.4%) were present in all three species (such as miR156, miR160, miR162, miR164, miR319), four were present in both B. napus and B. oleracea (miR6034, mIR6035, miR9409 and miR9411) and 13 were present in both B. napus and B. rapa (Fig. 3). Three families were present in both B. oleracea and B. rapa (miR399, miR6036 and miR9410), but absent in B. napus. In total, three unique miRNA families (miR6028, miR5719 and miR5726) were found in B. napus, eight in B. rapa (miR5713, miR5714, miR5716, miR5724, miR9553, miR9556, miR9561 and miR9567) and none in B. oleracea.
Furthermore, novel miRNAs were discovered with miREvo35 and mirdeep 236 according to the hairpin structure of the miRNA precursors, the Dicer cleavage site and predicted free energy. A total of 69, 55 and 38 novel miRNAs were identified in B. napus, B. oleracea and B. rapa, respectively (Supplementary Table 4).
We analyzed the distribution of MIRNA (precursor genes of mature miRNA) in the Brassica genomes. A total of 168 MIRNA were found in B. napus, of which 32 (20.3%) were located in genes (Supplementary Table 5, Table 1). Of the 32 MIRNAs in genes, 12 (37.5%) were within 3′UTRs, seven (21.9%) were within introns, two (6.2%) were within exons, three (9.4%) were in 5′UTRs and eight (25.0%) were located in other genic regions. A total of 144 MIRNAs were found in B. oleracea, of which only five (3.5%) were located in genes, while 14 (9.8%) of the 143 MIRNAs in B. rapa were located in genes. Most MIRNA were found in intergenic regions.
qRT-PCR validation of lncRNAs and miRNAs
To validate the expression patterns of lncRNAs in Brassica, a total of nine lncRNAs were selected for qRT-PCR analysis (TCONS_00135228, TCONS_00137880 and TCONS_00518145 in B. napus, TCONS_00143251, TCONS_00194861 and TCONS_00243878 in B. oleracea, and TCONS_00340124, TCONS_00225121 and TCONS_00200766 in B. rapa). Expression patterns of all lncRNAs were confirmed to be highly similar between the lncRNA sequencing and qRT-PCR methods (Fig. 4A,B).
In addition, qRT-PCR analyses of 12 miRNAs in the three Brassica species were performed to validate the results of miRNA sequencing. Expression patterns for 11/12 miRNAs in the three Brassica species were similar between the two methods of miRNA sequencing and qRT-PCR, although miR408-5p in B. napus had different expression patterns between the two analytical tools (Fig. 4C,D).
Intermediate conservation of lncRNAs in Brassica
We analyzed the homology of lncRNAs in the three Brassica species with E < 1e-20 using BLASTn analysis: lncRNAs of B. napus were used as queries (1885) and lncRNA sequences from B. oleracea and B. rapa were used as subjects (Supplementary Table 6). Homologs for 23.1% (435/1885) and 11.0% (208/1885) of lncRNAs were identified in B. oleracea and B. rapa, respectively. In addition, lncRNAs of B. oleracea were used as queries and lncRNA sequences from B. rapa were used as subjects. In total, 176 of 1910 lncRNAs (9.2%) in B. oleracea were homologous to lncRNAs in B. rapa (Supplementary Table 6). A further 1174, 1321 and 916 lncRNAs were found to be specific to B. napus, B. oleracea and B. rapa, respectively. For protein-coding genes, 38,554 of 101,040 genes (38.2%) in B. napus were homologous to B. oleracea and 34,255 (33.9%) were homologous to B. rapa28. Therefore, Brassica lncRNAs showed intermediate conservation in comparison to protein-coding genes, which is consistent with the conservation of lncRNAs relative to protein-coding genes observed between A. thaliana and B. rapa (16%)37.
