Abstract
The emergence of carbapenemase-producing Citrobacter freundii poses a significant threat to public health worldwide. Here, we reported a C. freundii strain CWH001 which was resistant to all tested antimicrobials except tetracycline. Whole genome sequencing and analysis were performed. The strain, which belonged to a new sequence type ST139, showed close relationship with other foreign C. freundii strains through phylogenetic analysis. A novel variant of the intrinsic blaCMY gene located on the chromosome was identified and designated as blaCMY-152. Coexistence of blaNDM-1 with qnrS1 was found on a conjugative IncN plasmid, which had a backbone appearing in various plasmids. Other class A ESBL genes (blaVEB-3 and blaTEM-1) were also detected on two different novel plasmids. The emergence of multidrug-resistant C. freundii is of major concern, causing great challenges to the treatment of clinical infections. Great efforts need to be taken for the specific surveillance of this opportunistic pathogen.
Similar content being viewed by others
Introduction
Citrobacter freundii, a gram-negative bacterium of the Enterobacteriaceae family, is often the causative pathogen of a wide spectrum of nosocomial infections involving the respiratory tract1, urinary tract2 and bloodstream3. Previous studies have also reported its association with neonatal meningitis and brain abscess of high mortality4. Multidrug resistance in opportunistic pathogen C. freundii raised particular concern considering the severe dependence of immunocompromised patients on antibiotics5, and posed a significant threat to patient care and public health.
New Delhi metallo-β-lactamase 1 (NDM-1), a mediator of carbapenem resistance, had spread across different members of Enterobacteriaceae6 including C. freundii since its first identification in 20097. The occurrence of blaNDM-1-positive C. freundii has been increasingly reported in China8,9,10,11, India12,13, Denmark14 and South Africa15. The majority of C. freundii with NDM-1 were often co-resistant to multiple antimicrobial agents, but usually remained susceptible to amikacin, gentamicin and fosfomycin9,10,11.
In this study, we report an NDM-1-producing C. freundii strain, which showed extensive resistance to nearly all tested antibiotics. Whole genome sequencing and analysis were performed to gain an insight into its genetic features and plasmid profiles.
Results
Microbiological and genetic features of strain CWH001
Strain CWH001 was recovered from the blood sample of a patient through routine surveillance in Wuhan, China, in 2014. The strain was identified as C. freundii using Vitek 2 compact system and confirmed by 16S rDNA sequencing. CWH001 was resistant to nearly all tested antibiotics including aminoglycosides, cephalosporins, carbapenems, fluoroquinolones and sulfonamides, but remained susceptible to tetracycline (Table 1). PCR amplification and sequencing confirmed the presence of blaNDM-1. S1 pulsed field gel electrophoresis (PFGE) showed that CWH001 contained three different plasmids (~60 kb, ~105 kb and ~220 kb) (Fig. 1). Southern blotting revealed that the blaNDM-1 gene was carried by the ~60 kb plasmid, which was transferable to Escherichia coli J53 at a high transfer frequency of 2.21 × 10−2 per donor cell. The transconjugants acquired resistance to amoxicillin-clavulanic acid, piperacillin, imipenem and meropenem. Interestingly, subsequent sequencing and southern blotting revealed that there existed the blaVEB-3 gene on the ~220 kb plasmid, which was transferred to the transconjugants simultaneously. A BLAST search indicated that the 3.2 kb blaVEB-3-carrying contig was composed of a novel combination of Klebsiella pneumoniae JM45 plasmid p1 (CP006657, unpublished) and uncultured bacterium plasmid pKAZ516. The presence of the blaNDM-1 and blaVEB-3 genes in the transconjugants was further confirmed by PCR amplification and sequencing.
S1-PFGE pattern for strain CWH001 and southern blotting for the blaNDM-1, blaTEM-1 and blaVEB-3 genes. Lanes: Marker, Salmonella serotype Braenderup strain H9812 as a reference size standard; 1, PFGE result for S1-digested plasmid DNA of strain CWH001; 2–4, southern blot hybridization with the probes specific to blaNDM-1, blaTEM-1 and blaVEB-3, respectively. Full length S1-PFGE and southern blotting results are presented in Supplementary Fig. S1.
