Abstract
Microglia, the main resident immune cells in the CNS, are thought to participate in the pathogenesis of various neurological disorders. LPS and LPS + IFNγ are stimuli that are widely used to activate microglia. However, the transcriptomic profiles of microglia treated with LPS and LPS + IFNγ have not been properly compared. Here, we treated murine primary microglial cultures with LPS or LPS + IFNγ for 6 hours and then performed RNA-Sequencing. Gene expression patterns induced by the treatments were obtained by WGCNA and 11 different expression profiles were found, showing differential responses to LPS and LPS + IFNγ in many genes. Interestingly, a subset of genes involved in Parkinson’s, Alzheimer’s and Huntington’s disease were downregulated by both treatments. By DESeq analysis we found differentially upregulated and downregulated genes that confirmed LPS and LPS + IFNγ as inducers of microglial pro-inflammatory responses, but also highlighted their involvement in specific cell functions. In response to LPS, microglia tended to be more proliferative, pro-inflammatory and phagocytic; whereas LPS + IFNγ inhibited genes were involved in pain, cell division and, unexpectedly, production of some inflammatory mediators. In summary, this study provides a detailed description of the transcriptome of LPS- and LPS + IFNγ treated primary microglial cultures. It may be useful to determine whether these in vitro phenotypes resemble microglia in in vivo pathological conditions.
Similar content being viewed by others
Introduction
Microglia are the main cells of myeloid origin in the central nervous system (CNS). They are considered tissue-resident macrophages that constantly patrol the cerebral microenvironment and respond to pathogens and cell damage1. Microglia responses to changes in homeostasis are highly plastic and multifaceted: finely tuned by the nature of the disturbance2. One of the most widely used stimuli to elicit inflammatory microglia is lipopolysaccharide (LPS) alone or in combination with interferon gamma (IFNγ).
LPS is the main structural component of the outer membrane of Gram-negative bacteria. It is a heat-stable, amphiphilic molecule comprising three regions: lipid A, a core oligosaccharide and an O side chain3. The primary receptor of LPS is Tlr4, which is a pivotal contributor to the microglial response to endotoxins, and is involved in LPS-mediated neurodegeneration4. In vitro, LPS-treated microglia produce pro-inflammatory mediators, such as Tnf, Il-1β, prostaglandin E2, NO and reactive oxygen species (ROS) (reviewed in Schwartz et al.5). LPS has been used to study the involvement of neuroinflammation in experimental models of Parkinson’s and Alzheimer’s disease, as well as amyotrophic lateral sclerosis (reviewed in Ransohoff & Perry6 and Nayak et al.7). Specifically, neuron-microglia co-cultures treated with LPS are a powerful tool to study the mechanisms involved in the dopaminergic neuronal death observed in Parkinson’s disease. In vivo, injection of LPS into the substantia nigra also results in the progressive and irreversible loss of dopaminergic neurons. In both cases, LPS triggers rapid activation of microglia, which precedes neuronal death. Therefore, LPS-treated microglia constitute a suitable system for the screening of therapeutic targets against neurodegeneration8.
IFNγ mediates immune and inflammatory responses by activating transcription factor STAT1 through the JAK-STAT signalling pathway. Interestingly, many IFNγ functions are mediated by cross-regulation of cellular responses to other inflammatory stimuli9. Therefore, IFNγ has been used together with LPS to mimic a microglial activation state that could be induced in vivo by the co-presence of damage-associated molecular patterns acting on TLRs, together with IFNγ produced by CNS cells. LPS + IFNγ treatment induces the production of pro-inflammatory mediators in the BV2 microglial cell line and in primary microglial cells10. However, whereas in some cases IFNγ potentiates the effect of LPS, e.g., NO production; in others, IFNγ counteracts it, e.g., Ptges and Csf3 expression11. These results indicate that two stimuli are capable of activating different pathways and they are not redundant.
Since modelling neuroinflammation and microglial activation in vitro constitutes a valuable tool to study the role of microglia under normal and pathological conditions, it is important to know how LPS and LPS + IFNγ affect microglia. In order to analyse the expression pattern of genes affected by these stimuli, we performed RNA-Sequencing analysis (RNA-Seq) on primary microglial cells stimulated with LPS or LPS + IFNγ. RNA-Seq is increasingly used to determine gene expression, since it provides unbiased expression profiles, identifies novel transcribed regions and is extremely accurate in comparison to RNA microarrays12.
Different methodologies have been developed to analyze RNA-Seq data in recent years. Here, we use DESeq to detect differentially expressed genes (DEGs), i.e., genes that are significantly upegulated or downregulated in pairwise comparisons, in murine primary microglia treated with vehicle, LPS or LPS + IFNγ. Additionally, we also use a different data analysis approach termed weighted gene correlation network analysis (WGCNA), which measures the correlation among genes, regardless of significance. Furthermore, since this approach compares all the genes in the data, it can infer complex expression patterns that could easily be overlooked in the pairwise differential expression analysis. Both, DESeq and WGCNA revealed that not all genes are equally affected by LPS and LPS + IFNγ. Therefore, our results confirm that the two stimuli generate distinct gene expression patterns and can constitute useful tools to induce different profiles of inflammation in microglia.
Materials and Methods
Animals
Phenotypically wild-type C/EBPβfl/fl mice on a C57BL/6J background (Jackson Laboratories) were breed and housed under specific pathogen-free conditions in the animal facilities at the School of Medicine, University of Barcelona. They received food and water ad libitum. All procedures were approved by the Ethical Committee for Animal Experimentation of the University of Barcelona and by the Commission for Animal Experimentation of the Generalitat de Catalunya, with protocols numbers DAAM 5026, 7064 and 7065.
Primary microglial cultures
Microglial primary cultures were prepared as described previously11. Briefly, cortices from P1-P3 mice were dissected, carefully stripped of their meninges and digested with 0.25% trypsin for 30 min at 37 °C. Trypsinization was stopped by adding an equal volume of culture medium (Dulbecco’s modified Eagle medium-F-12 nutrient mixture, fetal bovine serum 10%, penicillin 100 U/mL, streptomycin 100 μg/mL and amphotericin B 0.5 μg/mL) with 160 µg/mL deoxyribonuclease I, all from Invitrogen. Cells were brought to a single cell suspension by repeated pipetting followed by passage through a 100 μm-pore mesh and pelleted (7 min, 200 g). Glial cells were resuspended in culture medium and seeded at a density of 3.5 × 105 cells/mL. Cultures were maintained at 37 °C in humidified 5% CO2–95% air. Medium was replaced once a week. At DIV 20, mixed glial cultures were treated with diluted trypsin13 to obtain >98% pure microglial cultures. Microglial cells were used 24 h after isolation.
Cell treatment and total RNA isolation
Three independent microglial cultures were treated for 6 h with vehicle (cell culture medium) and 100 ng/mL of LPS (026:B6 Escherichia coli serotype, Sigma Aldrich) with or without 1 ng/mL of recombinant mouse IFNγ (I4777, Sigma Aldrich). Then, total RNA was isolated by lysing pelleted microglial cells from one 75 cm2 flask per treatment condition with 1 mL of TriReagent (Sigma Aldrich) and 100 µl of 1-bromo-3-chloropropane (BCP, Sigma Aldrich). The aqueous phase containing total RNA was recovered after centrifuging for 15 min at 12.000 g at 4 °C, mixed with and equal volume of ice-cold 70% Ethanol and loaded onto a PureLinkTM Micro Kit column (Invitrogen). Total RNA was then purified following manufacturer’s instructions. Total RNA was quantified spectrophotometrically using a Nano Drop ND-1000 (Thermo Scientific) and its integrity and quality were assessed with the Bioanalyzer 2100 system (Agilent). Only samples with an RNA integrity number (RIN) greater than 8 were used subsequently for RNAseq and qRT-PCR analyses.
qRT-PCR
300 ng of total RNA were used to carry out reverse transcription reactions with random primers using Transcriptor Reverse Transcriptase (Roche). Then, cDNA was diluted 1/30 and 3 µl were used to perform quantitative real-time PCR (qRT-PCR) with qPCRBIO Sygreen Mix Lo-ROX (PCB-P20.11–50, Vitro) in 15 µl of final volume reaction using CFX96 Thermal Cycler equipment (Bio-Rad). Measurements were performed in duplicates. Primers, shown in Table 1, were used at final concentration of 300 nM. Actin-β and Rn18s expression was not altered by treatments (data not shown). Samples were run for 45 cycles (95 °C for 30 seconds, 60 °C or 62 °C for 1 minute and 72 °C for 30 seconds). Amplification specificity was confirmed by the analysis of melting curves and by agarose-gel electrophoresis. Relative gene expression values were calculated using Bio-Rad CFX Managing software (Bio-Rad) with the comparative Ct or ΔΔCt method.