Repetitive sequences in lncRNAs
Most identified lncRNAs were located in intergenic regions. To understand the origin of these lncRNAs we analyzed repetitive sequences, as repetitive sequences are the major factor driving the emergence of lncNRAs37,38. Repetitive sequences include tandem repeats (TRs) that can be classified as satellite, minisatellite and microsatellite repeats, and dispersed repeats that include transposable elements (TEs). We analyzed the locations of lncRNAs in Brassica species in relation to repetitive sequences (Supplementary Table 7). In total, 477 of 1885 (25.3%) lncRNAs were found within repetitive sequences in B. napus, 98.9% of which were located in TR sequences, three of which were located in long terminal repeat (LTR) retrotransposon sequences, and one of which was located within a SINE sequence. In B. oleracea, 421 of 1901 lncRNAs (22.0%) were located within repetitive sequences, 417 of which (99.0%) were found in TR sequences, and one of which was located in a SINE sequence. In B. rapa, 304 of 1299 lncRNAs (23.5%) were found within repetitive sequences, of which 243 (79.9%) were located within TR sequences, 67 (22.0%) of which were located in LTR retrotransposon sequences and 9 (2.9%) of which were located in DNA transposons. A total of 1174, 1321 and 916 lncRNAs were found to be specific to B. napus, B. oleracea and B. rapa, respectively. Overall, 278 of 1174 (23.7%), 260 of 1321 (19.7%) and 186 of 916 (20.3%) lncRNAs contained repetitive sequences in B. napus, B. oleracea and B. rapa, respectively. TRs were more likely than other types of TEs to be associated with lncRNAs.
lncRNAs functioned as precursors or targets of miRNAs
lncRNAs and miRNAs play important roles in the regulation of gene expression. To understand the relationship between lncRNAs and miNRAs, we aligned the precursor sequences of miRNAs to lncRNAs. We found 14 lncRNAs (0.74%) were precursors of 20 miRNAs from 15 miRNA families (10 known and 5 novel miRNA families) in B. napus, 7 lncRNAs (0.37%) were precursors of 9 miRNAs from 8 miRNA families (4 known and 4 novel miRNA families) in B. oleracea, and 15 lncRNAs (1.15%) were precursors of 19 miRNAs from 15 miRNA families (10 known and 5 novel miRNA families) in B. rapa (Table 2). In addition, we found that some conserved miRNAs (miR156, miR159, miR166, miR167, miR168, miR172, miR393, miR1885, miR5654 and miR5718) were produced by lncRNAs in either B. rapa or B. oleracea as well as in B. napus. In addition, lncRNAs were predicted to be the targets of miRNAs in Brassica. A total of 18 (0.95%), 26 (1.36%) and 33 (2.54%) lncRNAs were the targets of miRNAs in B. napus, B. oleracea and B. rapa (Table 2). Hence, a fraction of lncRNAs appear to function as either precursors or targets of miRNAs.
Expression of lncRNAs
To understand the expression of lncRNAs in Brassica, the expression levels (FPKM) were assessed. In all Brassica species, mRNAs tended to have higher expression than lncRNAs (Supplementary Figure 2, Table 3). However, some lncRNAs had an expression of more than 15 log2(FPKM) in Brassica, such as TCONS_00135228 in B. napus and TCONS_00225121 in B. rapa, indicating that these lncRNAs may have functional roles, rather than being solely “transcriptional noise”.
In addition, we investigated the expression of lncRNAs to determine if expression bias was present between the two subgenomes of B. napus. The average expression level of lncRNAs in B. napus was 6.36. The number of lncRNAs in the A subgenome (573) was less than that in the Cn subgenome (867), which was consistent with the observation of the two progenitor genomes, where 1299 lncRNAs were identified in B. rapa (AA genome) and 1910 in B. oleracea (CC genome) (Table 3). The average expression level (FPKM) of lncRNAs was slightly higher in the An subgenome (7.40) than in the Cn subgenome (4.77) (t-test, P > 0.05). In the progenitor species, the average expression of lncRNAs in B. napus (6.36) and B. rapa (7.12) was also slightly higher than that in B. oleracea (3.96) (t-test, P > 0.05). However, the expression of mRNAs in the An subgenome (5.60) and Cn subgenome (5.99) was almost the same.
The average expression levels of TR-related, TE-related and unclassified lncRNAs were also analyzed. In B. napus and B. rapa, unclassified lncRNAs showed the highest expression levels, 104.88 and 172.82 respectively. In B. oleracea, TR-related lncRNAs had a slightly higher expression (18.61) than unclassified lncRNA sequences (14.85) (P > 0.05). More TE-related lncRNAs were present in B. rapa, and they showed higher expression levels (25.18) than TR-related lncRNAs (17.19).