In addition to blaNDM-1 and blaVEB-3, other resistance genes were also identified in strain CWH001 including blaTEM-1, qnrS1, dfrA12, armA, fosA3, mphA, sul1, aac(3)-IId and a novel variant of the blaCMY gene. Analysis of the deduced protein sequence of the blaCMY variant revealed a single amino acid substitution at position 22 (Thr → Ala) relative to that of CMY-41. This variant protein was designated CMY-152 (http://www.lahey.org/Studies/webt.asp). BLAST search and southern blotting revealed the blaCMY-152 gene, together with its regulator gene ampR flanked by the upstream frd genes and the downstream blc gene, was located on the chromosome.
Molecular typing and phylogenetic analysis
The NDM-1-producing C. freundii CWH001 did not belong to an existing sequence type and was assigned to a new ST, ST139, using the multi-locus sequence typing (MLST) web server. Phylogenetic analysis revealed a high degree of genetic diversity of 84 available C. freundii genomes with that of CWH001. CWH001 was clustered into clades with overseas strains, and had a close relationship with strain 5-172-05_S1_C1 from Tanzania (Fig. 2). Only 543 SNPs were detected between the chromosomes of strain CWH001 and 5-172-05_S1_C1. However, CWH001 fell into different clades and showing distant phylogenetic relationship to other domestic strains. Sequence alignments revealed the average nucleotide identity (ANI) between CWH001 and other isolates from China ranged from 92.20% to 98.52%, while the ANI between CWH001 and 5-172-05_S1_C1 was 99.50%, indicating a different evolutionary pathway of CWH001 from other domestic strains in China.
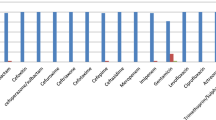
Phylogenetic tree and resistance gene profile of C. freundii strain CWH001 with other 84 available C. freundii genomes from GenBank. Strain CWH001 is indicated in red. Isolates from China are marked with a solid circle. NDM-harboring isolates are marked with an asterisk. The CWH001-including cluster is indicated with the dashed lines. Twenty-five closely related clones from the same geographical source (Houston, USA) are indicated as the RU2 cluster. Distribution of resistance genes is indicated by the heatmap according to the legend, which reflects percentage coverage of each gene sequence.
Characterization of bla NDM-1-carrying plasmid pNDM-CWH001
The 59-kb plasmid carrying blaNDM-1 was completely assembled and designated as pNDM-CWH001. pNDM-CWH001 belonged to the incompatibility type IncN. BLAST search against NCBI revealed that pNDM-CWH001 showed 100% coverage and >99% identity to the E. coli plasmid pNDM-BTR17 from China. Two single nucleotide deletions located within virB4 and virB8, respectively, were identified in pNDM-BTR. pNDM-CWH001 consisted of a blaNDM-1-containing transposon Tn6360 and a 42.3-kb backbone (Fig. 3a). Tn6360 was composed of an accessory region carrying blaNDM-1 and an intact Tn6292 element carrying qnrS1 (Fig. 3b). The accessory region comprised an IS26, a 427-bp truncated tnpA, and an 8.3-kb Tn3000 remnant (IS3000-ΔISAba125-blaNDM-1-ble-trpF-tat-ΔcutA1-groES-ΔgroEL). Compared with the prototype Tn300018, the remnant had undergone a deletion of the second copy of IS3000 together with the truncation of groEL in the 3′ extremity, suggesting a possible transposition event. The transposon Tn6292 had a quinolone resistance genetic platform organized as IS26-qnrS1-ISkpn19, which has been repeatedly reported in previous plasmids19,20 and was likely introduced due to the inter-plasmid transfer as a transposable element21.