cDNA library preparation and RNA-Seq
Following Illumina’s (San Diego, CA) protocols, cDNA library preparation and ultrasequencing were performed. The first step was removing transfer RNA (tRNA) and ribosomal RNA (rRNA) from 1 µg of total RNA using TruSeq Stranded Total RNA Sample Prep Kits (Illumina). Then, the RNA pool (mRNA, miRNA, lncRNA and other RNAs) was fragmented into pieces of approximately 200 bp. The cleaved RNA fragments were reverse transcribed into first strand cDNA using reverse transcriptase and random primers. Next, the second strand was synthesized using DNA polymerase I and RNAseH. These double-stranded cDNA fragments were end-repaired by T4 DNA polymerase and Klenow DNA polymerase, and phosphorylated by T4 polynucleotide kinase. The cDNA products were incubated with Klenow DNA polymerase to generate Adenine overhangs, therefore allowing ligation to Illumina indexing adapters to the double stranded cDNA ends. The adapter-ligated products were purified with Ampure XP magnetic beads (Agencourt Bioscience Corpo-ration, Beverly, MA, USA) and libraries were amplified by 15 cycles of PCR with Phusion DNA polymerase (Thermofisher, Finnzymes Reagents, Vantaa, Finland). Constructed libraries were validated and quantified using BioRad’s automated electrophoresis system Experion and qPCR respectively. Pools of 6 indexed libraries were mixed (multiplexed) at equimolar ratios to yield a total oligonucleotide mix concentration of 10 nM. Finally the resulting libraries were sequenced on the Genome Analyzer IIx platform (Illumina) to generate 150 bp single reads. Six pooled indexed libraries were sequenced in each flow cell lane. Raw sequence (FASTQ format) were processed through a series of sequential steps: (1) aggressive adapters removal; (2) alignment/mapping of RNA sequences to the mouse genome reference (Mus_musculus.mm10) using tophat software; and (3) sorting and cataloguing of the results by using Samtools software14 in BAM files.
Gene expression analysis using WGCNA
BAM files were processed using the Rsubread package15 in the R environment16 and summarized to the mm10 version of mouse genome with the function featureCounts. Summarized readings by gene were then normalized using voom normalization17 to fit the count matrix into linear modelling with the package limma18 and to obtain log2 RPKM values. Whole read counts normalized were used to cluster samples with package hcluster using standard hierarchical cluster with average linkage. Heatmap visualization and clustering of DEG’s was done with Genesis software after expressing gene values in standard deviation and using hierarchical clustering with average linkage by genes and samples. Genecodis19 tools (http://genecodis.cnb.csic.es/) were used to obtain Gene Ontology and KEGG pathway terms enrichment using the adjusted p-value < 0.01 after an hypergeometric test. Finally, Reads per kilobase per million mapped reads (RPKM) filtered values (sum of RPKM > 2 for each gene) were introduced to the Weighted Correlation Gene Network Analysis (WGCNA) package in R, to perform WGCNA as previously described20. Briefly, combined samples were measured with the pickSoftThreshold function to obtain a correct power β considering the smallest value to give a free-scale topology. Using 18 as the power value, the blockwise function was used with parameters mergeCutHeight = 0.2 to obtain modules containing genes with high co-expression similarity. WGCNA software then extracted the first principal component of the expression values of each module to obtain the module eigengene.
Differential Expressed Genes Analysis
To identify DEGs, the BAM files were analyzed using R-software packages easyRNAseq21 and DESeq22. The “Coverage vectors” for all genes of each sample were obtained using “easyRNAseq” and the UCSC.mm10 Mus musculus reference genome. Samples were grouped by phenotype (WT, WT-L, WT-L + I) and the expression for each locus. To find DEGs, this count matrix was analyzed using “DESeq” software, a method based on the negative binomial distribution, with variance and mean linked by local regression. For each phenotype comparison, the group base expression, the fold Regulation, the uncorrected p-value, and q-value (false discovery rate corrected p-value) for each gene were calculated. For each comparison, the list of DEGs with a q-value < 0.05 was analyzed using Genecodis tools to obtain Gene Ontology and KEGG pathway23,24,25 enrichment as previously described. For the Signalling Pathway Impact Analysis (SPIA) of the DEGs, the Clusterview web software26 (https://pathview.uncc.edu/analysis) was used for data integration and visualization of alterations in several KEGG inflammatory pathways (Toll-like receptor signalling, NF-kappa B signalling, TNF signalling, Cytokine-cytokine receptor interactions) using default parameters.
Statistical and Power analyses
Statistical analyses of qRT-PCR data were performed using one-way ANOVA followed by Bonferroni post-test. Values of p < 0.05 were considered statistically significant. Results were represented as mean ± standard error of the mean (SEM). Experimental data were analyzed using GraphPad Prism 5.01 software.
Differential gene expression and WGCNA analyses were calculated using DEseq and WGCNA statistical R-packages software. For Statistical power calculation, the RNAseqPS software27 (https://cqs.mc.vanderbilt.edu/shiny/RnaSeqSampleSize/) was used. Assumptions made were that prior data indicated that the minimum average read counts among the prognostic genes in the control group was 5, the maximum dispersion was 0.5 and the ratio of the geometric mean of normalization factors was 1. It was also supposed that the total number of genes for testing was 14500 and the top 300 genes were prognostic. Therefore, the present RNA sequencing experiment with 3 experimental subjects in each group can identify differential gene expression between two groups if the desired minimum fold change was 23.5. Then, it will be possible to reject the null hypothesis by using exact test with a statistical power of 80%. The FDR associated with this test was 0.01.
Literature Search
On 26th May 2018, the query terms “microglia AND RNA sequencing”, “microglia AND transcriptome” and “microglia AND RNA seq” retrieved 278, 205 and 54 results, respectively, from the Pubmed database (https://www.ncbi.nlm.nih.gov/pubmed/). The search was limited to studies published in English. A total of 7 reports have been selected according to their similarity in terms of treatment paradigm and microglial source to our study.
Results
RNA-Seq transcriptional analysis of primary microglia treated with LPS or LPS + FNγ
It is known that LPS alone or in combination with IFNγ, two widely used pro-inflammatory treatments, triggers significant changes in the microglia transcriptome. As depicted in Fig. 1A, Tnf, Csf3, Il-6, Cybb, Ptgs2 and Nos2 expression is induced in microglia treated with LPS or LPS + IFNγ for 6 h, in comparison with vehicle-treated cells. However, other genes, such as Ptgs1 and Cmklr1 are downregulated. Interestingly, the addition of IFNγ to LPS seems to compensate the expression of some LPS-induced genes, such as Tnf, in a statistically significant manner; and of others, like Csf3, Cybb, Ptgs1 and Cmklr1, merely as a non-significant trend. This suggests that the two stimuli are not always additive; indeed, they can be compensatory. To establish a high-resolution transcriptome profile of primary murine microglial cells in response to LPS and LPS + IFNγ, we performed RNA-Seq analysis. Three independent microglial cultures were treated with a vehicle, LPS or LPS + IFNγ; total RNA was isolated 6 h after treatment and analysed by RNA-Seq. RNA libraries and sequencing were prepared as described in the “Materials and Methods” section. Normalized data represented on a heatmap for the abovementioned genes show the same expression pattern as that observed in qRT-PCR experiments (Fig. 1B).
Microglial gene expression is affected by LPS alone or together with IFNγ, 6 h after treatment. (A) Expression of Tnf, Csf3, Il6, Cybb, Ptgs2, Nos2, Ptgs1 and Cmklr1 was analysed by qRT-PCR in primary microglial cultures treated with vehicle (C), LPS (100 ng/mL) or LPS + IFNγ (1 ng/mL) for 6 h (n = 4 independent experiments). Data are shown as mean + SEM. *p < 0.05; **p < 0.01; ***p < 0.001 compared with the respective control condition. ##p < 0.01; ###p < 0.001 compared with the corresponding LPS condition. (B) Heatmap of RNA-Seq data showing Tnf, Csf3, Il6, Cybb, Ptgs2, Nos2, Ptgs1 and Cmklr1 expression profiles. Values are represented as SD from the normalized average for each gene. Hierarchical Clustering (average linkage, Euclidean distance) grouped the samples into three clusters corresponding to vehicle (pink), LPS (magenta) and LPS + IFNγ (purple) conditions. Furthermore, the genes were grouped into two different clusters depending on whether the treatment increases (yellow) or decreases (orange) their expression.
The strong effect of the treatments on microglial RNA expression was revealed by unsupervised cluster analysis using normalized counts (reads per kilobase million, RPKM). The 9 samples were grouped into three clearly differentiated clusters corresponding to the microglial cultures treated with vehicle (n = 3), LPS (n = 3) and LPS + IFNγ (n = 3). No outliers were detected (Fig. 2A).