Expression of miRNA
The progenitor species showed similar TPM for miRNA expression: 6369 in B. oleracea and 6211 in B. rapa. In B. napus the average TPM was 5376, which was a little lower than that in the diploid species (Table 3). The average expression level (log2TPM) of miRNAs in the An genome (7.64) was higher than that of miRNAs in the Cn genome (7.49) (t-test, P > 0.05) in B. napus. This was consistent with the expression of lncRNAs in the subgenomes of B. napus.
In addition, miRNAs in genic regions had higher average expression than miRNAs in non-genic regions in B. napus and B. oleracea, but this finding was solely related to the position of the MIRNA159 gene in Brassica species. When miRNA159 was excluded from the analysis, the expression of miRNAs in non-genic regions was higher than that of miRNAs in genic regions in Brassica. The expression level of conserved miR159 was the highest of any miRNA in all three species: 420561 in B. napus (located in a genic region), 212657 in B. oleracea (located in a genic region) and 407567 B. rapa (located in a non-genic region), respectively (Fig. 5). The predicted target genes of miR159 mainly encoded MYB TFs, such as MYB81, MYB101, MYB65, MYB97, MYB120 and sporocyteless (SPL).
The expression levels of 43 common conserved miRNAs were also compared in Brassica, and found to average 13830 in B. napus, 7113 in B. oleracea and 13665 in B. rapa. Differentially expressed families were identified based on the criteria log2(fold change) > 2. A total of 12 miRNA families showed differential expression between B. napus and B. oleracea, eight between B. napus and B. rapa, and 13 between B. oleracea and B. rapa. The expression of miR9558 and miR9560 was up-regulated in B. napus relative to in B. rapa and B. oleracea (Fig. 5). Only the target genes of miR9558 were found, and these mainly participated in DNA synthesis and post-translational modification.
Target gene identification for miRNAs
To understand the role of miRNAs in Brassica gene regulation, target genes were predicted by psRobot39. A total of 61 miRNA families were targeted to 1080 genes in B. napus, 45 miRNA families were targeted to 340 genes in B. oleracea and 67 miRNA families were targeted to 906 genes in B. rapa (Supplementary Table 8). Some of the potential targets were transcription factors, and some were genes related to biotic and abiotic stress responses. A total of 161 genes were orthologous between B. rapa and B. oleracea, while 356 genes (33.0%) in B. rapa and 241 genes (71.2%) in B. oleracea were orthologous to those in B. napus. Gene functions were also similar in the three Brassica species. GO enrichment analysis showed that the target genes of miRNAs in Brassica were mainly enriched in cell death, multicellular organismal processes, developmental processes, defense response and immune system processes (Supplementary Table 9).
Discussion
We analyzed lncRNAs and miRNAs at the whole genome level in B. napus and its two progenitor species, B. oleracea and B. rapa. Very few studies to date have analysed lncRNAs in Brassica: one study previously identified 2237 lncRNAs in B. rapa33, while another identified 3183 lncRNAs in B. napus expressed in response to Sclerotinia34. In the present study we identified a total of 1885, 1910 and 1299 lncRNAs in B. napus, B. oleracea and B. rapa, respectively, significantly adding to the number of lncRNAs identified in Brassica species. Most lncRNAs (more than 85.6%) were located in intergenic regions in our study, consistent with previous studies in other species14,15. The density of lncRNAs was much lower in B. napus (2.00 lncRNAs per Mb) relative to its diploid progenitors B. rapa (4.17 lncRNAs per Mb) and B. oleracea (4.31 lncRNAs per Mb): this finding supports previously identified dynamics of “genome downsizing” via loss of repetitive elements and high-copy number sequences as a result of allopolyploid formation40,41.
A total of 117, 102 and 123 conserved miRNAs belonging to 63, 50 and 67 miRNA families were found in B. napus, B. oleracea and B. rapa respectively, including 69, 55 and 38 novel miRNAs. Most MIRNAs (80–90%) were located in intergenic regions. In addition, many more miRNAs (20.3%) were located in genes in B. napus than in B. rapa (3.5%) or in B. oleracea (9.8%), suggesting that these miRNAs may assist in the more complex gene regulation required in B. napus post-polyploidization. This finding supports previous work suggesting that miRNAs played an important role in polyploidization and subsequent genome evolution in B. napus32,42.