The comparative schematic diagram of (a) plasmids R46, pIMP-GZ1058, pNDM-CWH001, pNDM-BTR and pMR3-OXA181; (b) the accessory modules Tn6292 in pIMP-GZ1058, Tn6360 in pNDM-CWH001 and pNDM-BTR, and Tn6361 in pMR3-OXA181. The open reading frames are indicated by arrows. The brown, cyan, purple, orange and green arrows represent genes associated with replication, antirestriction, stability, conjugation and type IV secretion system, respectively. The blaNDM-1 gene is shown in red. The qnrS1 gene is shown in golden. The accessory modules are shown in blue. The 1-kb inversion region and insertion of IS26 are shown in pink. Other genes of the backbone are shown in dark gray. Homology regions among different plasmids are denoted by light gray.
The backbone of pNDM-CWH001 also presented >98% identity to those of pMR3-OXA18122 (100% coverage) and pIMP-GZ105823 (92% coverage). The backbone contained a set of core genes for plasmid replication (repA), conjugation (tra genes), stability (stdB), antirestriction (ardA and klcA) and type IV secretion system (virB genes). However, there existed an inversion of a 1-kb region in plasmid pNDM-CWH001 and pNDM-BTR, which encoded aldehyde dehydrogenase and transcriptional regulator. An additional IS26 was inserted following this inversion region. The blaNDM-1-carrying transposon Tn6360 was integrated into the fipA gene, which was interrupted into two fragments in pNDM-CWH001 compared to plasmid R4624 and may serve as a “hotspot” for insertion of transposable elements.
Genetic features of plasmid pTEM-CWH001
The blaTEM-1 gene was located on a novel plasmid designated as pTEM-CWH001, which had the length of 107,391 bp and comprised a combination of C. freundii plasmid p112298-KPC9 and Salmonella enterica plasmid pF847525. pTEM-CWH001 could not be assigned to any known incompatibility group. The deduced replication initiator RepA presented >98% amino acid similarity with various IncFII family RepA proteins from Citrobacter. An insertion of ISEc42 between conjugal transfer genes tra and trb were observed, which was likely to impair the expression of the trbABC operon and may result in a non-transferable plasmid. pTEM-CWH001 harbored a Tn21-like structure bound by the transposition genes (tnpAR) and the mer operon in the 5′ and 3′ portion, respectively. The blaTEM-1 gene and an insertion sequence ISCfr1 were located upstream of the Tn21-like structure. Compared with the prototype Tn21, this structure had undergone the replacement of aadA1 by dfrA12 and an insertion of a macrolide resistance operon organized as mphA-mrx-mphR in the class 1 integron In2, suggesting possible frequent transposition events.
Discussion
The ability to produce NDM-1 carbapenemases has been acquired by diverse Enterobacteriaceae species and posed a significant threat to public health. Our study identified a blaNDM-1-positive C. freundii isolate with coexistence of other multiple resistant determinants (blaVEB-3, blaTEM-1 and blaCMY-152) and provided detailed genetic characteristics of the NDM-1-carrying IncN plasmid pNDM-CWH001. Plasmids belonging to the IncN group are typically broad-host-range and self-conjugative26. The high transfer frequency of pNDM-CWH001 demonstrated its great potential to transfer across species. The resistance-determining region in those pNDM-CWH001-like plasmids was all inserted within the fipA gene. Interestingly, the fipA-encoded protein was reported to inhibit the conjugal transfer of some plasmids27. The interruption of the fipA gene could facilitate the ability of the plasmids of the plasmids to accumulate in diverse hosts and may serve as a “hotspot” for integration of mobile elements. Comparative analysis revealed that the acquisition of Tn6292 and the Tn3000 remnant might be subsequently integrated into pNDM-CWH001-like plasmids, highlighting the urgency of further surveillance and genetic analysis of such flexible mobile units for better understanding of extensive resistance dissemination.