Treatment with LPS alone or together with IFNγ for 6 h induces 11 different expression patterns in primary microglia. (A) Hierarchical clustering of total RNA-Seq RPKM normalized values clearly separates the 9 samples according to treatment condition: Control, LPS and LPS + IFNγ. (B) WGCNA correlation dendrogram of genes. According to their expression pattern, the genes were grouped into 11 different modules identified by arbitrary colours, turquoise module was the largest and the green-yellow the smallest. Outliers or unclustered transcripts are grey. Genes grouped in the yellow, blue and green modules and half of the genes clustered in the black module are clustered separately from the genes present in the other modules. (C) Module eigengene barplots represent how expression of the genes clustered in the 12 modules was affected by treatments. (D) To account for the different patterns of gene expression, two models of treatment order were fitted to the WGCNA. Left column. First model of treatment order (C, LPS, LPS + IFNγ). Right column. Second model of treatment order (C, LPS + IFNγ, LPS). Correlation between gene expression and treatment effect is represented using a red-green gradient: dark-red, 1, being the most positive correlation and dark-green, −1, the most negative correlation. The correlation p-value for each module is shown in parentheses.
WGCNA reveals that LPS and LPS + IFNγ induce 11 different expression patterns in microglial cells
WGCNA is a method used in systems biology to find clusters or modules of highly interconnected genes and their correlation to a trait, e.g., treatment20. In our RNA-Seq study, we used WGCNA to analyze the gene expression patterns, in response to LPS and LPS + IFNγ, of all 14337 genes detected in cultured microglia. Our analysis clustered the genes into 11 distinct modules, corresponding to 11 different expression profiles, which were coded by arbitrarily assigned colours (Fig. 2B). The numbers of genes in the modules were, in descending order: 9343 (turquoise), 1169 (blue), 886 (brown), 532 (yellow), 516 (green), 441 (red), 432 (black), 334 (pink), 212 (magenta), 192 (purple) and 79 (green-yellow). Grey (201 genes) was reserved for outliers or unclustered genes that do not fit in any module. Branches in the hierarchical clustered dendrogram correspond to individual genes and the y-axis shows the distance between them, i.e. similarity, the more distant the less similar. Note that all genes in “blue”, “yellow” and “green” modules, as well as half of genes grouped in “black”, are clustered separately from the genes in the other modules. Furthermore, although some modules are well-defined, such as “yellow”, and most of “blue”, “brown” and “turquoise”, others are not.
The module eigengene can be used to summarize and represent the gene expression profile of a given module. Thus, barplots of the values of the module eigengene were used to represent the correlation between genes clustered within a module and the effect that LPS and LPS + IFNγ had on their expression (Fig. 2C). To account for the different patterns of gene expression, we fitted two models of treatment orders into the WGCNA. For the first one, we considered the pattern of genes that are modified by LPS, but even more by LPS + IFNγ (i.e., treatment order: C, LPS, LPS + IFNγ). This model showed the “yellow”, “blue”, “black” and “turquoise” modules to be the most strongly correlated (Fig. 2D, first column). Next, to obtain the gene patterns that were more strongly affected by LPS than by LPS + IFNγ, we fitted the reverse treatment order (C, LPS + IFNγ, LPS) and checked for the module correlation. In this case, the modules “blue” and “turquoise”, whose genes are upregulated or downregulated by both treatments, were again strongly correlated; but so were the “green” and “red” modules (Fig. 2D, second column). Therefore, genes grouped in the “yellow”, “blue”, “green”, “red”, “black” and “turquoise” modules were further analysed, to understand the biological significance of the gene expression patterns induced by LPS and LPS + IFNγ.
Ontological analysis of WGCNA differentially affected modules
The Genecodis platform19 was used to perform the ontological analysis of those genes clustered in the “yellow”, “blue”, “green”, “red”, “black” and “turquoise” modules. Specifically, we focused on Gene Ontology (GO) terms for Biological Process (BP) (Fig. 3) and on Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways (Fig. 4).
Enrichment analysis of Gene Ontology (GO) terms (biological processes) in selected WGCNA modules. This figure shows the 10 most significantly enriched GO terms in the yellow (A), blue (B), green (C), red (D), black (E) and turquoise (F) modules. Gene percentage annotation (i.e., the percentage of the total number of genes annotated in each category that were detected in this work) is depicted on the upper x-axis; whereas the p-value is represented on the lower x-axis. The biological processes and the number of genes annotated within each of them are presented on the y-axis.
Enrichment analysis of Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways in selected WGCNA modules. This figure shows the 10 most significantly enriched KEGG pathways23,24,25 in the yellow (A), blue (B), green (C), red (D), black (E) and turquoise (F) modules. Gene percentage annotation (i.e., the percentage of the total number of genes annotated in each category that were detected in this work) is depicted on the upper x-axis; whereas the p-value is represented on the lower x-axis. The biological processes and the number of genes annotated within each of them are presented on the y-axis.
Module “yellow” grouped together genes clearly upregulated only by LPS + IFNγ, such as the members of the GBP family: Gbp1, Gbp8 and Gbp10. These IFNγ-inducible GTPases are involved in defence responses to viruses and bacteria28. Interestingly, our analysis of this module resulted in an enrichment of “defence response to virus and protozoan” together with “cytokine-mediated signalling pathway” and “cellular response to IFNγ” GO terms (Fig. 3A). Furthermore, “cytokine-cytokine receptor interaction”, “chemokine signalling pathway” and “JAK-STAT signalling pathway” are among the most enriched KEGG pathways (Fig. 4A). These results suggest that genes in “yellow” module are affected by IFNγ, rather than LPS.
Both treatments, LPS and LPS + IFNγ, markedly upregulated genes clustered in the “blue” module. Our analysis revealed that these genes are mostly involved in inflammatory and immune response, since 8 out of the 10 most enriched GO terms were related to these biological processes (Fig. 3B). Actually, the “blue” module grouped together prototypical pro-inflammatory genes such as Tnf, Csf3, Il-6, Cybb, Ptgs2, Nos2, Il-1β, Il-12a and Il-12b. In this module, not only does IFNγ greatly affect gene expression, but so does LPS, since “cellular response to interferon-gamma”, “LPS-mediated signalling pathway” and “Toll-like receptor signalling pathway” were among the top-10 annotated GO terms and KEGG pathways (Figs 3B and 4B).
Genes found in the “green” module, i.e., genes upregulated by LPS but not by LPS + IFNγ, were also involved in inflammatory and immune responses (Fig. 3C). Since LPS alone greatly induces gene expression in this module, “response to lipopolysaccharide” and “Toll-like receptor signalling pathway” stand out among the most enriched GO terms and KEGG pathways (Figs 3C and 4C). In contrast, genes downregulated only in response to LPS, and clustered in the “red” module, play a role in non-inflammatory processes like “transcription”, “metabolism” and “phosphorylation” (Fig. 3D). Our analysis showed that there were no significant enriched KEGG pathways in the “red” module.
The “black” and “turquoise” modules include genes whose expression is decreased by LPS + IFNγ or by both treatments, respectively. “Transcription”, “translation”, “transport”, “metabolism” and “phosphorylation” were the main biological processes affected in these modules (Fig. 3E,F). Furthermore, the “black” module KEGG pathways annotation revealed that genes with decreased expression after LPS + IFNγ treatment play important roles in phagocytosis (Fig. 4D). In fact, previous work by our group has demonstrated reduced microglial capacity to phagocytose bacteria after LPS + IFNγ treatment11, which is in accordance with this result. Meanwhile, analysis of genes downregulated by both LPS and LPS + IFNγ, clustered in the “turquoise” module, revealed “Huntington’s, Alzheimer’s and Parkinson’s disease” to be among the most significant affected KEGG pathways (Fig. 4E). Note that genes identified in this study and classified within these three pathways represent more than 60% of all the genes annotated in these diseases, indicating the high relevance of the data presented here.
In microglia, LPS and LPS + IFNγ downregulate Huntington’s, Alzheimer’s and Parkinson’s disease-related genes
Among the genes downregulated by the combination of both treatments, and sorted into the “turquoise” module, 119, 101 and 84 genes were annotated in Huntington’s, Alzheimer’s and Parkinson’s disease, respectively (Fig. 4E). In-depth analysis revealed that in total these represented 169 different genes. Of these, 64 were involved in all three neurodegenerative disorders, as depicted in the Venn Diagram (Fig. 5). Within these 64 common genes, there were different members of the gene families of ATP synthases, cytochrome c oxidases, NADH dehydrogenases, succinate dehydrogenases and ubiquinol-cytochrome c reductases. The Venn diagram also shows 15, 35 and 48 genes related exclusively to Parkinson’s, Alzheimer’s and Huntington’s disease, respectively.
LPS and LPS + IFNγ downregulate 169 genes involved in Parkinson’s, Alzheimer’s and Huntington’s disease. The Venn diagram shows that as many as 64 of these genes are common to all three diseases. They belong to the gene families of ATP synthases, cytochrome c oxidases, NADH dehydrogenases, succinate dehydrogenases and ubiquinol-cytochrome c reductases. In turn, 15, 35 and 48 genes are specific to Parkinson’s, Alzheimer’s and Huntington’s disease respectively. Note the presence Park2, Park7 and Pink1 in Parkinson’s disease, Adam10, Apoe, App, Bace1 and Psen2 in Alzheimer’s disease and Bax, Hdac2 and Sp1 in Huntington’s disease.