In our study, we found that the expression of lncRNAs and miRNAs in the An subgenome was higher than in the Cn subgenome in B. napus. The average expression level of An genome lncRNAs was also higher than the average expression level of Cn genome lncRNAs, and higher in A genome species B. rapa than in C genome species B. oleracea. The same trend of higher expression in the A genome relative to the C genome was found for miRNA (P > 0.05), as well as a higher abundance of miRNA reads. This is consistent with previous results identifying biased subgenome expression of miRNA and lncRNAs in B. napus and cotton, respectively32,43,44. This result supports the putative genome-wide homoeolog expression level bias45 between the A and the C genomes. The C subgenome of B. napus tends to be more readily lost than the A genome28,46, and is putatively more heavily silenced due to its increased burden of repetitive sequences relative to the A genome30; the A genome is therefore proposed to be “dominant” to the C genome in overall gene expression.
In general, miRNAs are moderately conserved between plant species32,47. Our study supported this result, with the identification of 43 of 74 conserved miRNA families (75.4%) present in all three Brassica species. In addition, we found some miRNAs that were putatively newly generated or lost after polyploidization: target genes for these miRNAs in B. napus were related to cell wall proteins and stress response, possible candidates for species adaptation processes. In contrast to miRNAs, lncRNAs showed poor conservation, with only 23 homologous lncRNAs out of the >5000 identified in the three Brassica species. This finding is consistent with previous studies, which support lncRNAs diversity and rapid evolution in plant species13,48. Although lncRNAs show rapid evolution, some conservation of function is predicted. lncRNA has previously been shown to be involved in plant developmental processes: FLC (FLOWERING LOCUS C) regulates flowering time in A. thaliana, and three lncRNAs, COOLAIR (CILD INDUCED LONG ANTISENSE INTRAGENCI RNAs), COLDAIR (COLD ASSISTED INTRONIC NONCODING RNA) and ASL (Antisense Long), modify FLC through epigenetic regulation5,49,50. lncRNAs may also be involved in phosphate (essential for plant growth and development) homeostasis in A. thaliana and rice51,52, and were also associated with putative phosphate metabolic process genes in our study. In addition, lncRNAs play a major role in various stress responses7,15, which might contribute to environmental adaptation in speciation. Wang et al. (2015c)53 identified lncRNAs expressed under osmotic and salt stress conditions in Medicago truncatula which were likely involved in adaptation to abiotic stresses.
Transcription factors were the main putative target genes of miRNAs in our study. Several of these transcription factors could be important in speciation. MYB101 functions in pollen tube reception in A. thaliana54, while ARF6 and ARF8 promote flower maturation in A. thaliana55. In addition, resistance genes accounted for a high proportion of genes associated with miRNAs in our study. Disease resistance genes have also been implicated in speciation via reproductive isolation, due to their potential relationship to hybrid necrosis56,57, and may contribute to differential environmental adaptation in newly formed species. However, a great deal more research still needs to be done to fully elucidate the evolutionary and regulatory functionality of miRNAs and particularly lncRNAs in the Brassica genus. Our research offers a tantalizing glimpse at possibilities for how these two classes of small RNAs may interact in polyploidization and speciation processes.
Materials and Methods
lncRNA sequencing and small RNA sequencing
Young leaves from single accessions of B. rapa, B. oleracea and B. napus (five lines per accession) were collected, pooled together and immediately frozen in liquid nitrogen for lncRNA sequencing and small RNA sequencing, with two biological replicates per accession. The B. oleracea (kale) sample was a DH line generated from accession “15M2143”, with black seeds and a long growth period. Semi-winter B. rapa “Yaanhuangyoucai” was a sixth generation self-pollinated inbred line from a local variety in Sichuan Agricultural University at Ya’an (187–190 day growth period and 30 day flowering period, 87% yellow-seeded/13% brown-seeded). Semi-winter brown-seeded B. napus “G184–189” was an eighth-generation self-pollinated inbred selection from the Sichuan Academy of Agricultural Sciences (226–229 day growth period, 30 day flowering period).