Recent studies have reported the simultaneous presence of multiple resistance genes in C. freundii strains isolated in China8,9,11. However, CWH001 showed long-distance dispersals from other C. freundii isolates in China and gained some resistance determinants (blaVEB-3 and fosA3) that were rarely identified in other C. freundii isolates. Previous study has reported C. freundii strain WCHCF65 from China clustered with strains from Denmark8. Phylogenetic analysis revealed that domestic C. freundii isolates showed close relationship with overseas ones but fell into distinct clusters, indicating different evolution and dissemination route. Plasmid pNDM-BTR was isolated from Beijing in 2013. Though lacking of epidemiological association, the close spatial and temporal proximity between pNDM-CWH001 and pNDM-BTR in China suggested possible dissemination of this novel plasmid, and more attention should be devoted to monitoring the epidemic spread of such blaNDM-1-carrying IncN plasmids among Enterobacteriaceae.
In summary, our study characterized a multidrug-resistant C. freundii isolate harboring multiple ESBL-encoding genes. Strain CWH001 belonged to a novel sequence type ST139 with a self-transferable plasmid pNDM-CWH001, which may facilitate the blaNDM-1 gene dissemination. Phylogenetic analysis revealed that CWH001 had different origin from domestic isolates but gained multidrug resistance. Our findings further emphasize the threat of NDM-1 carbapenemase circulation among diverse species, and urgent actions should be taken to control the potential rapid spread of such plasmids.
Materials and Methods
Bacterial isolation and identification
The blaNDM-1-positive C. freundii strain CWH001 was recovered from the blood sample of a 63-year-old male patient through routine surveillance in Wuhan, China, in 2014. The species level identification was performed by using Vitek 2 compact system (bioMérieux, France) and confirmed by 16S rDNA sequencing28. The presence of genes encoding carbapenemases and ESBLs was determined by PCR and sequencing29,30,31. The entire blaNDM, blaTEM, blaVEB and blaCMY genes were amplified with previously described primers32,33,34,35. Positive PCR results were further confirmed by sequencing. The informed consent was obtained from the patient. All experimental protocols were approved by Institutes of Military Medicine, Academy of Military Sciences. The methods were carried out in accordance with relevant guidelines.
Antimicrobial susceptibility testing
The minimal inhibitory concentrations (MICs) of amoxicillin/clavulanic acid (AMC), piperacillin (PIP), cefazolin (FAZ), ceftazidime (CAZ), ceftriaxone (CTR), cefepime (FEP), aztreonam (AZT), imipenem (IMI), meropenem (MEC), amikacin (AMI), gentamicin (GEN), ciprofloxacin (CIP), levofloxacin (LVX), tetracycline (TET), nitrofurantoin (NIT) and sulfamethoxazole/trimethoprim (SXT) were determined by Vitek 2 compact system (BioMérieux, France) following the manufacturer’s instructions. The results were interpreted following the Clinical and Laboratory Standards Institute (CLSI) guidelines36.
Southern blotting and Conjugation experiment
Genomic DNA from strain CWH001 was prepared in agarose plugs and digested with the S1 endonuclease (Takara, Dalian, China). DNA fragments were separated by PFGE through a CHEF-DR III system (Bio-Rad, Hercules, USA). The plasmid DNA was transferred to a positively charged nylon membrane (Roche) and hybridized with the digoxigenin-labeled probes specific to blaNDM-1, blaTEM-1, blaVEB-3 and blaCMY-152.
Conjugation experiment was carried out by broth and filter mating using the clinical strain CWH001 as donors and azide-resistant E. coli strain J53 as the recipient. The donor and recipient cultures were mixed at a ratio of 1:3 in LB broth and incubated at 37 °C for 18 hours. The mixture was inoculated into MacConkey agar plates containing 4 μg/ml meropenem and 150 μg/ml sodium azide. The transconjugants were selected after 12 h of incubation. Horizontal transferability of drug resistance was assessed by antimicrobial susceptibility testing and the transconjugants carrying resistant markers (blaNDM-1, blaVEB-3) were confirmed by PCR amplification.
Whole genome sequencing and phylogenetic analysis
Total DNA was extracted from cultured bacterium using the QIAamp DNA minikit (Qiagen, Inc., Valencia, CA). Sequencing was carried out using an Illumina HiSeq. 2500 platform with a 350-bp insert size at Novogene Company (Beijing, China). The genome was assembled de novo using SOAPdenovo (v2.04)37 with an average 110-fold coverage. Scaffolding and gap filling were performed using SSPACE and GapFiller38,39. Plasmids pNDM-BTR, p112298-KPC and pF8475 were selected as reference. Gaps were closed using reference-guided assembly and manually checked by re-mapping raw reads against the plasmids. Genome sequences were annotated using RAST40. Plasmid replicon type was identified using PlasmidFinder41 (Enterobacteriaceae).