Park2, Park7 and Pink1 stand out among Parkinson’s disease-specific genes downregulated by both treatments, since they are involved in biological processes characteristic of this disease, like dopamine uptake and neurotransmission, autophagy and degradation of mitochondria. Furthermore, our data reveal that Adam10, Apoe, App, Bace1 and Psen2 expression is decreased in microglial cells treated with LPS or LPS + IFNγ. These genes play a pivotal role in Nocht signalling, protein processing, apoptosis and beta-amyloid metabolic processes which are altered in Alzheimer’s disease. Finally, Bax, Hdac2 and Sp1, involved in apoptosis and transcription, are found among the Huntington’s disease-specific genes affected by LPS and LPS + IFNγ.
Differential expression of coding and non-coding RNAs induced by LPS and LPS + IFNγ
WGCNA revealed that although some genes respond equally to LPS and LPS + IFNγ, others do not. In order to further characterize microglial response to both treatments, we studied those RNAs that are differentially expressed using DESeq analysis. This is a straightforward approach to detect genes differentially upregulated or downregulated by direct pairwise comparisons.
Taking into account all the samples, 14,337 coding and non-coding RNA species were detected. When analysed separately, 11,225, 12,694 and 13,004 protein-coding RNAs were identified in Control, LPS and LPS + IFNγ samples, respectively (Table 2). Among non-coding RNAs, long non-coding RNAs (lncRNAs) were particularly abundant. Micro-RNAs (miRNAs) represented only 6% of the non-coding RNAs detected. Finally, pseudogenes, small nucleolar RNAs (snoRNAs) and small nuclear RNAs (snRNAs) were also found. Note that treatments can turn gene expression on or off; therefore, the number of RNAs detected is different for each Control, LPS and LPS + IFNγ condition.
Data analysis revealed that LPS greatly influences protein-coding RNA expression, since 37% of these RNAs detected in primary microglia show significant changes compared to Control (Table 3). Upregulation versus downregulation is split roughly equally: 52.4% and 47.6% respectively. In addition, LPS turns on the expression of 32 protein-coding RNAs and turns off 27. LPS also tends to induce overexpression of non-coding RNAs, particularly snoRNAs and miRNAs, since none of them were downregulated by LPS.
Regarding expression changes induced by LPS + IFNγ when compared to Control, our results do not differ, in general terms, from those reported above for LPS: the percentage of affected RNAs and the numbers of those increased or decreased are similar, though snoRNAs are an exception. Comparison between LPS and LPS + IFNγ shows that 10% of coding RNAs are altered by IFNγ addition. Its effect can be synergistic or antagonistic to LPS, since the number of RNAs increased or decreased is similar.
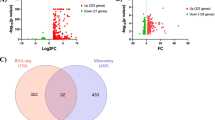
Venn diagrams (Fig. 6) also prove to be a useful tool to evaluate overall similarities and differences between the changes induced by LPS and by LPS + IFNγ in primary microglia. The response of protein-coding RNAs to LPS is stronger than that to LPS + IFNγ, since 4,684 and 4,397 genes are affected, respectively. However, most of them are common, specifically 3011. An additional 935 are affected by LPS alone, whereas only 644 are affected by LPS + IFNγ alone. In contrast, when comparing LPS + IFNγ directly with LPS (not each treatment with Control) our results show that out of 1,086 protein-coding RNAs altered. Of them, 80 are exclusively affected by LPS + IFNγ compared to LPS. Interestingly, there are 474 protein-coding RNAs whose expression changes in all three comparisons: LPS versus Control, LPS + IFNγ versus Control and LPS + IFNγ versus LPS.
Differentially expressed RNAs affected by either or both of LPS and LPS + IFNγ. Venn diagrams showing the number of protein-coding RNAs (A), pseudogenes (B), lncRNAs (C) and miRNAs (D) significantly affected by LPS or LPS + IFNγ in primary microglia as assessed by DESeq analysis. Note the large number of protein coding genes significantly affected by both LPS and LPS + IFNγ; but also the existence of many genes specifically affected by LPS or LPS + IFNγ.
The effect of LPS activation is also notable in pseudogenes and lncRNAs, where once again many LPS-altered RNAs also respond to LPS + IFNγ co-treatment. miRNAs are quite particular, since only 6 are differentially expressed: mir-155, mir-221, mir-222, mir-22hg, mir-5111 and mir-1957a. Five of these are induced by both LPS and LPS + IFNγ, but only mir-1957a expression increases due to LPS + IFNγ. As opposed to protein-coding RNAs, pseudogenes and lncRNAs, no miRNA changed when comparing the two treatments directly.
LPS- and LPS + IFNγ-treated microglia share common features, but have distinct phenotypes
Little is known about the biological functions of non-coding RNAs in microglia, except for a few miRNAs, such as mir-155, which was upregulated by both treatments in our analysis. Therefore, we focused on protein-coding RNAs to determine the similarities and differences between LPS- and LPS + IFNγ-activated microglia.
SPIA analysis of the DEGs26 provides the first clues to this question. We analysed separately all 4,684, 4,397 and 1,086 DEGs affected in the LPS vs Control, LPS + IFNγ vs Control and LPS + IFNγ vs LPS comparisons, respectively. The Toll-like receptor signalling pathway was significantly affected in all of them. Interestingly, many genes involved in this pathway were upregulated by both LPS and LPS + IFNγ treatments (Fig. 7A). However, others including Tlr2, Tlr7 and Tlr8 were downregulated by LPS + IFNγ. A dampening effect of IFNγ was also observed in the case of inflammatory genes such as Tnf, Il-1β, IKKβ and Irf7, when comparing LPS + IFNγ to LPS (Fig. 7B).
Toll-like receptor signalling pathway shows similarities, but also differences between LPS and LPS + IFNγ. (A) Genes involved in Toll-like receptor signalling affected by LPS (left side of the square) or LPS + IFNγ (right side of the square) compared to control. (B) This graph shows the genes within the pathway that are significantly altered when comparing LPS + IFNγ to LPS. Red indicates upregulation; green downregulation. Data on KEGG graph23,24,25 were rendered using Pathview.
More significant results were obtained when considering common and non-common DEGs separately. Nos2, Tnf, Tlr2, Tlr3, Il-1β, Il-18, Il-23a, Irf1, Irf7 and Irf8 were among the 474 coding RNAs differentially expressed in all three comparisons. These RNAs did not share the same expression profile, e.g., Nos2 expression increases due to both treatments but particularly with LPS + IFNγ; while LPS alone induced the highest Tnf expression. These findings are in accordance with the qRT-PCR results (shown in Fig. 1A). This expression heterogeneity is also confirmed by WGCNA: 30%, 26%, 20%, 16%, 5% and 3% of the DEGs detected by DESeq are clustered within the “yellow”, “green”, “blue”, “turquoise”, “pink” and “red” modules, respectively. Even so, most of them are involved in inflammatory and immune responses, since 7 of the 10 most significant annotated GO terms were related to these processes (Fig. 8A). Thus, both treatments, LPS and LPS + IFNγ, trigger an inflammatory microglial phenotype.
LPS and LPS + IFNγ are involved in similar biological functions, but some differences exist. This figure shows the 10 most enriched GO terms in: (A) genes common to LPS vs Control, LPS + IFNγ vs Control and LPS + IFNγ vs LPS (n = 474); (B) Genes only downregulated in LPS vs Control (n = 502). (C) Genes only upregulated in LPS vs Control (n = 433). (D) Genes only downregulated in LPS + IFNγ vs Control (n = 377). (E) Genes only upregulated in LPS + IFNγ vs Control (n = 267). Note that 7 of the 10 GO terms in (A) are related to immune and inflammatory responses. ((B–E) Share metabolism and transcription as the biological processes affected; but LPS tends to alter cell cycle (B) and LPS + IFNγ chromatin modifications (E). Gene percentage annotation (i.e., the percentage of the total number of genes annotated in each category detected in this work) is depicted on the upper x-axis, whereas the p-value is represented on the lower x-axis. The GO terms and the number of genes annotated within each of them are presented on the y-axis.
Regarding non-common DEGs, 935 RNAs were affected only by LPS alone and 644 by LPS + IFNγ compared to Control. Both treatments seem to be more prone to inhibit expression of these RNAs than to promote it. Thus, 502 RNAs were downregulated and 433 were upregulated due to LPS, whereas expression of 377 RNAs was decreased and that of 267 was increased in response to LPS + IFNγ (Fig. 6A). Once again, these expression patterns were in accordance with WGCNA results. 97% and 3% of genes downregulated only by LPS were found within the “turquoise” and “red” modules, whilst most of the upregulated genes were clustered in the “green” and “blue” modules. Meanwhile, the “turquoise” and “black” modules included all the genes whose expression decreased only as a result of LPS + IFNγ treatment; whereas the genes upregulated only by this treatment were clustered in the “blue”, “yellow” and “pink” modules. According to the GO analysis, most of the DEGs only affected by LPS or only by LPS + IFNγ were involved in transcription and regulation of transcription processes (Fig. 8B-E). Furthermore, since inflammatory and immune responses are also associated with shifts in metabolism29, metabolic processes were also greatly altered by LPS and LPS + IFNγ.