For lncRNA sequencing, ribosomal RNA was removed by an Epicentre Ribo-zerpTM rRNA Removal Kit (Epicentre, USA). Subsequently, sequencing libraries were generated using the rRNA-depleted RNA by NEBNext® Ultra™ Directional RNA Library Prep Kit for Illumina® (NEB, USA) following the manufacturer’s recommendations. The libraries were sequenced on an Illumina Hiseq2000 platform and 100 bp paired-end reads were generated.
Small RNA sequencing libraries were generated using NEBNext® Multiplex Small RNA Library Prep Set for Illumina® (NEB, USA.) following manufacturer’s recommendations, and index codes were added to attribute sequences to each sample. The libraries were sequenced on an Illumina Hiseq2500/2000 platform and 50 bp single-end reads were generated.
Identification of lncRNAs
Indices of the reference genomes of B. napus, B. oleracea and B. rapa25,29,58 were built using Bowtie v2.0.6 (Broad Institute, Cambridge, MA, USA)59. Sequencing reads were aligned to the reference genome using TopHat v2.0.960, and assembled by both Scripture (beta2)61 and cufflinks (v2.1.1)62.
We selected transcripts which met the following criteria: length ≥ 200 bp; read coverage > 3; presence in both sample replicates and both assemblies (Cufflinks and scripture). We then filtered for known non-lncRNA annotation and classified remaining transcripts as candidate lncRNAs. We subsequently performed coding potential filtering using Coding Potential Calculator (CPC) and Pfam-scan. CPC (Coding Potential Calculator) (0.9-r2) assesses the extent and quality of the open reading frame (ORF) in a transcript, and attempts to match sequences with a known protein sequence database to classify transcripts as coding vs. non-coding. We used the NCBI eukaryote protein database and set the e-value cut-off to 1e−10 in our analysis63. Pfam Scan (v1.3) was used to identify occurrences of any of the known protein family domains documented in the Pfam database (release 27; used both Pfam A and Pfam B)64. Any transcript with a Pfam hit was excluded from the following analysis steps. Pfam searches used default parameters65.
Identification of known and novel miRNA
Small RNA tags were mapped to the reference genome by Bowtie59 without permitting mismatches. We then removed the tags originating from protein-coding genes, repeat sequences, rRNA, tRNA, snRNA, snoRNA and other small RNA tags. miRBase20.0 was used as the reference database for known miRNA. Modified software “mirdeep2”36 and “srna-tools-cli” were used to obtain the potential miRNAs and draw the predicted secondary structures.
The hairpin structure of miRNA precursors can be used to predict novel miRNAs. Novel miRNAs were identified with “miREvo”35 and “mirdeep2”36 through secondary structure, minimum free energy and Dicer cleavage site characteristics. Briefly, the secondary structures of miRNA precursors were detected using Mfold66, and the structure with the minimum free energy was selected67.
Characterization of lncRNAs and miRNAs
The distribution of lncRNA and MIRNA genes in the genome was visualized using Circos68. To evaluate the lncRNAs that may act as precursors of miRNAs, we aligned the lncRNAs with identified miRNAs (e-value = 1e-5). lncRNAs as targets of miRNAs were predicted by psRNATarget (http://plantgrn.noble.org/psRNATarget/) with default values69. The rate of non-synonymous substitutions (Ka) and the rate of synonymous substitutions (Ks) of coding genes were determined by PAML-condem70.
Expression of lncRNAs and miRNAs
The expression levels of lncRNAs and coding genes were estimated using FPKM (fragments per kilobase of exon per million mapped fragments). The expression of miRNAs was normalized to TPM (transcripts per million clean tags)71 using the formula: TPM = mapped read count/total reads*1000000. Differentially expressed miRNAs between species were identified based on log2(fold change) ≥ 2.
qRT-PCR
To validate the results of miRNA and lncRNA sequencing, qRT-PCR was conducted in the three Brassica species. Primers are listed in Supplementary Table 10 (RiboBio Co.). PCR reactions contained 10 μL SSoAdvanced SYBR Green Supermix (Bio-Rad), 2.0 μL cDNA, 1 µL primer, and distilled water to a final volume of 20 µL. Two independent biological replicates, each with three technical replicates, were run for test genes. The cycle threshold (Ct) was determined using the default settings.