Seven housekeeping genes (arca-aspc-clpx-dnag-fadd-lysp-mdh) were extracted from the genome of CWH001 and used in MLST typing through the MLST web server42. Genome sequences of 84 currently available C. freundii isolates were downloaded from the NCBI database for phylogenetic analysis (accessed 12th June 2017). C. freundii strain B38 (GenBank accession number CP016762) was used as the reference for comparison. Reads mapping was performed using BWA (v0.7.12)43. SNPs were identified using SAMtools (v1.3)44. The resulting 27366 SNPs were concatenated and aligned to construct the Maximum-Likelihood phylogenetic tree using RAxML (v8.2.4) with the general time reversible (GTR) model and a gamma distribution45. ANIs between CWH001 and other genomes were calculated using JSpeciesWS46 to evaluate the genome similarity.
Nucleotide sequence accession number
The shotgun whole genome sequence of strain CWH001 and complete sequence of plasmids pNDM-CWH001 and pTEM-CWH001 have been deposited in NCBI GenBank under accession number PEHH00000000.
References
Fujii, T. et al. Coexisting respiratory tract infection and bacteremia or sepsis caused by the same bacterium. Kansenshogaku Zasshi 68, 217–225 (1994).
Gill, M. A. & Schutze, G. E. Citrobacter urinary tract infections in children. Pediatr. Infect. Dis. J. 18, 889–892 (1999).
Pfaller, M. A. et al. Inducible amp C beta-lactamase producing gram-negative bacilli from blood stream infections: frequency, antimicrobial susceptibility, and molecular epidemiology in a national surveillance program (SCOPE). Diagn. Microbiol. Infect. Dis. 28, 211–219 (1997).
Joaquin, A., Khan, S., Russel, N. & al Fayez, N. Neonatal meningitis and bilateral cerebellar abscesses due to Citrobacter freundii. Pediatr Neurosurg 17, 23–24 (1991).
Bodey, G. P. Managing infections in the immunocompromised patient. Clinical Infectious Diseases 40, S239 (2005).
Wailan, A. M. & Paterson, D. L. The spread and acquisition of NDM-1: a multifactorial problem. Expert Rev Anti Infect Ther 12, 91–115 (2014).
Yong, D. et al. Characterization of a new metallo-beta-lactamase gene, bla(NDM-1), and a novel erythromycin esterase gene carried on a unique genetic structure in Klebsiella pneumoniae sequence type 14 from India. Antimicrob. Agents Chemother. 53, 5046–5054 (2009).
Wu, W., Espedido, B., Feng, Y. & Zong, Z. Citrobacter freundii carrying blaKPC-2 and blaNDM-1: characterization by whole genome sequencing. Sci Rep 6, 30670 (2016).
Feng, J. et al. Coexistence of a novel KPC-2-encoding MDR plasmid and an NDM-1-encoding pNDM-HN380-like plasmid in a clinical isolate of Citrobacter freundii. J. Antimicrob. Chemother. 70, 2987–2991 (2015).
Du, X.-X. et al. Genetic characteristics of blaNDM-1-positive plasmid in Citrobacter freundii isolate separated from a clinical infectious patient. J. Med. Microbiol. 62, 1332–1337 (2013).
Huang, Y.-M. et al. NDM-1-Producing Citrobacter freundii, Escherichia coli, and Acinetobacter baumannii Identified from a Single Patient in China. Antimicrob. Agents Chemother. 59, 5073–5077 (2015).
Dolejska, M., Villa, L., Poirel, L., Nordmann, P. & Carattoli, A. Complete sequencing of an IncHI1 plasmid encoding the carbapenemase NDM-1, the ArmA 16S RNA methylase and a resistance-nodulation-cell division/multidrug efflux pump. J. Antimicrob. Chemother. 68, 34–39 (2013).