Although these results suggest that LPS and LPS + IFNγ-treated primary microglial cultures are similar, note that the genes involved in the biological processes are different for each condition. On one hand, we found that LPS alone significantly downregulates the transcription factors Runx1, Klf10 and Klf11, but upregulates the small GTPase RhoA, oncogenes Myc and Bmi1, chromatin remodelling factor Mta2 and membrane receptor Trem1 (Fig. 9A). On the other hand, LPS + IFNγ significantly decreases expression of the small GTPase RhoB, membrane receptor P2x4r, transcription factor Mef2d and non-histone nuclear DNA-binding protein Hmgb1. Additionally, LPS + IFNγ also increases the expression of the moonlighting protein Gapdh, sirtuin Sirt1, histone deacetylase Hdac9 and epigenetic regulator Mecp2 (Fig. 9B).
A model of specific signalling pathways triggered by LPS and LPS + IFNγ in primary microglia. (A). By upregulating genes like Mta2, Bmi1, Myc, RhoA and Trem1 and downregulating Runx1, Klf10 and Klf11, LPS promotes microglial proliferation, phagocytosis and the release of pro-inflammatory cytokines. (B). LPS + IFNγ induces expression of Gapdh, Sirt1, Hdac9 and Mecp2, whereas it reduces that of Mef2d, P2xr4, RhoB and Hmgb1. This suggests that LPS + IFNγ-treated microglia have less proliferative potential, show reduced expression of genes involved in neuropathic pain and are more prone to resolve inflammation by activation of oxidative respiration and blockade of pro-inflammatory cytokine production.
Finally, only 80 RNAs were differentially expressed when directly comparing LPS + IFNγ to LPS. Of these, 39 were downregulated and clustered in the “black” (82%) and “turquoise” (18%) modules. The remaining 41 were upregulated and found within the “red” (94%) and “pink” (6%) modules (Fig. 6A). Because of the small number of affected genes, little can be inferred from analysis of the GO terms associated with these genes (data not shown).
Discussion
LPS alone or together with IFNγ is widely used to trigger microglial activation. However, to the best of our knowledge, similarities and differences between the effects of these two treatments on microglial gene expression have not been fully characterized. In the present study, to address this, we analysed the transcriptomic profile of primary microglial cells in response to LPS and LPS + IFNγ by RNA-Seq.
Determination of the time point used in this study was a significant initial step. Evidence shows that not all genes are expressed at once, but each has its own specific temporal expression pattern. Using LPS-treated primary microglia and BV2 cells, Das and colleagues12,30 reported that the most significant expression changes occur within the first and the eighth hour after treatment. We also previously described a marked gene upregulation in cultured microglia treated with LPS and LPS + IFNγ for 6 h11,31,32. In this study, our qRT-PCR results showed significant alterations of canonical pro-inflammatory genes, such as Tnf, Csf3, Il-6, Cybb, Ptgs2 and Nos2, 6 hours after treatment. The expression profile of these genes was also observed after analysing RNA-Seq data, thus validating the high-throughput sequencing.
We are aware that analysing a single time point is a limitation of this study. Nevertheless, the data presented here for 6 hours after treatment offer a good picture of the first wave of transcriptional changes induced by LPS or LPS + IFNγ which are mainly regulated by latent transcription factors33. The relative timing of IFNγ and LPS exposure was also important. We decided to use co-treatment because this procedure results in a strong potentiation of LPS-induced NO production and neurotoxic effects10,34. Priming with IFNγ could have resulted in a somewhat different pattern of modulation, including a stronger potentiation on the production of some pro-inflammatory cytokines like TNFα35.
Genes or proteins that are directly linked might have a similar function and therefore may be part of the same biological pathway. More and more, network methods such as WGCNA are chosen to analyse these interactions20. This approach classifies the whole transcriptome into discernible expression patterns. Furthermore, it compares all genes in the data and takes into account all changes through all samples at the same time. Therefore, it can extract complex expression profiles that could be easily missed if only DESeq pairwise differential expression analysis was used to examine the biological relevance of RNA-Seq data. Therefore, we used WGCNA to organize gene expression data in order to establish clusters or modules of genes with similar responses to LPS and LPS + IFNγ treatments. Our results show that these two stimuli, both considered pro-inflammatory, do not affect all transcripts in the same way, since 11 different gene expression patterns were detected. The results also show that while some genes are affected by both treatments, others are only affected by LPS or LPS + IFNγ, which means that LPS and LPS + IFNγ do not always affect a given gene in the same way. We consider these observations to be relevant findings. Frequently, reports only focus on gene upregulation due to pro-inflammatory stimuli12,36. However, according to our data, 9,343 genes out of 14,337 tend to be downregulated, without this trend emerging as statistically significantresult, by both treatments. Furthermore, in other studies, microglia are only treated with LPS + IFNγ, as strong pro-inflammatory stimuli, without considering a potential antagonistic effect between LPS and IFNγ37,38.
According to GO analysis, the genes that are upregulated by pro-inflammatory stimuli in cultured microglia show a significant enrichment of GO terms related to inflammatory and immune responses. The annotation of these functions in response to LPS and/or LPS + IFNγ is interesting and indicates that our approach of comparing both treatments is strong but rather predictable. To the best of our knowledge, a comprehensive transcriptional profile of LPS + IFNγ-treated microglia has not been derived using RNA-seq, though the mouse embryonic stem cell-derived microglia response to this stimulus has been analysed via RNA microarrays39. Those data showed an exacerbated immune response and increased pro-inflammatory gene expression. Other reports also identify alterations in inflammatory and immune system processes, as well as a marked upregulation of pro-inflammatory genes, such as Ptgs2, Tnf, Il-12a and Il-1β, in the BV2 microglial cell line, primary microglial cultures and acutely isolated microglia from adult CNS treated with LPS (Table 4). Altogether, this is in agreement with our results.
Downregulated genes due to pro-inflammatory treatments are often overlooked because they are not thought to be involved in inflammation12,36. Here, we show that most genes expressed in non-activated primary microglia are repressed when these cells are activated by LPS or LPS + IFNγ. Transcription, translation, metabolism, transport, apoptosis and phosphorylation were the main biological processes in which the genes that are downregulated by one or both treatments were involved. These results relativize the concept of resting microglia in physiological conditions.
Interestingly, 169 genes downregulated by both treatments were involved in three neurodegenerative pathologies: Huntington’s, Alzheimer’s and Parkinson’s disease. Common features of these disorders include abnormal protein aggregation, oxidative stress and mitochondrial dysfunction40. Indeed, 64 downregulated genes play a role in all three diseases; most of them belong to gene families involved in ATP synthesis through the mitochondrial electron transport chain. According to previous reports, inhibition of the mitochondrial respiratory chain enhances mitochondrial ROS production41. Therefore, downregulation of genes involved in the electron transport chain due to LPS and LPS + IFNγ treatments in microglia could lead to cellular damage and death of neighbouring cells due to ROS overproduction. Thus, activating primary microglia with both stimuli could be an experimental model to study this process.
CNS inflammation plays a role in the progression of chronic neurodegenerative diseases and this inflammatory response is mostly governed by microglia42. In agreement with our results, LPS downregulates the E3 ubiquitin ligase Park2 in microglial cells43, which leads to the accumulation of Tnf, Il-1β and Nos2. Park7-deficient microglia also show higher levels of Tnf, Nos2 and Ptgs244. Finally, Pink1 deletion exacerbates microglial activation45. Thus, downregulation of these genes could promote a pro-inflammatory microglial phenotype, thereby contributing to neurodegeneration in Parkinson’s disease.
Activated microglia induce neuronal cell death through the release of cytotoxic factors including cytokines and ROS in Alzheimer’s disease46. Interestingly, the “cytokine activity”, “defence response” and “T-cell activation” GO terms were associated with upregulated genes in an RNA-Seq analysis performed in sorted microglia from 15 to 18 month-old APP/PS1 transgenic mice47. Furthermore, that report showed downregulation of ApoE expression, similar to our data presented here. It was possible to revert this process, which is highly impaired in Alzheimer’s disease, by co-culturing neonatal microglia in brain slices of 20 month-old APP/PS1 mice. Therefore, a comparison between young and old microglia transcriptomic profiles could provide some clues as to which processes are impaired with age and lead to disease. Little is known about Adam10, App, Bace1 and Psen2 microglial expression. Nevertheless, due to their involvement in Alzheimer’s disease pathophysiology, it would be worth further studying the implications of their downregulation in response to LPS and LPS + IFNγ. Similarly, decreased expression of the Huntington’s disease-related genes Sp1, Hdac2 and Bax, as a result of both treatments could provide some clues about the underlying disease mechanisms.