References
Mattick, J. S. & Makunin, I. V. Non-coding RNA. Human Molecular Genetics 15, R17–29 (2006).
Iwasaki, Y. W., Siomi, M. C. & Siomi, H. PIWI-Interacting RNA: Its Biogenesis and Functions. Annual Review of Biochemistry 84, 405–433 (2015).
Zhang, Y. C. & Chen, Y. Q. Long noncoding RNAs: new regulators in plant development. Biochemical and Biophysical Research Communications 436, 111–114 (2013).
St Laurent, G., Wahlestedt, C. & Kapranov, P. The Landscape of long noncoding RNA classification. Trends in Genetics: TIG 31, 239–251 (2015).
Chekanova, J. A. Long non-coding RNAs and their functions in plants. Current Opinion in Plant Biology 27, 207–216 (2015).
Zhu, Q. H. & Wang, M. B. Molecular functions of long non-coding RNAs in plants. Genes 3, 176–190 (2012).
Liu, J. et al. Genome-wide analysis uncovers regulation of long intergenic noncoding RNAs in Arabidopsis. The Plant Cell 24, 4333–4345 (2012).
Di, C. et al. Characterization of stress-responsive lncRNAs in Arabidopsis thaliana by integrating expression, epigenetic and structural features. The Plant Journal 80, 848–861 (2014).
Xin, M. et al. Identification and characterization of wheat long non-protein coding RNAs responsive to powdery mildew infection and heat stress by using microarray analysis and SBS sequencing. BMC Plant Biology 11, 61 (2011).
Ding, J. et al. A long noncoding RNA regulates photoperiod-sensitive male sterility, an essential component of hybrid rice. Proc Natl Acad Sci USA 109, 2654–2659 (2012).
Zhang, Y. C. et al. Genome-wide screening and functional analysis identify a large number of long noncoding RNAs involved in the sexual reproduction of rice. Genome Biology 15, 512 (2014).
Kang, C. & Liu, Z. Global identification and analysis of long non-coding RNAs in diploid strawberry Fragaria vesca during flower and fruit development. BMC Genomics 16, 815 (2015).
Wang, X. et al. Expression and diversification analysis reveals transposable elements play important roles in the origin of Lycopersicon-specific lncRNAs in tomato. New Phytologist (2015).
Li, L. et al. Genome-wide discovery and characterization of maize long non-coding RNAs. Genome Biology 15, R40 (2014).
Wang, J. et al. Genome-wide analysis of tomato long non-coding RNAs and identification as endogenous target mimic for microRNA in response to TYLCV infection. Scientific Reports 5, 16946 (2015).
Eckardt, N. A. The plant cell reviews aspects of microRNA and PhasiRNA regulatory function. The Plant Cell 25, 2382 (2013).
Carthew, R. W. & Sontheimer, E. J. Origins and Mechanisms of miRNAs and siRNAs. Cell 136, 642–655 (2009).
Griffiths-Jones, S. miRBase: the microRNA sequence database. Methods in Molecular Biology 342, 129–138 (2006).
Griffiths-Jones, S. The microRNA Registry. Nucleic Acids Research 32, D109–111 (2004).
Griffiths-Jones, S., Saini, H. K., van Dongen, S. & Enright, A. J. miRBase: tools for microRNA genomics. Nucleic Acids Research 36, D154–158 (2008).
Kozomara, A. & Griffiths-Jones, S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Research 42, D68–73 (2014).
Mallory, A. C. & Vaucheret, H. Functions of microRNAs and related small RNAs in plants. Nature Genetics 38, Suppl, S31–36 (2006).
Zhang, B. MicroRNA: a new target for improving plant tolerance to abiotic stress. Journal of Experimental Botany 66, 1749–1761 (2015).
Dukowic-Schulze, S. et al. Novel Meiotic miRNAs and Indications for a Role of PhasiRNAs in Meiosis. Frontiers in Plant Science 7, 762 (2016).
Chalhoub, B. et al. Plant genetics. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345, 950–953 (2014).
U, N. Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Japanese Journal of Botany 7, 389-452 (1935).