Poirel, L. et al. Extremely drug-resistant Citrobacter freundii isolate producing NDM-1 and other carbapenemases identified in a patient returning from India. Antimicrob. Agents Chemother. 55, 447–448 (2011).
Hammerum, A. M. et al. Use of WGS data for investigation of a long-term NDM-1-producing Citrobacter freundii outbreak and secondary in vivo spread of blaNDM-1 to Escherichia coli, Klebsiella pneumoniae and Klebsiella oxytoca. J. Antimicrob. Chemother. 71, 3117–3124 (2016).
Rubin, J. E., Peirano, G., Peer, A. K., Govind, C. N. & Pitout, J. D. D. NDM-1-producing Enterobacteriaceae from South Africa: moving towards endemicity? Diagn. Microbiol. Infect. Dis. 79, 378–380 (2014).
Flach, C.-F. et al. Isolation of novel IncA/C and IncN fluoroquinolone resistance plasmids from an antibiotic-polluted lake. J. Antimicrob. Chemother. 70, 2709–2717 (2015).
Zhao, Y. et al. Structural genomics of pNDM-BTR harboring In191 and Tn6360, and other bla NDM-carrying IncN1 plasmids. Future Microbiol 12, 1271–1281 (2017).
Campos, J. C. et al. Characterization of Tn3000, a Transposon Responsible for blaNDM-1 Dissemination among Enterobacteriaceae in Brazil, Nepal, Morocco, and India. Antimicrob. Agents Chemother. 59, 7387–7395 (2015).
Chen, L. et al. Complete sequence of a KPC-producing IncN multidrug-resistant plasmid from an epidemic Escherichia coli sequence type 131 strain in China. Antimicrob. Agents Chemother. 58, 2422–2425 (2014).
Liu, Y. et al. First Report of OXA-181-Producing Escherichia coli in China and Characterization of the Isolate Using Whole-Genome Sequencing. Antimicrob. Agents Chemother. 59, 5022–5025 (2015).
Sumrall, E. T., Gallo, E. B., Aboderin, A. O., Lamikanra, A. & Okeke, I. N. Dissemination of the transmissible quinolone-resistance gene qnrS1 by IncX plasmids in Nigeria. PLoS ONE 9, e110279 (2014).
McGann, P. et al. War wound treatment complications due to transfer of an IncN plasmid harboring bla(OXA-181) from Morganella morganii to CTX-M-27-producing sequence type 131 Escherichia coli. Antimicrob. Agents Chemother. 59, 3556–3562 (2015).
Wang, Y. et al. IncN ST7 epidemic plasmid carrying blaIMP-4 in Enterobacteriaceae isolates with epidemiological links to multiple geographical areas in China. J. Antimicrob. Chemother. 72, 99–103 (2017).
Hall, R. M. & Vockler, C. The region of the IncN plasmid R46 coding for resistance to beta-lactam antibiotics, streptomycin/spectinomycin and sulphonamides is closely related to antibiotic resistance segments found in IncW plasmids and in Tn21-like transposons. Nucleic Acids Res. 15, 7491–7501 (1987).
Kubasova, T. et al. Antibiotic Resistance, Core-Genome and Protein Expression in IncHI1 Plasmids in Salmonella Typhimurium. Genome Biol Evol 8, 1661–1671 (2016).
Götz, A. et al. Detection and characterization of broad-host-range plasmids in environmental bacteria by PCR. Appl. Environ. Microbiol. 62, 2621–2628 (1996).
Santini, J. M. & Stanisich, V. A. Both the fipA gene of pKM101 and the pifC gene of F inhibit conjugal transfer of RP1 by an effect on traG. J. Bacteriol. 180, 4093–4101 (1998).
Frank, J. A. et al. Critical evaluation of two primers commonly used for amplification of bacterial 16S rRNA genes. Appl. Environ. Microbiol. 74, 2461–2470 (2008).