This study was performed with primary microglial cultures isolated from neonatal mouse cortex. Although this model is widely used and has contributed considerably to our current knowledge on microglial biology, it is important to be aware of its limitations. As with most cell types, in vitro culture has an impact on microglial biology. Gosselin and colleagues48 reported significant transcriptomic changes in human and mouse brain microglia when cultured. The transfer of cells to the in vitro environment resulted in 10-fold induction of more than 300 genes and 10-fold reduction of more than 700 genes after 7 days. Although cultured microglial cells show an activated phenotype, they become less responsive after 13–16 days in vitro (DIV)49,50. The microglial cultures in this study were used at 21 DIV. This relatively long time in culture allows microglia to mature and become less reactive, particularly because of their interactions with astrocytes. Be that as it may, it is important to acknowledge that the lack of neuronal inhibitory inputs probably results in microglia being in a more activated state than in the adult CNS in vivo. Another caveat to be added to this study is that microglial cells were isolated from the brains of early neonates. This allows for the preparation of microglial cultures with a high yield in a simple and reproducible manner. However, since early postnatal microglial cells differ from adult or aged microglia in terms of morphology, function and gene expression51, caution must be exercised when interpreting our data. Recent improvements in the isolation of fresh microglia using fluorescence-activated cell sorting and laser capture microdissection methods have been reported52. Such methods have the advantage over cultures that microglial cells live in their physiological environment until the moment of extraction, but they are not free from artefacts. The isolation procedure for fluorescence-activated cell sorting requires 3 to 5 hours, which may affect the expression profile of microglia. In contrast, cellular lysis in cultured microglia is immediate and the transcriptome obtained faithfully reflects the transcriptome of the living cells, which also occurs with RNA from microglia isolated by laser capture microdissection. A main drawback of RNA from microglia isolated by laser capture microdissection is the probable presence of RNA from surrounding cells. In this respect, the purity of cultured microglia RNAs is far superior. As has been stated before, it is critical to be aware of the limitations of the protocol used and choose the method most appropriate for the scientific question addressed53. Despite their limitations, LPS- or LPS + IFNγ-treated microglial cultures allow a considerable experimental versatility and provide valuable information on the potential phenotypes of activated microglia.
Apart from revealing the inhibition of important genes implicated in neurodegenerative diseases in cultured microglial cells treated with LPS or LPS + IFNγ, the present RNA-Seq analysis also allowed us to gain further understanding of how both stimuli affect microglial gene expression. Thus, DEGs were obtained using DESeq analysis. Both protein-coding and non-protein-coding RNAs were found; although protein-coding transcripts represented 90% of all RNAs detected. Among the non-protein-coding RNAs, few miRNAs were detected because the sequencing protocol used here only allows for the detection of pre-miRNAs. Thus, the data presented here show only a small fraction of the miRNAs present in primary microglial cells. To conduct a specific study of miRNAs, another sequencing library preparation protocol should be used.
LPS and LPS + IFNγ have a slightly greater tendency to upregulate gene expression than to downregulate it in all the transcripts differentially expressed. This slight difference between the number of genes upregulated and those downregulated, though in accordance with Hirbec et al.54, differs from most previous microglial RNA-Seq analysis, where most of the affected genes were upregulated (Table 4). These differences may be due to different time points, treatment doses or microglia sources being used. The number of samples can also lead to discrepancies30.
Because information on non-protein-coding RNAs is scant, the functional significance of their expression changes due to treatments in microglial biology was not further considered in this work. Instead, we focused on protein-coding RNAs. LPS versus Control and LPS + IFNγ versus Control comparisons shared more than 3000 DEGs. Moreover, another 474 DEGs were common to these two comparisons and also to the LPS + IFNγ versus LPS comparison. Most of these genes, including Nos2, Tnf, Tlr2, Tlr3, Il-1β, Il-18, Il-23a, Irf1, Irf7 and Irf8, are involved in inflammatory and immune responses, thus validating the usefulness of LPS and LPS + IFNγ to induce and study microglial activation in vitro. For this reason, they are sometimes used interchangeably. A detailed look into the genes differentially affected only by LPS and LPS + IFNγ, together with a comprehensive review of the literature, has allowed us to better understand the different signalling pathways affected under each condition.
The transcription factor Runx1 is sufficient to inhibit microglia activation in response to LPS treatment. Runx1 acts in amoeboid microglia to limit the length of the activated phase and guide the cells toward a deactivated phenotype. Moreover, this transcription factor inhibits the proliferative capacity of amoeboid microglia55. Through its target Bmi1, Runx1 is involved in cell self-renewal and its depletion increases Bmi1, thereby inducing cell proliferation56. It seems probable that, LPS downregulates Runx1 expression by increasing Mta2: a member of NuRD complex57. Therefore, Bmi1 expression is upregulated, which suggests the existence of a proliferative microglial phenotype in response to this treatment (Fig. 9A). The activation of the oncogene Myc also supports this hypothesis. Klf10 and Klf11 decrease cell proliferation and induce apoptosis by targeting Myc58. LPS treatment not only downregulates Klf10 and Klf11 genes in microglia, but also in bone marrow-derived macrophages. This inhibition enhances Il-12b expression59. Inflammation is also elicited by the upregulation of the small GTPase RhoA, since it promotes superoxide release60. RhoA also induces microglial phagocytosis of apoptotic neurons61, whereas Trem1 favours microglial Aβ uptake62. Therefore, LPS triggers a proliferative, pro-inflammatory and phagocytic microglial phenotype.
The NAD-dependent deacetylase Sirt1 is a crucial regulator of energy metabolism and cell survival. It supports oxidative respiration and anti-inflammatory responses, thus enhancing the resolution of inflammation63. Gapdh promotes Sirt1 activation and autophagy initiation which protect cells from apoptosis induced by glucose withdrawal64. Thus, through Gapdh, LPS + IFNγ could enhance Sirt1 expression (Fig. 9B). LPS + IFNγ-activated microglia seem to be non-proliferative and non-motile, since Sirt1 blocks microglial proliferation65 and the small GTPase RhoB, which induces cell migration, is downregulated by the co-treatment66. After spinal cord injury, RhoB is co-expressed with the purinergic receptor P2x4r in microglial cells. This receptor is a major contributor to the development of neuropathic pain. Therefore, P2x4r downregulation due to LPS + IFNγ could be related to pain reduction. This stimulus is also involved in chromatin structure. Sirt1 and Hdac9 are histone deacetylases and Mecp2 participates in histone methylation. All three upregulated genes participate in inflammatory responses. Sirt1 suppresses the release of pro-inflammatory cytokines65. Meanwhile, Hdac9 binds and represses the transcription factor Mef267, which in later stages of microglial activation promotes IL-10 release68. Finally, Hmgb1 interacts with and represses Mecp269. However, LPS + IFNγ downregulates Hmgb1, thus increasing Mecp2 expression and blocking secretion of pro-inflammatory mediators70,71. Thus, these results suggest that LPS + IFNγ treatment reduces genes involved in neuropathic pain, as well as microglial migration and proliferation, whereas it also promotes some anti-inflammatory responses.
Altogether, it seems that the addition of IFNγ could, somehow, dampen some pro-inflammatory genes. Downregulation of inflammatory mediators and effectors, such as that of Tnf, Il-1β, IKKβ and Irf7 observed in Toll-like receptor signalling pathway analysis when comparing LPS + IFNγ and LPS, supports this hypothesis. To confirm it, an additional experimental group, IFNγ-treated microglia, should have been included. This is certainly a limitation of the study.
Conclusions
In summary, by using RNA-seq, we have compared the transcriptomic changes induced by LPS and LPS + IFNγ in primary murine microglia. Our study demonstrates that both treatments greatly affect microglial gene expression. However, not all genes are equally affected by LPS and LPS + IFNγ. Interestingly, both treatments downregulate genes involved in Parkinson’s, Alzheimer’s and Huntington’s disease. These data break new ground in the study of the role of microglia in these disorders. Furthermore, a detailed look at the genes differentially expressed only by LPS or LPS + IFNγ has revealed that LPS-treated microglia are proliferative, phagocytic and release pro-inflammatory cytokines. In contrast, LPS + IFNγ reduces cell proliferation, downregulates genes involved in pain and favours resolution of inflammation. These different profiles must be taken into account when using LPS or LPS + IFNγ in in vitro models of microglial activation.
Ethics approval
All procedures involving animals were approved by the Ethical Committee for Animal Experimentation of the Universitat de Barcelona and by the Commission for Animal Experimentation of the Generalitat de Catalunya, with protocols numbers DAAM 5026, 7064 and 7065.
Availability of Data and Materials
RNAseq datasets were deposited in the Gene Expression Omnibus database under dataset accession number GSE90046. Note that this dataset also includes samples from our previous work11. However, no overlap analysis exists between both reports. Samples here analyzed can be found as C/EBPβfl/fl genotype, considered wild-type. Other data supporting the conclusions of this study will be available upon reasonable request.
References
Soulet, D. & Rivest, S. Microglia. Current biology: CB 18, R506–508, https://doi.org/10.1016/j.cub.2008.04.047 (2008).