Mason, A. S. & Snowdon, R. J. Oilseed rape: learning about ancient and recent polyploid evolution from a recent crop species. Plant Biology (2016).
Chalhoub, B. Early allopolyploid evolution in the post-Neolithic Brassica napus oilseed genome. Science 345, 1255–1255 (2014).
Liu, S. et al. The Brassica oleracea genome reveals the asymmetrical evolution of polyploid genomes. Nature Communications 5, 3930 (2014).
Parkin, I. A. P. et al. Transcriptome and methylome profiling reveals relics of genome dominance in the mesopolyploid Brassica oleracea. Genome Biology 15, R77 (2014).
Sun, C. et al. Impacts of whole-genome triplication on MIRNA evolution in Brassica rapa. Genome Biology and Evolution 7, 3085–3096 (2015).
Shen, E. et al. Identification, evolution, and expression partitioning of miRNAs in allopolyploid Brassica napus. Journal of Experimental Botany 66, 7241–7253 (2015).
Paul, P., Dhandapani, V., Choi, S. R. & Lim, Y. P. Genome wide identification and functional prediction of long non-coding RNAs in Brassica rapa. Genes Genom 38, 547–555 (2016).
Joshi, R. K., Megha, S., Basu, U., Rahman, M. H. & Kav, N. N. Genome wide identification and functional prediction of long non-coding RNAs responsive to Sclerotinia sclerotiorum Infection in Brassica napus. PloS One 11, e0158784 (2016).
Wen, M., Shen, Y., Shi, S. & Tang, T. miREvo: an integrative microRNA evolutionary analysis platform for next-generation sequencing experiments. BMC Bioinformatics 13, 140 (2012).
Friedlander, M. R., Mackowiak, S. D., Li, N., Chen, W. & Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Research 40, 37–52 (2012).
Nelson, A. D. et al. A genomic analysis of factors driving lincRNA diversification: lessons from plants. G3 6, 2881–2891 (2016).
Kapusta, A. et al. Transposable elements are major contributors to the origin, diversification, and regulation of vertebrate long noncoding RNAs. Plos Genet 9, e1003470 (2013).
Wu, H. J., Ma, Y. K., Chen, T., Wang, M. & Wang, X. J. PsRobot: a web-based plant small RNA meta-analysis toolbox. Nucleic Acids Research 40, W22–28 (2012).
Leitch, I. J. & Bennett, M. D. Genome downsizing in polyploid plants. Biological Journal of the Linnean Society 82, 651–663 (2004).
Renny-Byfield, S. et al. Diploidization and genome size change in allopolyploids is associated with differential dynamics of low- and high-copy sequences. The Plant Journal 74, 829–839 (2013).
Xie, F. & Zhang, B. microRNA evolution and expression analysis in polyploidized cotton genome. Plant Biotechnol J 13, 421–434 (2015).
Xie, F. L. & Zhang, B. H. microRNA evolution and expression analysis in polyploidized cotton genome. Plant Biotechnol J 13, 421–434 (2015).
Wang, M. J. et al. Long noncoding RNAs and their proposed functions in fibre development of cotton (Gossypium spp.). New Phytologist 207, 1181–1197 (2015).
Grover, C. E. et al. Homoeolog expression bias and expression level dominance in allopolyploids. New Phytologist 196, 966–971 (2012).
Samans, B., Chalhoub, B. & Snowdon, R. J. Surviving a genome collision: genomic signatures of allopolyploidization in the recent crop species Brassica napus. The Plant Genome 10 (2017).
Sunkar, R. & Jagadeeswaran, G. In silico identification of conserved microRNAs in large number of diverse plant species. BMC Plant Biology 8, 37 (2008).
Tian, J. et al. Population genomic analysis of gibberellin-responsive long non-coding RNAs in Populus. Journal of Experimental Botany https://doi.org/10.1093/jxb/erw057 (2016).
Liu, X., Hao, L., Li, D., Zhu, L. & Hu, S. Long non-coding RNAs and their biological roles in plants. Genomics, Proteomics & Bioinformatics 13, 137–147 (2015).
Shin, J. H. & Chekanova, J. A. Arabidopsis RRP6L1 and RRP6L2 function in FLOWERING LOCUS C silencing via regulation of antisense RNA synthesis. Plos Genet 10, e1004612 (2014).