Zong, Z. & Zhang, X. blaNDM-1-carrying Acinetobacter johnsonii detected in hospital sewage. J. Antimicrob. Chemother. 68, 1007–1010 (2013).
Mendes, R. E. et al. Rapid Detection and Identification of Metallo-β-Lactamase-Encoding Genes by Multiplex Real-Time PCR Assay and Melt Curve Analysis. J Clin Microbiol 45, 544–547 (2007).
Cai, J. C., Zhang, R., Hu, Y. Y., Zhou, H. W. & Chen, G.-X. Emergence of Escherichia coli Sequence Type 131 Isolates Producing KPC-2 Carbapenemase in China. Antimicrob Agents Chemother 58, 1146–1152 (2014).
Zou, D. et al. A novel New Delhi metallo-β-lactamase variant, NDM-14, isolated in a Chinese Hospital possesses increased enzymatic activity against carbapenems. Antimicrob. Agents Chemother. 59, 2450–2453 (2015).
Pasterán, F. et al. Emergence of PER-2 and VEB-1a in Acinetobacter baumannii Strains in the Americas. Antimicrob. Agents Chemother. 50, 3222–3224 (2006).
Briñas, L., Zarazaga, M., Sáenz, Y., Ruiz-Larrea, F. & Torres, C. Beta-lactamases in ampicillin-resistant Escherichia coli isolates from foods, humans, and healthy animals. Antimicrob. Agents Chemother. 46, 3156–3163 (2002).
Zhang, X. et al. First identification of coexistence of blaNDM-1 and blaCMY-42 among Escherichia coli ST167 clinical isolates. BMC Microbiol. 13, 282 (2013).
Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing: twenty-third informational supplement. M100-S23. Clinical and Laboratory Standards Institute, Wayne, PA (2013).
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010).
Boetzer, M., Henkel, C. V., Jansen, H. J., Butler, D. & Pirovano, W. Scaffolding pre-assembled contigs using SSPACE. Bioinformatics 27, 578–579 (2011).
Boetzer, M. & Pirovano, W. Toward almost closed genomes with GapFiller. Genome Biol. 13, R56 (2012).
Aziz, R. K. et al. The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9, 75 (2008).
Carattoli, A. et al. In silico detection and typing of plasmids using Plasmid Finder and plasmid multilocus sequence typing. Antimicrob. Agents Chemother. 58, 3895–3903 (2014).
Larsen, M. V. et al. Multilocus sequence typing of total-genome-sequenced bacteria. J. Clin. Microbiol. 50, 1355–1361 (2012).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Richter, M., Rosselló-Móra, R., Oliver Glöckner, F. & Peplies, J. JSpeciesWS: a web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics 32, 929–931 (2016).
Acknowledgements
This work was supported by grants from the Mega-projects of Science and Technology Research (No. 2017ZX10303405-003), the Beijing Nova program (No. xx2018042), the Beijing Natural Science Foundation (No. 5172029) and the National Nature Science Foundation of China (No. 31200942).
Author information
Authors and Affiliations
Contributions
L.Y., P.L., B.L. and X.H. performed genome analysis and experiment. J.L. collected samples. J.X., C.Y. and R.H. performed bacterial culture and DNA extraction. L.W. and L.J. performed library construction and genome sequencing. L.Y. and P.L. prepared the manuscript. P.L., S.Q. and H.S. designed the study and revised the manuscript. All authors contributed to review and revision, and approved the final version.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yang, L., Li, P., Liang, B. et al. Multidrug-resistant Citrobacter freundii ST139 co-producing NDM-1 and CMY-152 from China. Sci Rep 8, 10653 (2018). https://doi.org/10.1038/s41598-018-28879-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-28879-9
- Springer Nature Limited
This article is cited by
-
Complete-genome sequencing and comparative genomic characterization of blaNDM-5 carrying Citrobacter freundii isolates from a patient with multiple infections
BMC Genomics (2023)
-
Genomic characterization of Citrobacter freundii strains coproducing OXA-48 and VIM-1 carbapenemase enzymes isolated in leukemic patient in Spain
Antimicrobial Resistance & Infection Control (2019)