Gomez-Nicola, D. & Perry, V. H. Microglial dynamics and role in the healthy and diseased brain: a paradigm of functional plasticity. The Neuroscientist: a review journal bringing neurobiology, neurology and psychiatry 21, 169–184, https://doi.org/10.1177/1073858414530512 (2015).
Lu, Y. C., Yeh, W. C. & Ohashi, P. S. LPS/TLR4 signal transduction pathway. Cytokine 42, 145–151, https://doi.org/10.1016/j.cyto.2008.01.006 (2008).
Block, M. L., Zecca, L. & Hong, J. S. Microglia-mediated neurotoxicity: uncovering the molecular mechanisms. Nature reviews. Neuroscience 8, 57–69, https://doi.org/10.1038/nrn2038 (2007).
Schwartz, M., Kipnis, J., Rivest, S. & Prat, A. How do immune cells support and shape the brain in health, disease, and aging? The Journal of neuroscience: the official journal of the Society for Neuroscience 33, 17587–17596, https://doi.org/10.1523/JNEUROSCI.3241-13.2013 (2013).
Ransohoff, R. M. & Perry, V. H. Microglial physiology: unique stimuli, specialized responses. Annual review of immunology 27, 119–145, https://doi.org/10.1146/annurev.immunol.021908.132528 (2009).
Nayak, D., Roth, T. L. & McGavern, D. B. Microglia development and function. Annual review of immunology 32, 367–402, https://doi.org/10.1146/annurev-immunol-032713-120240 (2014).
Dutta, G., Zhang, P. & Liu, B. The lipopolysaccharide Parkinson’s disease animal model: mechanistic studies and drug discovery. Fundamental & clinical pharmacology 22, 453–464, https://doi.org/10.1111/j.1472-8206.2008.00616.x (2008).
Hu, X. & Ivashkiv, L. B. Cross-regulation of signaling pathways by interferon-gamma: implications for immune responses and autoimmune diseases. Immunity 31, 539–550, https://doi.org/10.1016/j.immuni.2009.09.002 (2009).
Gresa-Arribas, N. et al. Modelling neuroinflammation in vitro: a tool to test the potential neuroprotective effect of anti-inflammatory agents. PloS one 7, e45227, https://doi.org/10.1371/journal.pone.0045227 (2012).
Pulido-Salgado, M. et al. Myeloid C/EBPbeta deficiency reshapes microglial gene expression and is protective in experimental autoimmune encephalomyelitis. Journal of neuroinflammation 14, 54, https://doi.org/10.1186/s12974-017-0834-5 (2017).
Das, A. et al. Dual RNA sequencing reveals the expression of unique transcriptomic signatures in lipopolysaccharide-induced BV-2 microglial cells. PloS one 10, e0121117, https://doi.org/10.1371/journal.pone.0121117 (2015).
Saura, J., Tusell, J. M. & Serratosa, J. High-yield isolation of murine microglia by mild trypsinization. Glia 44, 183–189, https://doi.org/10.1002/glia.10274 (2003).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079, https://doi.org/10.1093/bioinformatics/btp352 (2009).
Liao, Y., Smyth, G. K. & Shi, W. The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic acids research 41, e108, https://doi.org/10.1093/nar/gkt214 (2013).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930, https://doi.org/10.1093/bioinformatics/btt656 (2014).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome biology 15, R29, https://doi.org/10.1186/gb-2014-15-2-r29 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic acids research 43, e47, https://doi.org/10.1093/nar/gkv007 (2015).
Carmona-Saez, P., Chagoyen, M., Tirado, F., Carazo, J. M. & Pascual-Montano, A. GENECODIS: a web-based tool for finding significant concurrent annotations in gene lists. Genome biology 8, R3, https://doi.org/10.1186/gb-2007-8-1-r3 (2007).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC bioinformatics 9, 559, https://doi.org/10.1186/1471-2105-9-559 (2008).
Delhomme, N., Padioleau, I., Furlong, E. E. & Steinmetz, L. M. easyRNASeq: a bioconductor package for processing RNA-Seq data. Bioinformatics 28, 2532–2533, https://doi.org/10.1093/bioinformatics/bts477 (2012).
Anders, S. et al. Count-based differential expression analysis of RNA sequencing data using R and Bioconductor. Nature protocols 8, 1765–1786, https://doi.org/10.1038/nprot.2013.099 (2013).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic acids research 45, D353–D361, https://doi.org/10.1093/nar/gkw1092 (2017).
Kanehisa, M., Sato, Y., Kawashima, M., Furumichi, M. & Tanabe, M. KEGG as a reference resource for gene and protein annotation. Nucleic acids research 44, D457–462, https://doi.org/10.1093/nar/gkv1070 (2016).
Kanehisa, M. & Goto, S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic acids research 28, 27–30 (2000).
Luo, W. et al. Pathview Web: user friendly pathway visualization and data integration. Nucleic acids research, https://doi.org/10.1093/nar/gkx372 (2017).
Guo, Y., Zhao, S., Li, C. I., Sheng, Q. & Shyr, Y. RNAseqPS: A Web Tool for Estimating Sample Size and Power for RNAseq Experiment. Cancer informatics 13, 1–5, https://doi.org/10.4137/CIN.S17688 (2014).
Vestal, D. J. & Jeyaratnam, J. A. The guanylate-binding proteins: emerging insights into the biochemical properties and functions of this family of large interferon-induced guanosine triphosphatase. Journal of interferon & cytokine research: the official journal of the International Society for Interferon and Cytokine Research 31, 89–97, https://doi.org/10.1089/jir.2010.0102 (2011).
Kominsky, D. J., Campbell, E. L. & Colgan, S. P. Metabolic shifts in immunity and inflammation. J Immunol 184, 4062–4068, https://doi.org/10.4049/jimmunol.0903002 (2010).
Das, A. et al. Transcriptome sequencing reveals that LPS-triggered transcriptional responses in established microglia BV2 cell lines are poorly representative of primary microglia. Journal of neuroinflammation 13, 182, https://doi.org/10.1186/s12974-016-0644-1 (2016).
Valente, T. et al. CCAAT/enhancer binding protein delta regulates glial proinflammatory gene expression. Neurobiology of aging 34, 2110–2124, https://doi.org/10.1016/j.neurobiolaging.2013.02.007 (2013).
Straccia, M. et al. Pro-inflammatory gene expression and neurotoxic effects of activated microglia are attenuated by absence of CCAAT/enhancer binding protein beta. Journal of neuroinflammation 8, 156, https://doi.org/10.1186/1742-2094-8-156 (2011).
Litvak, V. et al. Function of C/EBPdelta in a regulatory circuit that discriminates between transient and persistent TLR4-induced signals. Nature immunology 10, 437–443, https://doi.org/10.1038/ni.1721 (2009).
Papageorgiou, I. E. et al. TLR4-activated microglia require IFN-gamma to induce severe neuronal dysfunction and death in situ. Proceedings of the National Academy of Sciences of the United States of America 113, 212–217, https://doi.org/10.1073/pnas.1513853113 (2016).
Hausler, K. G. et al. Interferon-gamma differentially modulates the release of cytokines and chemokines in lipopolysaccharide- and pneumococcal cell wall-stimulated mouse microglia and macrophages. The European journal of neuroscience 16, 2113–2122 (2002).
Das, A. et al. RNA sequencing reveals resistance of TLR4 ligand-activated microglial cells to inflammation mediated by the selective jumonji H3K27 demethylase inhibitor. Scientific reports 7, 6554, https://doi.org/10.1038/s41598-017-06914-5 (2017).
Gimeno-Bayon, J., Lopez-Lopez, A., Rodriguez, M. J. & Mahy, N. Glucose pathways adaptation supports acquisition of activated microglia phenotype. Journal of neuroscience research 92, 723–731, https://doi.org/10.1002/jnr.23356 (2014).
Zhang, S. et al. Resveratrol Attenuates Microglial Activation via SIRT1-SOCS1 Pathway. Evidence-based complementary and alternative medicine: eCAM 2017, 8791832, https://doi.org/10.1155/2017/8791832 (2017).
Beins, E. et al. Characterization of inflammatory markers and transcriptome profiles of differentially activated embryonic stem cell-derived microglia. Glia 64, 1007–1020, https://doi.org/10.1002/glia.22979 (2016).
Carvalho, C. et al. The role of mitochondrial disturbances in Alzheimer, Parkinson and Huntington diseases. Expert review of neurotherapeutics 15, 867–884, https://doi.org/10.1586/14737175.2015.1058160 (2015).
Ye, J. et al. Electron transport chain inhibitors induce microglia activation through enhancing mitochondrial reactive oxygen species production. Experimental cell research 340, 315–326, https://doi.org/10.1016/j.yexcr.2015.10.026 (2016).
Cunningham, C. Microglia and neurodegeneration: the role of systemic inflammation. Glia 61, 71–90, https://doi.org/10.1002/glia.22350 (2013).
Tran, T. A. et al. Lipopolysaccharide and tumor necrosis factor regulate Parkin expression via nuclear factor-kappa B. PloS one 6, e23660, https://doi.org/10.1371/journal.pone.0023660 (2011).