Franco-Zorrilla, J. M. et al. Target mimicry provides a new mechanism for regulation of microRNA activity. Nature Genetics 39, 1033–1037 (2007).
Jabnoune, M. et al. A rice cis-natural antisense RNA acts as a translational enhancer for its cognate mRNA and contributes to phosphate homeostasis and plant fitness. The Plant Cell 25, 4166–4182 (2013).
Wang, T. Z., Liu, M., Zhao, M. G., Chen, R. & Zhang, W. H. Identification and characterization of long non-coding RNAs involved in osmotic and salt stress in Medicago truncatula using genome-wide high-throughput sequencing. BMC Plant Biology 15, 131 (2015).
Liang, Y. et al. MYB97, MYB101 and MYB120 function as male factors that control pollen tube-synergid interaction in Arabidopsis thaliana fertilization. Plos Genet 9, e1003933 (2013).
Nagpal, P. et al. Auxin response factors ARF6 and ARF8 promote jasmonic acid production and flower maturation. Development 132, 4107–4118 (2005).
Rieseberg, L. H. & Blackman, B. K. Speciation genes in plants. Ann Bot-London 106, 439–455 (2010).
Bomblies, K. & Weigel, D. Hybrid necrosis: autoimmunity as a potential gene-flow barrier in plant species. Nature Reviews Genetics 8, 382–393 (2007).
Wang, X. et al. The genome of the mesopolyploid crop species Brassica rapa. Nature Genetics 43, 1035–1039 (2011).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biology 10 (2009).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Guttman, M. et al. Ab initio reconstruction of cell type-specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincRNAs. Nature Biotechnology 28, 503–510 (2010).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nature Biotechnology 28, 511–515 (2010).
Kong, L. et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Research 35, W345–W349 (2007).
Punta, M. et al. The Pfam protein families database. Nucleic Acids Research 40, D290–301 (2012).
Bateman, A. et al. The Pfam protein families database. Nucleic Acids Research 30, 276–280 (2002).
Wiese, K. C. & Hendriks, A. Comparison of P-RnaPredict and mfold - algorithms for RNA secondary structure prediction. Bioinformatics 22, 934–942 (2006).
Jian, H. J. et al. Identification of rapeseed microRNAs involved in early stage seed germination under salt and drought stresses. Frontiers in Plant Science 7 (2016).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Research 19, 1639–1645 (2009).
Dai, X. B. & Zhao, P. X. psRNATarget: a plant small RNA target analysis server. Nucleic Acids Research 39, W155–W159 (2011).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Molecular Biology and Evolution 24, 1586–1591 (2007).
Zhou, L. et al. Integrated profiling of microRNAs and mRNAs: microRNAs located on Xq27.3 associate with clear cell renal cell carcinoma. PloS One 5, e15224 (2010).
Acknowledgements
This work was supported financially by the National Key Research and Development Plan (code: 2016YFD0101305), the earmarked fund for Modern Agro-industry Technology Research System of China (CARS-13), Science and Technology “Three Aid” Project of Jiangxi Province (code: 20133BFB29005), Scientific observing and experimental station of oil crops in the upper Yangtze River, Ministry of Agriculture, P. R. China (09203020) and the Sichuan Crop Breeding Community. ASM is supported by an Emmy Noether DFG grant (MA6473/1-1).
Author information
Authors and Affiliations
Contributions
D.F. conceived the project; L.J. and D.F. designed the experiments; J.Z., L.W., J.J. and H.L. analyzed the transcriptome data; A.S.M. revised the manuscript; J.Z., L.W., C.C., L.C. and B.Z. analyzed the lncRNA data; Y.Z. and Q.X. performed the qRT-PCR; J.Z. and L.W. wrote the article; L.J. and D.F. supervised and complemented the writing.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, J., Wei, L., Jiang, J. et al. Genome-wide identification, putative functionality and interactions between lncRNAs and miRNAs in Brassica species. Sci Rep 8, 4960 (2018). https://doi.org/10.1038/s41598-018-23334-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-23334-1
- Springer Nature Limited
We’re sorry, something doesn't seem to be working properly.
Please try refreshing the page. If that doesn't work, please contact support so we can address the problem.