Kim, J. H. et al. DJ-1 facilitates the interaction between STAT1 and its phosphatase, SHP-1, in brain microglia and astrocytes: A novel anti-inflammatory function of DJ-1. Neurobiology of disease 60, 1–10, https://doi.org/10.1016/j.nbd.2013.08.007 (2013).
Flinn, L. J. et al. TigarB causes mitochondrial dysfunction and neuronal loss in PINK1 deficiency. Annals of neurology 74, 837–847, https://doi.org/10.1002/ana.23999 (2013).
Cai, Z., Hussain, M. D. & Yan, L. J. Microglia, neuroinflammation, and beta-amyloid protein in Alzheimer’s disease. The International journal of neuroscience 124, 307–321, https://doi.org/10.3109/00207454.2013.833510 (2014).
Orre, M. et al. Isolation of glia from Alzheimer’s mice reveals inflammation and dysfunction. Neurobiology of aging 35, 2746–2760, https://doi.org/10.1016/j.neurobiolaging.2014.06.004 (2014).
Gosselin, D. et al. An environment-dependent transcriptional network specifies human microglia identity. Science 356, https://doi.org/10.1126/science.aal3222 (2017).
Cristovao, A. C. et al. Microglia of rat ventral midbrain recovers its resting state over time in vitro: let microglia rest before work. Journal of neuroscience research 88, 552–562, https://doi.org/10.1002/jnr.22219 (2010).
Caldeira, C. et al. Microglia change from a reactive to an age-like phenotype with the time in culture. Frontiers in cellular neuroscience 8, 152, https://doi.org/10.3389/fncel.2014.00152 (2014).
Harry, G. J. Microglia during development and aging. Pharmacology & therapeutics 139, 313–326, https://doi.org/10.1016/j.pharmthera.2013.04.013 (2013).
Hirbec, H., Noristani, H. N. & Perrin, F. E. Microglia Responses in Acute and Chronic Neurological Diseases: What Microglia-Specific Transcriptomic Studies Taught (and did Not Teach) Us. Frontiers in aging neuroscience 9, 227, https://doi.org/10.3389/fnagi.2017.00227 (2017).
Carson, M. J., Crane, J. & Xie, A. X. Modeling CNS microglia: the quest to identify predictive models. Drug discovery today. Disease models 5, 19–25, https://doi.org/10.1016/j.ddmod.2008.07.006 (2008).
Hirbec, H. et al. The microglial reaction signature revealed by RNAseq from individual mice. Glia 66, 971–986, https://doi.org/10.1002/glia.23295 (2018).
Zusso, M. et al. Regulation of postnatal forebrain amoeboid microglial cell proliferation and development by the transcription factor Runx1. The Journal of neuroscience: the official journal of the Society for Neuroscience 32, 11285–11298, https://doi.org/10.1523/JNEUROSCI.6182-11.2012 (2012).
Motoda, L. et al. Runx1 protects hematopoietic stem/progenitor cells from oncogenic insult. Stem Cells 25, 2976–2986, https://doi.org/10.1634/stemcells.2007-0061 (2007).
Vishwakarma, B. A. et al. Runx1 repression by histone deacetylation is critical for Setbp1-induced mouse myeloid leukemia development. Leukemia 30, 200–208, https://doi.org/10.1038/leu.2015.200 (2016).
Ellenrieder, V. TGFbeta regulated gene expression by Smads and Sp1/KLF-like transcription factors in cancer. Anticancer research 28, 1531–1539 (2008).
Zhang, W. et al. Klf10 inhibits IL-12p40 production in macrophage colony-stimulating factor-induced mouse bone marrow-derived macrophages. European journal of immunology 43, 258–269, https://doi.org/10.1002/eji.201242697 (2013).
Moon, M. Y. et al. Involvement of small GTPase RhoA in the regulation of superoxide production in BV2 cells in response to fibrillar Abeta peptides. Cellular signalling 25, 1861–1869, https://doi.org/10.1016/j.cellsig.2013.05.023 (2013).
Jiang, T. et al. TREM1 facilitates microglial phagocytosis of amyloid beta. Acta neuropathologica 132, 667–683, https://doi.org/10.1007/s00401-016-1622-5 (2016).
Scheiblich, H. & Bicker, G. Regulation of Microglial Phagocytosis by RhoA/ROCK-Inhibiting Drugs. Cellular and molecular neurobiology 37, 461–473, https://doi.org/10.1007/s10571-016-0379-7 (2017).
Kauppinen, A., Suuronen, T., Ojala, J., Kaarniranta, K. & Salminen, A. Antagonistic crosstalk between NF-kappaB and SIRT1 in the regulation of inflammation and metabolic disorders. Cellular signalling 25, 1939–1948, https://doi.org/10.1016/j.cellsig.2013.06.007 (2013).
Chang, C. et al. AMPK-Dependent Phosphorylation of GAPDH Triggers Sirt1 Activation and Is Necessary for Autophagy upon Glucose Starvation. Molecular cell 60, 930–940, https://doi.org/10.1016/j.molcel.2015.10.037 (2015).
Li, L. et al. Overexpression of SIRT1 Induced by Resveratrol and Inhibitor of miR-204 Suppresses Activation and Proliferation of Microglia. Journal of molecular neuroscience: MN 56, 858–867, https://doi.org/10.1007/s12031-015-0526-5 (2015).
Schwab, J. M., Guo, L. & Schluesener, H. J. Spinal cord injury induces early and persistent lesional P2X4 receptor expression. Journal of neuroimmunology 163, 185–189, https://doi.org/10.1016/j.jneuroim.2005.02.016 (2005).
Smith, J. D. New role for histone deacetylase 9 in atherosclerosis and inflammation. Arteriosclerosis, thrombosis, and vascular biology 34, 1798–1799, https://doi.org/10.1161/ATVBAHA.114.304295 (2014).
Yang, S. et al. Transcription factor myocyte enhancer factor 2D regulates interleukin-10 production in microglia to protect neuronal cells from inflammation-induced death. Journal of neuroinflammation 12, 33, https://doi.org/10.1186/s12974-015-0258-z (2015).
Dintilhac, A. & Bernues, J. HMGB1 interacts with many apparently unrelated proteins by recognizing short amino acid sequences. The Journal of biological chemistry 277, 7021–7028, https://doi.org/10.1074/jbc.M108417200 (2002).
Cronk, J. C. et al. Methyl-CpG Binding Protein 2 Regulates Microglia and Macrophage Gene Expression in Response to Inflammatory Stimuli. Immunity 42, 679–691, https://doi.org/10.1016/j.immuni.2015.03.013 (2015).
Kim, I. D. & Lee, J. K. HMGB1-Binding Heptamer Confers Anti-Inflammatory Effects in Primary MicrogliaCulture. Experimental neurobiology 22, 301–307, https://doi.org/10.5607/en.2013.22.4.301 (2013).
Cortes, M. et al. RNAseq profiling of primary microglia and astrocyte cultures in near-term ovine fetus: A glial in vivo-in vitro multi-hit paradigm in large mammalian brain. Journal of neuroscience methods 276, 23–32, https://doi.org/10.1016/j.jneumeth.2016.11.008 (2017).
Healy, L. M. et al. MerTK Is a Functional Regulator of Myelin Phagocytosis by Human Myeloid Cells. J Immunol 196, 3375–3384, https://doi.org/10.4049/jimmunol.1502562 (2016).
Holtman, I. R. et al. Induction of a common microglia gene expression signature by aging and neurodegenerative conditions: a co-expression meta-analysis. Acta neuropathologica communications 3, 31, https://doi.org/10.1186/s40478-015-0203-5 (2015).
Acknowledgements
The authors thank Dr. Joan Serratosa and Dr. Eva Serrano for their revision and helpful comments on this work. The authors would also thank the staff of the Animal Facilities of the School of Medicine (University of Barcelona) for the professional care of the mouse colonies and Miguel Morales, Maria Iñíguez and Alberto de Luis (Genomic Plataform, CIBIR) for sequencing of RNA samples. MPS was a recipient of a FPU grant AP2010-5428 from the Spanish Ministerio de Educacion, Cultura y Deporte. This study was supported by grants PI07/455, PI10/378, PI12/709 and PI14/302 from the Instituto de Salud Carlos III, Spain, cofinanced with FEDER funds, and a grant from La Marató-TV3.
Author information
Authors and Affiliations
Contributions
M.P.S. performed most of the experimental work and wrote the manuscript. J.M.V.T. and A.G.G.D.B. did the RNA-seq analysis. C.S. participated in data analysis. J.S. conceived the project. M.P.S., J.M.V.T., C.S. and J.S. provided input and ideas throughout the process. A.G.G.D.B. wrote parts of the manuscript. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pulido-Salgado, M., Vidal-Taboada, J.M., Barriga, G.GD. et al. RNA-Seq transcriptomic profiling of primary murine microglia treated with LPS or LPS + IFNγ. Sci Rep 8, 16096 (2018). https://doi.org/10.1038/s41598-018-34412-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-018-34412-9
- Springer Nature Limited